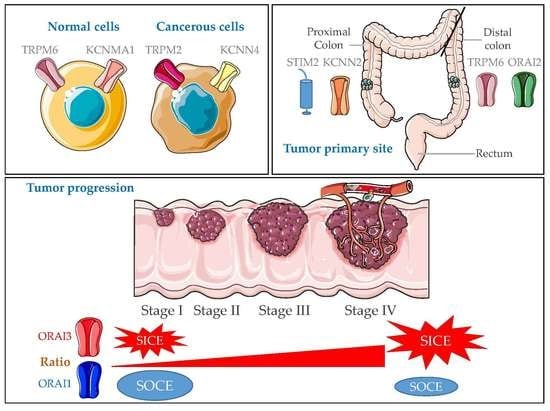
Expression Profiling of Calcium Channels and Calcium-Activated Potassium Channels in Colorectal Cancer
Abstract
:1. Introduction
2. Results
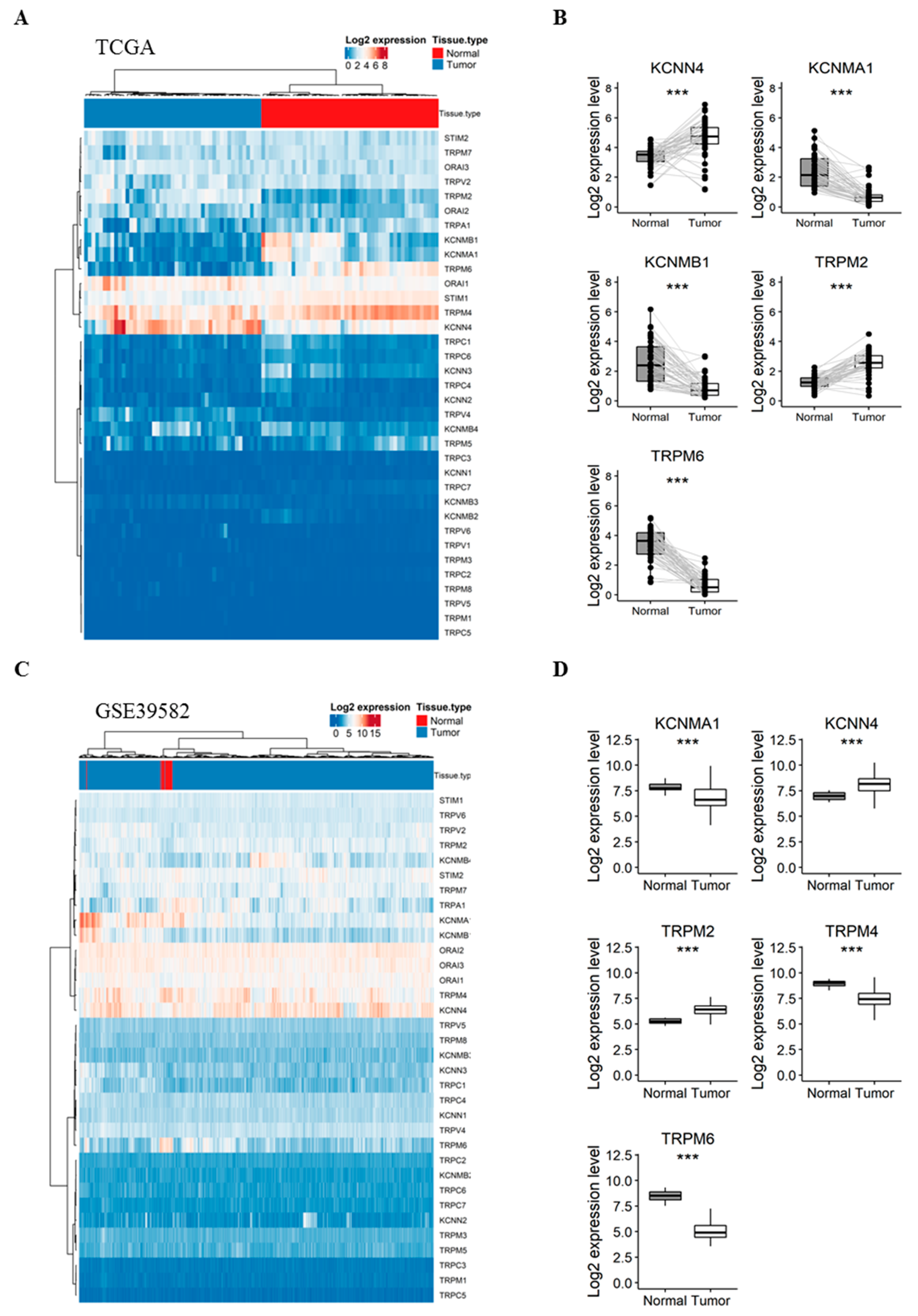
2.1. Gene Expression of KCa and Ca2+ Channels in Normal Mucosa versus CRC Tissues
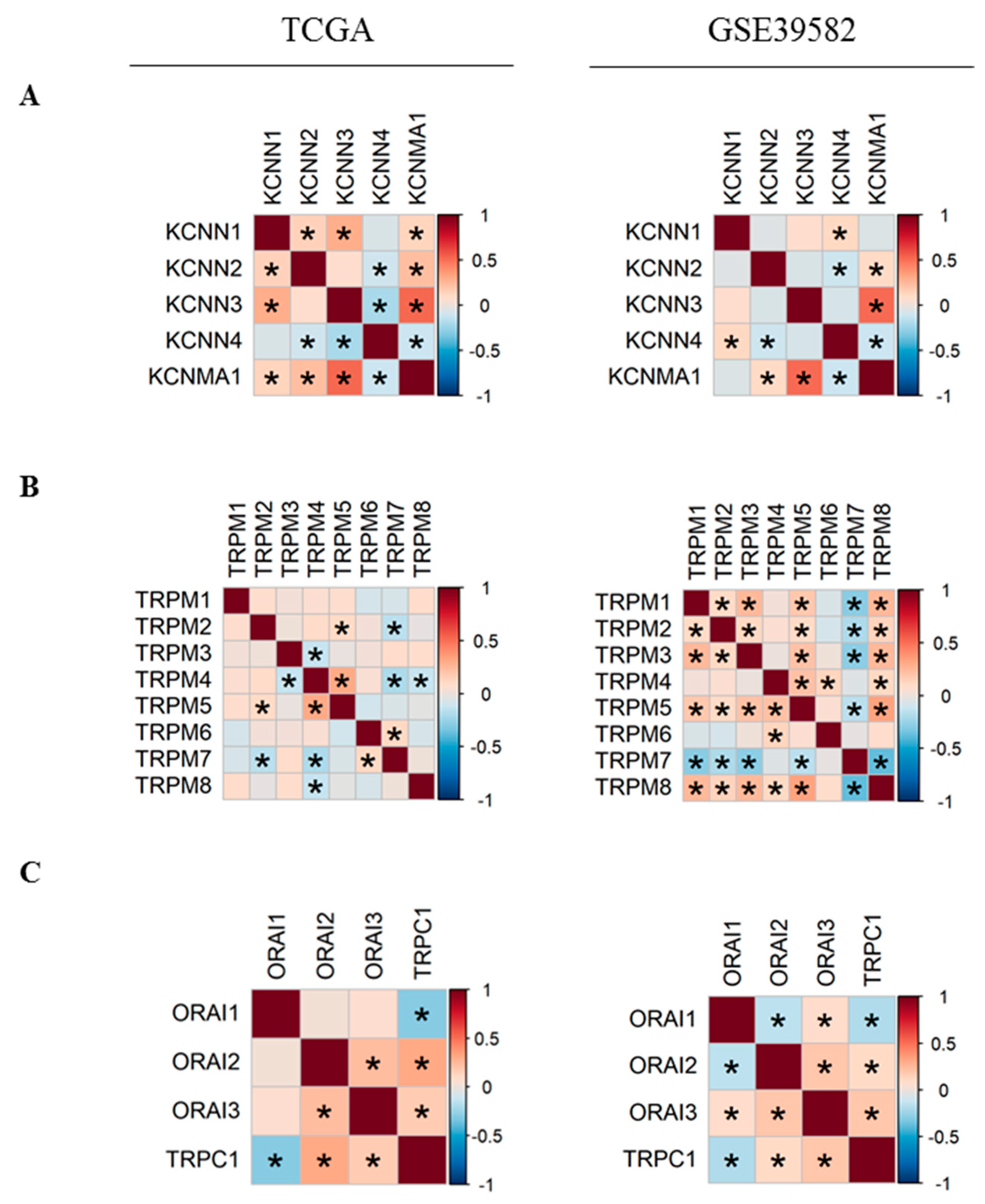
2.2. Co-Expression of Ca2+ and KCa Channels in CRC
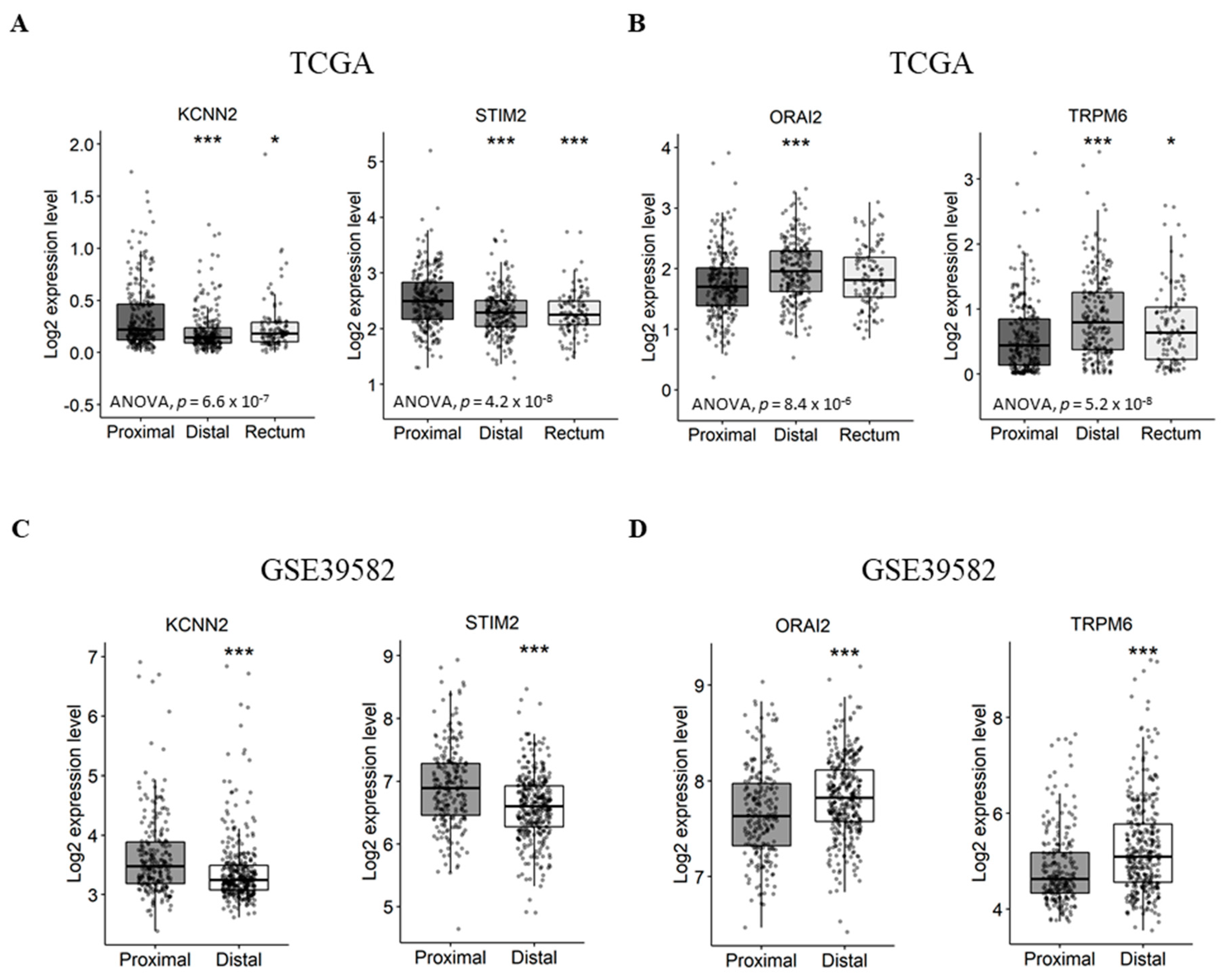
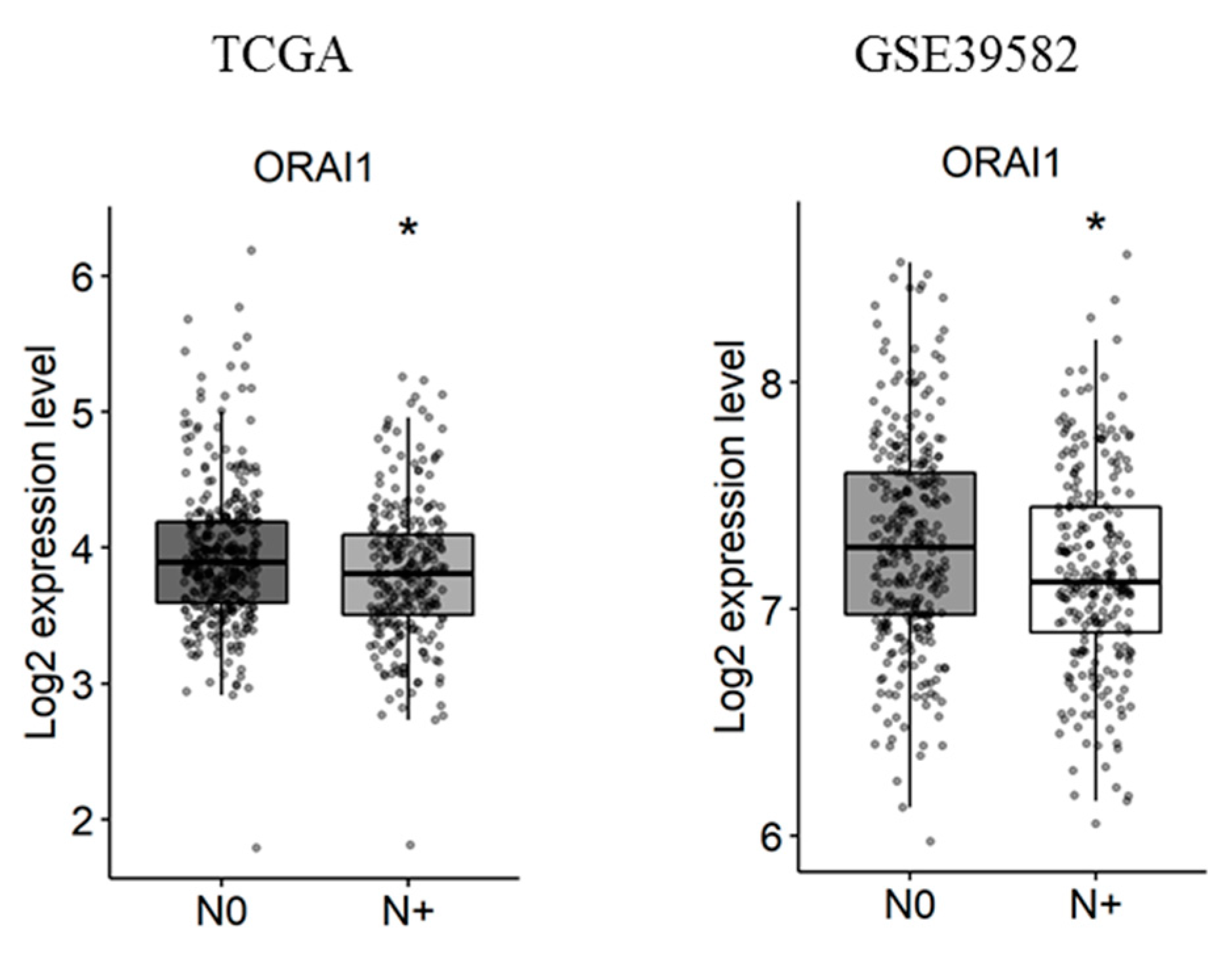
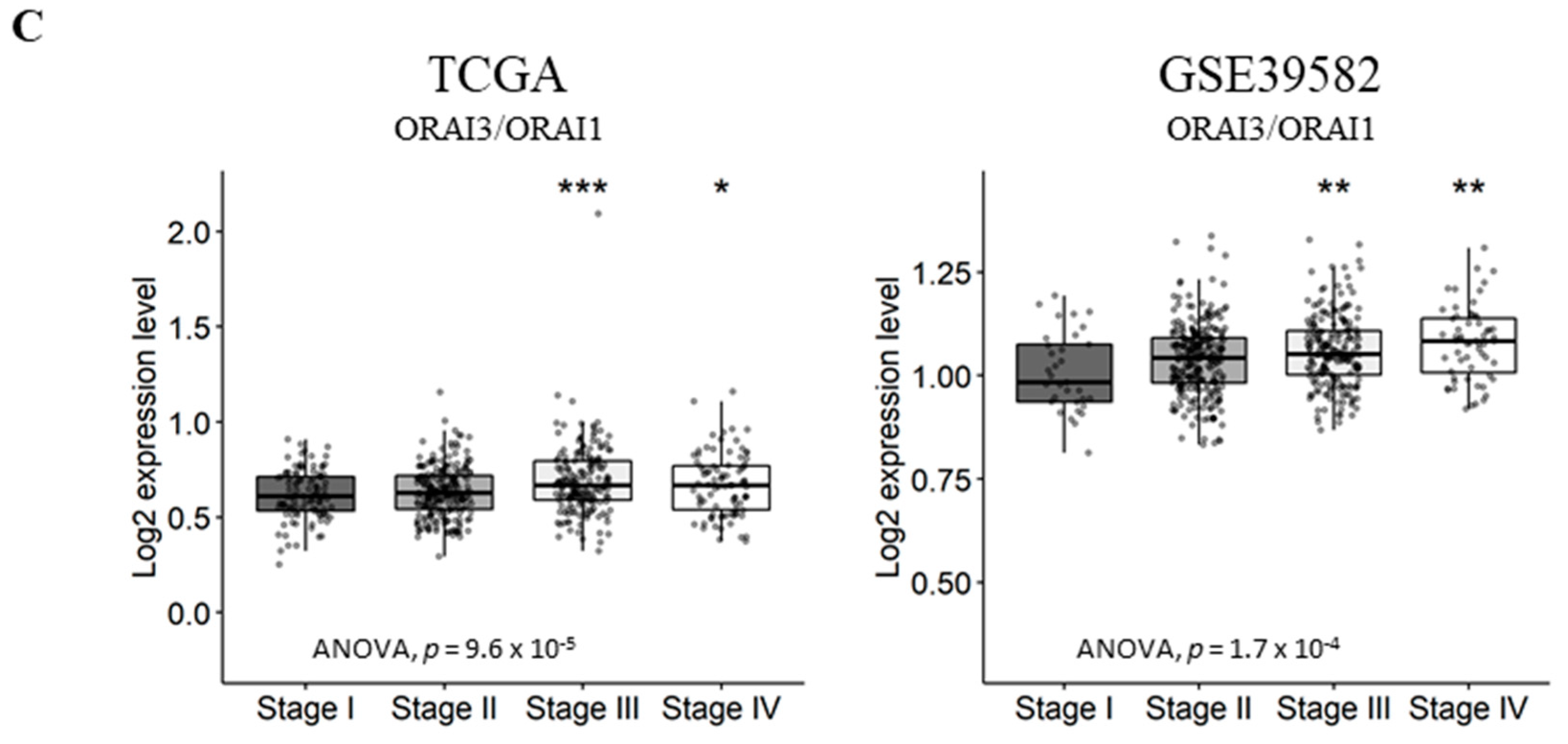
2.3. Gene Expression Analysis According to Tumor Primary Site, Lymph Node Metastases and Tumor Stage
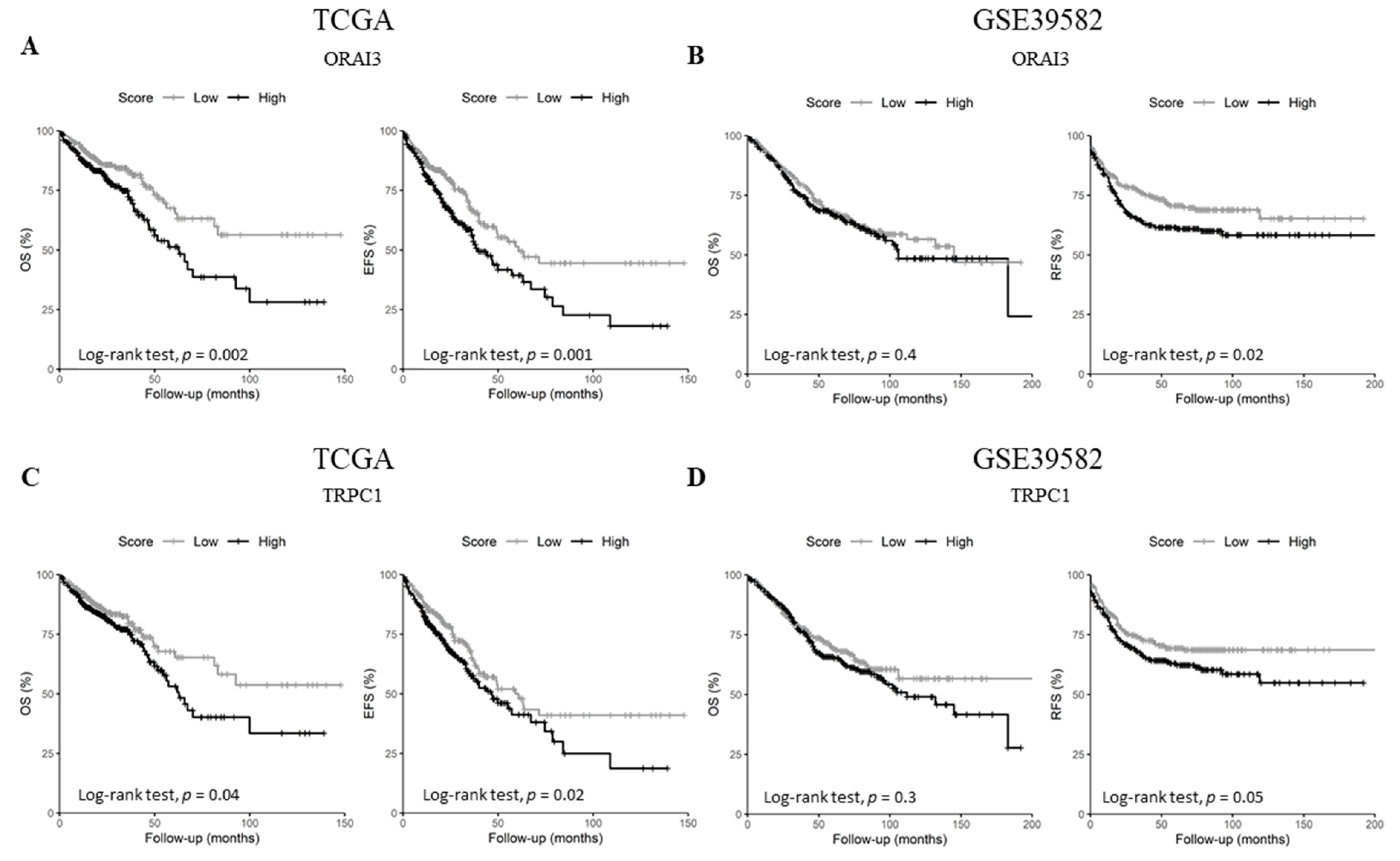
2.4. ORAI3 and TRPC1 Expression Predict Poor Prognosis of CRC Patients
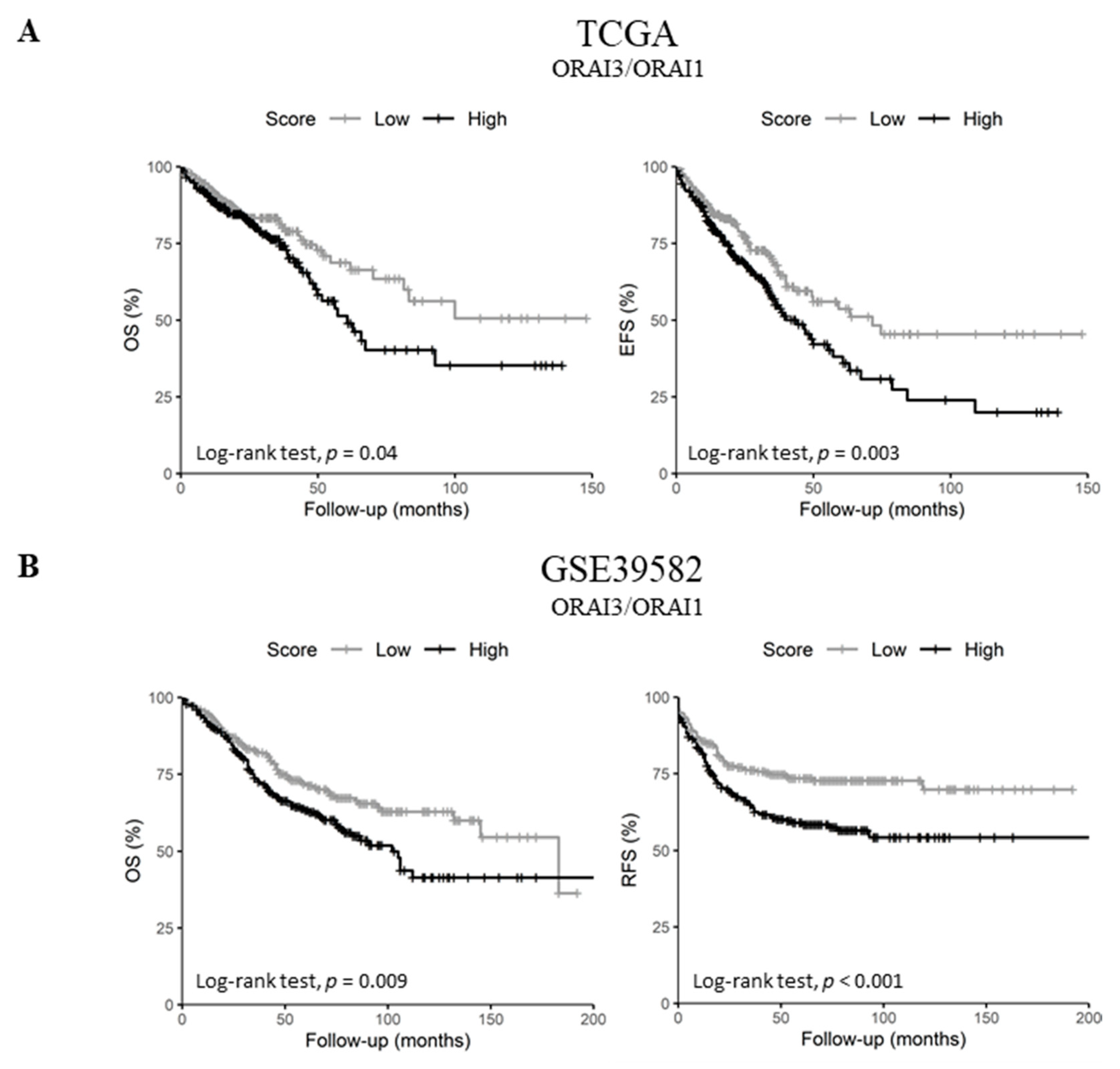
2.5. Upregulated ORAI3/ORAI1 Ratio Predicts Poor Prognosis of CRC Patients
3. Discussion
4. Materials and Methods
4.1. Public Datasets
4.2. Statistical Analysis
4.3. Survival Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef]
- Morris, E.J.A.; Forman, D.; Thomas, J.D.; Quirke, P.; Taylor, E.F.; Fairley, L.; Cottier, B.; Poston, G. Surgical management and outcomes of colorectal cancer liver metastases. Br. J. Surg. 2010, 97, 1110–1118. [Google Scholar] [CrossRef]
- Schwab, A.; Stock, C. Ion channels and transporters in tumour cell migration and invasion. Philos. Trans. R. Soc. 2014, 369, 1–8. [Google Scholar] [CrossRef]
- Déliot, N.; Constantin, B. Plasma membrane calcium channels in cancer: Alterations and consequences for cell proliferation and migration. Biochim. Biophys. Acta Biomembr. 2015, 1848, 2512–2522. [Google Scholar] [CrossRef]
