Expression of the Major and Pro-Oncogenic H3K9 Lysine Methyltransferase SETDB1 in Non-Small Cell Lung Cancer
Abstract
1. Introduction
2. Results
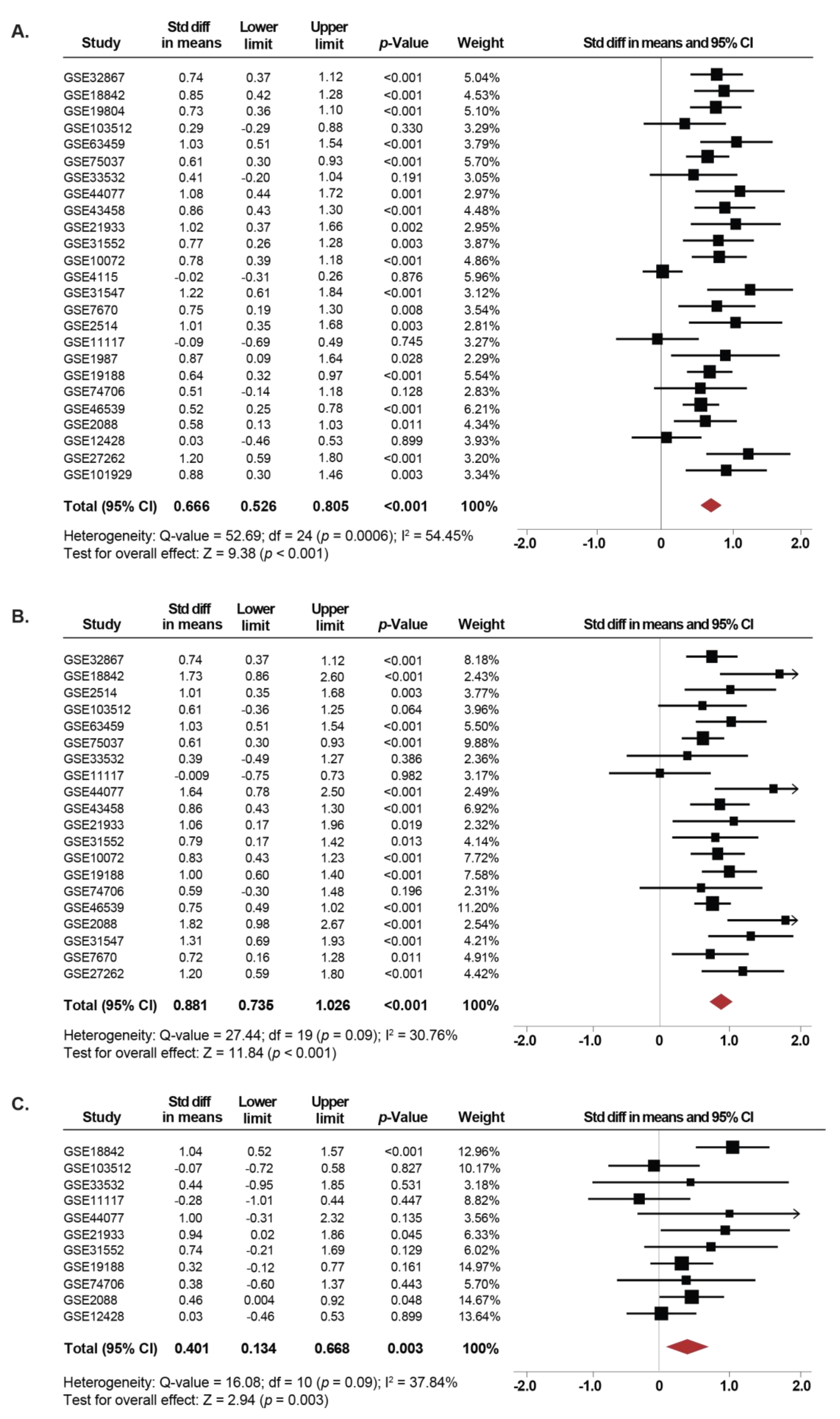
2.1. Association Between SETDB1 Expression and NSCLC
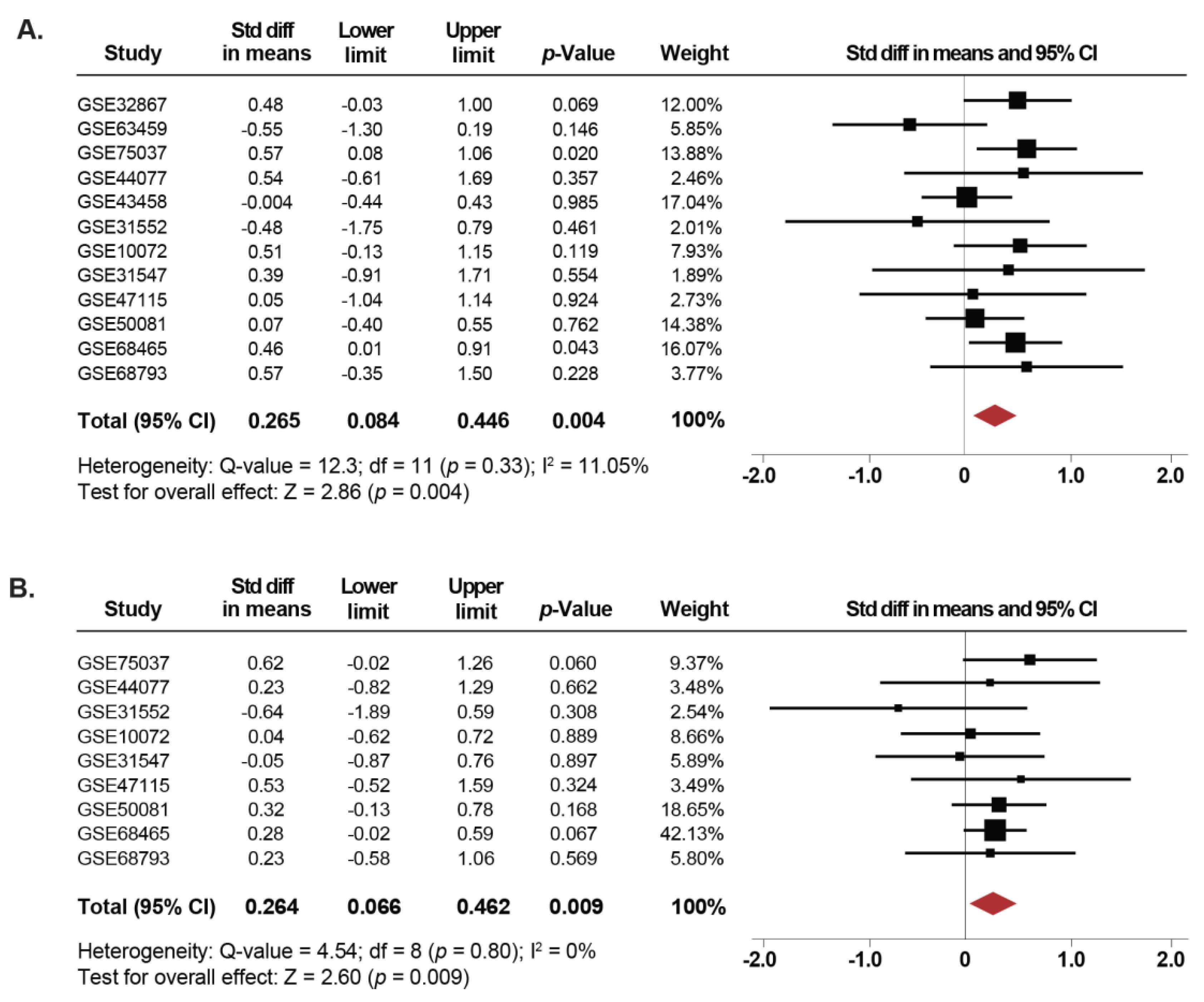
2.2. Expression of SETDB1 Is Increased in Current and Former Smokers Compared to NSCLC Non-Smoker Patients
2.3. Association Between SETDB1 Levels and Pathological Characteristics of the NSCLC Samples
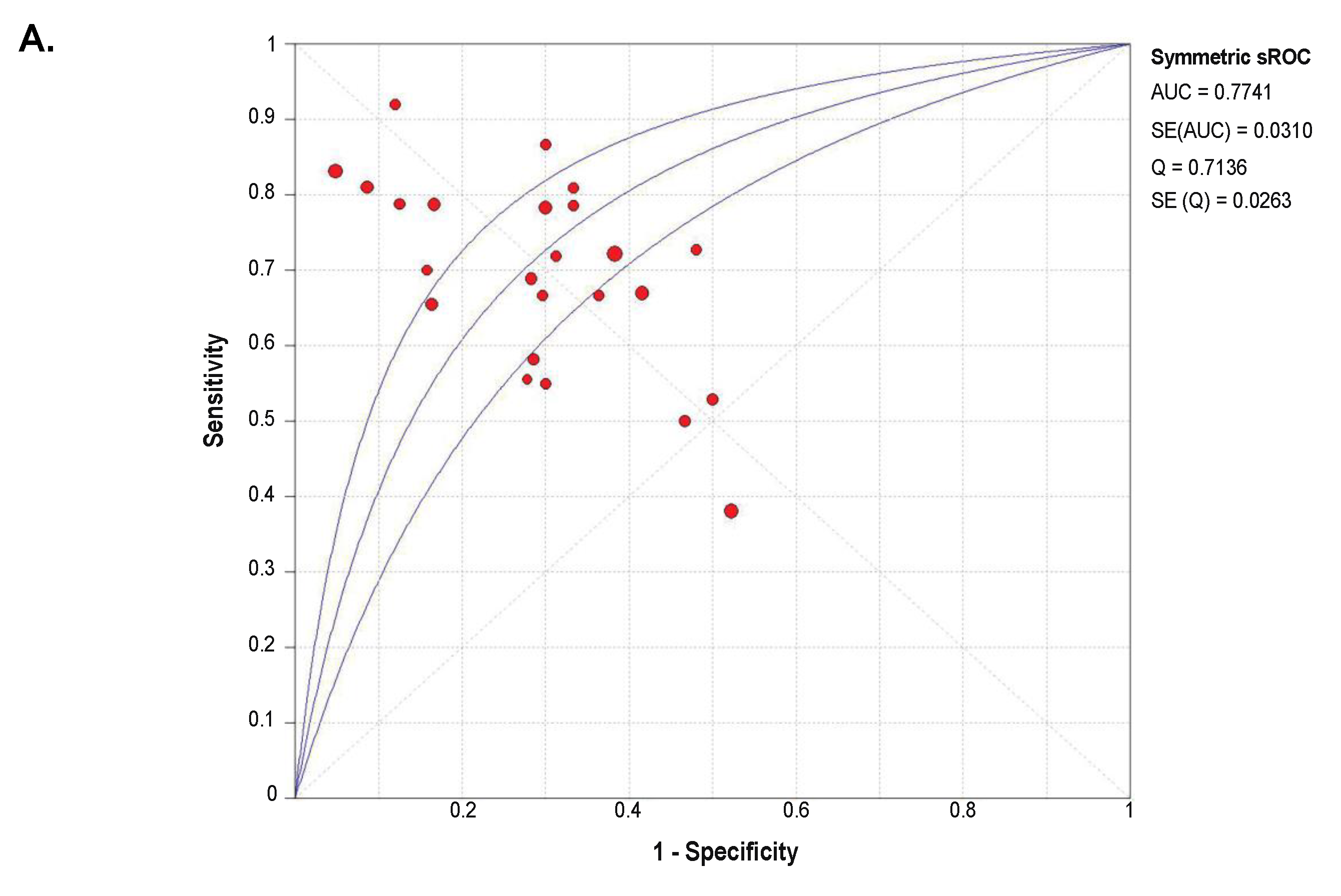
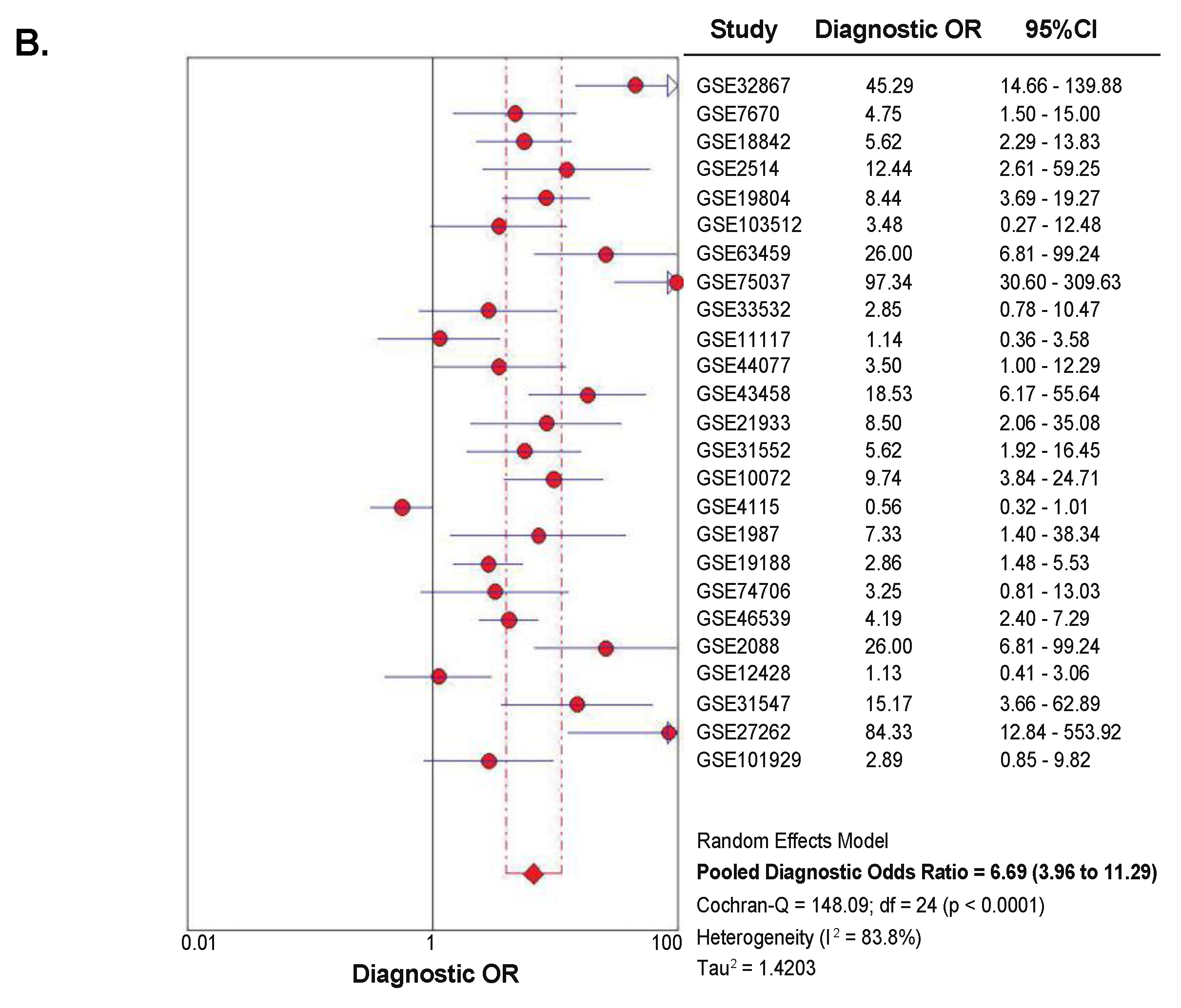
2.4. Diagnostic Value of SETDB1 in NSCLC, Based on Meta-Analysis
2.5. Literature Review on SETDB1 in NSCLC
3. Discussion
4. Materials and Methods
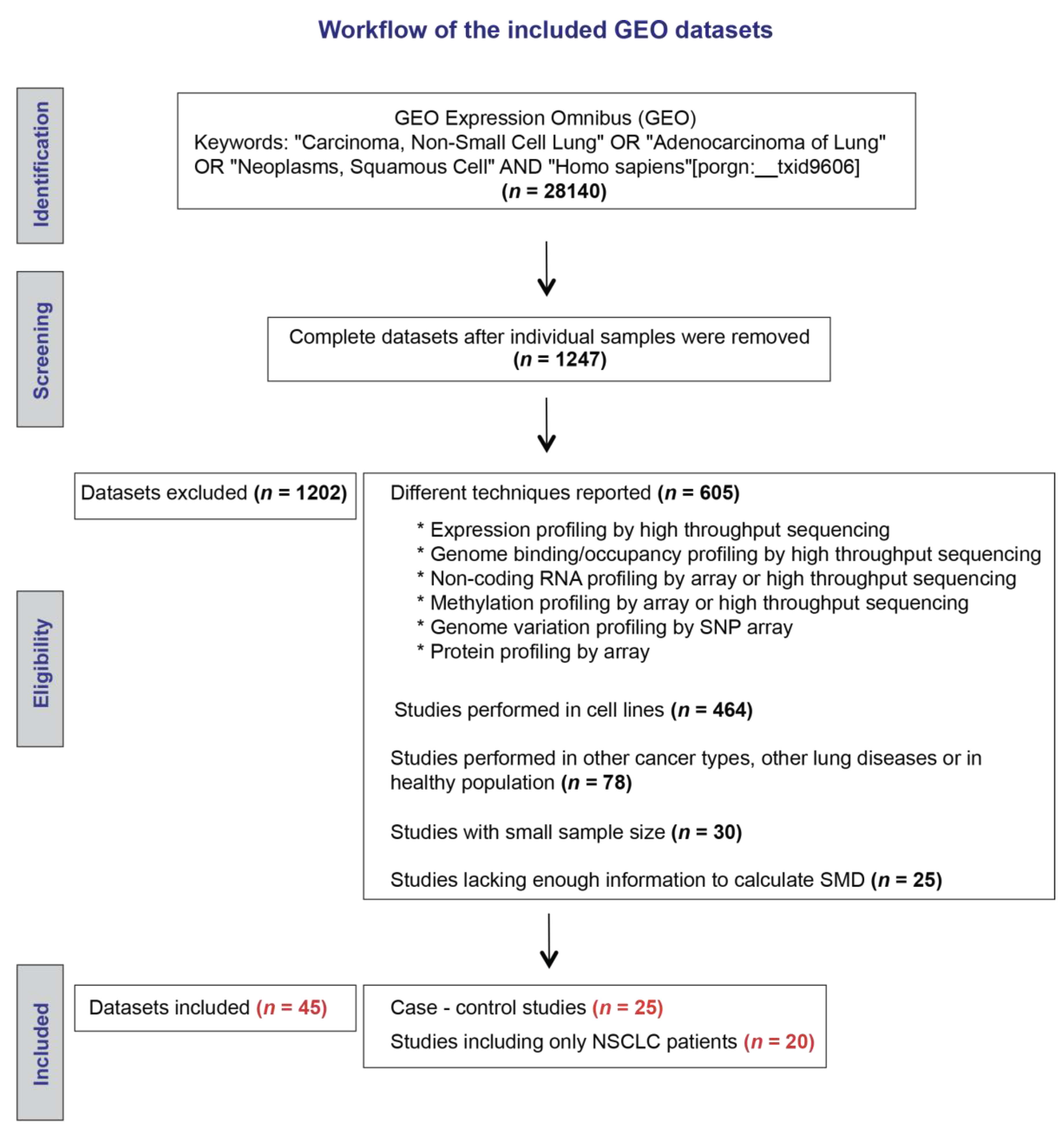
4.1. Search Strategy for Microarray Databases in the Gene Expression Omnibus (GEO) Repository
4.2. Data Extraction
4.3. Statistical Analysis
4.4. Search Strategy for Peer-Reviewed Journals
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Barta, J.A.; Powell, C.A.; Wisnivesky, J.P. Global Epidemiology of Lung Cancer. Ann. Glob. Heal. 2019, 85. [Google Scholar] [CrossRef] [PubMed]
- Ellis, P.M.; Vandermeer, R. Delays in the diagnosis of lung cancer. J. Thorac. Dis. 2011, 3, 183. [Google Scholar] [PubMed]
- Zhao, H.; Wang, J.; Kong, X.; Li, E.; Liu, Y.; Du, X.; Kang, Z.; Tang, Y.; Kuang, Y.; Yang, Z.; et al. CD47 Promotes Tumor Invasion and Metastasis in Non-small Cell Lung Cancer. Sci. Rep. 2016, 6, 29719. [Google Scholar] [CrossRef] [PubMed]
- Langevin, S.M.; Kratzke, R.A.; Kelsey, K.T. Epigenetics of lung cancer. Transl. Res. 2015, 165, 74–90. [Google Scholar] [CrossRef] [PubMed]
- Cohen, I.; Poręba, E.; Kamieniarz, K.; Schneider, R. Histone modifiers in cancer: Friends or foes? Genes Cancer 2011, 2, 631–647. [Google Scholar] [CrossRef] [PubMed]
- Mozzetta, C.; Boyarchuk, E.; Pontis, J.; Ait-Si-Ali, S. Sound of silence: The properties and functions of repressive Lys methyltransferases. Nat. Rev. Mol. Cell Biol. 2015, 16, 499–513. [Google Scholar] [CrossRef]
- Greer, E.L.; Shi, Y. Histone methylation: A dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 2012, 13, 343–357. [Google Scholar] [CrossRef]
- Ellinger, J.; Kahl, P.; von der Gathen, J.; Rogenhofer, S.; Heukamp, L.C.; Gütgemann, I.; Walter, B.; Hofstädter, F.; Büttner, R.; Müller, S.C.; et al. Global levels of histone modifications predict prostate cancer recurrence. Prostate 2010, 70, 61–69. [Google Scholar] [CrossRef]
- Seligson, D.B.; Horvath, S.; McBrian, M.A.; Mah, V.; Yu, H.; Tze, S.; Wang, Q.; Chia, D.; Goodglick, L.; Kurdistani, S.K.; et al. Global levels of histone modifications predict prognosis in different cancers. Am. J. Pathol. 2009, 174, 1619–1628. [Google Scholar] [CrossRef]
- Song, J.S.; Kim, Y.S.; Kim, D.K.; Park, S.I.; Jang, S.J. Global histone modification pattern associated with recurrence and disease-free survival in non-small cell lung cancer patients. Pathol. Int. 2012, 62, 182–190. [Google Scholar] [CrossRef]
- Van Rechem, C.; Whetstine, J.R. Examining the impact of gene variants on histone lysine methylation. Biochim. Biophys. Acta 2014, 1839, 1463–1476. [Google Scholar] [CrossRef] [PubMed]
- Dodge, J.E.; Kang, Y.-K.; Beppu, H.; Lei, H.; Li, E. Histone H3-K9 methyltransferase ESET is essential for early development. Mol. Cell. Biol. 2004, 24, 2478–2486. [Google Scholar] [CrossRef] [PubMed]
- Lawson, K.A.; Teteak, C.J.; Gao, J.; Li, N.; Hacquebord, J.; Ghatan, A.; Zielinska-Kwiatkowska, A.; Song, G.; Chansky, H.A.; Yang, L.; et al. ESET histone methyltransferase regulates osteoblastic differentiation of mesenchymal stem cells during postnatal bone development. FEBS Lett. 2013, 587, 3961–3967. [Google Scholar] [CrossRef] [PubMed]
