Control of Tyrosine Kinase Signalling by Small Adaptors in Colorectal Cancer
Abstract
1. Colorectal Cancer
2. TK Signalling in CRC
3. The Control of SRC Oncogenic Signalling by SLAP in CRC
3.1. SRC Tumour Activity in CRC
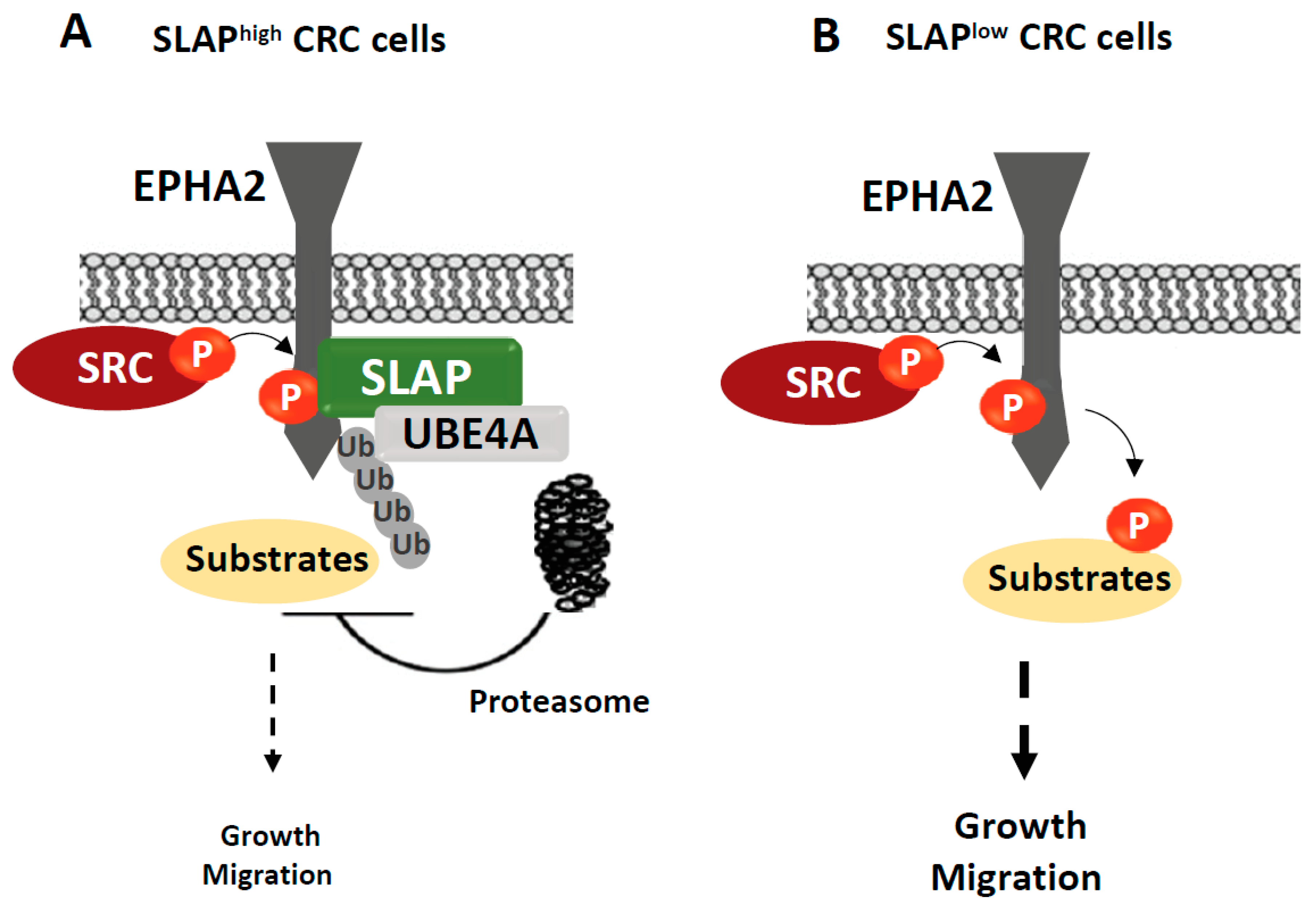
3.2. SLAP Tumour Suppressor Activity in CRC
4. The Control of JAK Oncogenic Signalling by SOCS in CRC
4.1. JAK/STAT Activities in CRC
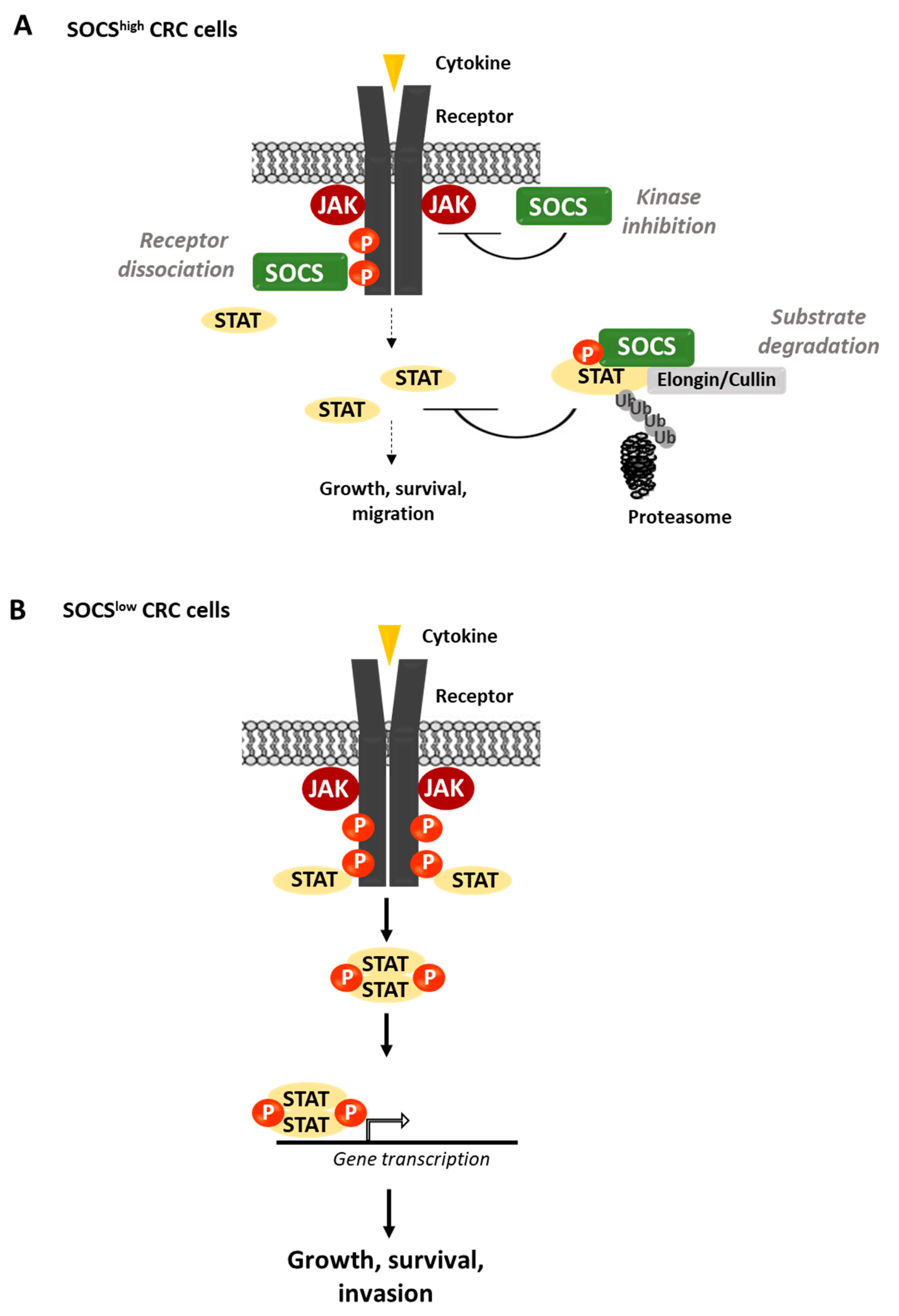
4.2. SOCSs Activity in CRC
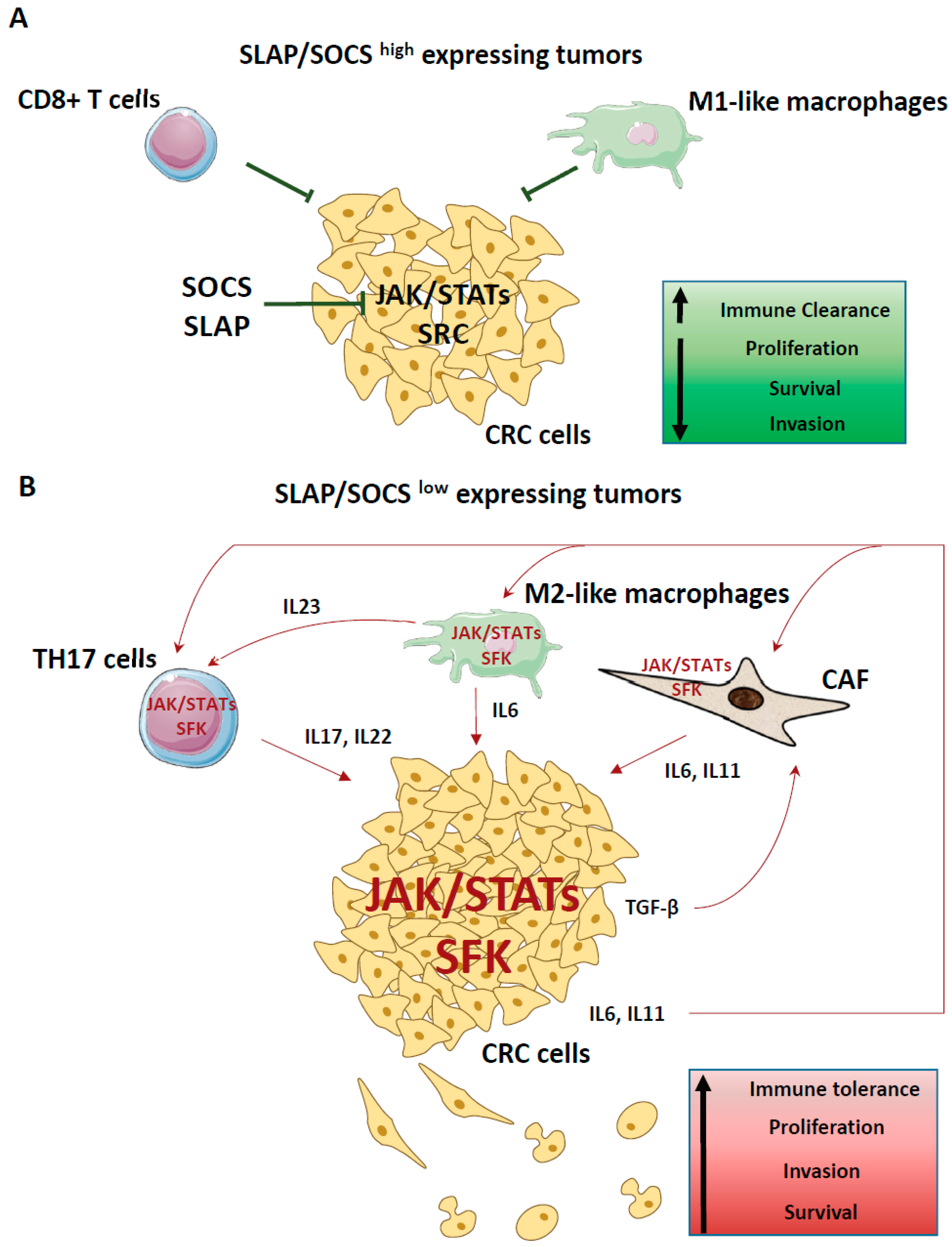
5. Discussion
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Brenner, H.; Kloor, M.; Pox, C.P. Colorectal cancer. Lancet 2014, 383, 1490–1502. [Google Scholar] [CrossRef]
- Vanharanta, S.; Massagué, J. Origins of metastatic traits. Cancer Cell 2013, 24, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Pohl, M.; Schmiegel, W. Therapeutic strategies in diseases of the digestive tract—2015 and beyond targeted therapies in colon cancer today and tomorrow. Dig. Dis. 2016, 34, 574–579. [Google Scholar] [CrossRef] [PubMed]
- Lièvre, A.; Laurent-Puig, P. Genetics: Predictive value of KRAS mutations in chemoresistant CRC. Nat. Rev. Clin. Oncol. 2009, 6, 306–307. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reyniès, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef] [PubMed]
- Lim, W.A.; Pawson, T. Phosphotyrosine signaling: Evolving a new cellular communication system. Cell 2010, 142, 661–667. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Pawson, T. Modular evolution of phosphorylation-based signalling systems. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 2540–2555. [Google Scholar] [CrossRef]
- Liu, B.A.; Nash, P.D. Evolution of SH2 domains and phosphotyrosine signalling networks. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 2556–2573. [Google Scholar] [CrossRef] [PubMed]
- Blume-Jensen, P.; Hunter, T. Oncogenic kinase signalling. Nature 2001, 411, 355–365. [Google Scholar] [CrossRef]
- Savage, D.G.; Antman, K.H. Imatinib mesylate—A new oral targeted therapy. N. Engl. J. Med. 2002, 346, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Krause, D.S.; Van Etten, R.A. Tyrosine kinases as targets for cancer therapy. N. Engl. J. Med. 2005, 353, 172–187. [Google Scholar] [CrossRef] [PubMed]
- Frame, M.C. Newest findings on the oldest oncogene; how activated src does it. J. Cell Sci. 2004, 117, 989–998. [Google Scholar] [CrossRef]
- García-Aranda, M.; Redondo, M. Targeting Receptor Kinases in Colorectal Cancer. Cancers 2019, 11, 433. [Google Scholar] [CrossRef] [PubMed]
- Jeitany, M.; Leroy, C.; Tosti, P.; Lafitte, M.; Le Guet, J.; Simon, V.; Bonenfant, D.; Robert, B.; Grillet, F.; Mollevi, C.; et al. Inhibition of DDR1-BCR signalling by nilotinib as a new therapeutic strategy for metastatic colorectal cancer. EMBO Mol. Med. 2018, 10, e7918. [Google Scholar] [CrossRef] [PubMed]
- Cohen, R.B. Epidermal growth factor receptor as a therapeutic target in colorectal cancer. Clin. Colorectal Cancer 2003, 2, 246–251. [Google Scholar] [CrossRef]
- Saltz, L.B.; Meropol, N.J.; Loehrer, P.J.; Needle, M.N.; Kopit, J.; Mayer, R.J. Phase II trial of cetuximab in patients with refractory colorectal cancer that expresses the epidermal growth factor receptor. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2004, 22, 1201–1208. [Google Scholar] [CrossRef]
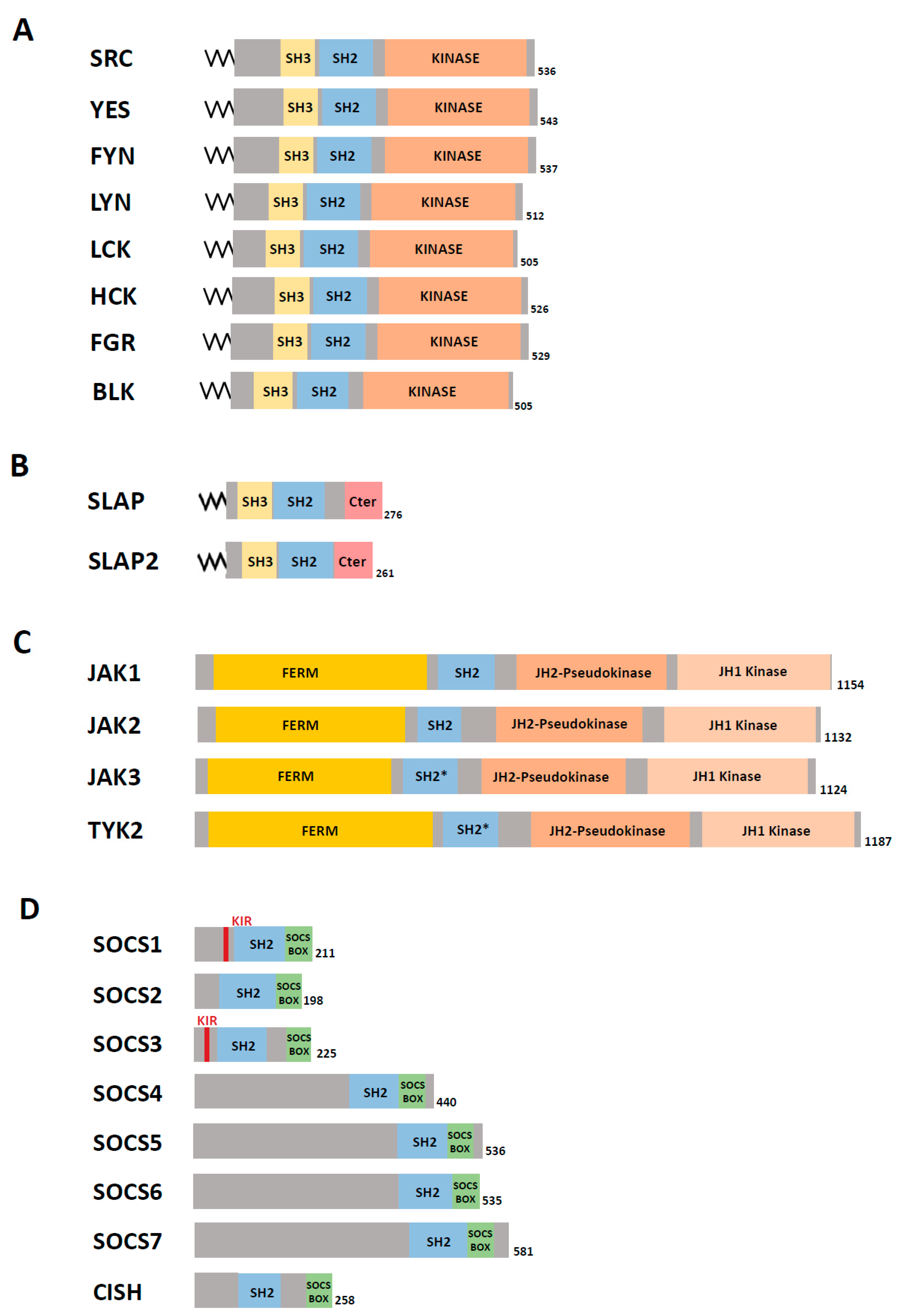
- Naudin, C.; Chevalier, C.; Roche, S. The role of small adaptor proteins in the control of oncogenic signaling driven by tyrosine kinases in human cancer. Oncotarget 2016, 7, 11033–11055. [Google Scholar] [CrossRef] [PubMed]
- Trengove, M.C.; Ward, A.C. SOCS proteins in development and disease. Am. J. Clin. Exp. Immunol. 2013, 2, 1–29. [Google Scholar]
- Inagaki-Ohara, K.; Kondo, T.; Ito, M.; Yoshimura, A. SOCS, inflammation, and cancer. JAK-STAT 2013, 2, e24053. [Google Scholar] [CrossRef]
- Thomas, S.M.; Brugge, J.S. Cellular functions regulated by Src family kinases. Annu. Rev. Cell Dev. Biol. 1997, 13, 513–609. [Google Scholar] [CrossRef]
- Cordero, J.B.; Ridgway, R.A.; Valeri, N.; Nixon, C.; Frame, M.C.; Muller, W.J.; Vidal, M.; Sansom, O.J. c-Src drives intestinal regeneration and transformation. EMBO J. 2014, 33, 1474–1491. [Google Scholar] [CrossRef] [PubMed]
- Kohlmaier, A.; Fassnacht, C.; Jin, Y.; Reuter, H.; Begum, J.; Dutta, D.; Edgar, B.A. Src kinase function controls progenitor cell pools during regeneration and tumor onset in the Drosophila intestine. Oncogene 2015, 34, 2371–2384. [Google Scholar] [CrossRef]
- Sirvent, A.; Benistant, C.; Roche, S. Oncogenic signaling by tyrosine kinases of the SRC family in advanced colorectal cancer. Am. J. Cancer Res. 2012, 2, 357–371. [Google Scholar]