- Prevarskaya, N.; Skryma, R.; Shuba, Y. Calcium in tumour metastasis: New roles for known actors. Nat. Rev. Cancer 2011, 11, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Pan, H.; Yao, J.; Zhou, Y.; Han, W. SOCE and cancer: Recent progress and new perspectives. Int. J. Cancer 2016, 138, 2067–2077. [Google Scholar] [CrossRef] [PubMed]
- Putney, J.W. A model for receptor-regulated calcium entry. Cell Calcium 1986, 7, 1–12. [Google Scholar] [CrossRef]
- Berridge, M.J.; Lipp, P.; Bootman, M.D. The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 2000, 1, 11–21. [Google Scholar] [CrossRef]
- Prakriya, M.; Lewis, R.S. Store-Operated Calcium Channels. Physiol. Rev. 2015, 95, 1383–1436. [Google Scholar] [CrossRef]
- Mignen, O.; Constantin, B.; Potier-Cartereau, M.; Penna, A.; Gautier, M.; Guéguinou, M.; Renaudineau, Y.; Shoji, K.F.; Félix, R.; Bayet, E.; et al. Constitutive calcium entry and cancer: Updated views and insights. Eur. Biophys. J. 2017, 46, 395–413. [Google Scholar] [CrossRef]
- Shuttleworth, T.J.; Thompson, J.L. Discriminating between capacitative and arachidonate-activated Ca(2+) entry pathways in HEK293 cells. J. Biol. Chem. 1999, 274, 31174–31178. [Google Scholar] [CrossRef]
- Mignen, O.; Thompson, J.L.; Yule, D.I.; Shuttleworth, T.J. Agonist activation of arachidonate-regulated Ca2+-selective (ARC) channels in murine parotid and pancreatic acinar cells. J. Physiol. 2005, 564, 791–801. [Google Scholar] [CrossRef] [PubMed]
- Mignen, O.; Thompson, J.L.; Shuttleworth, T.J. Both Orai1 and Orai3 are essential components of the arachidonate-regulated Ca2+-selective (ARC) channels. J. Physiol. 2008, 586, 185–195. [Google Scholar] [CrossRef]
- Mignen, O.; Thompson, J.L.; Shuttleworth, T.J. STIM1 regulates Ca2+ entry via arachidonate-regulated Ca2+-selective (ARC) channels without store depletion or translocation to the plasma membrane. J. Physiol. 2007, 579, 703–715. [Google Scholar] [CrossRef]