- Beyer, S.; Pontis, J.; Schirwis, E.; Battisti, V.; Rudolf, A.; Le Grand, F.; Ait-Si-Ali, S. Canonical Wnt signalling regulates nuclear export of Setdb1 during skeletal muscle terminal differentiation. Cell Discov. 2016, 2, 16037. [Google Scholar] [CrossRef] [PubMed]
- Yeap, L.-S.; Hayashi, K.; Surani, M.A. ERG-associated protein with SET domain (ESET)-Oct4 interaction regulates pluripotency and represses the trophectoderm lineage. Epigenet. Chromatin 2009, 2, 12. [Google Scholar] [CrossRef]
- Bilodeau, S.; Kagey, M.H.; Frampton, G.M.; Rahl, P.B.; Young, R.A. SetDB1 contributes to repression of genes encoding developmental regulators and maintenance of ES cell state. Genes Dev. 2009, 23, 2484–2489. [Google Scholar] [CrossRef]
- Lohmann, F.; Loureiro, J.; Su, H.; Fang, Q.; Lei, H.; Lewis, T.; Yang, Y.; Labow, M.; Li, E.; Chen, T.; et al. KMT1E mediated H3K9 methylation is required for the maintenance of embryonic stem cells by repressing trophectoderm differentiation. Stem. Cells 2010, 28, 201–212. [Google Scholar] [CrossRef]
- Reya, T.; Morrison, S.J.; Clarke, M.F.; Weissman, I.L. Stem cells, cancer, and cancer stem cells. Nature 2001, 414, 105–111. [Google Scholar] [CrossRef]
- Ceol, C.J.; Houvras, Y.; Jane-Valbuena, J.; Bilodeau, S.; Orlando, D.A.; Battisti, V.; Fritsch, L.; Lin, W.M.; Hollmann, T.J.; Ferré, F.; et al. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature 2011, 471, 513–517. [Google Scholar] [CrossRef]
- Watanabe, H.; Soejima, K.; Yasuda, H.; Kawada, I.; Nakachi, I.; Yoda, S.; Naoki, K.; Ishizaka, A. Deregulation of histone lysine methyltransferases contributes to oncogenic transformation of human bronchoepithelial cells. Cancer Cell Int. 2008, 8, 15. [Google Scholar] [CrossRef]
- Li, H.; Rauch, T.; Chen, Z.-X.; Szabó, P.E.; Riggs, A.D.; Pfeifer, G.P. The histone methyltransferase SETDB1 and the DNA methyltransferase DNMT3A interact directly and localize to promoters silenced in cancer cells. J. Biol. Chem. 2006, 281, 19489–19500. [Google Scholar] [CrossRef] [PubMed]
- Kokura, K.; Sun, L.; Bedford, M.T.; Fang, J. Methyl-H3K9-binding protein MPP8 mediates E-cadherin gene silencing and promotes tumour cell motility and invasion. EMBO J. 2010, 29, 3673–3687. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Paredes, M.; de Paz, A.M.; Simó-Riudalbas, L.; Sayols, S.; Moutinho, C.; Moran, S.; Villanueva, A.; Vázquez-Cedeira, M.; Lazo, P.A.; Carneiro, F.; et al. Gene amplification of the histone methyltransferase SETDB1 contributes to human lung tumorigenesis. Oncogene 2014, 33, 2807–2813. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.-Y.; Ding, L.-W.; Xiao, J.-F.; Chien, W.; Lim, S.-L.; Hattori, N.; Goodglick, L.; Chia, D.; Mah, V.; Alavi, M.; et al. SETDB1 accelerates tumourigenesis by regulating the WNT signalling pathway. J. Pathol. 2015, 235, 559–570. [Google Scholar] [CrossRef] [PubMed]
- Selamat, S.A.; Chung, B.S.; Girard, L.; Zhang, W.; Zhang, Y.; Campan, M.; Siegmund, K.D.; Koss, M.N.; Hagen, J.A.; Lam, W.L.; et al. Genome-scale analysis of DNA methylation in lung adenocarcinoma and integration with mRNA expression. Genome Res. 2012, 22, 1197–1211. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Palencia, A.; Gomez-Morales, M.; Gomez-Capilla, J.A.; Pedraza, V.; Boyero, L.; Rosell, R.; Fárez-Vidal, M.E. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int. J. Cancer 2011, 129, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.-P.; Tsai, M.-H.; Lee, J.-M.; Hsu, C.-P.; Chen, P.-C.; Lin, C.-W.; Shih, J.Y.; Yang, P.C.; Hsiao, C.K.; Lai, L.C.; et al. Identification of a novel biomarker, SEMA5A, for non-small cell lung carcinoma in nonsmoking women. Cancer Epidemiol. Biomark. Prev. 2010, 19, 2590–2597. [Google Scholar] [CrossRef] [PubMed]
