A Multi-Center Study of BRCA1 and BRCA2 Germline Mutations in Mexican-Mestizo Breast Cancer Families Reveals Mutations Unreported in Latin American Population
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Patient Cohort
4.2. DNA Isolation
4.3. Ion Semiconductor Sequencing
4.4. Data Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 49, 394–424. [Google Scholar] [CrossRef]
- Antoniou, A.; Pharoah, P.; Narod, S.; Risch, H.; Eyfjord, J.; Hopper, J.; Loman, N.; Olsson, H.; Johannsson, O.; Borg, Å.; et al. Average Risks of Breast and Ovarian Cancer Associated with BRCA1 or BRCA2 Mutations Detected in Case Series Unselected for Family History: A Combined Analysis of 22 Studies. Am. J. Hum. Genet. 2003, 72, 1117–1130. [Google Scholar] [CrossRef] [PubMed]
- Petrucelli, N.; Daly, M.B.; Feldman, G.L. Hereditary breast and ovarian cancer due to mutations in BRCA1 and BRCA2. Genet. Med. 2010, 12, 245–259. [Google Scholar] [CrossRef] [PubMed]
- Rebbeck, T.R.; Friebel, T.M.; Friedman, E.; Hamann, U.; Huo, D.; Kwong, A.; Olah, E.; Olopade, O.I.; Solano, A.R.; Teo, S.; et al. Mutational spectrum in a worldwide study of 29,700 families with BRCA1 or BRCA2 mutations. Hum. Mutat. 2018, 39, 593–620. [Google Scholar] [CrossRef] [PubMed]
- Corso, G.; Feroce, I.; Intra, M.; Toesca, A.; Magnoni, F.; Sargenti, M.; Naninato, P.; Caldarella, P.; Pagani, G.; Vento, A.; et al. BRCA1/2 germline missense mutations: A systematic review. Eur. J. Cancer Prev. 2017, 27, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Riahi, A.; Kharrat, M.; Ghourabi, M.; Khomsi, F.; Gamoudi, A.; Lariani, I.; May, A.; Rahal, K.; Chaabouni Bouhamed, H. Mutation spectrum and prevalence of BRCA1 and BRCA2 genes in patients with familial and early-Onset breast/ovarian cancer from Tunisia. Clin. Genet. 2015, 87, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Maistro, S.; Teixeira, N.; Encinas, G.; Katayama, M.; Niewiadonski, V.; Cabral, L.; Ribeiro, R.; Junior, N.; de Gouvêa, A.; Carraro, D.; et al. Germline mutations in BRCA1 and BRCA2 in epithelial ovarian cancer patients in Brazil. BMC Cancer 2016, 16, 934. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Yang, J.; Li, L.; Cao, D.; Yu, M.; Shen, K.; BGI Group. Germline and somatic mutations in homologous recombination genes among Chinese ovarian cancer patients detected using next-Generation sequencing. J. Gynecol. Oncol. 2017, 28, 39. [Google Scholar] [CrossRef]
- Calderón-Garcidueñas, A.; Parás-Barrientos, F.; Cárdenas-Ibarra, L.; González-Guerrero, J.; Villarreal-Ríos, E.; Staines-Boone, T.; Barrera-Saldaña, H. Risk factors of breast cancer in Mexican women. Salud Públ. Méx. 2000, 42, 26–33. [Google Scholar] [CrossRef]