- Vergara, C.; Latorre, R.; Marrion, N.V.; Adelman, J.P. Calcium-activated potassium channels. Curr. Opin. Neurobiol. 1998, 8, 321–329. [Google Scholar] [CrossRef]
- Guéguinou, M.; Gambade, A.; Félix, R.; Chantôme, A.; Fourbon, Y.; Bougnoux, P.; Weber, G.; Potier-Cartereau, M.; Vandier, C. Lipid rafts, KCa/ClCa/Ca2+ channel complexes and EGFR signaling: Novel targets to reduce tumor development by lipids? Biochim. Biophys. Acta Biomembr. 2015, 1848, 2603–2620. [Google Scholar] [CrossRef] [PubMed]
- Sobradillo, D.; Hernández-Morales, M.; Ubierna, D.; Moyer, M.P.; Núñez, L.; Villalobos, C. A reciprocal shift in transient receptor potential channel 1 (TRPC1) and stromal interaction molecule 2 (STIM2) contributes to Ca2+ remodeling and cancer hallmarks in colorectal carcinoma cells. J. Biol. Chem. 2014, 289, 28765–28782. [Google Scholar] [CrossRef] [PubMed]
- Villalobos, C.; Sobradillo, D.; Hernández-Morales, M.; Núñez, L. Calcium remodeling in colorectal cancer. Biochim. Biophys. Acta Mol. Cell Res. 2017, 1864, 843–849. [Google Scholar] [CrossRef] [PubMed]
- Harnois, T.; Crottes, D.; Uguen, A.; Deliot, N.; Gambade, A.; Chantôme, A.; Haelters, J.P.; Jaffrès, P.A.; Jourdan, M.L.; Weber, G.; et al. SK3/TRPC1/Orai1 complex regulates SOCE-dependent colon cancer cell migration: A novel opportunity to modulate anti-EGFR mAb action by the alkyl-lipid Ohmline. Oncotarget 2016, 7, 36168–36184. [Google Scholar]
- Gueguinou, M.; Crottès, D.; Chantôme, A.; Rapetti-Mauss, R.; Potier-Cartereau, M.; Clarysse, L.; Girault, A.; Fourbon, Y.; Jézéquel, P.; Guérin-Charbonnel, C.; et al. The SigmaR1 chaperone drives breast and colorectal cancer cell migration by tuning SK3-dependent Ca2+ homeostasis. Oncogene 2017, 36, 3640–3647. [Google Scholar] [CrossRef]