- Brouwer-Visser, J.; Cheng, W.-Y.; Bauer-Mehren, A.; Maisel, D.; Lechner, K.; Andersson, E.; Dudley, J.T.; Milletti, F. Regulatory T-cell Genes Drive Altered Immune Microenvironment in Adult Solid Cancers and Allow for Immune Contextual Patient Subtyping. Cancer Epidemiol. Biomark. Prev. 2018, 27, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Robles, A.I.; Arai, E.; Mathé, E.A.; Okayama, H.; Schetter, A.J.; Brown, D.; Petersen, D.; Bowman, E.D.; Noro, R.; Welsh, J.A.; et al. An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma Based on mRNA, microRNA, and DNA Methylation Biomarkers. J. Thorac. Oncol. 2015, 10, 1037–1048. [Google Scholar] [CrossRef]
- Pickup, M.E.; Dixon, J.S.; Lowe, J.R.; Wright, V. Serum histidine in rheumatoid arthritis: Changes induced by antirheumatic drug therapy. J. Rheumatol. 1980, 7, 71–76. [Google Scholar]
- Meister, M.; Belousov, A.; Xu, E.; Schnabel, P.; Warth, A.; Hoffmann, H.; Dienemann, H.; Riedlinger, J.; Bodenmueller, H.; Zolg, W.; et al. Intra-tumor Heterogeneity of Gene Expression Profiles in Early Stage Non-Small Cell Lung Cancer. J. Bioinf. Res. Stud. 2014, 1, 1. [Google Scholar]
- Kadara, H.; Fujimoto, J.; Yoo, S.-Y.; Maki, Y.; Gower, A.C.; Kabbout, M.; Garcia, M.M.; Chow, C.W.; Chu, Z.; Mendoza, G.; et al. Transcriptomic architecture of the adjacent airway field cancerization in non-small cell lung cancer. J. Natl. Cancer Inst. 2014, 106, dju004. [Google Scholar] [CrossRef] [PubMed]
- Kabbout, M.; Garcia, M.M.; Fujimoto, J.; Liu, D.D.; Woods, D.; Chow, C.-W.; Mendoza, G.; Momin, A.A.; James, B.P.; Solis, L.; et al. ETS2 mediated tumor suppressive function and MET oncogene inhibition in human non-small cell lung cancer. Clin. Cancer Res. 2013, 19, 3383–3395. [Google Scholar] [CrossRef] [PubMed]
- Lo, F.-Y.; Chang, J.-W.; Chang, I.-S.; Chen, Y.-J.; Hsu, H.-S.; Huang, S.-F.K.; Tsai, F.Y.; Jiang, S.S.; Kanteti, R.; Nandi, S.; et al. The database of chromosome imbalance regions and genes resided in lung cancer from Asian and Caucasian identified by array-comparative genomic hybridization. BMC Cancer 2012, 12, 235. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Marquardt, G.; Mullapudi, N.; Wang, T.; Han, W.; Shi, M.; Keller, S.; Zhu, C.; Locker, J.; Spivack, S.D. Lung cancer transcriptomes refined with laser capture microdissection. Am. J. Pathol. 2014, 184, 2868–2884. [Google Scholar] [CrossRef] [PubMed]
- Landi, M.T.; Dracheva, T.; Rotunno, M.; Figueroa, J.D.; Liu, H.; Dasgupta, A.; Mann, F.E.; Fukuoka, J.; Hames, M.; Bergen, A.W.; et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS ONE 2008, 3, e1651. [Google Scholar] [CrossRef] [PubMed]
- Spira, A.; Beane, J.E.; Shah, V.; Steiling, K.; Liu, G.; Schembri, F.; Gilman, S.; Dumas, Y.M.; Calner, P.; Sebastiani, P.; et al. Airway epithelial gene expression in the diagnostic evaluation of smokers with suspect lung cancer. Nat. Med. 2007, 13, 361–366. [Google Scholar] [CrossRef]
- Su, L.-J.; Chang, C.-W.; Wu, Y.-C.; Chen, K.-C.; Lin, C.-J.; Liang, S.-C.; Lin, C.H.; Whang-Peng, J.; Hsu, S.L.; Chen, C.H.; et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genom. 2007, 8, 140. [Google Scholar] [CrossRef]
- Stearman, R.S.; Dwyer-Nield, L.; Zerbe, L.; Blaine, S.A.; Chan, Z.; Bunn, P.A.; Johnson, G.L.; Hirsch, F.R.; Merrick, D.T.; Franklin, W.A.; et al. Analysis of orthologous gene expression between human pulmonary adenocarcinoma and a carcinogen-induced murine model. Am. J. Pathol. 2005, 167, 1763–1775. [Google Scholar] [CrossRef]
