miRNA Predictors of Pancreatic Cancer Chemotherapeutic Response: A Systematic Review and Meta-Analysis
Abstract
1. Introduction
2. Methods
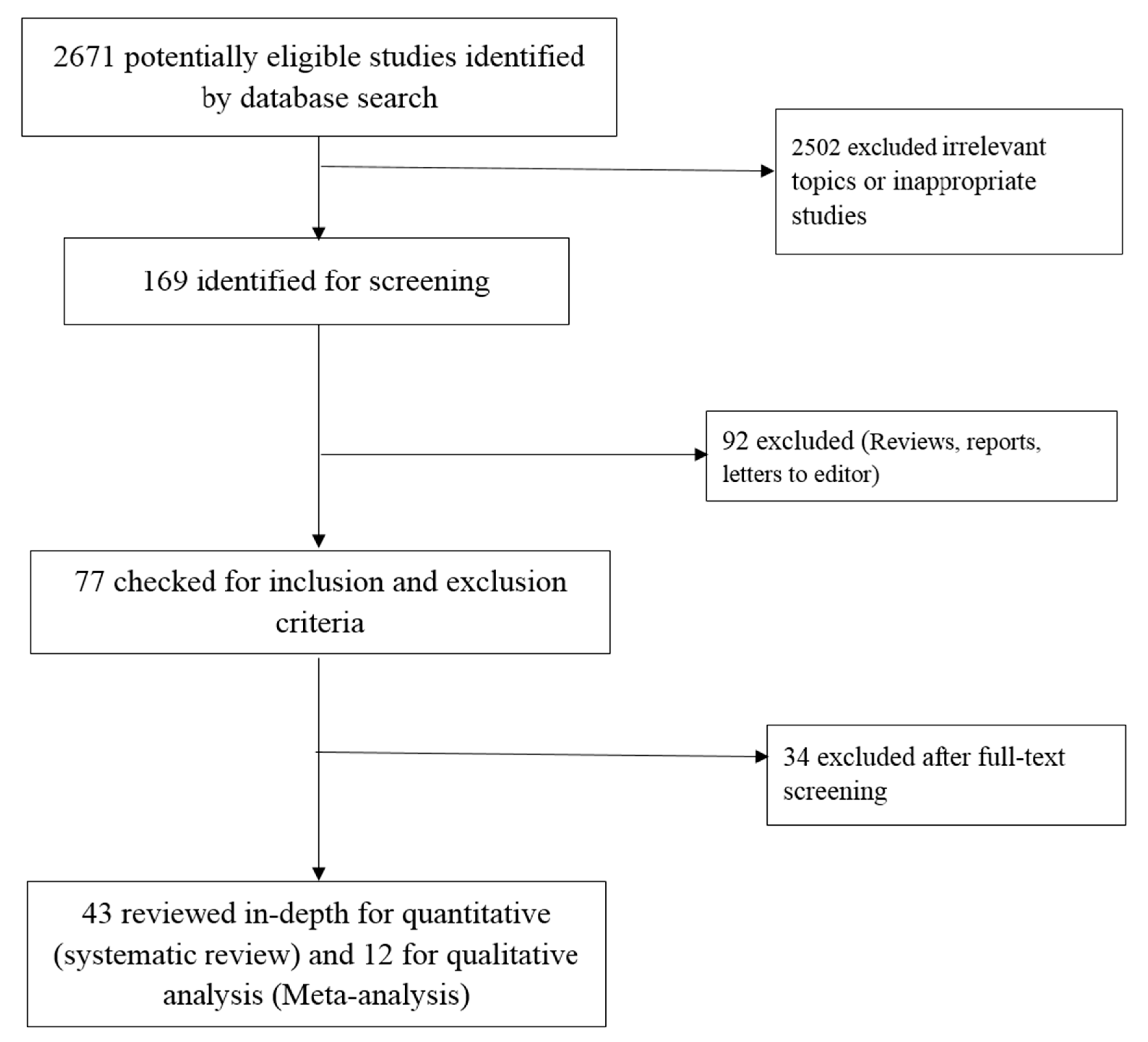
2.1. Search Strategy and Study Selection
2.2. Selection Criteria
2.3. Inclusion Criteria
- Studies involving miRNAs expression and PC.
- Studies involving clinical patient data or preclinical data.
- Studies focusing on the resistance to some form of chemotherapy.
- Studies that reported the miRNA profiling platforms.
- Articles using in vitro assays to analyse the expression of miRNAs or gene related studies.
2.4. Exclusion Criteria
- Studies published in non-English language and that do not involve drug resistance in PC were removed.
- Case reports, review articles, editorial, and studies with only in vitro or only PC patient samples data were excluded.
2.5. Data Extraction
2.6. Quality Assessment
2.7. Publication Bias
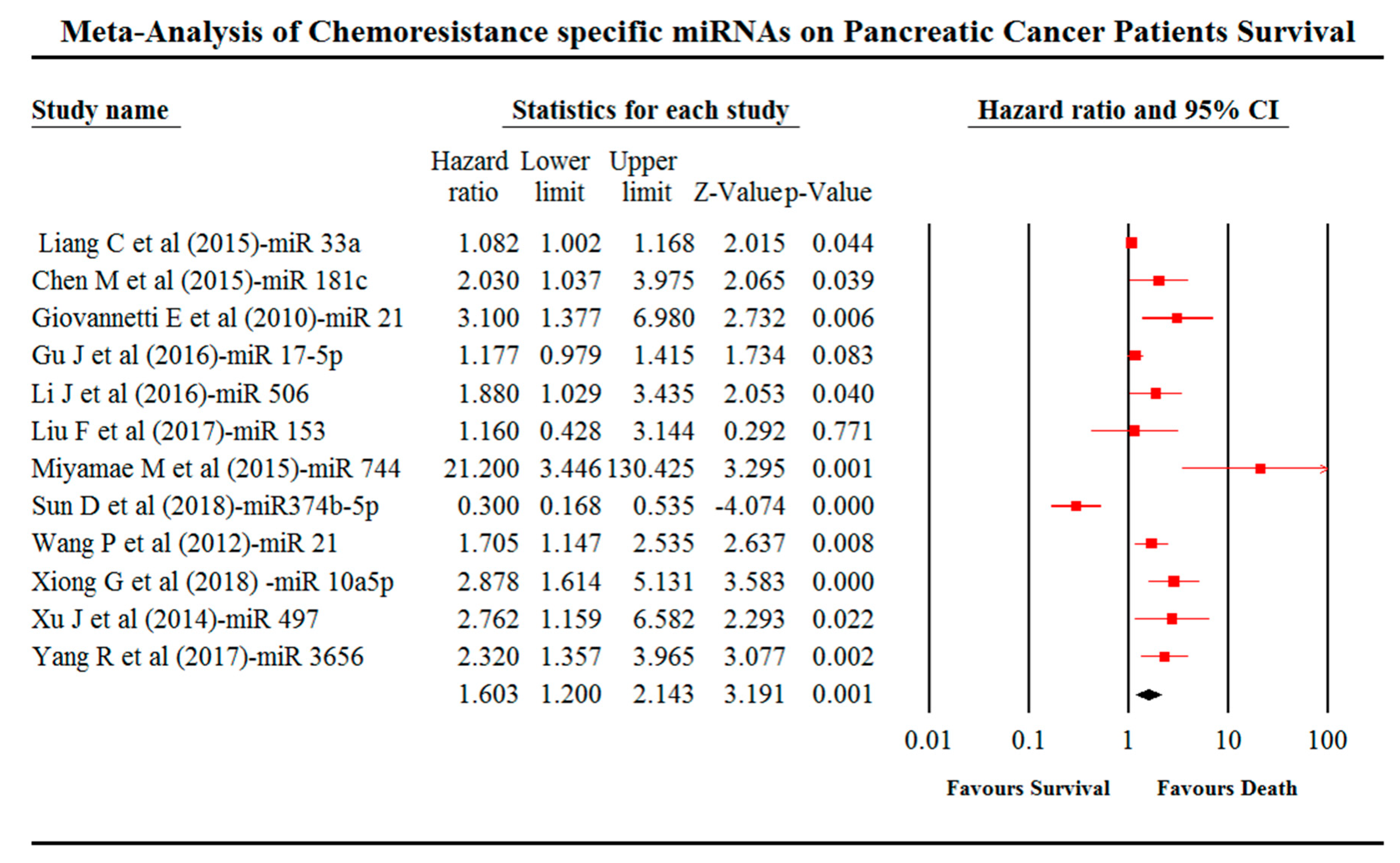
2.8. Meta-Analysis
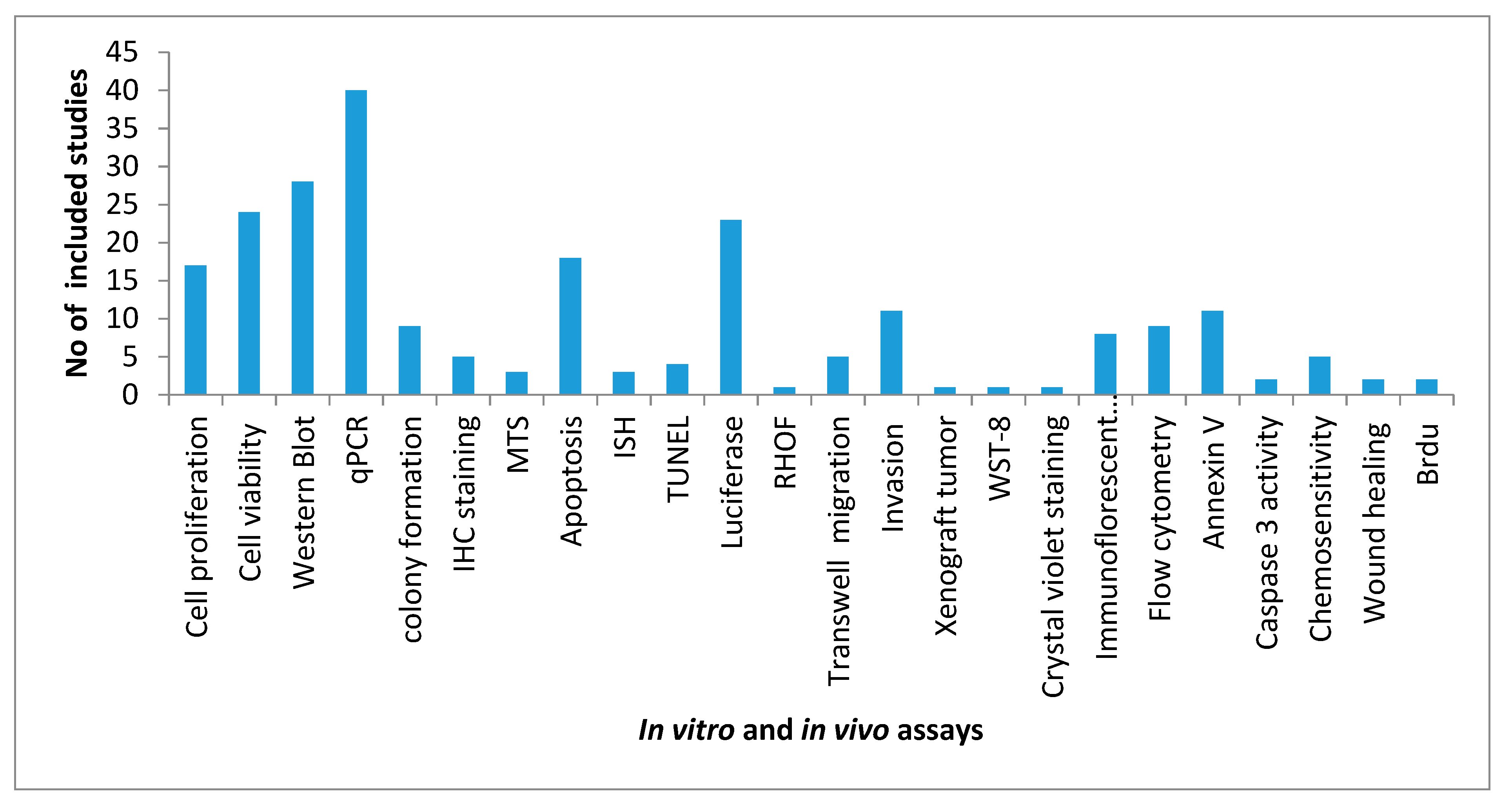
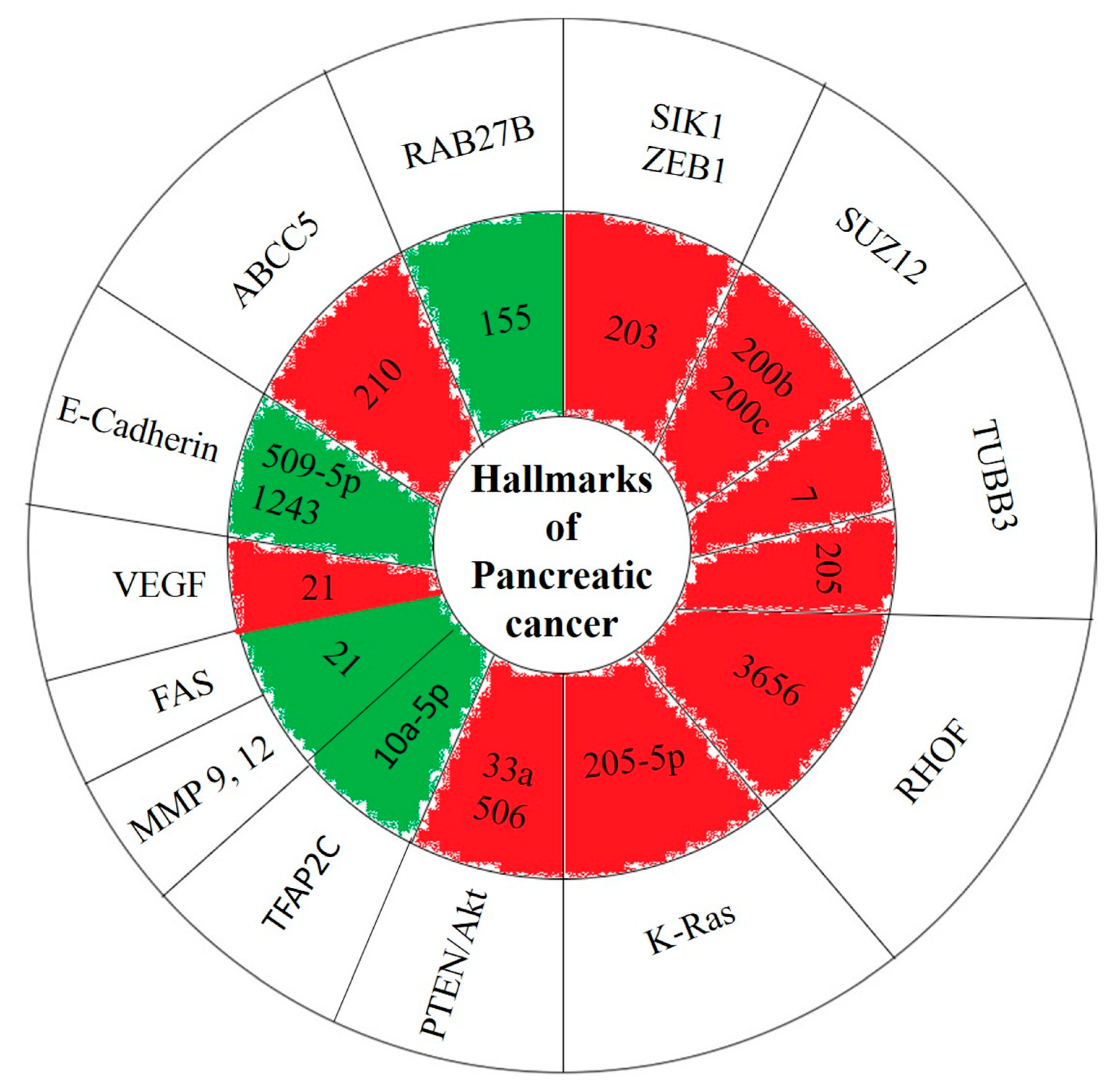
3. Results
4. Meta-Analysis
5. Publication Bias Indicators
5.1. Classic Fail-Safe N
5.2. Orwin Fail-Safe N
5.3. Begg and Mazumdar Rank Correlation Test
5.4. Egger′s Test of the Intercept
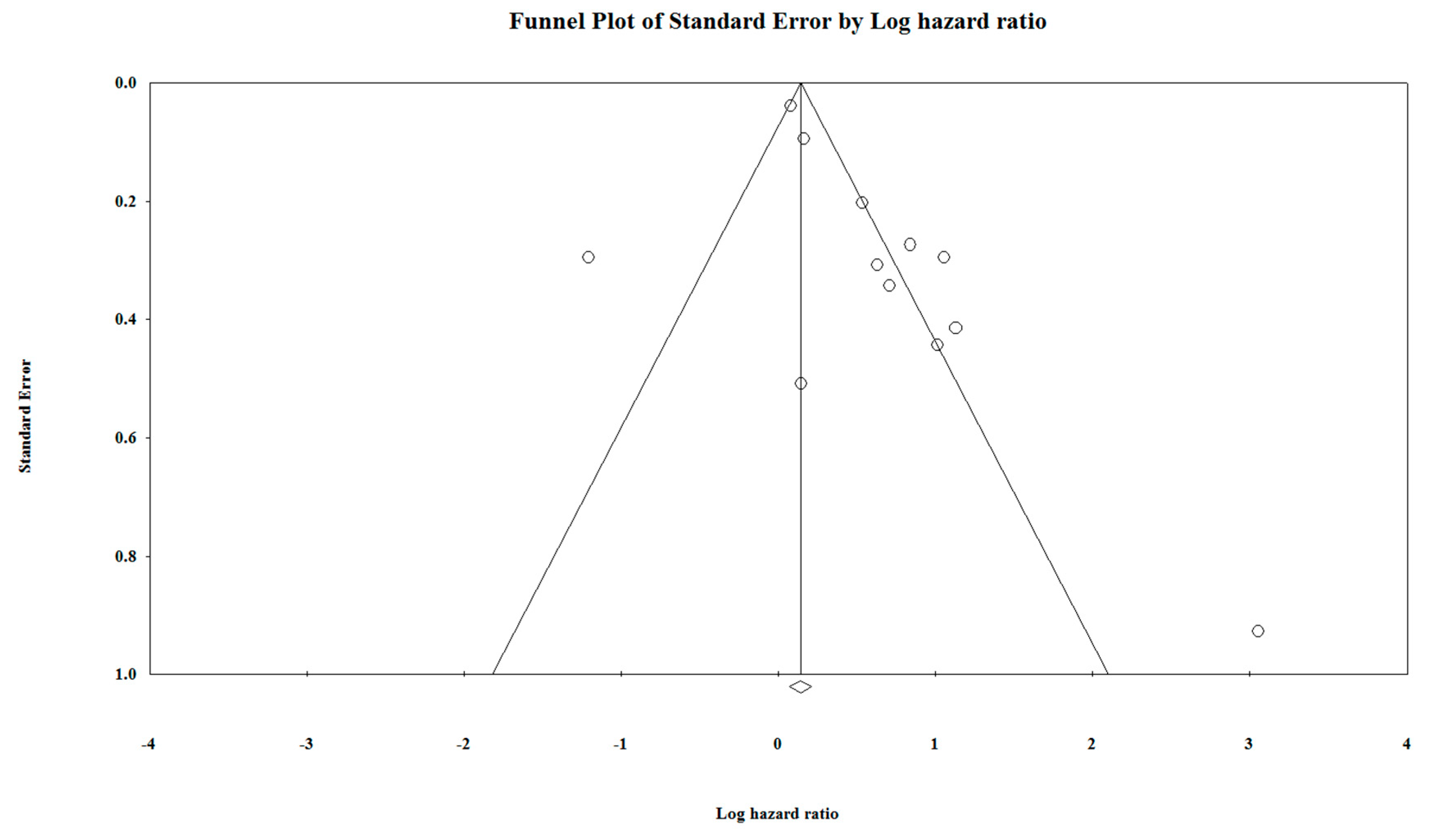
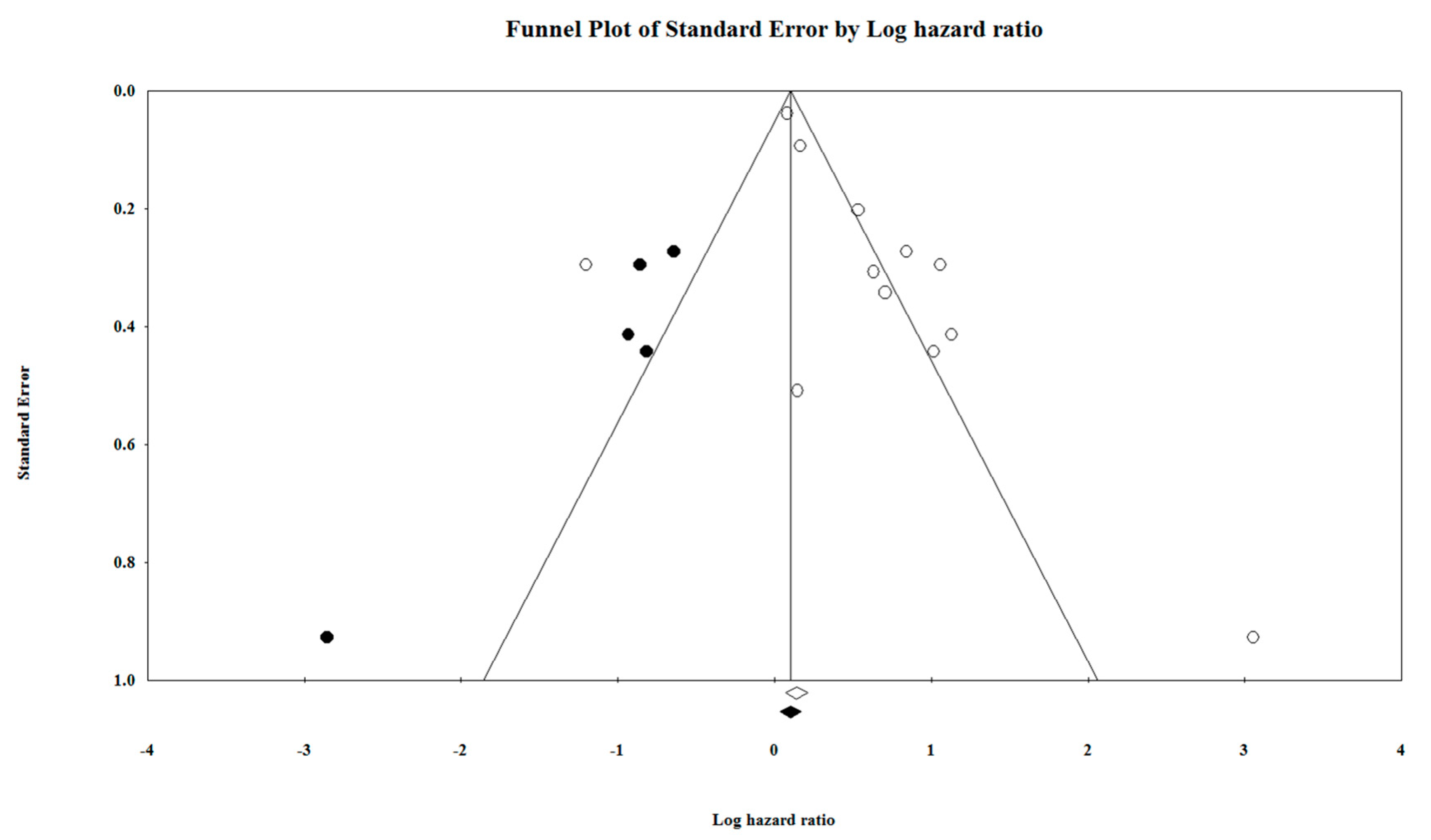
5.5. Duval and Tweedie′s Trim and Fill Test
6. Discussion
7. Limitations and Strengths of Our Study
8. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PC | Pancreatic cancer |
| HR | Hazard ratio |
| CIs | Confidence Intervals |
| GEM | Gemcitabine |
| 5-FU | 5-fluorouracil |
| CAP | Capecitabine |
| LAP | Lapatinib |
| GSI | γ-secretase inhibitor |
| OHP | Oxaliplatin |
| NM | Not Mentioned |
| RHOF | Ras homolog family member F |
| EMT | Epithelial–Mesenchymal Transition |
| CXCR4 | C-X- C chemokine receptor type 4 |
| HMGA2 | High-mobility group AT-hook 2 |
| DNA-PKcs | DNA- dependent protein kinase catalytic subunit |
| PTBP1 | Polypyrimidine tract binding protein 1 |
| PKM2 | Pyruvate Kinase |
| SPHK1 | Sphingosine Kinase 1 |
| NF-κB | Nuclear Factor kappa-light- chain-enhancer of activated B cells |
| PTEN | Phosphatase and Tensin Homolog |
| EGFR | Epidermal Growth Factor Receptor |
| HER2 | Human Epidermal growth factor Receptor 2 |
| RMM1 | Ribonucleotide eductase M1 |
| SIK1 | Salt-inducible Kinase 1 |
| MST1/2 | Mammalian STE20-like protein kinase 1/2 |
| LATS1/2 | Large Tumour Suppressor 1/2 |
| SAV1 | Salvador homolog 1 |
| MOB1 | MOB kinase activator 1 |
| SMURF2 | SMAD specific E3 ubiquitin protein ligase 2 |
| VIM | Vimentin |
| GSK-3 β | Glycogen Synthase Kinase 3 beta |
| SIRT1 | Silent mating type Information Regulation 2 homolog 1 |
| ZEB1 | Zinc finger e-box binding homeobox 1 |
| FGF | Fibroblast Growth Factor |
| FGFR3 | Fibroblast Growth Factor Receptor 3 |
| ATM | Ataxia Telangiectasia Mutated protein |
| ROCK2 | Rho associated coiled-coil containing protein kinase 2 |
| MTOR | Mechanistic Target of Rapamycin |
| RRM2 | Ribonucleotide Reductase M2 |
| TUBB3 | class III b-tubulin |
| VEGF | Vascular Endothelial Growth Factor |
| MMP | Matrix Metalloproteinases |
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- World Cancer Research Fund International. Pancreatic cancer statistics. Available online: http://www.wcrf.org/int/cancer-facts-figures/data-specific-cancers/pancreatic-cancer-statistics (accessed on 11 December 2017).
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global cancer statistics, 2012. CA A Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef] [PubMed]
- GLOBCAN. Cancer Incidence, Mortality and Prevalence Worldwide, World Pancreas New Cancer Cases. Available online: http://globocan.iarc.fr/old/burden.asp?selection_pop=224900&Text-p=World&selection_cancer=23090&Text-c=Pancreas&pYear=3&type=0&window=1&submit=%C2%A0Execute (accessed on 11 December 2017).
- GLOBCAN. Cancer Incidence, Mortality and Prevalence Worldwide, World Pancreas Deaths. Available online: http://globocan.iarc.fr/old/burden.asp?selection_pop=224900&Text-p=World&selection_cancer=23090&Text-c=Pancreas&pYear=3&type=1&window=1&submit=%C2%A0Execute (accessed on 11 December 2017).
- INDIA AGAINST CANCER and NICPR. Cancer Factsheet. Available online: http://cancerindia.org.in/cancer-factsheet/ (accessed on 11 December 2017).
- Hartwig, W.; Hackert, T.; Hinz, U.; Gluth, A.; Bergmann, F.; Strobel, O.; Büchler, M.W.; Werner, J. Pancreatic cancer surgery in the new millennium: Better prediction of outcome. Ann. Surg. 2011, 254, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Van Rijswijk, R.; Jeziorski, K.; Wagener, D.T.; Van Laethem, J.-L.; Reuse, S.; Baron, B.; Wils, J.; EORTC GastroIntestinal Tract Cancer Cooperative Group. Weekly high-dose 5-fluorouracil and folinic acid in metastatic pancreatic carcinoma: A phase II study of the EORTC GastroIntestinal Tract Cancer Cooperative Group. Eur. J. Cancer 2004, 40, 2077–2081. [Google Scholar] [CrossRef] [PubMed]
- Cullinan, S.A.; Moertel, C.G.; Fleming, T.R.; Rubin, J.R.; Krook, J.E.; Everson, L.K.; Windschitl, H.E.; Twito, D.I.; Marschke, R.F.; Foley, J.F. A comparison of three chemotherapeutic regimens in the treatment of advanced pancreatic and gastric carcinoma: Fluorouracil vs fluorouracil and doxorubicin vs fluorouracil, doxorubicin, and mitomycin. JAMA 1985, 253, 2061–2067. [Google Scholar] [CrossRef]
- Androulakis, N.; Kourousis, C.; Dimopoulos, M.A.; Samelis, G.; Kakolyris, S.; Tsavaris, N.; Genatas, K.; Aravantinos, G.; Papadimitriou, C.; Karabekios, S. Treatment of pancreatic cancer with docetaxel and granulocyte colony-stimulating factor: A multicenter phase II study. J. Clin. Oncol. 1999, 17, 1779. [Google Scholar] [CrossRef] [PubMed]