- Hernández-Morales, M.; Sobradillo, D.; Valero, R.A.; Muñoz, E.; Ubierna, D.; Moyer, M.P.; Núñez, L.; Villalobos, C.; Hernández-Morales, M.; Sobradillo, D.; et al. Mitochondria sustain store-operated currents in colon cancer cells but not in normal colonic cells: Reversal by non-steroidal anti-inflammatory drugs. Oncotarget 2017, 8, 55332–55352. [Google Scholar] [CrossRef]
- Zuccolo, E.; Laforenza, U.; Ferulli, F.; Pellavio, G.; Scarpellino, G.; Tanzi, M.; Turin, I.; Faris, P.; Lucariello, A.; Maestri, M.; et al. Stim and Orai mediate constitutive Ca2+ entry and control endoplasmic reticulum Ca2+ refilling in primary cultures of colorectal carcinoma cells. Oncotarget 2018, 9, 31098–31119. [Google Scholar] [CrossRef]
- Dubois, C.; Abeele, F.V.; Lehen’kyi, V.; Gkika, D.; Guarmit, B.; Lepage, G.; Slomianny, C.; Borowiec, A.S.; Bidaux, G.; Benahmed, M.; et al. Remodeling of Channel-Forming ORAI Proteins Determines an Oncogenic Switch in Prostate Cancer. Cancer Cell 2014, 26, 19–32. [Google Scholar] [CrossRef]
- Azimi, I.; Milevskiy, M.; Chalmers, S.; Yapa, K.; Robitaille, M.; Henry, C.; Baillie, G.; Thompson, E.; Roberts-Thomson, S.; Monteith, G.; et al. ORAI1 and ORAI3 in Breast Cancer Molecular Subtypes and the Identification of ORAI3 as a Hypoxia Sensitive Gene and a Regulator of Hypoxia Responses. Cancers 2019, 11, 208. [Google Scholar] [CrossRef]
- Litan, A.; Langhans, S.A. Cancer as a channelopathy: Ion channels and pumps in tumor development and progression. Front. Cell. Neurosci. 2015, 9, 86. [Google Scholar] [CrossRef] [PubMed]
- Prevarskaya, N.; Skryma, R.; Shuba, Y. Ion Channels in Cancer: Are Cancer Hallmarks Oncochannelopathies? Physiol. Rev. 2018, 98, 559–621. [Google Scholar] [CrossRef] [PubMed]
