MicroRNA in Lung Cancer Metastasis
Abstract
1. Introduction
2. EMT: The Key Mechanism of Lung Cancer Metastasis
3. MicroRNAs Regulate EMT in Lung Cancer
3.1. SNAI Family TF-Related miRNAs
3.2. ZEB1/ZEB2 TF-Related miRNAs
3.3. Twist TF-Related miRNAs
3.4. MiRNAs Modulate Other EMT-Associated Signaling Genes and Related Downstream Proteins
4. Role of miRNAs in Different Metastasis Sites (Bone, Brain and Lymph Nodes) in Lung Cancer
4.1. Role of miRNAs in Lung Cancer Bone Metastasis
4.2. Role of miRNAs in Lung Cancer Brain Metastasis
4.3. Role of miRNAs in Lung Cancer Lymph Node Metastasis
5. Potential of miRNA as a Therapeutic Target and Tool in Patients with Lung Cancer
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef] [PubMed]
- The National Lung Screening Trial Research Team; Church, T.R.; Black, W.C.; Aberle, D.R.; Berg, C.D.; Clingan, K.L.; Duan, F.; Fagerstrom, R.M.; Gareen, I.F.; Gierada, D.S.; et al. Results of initial low-dose computed tomographic screening for lung cancer. N. Engl. J. Med. 2013, 368, 1980–1991. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P.; Acloque, H.; Huang, R.Y.; Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef] [PubMed]
- Krol, J.; Loedige, I.; Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 2010, 11, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Lagos-Quintana, M.; Rauhut, R.; Yalcin, A.; Meyer, J.; Lendeckel, W.; Tuschl, T. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 2002, 12, 735–739. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Ruvkun, G. Clarifications on miRNA and cancer. Science 2006, 311, 36–37. [Google Scholar] [CrossRef] [PubMed]
- Stahlhut, C.; Slack, F.J. MicroRNAs and the cancer phenotype: Profiling, signatures and clinical implications. Genome Med. 2013, 5, 111. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, Y.; Tay, F.C.; Lam, D.H.; Sandanaraj, E.; Tang, C.; Ang, B.T.; Wang, S. Attenuated adenosine-to-inosine editing of microRNA-376a* promotes invasiveness of glioblastoma cells. J. Clin. Investig. 2012, 122, 4059–4076. [Google Scholar] [CrossRef] [PubMed]
- Bui, N.; Woodward, B.; Johnson, A.; Husain, H. Novel Treatment Strategies for Brain Metastases in Non-small-cell Lung Cancer. Curr. Treat. Options Oncol. 2016, 17, 25. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.G.; Sanders, A.J.; Katoh, M.; Ungefroren, H.; Gieseler, F.; Prince, M.; Thompson, S.K.; Zollo, M.; Spano, D.; Dhawan, P.; et al. Tissue invasion and metastasis: Molecular, biological and clinical perspectives. Semin. Cancer Biol. 2015, 35, S244–S275. [Google Scholar] [CrossRef] [PubMed]
- Woodhouse, E.C.; Chuaqui, R.F.; Liotta, L.A. General mechanisms of metastasis. Cancer 1997, 80, 1529–1537. [Google Scholar] [CrossRef]
- Cheung, K.J.; Ewald, A.J. A collective route to metastasis: Seeding by tumor cell clusters. Science 2016, 352, 167–169. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2002, 2, 442–454. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Weinberg, R.A. Epithelial-mesenchymal transition: At the crossroads of development and tumor metastasis. Dev. Cell 2008, 14, 818–829. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Acloque, H.; Adams, M.S.; Fishwick, K.; Bronner-Fraser, M.; Nieto, M.A. Epithelial-mesenchymal transitions: The importance of changing cell state in development and disease. J. Clin. Investig. 2009, 119, 1438–1449. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Neilson, E.G. Epithelial-mesenchymal transition and its implications for fibrosis. J. Clin. Investig. 2003, 112, 1776–1784. [Google Scholar] [CrossRef] [PubMed]
- Edme, N.; Downward, J.; Thiery, J.P.; Boyer, B. Ras induces NBT-II epithelial cell scattering through the coordinate activities of Rac and MAPK pathways. J. Cell Sci. 2002, 115, 2591–2601. [Google Scholar] [PubMed]
- Jenndahl, L.E.; Isakson, P.; Baeckstrom, D. c-erbB2-induced epithelial-mesenchymal transition in mammary epithelial cells is suppressed by cell-cell contact and initiated prior to E-cadherin downregulation. Int. J. Oncol. 2005, 27, 439–448. [Google Scholar] [CrossRef] [PubMed]
- Jeon, H.M.; Lee, J. MET: Roles in epithelial-mesenchymal transition and cancer stemness. Ann. Transl. Med. 2017, 5, 5. [Google Scholar] [CrossRef] [PubMed]
- Webb, C.P.; Taylor, G.A.; Jeffers, M.; Fiscella, M.; Oskarsson, M.; Resau, J.H.; Vande Woude, G.F. Evidence for a role of Met-HGF/SF during Ras-mediated tumorigenesis/metastasis. Oncogene 1998, 17, 2019–2025. [Google Scholar] [CrossRef] [PubMed]
- Wise, R.; Zolkiewska, A. Metalloprotease-dependent activation of EGFR modulates CD44(+)/CD24(-) populations in triple negative breast cancer cells through the MEK/ERK pathway. Breast Cancer Res. Treat. 2017, 166, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Claperon, A.; Mergey, M.; Nguyen Ho-Bouldoires, T.H.; Vignjevic, D.; Wendum, D.; Chretien, Y.; Merabtene, F.; Frazao, A.; Paradis, V.; Housset, C.; et al. EGF/EGFR axis contributes to the progression of cholangiocarcinoma through the induction of an epithelial-mesenchymal transition. J. Hepatol. 2014, 61, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Mamuya, F.A.; Duncan, M.K. aV integrins and TGF-beta-induced EMT: A circle of regulation. J. Cell. Mol. Med. 2012, 16, 445–455. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Massague, J. Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 2003, 113, 685–700. [Google Scholar] [CrossRef]
- Kokudo, T.; Suzuki, Y.; Yoshimatsu, Y.; Yamazaki, T.; Watabe, T.; Miyazono, K. Snail is required for TGFbeta-induced endothelial-mesenchymal transition of embryonic stem cell-derived endothelial cells. J. Cell Sci. 2008, 121, 3317–3324. [Google Scholar] [CrossRef] [PubMed]
- Jechlinger, M.; Grunert, S.; Beug, H. Mechanisms in epithelial plasticity and metastasis: Insights from 3D cultures and expression profiling. J. Mammary Gland Biol. Neoplasia 2002, 7, 415–432. [Google Scholar] [CrossRef] [PubMed]
