miR-10a-5p and miR-29b-3p as Extracellular Vesicle-Associated Prostate Cancer Detection Markers
Abstract
1. Introduction
2. Results
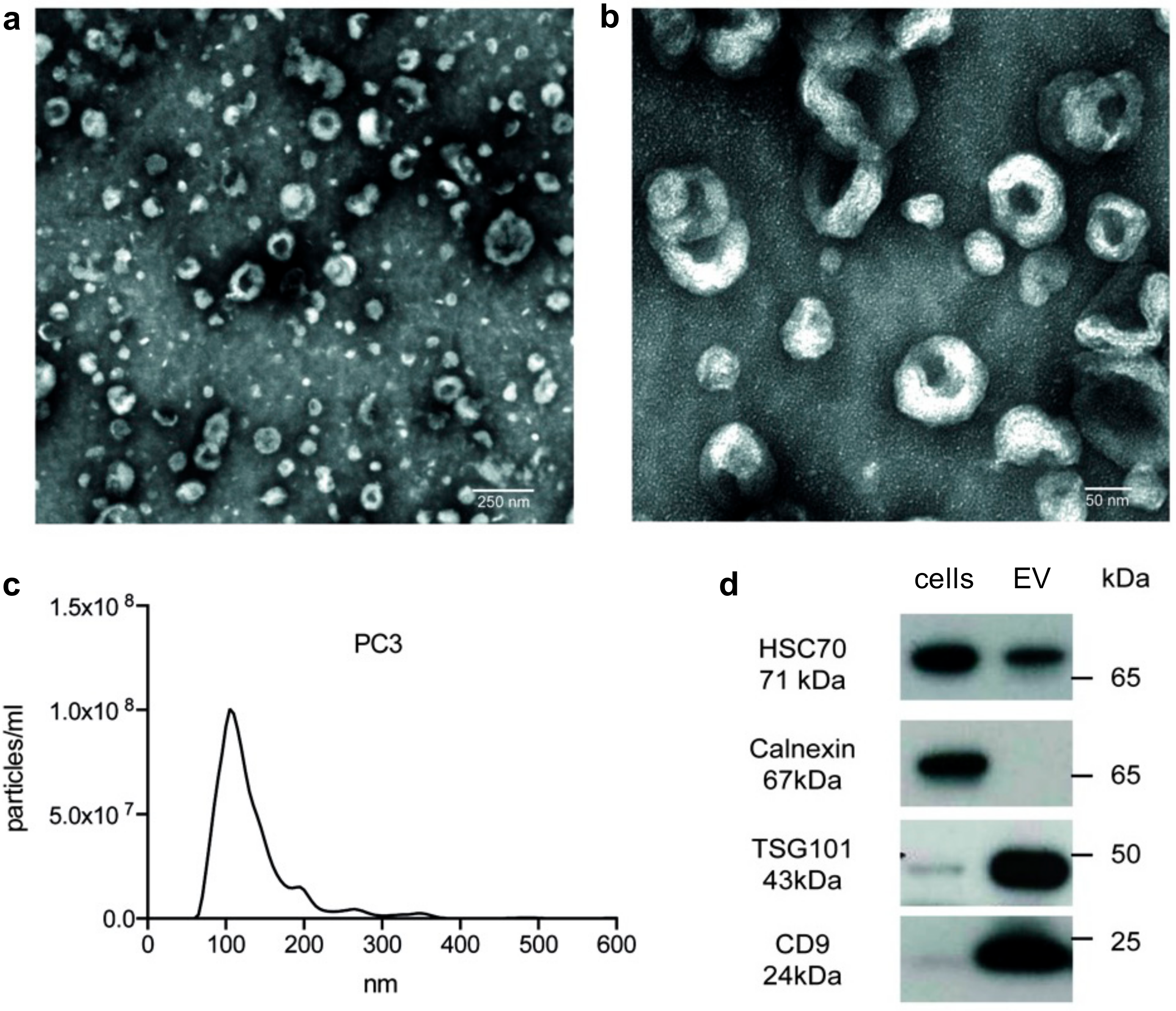
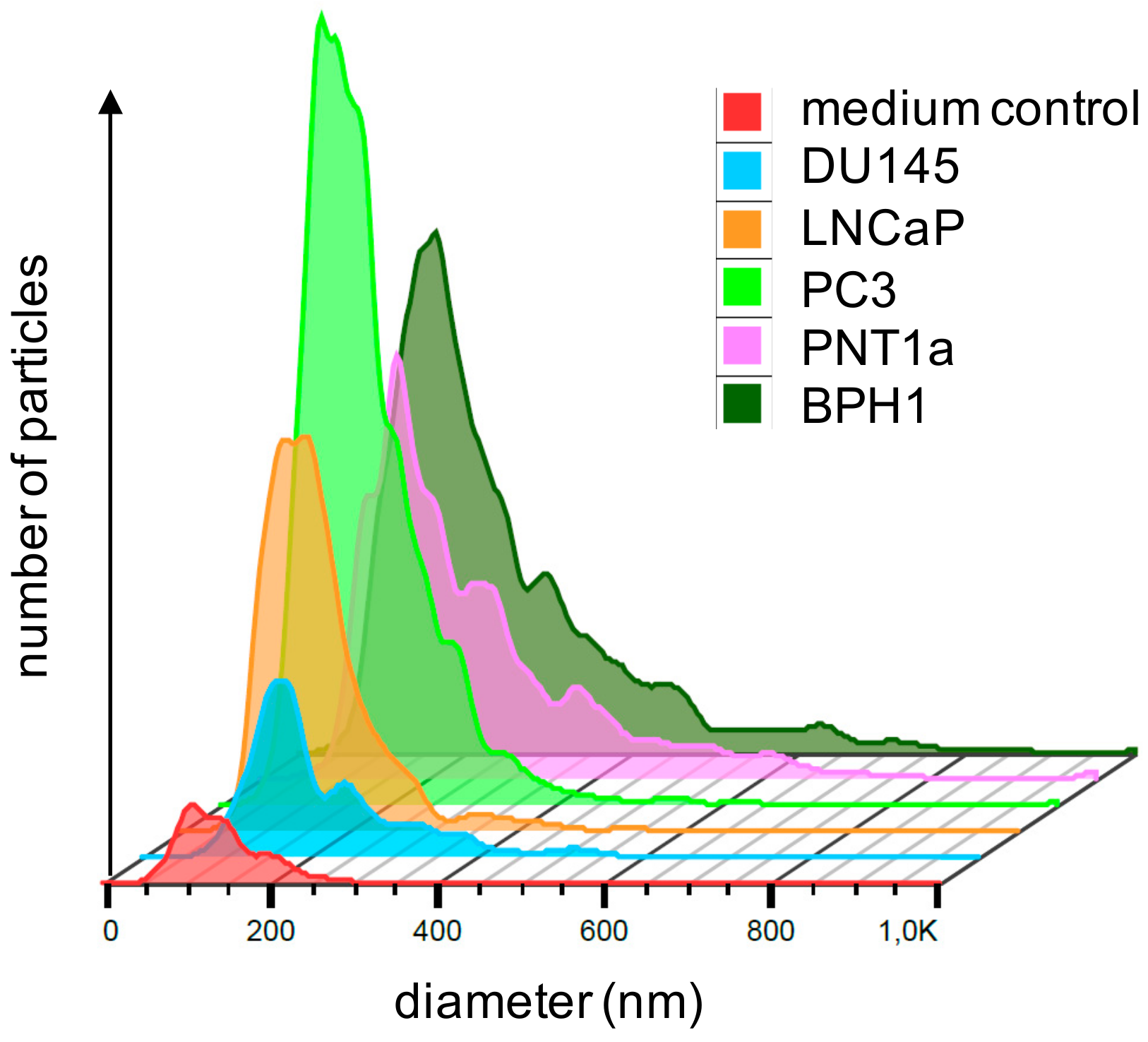
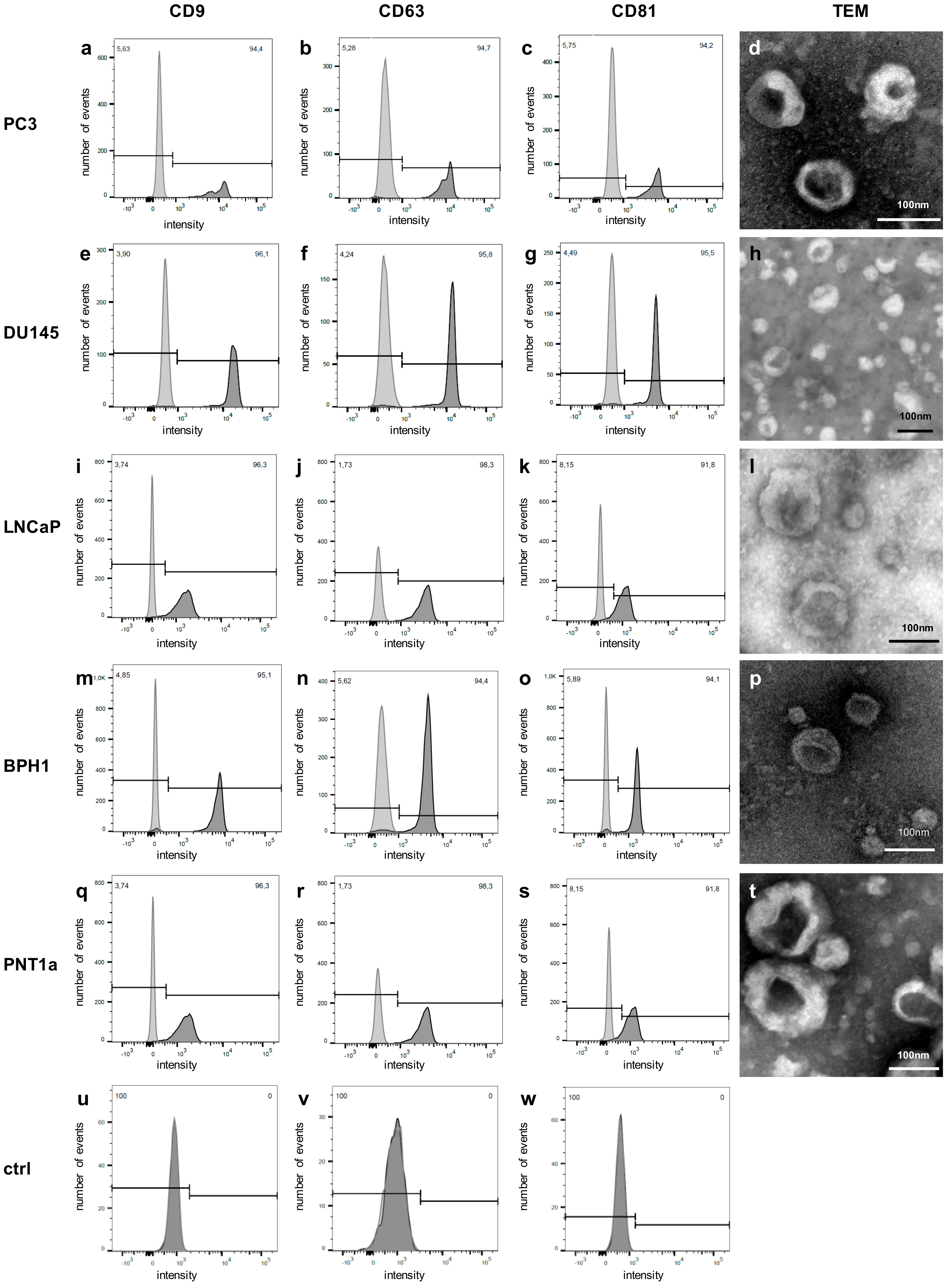
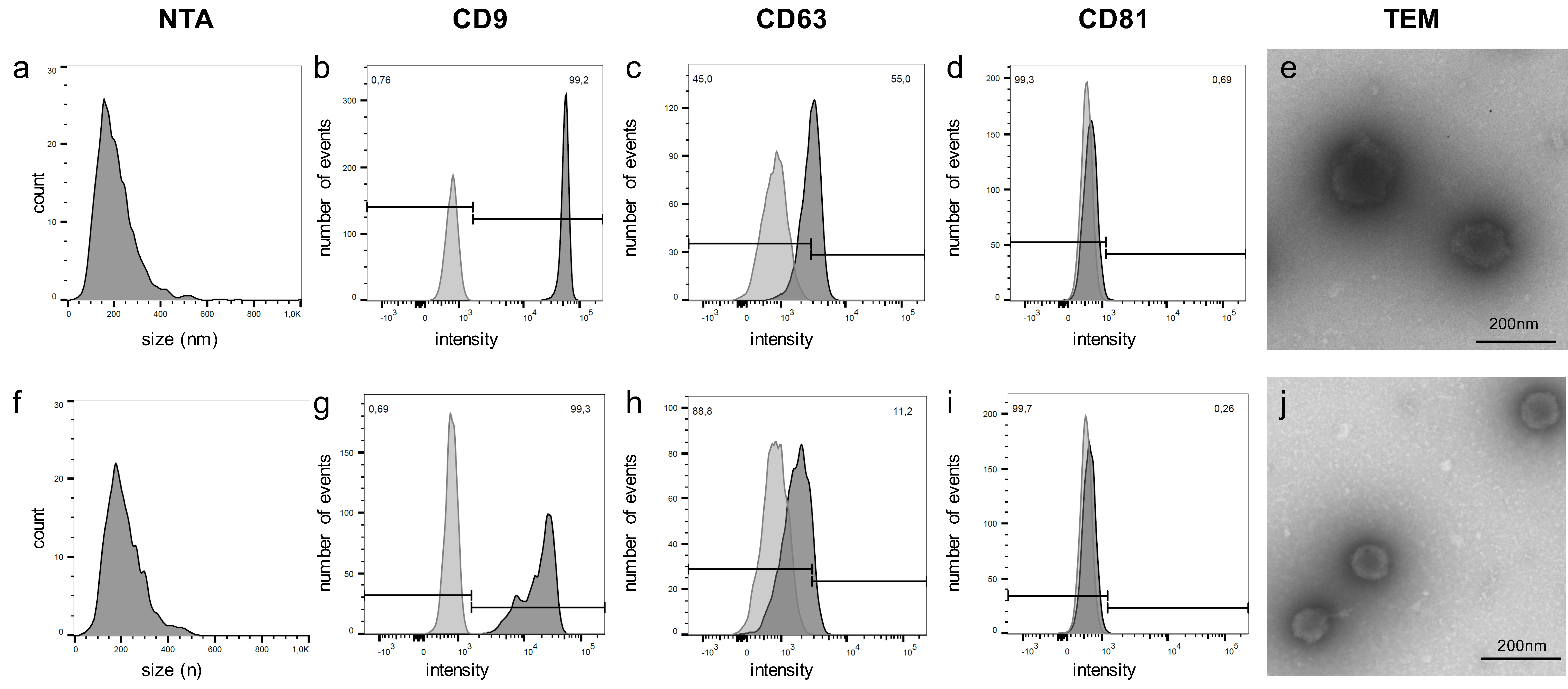
2.1. Isolated Particles Show Typical EV Features
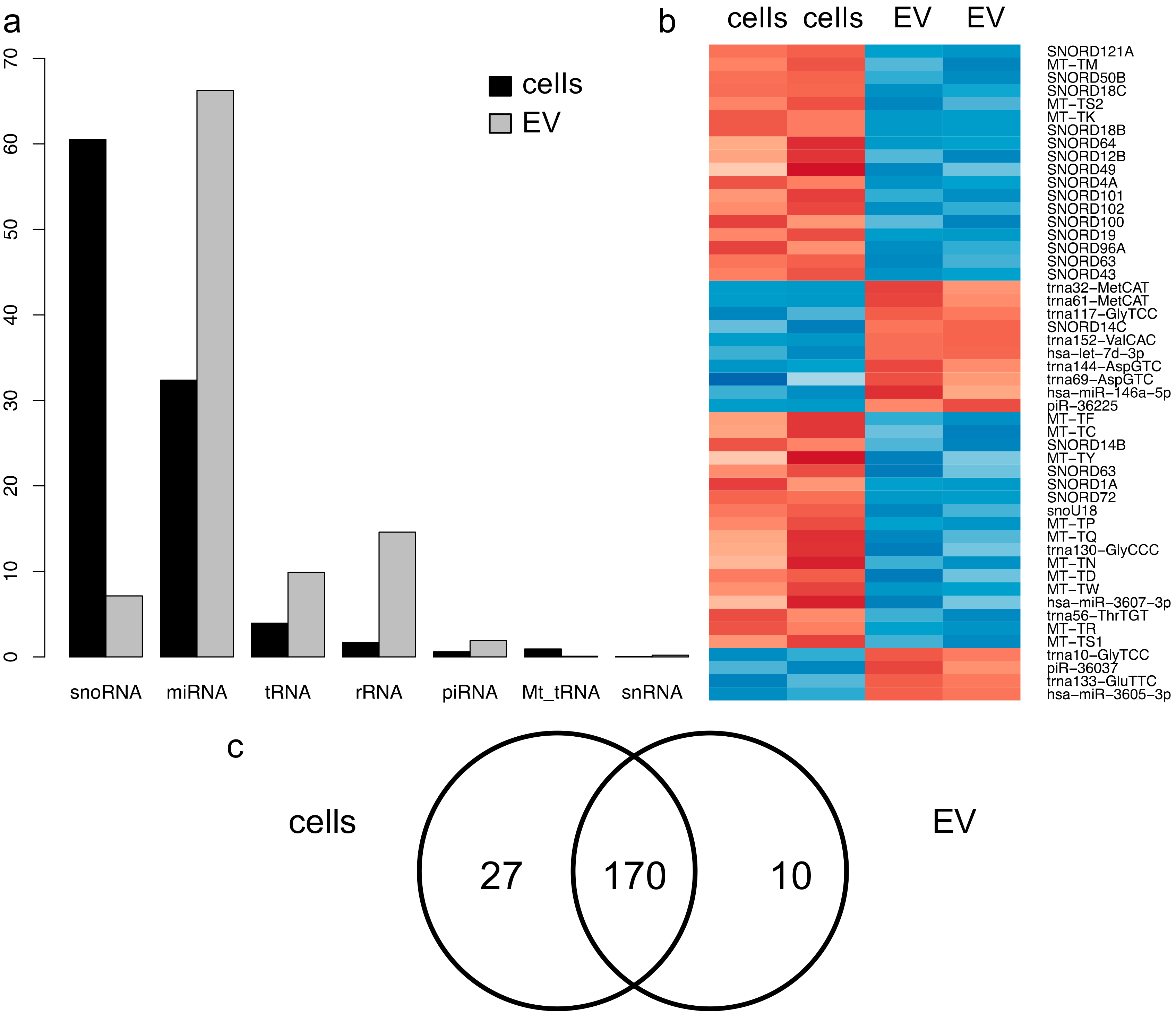
2.2. ncRNAs Are Differentially Enriched in PC3 EVs
2.3. Distinct ncRNA Distribution in PC3 Cells and EVs
2.4. PC3 Cells and EVs Have Distinct miRNA Profiles
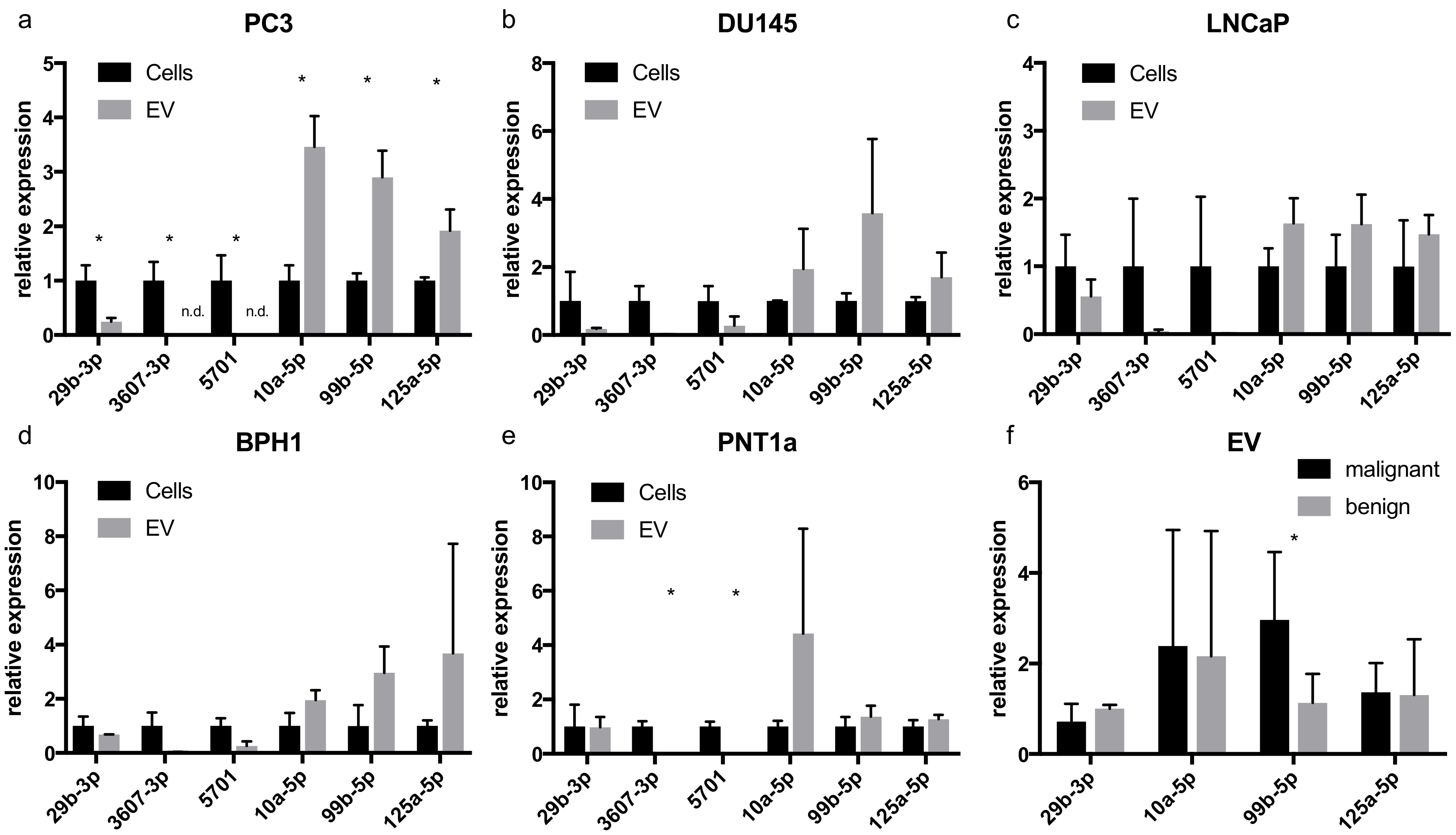
2.5. qRT-PCR Shows Similar miRNA Signatures in Other Prostate Cell Line EVs
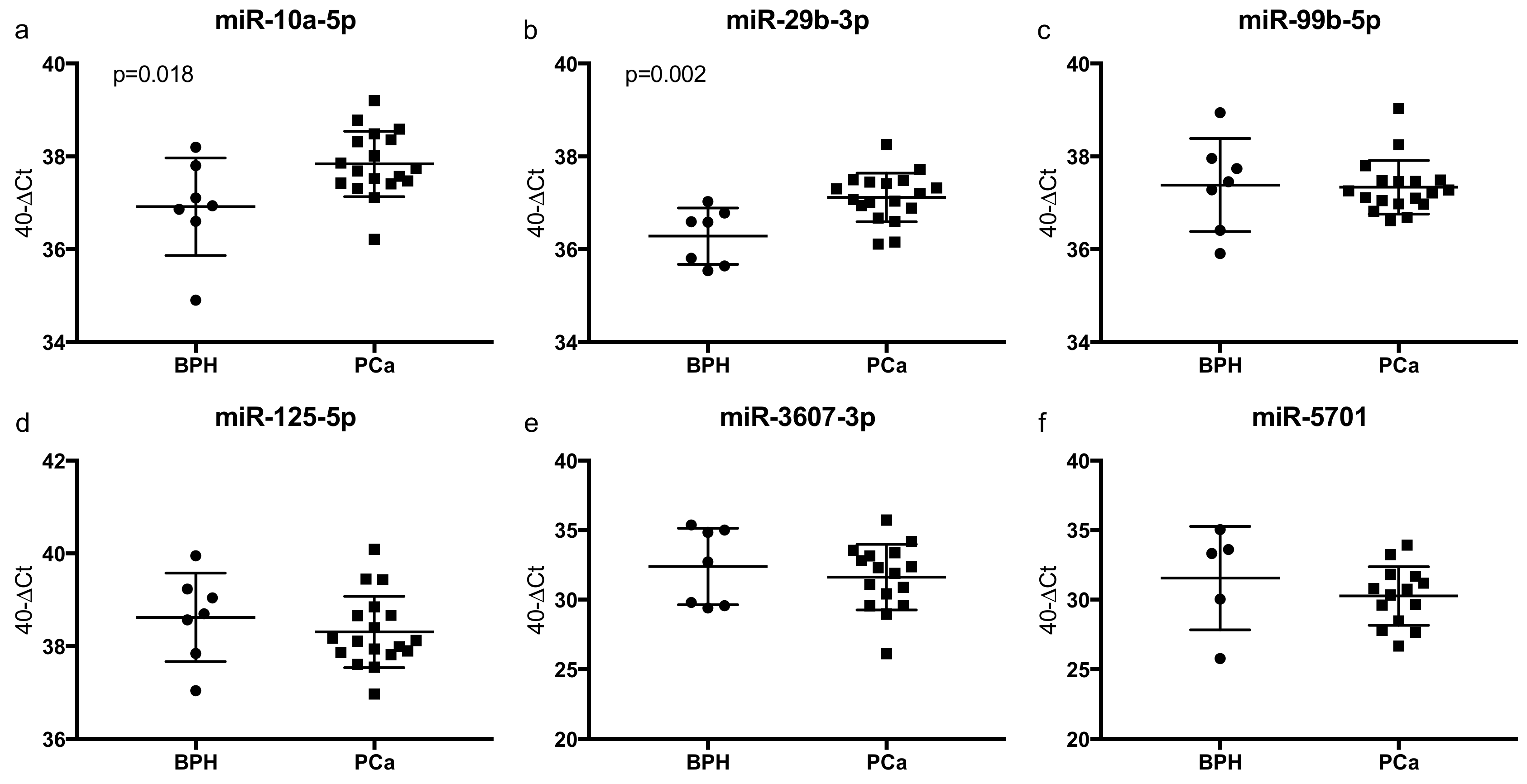
2.6. Validation Shows Overexpression of miR-10a-5p and miR-29b-3p in EVs Derived from PCa Patient Plasma
3. Discussion
4. Materials and Methods
4.1. Cell Culture and EV Isolation
4.2. Patient Samples
4.3. Sample Quality Control
4.4. RNA Extraction from EVs and NGS Library Preparation
4.5. Sequencing and Data Analysis
4.6. qRT-PCR Analyses of EVs from Cell Lines and Blood Plasma and of Tissue Samples
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the international society for extracellular vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, e1535750. [Google Scholar] [CrossRef] [PubMed]
- Kahlert, C.; Melo, S.A.; Protopopov, A.; Tang, J.; Seth, S.; Koch, M.; Zhang, J.; Weitz, J.; Chin, L.; Futreal, A.; et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J. Biol. Chem. 2014, 289, 3869–3875. [Google Scholar] [CrossRef] [PubMed]
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim. Biophys. Acta 2012, 1820, 940–948. [Google Scholar] [CrossRef] [PubMed]
- Rupp, A.-K.; Rupp, C.; Keller, S.; Brase, J.C.; Ehehalt, R.; Fogel, M.; Moldenhauer, G.; Marmé, F.; Sültmann, H.; Altevogt, P. Loss of EpCAM expression in breast cancer derived serum exosomes: Role of proteolytic cleavage. Gynecol. Oncol. 2011, 122, 437–446. [Google Scholar] [CrossRef]
- Skogberg, G.; Telemo, E.; Ekwall, O. Exosomes in the thymus: Antigen transfer and vesicles. Front. Immunol. 2015, 6, e366. [Google Scholar] [CrossRef]
- Gross, J.C.; Chaudhary, V.; Bartscherer, K.; Boutros, M. Active Wnt proteins are secreted on exosomes. Nat. Cell Biol. 2012, 14, 1036–1045. [Google Scholar] [CrossRef]
- Mulcahy, L.A.; Pink, R.C.; Carter, D.R.F. Routes and mechanisms of extracellular vesicle uptake. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef]