- Baty, F.; Facompré, M.; Kaiser, S.; Schumacher, M.; Pless, M.; Bubendorf, L.; Savic, S.; Marrer, E.; Budach, W.; Buess, M.; et al. Gene profiling of clinical routine biopsies and prediction of survival in non-small cell lung cancer. Am. J. Respir. Crit. Care Med. 2010, 181, 181–188. [Google Scholar] [CrossRef]
- Dehan, E.; Ben-Dor, A.; Liao, W.; Lipson, D.; Frimer, H.; Rienstein, S.; Simansky, D.; Krupsky, M.; Yaron, P.; Friedman, E.; et al. Chromosomal aberrations and gene expression profiles in non-small cell lung cancer. Lung Cancer 2007, 56, 175–184. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Aerts, J.; den Hamer, B.; van Ijcken, W.; den Bakker, M.; Riegman, P.; van der Leest, C.; van der Spek, P.; Foekens, J.A.; Hoogsteden, H.C.; et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS ONE 2010, 5, e10312. [Google Scholar] [CrossRef] [PubMed]
- Marwitz, S.; Depner, S.; Dvornikov, D.; Merkle, R.; Szczygieł, M.; Müller-Decker, K.; Lucarelli, P.; Wäsch, M.; Mairbäurl, H.; Rabe, K.F.; et al. Downregulation of the TGFβ Pseudoreceptor BAMBI in Non-Small Cell Lung Cancer Enhances TGFβ Signaling and Invasion. Cancer Res. 2016, 76, 3785–3801. [Google Scholar] [CrossRef] [PubMed]
- Chang, I.-S.; Jiang, S.S.; Yang, J.C.-H.; Su, W.-C.; Chien, L.-H.; Hsiao, C.-F.; Lee, J.H.; Chen, C.Y.; Chen, C.H.; Chang, G.C.; et al. Genetic Modifiers of Progression-Free Survival in Never-Smoking Lung Adenocarcinoma Patients Treated with First-Line Tyrosine Kinase Inhibitors. Am. J. Respir. Crit. Care Med. 2017, 195, 663–673. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, T.; Hiramatsu, M.; Isagawa, T.; Ninomiya, H.; Inamura, K.; Ishikawa, S.; Ushijima, M.; Matsuura, M.; Jones, M.H.; Shimane, M.; et al. ASCL1-coexpression profiling but not single gene expression profiling defines lung adenocarcinomas of neuroendocrine nature with poor prognosis. Lung Cancer 2012, 75, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Boelens, M.C.; van den Berg, A.; Fehrmann, R.S.N.; Geerlings, M.; de Jong, W.K.; te Meerman, G.J.; Sietsma, H.; Timens, W.; Postma, D.S.; Groen, H.J. Current smoking-specific gene expression signature in normal bronchial epithelium is enhanced in squamous cell lung cancer. J. Pathol. 2009, 218, 182–191. [Google Scholar] [CrossRef]
- Wei, T.-Y.W.; Juan, C.-C.; Hisa, J.-Y.; Su, L.-J.; Lee, Y.-C.G.; Chou, H.-Y.; Chen, J.M.; Wu, Y.C.; Chiu, S.C.; Hsu, C.P.; et al. Protein arginine methyltransferase 5 is a potential oncoprotein that upregulates G1 cyclins/cyclin-dependent kinases and the phosphoinositide 3-kinase/AKT signaling cascade. Cancer Sci. 2012, 103, 1640–1650. [Google Scholar] [CrossRef]
- Mitchell, K.A.; Zingone, A.; Toulabi, L.; Boeckelman, J.; Ryan, B.M. Comparative Transcriptome Profiling Reveals Coding and Noncoding RNA Differences in NSCLC from African Americans and European Americans. Clin. Cancer Res. 2017, 23, 7412–7425. [Google Scholar] [CrossRef]
- Inoue, Y.; Matsuura, S.; Kurabe, N.; Kahyo, T.; Mori, H.; Kawase, A.; Karayama, M.; Inui, N.; Funai, K.; Shinmura, K.; et al. Clinicopathological and Survival Analysis of Japanese Patients with Resected Non-Small-Cell Lung Cancer Harboring NKX2-1, SETDB1, MET, HER2, SOX2, FGFR1, or PIK3CA Gene Amplification. J. Thorac. Oncol. 2015, 10, 1590–1600. [Google Scholar] [CrossRef]
- Wang, G.; Long, J.; Gao, Y.; Zhang, W.; Han, F.; Xu, C.; Sun, L.; Yang, S.C.; Lan, J.; Hou, Z.; et al. SETDB1-mediated methylation of Akt promotes its K63-linked ubiquitination and activation leading to tumorigenesis. Nat. Cell Biol. 2019, 21, 214–225. [Google Scholar] [CrossRef]