- Summy, J.M.; Gallick, G.E. Src family kinases in tumor progression and metastasis. Cancer Metastasis Rev. 2003, 22, 337–358. [Google Scholar] [CrossRef] [PubMed]
- Aligayer, H.; Boyd, D.D.; Heiss, M.M.; Abdalla, E.K.; Curley, S.A.; Gallick, G.E. Activation of Src kinase in primary colorectal carcinoma: An indicator of poor clinical prognosis. Cancer 2002, 94, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Elfiky, A.; Han, M.; Chen, C.; Saif, M.W. The role of Src in colon cancer and its therapeutic implications. Clin. Colorectal Cancer 2014, 13, 5–13. [Google Scholar] [CrossRef]
- Sancier, F.; Dumont, A.; Sirvent, A.; Paquay de Plater, L.; Edmonds, T.; David, G.; Jan, M.; de Montrion, C.; Cogé, F.; Léonce, S.; et al. Specific oncogenic activity of the Src-family tyrosine kinase c-Yes in colon carcinoma cells. PLoS ONE 2011, 6, e17237. [Google Scholar] [CrossRef][Green Version]
- Rosenbluh, J.; Nijhawan, D.; Cox, A.G.; Li, X.; Neal, J.T.; Schafer, E.J.; Zack, T.I.; Wang, X.; Tsherniak, A.; Schinzel, A.C.; et al. β-Catenin-driven cancers require a YAP1 transcriptional complex for survival and tumorigenesis. Cell 2012, 151, 1457–1473. [Google Scholar] [CrossRef]
- Dubois, F.; Leroy, C.; Simon, V.; Benistant, C.; Roche, S. YES oncogenic activity is specified by its SH4 domain and regulates RAS/MAPK signaling in colon carcinoma cells. Am. J. Cancer Res. 2015, 5, 1972–1987. [Google Scholar]
- Parseghian, C.M.; Parikh, N.U.; Wu, J.Y.; Jiang, Z.-Q.; Henderson, L.; Tian, F.; Pastor, B.; Ychou, M.; Raghav, K.; Dasari, A.; et al. Dual inhibition of EGFR and c-Src by cetuximab and dasatinib combined with FOLFOX chemotherapy in patients with metastatic colorectal cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2017, 23, 4146–4154. [Google Scholar] [CrossRef]
- Perez, M.; Lucena-Cacace, A.; Marín-Gómez, L.M.; Padillo-Ruiz, J.; Robles-Frias, M.J.; Saez, C.; Garcia-Carbonero, R.; Carnero, A. Dasatinib, a Src inhibitor, sensitizes liver metastatic colorectal carcinoma to oxaliplatin in tumors with high levels of phospho-Src. Oncotarget 2016, 7, 33111–33124. [Google Scholar] [PubMed]
- Reddy, S.M.; Kopetz, S.; Morris, J.; Parikh, N.; Qiao, W.; Overman, M.J.; Fogelman, D.; Shureiqi, I.; Jacobs, C.; Malik, Z.; et al. Phase II study of saracatinib (AZD0530) in patients with previously treated metastatic colorectal cancer. Investig. New Drugs 2015, 33, 977–984. [Google Scholar] [CrossRef]
- Leroy, C.; Fialin, C.; Sirvent, A.; Simon, V.; Urbach, S.; Poncet, J.; Robert, B.; Jouin, P.; Roche, S. Quantitative phosphoproteomics reveals a cluster of tyrosine kinases that mediates SRC invasive activity in advanced colon carcinoma cells. Cancer Res. 2009, 69, 2279–2286. [Google Scholar] [CrossRef]
- Taniguchi, K.; Moroishi, T.; de Jong, P.R.; Krawczyk, M.; Grebbin, B.M.; Luo, H.; Xu, R.-H.; Golob-Schwarzl, N.; Schweiger, C.; Wang, K.; et al. YAP-IL-6ST autoregulatory loop activated on APC loss controls colonic tumorigenesis. Proc. Natl. Acad. Sci. USA 2017, 114, 1643–1648. [Google Scholar] [CrossRef]
- Sirvent, A.; Vigy, O.; Orsetti, B.; Urbach, S.; Roche, S. Analysis of SRC oncogenic signaling in colorectal cancer by stable isotope labeling with heavy amino acids in mouse xenografts. Mol. Cell. Proteom. MCP 2012, 11, 1937–1950. [Google Scholar] [CrossRef]
- Boggon, T.J.; Eck, M.J. Structure and regulation of Src family kinases. Oncogene 2004, 23, 7918–7927. [Google Scholar] [CrossRef]
- Spassov, D.S.; Ruiz-Saenz, A.; Piple, A.; Moasser, M.M. A Dimerization Function in the Intrinsically Disordered N-Terminal Region of Src. Cell Rep. 2018, 25, 449–463.e4. [Google Scholar] [CrossRef]
- Ahler, E.; Register, A.C.; Chakraborty, S.; Fang, L.; Dieter, E.M.; Sitko, K.A.; Vidadala, R.S.R.; Trevillian, B.M.; Golkowski, M.; Gelman, H.; et al. A Combined approach reveals a regulatory mechanism coupling Src’s kinase activity, localization, and phosphotransferase-independent functions. Mol. Cell 2019, 74, 393–408. [Google Scholar] [CrossRef]
- Le Roux, A.-L.; Busquets, M.A.; Sagués, F.; Pons, M. Kinetics characterization of c-Src binding to lipid membranes: Switching from labile to persistent binding. Colloids Surf. B Biointerfaces 2016, 138, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Imada, S.; Murata, Y.; Kotani, T.; Hatano, M.; Sun, C.; Konno, T.; Park, J.-H.; Kitamura, Y.; Saito, Y.; Ohdan, H.; et al. Role of Src family kinases in regulation of intestinal epithelial homeostasis. Mol. Cell. Biol. 2016, 36, 2811–2823. [Google Scholar] [CrossRef]