- Skog, J.; Würdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef]
- Braicu, C.; Tomuleasa, C.; Monroig, P.; Cucuianu, A.; Berindan-Neagoe, I.; Calin, G.A. Exosomes as divine messengers: Are they the hermes of modern molecular oncology? Cell Death Differ. 2015, 22, 34–45. [Google Scholar] [CrossRef]
- Wahlgren, J.; Karlson, T.D.L.; Glader, P.; Telemo, E.; Valadi, H. Activated human T cells secrete exosomes that participate in IL-2 mediated immune response signaling. PLoS ONE 2012, 7, e49723. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.P.; Spary, L.K.; Sanders, A.J.; Chowdhury, R.; Jiang, W.G.; Steadman, R.; Wymant, J.; Jones, A.T.; Kynaston, H.; Mason, M.D.; et al. Differentiation of tumour-promoting stromal myofibroblasts by cancer exosomes. Oncogene 2015, 34, 290–302. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, R.; Webber, J.P.; Gurney, M.; Mason, M.D.; Tabi, Z.; Clayton, A. Cancer exosomes trigger mesenchymal stem cell differentiation into pro-angiogenic and pro-invasive myofibroblasts. Oncotarget 2015, 6, 715–731. [Google Scholar] [CrossRef]
- Peinado, H.; Alecković, M.; Lavotshkin, S.; Matei, I.; Costa-Silva, B.; Moreno-Bueno, G.; Hergueta-Redondo, M.; Williams, C.; García-Santos, G.; Ghajar, C.; et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 2012, 18, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Havrylov, S.; Park, M. MS/MS-based strategies for proteomic profiling of invasive cell structures. Proteomics 2015, 15, 272–286. [Google Scholar] [CrossRef] [PubMed]
- Cosme, J.; Liu, P.P.; Gramolini, A.O. The cardiovascular exosome: Current perspectives and potential. Proteomics 2013, 13, 1654–1659. [Google Scholar] [CrossRef]
- Melo, S.A.; Luecke, L.B.; Kahlert, C.; Fernandez, A.F.; Gammon, S.T.; Kaye, J.; LeBleu, V.S.; Mittendorf, E.A.; Weitz, J.; Rahbari, N.; et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015, 523, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, K.G.; Ronquist, G.; Larsson, A.; Carlsson, L. Proteomic analysis of prostate cancer metastasis-derived prostasomes. Anticancer Res. 2010, 30, 285–290. [Google Scholar] [PubMed]
- Duijvesz, D.; Burnum-Johnson, K.E.; Gritsenko, M.A.; Hoogland, A.M.; Vredenbregt-van den Berg, M.S.; Willemsen, R.; Luider, T.; Paša-Tolić, L.; Jenster, G. Proteomic profiling of exosomes leads to the identification of novel biomarkers for prostate cancer. PLoS ONE 2013, 8, e82589. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Skotland, T.; Berge, V.; Sandvig, K.; Llorente, A. Exosomal proteins as prostate cancer biomarkers in urine: From mass spectrometry discovery to immunoassay-based validation. Eur. J. Pharm. Sci. Off. J. Eur. Fed. Pharm. Sci. 2017, 98, 80–85. [Google Scholar] [CrossRef]
