Analysis of AR/ARV7 Expression in Isolated Circulating Tumor Cells of Patients with Metastatic Castration-Resistant Prostate Cancer (SAKK 08/14 IMPROVE Trial)
Abstract
1. Introduction
2. Results
2.1. Sensitivity and Specificity of the Adna ARV7 Test
2.2. Cross-Laboratory Variability
2.3. Reproducibility of the CTC Profile Measured at Baseline by the Adna ARV7 Test
2.4. Patient Cohort
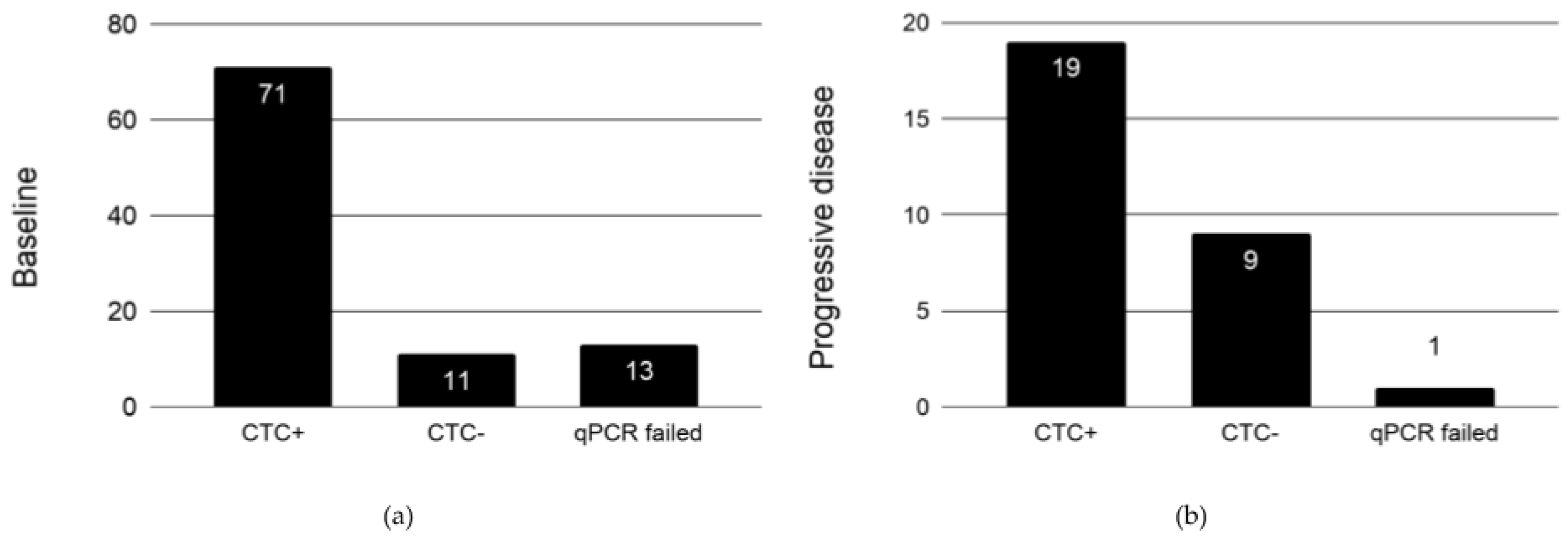
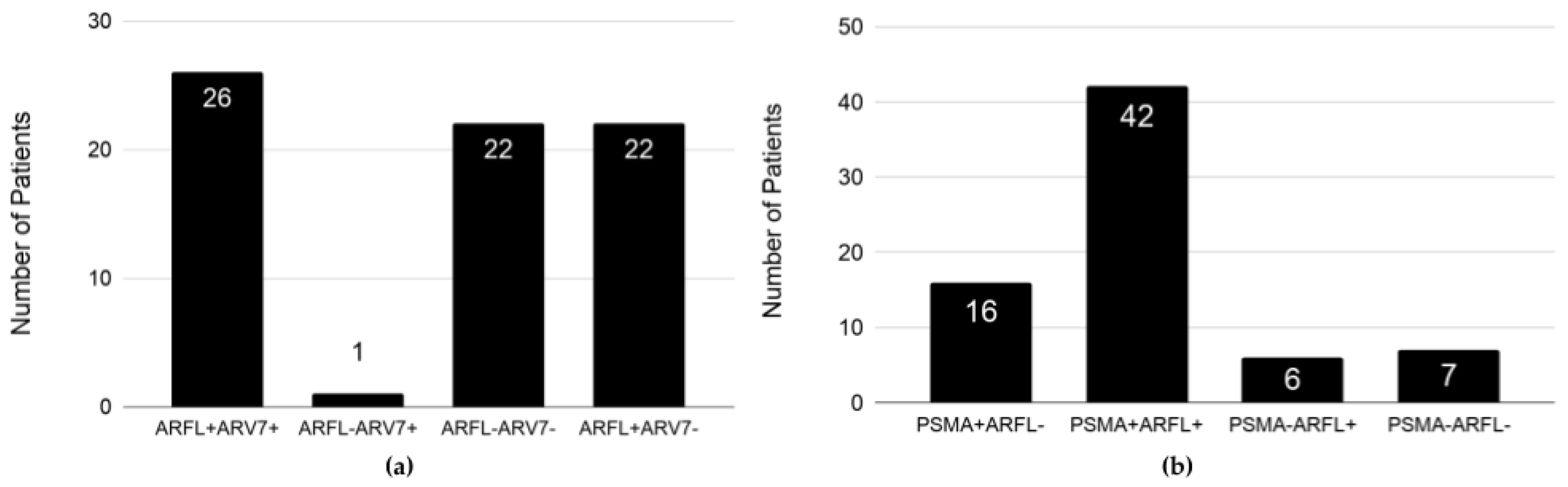
2.5. Expression Profile of EpCAM+ CTCs in Patients at Baseline
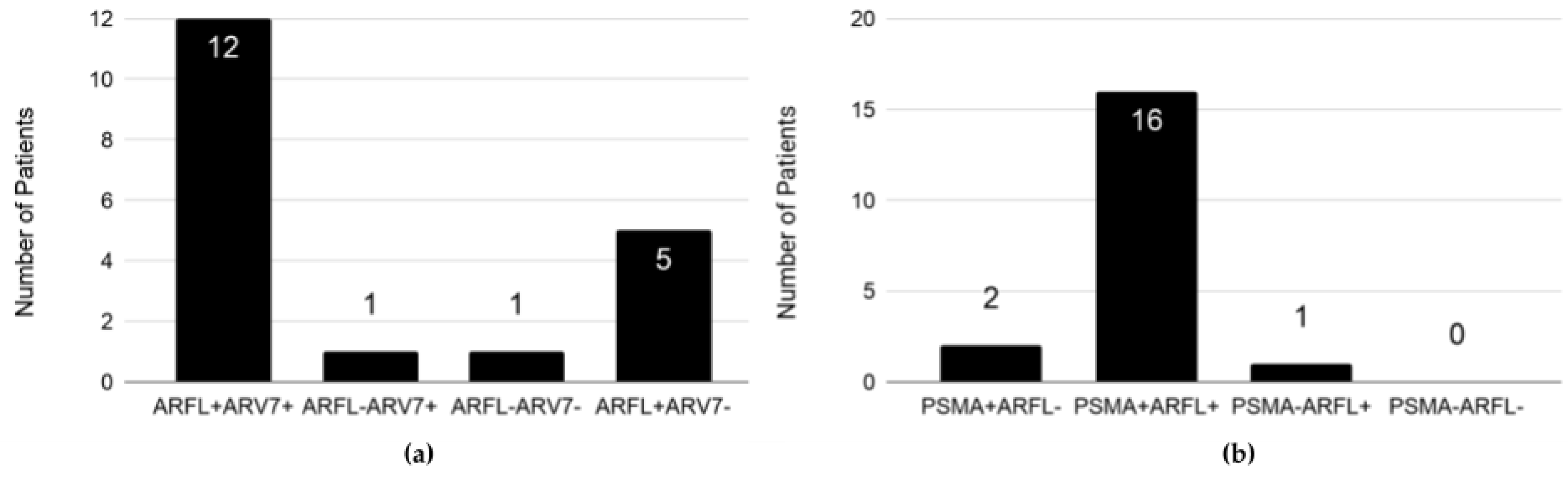
2.6. Expression Profile of EpCAM+ CTCs in Patients on Enzalutamide at Progressive Disease
2.7. ARFL/ARV7 Conversions at Progressive Disease
2.8. PSMA/ARFL Conversions at Progressive Disease
3. Discussion
4. Material and Methods
4.1. Study Design
4.2. Patient Characteristics
4.3. RNA Extraction from VCaP, LNCaP, PC3, and DU145 Cell Lines and Detection of PSA, PSMA, ARFL, and ARV7 Expression by RT-qPCR
4.4. Quantitative Real-Time PCR and PSA, PSMA, ARFL and ARV7 Detection in EpCAM+ CTCs
4.5. Analysis of PSA and PSMA Expression in EpCAM+ CTCs by Adna Detect
4.6. Cell Spiking—Validation Experiments
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Negoita, S.; Feuer, E.J.; Mariotto, A.; Cronin, K.A.; Petkov, V.I.; Hussey, S.K.; Benard, V.; Henley, S.J.; Anderson, R.N.; Fedewa, S.; et al. Annual Report to the Nation on the Status of Cancer, part II: Recent changes in prostate cancer trends and disease characteristics: Recent Changes in Prostate Cancer Trends. Cancer 2018, 124, 2801–2814. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Li, L.; Yang, Z.; Luo, J.; Yeh, S.; Chang, C. Androgen-deprivation therapy with enzalutamide enhances prostate cancer metastasis via decreasing the EPHB6 suppressor expression. Cancer Lett. 2017, 408, 155–163. [Google Scholar] [CrossRef] [PubMed]
- James, N.D.; Spears, M.R.; Clarke, N.W.; Dearnaley, D.P.; De Bono, J.S.; Gale, J.; Hetherington, J.; Hoskin, P.J.; Jones, R.J.; Laing, R.; et al. Survival with Newly Diagnosed Metastatic Prostate Cancer in the “Docetaxel Era”: Data from 917 Patients in the Control Arm of the STAMPEDE Trial (MRC PR08, CRUK/06/019). Eur. Urol. 2015, 67, 1028–1038. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Tangen, C.M.; Higano, C.; Schelhammer, P.F.; Faulkner, J.; Crawford, E.D.; Wilding, G.; Akdas, A.; Small, E.J.; Donnelly, B.; et al. Absolute Prostate-Specific Antigen Value After Androgen Deprivation Is a Strong Independent Predictor of Survival in New Metastatic Prostate Cancer: Data From Southwest Oncology Group Trial 9346 (INT-0162). J. Clin. Oncol. 2006, 24, 3984–3990. [Google Scholar] [CrossRef] [PubMed]