- Villarreal-Garza, C.; Aguila, C.; Magallanes-Hoyos, M.C.; Mohar, A.; Bargalló, E.; Meneses, A.; Cazap, E.; Gomez, H.; López-Carrillo, L.; Chávarri-Guerra, Y.; et al. Breast Cancer in Young Women in Latin America: An Unmet, Growing Burden. Oncology 2013, 18, 1298–1306. [Google Scholar] [CrossRef]
- Lara-Medina, F.; Pérez-Sánchez, V.; Saavedra-Pérez, D.; Blake-Cerda, M.; Arce, C.; Motola-Kuba, D.; Villarreal-Garza, C.; González-Angulo, A.; Bargalló, E.; Aguilar, J.; et al. Triple-Negative breast cancer in Hispanic patients. Cancer 2011, 117, 3658–3669. [Google Scholar] [CrossRef]
- Gonzalez-Angulo, A.M.; Timms, K.M.; Liu, S.; Chen, H.; Litton, J.K.; Potter, J.; Lanchbury, J.S.; Stemke-Hale, K.; Hennessy, B.T.; Arun, B.K.; et al. Incidence and Outcome of BRCA Mutations in Unselected Patients with Triple Receptor-Negative Breast Cancer. Clin. Cancer Res. 2011, 17, 1082–1089. [Google Scholar] [CrossRef]
- Vaca-Paniagua, F.; Alvarez-Gomez, R.; Fragoso-Ontiveros, V.; Vidal-Millan, S.; Herrera, L.; Cantú, D.; Bargallo-Rocha, E.; Mohar, A.; Lopez-Camarillo, C.; Pérez-Plasencia, C. Full-Exon pyrosequencing screening of BRCA germline mutations in Mexican women with inherited breast and ovarian cancer. PLoS ONE 2012, 7, 37432. [Google Scholar] [CrossRef]
- Ashton-Prolla, P.; Vargas, F. Prevalence and impact of founder mutations in hereditary breast cancer in Latin America. Genet. Mol. Biol. 2014, 37, 234–240. [Google Scholar] [CrossRef]
- Blay, P.; Santamaría, I.; Pitiot, A.S.; Luque, M.; Alvarado, M.G.; Lastra, A.; Fernández, Y.; Paredes, Á.; Freije, J.M.; Balbín, M. Mutational analysis of BRCA1 and BRCA2 in hereditary breast and ovarian cancer families from Asturias (Northern Spain). BMC Cancer 2013, 13, 243. [Google Scholar] [CrossRef]
- Weitzel, J.N.; Clague, J.; Martir-Negron, A.; Ogaz, R.; Herzog, J.; Ricker, C.; Jungbluth, C.; Cina, C.; Duncan, P.; Unzeitig, G.; et al. Prevalence and Type of BRCA Mutations in Hispanics Undergoing Genetic Cancer Risk Assessment in the Southwestern United States: A Report from the Clinical Cancer Genetics Community Research Network. J. Clin. Oncol. 2012, 31, 210–216. [Google Scholar] [CrossRef]
- Fernández-Lopez, J.; Romero-Córdoba, S.; Rebollar-Vega, R.; Alfaro-Ruiz, L.; Jiménez-Morales, S.; Beltrán-Anaya, F.; Arellano-Llamas, R.; Cedro-Tanda, A.; Rios-Romero, M.; Ramirez-Florencio, M.; et al. Population and breast cancer patients’ analysis reveals the diversity of genomic variation of the BRCA genes in the Mexican population. Hum. Genom. 2019, 13, 3. [Google Scholar] [CrossRef]
- Santos, C.; Peixoto, A.; Rocha, P.; Vega, A.; Soares, M.; Cerveira, N.; Bizarro, S.; Pinheiro, M.; Pereira, D.; Rodrigues, H.; et al. Haplotype and quantitative transcript analyses of Portuguese breast/ovarian cancer families with the BRCA1 R71G founder mutation of Galician origin. Fam. Cancer 2009, 8, 203–208. [Google Scholar] [CrossRef]