- Moustakas, A.; Heldin, C.H. Signaling networks guiding epithelial-mesenchymal transitions during embryogenesis and cancer progression. Cancer Sci. 2007, 98, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Yi, B.R.; Kim, N.H.; Choi, K.C. Role of the epithelial-mesenchymal transition and its effects on embryonic stem cells. Exp. Mol. Med. 2014, 46, e108. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.H.; Kim, H.S.; Li, X.Y.; Lee, I.; Choi, H.S.; Kang, S.E.; Cha, S.Y.; Ryu, J.K.; Yoon, D.; Fearon, E.R.; et al. A p53/miRNA-34 axis regulates Snail1-dependent cancer cell epithelial-mesenchymal transition. J. Cell Biol. 2011, 195, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Jia, Z.; Zhang, Y.; Xu, Q.; Guo, W.; Guo, A. miR-126 suppresses epithelial-to-mesenchymal transition and metastasis by targeting PI3K/AKT/Snail signaling of lung cancer cells. Oncol. Lett. 2018, 15, 7369–7375. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.C.; Li, S.J.; Yuan, Z.P.; Li, D.J. MicroRNA-346 facilitates cell growth and metastasis, and suppresses cell apoptosis in human non-small cell lung cancer by regulation of XPC/ERK/Snail/E-cadherin pathway. Aging (Albany N. Y.) 2016, 8, 2509–2524. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Li, X.Y.; Wang, Z.M.; Han, Z.F.; Zhao, Y.H. MiR-22 inhibits lung cancer cell EMT and invasion through targeting Snail. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3598–3604. [Google Scholar] [PubMed]
- Fan, M.J.; Zhong, Y.H.; Shen, W.; Yuan, K.F.; Zhao, G.H.; Zhang, Y.; Wang, S.K. MiR-30 suppresses lung cancer cell 95D epithelial mesenchymal transition and invasion through targeted regulating Snail. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 2642–2649. [Google Scholar] [PubMed]
- Kumarswamy, R.; Mudduluru, G.; Ceppi, P.; Muppala, S.; Kozlowski, M.; Niklinski, J.; Papotti, M.; Allgayer, H. MicroRNA-30a inhibits epithelial-to-mesenchymal transition by targeting Snai1 and is downregulated in non-small cell lung cancer. Int. J. Cancer 2012, 130, 2044–2053. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.W.; Chen, P.C.; Chen, J.M.; Wu, Y.M.; Liu, P.Y.; Lu, C.H.; Lin, Y.F.; Tang, C.H.; Chao, C.C. Periostin promotes epithelial-mesenchymal transition via the MAPK/miR-381 axis in lung cancer. Oncotarget 2017, 8, 62248–62260. [Google Scholar] [CrossRef] [PubMed]
- Tominaga, E.; Yuasa, K.; Shimazaki, S.; Hijikata, T. MicroRNA-1 targets Slug and endows lung cancer A549 cells with epithelial and anti-tumorigenic properties. Exp. Cell Res. 2013, 319, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Qu, J.; Li, M.; An, J.; Zhao, B.; Zhong, W.; Gu, Q.; Cao, L.; Yang, H.; Hu, C. MicroRNA-33b inhibits lung adenocarcinoma cell growth, invasion, and epithelial-mesenchymal transition by suppressing Wnt/beta-catenin/ZEB1 signaling. Int. J. Oncol. 2015, 47, 2141–2152. [Google Scholar] [CrossRef] [PubMed]
- You, J.; Li, Y.; Fang, N.; Liu, B.; Zu, L.; Chang, R.; Li, X.; Zhou, Q. MiR-132 suppresses the migration and invasion of lung cancer cells via targeting the EMT regulator ZEB2. PLoS ONE 2014, 9, e91827. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Yang, J.; Li, J.; Shen, X.; Le, Y.; Zhou, C.; Wang, S.; Zhang, S.; Xu, D.; Gong, Z. MircoRNA-33a inhibits epithelial-to-mesenchymal transition and metastasis and could be a prognostic marker in non-small cell lung cancer. Sci. Rep. 2015, 5, 13677. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yang, S.; Yan, W.; Yang, J.; Qin, Y.J.; Lin, X.L.; Xie, R.Y.; Wang, S.C.; Jin, W.; Gao, F.; et al. MicroRNA-19 triggers epithelial-mesenchymal transition of lung cancer cells accompanied by growth inhibition. Lab. Investig. 2015, 95, 1056–1070. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Zhu, Y.; Ma, T.; Pan, C.; Wang, J.; He, Z.; Li, Z.; Qi, X.; Chen, Y. miR-204 functions as a tumor suppressor by regulating SIX1 in NSCLC. FEBS Lett. 2014, 588, 3703–3712. [Google Scholar] [CrossRef] [PubMed]
- Cui, Z.; Hu, Y. MicroRNA-124 suppresses Slug-mediated lung cancer metastasis. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3802–3811. [Google Scholar] [PubMed]
- Hu, F.Y.; Cao, X.N.; Xu, Q.Z.; Deng, Y.; Lai, S.Y.; Ma, J.; Hu, J.B. miR-124 modulates gefitinib resistance through SNAI2 and STAT3 in non-small cell lung cancer. J. Huazhong Univ. Sci. Technol. Med. Sci. 2016, 36, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Ahn, Y.H.; Gibbons, D.L.; Chakravarti, D.; Creighton, C.J.; Rizvi, Z.H.; Adams, H.P.; Pertsemlidis, A.; Gregory, P.A.; Wright, J.A.; Goodall, G.J.; et al. ZEB1 drives prometastatic actin cytoskeletal remodeling by downregulating miR-34a expression. J. Clin. Investig. 2012, 122, 3170–3183. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Guan, L.; Song, Y.; Xiang, G.M.; Chen, S.X.; Gao, B. MicroRNA-138 regulates chemoresistance in human non-small cell lung cancer via epithelial mesenchymal transition. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 1080–1086. [Google Scholar] [PubMed]
- Liu, X.; Tian, X.D.; Liu, Y.; Zhang, T.; Chen, L. Regulation of Twist in the metastasis of non-small cell lung cancer by miR-92b. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 4003–4010. [Google Scholar] [PubMed]
- Yang, Y.; Meng, H.; Peng, Q.; Yang, X.; Gan, R.; Zhao, L.; Chen, Z.; Lu, J.; Meng, Q.H. Downregulation of microRNA-21 expression restrains non-small cell lung cancer cell proliferation and migration through upregulation of programmed cell death 4. Cancer Gene Ther. 2015, 22, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Larzabal, L.; de Aberasturi, A.L.; Redrado, M.; Rueda, P.; Rodriguez, M.J.; Bodegas, M.E.; Montuenga, L.M.; Calvo, A. TMPRSS4 regulates levels of integrin alpha5 in NSCLC through miR-205 activity to promote metastasis. Br. J. Cancer 2014, 110, 764–774. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.H.; Tsai, M.F.; Gow, C.H.; Wu, S.G.; Liu, Y.N.; Chang, Y.L.; Yu, S.L.; Tsai, H.C.; Lin, S.W.; Chen, Y.W.; et al. Upregulation of microRNA-137 expression by Slug promotes tumor invasion and metastasis of non-small cell lung cancer cells through suppression of TFAP2C. Cancer Lett. 2017, 402, 190–202. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Chen, W.; Xia, Y.; Song, Y.; Zhao, Z.; Cheng, H.; Jiang, T. MiR-101 inhibits the proliferation and metastasis of lung cancer by targeting zinc finger E-box binding homeobox 1. Am. J. Transl. Res. 2018, 10, 1172–1183. [Google Scholar] [PubMed]