- Mohler, J.L.; Armstrong, A.J.; Bahnson, R.R.; D’Amico, A.V.; Davis, B.J.; Eastham, J.A.; Enke, C.A.; Farrington, T.A.; Higano, C.S.; Horwitz, E.M.; et al. Prostate Cancer, Version 1.2016. J. Natl. Compr. Cancer Netw. JNCCN 2016, 14, 19–30. [Google Scholar] [CrossRef]
- Gillessen, S.; Attard, G.; Beer, T.M.; Beltran, H.; Bossi, A.; Bristow, R.; Carver, B.; Castellano, D.; Chung, B.H.; Clarke, N.; et al. Management of Patients with Advanced Prostate Cancer: The Report of the Advanced Prostate Cancer Consensus Conference APCCC 2017. Eur. Urol. 2018, 73, 178–211. [Google Scholar] [CrossRef] [PubMed]
- Galletti, G.; Leach, B.I.; Lam, L.; Tagawa, S.T. Mechanisms of resistance to systemic therapy in metastatic castration-resistant prostate cancer. Cancer Treat. Rev. 2017, 57, 16–27. [Google Scholar] [CrossRef]
- Guo, Z.; Yang, X.; Sun, F.; Jiang, R.; Linn, D.E.; Chen, H.; Chen, H.; Kong, X.; Melamed, J.; Tepper, C.G.; et al. A Novel Androgen Receptor Splice Variant Is Up-regulated during Prostate Cancer Progression and Promotes Androgen Depletion-Resistant Growth. Cancer Res. 2009, 69, 2305–2313. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.; Dunn, T.A.; Wei, S.; Isharwal, S.; Veltri, R.W.; Humphreys, E.; Han, M.; Partin, A.W.; Vessella, R.L.; Isaacs, W.B.; et al. Ligand-Independent Androgen Receptor Variants Derived from Splicing of Cryptic Exons Signify Hormone-Refractory Prostate Cancer. Cancer Res. 2009, 69, 16–22. [Google Scholar] [CrossRef]
- Antonarakis, E.S.; Lu, C.; Luber, B.; Wang, H.; Chen, Y.; Zhu, Y.; Silberstein, J.L.; Taylor, M.N.; Maughan, B.L.; Denmeade, S.R.; et al. Clinical Significance of Androgen Receptor Splice Variant-7 mRNA Detection in Circulating Tumor Cells of Men With Metastatic Castration-Resistant Prostate Cancer Treated With First- and Second-Line Abiraterone and Enzalutamide. J. Clin. Oncol. 2017, 35, 2149–2156. [Google Scholar] [CrossRef]
- Antonarakis, E.S.; Lu, C.; Wang, H.; Luber, B.; Nakazawa, M.; Roeser, J.C.; Chen, Y.; Mohammad, T.A.; Chen, Y.; Fedor, H.L.; et al. AR-V7 and Resistance to Enzalutamide and Abiraterone in Prostate Cancer. N. Engl. J. Med. 2014, 371, 1028–1038. [Google Scholar] [CrossRef] [PubMed]