- Wagener, D.T.; Verdonk, H.; Dirix, L.; Catimel, G.; Siegenthaler, P.; Buitenhuis, M.; Mathieu-Boue, A.; Verweij, J. Phase II trial of CPT-11 in patients with advanced pancreatic cancer, an EORTC early clinical trials group study. Ann. Oncol. 1995, 6, 129–132. [Google Scholar] [CrossRef]
- Oettle, H.; Post, S.; Neuhaus, P.; Gellert, K.; Langrehr, J.; Ridwelski, K.; Schramm, H.; Fahlke, J.; Zuelke, C.; Burkart, C. Adjuvant chemotherapy with gemcitabine vs observation in patients undergoing curative-intent resection of pancreatic cancer: A randomized controlled trial. Jama 2007, 297, 267–277. [Google Scholar] [CrossRef]
- Berlin, J.D.; Catalano, P.; Thomas, J.P.; Kugler, J.W.; Haller, D.G.; Benson III, A.B. Phase III study of gemcitabine in combination with fluorouracil versus gemcitabine alone in patients with advanced pancreatic carcinoma: Eastern Cooperative Oncology Group Trial E2297. J. Clin. Oncol. 2002, 20, 3270–3275. [Google Scholar] [CrossRef]
- Herrmann, R.; Bodoky, G.r.; Ruhstaller, T.; Glimelius, B.; Bajetta, E.; Schuller, J.; Saletti, P.; Bauer, J.; Figer, A.; Pestalozzi, B. Gemcitabine plus capecitabine compared with gemcitabine alone in advanced pancreatic cancer: A randomized, multicenter, phase III trial of the Swiss Group for Clinical Cancer Research and the Central European Cooperative Oncology Group. J. Clin. Oncol. 2007, 25, 2212–2217. [Google Scholar] [CrossRef]
- Ueno, H.; Ioka, T.; Ikeda, M.; Ohkawa, S.; Yanagimoto, H.; Boku, N.; Fukutomi, A.; Sugimori, K.; Baba, H.; Yamao, K. Randomized phase III study of gemcitabine plus S-1, S-1 alone, or gemcitabine alone in patients with locally advanced and metastatic pancreatic cancer in Japan and Taiwan: GEST study. J. Clin. Oncol. 2013, 31, 1640–1648. [Google Scholar] [CrossRef] [PubMed]
- Chao, Y.; Wu, C.-Y.; Wang, J.P.; Lee, R.-C.; Lee, W.-P.; Li, C.-P. A randomized controlled trial of gemcitabine plus cisplatin versus gemcitabine alone in the treatment of metastatic pancreatic cancer. Cancer Chemother. Pharmacol. 2013, 72, 637–642. [Google Scholar] [CrossRef] [PubMed]
- Stathopoulos, G.; Syrigos, K.; Aravantinos, G.; Polyzos, A.; Papakotoulas, P.; Fountzilas, G.; Potamianou, A.; Ziras, N.; Boukovinas, J.; Varthalitis, J. A multicenter phase III trial comparing irinotecan-gemcitabine (IG) with gemcitabine (G) monotherapy as first-line treatment in patients with locally advanced or metastatic pancreatic cancer. Br. J. Cancer 2006, 95, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Reni, M.; Cordio, S.; Milandri, C.; Passoni, P.; Bonetto, E.; Oliani, C.; Luppi, G.; Nicoletti, R.; Galli, L.; Bordonaro, R. Gemcitabine versus cisplatin, epirubicin, fluorouracil, and gemcitabine in advanced pancreatic cancer: A randomised controlled multicentre phase III trial. Lancet Oncol. 2005, 6, 369–376. [Google Scholar] [CrossRef]
- Poplin, E.; Feng, Y.; Berlin, J.; Rothenberg, M.L.; Hochster, H.; Mitchell, E.; Alberts, S.; O’Dwyer, P.; Haller, D.; Catalano, P. Phase III, randomized study of gemcitabine and oxaliplatin versus gemcitabine (fixed-dose rate infusion) compared with gemcitabine (30-minute infusion) in patients with pancreatic carcinoma E6201: A trial of the Eastern Cooperative Oncology Group. J. Clin. Oncol. 2009, 27, 3778–3785. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Goldstein, D.; Hamm, J.; Figer, A.; Hecht, J.R.; Gallinger, S.; Au, H.J.; Murawa, P.; Walde, D.; Wolff, R.A. Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: A phase III trial of the National Cancer Institute of Canada Clinical Trials Group. J. Clin. Oncol. 2007, 25, 1960–1966. [Google Scholar] [CrossRef]
- Von Hoff, D.D.; Ervin, T.; Arena, F.P.; Chiorean, E.G.; Infante, J.; Moore, M.; Seay, T.; Tjulandin, S.A.; Ma, W.W.; Saleh, M.N. Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N. Engl. J. Med. 2013, 369, 1691–1703. [Google Scholar] [CrossRef]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotechnol. 2007, 2, 751–760. [Google Scholar] [CrossRef]
- Wiebe, L.A. A myriad of symptoms: New approaches to optimizing palliative care of patients with advanced pancreatic cancer. Am. Soc. Clin. Oncol. Educ. Book 2012, 32, 243–248. [Google Scholar]
- Vanneman, M.; Dranoff, G. Combining immunotherapy and targeted therapies in cancer treatment. Nat. Rev. Cancer 2012, 12, 237–251. [Google Scholar] [CrossRef]
- Liu, R.; Chen, X.; Du, Y.; Yao, W.; Shen, L.; Wang, C.; Hu, Z.; Zhuang, R.; Ning, G.; Zhang, C. Serum microRNA expression profile as a biomarker in the diagnosis and prognosis of pancreatic cancer. Clin. Chem. 2012, 58, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Gusev, Y.; Jiang, J.; Nuovo, G.J.; Lerner, M.R.; Frankel, W.L.; Morgan, D.L.; Postier, R.G.; Brackett, D.J.; Schmittgen, T.D. Expression profiling identifies microRNA signature in pancreatic cancer. Int. J. Cancer 2007, 120, 1046–1054. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, J.; Chang, P.; LeBlanc, A.; Li, D.; Abbruzzesse, J.L.; Frazier, M.L.; Killary, A.M.; Sen, S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev. Res. 2009, 2, 807–813. [Google Scholar] [CrossRef] [PubMed]
- Dillhoff, M.; Liu, J.; Frankel, W.; Croce, C.; Bloomston, M. MicroRNA-21 is overexpressed in pancreatic cancer and a potential predictor of survival. J. Gastrointest. Surg. 2008, 12, 2171. [Google Scholar] [CrossRef] [PubMed]
- Greither, T.; Grochola, L.F.; Udelnow, A.; Lautenschläger, C.; Würl, P.; Taubert, H. Elevated expression of microRNAs 155, 203, 210 and 222 in pancreatic tumors is associated with poorer survival. Int. J. Cancer 2010, 126, 73–80. [Google Scholar] [CrossRef]