- Liu, Q.; Chen, J.; Wang, B.; Zheng, Y.; Wan, Y.; Wang, Y.; Zhou, L.; Liu, S.; Li, G.; Yan, Y. miR-145 modulates epithelial-mesenchymal transition and invasion by targeting ZEB2 in non-small cell lung cancer cell lines. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Huang, Z.; Chen, X.; Chen, S. miR-98 inhibits expression of TWIST to prevent progression of non-small cell lung cancers. Biomed. Pharmacother. 2017, 89, 1453–1461. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Wu, X.; Liu, B.; Wang, C.; Liu, Y.; Zhou, Q.; Xu, K. MiR-26a enhances metastasis potential of lung cancer cells via AKT pathway by targeting PTEN. Biochim. Biophys. Acta 2012, 1822, 1692–1704. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Zhu, J.; Shen, D.; Qin, H.; Lei, Z.; Li, W.; Huang, J.A.; Liu, Z. Repression of Smad4 by miR205 moderates TGF-beta-induced epithelial-mesenchymal transition in A549 cell lines. Int. J. Oncol. 2016, 49, 700–708. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.M.; Wang, L.; Shen, H.; Jiang, C.F.; Ge, X.; Li, D.M.; Wen, Y.Y.; Sun, H.R.; Pan, M.H.; Li, W.; et al. Downregulation of miR-218 contributes to epithelial-mesenchymal transition and tumor metastasis in lung cancer by targeting Slug/ZEB2 signaling. Oncogene 2017, 36, 2577–2588. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wang, X.; Li, W.; Wu, L.; Chang, L.; Chen, H. miRNA-124 modulates lung carcinoma cell migration and invasion. Int. J. Clin. Pharmacol. Ther. 2016, 54, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Yang, Z.; Zhang, P.; Liu, Y.; Shao, G. miR-154 inhibits migration and invasion of human non-small cell lung cancer by targeting ZEB2. Oncol. Lett. 2016, 12, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Yu, Y.; Zou, Q.; Wang, M.; Cui, Y.; Xie, J.; Wang, Z. MicroRNA-105 promotes epithelial-mesenchymal transition of nonsmall lung cancer cells through upregulating Mcl-1. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.Y.; Jiao, D.M.; Wang, J.; Hu, H.; Tang, X.; Chen, J.; Mou, H.; Lu, W. miR-206 regulates cisplatin resistance and EMT in human lung adenocarcinoma cells partly by targeting MET. Oncotarget 2016, 7, 24510–24526. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Jang, K.; Miller, P.; Picon-Ruiz, M.; Yeasky, T.M.; El-Ashry, D.; Slingerland, J.M. VEGFA links self-renewal and metastasis by inducing Sox2 to repress miR-452, driving Slug. Oncogene 2017, 36, 5199–5211. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.L.; Wen, Z.S.; Huang, Y.C.; Cheng, X.; Wang, G.Z.; Zhou, Y.C.; Wang, Z.Y.; Guo, Y.Q.; Cao, Y.; Zhou, G.B. Down-regulation of microRNA-144 in air pollution-related lung cancer. Sci. Rep. 2015, 5, 14331. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; An, H.; Fang, X. MicroRNA-144 regulates proliferation, invasion, and apoptosis of cells in malignant solitary pulmonary nodule via zinc finger E-box-binding homeobox 1. Int. J. Clin. Exp. Pathol. 2015, 8, 5960–5967. [Google Scholar] [PubMed]
- Lin, J.; Chen, Y.; Liu, L.; Shen, A.; Zheng, W. MicroRNA-155-5p suppresses the migration and invasion of lung adenocarcinoma A549 cells by targeting Smad2. Oncol. Lett. 2018, 16, 2444–2452. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, Z.; Li, Y.; Liu, S.; Gao, C.; Hou, X.; Yao, R.; Cui, L. The tumor suppressor miR-124 inhibits cell proliferation by targeting STAT3 and functions as a prognostic marker for postoperative NSCLC patients. Int. J. Oncol. 2015, 46, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Garofalo, M.; Di Leva, G.; Romano, G.; Nuovo, G.; Suh, S.S.; Ngankeu, A.; Taccioli, C.; Pichiorri, F.; Alder, H.; Secchiero, P.; et al. miR-221&222 regulate TRAIL resistance and enhance tumorigenicity through PTEN and TIMP3 downregulation. Cancer Cell 2009, 16, 498–509. [Google Scholar] [CrossRef] [PubMed]
- Jiao, A.; Sui, M.; Zhang, L.; Sun, P.; Geng, D.; Zhang, W.; Wang, X.; Li, J. MicroRNA-200c inhibits the metastasis of non-small cell lung cancer cells by targeting ZEB2, an epithelial-mesenchymal transition regulator. Mol. Med. Rep. 2016, 13, 3349–3355. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Liu, H.; Gu, Z.; Ji, C. Expression of microRNA-133 inhibits epithelial-mesenchymal transition in lung cancer cells by directly targeting FOXQ1. Arch. Bronconeumol. 2016, 52, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Ran, P.; Zhang, X.; Guo, X.; Yuan, Y.; Dong, T.; Zhu, B.; Zheng, S.; Xiao, C. Downregulation of N-Acetylglucosaminyltransferase GCNT3 by miR-302b-3p Decreases Non-Small Cell Lung Cancer (NSCLC) Cell Proliferation, Migration and Invasion. Cell. Physiol. Biochem. 2018, 50, 987–1004. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Banerjee, P.; Liu, X.; Yu, J.; Gibbons, D.L.; Wu, P.; Scott, K.L.; Diao, L.; Zheng, X.; Wang, J.; et al. The epithelial-to-mesenchymal transition activator ZEB1 initiates a prometastatic competing endogenous RNA network. J. Clin. Investig. 2018, 128, 1267–1282. [Google Scholar] [CrossRef] [PubMed]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.W.; Chang, Y.L.; Chang, Y.C.; Lin, J.C.; Chen, C.C.; Pan, S.H.; Wu, C.T.; Chen, H.Y.; Yang, S.C.; Hong, T.M.; et al. MicroRNA-135b promotes lung cancer metastasis by regulating multiple targets in the Hippo pathway and LZTS1. Nat. Commun. 2013, 4, 1877. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Wang, J.; Liu, S.; Wang, S.; Cheng, Y.; Zhou, W.; Duan, C.; Zhang, C. MicroRNA-361-3p suppresses tumor cell proliferation and metastasis by directly targeting SH2B1 in NSCLC. J. Exp. Clin. Cancer Res. 2016, 35, 76. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhou, F.; Yin, L.; Zhao, L.; Zhang, Y.; Wang, J. MicroRNA-199b targets the regulation of ZEB1 expression to inhibit cell proliferation, migration and invasion in nonsmall cell lung cancer. Mol. Med. Rep. 2017, 16, 5007–5014. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Fu, Z.; Gao, L.; Zhou, J.; Deng, X.; Luo, X.; Fang, W.; Luo, R. Direct interaction between miR-203 and ZEB2 suppresses epithelial-mesenchymal transition signaling and reduces lung adenocarcinoma chemoresistance. Acta Biochim. Biophys. Sin. (Shanghai) 2016, 48, 1042–1049. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Liu, L.; Cai, J.; Wu, J.; Guan, H.; Zhu, X.; Yuan, J.; Chen, S.; Li, M. Targeting Smad2 and Smad3 by miR-136 suppresses metastasis-associated traits of lung adenocarcinoma cells. Oncol. Res. 2013, 21, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, Y.; Sun, D.; Bu, J.; Ren, F.; Liu, B.; Zhang, S.; Xu, Z.; Pang, S.; Xu, S. miR-455-5p promotes cell growth and invasion by targeting SOCO3 in non-small cell lung cancer. Oncotarget 2017, 8, 114956–114965. [Google Scholar] [CrossRef] [PubMed]