- Sciarra, A.; Gentilucci, A.; Silvestri, I.; Salciccia, S.; Cattarino, S.; Scarpa, S.; Gatto, A.; Frantellizzi, V.; Von Heland, M.; Ricciuti, G.P.; et al. Androgen receptor variant 7 (AR-V7) in sequencing therapeutic agents for castratrion resistant prostate cancer: A critical review. Medicine (Baltimore) 2019, 98, e15608. [Google Scholar] [CrossRef] [PubMed]
- Steinestel, J.; Luedeke, M.; Arndt, A.; Schnoeller, T.J.; Lennerz, J.K.; Wurm, C.; Maier, C.; Cronauer, M.V.; Schrader, A.J. Detecting predictive androgen receptor modifications in circulating prostate cancer cells. Oncotarget 2015, 10, 4213–4223. [Google Scholar] [PubMed]
- Steinestel, J.; Bernemann, C.; Schrader, A.J.; Lennerz, J.K. Re: Emmanuel S. Antonarakis, Changxue Lu, Brandon Luber, et al. Clinical Significance of Androgen Receptor Splice Variant-7 mRNA Detection in Circulating Tumor Cells of Men with Metastatic Castration-resistant Prostate Cancer Treated with First- and Second-line Abiraterone and Enzalutamide. J Clin Oncol 2017; 35, 2149–56: AR-V7 Testing: What’s in it for the Patient? Eur. Urol. 2017, 72, e168–e169. [Google Scholar] [PubMed]
- Sharp, A.; Welti, J.C.; Lambros, M.B.K.; Dolling, D.; Rodrigues, D.N.; Pope, L.; Aversa, C.; Figueiredo, I.; Fraser, J.; Ahmad, Z.; et al. Clinical Utility of Circulating Tumour Cell Androgen Receptor Splice Variant-7 Status in Metastatic Castration-resistant Prostate Cancer. Eur. Urol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Dong, X. Complex Impacts of PI3K/AKT Inhibitors to Androgen Receptor Gene Expression in Prostate Cancer Cells. PLoS ONE 2014, 9, e108780. [Google Scholar] [CrossRef] [PubMed]
- Korenchuk, S.; Lehr, J.E.; MClean, L.; Lee, Y.G.; Whitney, S.; Vessella, R.; Lin, D.L.; Pienta, K.J. VCaP, a cell-based model system of human prostate cancer. In Vivo Athens Greece 2001, 15, 163–168. [Google Scholar]
- Scher, H.I.; Lu, D.; Schreiber, N.A.; Louw, J.; Graf, R.P.; Vargas, H.A.; Johnson, A.; Jendrisak, A.; Bambury, R.; Danila, D.; et al. Association of AR-V7 on Circulating Tumor Cells as a Treatment-Specific Biomarker With Outcomes and Survival in Castration-Resistant Prostate Cancer. JAMA Oncol. 2016, 2, 1441–1449. [Google Scholar] [CrossRef]
- Bernemann, C.; Schnoeller, T.J.; Luedeke, M.; Steinestel, K.; Boegemann, M.; Schrader, A.J.; Steinestel, J. Expression of AR-V7 in Circulating Tumour Cells Does Not Preclude Response to Next Generation Androgen Deprivation Therapy in Patients with Castration Resistant Prostate Cancer. Eur. Urol. 2017, 71, 1–3. [Google Scholar] [CrossRef]