- Weitzel, J.N.; Lagos, V.; Blazer, K.R.; Nelson, R.; Ricker, C.; Herzog, J.; McGuire, C.; Neuhausen, S. Prevalence of BRCA Mutations and Founder Effect in High-Risk Hispanic Families. Cancer Epidemiol. Biomark. 2005, 14, 1666–1671. [Google Scholar] [CrossRef]
- Torres-Mejía, G.; Royer, R.; Llacuachaqui, M.; Akbari, M.R.; Giuliano, A.R.; Martínez-Matsushita, L.; Angeles-Llerenas, A.; Ortega-Olvera, C.; Ziv, E.; Lazcano-Ponce, E.; et al. Recurrent BRCA1 and BRCA2 Mutations in Mexican Women with Breast Cancer. Cancer Epidemiol. Prev. Biomark. 2015, 24, 498–505. [Google Scholar] [CrossRef]
- Gonzalez-Hormazabal, P.; Gutierrez-Enriquez, S.; Gaete, D.; Reyes, J.M.; Peralta, O.; Waugh, E.; Gomez, F.; Margarit, S.; Bravo, T.; Blanco, R.; et al. Spectrum of BRCA1/2 point mutations and genomic rearrangements in high-Risk breast/ovarian cancer Chilean families. Breast Cancer Res. Treat. 2011, 126, 705–716. [Google Scholar] [CrossRef]
- de Jiménez, I.; Cardeñosa, E.; Suela, S.; González, E.; Carretero, I.; Gandía, B.; Bertran, A.; Maicas, M.; Ponce, C.; Heras, A.; et al. Low prevalence of BRCA1 and BRCA2 mutations in the sporadic breast cancer of Spanish population. Fam. Cancer 2012, 11, 49–56. [Google Scholar] [CrossRef]
- Silva, F.C.; Lisboa, B.C.; Figueiredo, M.C.; Torrezan, G.T.; Mntos, É.; Krepischi, A.C.; Rossi, B.M.; Achatz, M.I.; Carraro, D.M. Hereditary breast and ovarian cancer: Assessment of point mutations and copy number variations in Brazilian patients. BMC Med. Genet. 2014, 15, 55. [Google Scholar] [CrossRef]
- Carraro, D.; Folgueira, M.; Lisboa, B.; Olivieri, E.; Krepischi, A.; de Carvalho, A.; de Mota, L.; Puga, R.; do Maciel, M.; Michelli, R.; et al. Comprehensive Analysis of BRCA1, BRCA2 and TP53 Germline Mutation and Tumor Characterization: A Portrait of Early-Onset Breast Cancer in Brazil. PLoS ONE 2013, 8, 57581. [Google Scholar] [CrossRef]
- Caldes, T.; de la Hoya, M.; Tosar, A.; Sulleiro, S.; Godino, J.; Ibañez, D.; Martin, M.; Perez-Segura, P.; Diaz-Rubio, E. A breast cancer family from Spain with germline mutations in both the BRCA1 and BRCA2 genes. J. Med. Genet. 2002, 39, 44. [Google Scholar] [CrossRef]
- de la Hoya, M.; Osorio, A.; Godino, J.; Sulleiro, S.; Tosar, A.; Perez-Segura, P.; Fernandez, C.; Rodríguez, R.; Díaz-Rubio, E.; Benítez, J.; et al. Association between BRCA1 and BRCA2 mutations and cancer phenotype in Spanish breast/ovarian cancer families: Implications for genetic testing. Int. J. Cancer 2002, 97, 466–471. [Google Scholar] [CrossRef]