- Burk, U.; Schubert, J.; Wellner, U.; Schmalhofer, O.; Vincan, E.; Spaderna, S.; Brabletz, T. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 2008, 9, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Nishijima, N.; Seike, M.; Soeno, C.; Chiba, M.; Miyanaga, A.; Noro, R.; Sugano, T.; Matsumoto, M.; Kubota, K.; Gemma, A. miR-200/ZEB axis regulates sensitivity to nintedanib in non-small cell lung cancer cells. Int. J. Oncol. 2016, 48, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Zhen, J.; Xu, X.; Zhen, K.; Zhu, B.; Pan, R.; Zhao, C. miR-215 functions as a tumor suppressor and directly targets ZEB2 in human non-small cell lung cancer. Oncol. Lett. 2015, 10, 1985–1992. [Google Scholar] [CrossRef] [PubMed]
- Mo, D.; Yang, D.; Xiao, X.; Sun, R.; Huang, L.; Xu, J. MiRNA-145 suppresses lung adenocarcinoma cell invasion and migration by targeting N-cadherin. Biotechnol. Lett. 2017, 39, 701–710. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Cai, L.; Li, R.; Zheng, J.; Wu, H.; Yang, X.; Li, H.; Wang, Z. Down-regulation of miR-489 contributes into NSCLC cell invasion through targeting SUZ12. Tumour Biol. 2015, 36, 6497–6505. [Google Scholar] [CrossRef] [PubMed]
- Yin, Q.; Han, Y.; Zhu, D.; Li, Z.; Shan, S.; Jin, W.; Lu, Q.; Ren, T. miR-145 and miR-497 suppress TGF-beta-induced epithelial-mesenchymal transition of non-small cell lung cancer by targeting MTDH. Cancer Cell Int. 2018, 18, 105. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.T.; Xu, M.; Xu, C.X.; Song, Z.G.; Jin, H. Decreased expression of miR216a contributes to non-small-cell lung cancer progression. Clin. Cancer Res. 2014, 20, 4705–4716. [Google Scholar] [CrossRef] [PubMed]
- Hong-Yuan, W.; Xiao-Ping, C. miR-338-3p suppresses epithelial-mesenchymal transition and metastasis in human nonsmall cell lung cancer. Indian J. Cancer 2015, 52 (Suppl. 3), E168–E171. [Google Scholar] [CrossRef]
- Li, J.; Song, Y.; Wang, Y.; Luo, J.; Yu, W. MicroRNA-148a suppresses epithelial-to-mesenchymal transition by targeting ROCK1 in non-small cell lung cancer cells. Mol. Cell. Biochem. 2013, 380, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Mo, X.; Zhang, F.; Liang, H.; Liu, M.; Li, H.; Xia, H. miR-544a promotes the invasion of lung cancer cells by targeting cadherina 1 in vitro. Onco-Targets Ther. 2014, 7, 895–900. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.J.; Ping, C.; Tang, J.; Zhang, W. MicroRNA-455 suppresses non-small cell lung cancer through targeting ZEB1. Cell Biol. Int. 2016, 40, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.; Su, P.; Yang, H.; Chi, F.; Shen, L.; Feng, X.; Jiang, H.; Zhang, X.; Wang, Z. MicroRNA-598 inhibits the proliferation and invasion of non-small cell lung cancer cells by directly targeting ZEB2. Exp. Ther. Med. 2018, 16, 5417–5423. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Liang, C. MicroRNA-148b inhibits proliferation and the epithelial-mesenchymal transition and increases radiosensitivity in non-small cell lung carcinomas by regulating ROCK1. Exp. Ther. Med. 2018, 15, 3609–3616. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, F.; Xu, P. miR-590 accelerates lung adenocarcinoma migration and invasion through directly suppressing functional target OLFM4. Biomed. Pharmacother. 2017, 86, 466–474. [Google Scholar] [CrossRef] [PubMed]
- Diepenbruck, M.; Tiede, S.; Saxena, M.; Ivanek, R.; Kalathur, R.K.R.; Luond, F.; Meyer-Schaller, N.; Christofori, G. miR-1199-5p and Zeb1 function in a double-negative feedback loop potentially coordinating EMT and tumour metastasis. Nat. Commun. 2017, 8, 1168. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Ouyang, R.; Wang, Z.; Zhou, W.; Chen, H.; Jiang, Y.; Zhang, Y.; Li, H.; Liao, M.; Wang, W.; et al. MiR-150 promotes cellular metastasis in non-small cell lung cancer by targeting FOXO4. Sci. Rep. 2016, 6, 39001. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.F.; Wang, S.; Xue, W.H.; Cheng, J.L. microRNA-590 suppresses the tumorigenesis and invasiveness of non-small cell lung cancer cells by targeting ADAM9. Mol. Cell. Biochem. 2016, 423, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.B.; Gu, Z.F.; Ma, M.; Wang, J.Y.; Wang, H.N. MicroRNA-590-5p suppresses the proliferation and invasion of non-small cell lung cancer by regulating GAB1. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 5954–5963. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.L.; Zheng, X.L.; Ye, K.; Ge, H.; Sun, Y.N.; Lu, Y.F.; Fan, Q.X. MicroRNA-183 Acts as a Tumor Suppressor in Human Non-Small Cell Lung Cancer by Down-Regulating MTA1. Cell. Physiol. Biochem. 2018, 46, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Wei, K.; Qin, Z.; Liu, W.; Shao, C.; Wang, C.; Ma, L.; Xie, M.; Shu, Y.; Shen, H. MiR-598 Suppresses Invasion and Migration by Negative Regulation of Derlin-1 and Epithelial-Mesenchymal Transition in Non-Small Cell Lung Cancer. Cell. Physiol. Biochem. 2018, 47, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Luo, F.; Sun, B.; Ye, H.; Li, J.; Shi, L.; Liu, Y.; Lu, X.; Wang, B.; Wang, Q.; et al. HIF-2alpha, acting via miR-191, is involved in angiogenesis and metastasis of arsenite-transformed HBE cells. Toxicol. Res. (Camb.) 2016, 5, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Ju, S.; Yuan, F.; Chen, G.; Shu, Y.; Li, C.; Xu, Y.; Luo, J.; Xia, L. microRNA-664 enhances proliferation, migration and invasion of lung cancer cells. Exp. Ther. Med. 2017, 13, 3555–3562. [Google Scholar] [CrossRef] [PubMed]
- Yongchun, Z.; Linwei, T.; Xicai, W.; Lianhua, Y.; Guangqiang, Z.; Ming, Y.; Guanjian, L.; Yujie, L.; Yunchao, H. MicroRNA-195 inhibits non-small cell lung cancer cell proliferation, migration and invasion by targeting MYB. Cancer Lett. 2014, 347, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Bao, L.; Lv, L.; Feng, J.; Chen, Y.; Wang, X.; Han, S.; Zhao, H. MiR-876-5p suppresses epithelial-mesenchymal transition of lung cancer by directly down-regulating bone morphogenetic protein 4. J. Biosci. 2017, 42, 671–681. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.H.; Lu, K.H.; Wang, K.M.; Sun, M.; Zhang, E.B.; Yang, J.S.; Yin, D.D.; Liu, Z.L.; Zhou, J.; Liu, Z.J.; et al. MicroRNA-196a promotes non-small cell lung cancer cell proliferation and invasion through targeting HOXA5. BMC Cancer 2012, 12, 348. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Xu, X.; Huang, H.; Ma, Z.; Zhang, S.; Niu, P.; Chen, Y.; Ping, J.; Lu, P.; Yu, C.; et al. MiR-1260b promotes the migration and invasion in non-small cell lung cancer via targeting PTPRK. Pathol. Res. Pract. 2018, 214, 776–783. [Google Scholar] [CrossRef] [PubMed]