- Bénistant, C.; Bourgaux, J.F.; Chapuis, H.; Mottet, N.; Roche, S.; Bali, J.P. The COOH-terminal Src kinase Csk is a tumor antigen in human carcinoma. Cancer Res. 2001, 61, 1415–1420. [Google Scholar]
- Zhang, B.; Wang, J.; Wang, X.; Zhu, J.; Liu, Q.; Shi, Z.; Chambers, M.C.; Zimmerman, L.J.; Shaddox, K.F.; Kim, S.; et al. Proteogenomic characterization of human colon and rectal cancer. Nature 2014, 513, 382–387. [Google Scholar] [CrossRef]
- Sirvent, A.; Bénistant, C.; Pannequin, J.; Veracini, L.; Simon, V.; Bourgaux, J.-F.; Hollande, F.; Cruzalegui, F.; Roche, S. Src family tyrosine kinases-driven colon cancer cell invasion is induced by Csk membrane delocalization. Oncogene 2010, 29, 1303–1315. [Google Scholar] [CrossRef]
- Oneyama, C.; Hikita, T.; Enya, K.; Dobenecker, M.-W.; Saito, K.; Nada, S.; Tarakhovsky, A.; Okada, M. The lipid raft-anchored adaptor protein Cbp controls the oncogenic potential of c-Src. Mol. Cell 2008, 30, 426–436. [Google Scholar] [CrossRef]
- Suzuki, K.; Oneyama, C.; Kimura, H.; Tajima, S.; Okada, M. Down-regulation of the tumor suppressor C-terminal Src kinase (Csk)-binding protein (Cbp)/PAG1 is mediated by epigenetic histone modifications via the mitogen-activated protein kinase (MAPK)/phosphatidylinositol 3-kinase (PI3K) pathway. J. Biol. Chem. 2011, 286, 15698–15706. [Google Scholar] [CrossRef] [PubMed]
- Dragone, L.L.; Shaw, L.A.; Myers, M.D.; Weiss, A. SLAP, a regulator of immunoreceptor ubiquitination, signaling, and trafficking. Immunol. Rev. 2009, 232, 218–228. [Google Scholar] [CrossRef]
- Wybenga-Groot, L.E.; McGlade, C.J. RTK SLAP down: The emerging role of Src-like adaptor protein as a key player in receptor tyrosine kinase signaling. Cell. Signal. 2015, 27, 267–274. [Google Scholar] [CrossRef]
- Sosinowski, T.; Killeen, N.; Weiss, A. The Src-like adaptor protein downregulates the T cell receptor on CD4+CD8+ thymocytes and regulates positive selection. Immunity 2001, 15, 457–466. [Google Scholar] [CrossRef]
- Myers, M.D.; Sosinowski, T.; Dragone, L.L.; White, C.; Band, H.; Gu, H.; Weiss, A. Src-like adaptor protein regulates TCR expression on thymocytes by linking the ubiquitin ligase c-Cbl to the TCR complex. Nat. Immunol. 2006, 7, 57–66. [Google Scholar] [CrossRef]
- Roche, S.; Alonso, G.; Kazlauskas, A.; Dixit, V.M.; Courtneidge, S.A.; Pandey, A. Src-like adaptor protein (Slap) is a negative regulator of mitogenesis. Curr. Biol. 1998, 8, 975–978. [Google Scholar] [CrossRef]
- Sirvent, A.; Leroy, C.; Boureux, A.; Simon, V.; Roche, S. The Src-like adaptor protein regulates PDGF-induced actin dorsal ruffles in a c-Cbl-dependent manner. Oncogene 2008, 27, 3494–3500. [Google Scholar] [CrossRef]
- Manes, G.; Bello, P.; Roche, S. Slap negatively regulates Src mitogenic function but does not revert Src-induced cell morphology changes. Mol. Cell. Biol. 2000, 20, 3396–3406. [Google Scholar] [CrossRef]
- Naudin, C.; Sirvent, A.; Leroy, C.; Larive, R.; Simon, V.; Pannequin, J.; Bourgaux, J.-F.; Pierre, J.; Robert, B.; Hollande, F.; et al. SLAP displays tumour suppressor functions in colorectal cancer via destabilization of the SRC substrate EPHA2. Nat. Commun. 2014, 5, 3159. [Google Scholar] [CrossRef]
- Sakiyama, T.; Fujita, H.; Tsubouchi, H. Autoantibodies against ubiquitination factor E4A (UBE4A) are associated with severity of Crohn’s disease. Inflamm. Bowel Dis. 2008, 14, 310–317. [Google Scholar] [CrossRef]
- Kazi, J.U.; Rönnstrand, L. Src-like adaptor protein (SLAP) binds to the receptor tyrosine kinase Flt3 and modulates receptor stability and downstream signaling. PLoS ONE 2012, 7, e53509. [Google Scholar] [CrossRef]
- David, M.; Naudin, C.; Letourneur, M.; Polrot, M.; Renoir, J.-M.; Lazar, V.; Dessen, P.; Roche, S.; Bertoglio, J.; Pierre, J. Suppressor of cytokine signaling 1 modulates invasion and metastatic potential of colorectal cancer cells. Mol. Oncol. 2014, 8, 942–955. [Google Scholar] [CrossRef]
- Lin, S.-Y.; Yeh, K.-T.; Chen, W.T.-L.; Chen, H.-C.; Chen, S.-T.; Chiou, H.-Y.; Chang, J.-G. Promoter CpG methylation of tumor suppressor genes in colorectal cancer and its relationship to clinical features. Oncol. Rep. 2004, 11, 341–348. [Google Scholar] [CrossRef]
- Fujitake, S.; Hibi, K.; Okochi, O.; Kodera, Y.; Ito, K.; Akiyama, S.; Nakao, A. Aberrant methylation of SOCS-1 was observed in younger colorectal cancer patients. J. Gastroenterol. 2004, 39, 120–124. [Google Scholar] [CrossRef]
- Hibi, K.; Kodera, Y.; Ito, K.; Akiyama, S.; Nakao, A. Aberrant methylation of HLTF, SOCS-1, and CDH13 genes is shown in colorectal cancers without lymph node metastasis. Dis. Colon Rectum 2005, 48, 1282–1286. [Google Scholar] [CrossRef]