- Ewald, I.P.; Izetti, P.; Vargas, F.R.; Moreira, M.A.; Moreira, A.S.; Moreira-Filho, C.A.; Cunha, D.R.; Hamaguchi, S.; Camey, S.A.; Schmidt, A.; et al. Prevalence of the BRCA1 founder mutation c.5266dupin Brazilian individuals at-Risk for the hereditary breast and ovarian cancer syndrome. Hered. Cancer Clin. Pract. 2011, 9, 12. [Google Scholar] [CrossRef]
- De Jiménez, I.; Casado, Z.; Suela, S.; Cardeñosa, E.; Guerrero, J.; Huerta, Á.; González, I.; Heras, A.; Fita, M.; García, I.; et al. Novel and recurrent BRCA1/BRCA2 mutations in early onset and familial breast and ovarian cancer detected in the Program of Genetic Counseling in Cancer of Valencian Community (eastern Spain). Relationship of family phenotypes with mutation prevalence. Fam. Cancer 2013, 12, 767–777. [Google Scholar]
- Llort, G.; Muñoz, C.; Tuser, M.; Guillermo, I.; Lluch, J.; Bale, A.E.; Franco, M. Low frequency of recurrent BRCA1 and BRCA2 mutations in Spain. Hum. Mutat. 2002, 19, 307. [Google Scholar] [CrossRef]
- Rodriguez, R.; Esperon, A.; Ropero, R.; Rubio, M.; Rodriguez, R.; Ortiz, R.M.; Anta, J.J.; de los Rios, M.; Carnesolta, D.; del Olivera, M.C.; et al. Prevalence of BRCA1 and BRCA2 mutations in breast cancer patients from Cuba. Fam. Cancer 2008, 7, 275–279. [Google Scholar] [CrossRef]
- Cardeñosa, E.; Gilabert, P.; de Jimenez, I.; Suela, S.; González, E.; González, I.; Huerta, Á.; Ponce, C.; de Dueñas, E. Broad BRCA1 and BRCA2 mutational spectrum and high incidence of recurrent and novel mutations in the eastern Spain population. Breast Cancer Res. Treat. 2010, 121, 257–260. [Google Scholar] [CrossRef]
- Villarreal-Garza, C.; Alvarez-Gomez, R.; Pérez-Plasencia, C.; Herrera, L.A.; Herzog, J.; Castillo, D.; Mohar, A.; Castro, C.; Gallardo, L.N.; Gallardo, D.; et al. Significant clinical impact of recurrent BRCA1and BRCA2mutations in Mexico. Cancer 2014, 121, 372–378. [Google Scholar] [CrossRef]
- Rodríguez, A.; Llacuachaqui, M.; Pardo, G.; Royer, R.; Larson, G.; Weitzel, J.N.; Narod, S.A. BRCA1 and BRCA2 mutations among ovarian cancer patients from Colombia. Gynecol. Oncol. 2012, 124, 236–243. [Google Scholar] [CrossRef]
- Ruiz-Flores, P.; Sinilnikova, O.M.; Badzioch, M.; Calderon-Garcidueñas, A.L.; Chopin, S.; Fabrice, O.; González-Guerrero, J.F.; Szabo, C.; Lenoir, G.; Goldgar, D.E.; et al. BRCA1 and BRCA2 mutation analysis of early-Onset and familial breast cancer cases in Mexico. Hum. Mutat. 2002, 20, 474–475. [Google Scholar] [CrossRef]
- Vidal-Millán, S.; Taja-Chayeb, L.; Gutiérrez-Hernández, O.; Ugalde, R.M.; Robles-Vidal, C.; Bargallo-Rocha, E.; Mohar-Betancourt, A.; Dueñas-González, A. Mutational analysis of BRCA1 and BRCA2 genes in Mexican breast cancer patients. Eur. J. Gynaecol. Oncol. 2009, 30, 527–530. [Google Scholar]