- Krebs, A.M.; Mitschke, J.; Lasierra Losada, M.; Schmalhofer, O.; Boerries, M.; Busch, H.; Boettcher, M.; Mougiakakos, D.; Reichardt, W.; Bronsert, P.; et al. The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nat. Cell Biol. 2017, 19, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Sun, Y.; Ma, L. ZEB1: At the crossroads of epithelial-mesenchymal transition, metastasis and therapy resistance. Cell Cycle 2015, 14, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Vandewalle, C.; Comijn, J.; De Craene, B.; Vermassen, P.; Bruyneel, E.; Andersen, H.; Tulchinsky, E.; Van Roy, F.; Berx, G. SIP1/ZEB2 induces EMT by repressing genes of different epithelial cell-cell junctions. Nucleic Acids Res. 2005, 33, 6566–6578. [Google Scholar] [CrossRef] [PubMed]
- Larsen, J.E.; Nathan, V.; Osborne, J.K.; Farrow, R.K.; Deb, D.; Sullivan, J.P.; Dospoy, P.D.; Augustyn, A.; Hight, S.K.; Sato, M.; et al. ZEB1 drives epithelial-to-mesenchymal transition in lung cancer. J. Clin. Investig. 2016, 126, 3219–3235. [Google Scholar] [CrossRef] [PubMed]
- Humphries, B.; Yang, C. The microRNA-200 family: Small molecules with novel roles in cancer development, progression and therapy. Oncotarget 2015, 6, 6472–6498. [Google Scholar] [CrossRef] [PubMed]
- Kolesnikoff, N.; Attema, J.L.; Roslan, S.; Bert, A.G.; Schwarz, Q.P.; Gregory, P.A.; Goodall, G.J. Specificity protein 1 (Sp1) maintains basal epithelial expression of the miR-200 family: Implications for epithelial-mesenchymal transition. J. Biol. Chem. 2014, 289, 11194–11205. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Veronese, A.; Pichiorri, F.; Lee, T.J.; Jeon, Y.J.; Volinia, S.; Pineau, P.; Marchio, A.; Palatini, J.; Suh, S.S.; et al. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J. Exp. Med. 2011, 208, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.J.; Chao, C.H.; Xia, W.; Yang, J.Y.; Xiong, Y.; Li, C.W.; Yu, W.H.; Rehman, S.K.; Hsu, J.L.; Lee, H.H.; et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat. Cell Biol. 2011, 13, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Tan, Q.; Yan, M.; Liu, L.; Lin, H.; Zhao, F.; Bao, G.; Kong, H.; Ge, C.; Zhang, F.; et al. miRNA-200c inhibits invasion and metastasis of human non-small cell lung cancer by directly targeting ubiquitin specific peptidase 25. Mol. Cancer 2014, 13, 166. [Google Scholar] [CrossRef] [PubMed]
- Hurteau, G.J.; Carlson, J.A.; Spivack, S.D.; Brock, G.J. Overexpression of the microRNA hsa-miR-200c leads to reduced expression of transcription factor 8 and increased expression of E-cadherin. Cancer Res. 2007, 67, 7972–7976. [Google Scholar] [CrossRef] [PubMed]
- Mutlu, M.; Raza, U.; Saatci, O.; Eyupoglu, E.; Yurdusev, E.; Sahin, O. miR-200c: A versatile watchdog in cancer progression, EMT, and drug resistance. J. Mol. Med. (Berl.) 2016, 94, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Hurteau, G.J.; Spivack, S.D.; Brock, G.J. Potential mRNA degradation targets of hsa-miR-200c, identified using informatics and qRT-PCR. Cell Cycle 2006, 5, 1951–1956. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Gibbons, D.L.; Goswami, S.; Cortez, M.A.; Ahn, Y.H.; Byers, L.A.; Zhang, X.; Yi, X.; Dwyer, D.; Lin, W.; et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat. Commun. 2014, 5, 5241. [Google Scholar] [CrossRef] [PubMed]
- Park, K.S.; Moon, Y.W.; Raffeld, M.; Lee, D.H.; Wang, Y.; Giaccone, G. High cripto-1 and low miR-205 expression levels as prognostic markers in early stage non-small cell lung cancer. Lung Cancer 2018, 116, 38–45. [Google Scholar] [CrossRef] [PubMed]
- Park, K.S.; Raffeld, M.; Moon, Y.W.; Xi, L.; Bianco, C.; Pham, T.; Lee, L.C.; Mitsudomi, T.; Yatabe, Y.; Okamoto, I.; et al. CRIPTO1 expression in EGFR-mutant NSCLC elicits intrinsic EGFR-inhibitor resistance. J. Clin. Investig. 2014, 124, 3003–3015. [Google Scholar] [CrossRef] [PubMed]
- Wellner, U.; Schubert, J.; Burk, U.C.; Schmalhofer, O.; Zhu, F.; Sonntag, A.; Waldvogel, B.; Vannier, C.; Darling, D.; zur Hausen, A.; et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat. Cell. Biol. 2009, 11, 1487–1495. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kaminska, B.; Wesolowska, A.; Danilkiewicz, M. TGF beta signalling and its role in tumour pathogenesis. Acta Biochim. Pol. 2005, 52, 329–337. [Google Scholar] [PubMed]
- Pain, M.; Bermudez, O.; Lacoste, P.; Royer, P.J.; Botturi, K.; Tissot, A.; Brouard, S.; Eickelberg, O.; Magnan, A. Tissue remodelling in chronic bronchial diseases: From the epithelial to mesenchymal phenotype. Eur. Respir. Rev. 2014, 23, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Zaravinos, A. The Regulatory Role of MicroRNAs in EMT and Cancer. J. Oncol. 2015, 2015, 865816. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.J.; Wang, H.Y.; Zhang, H.T.; Su, J.M.; Zhu, J.; Wang, H.B.; Zhou, W.Y.; Zhang, H.; Zhao, M.C.; Zhang, L.; et al. Transforming growth factor-beta1 promotes lung adenocarcinoma invasion and metastasis by epithelial-to-mesenchymal transition. Mol. Cell. Biochem. 2011, 355, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, Y.; Takanashi, S.; Kanehira, Y.; Tsushima, T.; Imai, T.; Okumura, K. Transforming growth factor-beta1 level correlates with angiogenesis, tumor progression, and prognosis in patients with nonsmall cell lung carcinoma. Cancer 2001, 91, 964–971. [Google Scholar] [CrossRef]
- Jakubowska, K.; Naumnik, W.; Niklinska, W.; Chyczewska, E. Clinical Significance of HMGB-1 and TGF-beta Level in Serum and BALF of Advanced Non-Small Cell Lung Cancer. Adv. Exp. Med. Biol. 2015, 852, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Soucheray, M.; Capelletti, M.; Pulido, I.; Kuang, Y.; Paweletz, C.P.; Becker, J.H.; Kikuchi, E.; Xu, C.; Patel, T.B.; Al-Shahrour, F.; et al. Intratumoral Heterogeneity in EGFR-Mutant NSCLC Results in Divergent Resistance Mechanisms in Response to EGFR Tyrosine Kinase Inhibition. Cancer Res. 2015, 75, 4372–4383. [Google Scholar] [CrossRef] [PubMed]
- Sequist, L.V.; Waltman, B.A.; Dias-Santagata, D.; Digumarthy, S.; Turke, A.B.; Fidias, P.; Bergethon, K.; Shaw, A.T.; Gettinger, S.; Cosper, A.K.; et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci. Transl. Med. 2011, 3, 75ra26. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Yang, H.; He, L.; Zhao, J.J.; Coppola, D.; Dalton, W.S.; Cheng, J.Q. MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol. Cell. Biol. 2008, 28, 6773–6784. [Google Scholar] [CrossRef] [PubMed]