- Basile, M.S.; Fagone, P.; Mangano, K.; Mammana, S.; Magro, G.; Salvatorelli, L.; Destri, G.L.; la Greca, G.; Nicoletti, F.; Puleo, S.; et al. KCNMA1 Expression is Downregulated in Colorectal Cancer via Epigenetic Mechanisms. Cancers 2019, 11, 245. [Google Scholar] [CrossRef] [PubMed]
- Ousingsawat, J.; Spitzner, M.; Schreiber, R.; Kunzelmann, K. Upregulation of colonic ion channels in APC Min/+ mice. Pflügers Arch. Eur. J. Physiol. 2008, 456, 847–855. [Google Scholar] [CrossRef] [PubMed]
- Mohr, C.J.; Steudel, F.A.; Gross, D.; Ruth, P.; Lo, W.-Y.; Hoppe, R.; Schroth, W.; Brauch, H.; Huber, S.M.; Lukowski, R.; et al. Cancer-Associated Intermediate Conductance Ca2+-Activated K+ Channel KCa3.1. Cancers 2019, 11, 109. [Google Scholar] [CrossRef]
- Köhler, R.; Wulff, H.; Eichler, I.; Kneifel, M.; Neumann, D.; Knorr, A.; Grgic, I.; Kämpfe, D.; Si, H.; Wibawa, J.; et al. Blockade of the Intermediate-Conductance Calcium-Activated Potassium Channel as a New Therapeutic Strategy for Restenosis. Circulation 2003, 108, 1119–1125. [Google Scholar] [CrossRef]
- De Marchi, U.; Sassi, N.; Fioretti, B.; Catacuzzeno, L.; Cereghetti, G.M.; Szabò, I.; Zoratti, M. Intermediate conductance Ca2+-activated potassium channel (KCa3.1) in the inner mitochondrial membrane of human colon cancer cells. Cell Calcium 2009, 45, 509–516. [Google Scholar] [CrossRef] [PubMed]
- Lai, W.; Liu, L.; Zeng, Y.; Wu, H.; Xu, H.; Chen, S.; Chu, Z. KCNN4 Channels participate in the EMT induced by PRL-3 in colorectal cancer. Med. Oncol. 2013, 30, 566. [Google Scholar] [CrossRef]
- Luongo, F.; Pietropaolo, G.; Gautier, M.; Dhennin-Duthille, I.; Ouadid-Ahidouch, H.; Wolf, F.I.; Trapani, V. TRPM6 is Essential for Magnesium Uptake and Epithelial Cell Function in the Colon. Nutrients 2018, 10, 784. [Google Scholar] [CrossRef]