- Armstrong, A.J.; Halabi, S.; Luo, J.; Nanus, D.M.; Giannakakou, P.; Szmulewitz, R.Z.; Danila, D.C.; Healy, P.; Anand, M.; Rothwell, C.J.; et al. Prospective Multicenter Validation of Androgen Receptor Splice Variant 7 and Hormone Therapy Resistance in High-Risk Castration-Resistant Prostate Cancer: The PROPHECY Study. J. Clin. Oncol. 2019, 37, 1120–1129. [Google Scholar] [CrossRef]
- Scher, H.I.; Graf, R.P.; Schreiber, N.A.; McLaughlin, B.; Lu, D.; Louw, J.; Danila, D.C.; Dugan, L.; Johnson, A.; Heller, G.; et al. Nuclear-specific AR-V7 Protein Localization is Necessary to Guide Treatment Selection in Metastatic Castration-resistant Prostate Cancer. Eur. Urol. 2017, 71, 874–882. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S. Predicting treatment response in castration-resistant prostate cancer: Could androgen receptor variant-7 hold the key? Expert Rev. Anticancer Ther. 2015, 15, 143–145. [Google Scholar] [CrossRef] [PubMed]
- Onstenk, W.; Sieuwerts, A.M.; Kraan, J.; Van, M.; Nieuweboer, A.J.M.; Mathijssen, R.H.J.; Hamberg, P.; Meulenbeld, H.J.; De Laere, B.; Dirix, L.Y.; et al. Efficacy of Cabazitaxel in Castration-resistant Prostate Cancer Is Independent of the Presence of AR-V7 in Circulating Tumor Cells. Eur. Urol. 2015, 68, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Danila, D.C.; Samoila, A.; Patel, C.; Schreiber, N.; Herkal, A.; Anand, A.; Bastos, D.; Heller, G.; Fleisher, M.; Scher, H.I. Clinical Validity of Detecting Circulating Tumor Cells by AdnaTest Assay Compared With Direct Detection of Tumor mRNA in Stabilized Whole Blood, as a Biomarker Predicting Overall Survival for Metastatic Castration-Resistant Prostate Cancer Patients. Cancer J. 2016, 22, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Khetawat, G.; Faraday, N.; Nealen, M.L.; Vijayan, K.V.; Bolton, E.; Noga, S.J.; Bray, P.F. Human megakaryocytes and platelets contain the estrogen receptor beta and androgen receptor (AR): Testosterone regulates AR expression. Blood 2000, 95, 2289–2296. [Google Scholar] [PubMed]
- Van der Toom, E.E.; Axelrod, H.D.; de la Rosette, J.J.; de Reijke, T.M.; Pienta, K.J.; Valkenburg, K.C. Prostate-specific markers to identify rare prostate cancer cells in liquid biopsies. Nat. Rev. Urol. 2019, 16, 7–22. [Google Scholar] [CrossRef] [PubMed]