- Xu, S.B.; Liu, X.H.; Li, B.H.; Zhang, Y.; Yuan, J.; Yuan, Q.; Li, P.D.; Yang, X.Z.; Li, F.; Zhang, W.J. DNA methylation regulates constitutive expression of Stat6 regulatory genes SOCS-1 and SHP-1 in colon cancer cells. J. Cancer Res. Clin. Oncol. 2009, 135, 1791–1798. [Google Scholar] [CrossRef]
- Kang, X.-C.; Chen, M.-L.; Yang, F.; Gao, B.-Q.; Yang, Q.-H.; Zheng, W.-W.; Hao, S. Promoter methylation and expression of SOCS-1 affect clinical outcome and epithelial-mesenchymal transition in colorectal cancer. Biomed. Pharmacother. 2016, 80, 23–29. [Google Scholar] [CrossRef]
- Slattery, M.L.; Lundgreen, A.; Kadlubar, S.A.; Bondurant, K.L.; Wolff, R.K. JAK/STAT/SOCS-signaling pathway and colon and rectal cancer. Mol. Carcinog. 2013, 52, 155–166. [Google Scholar] [CrossRef]
- Ren, D.; Lin, B.; Zhang, X.; Peng, Y.; Ye, Z.; Ma, Y.; Liang, Y.; Cao, L.; Li, X.; Li, R.; et al. Maintenance of cancer stemness by miR-196b-5p contributes to chemoresistance of colorectal cancer cells via activating STAT3 signaling pathway. Oncotarget 2017, 8, 49807–49823. [Google Scholar] [CrossRef]
- Letellier, E.; Schmitz, M.; Baig, K.; Beaume, N.; Schwartz, C.; Frasquilho, S.; Antunes, L.; Marcon, N.; Nazarov, P.V.; Vallar, L.; et al. Identification of SOCS2 and SOCS6 as biomarkers in human colorectal cancer. Br. J. Cancer 2014, 111, 726–735. [Google Scholar] [CrossRef]
- Kim, J.-H.; Lee, M.-J.; Yu, G.-R.; Kim, S.-W.; Jang, K.-Y.; Yu, H.-C.; Cho, B.-H.; Kim, D.-G. Alterations in the p53-SOCS2 axis contribute to tumor growth in colon cancer. Exp. Mol. Med. 2018, 50, 3. [Google Scholar] [CrossRef]
- Liu, X.H.; Xu, S.B.; Yuan, J.; Li, B.H.; Zhang, Y.; Yuan, Q.; Li, P.D.; Li, F.; Zhang, W.J. Defective interleukin-4/Stat6 activity correlates with increased constitutive expression of negative regulators SOCS-3, SOCS-7, and CISH in colon cancer cells. J. Interferon Cytokine Res. Off. J. Int. Soc. Interferon Cytokine Res. 2009, 29, 809–816. [Google Scholar] [CrossRef]
- Chu, Q.; Shen, D.; He, L.; Wang, H.; Liu, C.; Zhang, W. Prognostic significance of SOCS3 and its biological function in colorectal cancer. Gene 2017, 627, 114–122. [Google Scholar] [CrossRef]
- Li, Y.; Deuring, J.; Peppelenbosch, M.P.; Kuipers, E.J.; de Haar, C.; van der Woude, C.J. IL-6-induced DNMT1 activity mediates SOCS3 promoter hypermethylation in ulcerative colitis-related colorectal cancer. Carcinogenesis 2012, 33, 1889–1896. [Google Scholar] [CrossRef]
- Su, M.; Qin, B.; Liu, F.; Chen, Y.; Zhang, R. miR-885-5p upregulation promotes colorectal cancer cell proliferation and migration by targeting suppressor of cytokine signaling. Oncol. Lett. 2018, 16, 65–72. [Google Scholar] [CrossRef]
- Ragusa, M.; Statello, L.; Maugeri, M.; Majorana, A.; Barbagallo, D.; Salito, L.; Sammito, M.; Santonocito, M.; Angelica, R.; Cavallaro, A.; et al. Specific alterations of the microRNA transcriptome and global network structure in colorectal cancer after treatment with MAPK/ERK inhibitors. J. Mol. Med. 2012, 90, 1421–1438. [Google Scholar] [CrossRef]
- Storojeva, I.; Boulay, J.-L.; Ballabeni, P.; Buess, M.; Terracciano, L.; Laffer, U.; Mild, G.; Herrmann, R.; Rochlitz, C. Prognostic and predictive relevance of DNAM-1, SOCS6 and CADH-7 genes on chromosome 18q in colorectal cancer. Oncology 2005, 68, 246–255. [Google Scholar] [CrossRef]
- Lai, R.-H.; Wang, M.-J.; Yang, S.-H.; Chen, J.-Y. Genomic organization and functional characterization of the promoter for the human suppressor of cytokine signaling 6 gene. Gene 2009, 448, 64–73. [Google Scholar] [CrossRef]
- Fang, Y.; Sun, B.; Xiang, J.; Chen, Z. MiR-301a promotes colorectal cancer cell growth and invasion by directly targeting SOCS6. Cell. Physiol. Biochem. 2015, 35, 227–236. [Google Scholar] [CrossRef]
- Poh, A.R.; Love, C.G.; Masson, F.; Preaudet, A.; Tsui, C.; Whitehead, L.; Monard, S.; Khakham, Y.; Burstroem, L.; Lessene, G.; et al. Inhibition of hematopoietic cell kinase activity suppresses myeloid cell-mediated colon cancer progression. Cancer Cell 2017, 31, 563–575.e5. [Google Scholar] [CrossRef]
- Roseweir, A.K.; Powell, A.G.M.T.; Horstman, S.L.; Inthagard, J.; Park, J.H.; McMillan, D.C.; Horgan, P.G.; Edwards, J. Src family kinases, HCK and FGR, associate with local inflammation and tumour progression in colorectal cancer. Cell. Signal. 2019, 56, 15–22. [Google Scholar] [CrossRef]
- Pakuts, B.; Debonneville, C.; Liontos, L.M.; Loreto, M.P.; McGlade, C.J. The Src-like adaptor protein 2 regulates colony-stimulating factor-1 receptor signaling and down-regulation. J. Biol. Chem. 2007, 282, 17953–17963. [Google Scholar] [CrossRef]
- Kazi, J.U.; Agarwal, S.; Sun, J.; Bracco, E.; Rönnstrand, L. Src-like-adaptor protein (SLAP) differentially regulates normal and oncogenic c-Kit signaling. J. Cell Sci. 2014, 127, 653–662. [Google Scholar] [CrossRef]
- Villarino, A.V.; Kanno, Y.; O’Shea, J.J. Mechanisms and consequences of Jak–STAT signaling in the immune system. Nat. Immunol. 2017, 18, 374–384. [Google Scholar] [CrossRef]
- Darnell, J.E.; Kerr, I.M.; Stark, G.R. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 1994, 264, 1415–1421. [Google Scholar] [CrossRef]
- Leonard, W.J.; O’Shea, J.J. JAKS AND STATS: Biological Implications. Annu. Rev. Immunol. 1998, 16, 293–322. [Google Scholar] [CrossRef]
- Firmbach-Kraft, I.; Byers, M.; Shows, T.; Dalla-Favera, R.; Krolewski, J.J. tyk2, prototype of a novel class of non-receptor tyrosine kinase genes. Oncogene 1990, 5, 1329–1336. [Google Scholar]
- Kawamura, M.; McVicar, D.W.; Johnston, J.A.; Blake, T.B.; Chen, Y.Q.; Lal, B.K.; Lloyd, A.R.; Kelvin, D.J.; Staples, J.E.; Ortaldo, J.R. Molecular cloning of L-JAK, a Janus family protein-tyrosine kinase expressed in natural killer cells and activated leukocytes. Proc. Natl. Acad. Sci. USA 1994, 91, 6374–6378. [Google Scholar] [CrossRef] [PubMed]
- Wilks, A.F.; Harpur, A.G.; Kurban, R.R.; Ralph, S.J.; Zürcher, G.; Ziemiecki, A. Two novel protein-tyrosine kinases, each with a second phosphotransferase-related catalytic domain, define a new class of protein kinase. Mol. Cell. Biol. 1991, 11, 2057–2065. [Google Scholar] [CrossRef] [PubMed]
- Vainchenker, W.; Constantinescu, S.N. JAK/STAT signaling in hematological malignancies. Oncogene 2013, 32, 2601–2613. [Google Scholar] [CrossRef] [PubMed]
- Lacout, C.; Pisani, D.F.; Tulliez, M.; Gachelin, F.M.; Vainchenker, W.; Villeval, J.-L. JAK2V617F expression in murine hematopoietic cells leads to MPD mimicking human PV with secondary myelofibrosis. Blood 2006, 108, 1652–1660. [Google Scholar] [CrossRef] [PubMed]
- Tiedt, R.; Hao-Shen, H.; Sobas, M.A.; Looser, R.; Dirnhofer, S.; Schwaller, J.; Skoda, R.C. Ratio of mutant JAK2-V617F to wild-type Jak2 determines the MPD phenotypes in transgenic mice. Blood 2008, 111, 3931–3940. [Google Scholar] [CrossRef]
- Wernig, G.; Gonneville, J.R.; Crowley, B.J.; Rodrigues, M.S.; Reddy, M.M.; Hudon, H.E.; Walz, C.; Reiter, A.; Podar, K.; Royer, Y.; et al. The Jak2V617F oncogene associated with myeloproliferative diseases requires a functional FERM domain for transformation and for expression of the Myc and Pim proto-oncogenes. Blood 2008, 111, 3751–3759. [Google Scholar] [CrossRef]
- Akada, H.; Yan, D.; Zou, H.; Fiering, S.; Hutchison, R.E.; Mohi, M.G. Conditional expression of heterozygous or homozygous Jak2V617F from its endogenous promoter induces a polycythemia vera-like disease. Blood 2010, 115, 3589–3597. [Google Scholar] [CrossRef]
- Li, J.; Spensberger, D.; Ahn, J.S.; Anand, S.; Beer, P.A.; Ghevaert, C.; Chen, E.; Forrai, A.; Scott, L.M.; Ferreira, R.; et al. JAK2 V617F impairs hematopoietic stem cell function in a conditional knock-in mouse model of JAK2 V617F-positive essential thrombocythemia. Blood 2010, 116, 1528–1538. [Google Scholar] [CrossRef]
- Marty, C.; Lacout, C.; Martin, A.; Hasan, S.; Jacquot, S.; Birling, M.-C.; Vainchenker, W.; Villeval, J.-L. Myeloproliferative neoplasm induced by constitutive expression of JAK2V617F in knock-in mice. Blood 2010, 116, 783–787. [Google Scholar] [CrossRef]
- Mullally, A.; Lane, S.W.; Ball, B.; Megerdichian, C.; Okabe, R.; Al-Shahrour, F.; Paktinat, M.; Haydu, J.E.; Housman, E.; Lord, A.M.; et al. Physiological Jak2V617F expression causes a lethal myeloproliferative neoplasm with differential effects on hematopoietic stem and progenitor cells. Cancer Cell 2010, 17, 584–596. [Google Scholar] [CrossRef]
- Jeong, E.G.; Kim, M.S.; Nam, H.K.; Min, C.K.; Lee, S.; Chung, Y.J.; Yoo, N.J.; Lee, S.H. Somatic mutations of JAK1 and JAK3 in acute leukemias and solid cancers. Clin. Cancer Res. 2008, 14, 3716–3721. [Google Scholar] [CrossRef]
- Bardelli, A.; Parsons, D.W.; Silliman, N.; Ptak, J.; Szabo, S.; Saha, S.; Markowitz, S.; Willson, J.K.V.; Parmigiani, G.; Kinzler, K.W.; et al. Mutational analysis of the tyrosine kinome in colorectal cancers. Science 2003, 300, 949. [Google Scholar] [CrossRef]
- Sveen, A.; Johannessen, B.; Tengs, T.; Danielsen, S.A.; Eilertsen, I.A.; Lind, G.E.; Berg, K.C.G.; Leithe, E.; Meza-Zepeda, L.A.; Domingo, E.; et al. Multilevel genomics of colorectal cancers with microsatellite instability—Clinical impact of JAK1 mutations and consensus molecular subtype 1. Genome Med. 2017, 9, 46. [Google Scholar] [CrossRef]