- Castiglioni, S.; Cazzaniga, A.; Trapani, V.; Cappadone, C.; Farruggia, G.; Merolle, L.; Wolf, F.I.; Iotti, S.; Maier, J.A.M. Magnesium homeostasis in colon carcinoma LoVo cells sensitive or resistant to doxorubicin. Sci. Rep. 2015, 5, 16538. [Google Scholar] [CrossRef]
- Miller, B.A. TRPM2 in Cancer. Cell Calcium 2019. [Google Scholar] [CrossRef] [PubMed]
- Csanády, L.; Törőcsik, B. Four Ca2+ Ions Activate TRPM2 Channels by Binding in Deep Crevices near the Pore but Intracellularly of the Gate. J. Gen. Physiol. 2009, 133, 189. [Google Scholar] [CrossRef]
- Du, J.; Xie, J.; Yue, L. Intracellular calcium activates TRPM2 and its alternative spliced isoforms. Proc. Natl. Acad. Sci. USA 2009, 106, 7239–7244. [Google Scholar] [CrossRef] [PubMed]
- Sadeghi, M.R.; Jeddi, F.; Soozangar, N.; Somi, M.H.; Samadi, N. The role of Nrf2-Keap1 axis in colorectal cancer, progression and chemoresistance. Tumor Biol. 2017, 39, 101042831770551. [Google Scholar] [CrossRef]
- Imamura, T.; Kikuchi, H.; Herraiz, M.-T.; Park, D.-Y.; Mizukami, Y.; Mino-Kenduson, M.; Lynch, M.P.; Rueda, B.R.; Benita, Y.; Xavier, R.J.; et al. HIF-1alpha and HIF-2alpha have divergent roles in colon cancer. Int. J. Cancer 2009, 124, 763–771. [Google Scholar] [CrossRef]
- Benedix, F.; Kube, R.; Meyer, F.; Schmidt, U.; Gastinger, I.; Lippert, H. Colon/Rectum Carcinomas (Primary Tumor) Study Group, Comparison of 17,641 patients with right- and left-sided colon cancer: Differences in epidemiology, perioperative course, histology and survival. Dis. Colon Rectum 2010, 53, 57–64. [Google Scholar] [CrossRef]