- Abu, E.O.; Horner, A.; Kusec, V.; Triffitt, J.T.; Compston, J.E. The Localization of Androgen Receptors in Human Bone. J. Clin. Endocrinol. Metab. 1997, 82, 3493–3497. [Google Scholar] [CrossRef]
- Danila, D.C.; Heller, G.; Gignac, G.A.; Gonzalez-Espinoza, R.; Anand, A.; Tanaka, E.; Lilja, H.; Schwartz, L.; Larson, S.; Fleisher, M.; et al. Circulating Tumor Cell Number and Prognosis in Progressive Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2007, 13, 7053–7058. [Google Scholar] [CrossRef]
- De Bono, J.S.; Scher, H.I.; Montgomery, R.B.; Parker, C.; Miller, M.C.; Tissing, H.; Doyle, G.V.; Terstappen, L.W.W.M.; Pienta, K.J.; Raghavan, D. Circulating Tumor Cells Predict Survival Benefit from Treatment in Metastatic Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2008, 14, 6302–6309. [Google Scholar] [CrossRef]
- Heller, G.; McCormack, R.; Kheoh, T.; Molina, A.; Smith, M.R.; Dreicer, R.; Saad, F.; de Wit, R.; Aftab, D.T.; Hirmand, M.; et al. Circulating Tumor Cell Number as a Response Measure of Prolonged Survival for Metastatic Castration-Resistant Prostate Cancer: A Comparison With Prostate-Specific Antigen Across Five Randomized Phase III Clinical Trials. J. Clin. Oncol. 2018, 36, 572–580. [Google Scholar] [CrossRef]
- Thalgott, M.; Heck, M.M.; Eiber, M.; Souvatzoglou, M.; Hatzichristodoulou, G.; Kehl, V.; Krause, B.J.; Rack, B.; Retz, M.; Gschwend, J.E.; et al. Circulating tumor cells versus objective response assessment predicting survival in metastatic castration-resistant prostate cancer patients treated with docetaxel chemotherapy. J. Cancer Res. Clin. Oncol. 2015, 141, 1457–1464. [Google Scholar] [CrossRef] [PubMed]
- Okegawa, T.; Itaya, N.; Hara, H.; Tambo, M.; Nutahara, K. Circulating tumor cells as a biomarker predictive of sensitivity to docetaxel chemotherapy in patients with castration-resistant prostate cancer. Anticancer Res. 2014, 34, 6705–6710. [Google Scholar] [PubMed]
- Perner, S.; Hofer, M.D.; Kim, R.; Shah, R.B.; Li, H.; Möller, P.; Hautmann, R.E.; Gschwend, J.E.; Kuefer, R.; Rubin, M.A. Prostate-specific membrane antigen expression as a predictor of prostate cancer progression. Hum. Pathol. 2007, 38, 696–701. [Google Scholar] [CrossRef] [PubMed]