- Buchert, M.; Burns, C.J.; Ernst, M. Targeting JAK kinase in solid tumors: Emerging opportunities and challenges. Oncogene 2016, 35, 939–951. [Google Scholar] [CrossRef]
- Wang, S.; Zhang, W. Genetic variants in IL-6/JAK/STAT3 pathway and the risk of CRC. Tumour Biol. J. Int. Soc. Oncodev. Biol. Med. 2016, 37, 6561–6569. [Google Scholar] [CrossRef]
- Kusaba, T.; Nakayama, T.; Yamazumi, K.; Yakata, Y.; Yoshizaki, A.; Nagayasu, T.; Sekine, I. Expression of p-STAT3 in human colorectal adenocarcinoma and adenoma; correlation with clinicopathological factors. J. Clin. Pathol. 2005, 58, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Tsareva, S.A.; Moriggl, R.; Corvinus, F.M.; Wiederanders, B.; Schütz, A.; Kovacic, B.; Friedrich, K. Signal transducer and activator of transcription 3 activation promotes invasive growth of colon carcinomas through matrix metalloproteinase induction. Neoplasia 2007, 9, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Corvinus, F.M.; Orth, C.; Moriggl, R.; Tsareva, S.A.; Wagner, S.; Pfitzner, E.B.; Baus, D.; Kaufmann, R.; Huber, L.A.; Zatloukal, K.; et al. Persistent STAT3 activation in colon cancer is associated with enhanced cell proliferation and tumor growth. Neoplasia 2005, 7, 545–555. [Google Scholar] [CrossRef]
- Xiong, H.; Zhang, Z.-G.; Tian, X.-Q.; Sun, D.-F.; Liang, Q.-C.; Zhang, Y.-J.; Lu, R.; Chen, Y.-X.; Fang, J.-Y. Inhibition of JAK1, 2/STAT3 signaling induces apoptosis, cell cycle arrest, and reduces tumor cell invasion in colorectal cancer cells. Neoplasia 2008, 10, 287–297. [Google Scholar] [CrossRef]
- Lee, J.; Kim, J.C.K.; Lee, S.-E.; Quinley, C.; Kim, H.; Herdman, S.; Corr, M.; Raz, E. Signal transducer and activator of transcription 3 (STAT3) protein suppresses adenoma-to-carcinoma transition in Apcmin/+ mice via regulation of Snail-1 (SNAI) protein stability. J. Biol. Chem. 2012, 287, 18182–18189. [Google Scholar] [CrossRef] [PubMed]
- Musteanu, M.; Blaas, L.; Mair, M.; Schlederer, M.; Bilban, M.; Tauber, S.; Esterbauer, H.; Mueller, M.; Casanova, E.; Kenner, L.; et al. Stat3 is a negative regulator of intestinal tumor progression in Apc(Min) mice. Gastroenterology 2010, 138, 1003–1011.e5. [Google Scholar] [CrossRef]
- Rodig, S.J.; Meraz, M.A.; White, J.M.; Lampe, P.A.; Riley, J.K.; Arthur, C.D.; King, K.L.; Sheehan, K.C.F.; Yin, L.; Pennica, D.; et al. Disruption of the Jak1 gene demonstrates obligatory and nonredundant roles of the Jaks in cytokine-induced biologic responses. Cell 1998, 93, 373–383. [Google Scholar] [CrossRef]
- Neubauer, H.; Cumano, A.; Müller, M.; Wu, H.; Huffstadt, U.; Pfeffer, K. Jak2 deficiency defines an essential developmental checkpoint in definitive hematopoiesis. Cell 1998, 93, 397–409. [Google Scholar] [CrossRef]
- Parganas, E.; Wang, D.; Stravopodis, D.; Topham, D.J.; Marine, J.C.; Teglund, S.; Vanin, E.F.; Bodner, S.; Colamonici, O.R.; van Deursen, J.M.; et al. Jak2 is essential for signaling through a variety of cytokine receptors. Cell 1998, 93, 385–395. [Google Scholar] [CrossRef]
- Aringer, M.; Cheng, A.; Nelson, J.W.; Chen, M.; Sudarshan, C.; Zhou, Y.J.; O’Shea, J.J. Janus kinases and their role in growth and disease. Life Sci. 1999, 64, 2173–2186. [Google Scholar] [CrossRef]
- Baird, A.M.; Thomis, D.C.; Berg, L.J. T cell development and activation in Jak3-deficient mice. J. Leukoc. Biol. 1998, 63, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Shimoda, K.; Kamesaki, K.; Numata, A.; Aoki, K.; Matsuda, T.; Oritani, K.; Tamiya, S.; Kato, K.; Takase, K.; Imamura, R.; et al. Cutting edge: tyk2 is required for the induction and nuclear translocation of Daxx which regulates IFN-alpha-induced suppression of B lymphocyte formation. J. Immunol. 2002, 169, 4707–4711. [Google Scholar] [CrossRef]
- Shimoda, K.; Tsutsui, H.; Aoki, K.; Kato, K.; Matsuda, T.; Numata, A.; Takase, K.; Yamamoto, T.; Nukina, H.; Hoshino, T.; et al. Partial impairment of interleukin-12 (IL-12) and IL-18 signaling in Tyk2-deficient mice. Blood 2002, 99, 2094–2099. [Google Scholar] [CrossRef] [PubMed]
- Shimoda, K.; Kato, K.; Aoki, K.; Matsuda, T.; Miyamoto, A.; Shibamori, M.; Yamashita, M.; Numata, A.; Takase, K.; Kobayashi, S.; et al. Tyk2 plays a restricted role in IFN alpha signaling, although it is required for IL-12-mediated T cell function. Immunity 2000, 13, 561–571. [Google Scholar] [CrossRef]
- Hainzl, E.; Stockinger, S.; Rauch, I.; Heider, S.; Berry, D.; Lassnig, C.; Schwab, C.; Rosebrock, F.; Milinovich, G.; Schlederer, M.; et al. Intestinal epithelial cell tyrosine kinase 2 transduces IL-22 signals to protect from acute colitis. J. Immunol. 2015, 195, 5011–5024. [Google Scholar] [CrossRef]