- O’donnell, K.A.; Wentzel, E.A.; Zeller, K.I.; Dang, C.V.; Mendell, J.T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005, 435, 839–843. [Google Scholar] [CrossRef]
- Tagawa, H.; Karube, K.; Tsuzuki, S.; Ohshima, K.; Seto, M. Synergistic action of the microRNA-17 polycistron and Myc in aggressive cancer development. Cancer Sci. 2007, 98, 1482–1490. [Google Scholar] [CrossRef]
- Shi, X.; Liu, S.; Kleeff, J.; Friess, H.; Büchler, M.W. Acquired resistance of pancreatic cancer cells towards 5-Fluorouracil and gemcitabine is associated with altered expression of apoptosis-regulating genes. Oncology 2002, 62, 354–362. [Google Scholar] [CrossRef]
- Yu, S.; Lu, Z.; Liu, C.; Meng, Y.; Ma, Y.; Zhao, W.; Liu, J.; Yu, J.; Chen, J. miRNA-96 suppresses KRAS and functions as a tumor suppressor gene in pancreatic cancer. Cancer Res. 2010, 70, 6015–6025. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, M.; Wang, H.; Fisher, W.E.; Lin, P.H.; Yao, Q.; Chen, C. Profiling of 95 microRNAs in pancreatic cancer cell lines and surgical specimens by real-time PCR analysis. World J. Surg. 2009, 33, 698. [Google Scholar] [CrossRef]
- Li, Y.; VandenBoom, T.G.; Kong, D.; Wang, Z.; Ali, S.; Philip, P.A.; Sarkar, F.H. Up-regulation of miR-200 and let-7 by natural agents leads to the reversal of epithelial-to-mesenchymal transition in gemcitabine-resistant pancreatic cancer cells. Cancer Res. 2009, 69, 6704–6712. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Wu, H.; Liu, X.; Evans, B.R.; Medina, D.J.; Liu, C.-G.; Yang, J.-M. Role of MicroRNA miR-27a and miR-451 in the regulation of MDR1/P-glycoprotein expression in human cancer cells. Biochem. Pharmacol. 2008, 76, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Kovalchuk, O.; Filkowski, J.; Meservy, J.; Ilnytskyy, Y.; Tryndyak, V.P.; Vasyl’F, C.; Pogribny, I.P. Involvement of microRNA-451 in resistance of the MCF-7 breast cancer cells to chemotherapeutic drug doxorubicin. Mol. Cancer Ther. 2008, 7, 2152–2159. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Dong, C.; Ji, C. MicroRNA and drug resistance. Cancer Gene Ther. 2010, 17, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, B.B. MicroRNA-in drug resistance. Oncoscience 2014, 1, 3. [Google Scholar]
- Zheng, T.; Wang, J.; Chen, X.; Liu, L. Role of microRNA in anticancer drug resistance. Int. J. Cancer 2010, 126, 2–10. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Ahmad, A.; Banerjee, S.; Padhye, S.; Dominiak, K.; Schaffert, J.M.; Wang, Z.; Philip, P.A.; Sarkar, F.H. Gemcitabine sensitivity can be induced in pancreatic cancer cells through modulation of miR-200 and miR-21 expression by curcumin or its analogue CDF. Cancer Res. 2010, 70, 3606–3617. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Y.; Kong, D.; Banerjee, S.; Ahmad, A.; Azmi, A.S.; Ali, S.; Abbruzzese, J.L.; Gallick, G.E.; Sarkar, F.H. Acquisition of epithelial-mesenchymal transition phenotype of gemcitabine-resistant pancreatic cancer cells is linked with activation of the notch signaling pathway. Cancer Res. 2009, 69, 2400–2407. [Google Scholar] [CrossRef]
- Hong, S.P.; Wen, J.; Bang, S.; Park, S.; Song, S.Y. CD44-positive cells are responsible for gemcitabine resistance in pancreatic cancer cells. Int. J. Cancer 2009, 125, 2323–2331. [Google Scholar] [CrossRef]
- Jayaraj, R.; Kumarasamy, C. Systematic review and meta-analysis of cancer studies evaluating diagnostic test accuracy and prognostic values: Approaches to improve clinical interpretation of results. Cancer Manag. Res. 2018, 10, 4669–4670. [Google Scholar] [CrossRef]
- Jayaraj, R.; Kumarasamy, C.; Ramalingam, S.; Devi, A. Systematic review and meta-analysis of risk-reductive dental strategies for medication related osteonecrosis of the jaw among cancer patients: Approaches and strategies. Oral Oncol. 2018, 86, 312–313. [Google Scholar] [CrossRef] [PubMed]
- Sabarimurugan, S.; Royam, M.M.; Das, A.; Das, S.; Gothandam, K.M.; Jayaraj, R. Systematic Review and Meta-analysis of the Prognostic Significance of miRNAs in Melanoma Patients. Mol. Diagn. Ther. 2018, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Jayaraj, R.; Kumarasamy, C.; Madhav, M.R.; Pandey, V.; Sabarimurugan, S.; Ramesh, N.; Gothandam, K.M.; Baxi, S. Comment on “Systematic Review and Meta-Analysis of Diagnostic Accuracy of miRNAs in Patients with Pancreatic Cancer”. Dis. Markers 2018, 2018, 2. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Group, P. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Stroup, D.F.; Berlin, J.A.; Morton, S.C.; Olkin, I.; Williamson, G.D.; Rennie, D.; Moher, D.; Becker, B.J.; Sipe, T.A.; Thacker, S.B. Meta-analysis of observational studies in epidemiology: A proposal for reporting. JAMA 2000, 283, 2008–2012. [Google Scholar] [CrossRef] [PubMed]
- Jayaraj, R.; Kumarasamy, C. Comment on “Prognostic biomarkers for oral tongue squamous cell carcinoma: A systematic review and meta-analysis”. Br. J. Cancer 2018, 118, e11. [Google Scholar] [CrossRef] [PubMed]
- Jayaraj, R.; Kumarasamy, C. Comment on “Survival for HPV-positive oropharyngeal squamous cell carcinoma with surgical versus non-surgical treatment approach: A systematic review and meta-analysis”. Oral Oncol. 2019, 90, 137. [Google Scholar] [CrossRef]
- Jayaraj, R.; Kumarasamy, C.; Sabarimurugan, S.; Baxi, S. Commentary: Blood-Derived microRNAs for Pancreatic Cancer Diagnosis: A Narrative Review and Meta-Analysis. Front. Physiol. 2018, 9, 1896. [Google Scholar] [CrossRef]