- Santibanez, J.F.; Obradovic, H.; Kukolj, T.; Krstic, J. Transforming growth factor-beta, matrix metalloproteinases, and urokinase-type plasminogen activator interaction in the cancer epithelial to mesenchymal transition. Dev. Dyn. 2018, 247, 382–395. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Shen, H.; Zhou, Y.; Yang, Z.; Hu, T. MicroRNA-215 suppresses the proliferation, migration and invasion of non-small cell lung carcinoma cells through the downregulation of matrix metalloproteinase-16 expression. Exp. Ther. Med. 2018, 15, 3239–3246. [Google Scholar] [CrossRef] [PubMed]
- Li, T.T.; Gao, X.; Gao, L.; Gan, B.L.; Xie, Z.C.; Zeng, J.J.; Chen, G. Role of upregulated miR-136-5p in lung adenocarcinoma: A study of 1242 samples utilizing bioinformatics analysis. Pathol. Res. Pract. 2018, 214, 750–766. [Google Scholar] [CrossRef] [PubMed]
- Jin, G.; Kim, M.J.; Jeon, H.S.; Choi, J.E.; Kim, D.S.; Lee, E.B.; Cha, S.I.; Yoon, G.S.; Kim, C.H.; Jung, T.H.; et al. PTEN mutations and relationship to EGFR, ERBB2, KRAS, and TP53 mutations in non-small cell lung cancers. Lung Cancer 2010, 69, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Kohno, T.; Takahashi, M.; Manda, R.; Yokota, J. Inactivation of the PTEN/MMAC1/TEP1 gene in human lung cancers. Genes Chromosomes Cancer 1998, 22, 152–156. [Google Scholar] [CrossRef]
- Yu, Y.X.; Wang, Y.; Liu, H. Overexpression of PTEN suppresses non-small-cell lung carcinoma metastasis through inhibition of integrin alphaVbeta6 signaling. Am. J. Transl. Res. 2017, 9, 3304–3314. [Google Scholar] [PubMed]
- Xiao, J.; Hu, C.P.; He, B.X.; Chen, X.; Lu, X.X.; Xie, M.X.; Li, W.; He, S.Y.; You, S.J.; Chen, Q. PTEN expression is a prognostic marker for patients with non-small cell lung cancer: A systematic review and meta-analysis of the literature. Oncotarget 2016, 7, 57832–57840. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Shi, X.; Wang, L.; Wu, Y.; Jin, F.; Bai, C.; Song, Y. SUZ12 is involved in progression of non-small cell lung cancer by promoting cell proliferation and metastasis. Tumour Biol. 2014, 35, 6073–6082. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Zhang, S.; Chen, J.; Jiang, K.; Zhang, Q.; Guo, K.; Liu, Y. The transcriptional profiling of glycogenes associated with hepatocellular carcinoma metastasis. PLoS ONE 2014, 9, e107941. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.C.; Chen, H.Y.; Huang, H.C.; Huang, J.; Liang, J.T.; Shen, T.L.; Lin, N.Y.; Ho, C.C.; Cho, I.M.; Hsu, S.M. C2GnT-M is downregulated in colorectal cancer and its re-expression causes growth inhibition of colon cancer cells. Oncogene 2006, 25, 3267–3276. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.L.; Chen, H.Y.; Chang, G.C.; Chen, C.Y.; Chen, H.W.; Singh, S.; Cheng, C.L.; Yu, C.J.; Lee, Y.C.; Chen, H.S.; et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell 2008, 13, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Aiso, T.; Ohtsuka, K.; Ueda, M.; Karita, S.; Yokoyama, T.; Takata, S.; Matsuki, N.; Kondo, H.; Takizawa, H.; Okada, A.A.; et al. Serum levels of candidate microRNA diagnostic markers differ among the stages of non-small-cell lung cancer. Oncol. Lett. 2018, 16, 6643–6651. [Google Scholar] [CrossRef] [PubMed]
- Pastorkova, Z.; Skarda, J.; Andel, J. The role of microRNA in metastatic processes of non-small cell lung carcinoma. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech. Repub. 2016, 160, 343–357. [Google Scholar] [CrossRef] [PubMed]
- Roodman, G.D. Mechanisms of bone metastasis. N. Engl. J. Med. 2004, 350, 1655–1664. [Google Scholar] [CrossRef] [PubMed]
- Mundy, G.R. Metastasis to bone: Causes, consequences and therapeutic opportunities. Nat. Rev. Cancer 2002, 2, 584–593. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.; Yang, Z.; Li, G.; Shen, L.; Xiang, X.; Liu, X.; Xu, D.; Xu, L.; Chen, Y.; Tian, Z.; et al. Genome-wide identification of bone metastasis-related microRNAs in lung adenocarcinoma by high-throughput sequencing. PLoS ONE 2013, 8, e61212. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Zhang, H.; Zhang, L.; He, Y.; Weng, S.; Dong, Z.; Wang, J.; Zhang, P.; Nao, R. MicroRNA-21 regulates non-small cell lung cancer cell proliferation by affecting cell apoptosis via COX-19. Int. J. Clin. Exp. Med. 2015, 8, 8835–8841. [Google Scholar] [PubMed]
- Srinivasan, S.; Avadhani, N.G. Cytochrome c oxidase dysfunction in oxidative stress. Free Radic. Biol. Med. 2012, 53, 1252–1263. [Google Scholar] [CrossRef] [PubMed]
- Zou, H.; Li, Y.; Liu, X.; Wang, X. An APAF-1.cytochrome c multimeric complex is a functional apoptosome that activates procaspase-9. J. Biol. Chem. 1999, 274, 11549–11556. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Liu, X.; Wang, H.; Li, J.; Dai, L.; Li, J.; Dong, C. Lung adenocarcinoma cell-derived exosomal miR-21 facilitates osteoclastogenesis. Gene 2018, 666, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Ye, Y.; Jiang, Q.; Chen, Y.; Lyu, X.; Li, J.; Wang, S.; Liu, T.; Cai, H.; Yao, K.; et al. Epstein-Barr virus-encoded microRNA BART1 induces tumour metastasis by regulating PTEN-dependent pathways in nasopharyngeal carcinoma. Nat. Commun. 2015, 6, 7353. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Bertke, A.S.; Patel, A.; Wang, K.; Cohen, J.I.; Krause, P.R. An acutely and latently expressed herpes simplex virus 2 viral microRNA inhibits expression of ICP34.5, a viral neurovirulence factor. Proc. Natl. Acad. Sci. USA 2008, 105, 10931–10936. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Liu, S.; Zhou, Z.; Yan, H.; Xiao, J. A herpes simplex virus type 2-encoded microRNA promotes tumor cell metastasis by targeting suppressor of cytokine signaling 2 in lung cancer. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Sen, B.; Peng, S.; Woods, D.M.; Wistuba, I.; Bell, D.; El-Naggar, A.K.; Lai, S.Y.; Johnson, F.M. STAT5A-mediated SOCS2 expression regulates Jak2 and STAT3 activity following c-Src inhibition in head and neck squamous carcinoma. Clin. Cancer Res. 2012, 18, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Wikman, H.; Kettunen, E.; Seppanen, J.K.; Karjalainen, A.; Hollmen, J.; Anttila, S.; Knuutila, S. Identification of differentially expressed genes in pulmonary adenocarcinoma by using cDNA array. Oncogene 2002, 21, 5804–5813. [Google Scholar] [CrossRef] [PubMed]