- Lafuente-Sanchis, A.; Zúñiga, Á.; Galbis, J.M.; Cremades, A.; Estors, M.; Martínez-Hernández, N.J.; Carretero, J. Prognostic value of ERCC1, RRM1, BRCA1 and SETDB1 in early stage of non-small cell lung cancer. Clin. Transl. Oncol. 2016, 18, 798–804. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.-C.; Lu, J.-W.; Yang, J.-Y.; Lin, I.-H.; Ou, D.-L.; Lin, Y.-H.; Chou, K.H.; Huang, W.F.; Wang, W.P.; Huang, Y.L.; et al. H3K9 histone methyltransferase, KMT1E/SETDB1, cooperates with the SMAD2/3 pathway to suppress lung cancer metastasis. Cancer Res. 2014, 74, 7333–7343. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-K.; Kim, K.-C. DZNep, inhibitor of S-adenosylhomocysteine hydrolase, down-regulates expression of SETDB1 H3K9me3 HMTase in human lung cancer cells. Biochem. Biophys. Res. Commun. 2013, 438, 647–652. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Wang, J.; Wang, J.; Wang, H.; Gu, X.; Tang, L.; Feng, X. A regulatory circuitry comprising TP53, miR-29 family, and SETDB1 in non-small cell lung cancer. Biosci. Rep. 2018, 38, BSR20180678. [Google Scholar] [CrossRef] [PubMed]
- Na, H.-H.; Kim, K.-C. SETDB1-mediated FosB regulation via ERK2 is associated with an increase in cell invasiveness during anticancer drug treatment of A549 human lung cancer cells. Biochem. Biophys. Res. Commun. 2018, 495, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Al Emran, A.; Marzese, D.M.; Menon, D.R.; Stark, M.S.; Torrano, J.; Hammerlindl, H.; Zhang, G.; Brafford, P.; Salomon, M.P.; Nelson, N.; et al. Distinct histone modifications denote early stress-induced drug tolerance in cancer. Oncotarget 2018, 9, 8206–8222. [Google Scholar] [CrossRef] [PubMed]
- Noh, H.-J.; Kim, K.-A.; Kim, K.-C. p53 down-regulates SETDB1 gene expression during paclitaxel induced-cell death. Biochem. Biophys. Res. Commun. 2014, 446, 43–48. [Google Scholar] [CrossRef]
- Kruse, J.-P.; Gu, W. Modes of p53 Regulation. Cell 2009, 137, 609–622. [Google Scholar] [CrossRef]
- Chen, X.; El Gazzar, M.; Yoza, B.K.; McCall, C.E. The NF-κB Factor RelB and Histone H3 Lysine Methyltransferase G9a Directly Interact to Generate Epigenetic Silencing in Endotoxin Tolerance. J. Biol. Chem. 2009, 284, 27857–27865. [Google Scholar] [CrossRef]
- Fang, L.; Teng, H.; Wang, Y.; Liao, G.; Weng, L.; Li, Y.; Wang, X.; Jin, J.; Jiao, C.; Chen, L.; et al. SET1A-Mediated Mono-Methylation at K342 Regulates YAP Activation by Blocking Its Nuclear Export and Promotes Tumorigenesis. Cancer Cell 2018, 34, 103–118. [Google Scholar] [CrossRef]
- Kim, E.; Kim, M.; Woo, D.-H.; Shin, Y.; Shin, J.; Chang, N.; Oh, Y.T.; Kim, H.; Rheey, J.; Nakano, I.; et al. Phosphorylation of EZH2 Activates STAT3 Signaling via STAT3 Methylation and Promotes Tumorigenicity of Glioblastoma Stem-like Cells. Cancer Cell 2013, 23, 839–852. [Google Scholar] [CrossRef] [PubMed]
- Fei, Q.; Shang, K.; Zhang, J.; Chuai, S.; Kong, D.; Zhou, T.; Fu, S.; Liang, Y.; Li, C.; Chen, Z.; et al. Histone methyltransferase SETDB1 regulates liver cancer cell growth through methylation of p53. Nat. Commun. 2015, 6, 8651. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.-M.; Wei, L.; Law, C.-T.; Ho, D.W.-H.; Tsang, F.H.-C.; Au, S.L.-K.; Sze, K.M.; Lee, J.M.; Wong, C.C.; Ng, I.O.; et al. Up-regulation of histone methyltransferase SETDB1 by multiple mechanisms in hepatocellular carcinoma promotes cancer metastasis. Hepatology 2016, 63, 474–487. [Google Scholar] [CrossRef] [PubMed]
- Robbez-Masson, L.; Tie, C.H.C.; Rowe, H.M. Cancer cells, on your histone marks, get SETDB1, silence retrotransposons, and go! J. Cell Biol. 2017, 216, 3429–3431. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Wei, M.; Ren, S.-C.; Chen, R.; Xu, W.-D.; Wang, F.-B.; Lu, J.; Shen, J.; Yu, Y.W.; Hou, J.G.; et al. Histone methyltransferase SETDB1 is required for prostate cancer cell proliferation, migration and invasion. Asian. J. Androl. 2014, 16, 319–324. [Google Scholar] [CrossRef]