- Buchon, N.; Broderick, N.A.; Poidevin, M.; Pradervand, S.; Lemaitre, B. Drosophila intestinal response to bacterial infection: Activation of host defense and stem cell proliferation. Cell Host Microbe 2009, 5, 200–211. [Google Scholar] [CrossRef]
- Jiang, H.; Patel, P.H.; Kohlmaier, A.; Grenley, M.O.; McEwen, D.G.; Edgar, B.A. Cytokine/Jak/Stat signaling mediates regeneration and homeostasis in the Drosophila midgut. Cell 2009, 137, 1343–1355. [Google Scholar] [CrossRef] [PubMed]
- Beebe, K.; Lee, W.-C.; Micchelli, C.A. JAK/STAT signaling coordinates stem cell proliferation and multilineage differentiation in the Drosophila intestinal stem cell lineage. Dev. Biol. 2010, 338, 28–37. [Google Scholar] [CrossRef]
- Cordero, J.B.; Stefanatos, R.K.; Myant, K.; Vidal, M.; Sansom, O.J. Non-autonomous crosstalk between the Jak/Stat and Egfr pathways mediates Apc1-driven intestinal stem cell hyperplasia in the Drosophila adult midgut. Development 2012, 139, 4524–4535. [Google Scholar] [CrossRef] [PubMed]
- Karpowicz, P.; Perez, J.; Perrimon, N. The Hippo tumor suppressor pathway regulates intestinal stem cell regeneration. Development 2010, 137, 4135–4145. [Google Scholar] [CrossRef]
- Taniguchi, K.; Wu, L.-W.; Grivennikov, S.I.; de Jong, P.R.; Lian, I.; Yu, F.-X.; Wang, K.; Ho, S.B.; Boland, B.S.; Chang, J.T.; et al. A gp130-Src-YAP module links inflammation to epithelial regeneration. Nature 2015, 519, 57–62. [Google Scholar] [CrossRef]
- Grivennikov, S.; Karin, E.; Terzic, J.; Mucida, D.; Yu, G.-Y.; Vallabhapurapu, S.; Scheller, J.; Rose-John, S.; Cheroutre, H.; Eckmann, L.; et al. IL-6 and Stat3 are required for survival of intestinal epithelial cells and development of colitis-associated cancer. Cancer Cell 2009, 15, 103–113. [Google Scholar] [CrossRef]
- Grivennikov, S.I.; Wang, K.; Mucida, D.; Stewart, C.A.; Schnabl, B.; Jauch, D.; Taniguchi, K.; Yu, G.-Y.; Osterreicher, C.H.; Hung, K.E.; et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 2012, 491, 254–258. [Google Scholar] [CrossRef]
- Liu, J.; Duan, Y.; Cheng, X.; Chen, X.; Xie, W.; Long, H.; Lin, Z.; Zhu, B. IL-17 is associated with poor prognosis and promotes angiogenesis via stimulating VEGF production of cancer cells in colorectal carcinoma. Biochem. Biophys. Res. Commun. 2011, 407, 348–354. [Google Scholar] [CrossRef]
- Tosolini, M.; Kirilovsky, A.; Mlecnik, B.; Fredriksen, T.; Mauger, S.; Bindea, G.; Berger, A.; Bruneval, P.; Fridman, W.-H.; Pagès, F.; et al. Clinical impact of different classes of infiltrating t cytotoxic and helper cells (Th1, Th2, Treg, Th17) in patients with colorectal cancer. Cancer Res. 2011, 71, 1263–1271. [Google Scholar] [CrossRef]
- Schetter, A.J.; Nguyen, G.H.; Bowman, E.D.; Mathé, E.A.; Yuen, S.T.; Hawkes, J.E.; Croce, C.M.; Leung, S.Y.; Harris, C.C. Association of inflammation-related and microRNA gene expression with cancer-specific mortality of colon adenocarcinoma. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 5878–5887. [Google Scholar] [CrossRef]
- Calon, A.; Espinet, E.; Palomo-Ponce, S.; Tauriello, D.V.F.; Iglesias, M.; Céspedes, M.V.; Sevillano, M.; Nadal, C.; Jung, P.; Zhang, X.H.-F.; et al. Dependency of colorectal cancer on a TGF-beta-driven programme in stromal cells for metastasis initiation. Cancer Cell 2012, 22, 571–584. [Google Scholar] [CrossRef]
- Zhang, Y.; Turkson, J.; Carter-Su, C.; Smithgall, T.; Levitzki, A.; Kraker, A.; Krolewski, J.J.; Medveczky, P.; Jove, R. Activation of Stat3 in v-Src-transformed fibroblasts requires cooperation of Jak1 kinase activity. J. Biol. Chem. 2000, 275, 24935–24944. [Google Scholar] [CrossRef]
- Phesse, T.J.; Buchert, M.; Stuart, E.; Flanagan, D.J.; Faux, M.; Afshar-Sterle, S.; Walker, F.; Zhang, H.-H.; Nowell, C.J.; Jorissen, R.; et al. Partial inhibition of gp130-Jak-Stat3 signaling prevents Wnt–β-catenin–mediated intestinal tumor growth and regeneration. Sci. Signal. 2014, 7, ra92. [Google Scholar] [CrossRef]
- Ernst, M.; Phesse, T.J.; Thiem, S.; Buchert, M. 78: Therapeutic inhibition of Gp130/Jak/Stat3-dependent cytokine signaling suppresses WNT-dependent colon cancer formation. Cytokine 2013, 63, 261. [Google Scholar] [CrossRef]
- Naka, T.; Narazaki, M.; Hirata, M.; Matsumoto, T.; Minamoto, S.; Aono, A.; Nishimoto, N.; Kajita, T.; Taga, T.; Yoshizaki, K.; et al. Structure and function of a new STAT-induced STAT inhibitor. Nature 1997, 387, 924–929. [Google Scholar] [CrossRef]
- Starr, R.; Willson, T.A.; Viney, E.M.; Murray, L.J.; Rayner, J.R.; Jenkins, B.J.; Gonda, T.J.; Alexander, W.S.; Metcalf, D.; Nicola, N.A.; et al. A family of cytokine-inducible inhibitors of signalling. Nature 1997, 387, 917–921. [Google Scholar] [CrossRef]