- Bilen, M.A.; Pan, T.; Lee, Y.-C.; Lin, S.-C.; Yu, G.; Pan, J.; Hawke, D.; Pan, B.-F.; Vykoukal, J.; Gray, K.; et al. Proteomics profiling of exosomes from primary Mouse osteoblasts under proliferation versus mineralization conditions and characterization of their uptake into prostate cancer cells. J. Proteome Res. 2017, 16, 2709–2728. [Google Scholar] [CrossRef] [PubMed]
- Worst, T.S.; von Hardenberg, J.; Gross, J.C.; Erben, P.; Schnoelzer, M.; Hausser, I.; Bugert, P.; Michel, M.S.; Boutros, M. A database-augmented, exosome-based mass spectrometry approach exemplarily identifies circulating claudin 3 as biomarker in prostate cancer. Mol. Cell. Proteom. MCP 2017. [Google Scholar] [CrossRef] [PubMed]
- Fish, R.N.; Bostick, M.; Lehman, A.; Farmer, A. Transcriptome analysis at the single-cell level using SMART technology. Curr. Protoc. Mol. Biol. 2016, 116, 4.26.1–4.26.24. [Google Scholar] [CrossRef] [PubMed]
- Matullo, G.; Naccarati, A.; Pardini, B. MicroRNA expression profiling in bladder cancer: The challenge of next-generation sequencing in tissues and biofluids. Int. J. Cancer 2016, 138, 2334–2345. [Google Scholar] [CrossRef]
- Miranda, K.C.; Bond, D.T.; Levin, J.Z.; Adiconis, X.; Sivachenko, A.; Russ, C.; Brown, D.; Nusbaum, C.; Russo, L.M. Massively parallel sequencing of human urinary exosome/microvesicle RNA reveals a predominance of non-coding RNA. PLoS ONE 2014, 9, e96094. [Google Scholar] [CrossRef]
- Rodríguez, M.; Bajo-Santos, C.; Hessvik, N.P.; Lorenz, S.; Fromm, B.; Berge, V.; Sandvig, K.; Linē, A.; Llorente, A. Identification of non-invasive miRNAs biomarkers for prostate cancer by deep sequencing analysis of urinary exosomes. Mol. Cancer 2017, 16, e156. [Google Scholar] [CrossRef]
- Huang, X.; Yuan, T.; Liang, M.; Du, M.; Xia, S.; Dittmar, R.; Wang, D.; See, W.; Costello, B.A.; Quevedo, F.; et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant Prostate Cancer. Eur. Urol. 2014, 67, 33–41. [Google Scholar] [CrossRef]
- Sánchez, C.A.; Andahur, E.I.; Valenzuela, R.; Castellón, E.A.; Fullá, J.A.; Ramos, C.G.; Triviño, J.C. Exosomes from bulk and stem cells from human prostate cancer have a differential microRNA content that contributes cooperatively over local and pre-metastatic niche. Oncotarget 2016, 7, 3993–4008. [Google Scholar] [CrossRef]