- Zhang, H.; Cai, K.; Wang, J.; Wang, X.; Cheng, K.; Shi, F.; Jiang, L.; Zhang, Y.; Dou, J. MiR-7, inhibited indirectly by lincRNA HOTAIR, directly inhibits SETDB1 and reverses the EMT of breast cancer stem cells by downregulating the STAT3 pathway. Stem. Cells 2014, 32, 2858–2868. [Google Scholar] [CrossRef]
- Vaughan, A.E.; Halbert, C.L.; Wootton, S.K.; Miller, A.D. Lung cancer in mice induced by the jaagsiekte sheep retrovirus envelope protein is not maintained by rare cancer stem cells, but tumorigenicity does correlate with Wnt pathway activation. Mol. Cancer Res. 2012, 10, 86–95. [Google Scholar] [CrossRef]
- Stewart, D.J. Wnt Signaling Pathway in Non-Small Cell Lung Cancer. JNCI J. Natl. Cancer Inst. 2014, 106, djt356. [Google Scholar] [CrossRef]
- Shang, S.; Hua, F.; Hu, Z.-W. The regulation of β-catenin activity and function in cancer: Therapeutic opportunities. Oncotarget 2017, 8, 33972–33989. [Google Scholar] [CrossRef]
- Stepanenko, A.A.; Vassetzky, Y.S.; Kavsan, V.M. Antagonistic functional duality of cancer genes. Gene 2013, 529, 199–207. [Google Scholar] [CrossRef]
| GEO Dataset | Country | Year | ADC | SCC | NSCLC * | Controls | Sample Type in Patients | Sample Type in Controls | Platform | SETDB1 ID | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|
| GSE32867 | Canada | 2012 | 58 | 58 | Cancer tissue | Adjacent non-cancerous tissues | Illumina Human WG-6 v3.0 Expression BeadChips | ILMN_1718207 | [25] | ||
| GSE18842 | Spain | 2010 | 14 | 31 | 46 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133 Plus 2.0 Array | 203155_at | [26] | |
| GSE19804 | Taiwan | 2010 | 60 | 60 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix GeneChip Human Genome U133 plus array | 203155_at | [27] | ||
| GSE103512 | Germany | 2017 | 30 | 25 | 14 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix HT-U133plus-2-PM microarrays | 203155_PM_at | [28] | |
| GSE63459 | USA/Japan | 2015 | 33 | 32 | Cancer tissue | Adjacent non-cancerous tissues | Illumina HumanRef-8 v3 Expression Beadchip arrays | ILMN_1718207 | [29] | ||
| GSE75037 | USA | 2016 | 83 | 83 | Cancer tissue | Adjacent non-cancerous tissues | Illumina BeadChip array HumanWG-6 V3 | ILMN_1718207 | [30] | ||
| GSE33532 | Germany | 2014 | 10 | 4 | 6 | 20 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix U133 Plus 2.0 arrays | 203155_at | [31] |
| GSE44077 | USA | 2013 | 14 | 5 | 2 | 21 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Gene 1.0 ST Array [transcript (gene) version] | 7905258 | [32] |
| GSE43458 | USA | 2013 | 80 | 30 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Gene 1.0 ST Array [transcript (gene) version] | 7905258 | [33] | ||
| GSE21933 | USA/Taiwan | 2012 | 11 | 10 | 21 | Cancer tissue | Adjacent non-cancerous tissues | Phalanx Human OneArray | PH_hs_0023897 | [34] | |
| GSE31552 | USA | 2014 | 21 | 9 | 2 | 32 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Gene 1.0 ST Array [transcript (gene) version] | 7905258 | [35] |
| GSE10072 | Italy | 2008 | 58 | 49 | Cancer tissue | Healthy lung tissues | Affymetrix Human Genome U133A Array | 203155_at | [36] | ||
| GSE4115 | USA | 2007 | 90 | Cancer tissue | Healthy lung tissues | Affymetrix Human Genome U133A Array | 203155_at | [37] | |||
| GSE31547 | USA | 2018 | 30 | 20 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133A Array | 203155_at | Notpublished | ||
| GSE7670 | Taiwan | 2007 | 26 | 27 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133A Array | 203155_at | [38] | ||
| GSE2514 | USA | 2005 | 20 | 19 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix GeneChip microarray (HGU95Av2) | 34189_at | [39] | ||
| GSE11117 | Switzerland | 2010 | 13 | 14 | 15 | Cancer tissue | Chronic inflammatory lung disease tissues | Nova Chip microarrays (Novartis) | H200002955 | [40] | |
| GSE1987 | Israel | 2006 | 28 | 9 | Cancer tissue | Healthy lung tissues | Affymetrix Human Genome U95A Array | 34189_at | [41] | ||
| GSE19188 | Netherlands | 2010 | 45 | 27 | 65 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133 Plus 2.0 Array | 203155_at | [42] | |
| GSE74706 | Germany | 2016 | 10 | 8 | 18 | Cancer tissue | Adjacent non-cancerous tissues | Agilent-026652 Whole Human Genome Microarray 4 × 44K v2 | A_23_P126393 | [43] | |
| GSE46539 | Taiwan | 2015 | 115 | 115 | Cancer tissue | Adjacent non-cancerous tissues | Illumina WG-DASL HumanRef8 v3 or HumanHT12 v4 BeadChip | ILMN_1718207 | [44] | ||
| GSE2088 | Japan | 2011 | 9 | 48 | 30 | Cancer tissue | Healthy lung tissues | CHUGAI 41K microarray | 11758 | [45] | |
| GSE12428 | Netherlands | 2008 | 34 | 28 | Cancer tissue | Adjacent non-cancerous tissues/Healthy lung tissues | Agilent-012391 Whole Human Genome Oligo Microarray G4112A | 7231 | [46] | ||
| GSE27262 | Taiwan | 2012 | 25 | 25 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133 Plus 2.0 Array | 203155_at | [47] | ||
| GSE101929 | USA | 2017 | 25 | 25 | Cancer tissue | Adjacent non-cancerous tissues | Affymetrix Human Genome U133 Plus 2.0 Array | 203155_at | [48] |
| Characteristics | Std Diff in Means | Lower Limit | Upper Limit | p-Value |
|---|---|---|---|---|
| Clinical Characteristics | ||||
| Size and extent of the main tumor (T) | ||||
| ADC T1 vs. T2/T3/T4 | −0.03 | −0.156 | 0.096 | 0.636 |
| SCC T1 vs. T2/T3/T4 | 0.077 | −0.192 | 0.346 | 0.574 |
| Spread to nearby lymph nodes (N) | ||||
| ADC N0 vs. N1/N2/N3 | 0.028 | −0.21 | 0.266 | 0.82 |
| SCC N0 vs. N1/N2/N3 | 0.075 | −0.137 | 0.287 | 0.489 |
| Stage of cancer | ||||
| ADC 1 vs. 2/3/4 | −0.023 | −0.129 | 0.083 | 0.666 |
| SCC 1 vs. 2/3/4 | −0.038 | −0.179 | 0.103 | 0.596 |
| Presence of mutations in genes | ||||
| EGFR | 0.088 | −0.116 | 0.291 | 0.399 |
| KRAS | 0.114 | −0.114 | 0.342 | 0.326 |
| Smoking Status | ||||
| NSCLC—Age | 0.007 | −0.125 | 0.139 | 0.918 |
| NSCLC—Gender | 0.013 | −0.131 | 0.158 | 0.857 |
| ADC—Age | 0.011 | −0.106 | 0.128 | 0.853 |
| ADC—Gender | 0.012 | −0.184 | 0.208 | 0.906 |
| ADC—Stage 1 vs. 2/3/4 | −0.106 | −0.239 | 0.027 | 0.119 |
| SCC—Age | 0.07 | −0.137 | 0.277 | 0.505 |
| SC—Gender | −0.096 | −0.339 | 0.148 | 0.441 |
| SCC—Stage 1 vs. 2/3/4 | 0.068 | −0.201 | 0.338 | 0.619 |
| Study | Study Design and Subjects | Main Findings | Methods and Analysis | Conclusions |
|---|---|---|---|---|
| Wu et al. 2014 [52] | Primary tumors of lung cancer patients at different clinical stages (n = 192) and adjacent normal tissues (n = 16). | SETDB1 protein levels were elevated in lung cancer tissues compared to non-tumor tissues. Possible association with early stages. | IHC, tissue microarray | SETDB1 is highly expressed in lung cancer. |
| Rodriguez-Paredes et al. 2014 [23] | Primary ADC (n = 20), SCC (n = 20), SCLC (n = 19) tissues. | Amplification of the SETDB1 gene correlates with elevated SETDB1 transcripts and protein overexpression in tissues from patients with NSCLC and SCLC. | FISH, qPCR, IHC | SETDB1 is amplified and highly expressed in NSCLC and SCLC. |
| Sun et al. 2015 [24] | TCGA ADC dataset. | Amplification of SETDB1 loci correlates with elevated SETDB1 transcripts. | Bioinformatics | SETDB1 is amplified and highly expressed in NSCLC. |