- Kuo, P.L.; Liao, S.H.; Hung, J.Y.; Huang, M.S.; Hsu, Y.L. MicroRNA-33a functions as a bone metastasis suppressor in lung cancer by targeting parathyroid hormone related protein. Biochim. Biophys. Acta 2013, 1830, 3756–3766. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Yang, F.; Liu, R.; Li, X.; Fan, H.; Liu, J.; Wei, S.; Chen, G.; Chen, J.; Da, Y. Serum microRNA-139-5p is downregulated in lung cancer patients with lytic bone metastasis. Oncol. Rep. 2018, 39, 2376–2384. [Google Scholar] [CrossRef] [PubMed]
- Valencia, K.; Luis-Ravelo, D.; Bovy, N.; Anton, I.; Martinez-Canarias, S.; Zandueta, C.; Ormazabal, C.; Struman, I.; Tabruyn, S.; Rebmann, V.; et al. miRNA cargo within exosome-like vesicle transfer influences metastatic bone colonization. Mol. Oncol. 2014, 8, 689–703. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Li, D.; Wang, D.; Qiu, T.; Liu, K. Evaluation of microRNA-203 in bone metastasis of patients with non-small cell lung cancer through TGF-beta/SMAD2 expression. Oncol. Rep. 2017. [Google Scholar] [CrossRef] [PubMed]
- Grinberg-Rashi, H.; Ofek, E.; Perelman, M.; Skarda, J.; Yaron, P.; Hajduch, M.; Jacob-Hirsch, J.; Amariglio, N.; Krupsky, M.; Simansky, D.A.; et al. The expression of three genes in primary non-small cell lung cancer is associated with metastatic spread to the brain. Clin. Cancer Res. 2009, 15, 1755–1761. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Xu, Y.; Zhang, Y.; Tan, W.; Xue, J.; Yang, Z.; Zhang, Y.; Lu, Y.; Hu, X. Downregulation of miR-145 contributes to lung adenocarcinoma cell growth to form brain metastases. Oncol. Rep. 2013, 30, 2027–2034. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Govindan, R.; Wang, L.; Liu, P.Y.; Goodgame, B.; Wen, W.; Sezhiyan, A.; Pfeifer, J.; Li, Y.F.; Hua, X.; et al. MicroRNA profiling and prediction of recurrence/relapse-free survival in stage I lung cancer. Carcinogenesis 2012, 33, 1046–1054. [Google Scholar] [CrossRef] [PubMed]
- Donzelli, S.; Mori, F.; Bellissimo, T.; Sacconi, A.; Casini, B.; Frixa, T.; Roscilli, G.; Aurisicchio, L.; Facciolo, F.; Pompili, A.; et al. Epigenetic silencing of miR-145-5p contributes to brain metastasis. Oncotarget 2015, 6, 35183–35201. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Garg, N.; Venugopal, C.; Hallett, R.; Tokar, T.; McFarlane, N.; Mahendram, S.; Bakhshinyan, D.; Manoranjan, B.; Vora, P.; et al. STAT3 pathway regulates lung-derived brain metastasis initiating cell capacity through miR-21 activation. Oncotarget 2015, 6, 27461–27477. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.J.; Lee, H.W.; Kim, H.R.; Song, H.J.; Lee, D.H.; Lee, H.; Shin, C.H.; Joung, J.G.; Kim, D.H.; Joo, K.M.; et al. Overexpression of microRNA-95-3p suppresses brain metastasis of lung adenocarcinoma through downregulation of cyclin D1. Oncotarget 2015, 6, 20434–20448. [Google Scholar] [CrossRef] [PubMed]
- Berrout, J.; Kyriakopoulou, E.; Moparthi, L.; Hogea, A.S.; Berrout, L.; Ivan, C.; Lorger, M.; Boyle, J.; Peers, C.; Muench, S.; et al. TRPA1-FGFR2 binding event is a regulatory oncogenic driver modulated by miRNA-142-3p. Nat. Commun. 2017, 8, 947. [Google Scholar] [CrossRef] [PubMed]
- Remon, J.; Alvarez-Berdugo, D.; Majem, M.; Moran, T.; Reguart, N.; Lianes, P. miRNA-197 and miRNA-184 are associated with brain metastasis in EGFR-mutant lung cancers. Clin. Transl. Oncol. 2016, 18, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Yu, J.; Wang, L. Machine Learning Based Prediction of Brain Metastasis of Patients with IIIA-N2 Lung Adenocarcinoma by a Three-miRNA Signature. Transl. Oncol. 2018, 11, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Arora, S.; Ranade, A.R.; Tran, N.L.; Nasser, S.; Sridhar, S.; Korn, R.L.; Ross, J.T.; Dhruv, H.; Foss, K.M.; Sibenaller, Z.; et al. MicroRNA-328 is associated with (non-small) cell lung cancer (NSCLC) brain metastasis and mediates NSCLC migration. Int. J. Cancer 2011, 129, 2621–2631. [Google Scholar] [CrossRef] [PubMed]
- Wei, C.H.; Wu, G.; Cai, Q.; Gao, X.C.; Tong, F.; Zhou, R.; Zhang, R.G.; Dong, J.H.; Hu, Y.; Dong, X.R. MicroRNA-330-3p promotes cell invasion and metastasis in non-small cell lung cancer through GRIA3 by activating MAPK/ERK signaling pathway. J. Hematol. Oncol. 2017, 10, 125. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.P.; Zhu, Z.T.; Zhang, Y.; He, C.Y. Downregulation of MicroRNA-330 Correlates with the Radiation Sensitivity and Prognosis of Patients with Brain Metastasis from Lung Cancer. Cell. Physiol. Biochem. 2017, 42, 2220–2229. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.J.; Li, X.Y.; Zhao, Y.Q.; Liu, W.J.; Wu, H.J.; Liu, J.; Mu, X.Q.; Wu, H.B. Down-regulated microRNA-375 expression as a predictive biomarker in non-small cell lung cancer brain metastasis and its prognostic significance. Pathol. Res. Pract. 2017, 213, 882–888. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.T.; Xu, S.D.; Xu, H.; Zhang, J.F.; Ning, J.F.; Wang, S.F. MicroRNA-378 is associated with non-small cell lung cancer brain metastasis by promoting cell migration, invasion and tumor angiogenesis. Med. Oncol. 2012, 29, 1673–1680. [Google Scholar] [CrossRef] [PubMed]
- Sun, G.; Ding, X.; Bi, N.; Wu, L.; Wang, J.; Zhang, W.; Dong, X.; Lv, N.; Song, Y.; Zhan, Q.; et al. MiR-423-5p in brain metastasis: Potential role in diagnostics and molecular biology. Cell Death Dis. 2018, 9, 936. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Feng, Q.; Wei, X.; Yu, Y. MicroRNA-490 regulates lung cancer metastasis by targeting poly r(C)-binding protein 1. Tumour Biol. 2016, 37, 15221–15228. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Chen, S.Y.; Yang, Z.; Zhang, J.; Wang, W.; Liu, M.Y.; Niu, Y.; Wei, X.M.; Li, H.M.; Hu, W.N.; et al. miR-4317 suppresses non-small cell lung cancer (NSCLC) by targeting fibroblast growth factor 9 (FGF9) and cyclin D2 (CCND2). J. Exp. Clin. Cancer Res. 2018, 37, 230. [Google Scholar] [CrossRef] [PubMed]
- Loffler, D.; Brocke-Heidrich, K.; Pfeifer, G.; Stocsits, C.; Hackermuller, J.; Kretzschmar, A.K.; Burger, R.; Gramatzki, M.; Blumert, C.; Bauer, K.; et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 2007, 110, 1330–1333. [Google Scholar] [CrossRef] [PubMed]