- Villarreal-Garza, C.; Weitzel, J.; Llacuachaqui, M.; Sifuentes, E.; Magallanes-Hoyos, M.; Gallardo, L.; Alvarez-Gómez, R.; Herzog, J.; Castillo, D.; Royer, R.; et al. The prevalence of BRCA1 and BRCA2 mutations among young Mexican women with triple-Negative breast cancer. Breast Cancer Res. Treat. 2015, 150, 389–394. [Google Scholar] [CrossRef]
- Bu, R.; Siraj, A.K.; Al-Obaisi, K.; Beg, S.; Hazmi, M.; Ajarim, D.; Tulbah, A.; Al-Dayel, F.; Al-Kuraya, K.S. Identification of novel BRCA founder mutations in Middle Eastern breast cancer patients using capture and Sanger sequencing analysis. Int. J. Cancer 2016, 139, 1091–1097. [Google Scholar] [CrossRef]
- Lang, G.; Shi, J.; Hu, X.; Zhang, C.; Shan, L.; Song, C.; Zhuang, Z.; Cao, A.; Ling, H.; Yu, K.; et al. The spectrum of BRCA mutations and characteristics of BRCA-associated breast cancers in China: Screening of 2,991 patients and 1,043 controls by next-Generation sequencing. Int. J. Cancer 2017, 141, 129–142. [Google Scholar] [CrossRef]
- Bernstein-Molho, R.; Singer, A.; Laitman, Y.; Netzer, I.; Zalmanoviz, S.; Friedman, E. Multigene panel testing in unselected Israeli breast cancer cases: Mutational spectrum and use of BRCA1/2 mutation prediction algorithms. Breast Cancer Res. Treat. 2019, 176, 165–170. [Google Scholar] [CrossRef]
- Alemar, B.; Herzog, J.; Netto, C.; Artigalás, O.; Schwartz, I.D.; Bittar, C.; Ashton-Prolla, P.; Weitzel, J.N. Prevalence of Hispanic BRCA1 and BRCA2 mutations among hereditary breast and ovarian cancer patients from Brazil reveals differences among Latin American populations. Cancer Genet. 2016, 209, 417–422. [Google Scholar] [CrossRef]
- Neamatzadeh, H.; Shiryazdi, S.; Kalantar, S. BRCA1 and BRCA2 mutations in Iranian breast cancer patients: A systematic review. J. Res. Med. Sci. 2015, 20, 284–293. [Google Scholar]
- Kumar, P.; Aggarwal, R. An overview of triple-Negative breast cancer. Arch. Gynecol. Obstet. 2016, 293, 247–269. [Google Scholar] [CrossRef]
- Wang, C.; Kar, S.; Lai, X.; Cai, W.; Arfuso, F.; Sethi, G.; Lobie, P.E.; Goh, B.C.; Lim, L.; Hartman, M.; et al. Triple negative breast cancer in Asia: An insider’s view. Cancer Treat. Rev. 2018, 62, 29–38. [Google Scholar] [CrossRef]
- Wong-Brown, M.W.; Meldrum, C.J.; Carpenter, J.E.; Clarke, C.L.; Narod, S.A.; Jakubowska, A.; Rudnicka, H.; Lubinski, J.; Scott, R.J. Prevalence of BRCA1 and BRCA2 germline mutations in patients with triple-negative breast cancer. Breast Cancer Res. Treat. 2015, 150, 71–80. [Google Scholar] [CrossRef]
- Baretta, Z.; Mocellin, S.; Goldin, E.; Olopade, O.I.; Huo, D. Effect of BRCA germline mutations on breast cancer prognosis. Medicine 2016, 95, 4975. [Google Scholar] [CrossRef]