- Jayaraj, R.; Kumarasamy, C. Conceptual interpretation of analysing and reporting of results on systematic review and meta-analysis of optimal extent of lateral neck dissection for well-differentiated thyroid carcinoma with metastatic lateral neck lymph nodes. Oral Oncol. 2019, 89, 153. [Google Scholar] [CrossRef]
- Higgins, J.; Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 2002, 21, 1539–1558. [Google Scholar] [CrossRef]
- Yang, R.-M.; Zhan, M.; Xu, S.-W.; Long, M.-M.; Yang, L.-H.; Chen, W.; Huang, S.; Liu, Q.; Zhou, J.; Zhu, J. miR-3656 expression enhances the chemosensitivity of pancreatic cancer to gemcitabine through modulation of the RHOF/EMT axis. Cell Death Dis. 2017, 8, e3129. [Google Scholar] [CrossRef] [PubMed]
- Xiao, G.; Wang, X.; Yu, Y. CXCR4/Let-7a Axis Regulates Metastasis and Chemoresistance of Pancreatic Cancer Cells Through Targeting HMGA2. Cell. Physiol. Biochem. 2017, 43, 840–851. [Google Scholar] [CrossRef] [PubMed]
- Hiramoto, H.; Muramatsu, T.; Ichikawa, D.; Tanimoto, K.; Yasukawa, S.; Otsuji, E.; Inazawa, J. miR-509-5p and miR-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer. Sci. Rep. 2017, 7, 4002. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, A.K.; Mondal, G.; Kumar, V.; Kattel, K.; Mahato, R.I. Chemosensitization and inhibition of pancreatic cancer stem cell proliferation by overexpression of microRNA-205. Cancer Lett. 2017, 402, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Liu, B.; Qian, J.; Wu, G.; Li, J.; Ma, Z. miR-153 enhances the therapeutic effect of gemcitabine by targeting Snail in pancreatic cancer. Acta Biochim. Biophys. Sin. 2017, 49, 520–529. [Google Scholar] [CrossRef] [PubMed]
- Mikamori, M.; Yamada, D.; Eguchi, H.; Hasegawa, S.; Kishimoto, T.; Tomimaru, Y.; Asaoka, T.; Noda, T.; Wada, H.; Kawamoto, K. MicroRNA-155 controls exosome synthesis and promotes gemcitabine resistance in pancreatic ductal adenocarcinoma. Sci. Rep. 2017, 7, 42339. [Google Scholar] [CrossRef]
- Hu, H.; He, Y.; Wang, Y.; Chen, W.; Hu, B.; Gu, Y. micorRNA-101 silences DNA-PKcs and sensitizes pancreatic cancer cells to gemcitabine. Biochem. Biophys. Res. Commun. 2017, 483, 725–731. [Google Scholar] [CrossRef]
- Amponsah, P.S.; Fan, P.; Bauer, N.; Zhao, Z.; Gladkich, J.; Fellenberg, J.; Herr, I. microRNA-210 overexpression inhibits tumor growth and potentially reverses gemcitabine resistance in pancreatic cancer. Cancer Lett. 2017, 388, 107–117. [Google Scholar] [CrossRef]
- Li, C.; Zhao, Z.; Zhou, Z.; Liu, R. Linc-ROR confers gemcitabine resistance to pancreatic cancer cells via inducing autophagy and modulating the miR-124/PTBP1/PKM2 axis. Cancer Chemother. Pharmacol. 2016, 78, 1199–1207. [Google Scholar] [CrossRef]
- Li, J.; Wu, H.; Li, W.; Yin, L.; Guo, S.; Xu, X.; Ouyang, Y.; Zhao, Z.; Liu, S.; Tian, Y. Downregulated miR-506 expression facilitates pancreatic cancer progression and chemoresistance via SPHK1/Akt/NF-κB signaling. Oncogene 2016, 35, 5501. [Google Scholar] [CrossRef]
- Gu, J.; Wang, D.; Zhang, J.; Zhu, Y.; Li, Y.; Chen, H.; Shi, M.; Wang, X.; Shen, B.; Deng, X. GFRα2 prompts cell growth and chemoresistance through down-regulating tumor suppressor gene PTEN via Mir-17-5p in pancreatic cancer. Cancer Lett. 2016, 380, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; Shivapurkar, N.; Wu, Z.; Hwang, J.J.; Pishvaian, M.J.; Weiner, L.M.; Ley, L.; Zhou, D.; Zhi, X.; Wellstein, A. Circulating microRNA profile predicts disease progression in patients receiving second-line treatment of lapatinib and capecitabine for metastatic pancreatic cancer. Oncology Lett. 2016, 11, 1645–1650. [Google Scholar] [CrossRef] [PubMed]
- Fan, P.; Liu, L.; Yin, Y.; Zhao, Z.; Zhang, Y.; Amponsah, P.S.; Xiao, X.; Bauer, N.; Abukiwan, A.; Nwaeburu, C.C. MicroRNA-101-3p reverses gemcitabine resistance by inhibition of ribonucleotide reductase M1 in pancreatic cancer. Cancer Lett. 2016, 373, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.; Li, Z.; Wang, X.; Xu, P.; Zhao, L.; Qian, J. MiR-125a regulates chemo-sensitivity to gemcitabine in human pancreatic cancer cells through targeting A20. Acta Biochim. Biophys. Sin. 2016, 48, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.-G.; Dong, S.-X.; Han, P.; Qi, J. miR-203 promotes proliferation, migration and invasion by degrading SIK1 in pancreatic cancer. Oncol. Rep. 2016, 35, 1365–1374. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wang, M.; Xu, S.; Guo, X.; Jiang, J. Upregulation of miR-181c contributes to chemoresistance in pancreatic cancer by inactivating the Hippo signaling pathway. Oncotarget 2015, 6, 44466. [Google Scholar] [CrossRef] [PubMed]
- Miyamae, M.; Komatsu, S.; Ichikawa, D.; Kawaguchi, T.; Hirajima, S.; Okajima, W.; Ohashi, T.; Imamura, T.; Konishi, H.; Shiozaki, A. Plasma microRNA profiles: Identification of miR-744 as a novel diagnostic and prognostic biomarker in pancreatic cancer. Br. J. Cancer 2015, 113, 1467. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.-L.; Zhang, J.-H.; Wu, X.-Z.; Yan, T.; Lv, W. miR-15b promotes epithelial-mesenchymal transition by inhibiting SMURF2 in pancreatic cancer. Int. J. Oncol. 2015, 47, 1043–1053. [Google Scholar] [CrossRef]
- Yu, C.; Wang, M.; Chen, M.; Huang, Y.; Jiang, J. Upregulation of microRNA-138-5p inhibits pancreatic cancer cell migration and increases chemotherapy sensitivity. Mol. Med. Rep. 2015, 12, 5135–5140. [Google Scholar] [CrossRef]
- Liang, C.; Yu, X.-J.; Guo, X.-Z.; Sun, M.-H.; Wang, Z.; Song, Y.; Ni, Q.-X.; Li, H.-Y.; Mukaida, N.; Li, Y.-Y. MicroRNA-33a-mediated downregulation of Pim-3 kinase expression renders human pancreatic cancer cells sensitivity to gemcitabine. Oncotarget 2015, 6, 14440. [Google Scholar] [CrossRef]