- Endo, T.A.; Masuhara, M.; Yokouchi, M.; Suzuki, R.; Sakamoto, H.; Mitsui, K.; Matsumoto, A.; Tanimura, S.; Ohtsubo, M.; Misawa, H.; et al. A new protein containing an SH2 domain that inhibits JAK kinases. Nature 1997, 387, 921–924. [Google Scholar] [CrossRef]
- Metcalf, D.; Greenhalgh, C.J.; Viney, E.; Willson, T.A.; Starr, R.; Nicola, N.A.; Hilton, D.J.; Alexander, W.S. Gigantism in mice lacking suppressor of cytokine signalling-2. Nature 2000, 405, 1069–1073. [Google Scholar] [CrossRef]
- Lee, S.; Cho, N.-Y.; Yoo, E.J.; Kim, J.H.; Kang, G.H. CpG island methylator phenotype in colorectal cancers: Comparison of the new and classic CpG island methylator phenotype marker panels. Arch. Pathol. Lab. Med. 2008, 132, 1657–1665. [Google Scholar]
- Hashimoto, M.; Ayada, T.; Kinjyo, I.; Hiwatashi, K.; Yoshida, H.; Okada, Y.; Kobayashi, T.; Yoshimura, A. Silencing of SOCS1 in macrophages suppresses tumor development by enhancing antitumor inflammation. Cancer Sci. 2009, 100, 730–736. [Google Scholar] [CrossRef]
- Tobelaim, W.S.; Beaurivage, C.; Champagne, A.; Pomerleau, V.; Simoneau, A.; Chababi, W.; Yeganeh, M.; Thibault, P.; Klinck, R.; Carrier, J.C.; et al. Tumour-promoting role of SOCS1 in colorectal cancer cells. Sci. Rep. 2015, 5, 14301. [Google Scholar] [CrossRef]
- Hanada, T.; Kobayashi, T.; Chinen, T.; Saeki, K.; Takaki, H.; Koga, K.; Minoda, Y.; Sanada, T.; Yoshioka, T.; Mimata, H.; et al. IFNγ-dependent, spontaneous development of colorectal carcinomas in SOCS1-deficient mice. J. Exp. Med. 2006, 203, 1391–1397. [Google Scholar] [CrossRef]
- Newton, V.A.; Ramocki, N.M.; Scull, B.P.; Simmons, J.G.; McNaughton, K.; Lund, P.K. Suppressor of cytokine signaling-2 gene disruption promotes ApcMin/+ tumorigenesis and activator protein-1 activation. Am. J. Pathol. 2010, 176, 2320–2332. [Google Scholar] [CrossRef]
- Zhang, L.; Li, J.; Li, L.; Zhang, J.; Wang, X.; Yang, C.; Li, Y.; Lan, F.; Lin, P. IL-23 selectively promotes the metastasis of colorectal carcinoma cells with impaired Socs3 expression via the STAT5 pathway. Carcinogenesis 2014, 35, 1330–1340. [Google Scholar] [CrossRef]
- Rigby, R.J.; Simmons, J.G.; Greenhalgh, C.J.; Alexander, W.S.; Lund, P.K. Suppressor of cytokine signaling 3 (SOCS3) limits damage-induced crypt hyper-proliferation and inflammation-associated tumorigenesis in the colon. Oncogene 2007, 26, 4833–4841. [Google Scholar] [CrossRef]
- Dong, X.; Wang, J.; Tang, B.; Hao, Y.-X.; Li, P.-Y.; Li, S.-Y.; Yu, P.-W. The role and gene expression profile of SOCS3 in colorectal carcinoma. Oncotarget 2017, 9, 15984–15996. [Google Scholar] [CrossRef]
- Pilling, C.; Cooper, J.A. SOCS2 binds to and regulates EphA2 through multiple mechanisms. Sci. Rep. 2017, 7, 10838. [Google Scholar] [CrossRef]
- Shin, D.S.; Zaretsky, J.M.; Escuin-Ordinas, H.; Garcia-Diaz, A.; Hu-Lieskovan, S.; Kalbasi, A.; Grasso, C.S.; Hugo, W.; Sandoval, S.; Torrejon, D.Y.; et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov. 2017, 7, 188–201. [Google Scholar] [CrossRef] [PubMed]

| Adaptor | Status in CRC | References |
|---|---|---|
| SLAP | Downregulation | [56] |
| [54] | ||
| SOCS1 | Downregulation | [57] |
| Hypermethylation | [58,59,60,61,62] | |
| SNPs association with CRC | [63] | |
| Downregulation by miRNA | [64] | |
| SOCS2 | SNPs association with CRC | [63] |
| Downregulation | [65] | |
| Upregulation | [66] | |
| SOCS3 | Hypermethylation | [67] |
| Downregulation | [68,69] | |
| Downregulation by miRNA | [64] | |
| SOCS5 | Downregulation by miRNA | [70,71] |
| SOCS6 | Gene loss | [72] |
| Hypermethylation | [73] | |
| Downregulation | [65] | |
| Downregulation by miRNA | [70,74] | |
| SOCS7 | Hypermethylation | [67] |
| Downregulation by miRNA | [70] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mevizou, R.; Sirvent, A.; Roche, S. Control of Tyrosine Kinase Signalling by Small Adaptors in Colorectal Cancer. Cancers 2019, 11, 669. https://doi.org/10.3390/cancers11050669
Mevizou R, Sirvent A, Roche S. Control of Tyrosine Kinase Signalling by Small Adaptors in Colorectal Cancer. Cancers. 2019; 11(5):669. https://doi.org/10.3390/cancers11050669
Chicago/Turabian StyleMevizou, Rudy, Audrey Sirvent, and Serge Roche. 2019. "Control of Tyrosine Kinase Signalling by Small Adaptors in Colorectal Cancer" Cancers 11, no. 5: 669. https://doi.org/10.3390/cancers11050669
APA StyleMevizou, R., Sirvent, A., & Roche, S. (2019). Control of Tyrosine Kinase Signalling by Small Adaptors in Colorectal Cancer. Cancers, 11(5), 669. https://doi.org/10.3390/cancers11050669