- Gardiner, C.; Ferreira, Y.J.; Dragovic, R.A.; Redman, C.W.G.; Sargent, I.L. Extracellular vesicle sizing and enumeration by nanoparticle tracking analysis. J. Extracell. Vesicles 2013, 2. [Google Scholar] [CrossRef]
- Gross, J.C.; Boutros, M. Secretion and extracellular space travel of Wnt proteins. Curr. Opin. Genet. Dev. 2013, 23, 385–390. [Google Scholar] [CrossRef]
- Tehler, D.; Høyland-Kroghsbo, N.M.; Lund, A.H. The miR-10 microRNA precursor family. RNA Biol. 2011, 8, 728–734. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Lee, Y.S.; Malhotra, A.; Kim, H.K.; Matecic, M.; Evans, C.; Jensen, R.V.; Moskaluk, C.A.; Dutta, A. miR-99 family of MicroRNAs suppresses the expression of prostate-specific antigen and prostate cancer cell proliferation. Cancer Res. 2011, 71, 1313–1324. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Chen, Z.; Tan, X.; Zhou, F.; Tan, F.; Gao, Y.; Sun, N.; Xu, X.; Shao, K.; He, J. MicroRNA-99a/100 promotes apoptosis by targeting mTOR in human esophageal squamous cell carcinoma. Med. Oncol. Northwood Lond. Engl. 2013, 30, e411. [Google Scholar] [CrossRef] [PubMed]
- Hassel, D.; Cheng, P.; White, M.P.; Ivey, K.N.; Kroll, J.; Augustin, H.G.; Katus, H.A.; Stainier, D.Y.R.; Srivastava, D. MicroRNA-10 regulates the angiogenic behavior of zebrafish and human endothelial cells by promoting vascular endothelial growth factor signaling. Circ. Res. 2012, 111, 1421–1433. [Google Scholar] [CrossRef] [PubMed]
- Jönsson, M.E.; Nelander Wahlestedt, J.; Åkerblom, M.; Kirkeby, A.; Malmevik, J.; Brattaas, P.L.; Jakobsson, J.; Parmar, M. Comprehensive analysis of microRNA expression in regionalized human neural progenitor cells reveals microRNA-10 as a caudalizing factor. Dev. Camb. Engl. 2015, 142, 3166–3177. [Google Scholar] [CrossRef] [PubMed]
- Heinzelmann, J.; Henning, B.; Sanjmyatav, J.; Posorski, N.; Steiner, T.; Wunderlich, H.; Gajda, M.R.; Junker, K. Specific miRNA signatures are associated with metastasis and poor prognosis in clear cell renal cell carcinoma. World J. Urol. 2011, 29, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Jin, Y.; Yu, D.; Wang, A.; Mahjabeen, I.; Wang, C.; Liu, X.; Zhou, X. Down-regulation of the microRNA-99 family members in head and neck squamous cell carcinoma. Oral Oncol. 2012, 48, 686–691. [Google Scholar] [CrossRef]
- Wang, Z.; Lv, J.; Zou, X.; Huang, Z.; Zhang, H.; Liu, Q.; Jiang, L.; Zhou, X.; Zhu, W. A three plasma microRNA signature for papillary thyroid carcinoma diagnosis in Chinese patients. Gene 2019, 693, 37–45. [Google Scholar] [CrossRef]
- Guo, X.; Qiu, W.; Liu, Q.; Qian, M.; Wang, S.; Zhang, Z.; Gao, X.; Chen, Z.; Xue, H.; Li, G. Immunosuppressive effects of hypoxia-induced glioma exosomes through myeloid-derived suppressor cells via the miR-10a/Rora and miR-21/Pten Pathways. Oncogene 2018, 37, 4239–4259. [Google Scholar] [CrossRef]
- Muntión, S.; Ramos, T.L.; Diez-Campelo, M.; Rosón, B.; Sánchez-Abarca, L.I.; Misiewicz-Krzeminska, I.; Preciado, S.; Sarasquete, M.-E.; de Las Rivas, J.; González, M.; et al. Microvesicles from mesenchymal stromal cells are involved in HPC-microenvironment crosstalk in myelodysplastic patients. PLoS ONE 2016, 11, e0146722. [Google Scholar] [CrossRef]
- Ke, K.; Lou, T. MicroRNA-10a suppresses breast cancer progression via PI3K/Akt/mTOR pathway. Oncol. Lett. 2017, 14, 5994–6000. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhou, Y.; Huan, L.; Xu, L.; Shen, M.; Huang, S.; Liang, L. LncRNA MIR22HG inhibits growth, migration and invasion through regulating the miR-10a-5p/NCOR2 axis in hepatocellular carcinoma cells. Cancer Sci. 2019, 110, 973–984. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ren, L.; Yu, M.; Liu, X.; Ma, W.; Huang, L.; Li, X.; Ye, X. S-equol inhibits proliferation and promotes apoptosis of human breast cancer MCF-7 cells via regulating miR-10a-5p and PI3K/AKT pathway. Arch. Biochem. Biophys. 2019, 672, e108064. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, A.S.; Afshari, J.T.; Esmaeili, S.-A.; Saburi, E.; Joneidi, Z.; Momtazi-Borojeni, A.A. Cardioprotective microRNAs: Lessons from stem cell-derived exosomal microRNAs to treat cardiovascular disease. Atherosclerosis 2019, 285, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, B.; Zhang, A.; Hassounah, F.; Seow, Y.; Wood, M.; Ma, F.; Klein, J.D.; Price, S.R.; Wang, X.H. Exosome-mediated miR-29 transfer reduces muscle atrophy and kidney fibrosis in mice. Mol. Ther. J. Am. Soc. Gene Ther. 2019, 27, 571–583. [Google Scholar] [CrossRef]
- Yeh, Y.-Y.; Ozer, H.G.; Lehman, A.M.; Maddocks, K.; Yu, L.; Johnson, A.J.; Byrd, J.C. Characterization of CLL exosomes reveals a distinct microRNA signature and enhanced secretion by activation of BCR signaling. Blood 2015, 125, 3297–3305. [Google Scholar] [CrossRef]
- Nishikawa, R.; Goto, Y.; Kojima, S.; Enokida, H.; Chiyomaru, T.; Kinoshita, T.; Sakamoto, S.; Fuse, M.; Nakagawa, M.; Naya, Y.; et al. Tumor-suppressive microRNA-29s inhibit cancer cell migration and invasion via targeting LAMC1 in prostate cancer. Int. J. Oncol. 2014, 45, 401–410. [Google Scholar] [CrossRef]
- Takayama, K.; Misawa, A.; Suzuki, T.; Takagi, K.; Hayashizaki, Y.; Fujimura, T.; Homma, Y.; Takahashi, S.; Urano, T.; Inoue, S. TET2 repression by androgen hormone regulates global hydroxymethylation status and prostate cancer progression. Nat. Commun. 2015, 6, e8219. [Google Scholar] [CrossRef]
- Qi, Y.; Huang, Y.; Pang, L.; Gu, W.; Wang, N.; Hu, J.; Cui, X.; Zhang, J.; Zhao, J.; Liu, C.; et al. Prognostic value of the MicroRNA-29 family in multiple human cancers: A meta-analysis and systematic review. Clin. Exp. Pharmacol. Physiol. 2017, 44, 441–454. [Google Scholar] [CrossRef]
- Siriwardhana, C.; Khadka, V.S.; Chen, J.J.; Deng, Y. Development of a miRNA-seq based prognostic signature in lung adenocarcinoma. BMC Cancer 2019, 19, e34. [Google Scholar] [CrossRef]
- Teng, Y.; Ren, Y.; Hu, X.; Mu, J.; Samykutty, A.; Zhuang, X.; Deng, Z.; Kumar, A.; Zhang, L.; Merchant, M.L.; et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 2017, 8, e14448. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 2006, 30, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Veneziano, D.; Di Bella, S.; Nigita, G.; Laganà, A.; Ferro, A.; Croce, C.M. Non-coding RNA: Current Deep Sequencing Data Analysis Approaches and Challenges. Hum. Mutat. 2016, 37, 1283–1298. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, e550. [Google Scholar] [CrossRef]
- Jenjaroenpun, P.; Kremenska, Y.; Nair, V.M.; Kremenskoy, M.; Joseph, B.; Kurochkin, I.V. Characterization of RNA in exosomes secreted by human breast cancer cell lines using next-generation sequencing. PeerJ 2013, 1. [Google Scholar] [CrossRef]
- Huang, X.; Yuan, T.; Tschannen, M.; Sun, Z.; Jacob, H.; Du, M.; Liang, M.; Dittmar, R.L.; Liu, Y.; Liang, M.; et al. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genom. 2013, 14, e319. [Google Scholar] [CrossRef]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-‘t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2013, 2. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- FASTX-Toolkit. Available online: http://hannonlab.cshl.edu/fastx_toolkit/index.html (accessed on 25 February 2016).
- Homer Software and Data Download. Available online: http://homer.salk.edu/homer/ngs/homerTools.html (accessed on 25 February 2016).
- Rosenkranz, D. piRNA cluster database: A web resource for piRNA producing loci. Nucleic Acids Res. 2016, 44, 223–230. [Google Scholar] [CrossRef]
- Flicek, P.; Amode, M.R.; Barrell, D.; Beal, K.; Billis, K.; Brent, S.; Carvalho-Silva, D.; Clapham, P.; Coates, G.; Fitzgerald, S.; et al. Ensembl 2014. Nucleic Acids Res. 2014, 42, 749–755. [Google Scholar] [CrossRef]
- Nawrocki, E.P.; Burge, S.W.; Bateman, A.; Daub, J.; Eberhardt, R.Y.; Eddy, S.R.; Floden, E.W.; Gardner, P.P.; Jones, T.A.; Tate, J.; et al. Rfam 12.0: Updates to the RNA families database. Nucleic Acids Res. 2015, 43, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Bioconductor-Home. Available online: https://bioconductor.org/ (accessed on 25 February 2016).
- Breyer, J.; Wirtz, R.M.; Otto, W.; Erben, P.; Worst, T.S.; Stoehr, R.; Eckstein, M.; Denzinger, S.; Burger, M.; Hartmann, A. High PDL1 mRNA expression predicts better survival of stage pT1 non-muscle-invasive bladder cancer (NMIBC) patients. Cancer Immunol. Immunother. 2017, 67, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Worst, T.S.; Weis, C.-A.; Stöhr, R.; Bertz, S.; Eckstein, M.; Otto, W.; Breyer, J.; Hartmann, A.; Bolenz, C.; Wirtz, R.M.; et al. CDKN2A as transcriptomic marker for muscle-invasive bladder cancer risk stratification and therapy decision-making. Sci. Rep. 2018, 8, e14383. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Nitschke, K.; Heinkele, J.; Weis, C.-A.; Worst, T.S.; Eckstein, M.; Porubsky, S.; Erben, P. ANLN and TLE2 in muscle invasive bladder cancer: A functional and clinical evaluation based on in silico and in vitro data. Cancers 2019, 11, 1840. [Google Scholar] [CrossRef] [PubMed]
| Sample | snoRNA (%) | miRNA (%) | tRNA (%) | rRNA (%) | piRNA (%) | Mt_tRNA (%) | snRNA (%) |
|---|---|---|---|---|---|---|---|
| cell 1 | 3,540,368 (56.40) | 2,374,035 (37.82) | 124,497 (1.98) | 148,002 (2.36) | 40,914 (0.65) | 48,378 (0.77) | 1511 (0.02) |
| cell 2 | 4,708,633 (64.01) | 2,040,209 (27.73) | 411,819 (5.60) | 78,915 (1.07) | 40,408 (0.55) | 73,719 (1.00) | 2722 (0.04) |
| average | 4,124,500 (60.50) | 2,207,122 (32.38) | 268,158 (3.93) | 113,458.5 (1.66) | 40,661 (0.60) | 61,048.5 (0.90) | 2116.5 (0.03) |
| EV 1 | 138,716 (6.26) | 1,514,573 (68.39) | 173,397 (7.83) | 350,202 (15.81) | 31,573 (1.43) | 1649 (0.07) | 4459 (0.20) |
| EV 2 | 132,583 (8.35) | 1,004,736 (63.27) | 202,431 (12.75) | 203,911 (12.84) | 40,494 (2.55) | 1279 (0.08) | 2467 (0.16) |
| average | 135,649.5 (7.13) | 1,259,654 (66.25) | 187,914 (9.88) | 277,056.5 (14.57) | 36,033.5 (1.9) | 1464 (0.08) | 3463 (0.18) |
| miRNA | log2 Fold Change | p-Value | FDR-Adjusted p-Value |
|---|---|---|---|
| hsa-miR-3605-3p | 6.06 | 1.53 × 10−32 | 3.95 × 10−31 |
| hsa-let-7d-3p | 5.09 | 4.32 × 10−39 | 1.34 × 10−37 |
| hsa-miR-146a-5p | 4.20 | 4.22 × 10−18 | 3.06 × 10−17 |
| hsa-miR-937-3p | 3.72 | 3.94 × 10−16 | 2.38 × 10−15 |
| hsa-let-7b-3p | 3.67 | 2.65 × 10−19 | 2.16 × 10−18 |
| hsa-miR-99b-5p | 3.67 | 5.12 × 10−27 | 7.93 × 10−26 |
| hsa-miR-92b-3p | 3.66 | 1.32 × 10−32 | 3.61 × 10−31 |
| hsa-miR-125a-5p | 3.59 | 2.35 × 10−19 | 1.95 × 10−18 |
| hsa-miR-873-3p | 3.50 | 1.14 × 10−17 | 7.91 × 10−17 |
| hsa-miR-320b | 3.49 | 2.35 × 10−24 | 3.12 × 10−23 |
| hsa-miR-625-3p | 3.48 | 1.83 × 10−22 | 2.13 × 10−21 |
| hsa-miR-320c | 3.29 | 3.23 × 10−19 | 2.54 × 10−18 |
| hsa-miR-1247-5p | 3.24 | 1.54 × 10−16 | 9.97 × 10−16 |
| hsa-miR-2110 | 3.15 | 6.22 × 10−11 | 2.39 × 10−10 |
| hsa-miR-105-5p | 3.09 | 8.05 × 10−16 | 4.80 × 10−15 |
| miRNA | log2 Fold Change | p-Value | FDR-Adjusted p-Value |
|---|---|---|---|
| hsa-miR-3607-3p | −5.07 | 5.59 × 10−11 | 2.19 × 10−10 |
| hsa-miR-5701 | −4.16 | 1.13 × 10−08 | 3.59 × 10−8 |