- Liu, Y.; Li, X.; Zhu, S.; Zhang, J.; Yang, M.; Qin, Q.; Deng, S.; Wang, B.; Tian, K.; Liu, L. Ectopic expression of miR-494 inhibited the proliferation, invasion and chemoresistance of pancreatic cancer by regulating SIRT1 and c-Myc. Gene Ther. 2015, 22, 729. [Google Scholar] [CrossRef] [PubMed]
- Meidhof, S.; Brabletz, S.; Lehmann, W.; Preca, B.T.; Mock, K.; Ruh, M.; Schüler, J.; Berthold, M.; Weber, A.; Burk, U. ZEB1-associated drug resistance in cancer cells is reversed by the class I HDAC inhibitor mocetinostat. EMBO Mol. Med. 2015, 7, 831–847. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhao, L.; Ischenko, I.; Bao, Q.; Schwarz, B.; Nieß, H.; Wang, Y.; Renner, A.; Mysliwietz, J.; Jauch, K.-W. Antisense inhibition of microRNA-21 and microRNA-221 in tumor-initiating stem-like cells modulates tumorigenesis, metastasis, and chemotherapy resistance in pancreatic cancer. Target. Oncol. 2015, 10, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, X.; Yu, C.; Wang, M.; Peng, F.; Xiao, J.; Tian, R.; Jiang, J.; Sun, C. MicroRNA-100 regulates pancreatic cancer cells growth and sensitivity to chemotherapy through targeting FGFR3. Tumor Bio. 2014, 35, 11751–11759. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Wang, T.; Cao, Z.; Huang, H.; Li, J.; Liu, W.; Liu, S.; You, L.; Zhou, L.; Zhang, T. MiR-497 downregulation contributes to the malignancy of pancreatic cancer and associates with a poor prognosis. Oncotarget 2014, 5, 6983. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, S.; Eguchi, H.; Nagano, H.; Konno, M.; Tomimaru, Y.; Wada, H.; Hama, N.; Kawamoto, K.; Kobayashi, S.; Nishida, N. MicroRNA-1246 expression associated with CCNG2-mediated chemoresistance and stemness in pancreatic cancer. Br. J. Cancer 2014, 111, 1572. [Google Scholar] [CrossRef] [PubMed]
- Lai, I.-L.; Chou, C.-C.; Lai, P.-T.; Fang, C.-S.; Shirley, L.A.; Yan, R.; Mo, X.; Bloomston, M.; Kulp, S.K.; Bekaii-Saab, T. Targeting the Warburg effect with a novel glucose transporter inhibitor to overcome gemcitabine resistance in pancreatic cancer cells. Carcinogenesis 2014, 35, 2203–2213. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Song, W.-F.; Wang, L.; Huang, W.-Y.; Cai, X.; Cui, J.-J.; Wang, L.-W. MiR-21 upregulation induced by promoter zone histone acetylation is associated with chemoresistance to gemcitabine and enhanced malignancy of pancreatic cancer cells. Asian Pac. J. Cancer Prev. 2013, 14, 7529–7536. [Google Scholar] [CrossRef]
- Peng, F.; Jiang, J.; Yu, Y.; Tian, R.; Guo, X.; Li, X.; Shen, M.; Xu, M.; Zhu, F.; Shi, C. Direct targeting of SUZ12/ROCK2 by miR-200b/c inhibits cholangiocarcinoma tumourigenesis and metastasis. Br. J. Cancer 2013, 109, 3092. [Google Scholar] [CrossRef]
- Nagano, H.; Tomimaru, Y.; Eguchi, H.; Hama, N.; Wada, H.; Kawamoto, K.; Kobayashi, S.; Mori, M.; Doki, Y. MicroRNA-29a induces resistance to gemcitabine through the Wnt/β-catenin signaling pathway in pancreatic cancer cells. Int. J. Oncol. 2013, 43, 1066–1072. [Google Scholar] [CrossRef]
- Wei, F.; Liu, Y.; Guo, Y.; Xiang, A.; Wang, G.; Xue, X.; Lu, Z. miR-99b-targeted mTOR induction contributes to irradiation resistance in pancreatic cancer. Mol. Cancer 2013, 12, 81. [Google Scholar] [CrossRef] [PubMed]
- Iwagami, Y.; Eguchi, H.; Nagano, H.; Akita, H.; Hama, N.; Wada, H.; Kawamoto, K.; Kobayashi, S.; Tomokuni, A.; Tomimaru, Y. miR-320c regulates gemcitabine-resistance in pancreatic cancer via SMARCC1. Br. J. Cancer 2013, 109, 502. [Google Scholar] [CrossRef] [PubMed]
- Bhutia, Y.D.; Hung, S.W.; Krentz, M.; Patel, D.; Lovin, D.; Manoharan, R.; Thomson, J.M.; Govindarajan, R. Differential processing of let-7a precursors influences RRM2 expression and chemosensitivity in pancreatic cancer: Role of LIN-28 and SET oncoprotein. PLoS ONE 2013, 8, e53436. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zhuang, L.; Zhang, J.; Fan, J.; Luo, J.; Chen, H.; Wang, K.; Liu, L.; Chen, Z.; Meng, Z. The serum miR-21 level serves as a predictor for the chemosensitivity of advanced pancreatic cancer, and miR-21 expression confers chemoresistance by targeting FasL. Mol. Oncol. 2013, 7, 334–345. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Chitkara, D.; Kumar, V.; Behrman, S.W.; Mahato, R.I. miRNA profiling in pancreatic cancer and restoration of chemosensitivity. Cancer Lett. 2013, 334, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Brabletz, S.; Bajdak, K.; Meidhof, S.; Burk, U.; Niedermann, G.; Firat, E.; Wellner, U.; Dimmler, A.; Faller, G.; Schubert, J. The ZEB1/miR-200 feedback loop controls Notch signalling in cancer cells. EMBO J. 2011, 30, 770–782. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Almhanna, K.; Chen, W.; Philip, P.A.; Sarkar, F.H. Differentially expressed miRNAs in the plasma may provide a molecular signature for aggressive pancreatic cancer. Am. J. Transl. Res. 2011, 3, 28. [Google Scholar]
- Hwang, J.-H.; Voortman, J.; Giovannetti, E.; Steinberg, S.M.; Leon, L.G.; Kim, Y.-T.; Funel, N.; Park, J.K.; Kim, M.A.; Kang, G.H. Identification of microRNA-21 as a biomarker for chemoresistance and clinical outcome following adjuvant therapy in resectable pancreatic cancer. PloS ONE 2010, 5, e10630. [Google Scholar] [CrossRef]
- Giovannetti, E.; Funel, N.; Peters, G.J.; Del Chiaro, M.; Erozenci, L.A.; Vasile, E.; Leon, L.G.; Pollina, L.E.; Groen, A.; Falcone, A. MicroRNA-21 in pancreatic cancer: Correlation with clinical outcome and pharmacologic aspects underlying its role in the modulation of gemcitabine activity. Cancer Res. 2010, 70, 4528–4538. [Google Scholar] [CrossRef]