- Iliopoulos, D.; Jaeger, S.A.; Hirsch, H.A.; Bulyk, M.L.; Struhl, K. STAT3 activation of miR-21 and miR-181b-1 via PTEN and CYLD are part of the epigenetic switch linking inflammation to cancer. Mol. Cell 2010, 39, 493–506. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, J.C.; Scrideli, C.A.; Brassesco, M.S.; Morales, A.G.; Pezuk, J.A.; Queiroz Rde, P.; Yunes, J.A.; Brandalise, S.R.; Tone, L.G. Differential miRNA expression in childhood acute lymphoblastic leukemia and association with clinical and biological features. Leuk. Res. 2012, 36, 293–298. [Google Scholar] [CrossRef] [PubMed]
- Subra, C.; Grand, D.; Laulagnier, K.; Stella, A.; Lambeau, G.; Paillasse, M.; De Medina, P.; Monsarrat, B.; Perret, B.; Silvente-Poirot, S.; et al. Exosomes account for vesicle-mediated transcellular transport of activatable phospholipases and prostaglandins. J. Lipid Res. 2010, 51, 2105–2120. [Google Scholar] [CrossRef] [PubMed]
- Bang, C.; Thum, T. Exosomes: New players in cell-cell communication. Int. J. Biochem. Cell Biol. 2012, 44, 2060–2064. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, J.; Jack, H.M. Serum microRNAs as powerful cancer biomarkers. Biochim. Biophys. Acta 2010, 1806, 200–207. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Aleckovic, M.; Lavotshkin, S.; Matei, I.; Costa-Silva, B.; Moreno-Bueno, G.; Hergueta-Redondo, M.; Williams, C.; Garcia-Santos, G.; Ghajar, C.; et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 2012, 18, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Rana, S.; Malinowska, K.; Zoller, M. Exosomal tumor microRNA modulates premetastatic organ cells. Neoplasia 2013, 15, 281–295. [Google Scholar] [CrossRef] [PubMed]
- Carr, I. Lymphatic metastasis. Cancer Metastasis Rev. 1983, 2, 307–317. [Google Scholar] [CrossRef] [PubMed]
- Eisenhauer, E.A.; Therasse, P.; Bogaerts, J.; Schwartz, L.H.; Sargent, D.; Ford, R.; Dancey, J.; Arbuck, S.; Gwyther, S.; Mooney, M.; et al. New response evaluation criteria in solid tumours: Revised RECIST guideline (version 1.1). Eur. J. Cancer 2009, 45, 228–247. [Google Scholar] [CrossRef] [PubMed]
- Asamura, H.; Suzuki, K.; Kondo, H.; Tsuchiya, R. Where is the boundary between N1 and N2 stations in lung cancer? Ann. Thorac. Surg. 2000, 70, 1839–1845; discussion 1845–1836. [Google Scholar] [CrossRef]
- Wu, J.; Ohta, Y.; Minato, H.; Tsunezuka, Y.; Oda, M.; Watanabe, Y.; Watanabe, G. Nodal occult metastasis in patients with peripheral lung adenocarcinoma of 2.0 cm or less in diameter. Ann. Thorac. Surg. 2001, 71, 1772–1777; discussion 1777–1778. [Google Scholar] [CrossRef]
- Ong, P.; Grosu, H.; Eapen, G.A.; Rodriguez, M.; Lazarus, D.; Ost, D.; Jimenez, C.A.; Morice, R.; Bandi, V.; Tamara, L.; et al. Endobronchial ultrasound-guided transbronchial needle aspiration for systematic nodal staging of lung cancer in patients with N0 disease by computed tomography and integrated positron emission tomography-computed tomography. Ann. Am. Thorac. Soc. 2015, 12, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Yoon, J.K.; Park, K.J.; Lee, S.J. Prediction of occult lymph node metastasis using volume-based PET parameters in small-sized peripheral non-small cell lung cancer. Cancer Imaging 2015, 15, 21. [Google Scholar] [CrossRef] [PubMed]
- Pieterman, R.M.; van Putten, J.W.; Meuzelaar, J.J.; Mooyaart, E.L.; Vaalburg, W.; Koeter, G.H.; Fidler, V.; Pruim, J.; Groen, H.J. Preoperative staging of non-small-cell lung cancer with positron-emission tomography. N. Engl. J. Med. 2000, 343, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Capodanno, A.; Boldrini, L.; Proietti, A.; Ali, G.; Pelliccioni, S.; Niccoli, C.; D’Incecco, A.; Cappuzzo, F.; Chella, A.; Lucchi, M.; et al. Let-7g and miR-21 expression in non-small cell lung cancer: Correlation with clinicopathological and molecular features. Int. J. Oncol. 2013, 43, 765–774. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zhang, B.; Shao, Y.; Chen, L.; Wang, X.; Zhang, Z.; Shu, Y.; Guo, R. Correlation between the expression levels of miR-1 and PIK3CA in non-small-cell lung cancer and their relationship with clinical characteristics and prognosis. Future Oncol. 2014, 10, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, D.; Wei, Y. Overexpressed CDR1as functions as an oncogene to promote the tumor progression via miR-7 in non-small-cell lung cancer. Onco-Targets Ther. 2018, 11, 3979–3987. [Google Scholar] [CrossRef] [PubMed]
- Cheng, M.W.; Shen, Z.T.; Hu, G.Y.; Luo, L.G. Prognostic Significance of microRNA-7 and its Roles in the Regulation of Cisplatin Resistance in Lung Adenocarcinoma. Cell. Physiol. Biochem. 2017, 42, 660–672. [Google Scholar] [CrossRef] [PubMed]
- Muraoka, T.; Soh, J.; Toyooka, S.; Maki, Y.; Shien, K.; Furukawa, M.; Ueno, T.; Tanaka, N.; Yamamoto, H.; Asano, H.; et al. Impact of aberrant methylation of microRNA-9 family members on non-small cell lung cancers. Mol. Clin. Oncol. 2013, 1, 185–189. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Liu, X.; Han, L.; Shen, H.; Liu, L.; Shu, Y. Up-regulation of miR-9 expression as a poor prognostic biomarker in patients with non-small cell lung cancer. Clin. Transl. Oncol. 2014, 16, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Liu, L.; Li, J.; Yan, M.; Lin, H.; Liu, Y.; Chu, D.; Tu, H.; Gu, A.; Yao, M. MiRNA-10a is upregulated in NSCLC and may promote cancer by targeting PTEN. Oncotarget 2015, 6, 30239–30250. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xu, L.; Yang, Z.; Lu, H.; Hu, D.; Li, W.; Zhang, Z.; Liu, B.; Ma, S. MicroRNA-10b indicates a poor prognosis of non-small cell lung cancer and targets E-cadherin. Clin. Transl. Oncol. 2015, 17, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.L.; Xu, L.P.; Zhuo, F.L.; Wang, T.Y. Prognostic value of microRNA-10b overexpression in peripheral blood mononuclear cells of nonsmall-cell lung cancer patients. Tumour Biol. 2015, 36, 7069–7075. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.L.; Wang, W.; Xu, L.P. Predictive value of microRNA-10b expression in peripheral blood mononuclear cells in evaluating short- and long-term efficacy of chemotherapy for patients with advanced non-small-cell lung cancer. Neoplasma 2018, 65, 610–619. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, Y.; Liu, J.; Fan, Y.; Li, X.; Dong, M.; Liu, H.; Chen, J. Expression levels of microRNA-145 and microRNA-10b are associated with metastasis in non-small cell lung cancer. Cancer Biol. Ther. 2016, 17, 272–279. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; An, X.; Zhao, H.; Zhang, