A Radiogenomic Approach for Decoding Molecular Mechanisms Underlying Tumor Progression in Prostate Cancer
Abstract
:1. Introduction
2. Results and Discussion
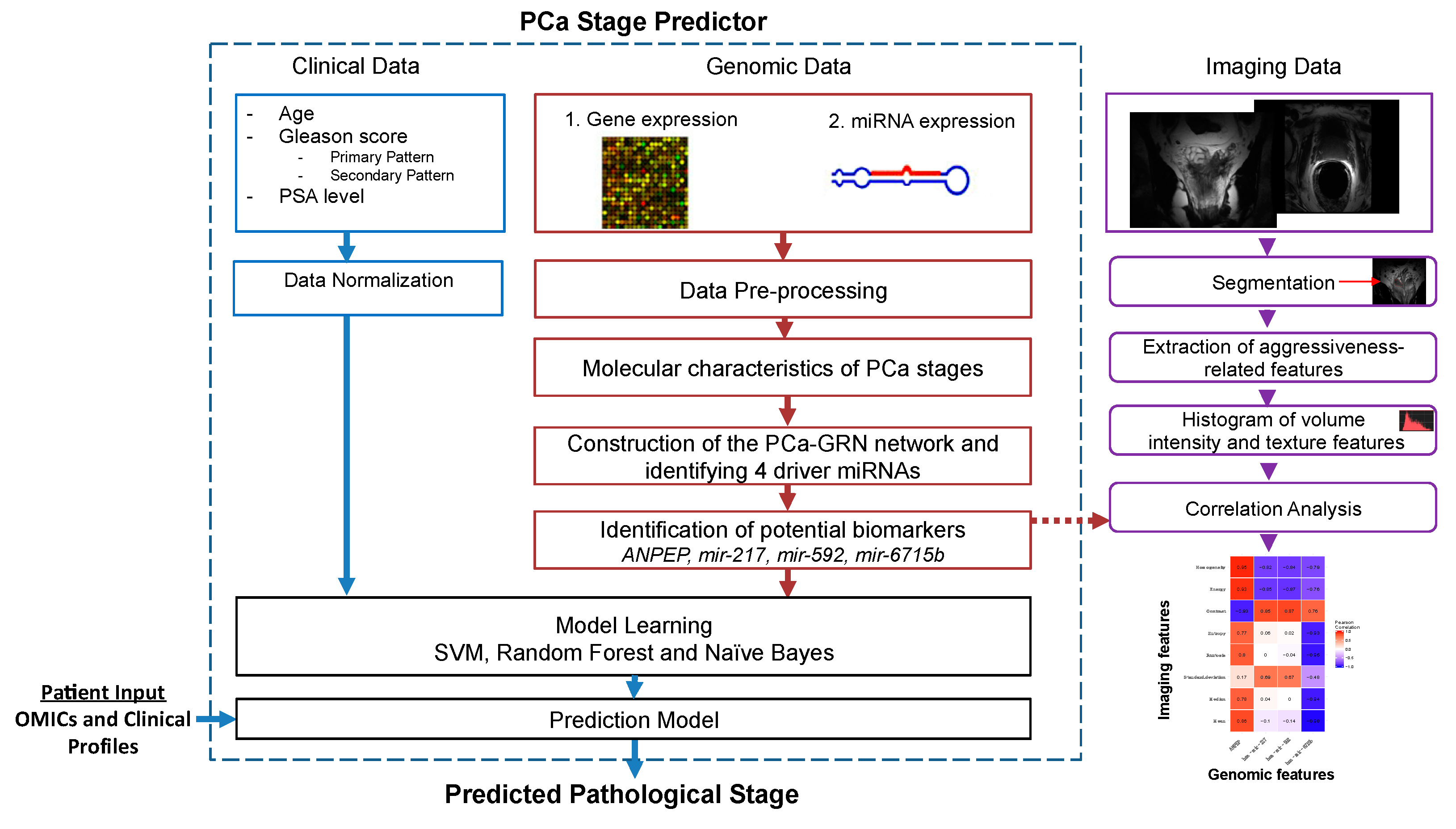
2.1. Description of the Radiogenomic Approach
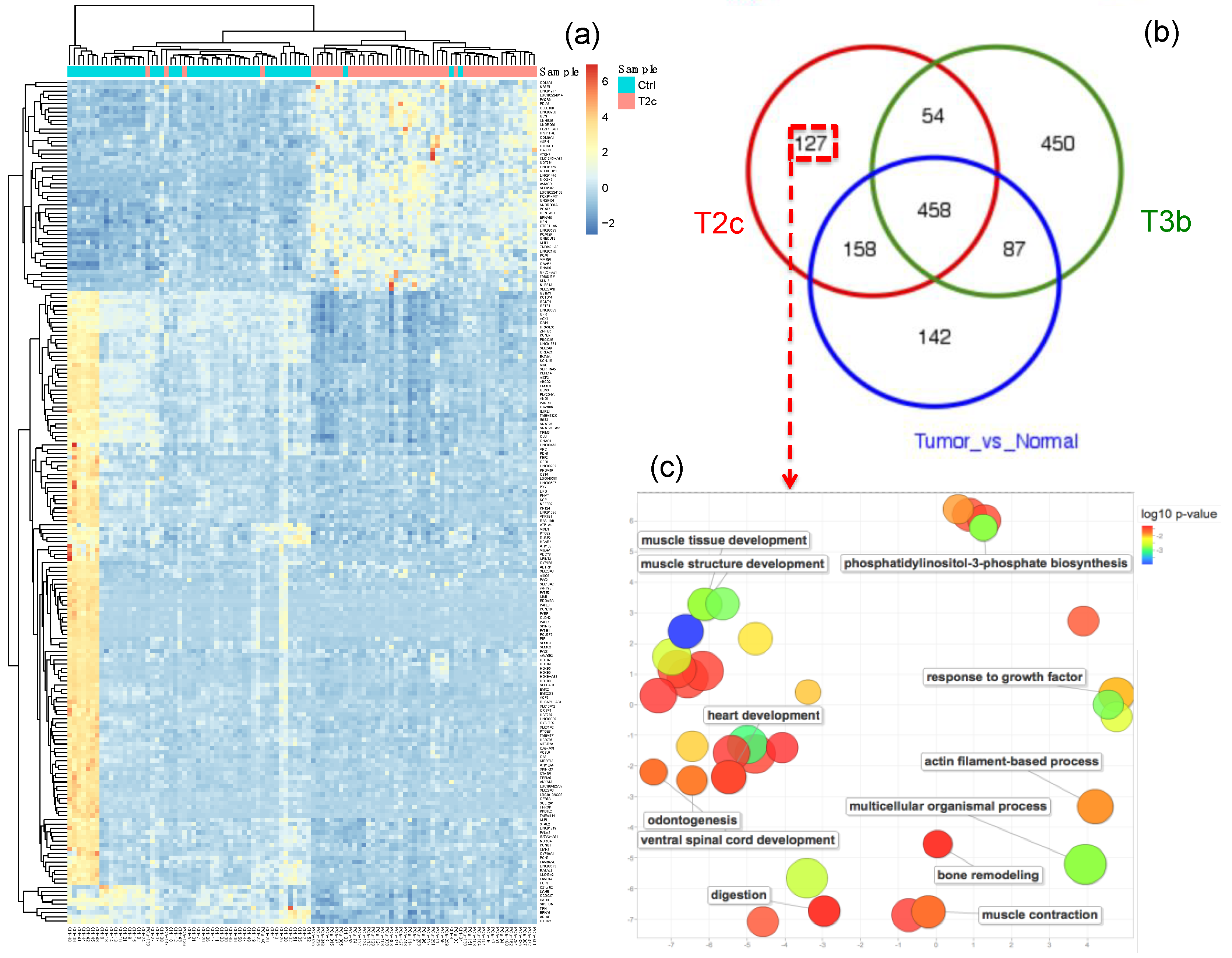
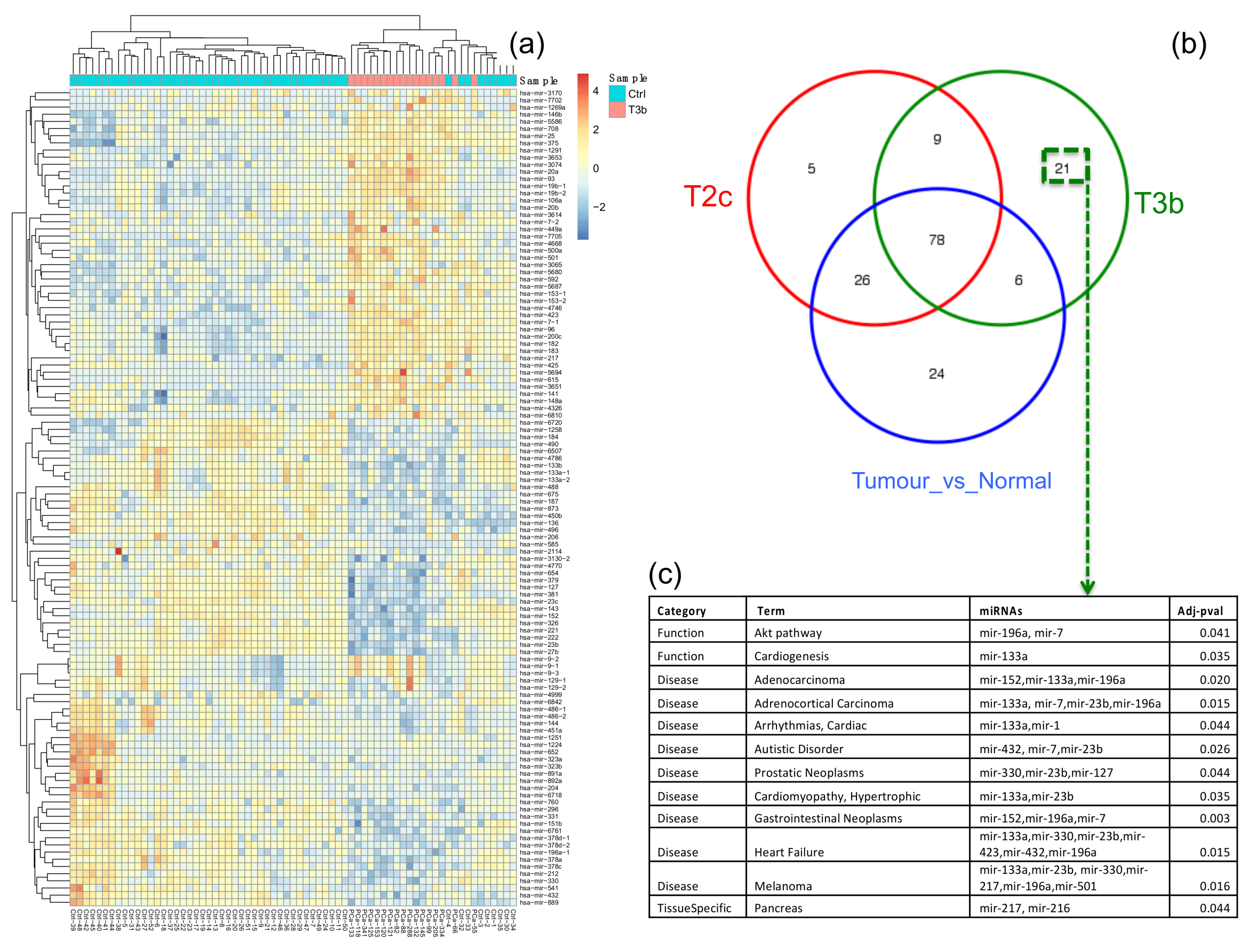
2.2. Functional Characteristics of T2c and T3b Stages
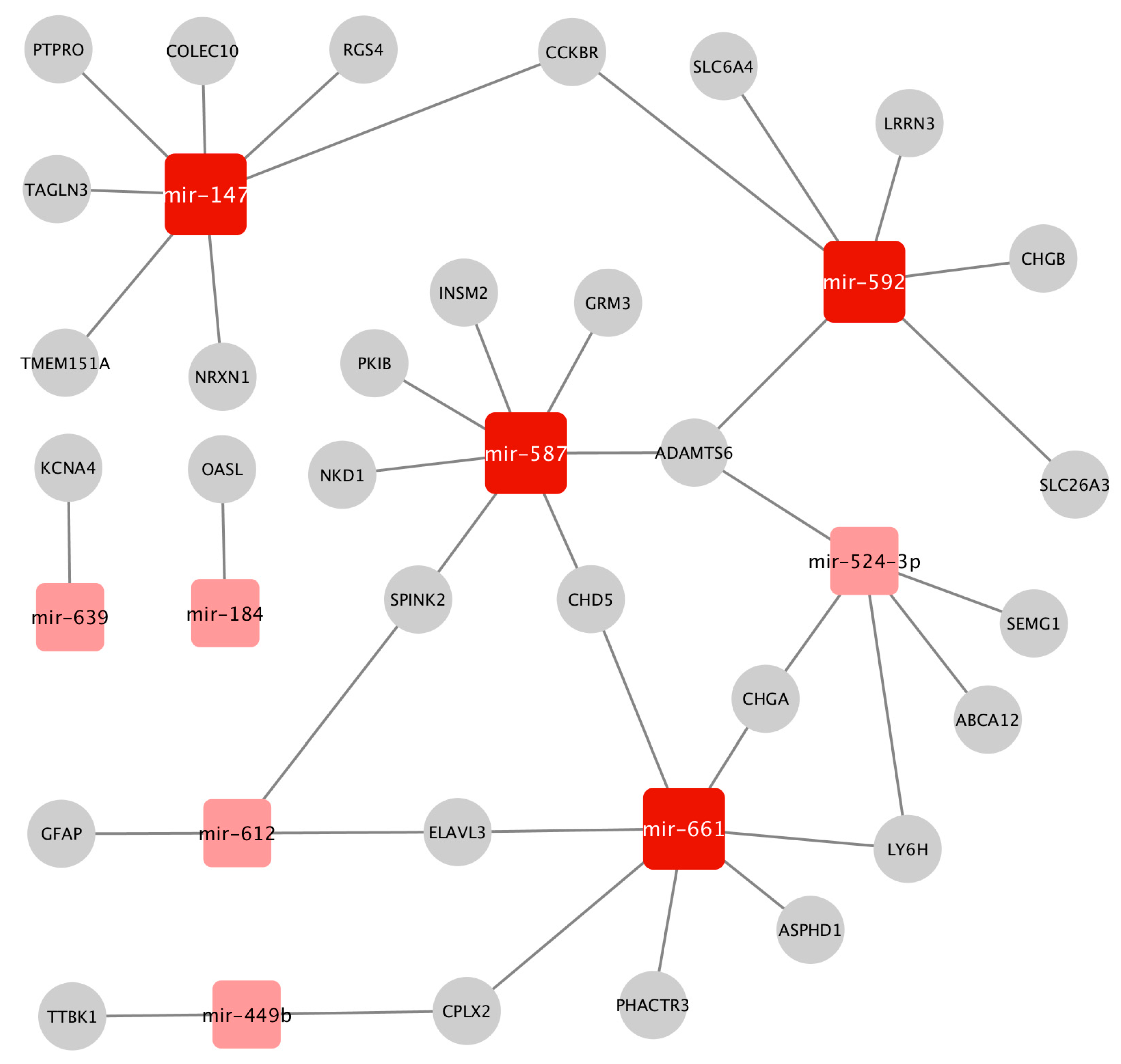
2.3. Construction of the Prostate Cancer (PCa-GRN) Network
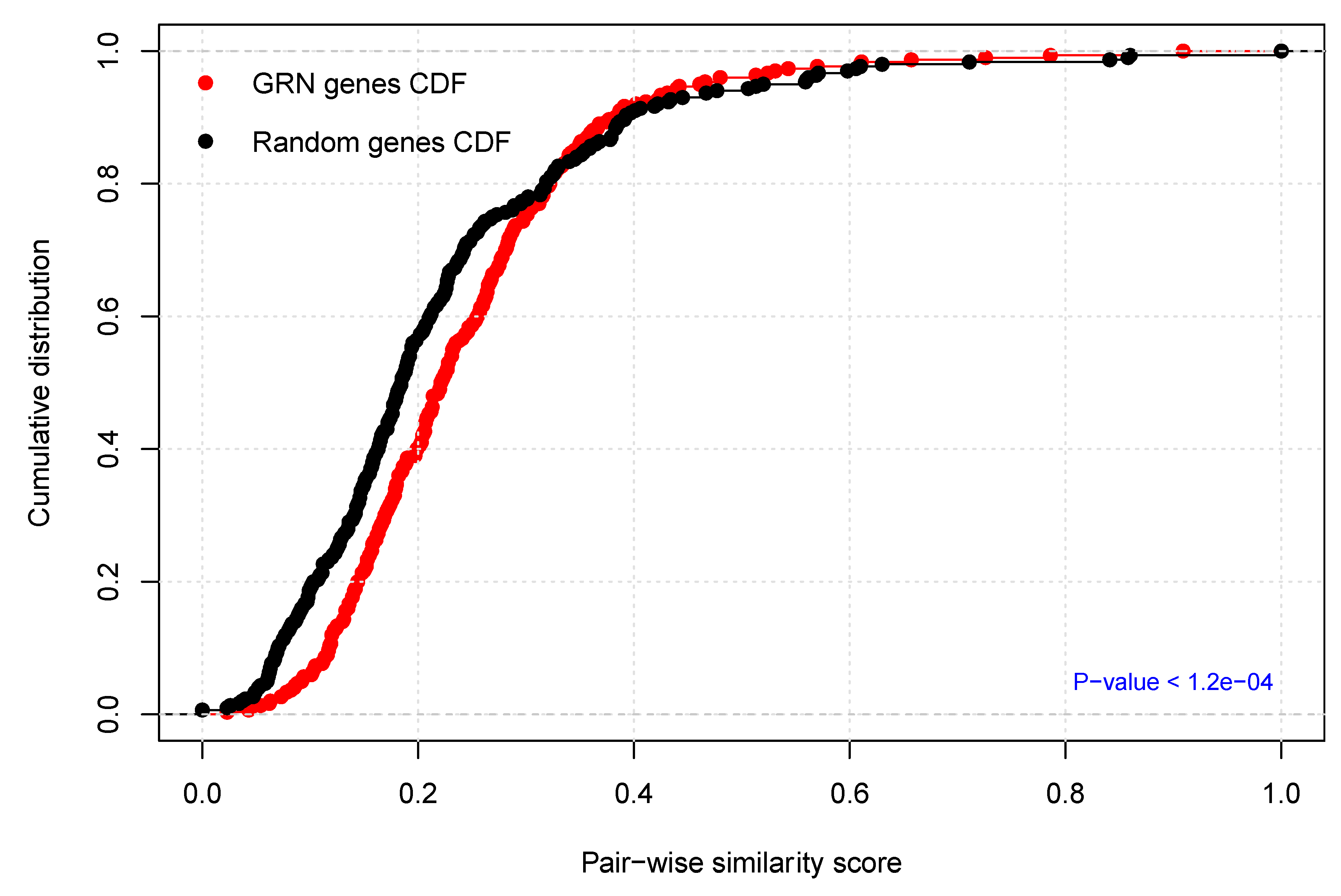
2.4. Functional Homogeneity within the Prostate Cancer (PCa-GRN) Network
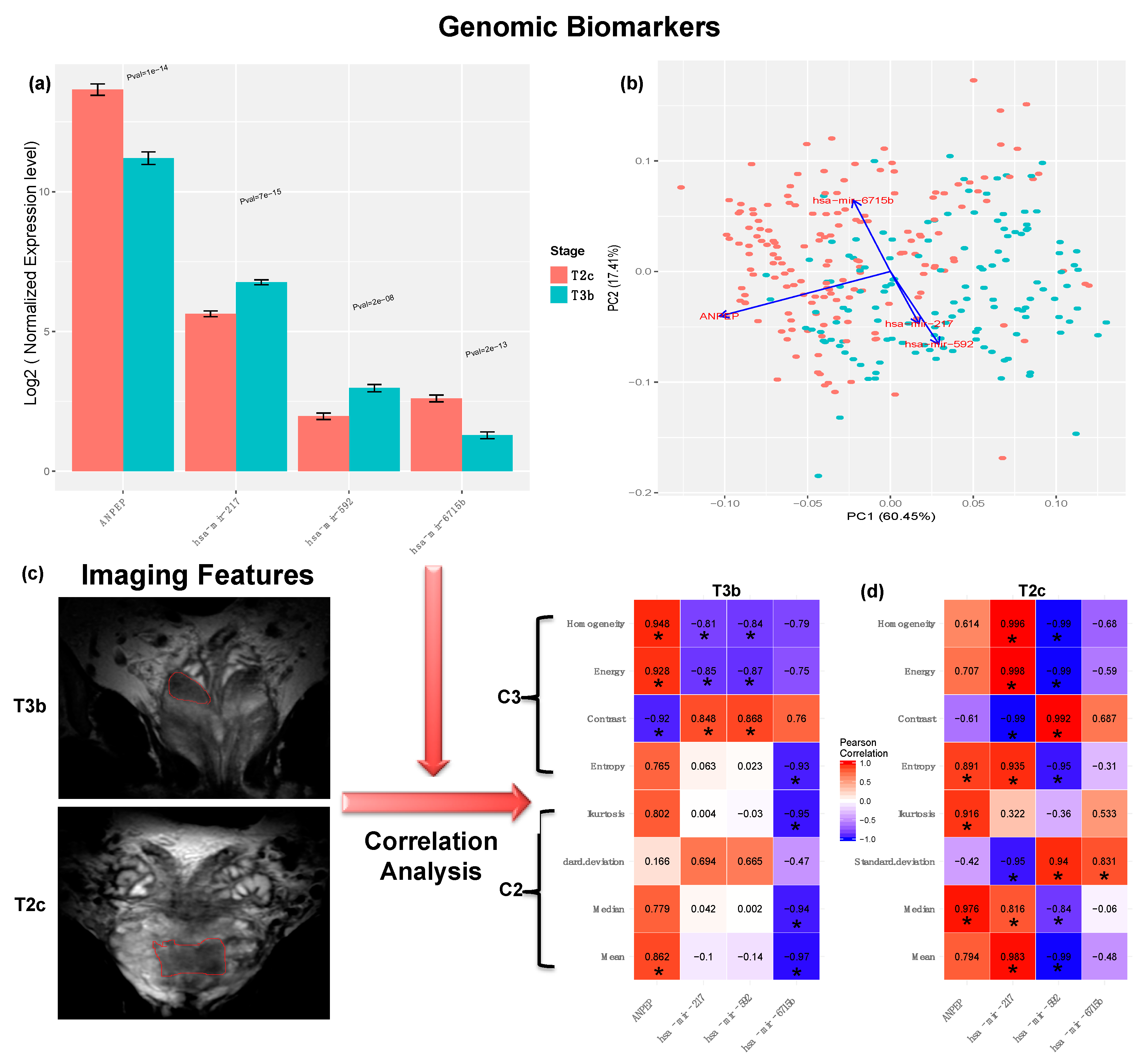
2.5. Biomarker Identification
2.6. Correlation Analysis with Aggressiveness-Related Imaging Features
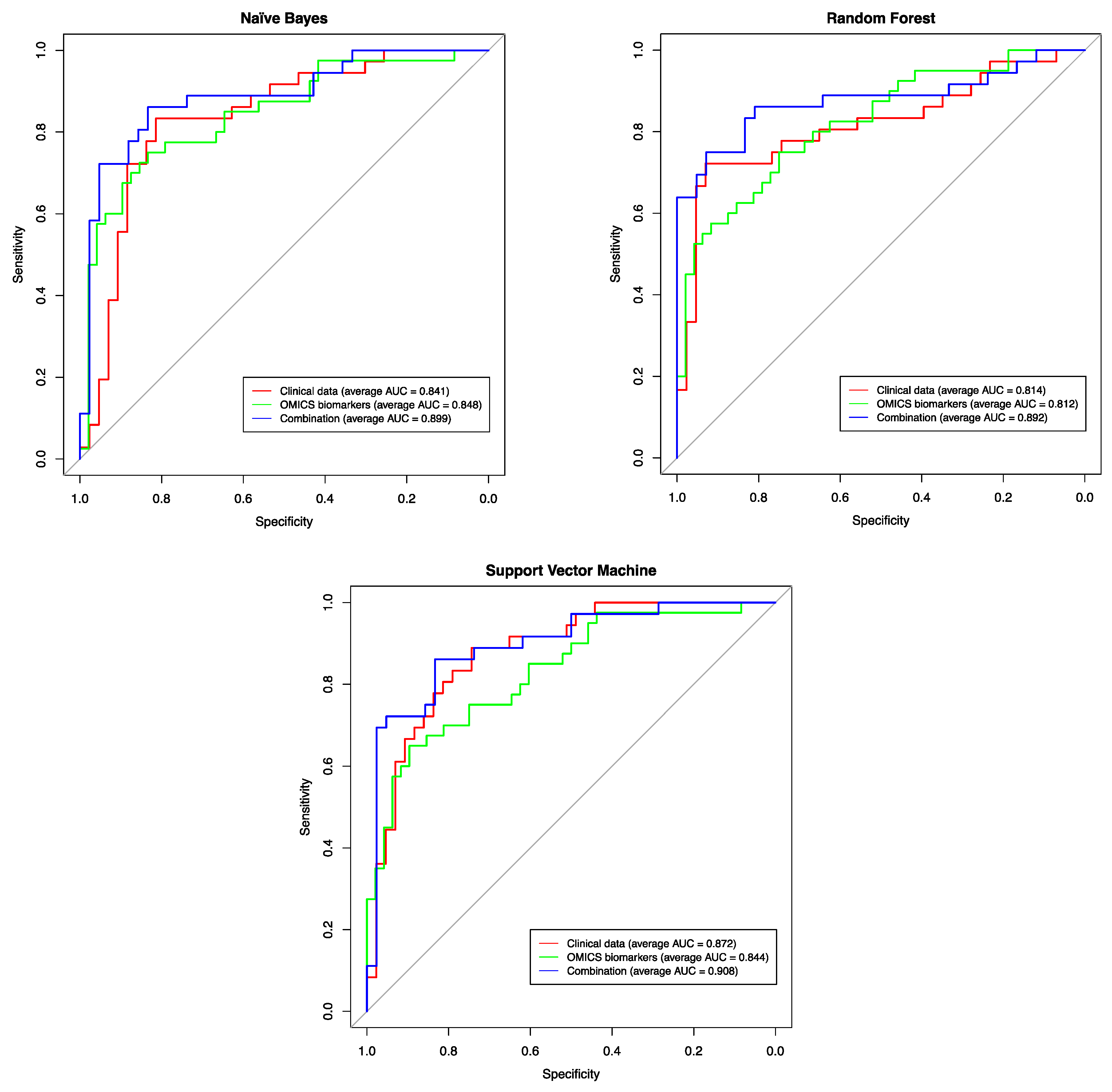
2.7. Assessing the Predictive Power of the Identified Biomarkers
3. Materials and Methods
3.1. Datasets
3.2. Data Pre-Processing
3.2.1. Genomic Data
3.2.2. Clinical Data
3.2.3. Imaging Data
3.3. Differential Expression and Association Analysis
3.4. Construction of Prostate-Specific GRN Network (PRAD-GRN)
3.5. Assessment of the Constructed Prostate Cancer (PCa-GRN) Network
3.5.1. Functional Homogeneity within the PCa-GRN Genes (Semantic Validation)
3.5.2. Enrichment Analysis of Genes and miRNAs
3.6. Correlation Analysis
3.7. Prediction Models
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Abeshouse, A.; Ahn, J.; Akbani, R.; Ally, A.; Amin, S.; Andry, C.D.; Annala, M.; Aprikian, A.; Armenia, J.; Arora, A. The molecular taxonomy of primary prostate cancer. Cell 2015, 163, 1011–1025. [Google Scholar] [CrossRef]
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef]
- Stoyanova, R.; Takhar, M.; Tschudi, Y.; Ford, J.C.; Solórzano, G.; Erho, N.; Balagurunathan, Y.; Punnen, S.; Davicioni, E.; Gillies, R.J. Prostate cancer radiomics and the promise of radiogenomics. Transl. Cancer Res. 2016, 5, 432. [Google Scholar] [CrossRef]
- Walz, J.; Joniau, S.; Chun, F.K.; Isbarn, H.; Jeldres, C.; Yossepowitch, O.; Chao-Yu, H.; Klein, E.A.; Scardino, P.T.; Reuther, A.; et al. Pathological results and rates of treatment failure in high-risk prostate cancer patients after radical prostatectomy. BJU Int. 2011, 107, 765–770. [Google Scholar] [CrossRef]
- Incoronato, M.; Aiello, M.; Infante, T.; Cavaliere, C.; Grimaldi, A.M.; Mirabelli, P.; Monti, S.; Salvatore, M. Radiogenomic analysis of oncological data: A technical survey. Int. J. Mol. Sci. 2017, 18, 805. [Google Scholar] [CrossRef]
- Barbieri, C.E.; Demichelis, F.; Rubin, M.A. Molecular genetics of prostate cancer: Emerging appreciation of genetic complexity. Histopathology 2012, 60, 187–198. [Google Scholar] [CrossRef]
- Jansen, F.H.; van Schaik, R.H.; Kurstjens, J.; Horninger, W.; Klocker, H.; Bektic, J.; Wildhagen, M.F.; Roobol, M.J.; Bangma, C.H.; Bartsch, G. Prostate-specific antigen (PSA) isoform p2PSA in combination with total PSA and free PSA improves diagnostic accuracy in prostate cancer detection. Eur. Urol. 2010, 57, 921–927. [Google Scholar] [CrossRef]
- Cookson, M.S.; Fleshner, N.E.; Soloway, S.M.; Fair, W.R. Correlation between Gleason score of needle biopsy and radical prostatectomy specimen: Accuracy and clinical implications. J. Urol. 1997, 157, 559–562. [Google Scholar] [CrossRef]
- Epstein, J.I.; Walsh, P.C.; Carmichael, M.; Brendler, C.B. Pathologic and clinical findings to predict tumor extent of nonpalpable (stage t1 c) prostate cancer. JAMA 1994, 271, 368–374. [Google Scholar] [CrossRef]
- Epstein, J.I.; Pizov, G.; Walsh, P.C. Correlation of pathologic findings with progression after radical retropubic prostatectomy. Cancer 1993, 71, 3582–3593. [Google Scholar] [CrossRef]
- Tosoian, J.J.; Chappidi, M.; Feng, Z.; Humphreys, E.B.; Han, M.; Pavlovich, C.P.; Epstein, J.I.; Partin, A.W.; Trock, B.J. Prediction of pathological stage based on clinical stage, serum prostate-specific antigen, and biopsy Gleason score: Partin Tables in the contemporary era. BJU Int. 2017, 119, 676–683. [Google Scholar] [CrossRef]
- Gospodarowicz, M.K.; Brierley, J.D.; Wittekind, C. TNM Classification of Malignant Tumours; John Wiley & Sons: Hoboken, NJ, USA, 2017. [Google Scholar]
- Mohler, J.L.; Armstrong, A.J.; Bahnson, R.R.; D’Amico, A.V.; Davis, B.J.; Eastham, J.A.; Enke, C.A.; Farrington, T.A.; Higano, C.S.; Horwitz, E.M. Prostate cancer, version 1.2016. J. Natl. Compr. Cancer Netw. 2016, 14, 19–30. [Google Scholar] [CrossRef]
- Dotan, Z.A.; Ramon, J. Nomograms as a tool in predicting prostate cancer prognosis. Eur. Urol. Suppl. 2009, 8, 721–724. [Google Scholar] [CrossRef]
- MSK. Memorial Sloan Kettering Cancer Center. Available online: https://www.mskcc.org/nomograms/prostate (accessed on 15 August 2019).
- Harada, T.; Abe, T.; Kato, F.; Matsumoto, R.; Fujita, H.; Murai, S.; Miyajima, N.; Tsuchiya, K.; Maruyama, S.; Kudo, K.; et al. Five-point Likert scaling on MRI predicts clinically significant prostate carcinoma. BMC Urol. 2015, 15, 91. [Google Scholar] [CrossRef]
- Briganti, A.; Karakiewicz, P.I.; Joniau, S.; Van Poppel, H. The motion: Nomograms should become a routine tool in determining prostate cancer prognosis. Eur. Urol. 2009, 55, 743–747. [Google Scholar] [CrossRef]
- Chun, F.K.H.; Karakiewicz, P.I.; Briganti, A.; Walz, J.; Kattan, M.W.; Huland, H.; Graefen, M. A critical appraisal of logistic regression-based nomograms, artificial neural networks, classification and regression-tree models, look-up tables and risk-group stratification models for prostate cancer. BJU Int. 2007, 99, 794–800. [Google Scholar] [CrossRef]
- Cosma, G.; Acampora, G.; Brown, D.; Rees, R.C.; Khan, M.; Pockley, A.G. Prediction of pathological stage in patients with prostate cancer: A neuro-fuzzy model. PLoS ONE 2016, 11, e0155856. [Google Scholar] [CrossRef]
- Hariri, A.R.; Weinberger, D.R. Imaging genomics. Br. Med. Bull. 2003, 65, 259–270. [Google Scholar] [CrossRef] [Green Version]
- Bai, H.X.; Lee, A.M.; Yang, L.; Zhang, P.; Davatzikos, C.; Maris, J.M.; Diskin, S.J. Imaging genomics in cancer research: Limitations and promises. Br. J. Radiol. 2016, 89, 20151030. [Google Scholar] [CrossRef]
- Bertoli, G.; Cava, C.; Castiglioni, I. MicroRNAs as biomarkers for diagnosis, prognosis and theranostics in prostate cancer. Int. J. Mol. Sci. 2016, 17, 421. [Google Scholar] [CrossRef]
- Bibikova, M.; Chudin, E.; Arsanjani, A.; Zhou, L.; Garcia, E.W.; Modder, J.; Kostelec, M.; Barker, D.; Downs, T.; Fan, J.B. Expression signatures that correlated with Gleason score and relapse in prostate cancer. Genomics 2007, 89, 666–672. [Google Scholar] [CrossRef] [Green Version]
- Agell, L.; Hernández, S.; Nonell, L.; Lorenzo, M.; Puigdecanet, E.; de Muga, S.; Juanpere, N.; Bermudo, R.; Fernández, P.L.; Lorente, J.A. A 12-gene expression signature is associated with aggressive histological in prostate cancer: SEC14L1 and TCEB1 genes are potential markers of progression. Am. J. Pathol. 2012, 181, 1585–1594. [Google Scholar] [CrossRef]
- Bismar, T.A.; Demichelis, F.; Riva, A.; Kim, R.; Varambally, S.; He, L.; Kutok, J.; Aster, J.C.; Tang, J.; Kuefer, R. Defining aggressive prostate cancer using a 12-gene model. Neoplasia 2006, 8, 59–68. [Google Scholar] [CrossRef]
- Cheville, J.C.; Karnes, R.J.; Therneau, T.M.; Kosari, F.; Munz, J.M.; Tillmans, L.; Basal, E.; Rangel, L.J.; Bergstralh, E.; Kovtun, I.V. Gene panel model predictive of outcome in men at high-risk of systemic progression and death from prostate cancer after radical retropubic prostatectomy. J. Clin. Oncol. 2008, 26, 3930. [Google Scholar] [CrossRef]
- Cuzick, J.; Swanson, G.P.; Fisher, G.; Brothman, A.R.; Berney, D.M.; Reid, J.E.; Mesher, D.; Speights, V.; Stankiewicz, E.; Foster, C.S. Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: A retrospective study. Lancet Oncol. 2011, 12, 245–255. [Google Scholar] [CrossRef]
- Larkin, S.; Holmes, S.; Cree, I.; Walker, T.; Basketter, V.; Bickers, B.; Harris, S.; Garbis, S.D.; Townsend, P.; Aukim-Hastie, C. Identification of markers of prostate cancer progression using candidate gene expression. Br. J. Cancer 2012, 106, 157. [Google Scholar] [CrossRef]
- Singh, D.; Febbo, P.G.; Ross, K.; Jackson, D.G.; Manola, J.; Ladd, C.; Tamayo, P.; Renshaw, A.A.; D’Amico, A.V.; Richie, J.P. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 2002, 1, 203–209. [Google Scholar] [CrossRef] [Green Version]
- Erho, N.; Crisan, A.; Vergara, I.A.; Mitra, A.P.; Ghadessi, M.; Buerki, C.; Bergstralh, E.J.; Kollmeyer, T.; Fink, S.; Haddad, Z. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS ONE 2013, 8, e66855. [Google Scholar] [CrossRef]
- Al-Kafaji, G.; Said, H.M.; Alam, M.A.; Al Naieb, Z.T. Blood-based microRNAs as diagnostic biomarkers to discriminate localized prostate cancer from benign prostatic hyperplasia and allow cancer-risk stratification. Oncol. Lett. 2018, 16, 1357–1365. [Google Scholar] [CrossRef]
- Stoyanova, R.; Pollack, A.; Takhar, M.; Lynne, C.; Parra, N.; Lam, L.L.; Alshalalfa, M.; Buerki, C.; Castillo, R.; Jorda, M.; et al. Association of multiparametric MRI quantitative imaging features with prostate cancer gene expression in MRI-targeted prostate biopsies. Oncotarget 2016, 7, 53362–53376. [Google Scholar] [CrossRef]
- Corn, P.G.; Wang, F.; McKeehen, W.; Navone, N. Targeting fibroblast growth factor pathways in prostate cancer. Clin. Cancer Res. 2013, 19, 5856–5866. [Google Scholar] [CrossRef]
- Chandran, U.R.; Ma, C.; Dhir, R.; Bisceglia, M.; Lyons-Weiler, M.; Liang, W.; Michalopoulos, G.; Becich, M.; Monzon, F.A. Gene expression profiles of prostate cancer reveal involvement of multiple molecular pathways in the metastatic process. BMC Cancer 2007, 7, 64. [Google Scholar] [CrossRef]
- Mazaris, E.; Tsiotras, A. Molecular pathways in prostate cancer. Nephro Urol. Mon. 2013, 5, 792. [Google Scholar] [CrossRef]
- Vivanco, I.; Sawyers, C.L. The phosphatidylinositol 3-kinase–AKT pathway in human cancer. Nat. Rev. Cancer 2002, 2, 489. [Google Scholar] [CrossRef]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 2011, 6, e21800. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, H.E.; Liu, Z. MicroRNA-147 suppresses proliferation, invasion and migration through the AKT/mTOR signaling pathway in breast cancer. Oncol. Lett. 2016, 11, 405–410. [Google Scholar] [CrossRef]
- Zhang, Y.; Talmon, G.; Wang, J. MicroRNA-587 antagonizes 5-FU-induced apoptosis and confers drug resistance by regulating PPP2R1B expression in colorectal cancer. Cell Death Dis. 2016, 7, e2525. [Google Scholar] [CrossRef]
- Liu, A.Y.; Roudier, M.P.; True, L.D. Heterogeneity in primary and metastatic prostate cancer as defined by cell surface CD profile. Am. J. Pathol. 2004, 165, 1543–1556. [Google Scholar] [CrossRef]
- Dall’Era, M.A.; True, L.D.; Siegel, A.F.; Porter, M.P.; Sherertz, T.M.; Liu, A.Y. Differential expression of CD10 in prostate cancer and its clinical implication. BMC Urol. 2007, 7, 3. [Google Scholar] [CrossRef]
- Sørensen, K.D.; Abildgaard, M.O.; Haldrup, C.; Ulhøi, B.P.; Kristensen, H.; Strand, S.; Parker, C.; Høyer, S.; Borre, M.; Ørntoft, T.F. Prognostic significance of aberrantly silenced ANPEP expression in prostate cancer. Br. J. Cancer 2013, 108, 420. [Google Scholar] [CrossRef]
- Lin, H.M.; Nikolic, I.; Yang, J.; Castillo, L.; Deng, N.; Chan, C.L.; Yeung, N.K.; Dodson, E.; Elsworth, B.; Spielman, C.; et al. MicroRNAs as potential therapeutics to enhance chemosensitivity in advanced prostate cancer. Sci. Rep. 2018, 8, 7820. [Google Scholar] [CrossRef]
- Lv, Z.; Rao, P.; Li, W. MiR-592 represses FOXO3 expression and promotes the proliferation of prostate cancer cells. Int. J. Clin. Exp. Med. 2015, 8, 15246–15253. [Google Scholar]
- Giulietti, M.; Occhipinti, G.; Principato, G.; Piva, F. Identification of candidate miRNA biomarkers for pancreatic ductal adenocarcinoma by weighted gene co-expression network analysis. Cell. Oncol. (Dordr.) 2017, 40, 181–192. [Google Scholar] [CrossRef]
- Rosset, A.; Spadola, L.; Ratib, O. OsiriX: An open-source software for navigating in multidimensional DICOM images. J. Digit. Imaging 2004, 17, 205–216. [Google Scholar] [CrossRef]
- Wibmer, A.; Hricak, H.; Gondo, T.; Matsumoto, K.; Veeraraghavan, H.; Fehr, D.; Zheng, J.; Goldman, D.; Moskowitz, C.; Fine, S.W.; et al. Haralick texture analysis of prostate MRI: Utility for differentiating non-cancerous prostate from prostate cancer and differentiating prostate cancers with different Gleason scores. Eur. Radiol. 2015, 25, 2840–2850. [Google Scholar] [CrossRef]
- Shen, J.; Hruby, G.W.; McKiernan, J.M.; Gurvich, I.; Lipsky, M.J.; Benson, M.C.; Santella, R.M. Dysregulation of circulating microRNAs and prediction of aggressive prostate cancer. Prostate 2012, 72, 1469–1477. [Google Scholar] [CrossRef] [Green Version]
- Tsao, C.W.; Liu, C.Y.; Cha, T.L.; Wu, S.T.; Sun, G.H.; Yu, D.S.; Chen, H.I.; Chang, S.Y.; Chen, S.C.; Hsu, C.Y. Artificial neural network for predicting pathological stage of clinically localized prostate cancer in a Taiwanese population. J. Chin. Med. Assoc. 2014, 77, 513–518. [Google Scholar] [CrossRef] [Green Version]
- Clark, K.; Vendt, B.; Smith, K.; Freymann, J.; Kirby, J.; Koppel, P.; Moore, S.; Phillips, S.; Maffitt, D.; Pringle, M. The Cancer Imaging Archive (TCIA): Maintaining and operating a public information repository. J. Digit. Imaging 2013, 26, 1045–1057. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas Prostate Adenocarcinoma (TCGA-PRAD) Data Collection. Available online: https://portal.gdc.cancer.gov/projects/TCGA-PRAD (accessed on 11 January 2019).
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Ihaka, R.; Gentleman, R. R: A language for data analysis and graphics. J. Comput. Graph. Stat. 1996, 5, 299–314. [Google Scholar]
- Gallón, S.; Loubes, J.M.; Maza, E. Statistical properties of the quantile normalization method for density curve alignment. Math. Biosci. 2013, 242, 129–142. [Google Scholar] [CrossRef] [Green Version]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671. [Google Scholar] [CrossRef]
- Hochberg, Y.; Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 1990, 9, 811–818. [Google Scholar] [CrossRef]
- Hamed, M.; Spaniol, C.; Nazarieh, M.; Helms, V. TFmiR: A web server for constructing and analyzing disease-specific transcription factor and miRNA co-regulatory networks. Nucleic Acids Res. 2015, 43, W283–W288. [Google Scholar] [CrossRef]
- Li, Y.; Qiu, C.; Tu, J.; Geng, B.; Yang, J.; Jiang, T.; Cui, Q. HMDD v2. 0: A database for experimentally supported human microRNA and disease associations. Nucleic Acids Res. 2013, 42, D1070–D1074. [Google Scholar] [CrossRef]
- Piñero, J.; Queralt-Rosinach, N.; Bravo, À.; Deu-Pons, J.; Bauer-Mehren, A.; Baron, M.; Sanz, F.; Furlong, L.I. DisGeNET: A discovery platform for the dynamical exploration of human diseases and their genes. Database 2015, 2015, bav028. [Google Scholar] [CrossRef]
- Piñero, J.; Bravo, À.; Queralt-Rosinach, N.; Gutiérrez-Sacristán, A.; Deu-Pons, J.; Centeno, E.; García-García, J.; Sanz, F.; Furlong, L.I. DisGeNET: A comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2016, gkw943. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Yu, G.; Li, F.; Qin, Y.; Bo, X.; Wu, Y.; Wang, S. GOSemSim: An R package for measuring semantic similarity among GO terms and gene products. Bioinformatics 2010, 26, 976–978. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2008, 4, 44. [Google Scholar] [CrossRef]
- Lu, M.; Shi, B.; Wang, J.; Cao, Q.; Cui, Q. TAM: A method for enrichment and depletion analysis of a microRNA category in a list of microRNAs. BMC Bioinform. 2010, 11, 419. [Google Scholar] [CrossRef]
- Hamed, M.; Trumm, J.; Spaniol, C.; Sethi, R.; Irhimeh, M.R.; Fuellen, G.; Paulsen, M.; Helms, V. Linking Hematopoietic Differentiation to Co-Expressed Sets of Pluripotency-Associated and Imprinted Genes and to Regulatory microRNA-Transcription Factor Motifs. PLoS ONE 2017, 12, e0166852. [Google Scholar] [CrossRef]
- Kuhn, M. Caret: Classification and Regression Training. Astrophys. Source Code Library. 2015. Available online: https://rdrr.io/rforge/caret/ (accessed on 23 March 2019).
- Robin, X.; Turck, N.; Hainard, A.; Tiberti, N.; Lisacek, F.; Sanchez, J.C.; Müller, M. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011, 12, 77. [Google Scholar] [CrossRef]
| Pathological STAGE | Count | Age Median (Min–Max) | PSA-Value Median (Min–Max) | Gleason Score | Count | Clinical Stage | Count | Biochemical Recurrence | Count | Ethnicity | Count |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Primary + Secondary | Stage | ||||||||||
| T2c | 164 | 59 (41–77) | 0.1 (0.01–14.69) | 3 + 3 | 25 | T1b | 0 | Yes | 6 | Black or African American | 3 |
| 3 + 4 | 84 | T1c | 84 | No | 131 | White | 58 | ||||
| 4 + 3 | 31 | T2 | 4 | Not available | 27 | Not available | 103 | ||||
| ≥8 | 24 | T2a | 16 | ||||||||
| T2b | 10 | ||||||||||
| T2c | 23 | ||||||||||
| T3a | 2 | ||||||||||
| T3b | 0 | ||||||||||
| T4 | 0 | ||||||||||
| Not available | 25 | ||||||||||
| T3b | 134 | 62 (46–78) | 0.1 (0.01–37.36) | 3 + 3 | 1 | T1b | 2 | Yes | 29 | White | 27 |
| 3 + 4 | 8 | T1c | 23 | No | 94 | Not available | 107 | ||||
| 4 + 3 | 21 | T2 | 6 | Not available | 11 | ||||||
| ≥8 | 104 | T2a | 14 | ||||||||
| T2b | 12 | ||||||||||
| T2c | 16 | ||||||||||
| T3a | 14 | ||||||||||
| T3b | 16 | ||||||||||
| T4 | 1 | ||||||||||
| Not available | 30 |
| Enriched Pathways Which Characterize the T2c Stage but Not the T3b Stage | ||
| Pathways | Involved Genes | Adj-Pval |
| hsa05218: Melanoma | FGF6, FGF8, FGF23, FGF3 | 3.00 × 10−3 |
| hsa04010: MAPK signaling pathway | FGF6, DUSP4, FGF8, FGF23, FGF3, PLA2G4D | 3.00 × 10−3 |
| hsa04810: Regulation of actin cytoskeleton | FGF6, FGF8, FGF23, MYLPF, FGF3 | 9.00 × 10−3 |
| hsa04151: PI3K-Akt signaling pathway | FGF6, FGF8, COL6A5, FGF23, FGF3, EIF4E1B | 1.00 × 10−2 |
| hsa04014: Ras signaling pathway | FGF6, FGF8, FGF23, FGF3, PLA2G4D | 1.1 × 10−2 |
| hsa04015: Rap1 signaling pathway | FGF6, FGF8, FGF23, FGF3 | 4.7 × 10−2 |
| Enriched Pathways Which Characterize the T3b Stage But Not the T2c Stage | ||
| Pathways | Involved Genes | Adj-Pval |
| hsa04080: Neuroactive ligand-receptor interaction | GABRD, MCHR1, GABRA2, GABRA3, GABRB2, ADCYAP1R1, GRIA3, NTSR2, GHRHR, HRH3, PRLR, GALR1, HRH2, P2RX2, NPFFR1, CHRNA1, ADRA1D, GABRQ | 2.23 × 10−5 |
| hsa05033: Nicotine addiction | GABRD, GABRA2, GABRB2, GABRA3, GRIA3, GABRQ | 9.70 × 10−4 |
| hsa04972: Pancreatic secretion | KCNMA1, CD38, ATP2B4, PLA2G2A, PLA2G2C, CPA1, ATP1A2, PRKCB | 2.09 × 10−3 |
| hsa05143: African trypanosomiasis | IL6, HBA2, HBB, SELE, PRKCB | 3.58 × 10−3 |
| hsa04727: GABAergic synapse | GABRD, PLCL1, GABRA2, GABRB2, GABRA3, GABRQ, PRKCB | 5.94 × 10−3 |
| hsa04510: Focal adhesion | CAV3, CAV1, RASGRF1, PAK3, RAC3, ACTN2, ITGB3, FLNC, COL4A6, PRKCB, FN1 | 6.53 × 10−3 |
| hsa04723: Retrograde endocannabinoid signaling | GABRD, GABRA2, GABRB2, GABRA3, GRIA3, GABRQ, PRKCB | 1.34 × 10−2 |
| hsa05144: Malaria | IL6, CXCL8, HBA2, HBB, SELE | 1.46 × 10−2 |
| hsa05146: Amoebiasis | GNAL, IL6, CXCL8, ACTN2, COL4A6, PRKCB, FN1 | 1.67 × 10−2 |
| hsa04020: Calcium signaling pathway | GNAL, CD38, ATP2B4, ERBB4, HRH2, PLN, P2RX2, ADRA1D, PRKCB | 2.27 × 10−2 |
| hsa04970: Salivary secretion | KCNMA1, CD38, ATP2B4, ATP1A2, ADRA1D, PRKCB | 2.53 × 10−2 |
| hsa04270: Vascular smooth muscle contraction | KCNMA1, ACTG2, PLA2G2A, PLA2G2C, ADRA1D, KCNMB1, PRKCB | 2.77 × 10−2 |
| hsa05032: Morphine addiction | GABRD, GABRA2, GABRB2, GABRA3, GABRQ, PRKCB | 3.14 × 10−2 |
| hsa05205: Proteoglycans in cancer | CAV3, MIR10B, WNT16, CAV1, ERBB4, ITGB3, FLNC, PRKCB, FN1 | 4.02 × 10−2 |
| hsa05412: Arrhythmogenic right ventricular cardiomyopathy (ARVC) | SGCG, DMD, ACTN2, ITGB3, CTNNA3 | 4.85 × 10−2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fischer, S.; Tahoun, M.; Klaan, B.; Thierfelder, K.M.; Weber, M.-A.; Krause, B.J.; Hakenberg, O.; Fuellen, G.; Hamed, M. A Radiogenomic Approach for Decoding Molecular Mechanisms Underlying Tumor Progression in Prostate Cancer. Cancers 2019, 11, 1293. https://doi.org/10.3390/cancers11091293
Fischer S, Tahoun M, Klaan B, Thierfelder KM, Weber M-A, Krause BJ, Hakenberg O, Fuellen G, Hamed M. A Radiogenomic Approach for Decoding Molecular Mechanisms Underlying Tumor Progression in Prostate Cancer. Cancers. 2019; 11(9):1293. https://doi.org/10.3390/cancers11091293
Chicago/Turabian StyleFischer, Sarah, Mohamed Tahoun, Bastian Klaan, Kolja M. Thierfelder, Marc-André Weber, Bernd J. Krause, Oliver Hakenberg, Georg Fuellen, and Mohamed Hamed. 2019. "A Radiogenomic Approach for Decoding Molecular Mechanisms Underlying Tumor Progression in Prostate Cancer" Cancers 11, no. 9: 1293. https://doi.org/10.3390/cancers11091293
APA StyleFischer, S., Tahoun, M., Klaan, B., Thierfelder, K. M., Weber, M.-A., Krause, B. J., Hakenberg, O., Fuellen, G., & Hamed, M. (2019). A Radiogenomic Approach for Decoding Molecular Mechanisms Underlying Tumor Progression in Prostate Cancer. Cancers, 11(9), 1293. https://doi.org/10.3390/cancers11091293