| hsa-miR-193a-3p | −3.60 | 8.39 × 10−07 | 2.23 × 10−6 |
| hsa-miR-1246 | −2.54 | 2.26 × 10−16 | 1.42 × 10−15 |
| hsa-miR-582-5p | −2.50 | 3.81 × 10−06 | 9.22 × 10−6 |
| hsa-miR-19b-3p | −2.49 | 7.81 × 10−11 | 2.93 × 10−10 |
| hsa-miR-561-5p | −2.36 | 1.74 × 10−06 | 4.37 × 10−6 |
| hsa-miR-26a-2-3p | −2.31 | 9.51 × 10−07 | 2.48 × 10−6 |
| hsa-miR-19a-3p | −2.20 | 2.00 × 10−07 | 5.65 × 10−7 |
| hsa-miR-29b-3p | −1.97 | 7.07 × 10−10 | 2.42 × 10−9 |
| hsa-miR-101-3p | −1.97 | 1.51 × 10−08 | 4.75 × 10−8 |
| hsa-miR-29c-3p | −1.92 | 3.80 × 10−06 | 9.22 × 10−6 |
| hsa-miR-32-5p | −1.73 | 8.96 × 10−06 | 2.09 × 10−5 |
| hsa-miR-17-3p | −1.69 | 8.17 × 10−05 | 1.73 × 10−4 |
| hsa-miR-590-3p | −1.65 | 2.16 × 10−05 | 4.25 × 10−4 |
| miRNA | log2 Fold Change | p-Value | FDR-Adjusted p-Value |
|---|---|---|---|
| hsa-miR-99b-5p | 3.67 | 5.12 × 10−27 | 7.93 × 10−26 |
| hsa-miR-125a-5p | 3.59 | 2.35 × 10−19 | 1.95 × 10−18 |
| hsa-miR-125b-5p | 2.48 | 2.75 × 10−09 | 9.01 × 10−09 |
| hsa-miR-125a-3p | 2.32 | 3.59 × 10−07 | 9.89 × 10−07 |
| hsa-miR-99a-5p | 2.03 | 4.08 × 10−09 | 1.32 × 10−08 |
| hsa-miR-10a-5p | 1.92 | 1.37 × 10−10 | 5.06 × 10−10 |
| hsa-miR-99b-3p | 1.87 | 7.41 × 10-06 | 1.76 × 10−05 |
| hsa-miR-10a-3p | 1.70 | 1.25 × 10−04 | 2.57 × 10−04 |
| hsa-miR-125b-1-3p | 1.28 | 1.53 × 10−04 | 3.09 × 10−04 |
| hsa-miR-10b-5p | 1.03 | 3.10 × 10−02 | 4.71 × 10−02 |
| Cell Line | Mean Particle Concentration ± SD | Mean Diameter of Particles ± SD |
|---|---|---|
| PC3 | 1.85 × 1012/mL ± 1.20 × 1012/mL | 179.5 nm ± 29.9 nm |
| DU145 | 4.15 × 1011/mL ± 4.20 × 1011/mL | 185.6 nm ± 7.7 nm |
| LNCaP | 2.54 × 1011/mL ± 1.14 × 1011/mL | 150.7 nm ± 21.0 nm |
| BPH1 | 1.67 × 1011/mL ± 1.08 × 1011/mL | 210.9 nm ± 10.4 nm |
| PNT1a | 1.47 × 1011/mL ± 1.23 × 1011/mL | 180.6 nm ± 31.8 nm |
| medium control | 1.04 × 109/mL ± 3.11 × 107/mL | 156.2 nm ± 26.4 nm |
| Parameter | PCa Patients (n = 18) | BPH Controls (n = 7) |
|---|---|---|
| Median age (years) | 65 (48–72) | 66 (59–80) |
| pT stage | ||
| 2c | 14 | |
| 3a | 2 | |
| 3b | 1 | |
| no surgery | 1 | |
| Gleason Score | ||
| 3 + 3 | 2 | |
| 3 + 4 | 7 | |
| 4 + 3 | 6 | |
| 4 + 4 | 1 * | |
| 4 + 5 | 1 | |
| 5 + 4 | 1 | |
| D’Amico risk group | ||
| low | 5 | |
| intermediate | 7 | |
| high | 6 | |
| gland volume (mL) | 33 (15–59) | 44 (20–120) |
| Median serum PSA level (ng/mL) | 7.2 (3.3–94.7) | 3.4 (0.7–10.0) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Worst, T.S.; Previti, C.; Nitschke, K.; Diessl, N.; Gross, J.C.; Hoffmann, L.; Frey, L.; Thomas, V.; Kahlert, C.; Bieback, K.; et al. miR-10a-5p and miR-29b-3p as Extracellular Vesicle-Associated Prostate Cancer Detection Markers. Cancers 2020, 12, 43. https://doi.org/10.3390/cancers12010043
Worst TS, Previti C, Nitschke K, Diessl N, Gross JC, Hoffmann L, Frey L, Thomas V, Kahlert C, Bieback K, et al. miR-10a-5p and miR-29b-3p as Extracellular Vesicle-Associated Prostate Cancer Detection Markers. Cancers. 2020; 12(1):43. https://doi.org/10.3390/cancers12010043
Chicago/Turabian StyleWorst, Thomas Stefan, Christopher Previti, Katja Nitschke, Nicolle Diessl, Julia Christina Gross, Lena Hoffmann, Lisa Frey, Vanessa Thomas, Christoph Kahlert, Karen Bieback, and et al. 2020. "miR-10a-5p and miR-29b-3p as Extracellular Vesicle-Associated Prostate Cancer Detection Markers" Cancers 12, no. 1: 43. https://doi.org/10.3390/cancers12010043
APA StyleWorst, T. S., Previti, C., Nitschke, K., Diessl, N., Gross, J. C., Hoffmann, L., Frey, L., Thomas, V., Kahlert, C., Bieback, K., Torres Crigna, A., Fricke, F., Porubsky, S., Westhoff, N., von Hardenberg, J., Nuhn, P., Erben, P., Michel, M. S., & Boutros, M. (2020). miR-10a-5p and miR-29b-3p as Extracellular Vesicle-Associated Prostate Cancer Detection Markers. Cancers, 12(1), 43. https://doi.org/10.3390/cancers12010043