- Missiaglia, E.; Jacobs, B.; D’Ario, G.; di Narzo, A.F.; Soneson, C.; Budinska, E.; Popovici, V.; Vecchione, L.; Gerster, S.; Yan, P.; et al. Distal and proximal colon cancers differ in terms of molecular, pathological and clinical features. Ann. Oncol. 2014, 25, 1995–2001. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Wang, J.; Zhang, J.; Cai, J.; Bai, Z.; Zhang, Z. Orai1, a Direct Target of microRNA-519, Promotes Progression of Colorectal Cancer via Akt/GSK3β Signaling Pathway. Dig. Dis. Sci. 2016, 61, 1553–1560. [Google Scholar] [CrossRef] [PubMed]
- Marisa, L.; de Reyniès, A.; Duval, A.; Selves, J.; Gaub, M.P.; Vescovo, L.; Etienne-Grimaldi, M.-C.; Schiappa, R.; Guenot, D.; Ayadi, M.; et al. Gene expression classification of colon cancer into molecular subtypes: Characterization, validation and prognostic value. PLoS Med. 2013, 10, e1001453. [Google Scholar] [CrossRef]
- Grossman, R.L.; Heath, A.P.; Ferretti, V.; Varmus, H.E.; Lowy, D.R.; Kibbe, W.A.; Staudt, L.M. Toward a Shared Vision for Cancer Genomic Data. N. Engl. J. Med. 2016, 375, 1109–1112. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to MultipleTesting. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Schoenfeld, D. Biometrika Trust Partial Residuals for The Proportional Hazards Regression Model Author(s): David Schoenfeld Published by: Oxford University Press on Behalf of Biometrika Trust Stable. Biometrika 1982, 69, 239–241. Available online: http://www.jstor.org/stable/2335876 (accessed on 8 April 2016). [CrossRef]
| Variable | Hazard Ratio | 95% CI | Wald’s p-Value |
|---|---|---|---|
| TCGA (OS n = 602) | |||
| ORAI3 (High vs. Low) | 1.72 | (1.21–2.45) | 0.003 |
| TRPC1 (High vs. Low) | 1.44 | (1.01–2.06) | 0.042 |
| TCGA (EFS n = 602) | |||
| ORAI3 (High vs. Low) | 1.6 | (1.2–2.13) | 0.001 |
| TRPC1 (High vs. Low) | 1.39 | (1.04–1.85) | 0.025 |
| GSE39582 (OS n = 562) | |||
| ORAI3 (High vs. Low) | 1.12 | (0.84–1.48) | 0.443 |
| TRPC1 (High vs. Low) | 1.18 | (0.89–1.57) | 0.258 |
| GSE39582 (RFS n = 557) | |||