- Moriyama, T.; Ohuchida, K.; Mizumoto, K.; Yu, J.; Sato, N.; Nabae, T.; Takahata, S.; Toma, H.; Nagai, E.; Tanaka, M. MicroRNA-21 modulates biological functions of pancreatic cancer cells including their proliferation, invasion, and chemoresistance. Mol. Cancer Ther. 2009, 8, 1067–1074. [Google Scholar] [CrossRef]
- Xiong, G.; Huang, H.; Feng, M.; Yang, G.; Zheng, S.; You, L.; Zheng, L.; Hu, Y.; Zhang, T.; Zhao, Y. MiR-10a-5p targets TFAP2C to promote gemcitabine resistance in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 2018, 37, 76. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, A.; Chattopadhyay, D.; Chakrabarti, G. miR-17-5p downregulation contributes to paclitaxel resistance of lung cancer cells through altering beclin1 expression. PLoS ONE 2014, 9, e95716. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Ohuchida, K.; Mizumoto, K.; Fujita, H.; Nakata, K.; Tanaka, M. MicroRNA miR-17-5p is overexpressed in pancreatic cancer, associated with a poor prognosis, and involved in cancer cell proliferation and invasion. Cancer Biol. Ther. 2010, 10, 748–757. [Google Scholar] [CrossRef] [PubMed]
- Garajová, I.; Le Large, T.Y.; Frampton, A.E.; Rolfo, C.; Voortman, J.; Giovannetti, E. Molecular mechanisms underlying the role of microRNAs in the chemoresistance of pancreatic cancer. BioMed. Res. Int. 2014. [Google Scholar] [CrossRef] [PubMed]
- Prislei, S.; Martinelli, E.; Mariani, M.; Raspaglio, G.; Sieber, S.; Ferrandina, G.; Shahabi, S.; Scambia, G.; Ferlini, C. MiR-200c and HuR in ovarian cancer. BMC Cancer 2013, 13, 72. [Google Scholar] [CrossRef] [PubMed]
- Cochrane, D.R.; Spoelstra, N.S.; Howe, E.N.; Nordeen, S.K.; Richer, J.K. MicroRNA-200c mitigates invasiveness and restores sensitivity to microtubule-targeting chemotherapeutic agents. Mol. Cancer Ther. 2009, 8, 1055–1066. [Google Scholar] [CrossRef] [PubMed]
- Jayaraj, R.; Kumarasamy, C.; Piedrafita, D. Systematic review and meta-analysis protocol for Fasciola DNA vaccines. Online J. Vet. Res. 2018, 22, 517. [Google Scholar]
- Kumarasamy, C.; Devi, A.; Jayaraj, R. Prognostic value of microRNAs in head and neck cancers: A systematic review and meta-analysis protocol. Syst. Rev. 2018, 7, 150. [Google Scholar] [CrossRef] [PubMed]
| S.No | Author | Ethnicity | Period of Study | Drug | No. of Samples (Cancer/Normal) | Cell Culture Type | Resistant Cells | miRNA | miRNA Profiling Platform | Pathways Associated with/Gene |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | Yang, R. et al. (2017) [55] | Chinese | 2013–2016 | GEM | 157/157 | Human PC cell lines Capan-2, HPAC, SW1990, PANC-1, CFPAC-1, BXPC-3, ASPC-1, PATU-8988, HPDE6-C7 and HPNE | PANC-1-GR and BXPC-3 GR | 3656 | Taqman microRNA Reverse Transcription kits (Thermo Fisher Scientific, Dreieich, Germany) | Ras Homolog Family Member F (RHOF)/Epithelial–mesenchymal transition (EMT) |
| 2 | Xiao, G. et al. (2017) [56] | Chinese | 2015–2016 | GEM | 15/15 | HPDE6-C7, PC cell lines Panc-1 and BxPc3 | NM | Let-7a | TAKARA PrimeScript Kit | C-X-C chemokine receptor type 4 (CXCR4)/let-7a/High-mobility group AT-hook 2 (HMGA2) |
| 3 | Hiramoto, H. et al. (2017) [57] | Japanese | 2000–2011 | GEM | 50 | Panc1, KP4-4, SU.86.86, BxPC3 and MDA-MB-231 | NM | 509-5p, 1243 | Custom Taqman miRNA Assays kit (Applied Biosystems, San Diego, CA, USA) | E-cadherin |
| 4 | Chaudhary, A.K. et al. (2017) [58] | American | NM | GEM | NM | HPDE | GEM-resistant MIA PaCa-2R cells | 205-5p | SYBR Green-based pathway-focused miScript miRNA PCR Array (catalog number 102ZF, Qiagen, MD, USA) using Roche Light Cycler 480® (Roche, Indianapolis, IN, USA) | K-ras, Caveolin-1, and Ki-67 |
| 5 | Liu. F. et al. (2017) [59] | Chinese | January 2010–December 2014 | GEM | 87 | BxPC-3, Panc-1, Capan-2, SW1990, Paca-2, AsPc-1, and CFPAC-1, HEK293T and HPDE | Capan-2, Panc-1, and AsPc-1 | 153 | SYBR Premix Ex Taq (TaKaRa, Dalian, China) and run with an Applied Biosystems ViiATM 7 Real-Time PCR System (Applied Biosystems) | Snail |
| 6 | Mikamori, M. et al. (2017) [60] | Japanese | March 2007–August 2015 | GEM | 45 | Panc1, MiaPaCa2, and PSN1 cell lines | Panc1-GR1, -GR3, and -GR4 cells | 155 | TaqMan MicroRNA Assays (Applied Biosystems) and the ABI7900HT system (Applied Biosystems) | Anti-apoptotic (RAB27B) |
| 7 | Hu, H. et al. (2016) [61] | Chinese | NM | GEM | 15/15 | PANC-1 | NM | 101 | TaqMan microRNA assay using ABI-7300 Real-Time machine (Shanghai, China) | DNA-dependent protein kinase catalytic subunit (DNA-PKcs) |
| 8 | Amponsah, P. et al. (2016) [62] | Deutsch | NM | GEM | 92/5 | ASAN-PaCa, BxPC-3, AsPC-1 and MIA-PaCa2 | Bx-GEM | 210 | Human HT-12 v4 Expression Bead Chip Kit or the Human miR Microarray (Release 19.0) | ABCC5 |
| 9 | Li, C. et al. (2016) [63] | Chinese | 2013-2015 | GEM | 31/31 | HPDE6, PANC-1, MIAPaCa-2 and SW1990 cells | NM | 124 | TaqMan microRNA assays (Applied Biosystems) | miR-124/polypyrimidine tract binding protein 1 (PTBP1)/Pyruvate kinase (PKM2) |
| 10 | Li, J. et al. (2016) [64] | Chinese | NM | GEM | 84/20 | HPC-Y5, AsPC-1, PANC-1, BxPC-3, Hs766t and CFPAC-1 | NM | 506 | Agilent Array | Sphingosine kinase 1 (SPHK1)/Protein kinase B(Akt)/nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) |
| 11 | Gu, J. et al. (2016) [65] | Chinese | 2008–2010 | GEM | 58 | PanC-1, Mia Paca-2 and HEK-293T | NM | 17-5p | SYBR Green Mix (Roche) using All-in-One miRNA qPCR Detection Kit (GeneCopoeia, Rock, MD, USA) | Phosphatase and tensin homolog (PTEN) |