| Stage | Percentage | Frequency |
|---|---|---|
| I | 13.5% | 34 |
| II | 29.7% | 75 |
| III | 38% | 95 |
| IV | 18.8% | 48 |
| Histology | Percentage | Frequency |
|---|---|---|
| Ductal | 83.7% | 211 |
| Lobular | 11.1% | 28 |
| Mixed | 5.2% | 13 |
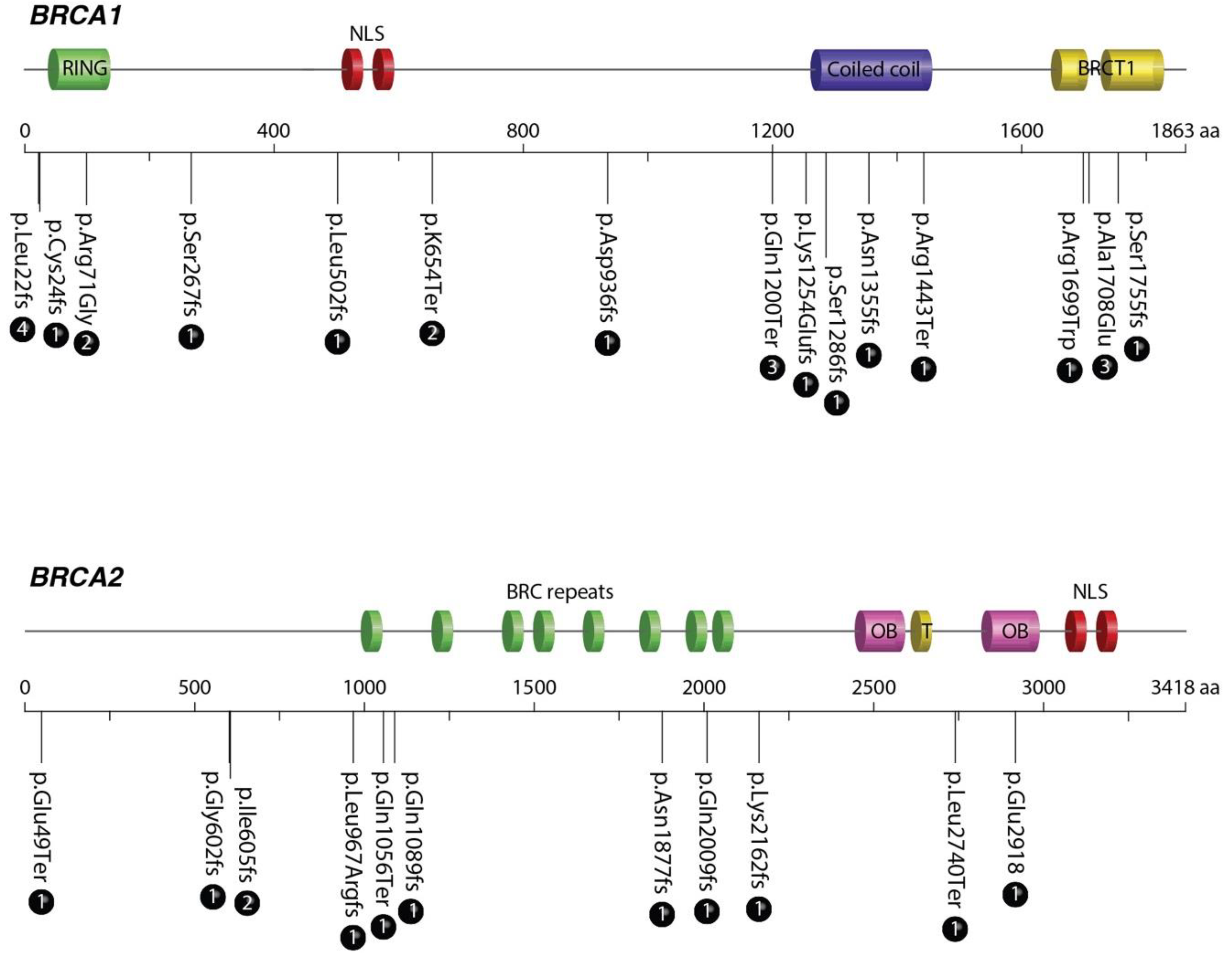
| Gene | Coding | Amino Acid Change | dbSNP | ClinVar/BIC Category | Frequency (Age of Diagnosis) | Ref. in LATAM Hispanic Population |
|---|---|---|---|---|---|---|
| BRCA1 | c.66_67delAG | p.Leu22fs | rs80357783 | Pathogenic/Class 5 | 4 (35, 36, 40, 44) | [14] [15] [16] |
| c.69_70insAG | p.Cys24fs | rs80357914 | Pathogenic/Class 5 | 1 (42) | [17] | |
| c.211A>G | p.Arg71Gly | rs80357382 | Pathogenic/Pending | 2 (49, 49) | [15] [17] [18] | |
| c.798_799delTT | p.Ser267fs | rs80357724 | Pathogenic/Class 5 | 1 (39) | Not reported | |
| c.1504_1508delTTAAA | p.Leu502fs | rs876659139 | Pathogenic/Class 5 | 1 (36) | Not reported | |
| c.1960A>T | p.K654Ter | rs80357355 | Pathogenic/Class 5 | 2 (32, 50) | [16] [19] | |
| c.2806_2809delGATA | p.Asp936fs | rs80357832 | Pathogenic/Class 5 | 1 (45) | Not reported | |
| c.3598C>T | p.Gln1200Ter | rs62625307 | Pathogenic/Class 5 | 3 (44, 45, 48) | [20] | |
| c.3759_3760delTA | p.Lys1254Glufs | rs80357520 | Pathogenic/Class 5 | 1 (37) | Not reported | |
| c.3858_3861delTGAG | p.Ser1286fs | rs80357842 | Pathogenic/Class 5 | 1 (34) | [21] | |
| c.4065_4068delTCAA | p.Asn1355fs | rs80357508 | Pathogenic/Class 5 | 1 (40) | Not reported | |
| c.4327C>T | p.Arg1443Ter | rs41293455 | Pathogenic/Class 5 | 1 (41) | [17] [19] | |
| c.5095C>T | p.Arg1699Trp | rs55770810 | Pathogenic/Pending | 1 (46) | [22] | |
| c.5123C>A | p.Ala1708Glu | rs28897696 | Pathogenic/Pending | 3 (40, 41, 44) | [23] [24] [25] [26] | |
| c.5263_5264insC | p.Ser1755fs | rs80357906 | Pathogenic/Class 5 | 1 (50) | [27] | |
| IVS5+1G>A c.212+1G>A | Splicing mutation | rs80358042 | Pathogenic/Pending | 1 (35) | [28] | |
| BRCA2 | c.145G>T | p.Glu49Ter | rs80358435 | Pathogenic/Class 5 | 1 (31) | [28] [29] |
| c.1806insA | p.Gly602fs | rs80359307 | Pathogenic/Class 5 | 1 (37) | Not reported | |
| c.1813delA | p.Ile605fs | rs80359306 | Pathogenic/Class 5 | 2 (33, 46) | Not reported | |
| c.2899_2900delCT | p.Leu967Argfs | rs80359361 | Pathogenic/Class 5 | 1 (35) | Not reported | |
| c.3166C>T | p.Gln1056Ter | rs79728106 | Pathogenic/Class 5 | 1 (37) | [30] | |
| c.3492insT | p.Gln1089fs | rs80359380 | Pathogenic/Class 5 | 1 (40) | [28] [31] | |
| c.5631delC | p.Asn1877fs | rs397507357 | Pathogenic/Class 5 | 1 (41) | [20] | |
| c.6244_6244delG | Pathogenic/Class 5 | 1 (47) | Not reported | |||
| c.6024_6025insG | p.Gln2009fs | rs80359554 | Pathogenic /Class 5 | 1 (40) | [32] [33] | |