| ORAI3 (High vs. Low) | 1.43 | (1.06–1.92) | 0.02 |
| TRPC1 (High vs. Low) | 1.34 | (1–1.81) | 0.05 |
| Variable | Hazard Ratio | 95% CI | Wald’s p-Value |
|---|---|---|---|
| TCGA (OS n = 602) | |||
| ORAI3/1 ratio (High vs. Low) | 1.45 | (1.02–2.07) | 0.037 |
| TCGA (EFS n = 602) | |||
| ORAI3/1 ratio (High vs. Low) | 1.54 | (1.15–2.06) | 0.003 |
| GSE39582 (OS n = 562) | |||
| ORAI3/1 ratio (High vs. Low) | 1.47 | (1.1–1.95) | 0.009 |
| GSE39582 (RFS n = 557) | |||
| ORAI3/1 ratio (High vs. Low) | 1.69 | (1.25–2.28) | 0.001 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ibrahim, S.; Dakik, H.; Vandier, C.; Chautard, R.; Paintaud, G.; Mazurier, F.; Lecomte, T.; Guéguinou, M.; Raoul, W. Expression Profiling of Calcium Channels and Calcium-Activated Potassium Channels in Colorectal Cancer. Cancers 2019, 11, 561. https://doi.org/10.3390/cancers11040561
Ibrahim S, Dakik H, Vandier C, Chautard R, Paintaud G, Mazurier F, Lecomte T, Guéguinou M, Raoul W. Expression Profiling of Calcium Channels and Calcium-Activated Potassium Channels in Colorectal Cancer. Cancers. 2019; 11(4):561. https://doi.org/10.3390/cancers11040561
Chicago/Turabian StyleIbrahim, Sajida, Hassan Dakik, Christophe Vandier, Romain Chautard, Gilles Paintaud, Frédéric Mazurier, Thierry Lecomte, Maxime Guéguinou, and William Raoul. 2019. "Expression Profiling of Calcium Channels and Calcium-Activated Potassium Channels in Colorectal Cancer" Cancers 11, no. 4: 561. https://doi.org/10.3390/cancers11040561
APA StyleIbrahim, S., Dakik, H., Vandier, C., Chautard, R., Paintaud, G., Mazurier, F., Lecomte, T., Guéguinou, M., & Raoul, W. (2019). Expression Profiling of Calcium Channels and Calcium-Activated Potassium Channels in Colorectal Cancer. Cancers, 11(4), 561. https://doi.org/10.3390/cancers11040561