| 12 | Tian, X. et al. (2016) [66] | American | March 2009–September 2013 | GEM, Lapatinib, and Capecitabine | 17 | PANC-1, MIA PaCa-2 and BXCP-3 cell lines | NM | 7, 21, 210, 221 | RT2 miRNA first strand kit (Qiagen, Inc.) and Applied Biosystems 7900HT Fast Real-Time PCR system (Thermo Fisher Scientifc, Inc., Waltham, MA, USA) | Epidermal growth factor receptor (EGFR)1 and human epidermal growth factor receptor (HER)2 pathways |
| 13 | Fan, P. et al. (2016) [67] | Deutsch | NM | GEM | 21 | ASAN-PaCa, AsPC-1, PANC-1, MIA-PaCa2 and BxPC-3 | Bx-GEM | 101-3p | Human HT-12 v4 Expression Bead Chip Kit or the Human miR Microarray (Release 19.0). | Ribonucleotide reductase M1 (RRM1) |
| 14 | Yao, J. et al. (2016) [68] | Chinese | NM | GEM | 26 | SW1990 and HEK293T cells | SW1990GZ | 125a | TaqMan MicroRNA Reverse Transcription Kit (Takara), miRscript SYBR Green PCR Kit and SYBR Green PCR Kit (Takara, Dalian, China) | TNF Alpha-Induced Protein 3 (A20) |
| 15 | Ren, Z. et al. (2016) [69] | Chinese | NM | GEM | 10/10 | L3.6pl, BxPC-3, CFPAC, MiaPaCa-2, ASPC-1, PANC-1, MPanc96, HPAC, SU86.86 and HS766T | NM | 203 | mirVana RT-qPCR miRNA Detection kit (cat no. AM7659; Ambion, Austin, TX, USA) | Salt-inducible kinase 1 (SIK1) |
| 16 | Chen, M. et al. (2015) [70] | Chinese | 2008–2011 | GEM | 124/10 | PANC-1 and BXPC3 | NM | 181c | miRNA-specific TaqMan MiRNA Assay Kit (Applied Biosystems). | Mammalian STE20-like protein kinase 1/2 (MST1/2), and large tumour suppressor 1/2 (LATS1/2), together with the adaptor proteins Salvador homolog 1 (SAV1) and MOB kinase activator 1 (MOB1) (Hippo signalling pathway) |
| 17 | Miyamae, M. et al. (2015) [71] | Japanese | January 2010–April 2013 | GEM | 94/68 | PK-45H, PANC-1, PK-59, KP4-1, and PK-1 | NM | 550a, 557, 575, 615-5p, 675, 744 | 3D-Gene miRNA microarray platform (Toray Industries, Kamakura, Japan and human TaqMan MicroRNA Assay Kit (Applied Biosystems, Foster City, CA, USA) | NM |
| 18 | Zhang, W. et al. (2015) [72] | Chinese | NM | GEM | 19 | HPAC, BxPC-3, Colo357, and L3.6pl | ASPC-1, Panc-1 and MiaPaCa-2 | 15b, 155, 212 | mirVana qRT-PCR miRNA detection kit (Ambion) | SMAD specific E3 ubiquitin protein ligase 2 (SMURF2) |
| 19 | Yu, C. et al. (2015) [73] | Chinese | 2013–2014 | 5-FU | 18 | AsPC-1, BxPc-3, Capan-1, Capan-2, CFPAC-1, PANC-1, MIA PaCa-2 & SW1990 | NM | 138-5p | Fluorescence-activated cell sorting (FACSnCanto II flow cytometer; BD Biosciences, San Jose, CA, USA) | Vimentin (VIM) |
| 20 | Liang, C. et al. (2015) [74] | Chinese | 2010–2012 | GEM | 106 | PCI35 & PCI55, SW1990, MiaPaca-2, PANC-1, BxPC-3, Capan-1 | NM | 33a | NM | AKT/Gsk-3β/β-catenin pathway |
| 21 | Liu, Y. et al. (2015) [75] | Chinese | 2007–2010 | 5-FU, GEM | 86/41 | AsPC-1, BXPC-3, SW1990, MIAPaCa-2, PANC-1 & HPDE | NM | 494 | NM | miR-494/c-Myc/SIRT1 pathway |
| 22 | Meidhof, S. et al. (2015) [76] | Deutsch | NM | GEM | 27/27 | Panc-1, MDA-MB-231, BxPC3, H358, DU-145, hPaca-1 and hPaca-2 | BxPC3 GEM-resistant cells, Tarceva-resistant H358 cells | 203 | Roche LightCycler 480 | ZEB-1 |
| 23 | Zhao, Y. et al. (2015) [77] | Deutsch | NM | GEM | 28/28 | L3.6pl | L3.6pl - GemR | 21,221 | miScript SYBR® Green PCR Kit (Qiagen, USA) | NM |
| 24 | Li, Z. et al. (2014) [78] | Chinese | 2013-2014 | GEM | 23/23 | AsPC1, BxPc-3, Capan-1, Capan-2, CFPAC-1, PANC-1, MIA PaCa-2, SW1990 | NM | 100 | TaqMan miRNA Assay (Applied Biosystems) | FGFR3 |
| 25 | Xu, J. et al. (2014) [79] | Chinese | NM | GEM | 87 | SW1990, MiaPaCa-2 | SW1990/GEM | 497 | NM | FGF/FGFR signalling pathway |
| 26 | Hasegawa, S. et al. (2014) [80] | Japanese | 2007-2010 | 5-FU, GEM | 24 | Panc1-P, Panc1-GR | Panc-1GemR | 1246 | Comparative CT method | CCNG2 |
| 27 | Lai, I.-L. et al. (2014) [81] | Americans | NM | GEM | NM | Panc-1, AsPC-1 and BxPC-3 | Panc-1GemR, BxPC3GemR and AsPC-1GemR | 520f | Bio-Rad CFX Manager 2.1 detection system and miScript PCR starter kit (Qiagen) | ATM/ATR checkpoint pathway |
| 28 | Song, W.-F. et al. (2013) [82] | Chinese | 2010–2012 | GEM | 41 | BxPc3, HPAF, HPAC, Capan, PANC-1 and PL-45 cell lines | HPAC and PANC-1/GEM | 21 | Specific Taqman MicroRNA assays (Applied Biosystems) | PTEN/Akt pathway |
| 29 | Peng, F. et al. (2013) [83] | Chinese | 2010–2011 | 5-FU | 14 | TFK-1, QBC939 cell line | NM | 220b, 200c and 429 | mirVana miRNA Isolation Kit (Ambion, Austin, TX, USA), Agilent Human miRNA Microarray Kit (V2) (Agilent Inc, Santa Clara, CA, USA) for analysis. | SUZ12, ROCK2 direct targets |
| 30 | Nagano, H. et al. (2013) [84] | Japanese | September 1999–February 2004 | GEM | 18 | MIAPaCa-2, PSN-1, BxPC-3, Panc-1 | NM | 29a | TRIzol agent (Invitrogen, Carlsbad, CA, USA) | Wnt/β-catenin signaling pathway |
| 31 | Wei, F. et al. (2013) [85] | Chinese | NM | Radiation and AZD8055 | NM | PANC-1, Capan-2, BxPC-3 | NM | 99b | NM | mTOR |
| 32 | Iwagami, Y. et al. (2013) [86] | Japanese | 1992–2008 | GEM | 66 | MiaPaCa2 and PSN1 | MiaPaCa2-RGs, PSN1-RGs | 320c | NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA | SMARCC1 mediated anti-cancer effect of GEM |
| 33 | Bhutia, Y.D. et al. (2013) [87] | Americans | NM | GEM | 10/2 | MIA PaCa-2 | L3.6pl and Capan-1/GEM | let-7a | miRNA Isolation Kit and the TaqManH MicroRNA Reverse Transcription Kit (Applied Biosystems) | RRM2 |
| 34 | Wang, P. et al. (2012) [88] | Chinese | Cohort1: 2003–2005 Cohort2: 2009–2010 | GEM | NM | Panc-1, BxPC3 | NM | 21 | RecoverAll Total Nucleic Acid Isolation Kit (Ambion) | FasL/Fas pathway |
| 35 | Singh, S. et al. (2013) [89] | NM | NM | GEM | NM | MIA PaCa-2 | MI PaCa-2/GEM | 7, 146, 205 | SYBR Green dye universal master mix on a Light Cycler 480 (Roche, Indianapolis) | Class III b-tubulin (TUBB3) |