- Minner, S.; Wittmer, C.; Graefen, M.; Salomon, G.; Steuber, T.; Haese, A.; Huland, H.; Bokemeyer, C.; Yekebas, E.; Dierlamm, J.; et al. High level PSMA expression is associated with early psa recurrence in surgically treated prostate cancer. Prostate 2011, 71, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Joung, J.Y.; Cho, K.S.; Chung, H.S.; Cho, I.-C.; Kim, J.E.; Seo, H.K.; Chung, J.; Park, W.S.; Choi, M.K.; Lee, K.H. Prostate Specific Membrane Antigen mRNA in Blood as a Potential Predictor of Biochemical Recurrence after Radical Prostatectomy. J. Korean Med. Sci. 2010, 25, 1291–1295. [Google Scholar] [CrossRef] [PubMed]
- Gorges, T.M.; Riethdorf, S.; von Ahsen, O.; Nastały, P.; Röck, K.; Boede, M.; Peine, S.; Kuske, A.; Schmid, E.; Kneip, C.; et al. Heterogeneous PSMA expression on circulating tumor cells—A potential basis for stratification and monitoring of PSMA-directed therapies in prostate cancer. Oncotarget 2016, 7, 34930–34941. [Google Scholar] [CrossRef]
- Lowes, L.E.; Goodale, D.; Xia, Y.; Postenka, C.; Piaseczny, M.M.; Paczkowski, F.; Allan, A.L. Epithelial-to-mesenchymal transition leads to disease-stage differences in circulating tumor cell detection and metastasis in pre-clinical models of prostate cancer. Oncotarget 2016, 7, 76125–76139. [Google Scholar] [CrossRef]
| Sample | GAPDH | PSA | PSMA | ARFL | ARV7 |
|---|---|---|---|---|---|
| DU145 (50 × 103)3 | + | − | − | − | − |
| DU145 (RNA)2 | + | − | − | − | − |
| PC3 (50 × 103)2 | + | − | + | − | − |
| Healthy 10 * | − | − | − | −/+ | − |
| LNCaP (200)1 | + | + | + | + | + |
| LNCaP (68)1 | + | + | + | + | + |
| LNCaP (10)4 | + | + | + | + | ++−− |
| LNCaP (5)3 | ++− | + | ++− | ++− | ++− |
| LNCaP95 (5)5 | ++++− | +++−− | + | + | + |
| VCaP (18 × 103)1 | + | − | + | + | + |
| VCaP (10)3 | +−+ | +−+ | + | + | ++− |
| VCaP (5)3 | − | +−− | + | + | + |
| Laboratories | GAPDH | PSA | PSMA | ARFL | ARV7 |
|---|---|---|---|---|---|
| Concordance | 90% (9/10) | 70% (7/10) | 100% (10/10) | 100% (10/10) | 100% (10/10) |
| Laboratories | GAPDH | PSA | PSMA | ARFL | ARV7 |
|---|---|---|---|---|---|
| Concordance | 92.86% (13/14) | 78% (11/14) | 100% (14/14) | 100% (14/14) | 100% (14/14) |
| Patients | PSMA | ARFL | ARV7 |
|---|---|---|---|
| P1 | 100% (2/2) | 100% (2/2) | 100% (2/2) |
| P2 | 100% (2/2) | 100% (2/2) | 100% (2/2) |
| P3 | 100% (2/2) | 100% (2/2) | 100% (2/2) |
| P4 | 100% (2/2) | 100% (2/2) | 100% (2/2) |
| P5 | 50% (1/2) | 50% (1/2) | 50% (1/2) |
| Progressive Disease (PD) | |||||
|---|---|---|---|---|---|
| Baseline | ARFL+ ARV7+ | ARFL+ ARV7− | ARFL− ARV7+ | ARFL− ARV7− | no CTCs |