| Eight microarrays from GEO and Expression Atlas databases. Primary NSCLC (n = 60) and their paired adjacent normal tissues (n = 60). | SETDB1 mRNA levels were higher in NSCLC tissues compared to non-tumor tissues. Elevated expression of SETDB1 correlates with advanced grade tumors. | Bioinformatics, RT-qPCR | ||
| Lung cancer tissues (n = 387) and normal bronchial epithelium cells (n = 106). | SETDB1 protein levels were elevated in lung cancer tissues compared to non-tumor tissues. Possible association with an advanced pathological stage. | IHC, tissue microarray | ||
| Inoue et al. 2015 [49] | Primary ADC (n = 164) and SCC (n = 99) tissues. | High-level amplification of the SETDB1 gene in ADC tissues was associated with an advanced pathological stage. | FISH | SETDB1 gene amplification is a marker of poor survival in ADC. |
| Low-level amplification of the SETDB1 gene was observed in SCC tissues. | FISH | |||
| SETDB1 gene amplification was associated with shorter postoperative overall survival in ADC patients. | Kaplan-Meier analysis | |||
| Lafuente-Sanchis et al. 2015 [51] | Stage I primary NSCLC (n = 64) and adjacent normal tissues. | SETDB1 mRNA was upregulated in primary NSCLC tissues. High SETDB1 mRNA levels were associated with a shorter disease-free survival in stage I NSCLC. | RT-qPCR, Kaplan-Meier analysis | High mRNA levels of SETDB1 as a prognostic marker of a shorter disease-free survival. |
| Chen et al. 2018 [54] | Stage III and IV primary NSCLC tissues (n = 30) and paired adjacent normal tissues (n = 30). | SETDB1 mRNA was upregulated in primary NSCLC tissues. SETDB1 and TP53 mRNA levels were negatively correlated. | RT-qPCR | SETDB1 is highly expressed in NSCLC. |
| Wang et al. 2019 [50] | Oncomine database: NSCLC tissues (n = 1926). | SETDB1 mRNA was upregulated in NSCLC tissues. Higher expression of SETDB1 mRNA was associated shorter overall survival in NSCLC patients. | Bioinformatics, Kaplan-Meier analysis | High mRNA levels of SETDB1 as a prognostic marker of poor survival in NSCLC patients. |
| Primary NSCLC tissues (n = 9) and paired adjacent normal tissues (n = 9). | SETDB1 was overexpressed in most paired NSCLC tumors compared to non-tumor tissues. | WB | SETDB1 is highly expressed in NSCLC. | |
| Primary NSCLC tissues (n = 156). | SETDB1 protein levels were elevated in NSCLC tissues (SETDB1 was detected in the nucleus and cytoplasm). | IHC |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cruz-Tapias, P.; Zakharova, V.; Perez-Fernandez, O.M.; Mantilla, W.; Ramírez-Clavijo, S.; Ait-Si-Ali, S. Expression of the Major and Pro-Oncogenic H3K9 Lysine Methyltransferase SETDB1 in Non-Small Cell Lung Cancer. Cancers 2019, 11, 1134. https://doi.org/10.3390/cancers11081134
Cruz-Tapias P, Zakharova V, Perez-Fernandez OM, Mantilla W, Ramírez-Clavijo S, Ait-Si-Ali S. Expression of the Major and Pro-Oncogenic H3K9 Lysine Methyltransferase SETDB1 in Non-Small Cell Lung Cancer. Cancers. 2019; 11(8):1134. https://doi.org/10.3390/cancers11081134
Chicago/Turabian StyleCruz-Tapias, Paola, Vlada Zakharova, Oscar M. Perez-Fernandez, William Mantilla, Sandra Ramírez-Clavijo, and Slimane Ait-Si-Ali. 2019. "Expression of the Major and Pro-Oncogenic H3K9 Lysine Methyltransferase SETDB1 in Non-Small Cell Lung Cancer" Cancers 11, no. 8: 1134. https://doi.org/10.3390/cancers11081134
APA StyleCruz-Tapias, P., Zakharova, V., Perez-Fernandez, O. M., Mantilla, W., Ramírez-Clavijo, S., & Ait-Si-Ali, S. (2019). Expression of the Major and Pro-Oncogenic H3K9 Lysine Methyltransferase SETDB1 in Non-Small Cell Lung Cancer. Cancers, 11(8), 1134. https://doi.org/10.3390/cancers11081134