| 36 | Brabletz, S. et al. (2011) [90] | NM | NM | GSI | NM | Panc1, HPAF2, MCF7, MiaPaCa2 | NM | 200 | Pfu Ultra Hotstart 2 Master Mix (Stratagene, Santa Clara) | Notch signaling |
| 37 | Ali, S. et al. (2010) [91] | NM | NM | GEM, OHP, tarceva | 50/10 | MIAPaCa-2, AsPc-1 | MIAPaCa-GR (GEM resistant), AsPc-1OR (oxaliplatin resistant), MIAPaCa-GTR, AsPc-1GTR (GEM and tarceva resistant) | 21, 146a, 200b, 200c, 221, let-7b and let-7d | TaqMan MicroRNA Assay kit (Applied Biosystems) | NM |
| 38 | Hwang, J.-H. et al. (2010) [92] | Korean, Italian | 1999–2007 and 2001–2004 | GEM, 5-FU | 127 | BxPc3, HPAF-II, HPAC, PANC1, PL45 | NM | 21 | TaqMan-microRNA assays and the 7900 HT-Fast RealTime PCR (Applied Biosystems, Foster City, CA, USA) | NM |
| 39 | Giovannetti, E. et al. (2010) [93] | Deutsch | 2001–2004 | GEM | 81 | hTERT-HPNE, Hhs27, LPc006, LPc033, LPc067, LPc111, LPc167, PP437 | NM | 21 | 7500HT sequence detection system (Applied Biosystems) | PTEN and PI3K-Akt pathway |
| 40 | Moriyama, T. et al. (2009) [94] | Japanese | 2000–2008 | GEM | 25/25 | AsPC-1, KP-1N, KP-2, KP-3, PANC-1, SUIT-2 MIA PaCa-2, CAPAN-1, CAPAN-2, CFPAC-1, H48N, HS766T, SW1990, NOR-P1 | NM | 21 | mirVana qRT-PCR miRNA Detection Kit, and mirVana Primer Sets (all from Ambion) | VEGF and MMP-2 and MMP-9 |
| 41 | Xiong G et al. (2018) [95] | Chinese | NM | GEM | 90/90 | AsPC-1, BxPC-3, MiaPaCa-2, PANC-1, Su86. 86, T3M4 | AsPC-1-Gem | 10a-5p | Genepharma (Shanghai, China) | TFPA2C |
| 42 | Sun, D. et al. (2018) | Chinese | January 2007–December 2015 | GEM | 87/8 | BxPC-3, PANC-1, AsPC-1, SW1990, Capan-1, Capan-2, CFPAC-1 and MIA PaCa-2 | NM | 374b-5p | LightCycler® 480 SYBR-Green I Master (Roche Diagnostics, Basel, Switzerland). | BIAPRC-3 and XIAP |
| 43 | You, L. et al. (2018) | Chinese | NM | GEM | 10 | 293T, MIA PaCa-2, Su.86.86, Capan-1, PANC-1, SW1990, BxPC-3 and AsPC-1 | GEM-R cells BxPC-3 and PANC-1 | 1207 | (Bio-Rad, Hercules, CA, USA) | PVT1 |
| Chemoresistance | |||
|---|---|---|---|
| Downregulated | Upregulated | ||
| miRNA | Pathway | miRNA | Pathway |
| GEM | GEM | ||
| 210 | ABCC5 | 17-5p | PTEN |
| 124 | PTBP1/PKM2 | 221 | EGFR and HER pathways |
| 101-3p | RRM1 | 203 | SIK1 |
| 100 | FGFR3 | 181c | MST1/2, and LATS1/2, together with the adaptor proteins SAV1 and MOB1 (Hippo signalling pathway) |
| 497 | FGF/FGFR signalling pathway | 15b | SMURF2 |
| 7 | TUBB3 | 21 | NM |
| 205 | TUBB3 | 221 | NM |
| 374b-5p | BIAPRC-3 and XIAP | 1246 | CCNG2 |
| 5-FU | 21 | PTEN/Akt pathway | |
| 200c | SUZ12, ROCK2 direct targets | 320c | SMARCC1 mediated the anti-cancer effect of GEM |
| 220b | SUZ12, ROCK2 direct targets | 155 | Anti-apoptotic (RAB27B) |
| 21 | NM | ||
| 221 | NM | ||
| 21 | VEGF and MMP-2 and MMP-9 | ||
| 10a-5p | TFAP2C | ||
| 5-FU | |||
| 1246 | CCNG2 | ||
| Chemosensitivity | |||
|---|---|---|---|
| Downregulated | Upregulated | ||
| miRNA | Pathway | miRNA | Pathway |
| GEM | GEM | ||
| 3656 | RHOF/EMT | 509-5p | E-cadherin |
| Let-7a | CXCR4/HMGA2 | 1243 | E-cadherin |
| 205-5p | K-ras, Caveolin-1 and Ki-67 | 33a | AKT/Gsk-3β/β-catenin pathway |
| 153 | SNAIL | 21 | FasL/Fas pathway |
| 101 | DNA-PKcs | 1207 | PVT1 |
| 506 | SPHK1/Akt/NF-κB | 5-FU | |
| 494 | c-Myc/SIRT1 pathway | 138-5p | vimentin |
| 203 | ZEB-1 | ||
| 5-FU | |||
| 494 | c-Myc/SIRT1 pathway | ||
| Groups | Clinical Outcomes | Classic Fail-Safe N | Orwin Fail-Safe N | Begg and Mazumdar | Egger′s Regression | Dual and Tweedie (Random Effects) | |||||||||
| z-Value | p-Value | HR in Observed | Tau | z-Value | p-Value | Intercept | p-Value | df | Observed | q-Value | Adjusted | q-Value | |||
| Main | Main Meta-analysis | 6.264 | 0 | 1.150 | 0.257 | 1.165 | 0.243 | 1.778 | 0.052 | 10 | 1.603 | 68.041 | 1.132 | 107.980 | |
| Fixed | Mixed/Random | Hypothesis Test | |||||||||||||
| Subgroups | Heterogeneity | HR | 95% CI | HR | 95% CI | Fixed effects model | Random effects model | ||||||||
| Q | P | I2 | Low | High | Low | High | Z | P | Studies | Z | P | Studies | |||
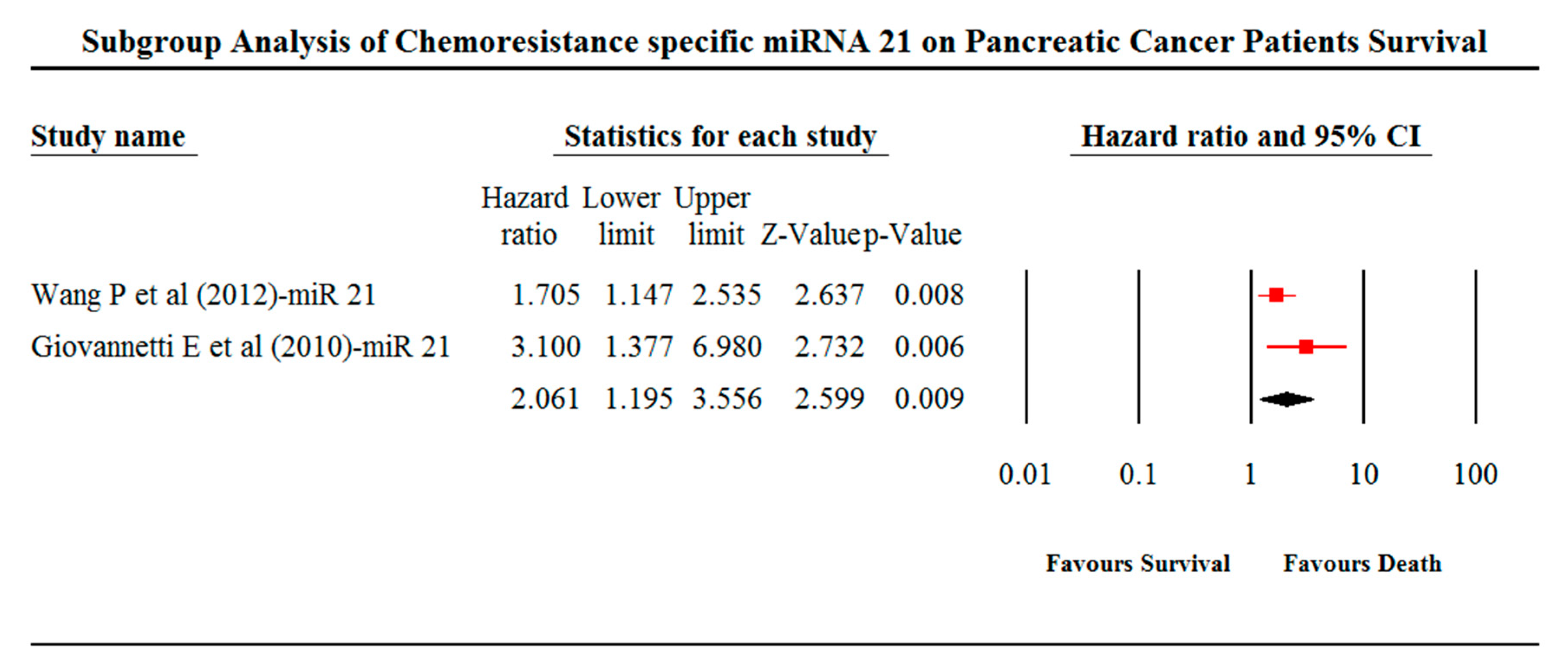
| miR-21 | 1.683 | 0.195 | 40.567 | 1.913 | 1.340 | 2.732 | 2.061 | 1.195 | 3.556 | 3.569 | 0 | 2 | 2.599 | 0.009 | 2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madurantakam Royam, M.; Ramesh, R.; Shanker, R.; Sabarimurugan, S.; Kumarasamy, C.; Ramesh, N.; Gothandam, K.M.; Baxi, S.; Gupta, A.; Krishnan, S.; et al. miRNA Predictors of Pancreatic Cancer Chemotherapeutic Response: A Systematic Review and Meta-Analysis. Cancers 2019, 11, 900. https://doi.org/10.3390/cancers11070900
Madurantakam Royam M, Ramesh R, Shanker R, Sabarimurugan S, Kumarasamy C, Ramesh N, Gothandam KM, Baxi S, Gupta A, Krishnan S, et al. miRNA Predictors of Pancreatic Cancer Chemotherapeutic Response: A Systematic Review and Meta-Analysis. Cancers. 2019; 11(7):900. https://doi.org/10.3390/cancers11070900
Chicago/Turabian StyleMadurantakam Royam, Madhav, Rithika Ramesh, Ritika Shanker, Shanthi Sabarimurugan, Chellan Kumarasamy, Nachimuthu Ramesh, Kodiveri Muthukalianan Gothandam, Siddharta Baxi, Ajay Gupta, Sunil Krishnan, and et al. 2019. "miRNA Predictors of Pancreatic Cancer Chemotherapeutic Response: A Systematic Review and Meta-Analysis" Cancers 11, no. 7: 900. https://doi.org/10.3390/cancers11070900
APA StyleMadurantakam Royam, M., Ramesh, R., Shanker, R., Sabarimurugan, S., Kumarasamy, C., Ramesh, N., Gothandam, K. M., Baxi, S., Gupta, A., Krishnan, S., & Jayaraj, R. (2019). miRNA Predictors of Pancreatic Cancer Chemotherapeutic Response: A Systematic Review and Meta-Analysis. Cancers, 11(7), 900. https://doi.org/10.3390/cancers11070900