| c.6486_6489delACAA | p.Lys2162fs | rs80359598 | Pathogenic/Class 5 | 1 (31) | [28] | |
| c.8219T>G | p.Leu2740Ter | rs80359070 | Pathogenic/Class 5 | 1 (33) | Not reported | |
| c.8754G>A | Splicing mutation p.Glu2918= | rs80359803 | Pathogenic, Likely pathogenic/Pending | 1 (38) | Not reported |
| Molecular Subtype | Mutations | |||||||
|---|---|---|---|---|---|---|---|---|
| BRCA1 | BRCA2 | TOTAL | ||||||
| Percentage | Frequency | Percentage | Frequency | Percentage | Frequency | Percentage | Frequency | |
| HR+, Her2− | 48% | 121 | 6.6% | 8 | 4.1% | 5 | 10.7% | 13 |
| Her2+ | 24.7% | 62 | 8% | 5 | 12.9% | 8 | 20.9% | 13 |
| Triple Negative | 23.8% | 60 | 20% | 12 | – | – | 20% | 12 |
| Unknown | 3.5% | 9 | – | – | – | – | – | – |
| Mutation Type | BRCA1 | BRCA2 |
|---|---|---|
| Frameshift | 8 | 8 |
| Amino acid substitution | 6 | 3 |
| Splicing mutation | 2 | 1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Millan Catalan, O.; Campos-Parra, A.D.; Vázquez-Romo, R.; Cantú de León, D.; Jacobo-Herrera, N.; Morales-González, F.; López-Camarillo, C.; Rodríguez-Dorantes, M.; López-Urrutia, E.; Pérez-Plasencia, C. A Multi-Center Study of BRCA1 and BRCA2 Germline Mutations in Mexican-Mestizo Breast Cancer Families Reveals Mutations Unreported in Latin American Population. Cancers 2019, 11, 1246. https://doi.org/10.3390/cancers11091246
Millan Catalan O, Campos-Parra AD, Vázquez-Romo R, Cantú de León D, Jacobo-Herrera N, Morales-González F, López-Camarillo C, Rodríguez-Dorantes M, López-Urrutia E, Pérez-Plasencia C. A Multi-Center Study of BRCA1 and BRCA2 Germline Mutations in Mexican-Mestizo Breast Cancer Families Reveals Mutations Unreported in Latin American Population. Cancers. 2019; 11(9):1246. https://doi.org/10.3390/cancers11091246
Chicago/Turabian StyleMillan Catalan, Oliver, Alma D. Campos-Parra, Rafael Vázquez-Romo, David Cantú de León, Nadia Jacobo-Herrera, Fermín Morales-González, César López-Camarillo, Mauricio Rodríguez-Dorantes, Eduardo López-Urrutia, and Carlos Pérez-Plasencia. 2019. "A Multi-Center Study of BRCA1 and BRCA2 Germline Mutations in Mexican-Mestizo Breast Cancer Families Reveals Mutations Unreported in Latin American Population" Cancers 11, no. 9: 1246. https://doi.org/10.3390/cancers11091246
APA StyleMillan Catalan, O., Campos-Parra, A. D., Vázquez-Romo, R., Cantú de León, D., Jacobo-Herrera, N., Morales-González, F., López-Camarillo, C., Rodríguez-Dorantes, M., López-Urrutia, E., & Pérez-Plasencia, C. (2019). A Multi-Center Study of BRCA1 and BRCA2 Germline Mutations in Mexican-Mestizo Breast Cancer Families Reveals Mutations Unreported in Latin American Population. Cancers, 11(9), 1246. https://doi.org/10.3390/cancers11091246