| ARFL+, ARV7+ (8) | 75% (6) | 12.5% (1) | 0 | 0 | 12.5% (1) |
| ARFL+, ARV7− (9) | 44.4% (4) | 22.2% (2) | 0 | 0 | 33.3% (3) |
| ARFL−, ARV7− (5) | 20% (1) | 20% (1) | 20% (1) | 20% (1) | 20% (1) |
| no CTCs (3) | 0 | 33.3% (1) | 0 | 0 | 66.7% (2) |
| Total (25) | 11 | 5 | 1 | 1 | 7 |
| Progressive Disease (PD) | |||||
|---|---|---|---|---|---|
| Baseline | PSMA+ ARFL+ | PSMA+ ARFL− | PSMA− ARFL+ | PSMA− ARFL− | no CTCs |
| PSMA+, ARFL+ (14) | 71.4% (10) | 0 | 7.1% (1) | 0 | 21.4% (3) |
| PSMA+, ARFL− (3) | 33.3% (1) | 66.7% (2) | 0 | 0 | 0 |
| PSMA−, ARFL+ (3) | 66.7% (2) | 0 | 0 | 0 | 33.3% (1) |
| PSMA−, ARFL− (2) | 50% (1) | 0 | 0 | 0 | 50% (1) |
| no CTCs (3) | 33.3% (1) | 0 | 0 | 0 | 66.7% (2) |
| Total (25) | 15 | 2 | 1 | 0 | 7 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bratic Hench, I.; Cathomas, R.; Costa, L.; Fischer, N.; Gillessen, S.; Hench, J.; Hermanns, T.; Kremer, E.; Mingrone, W.; Pereira Mestre, R.; et al. Analysis of AR/ARV7 Expression in Isolated Circulating Tumor Cells of Patients with Metastatic Castration-Resistant Prostate Cancer (SAKK 08/14 IMPROVE Trial). Cancers 2019, 11, 1099. https://doi.org/10.3390/cancers11081099
Bratic Hench I, Cathomas R, Costa L, Fischer N, Gillessen S, Hench J, Hermanns T, Kremer E, Mingrone W, Pereira Mestre R, et al. Analysis of AR/ARV7 Expression in Isolated Circulating Tumor Cells of Patients with Metastatic Castration-Resistant Prostate Cancer (SAKK 08/14 IMPROVE Trial). Cancers. 2019; 11(8):1099. https://doi.org/10.3390/cancers11081099
Chicago/Turabian StyleBratic Hench, Ivana, Richard Cathomas, Luigi Costa, Natalie Fischer, Silke Gillessen, Jürgen Hench, Thomas Hermanns, Eloïse Kremer, Walter Mingrone, Ricardo Pereira Mestre, and et al. 2019. "Analysis of AR/ARV7 Expression in Isolated Circulating Tumor Cells of Patients with Metastatic Castration-Resistant Prostate Cancer (SAKK 08/14 IMPROVE Trial)" Cancers 11, no. 8: 1099. https://doi.org/10.3390/cancers11081099
APA StyleBratic Hench, I., Cathomas, R., Costa, L., Fischer, N., Gillessen, S., Hench, J., Hermanns, T., Kremer, E., Mingrone, W., Pereira Mestre, R., Püschel, H., Rothermundt, C., Ruiz, C., Tolnay, M., Von Burg, P., Bubendorf, L., & Vlajnic, T. (2019). Analysis of AR/ARV7 Expression in Isolated Circulating Tumor Cells of Patients with Metastatic Castration-Resistant Prostate Cancer (SAKK 08/14 IMPROVE Trial). Cancers, 11(8), 1099. https://doi.org/10.3390/cancers11081099






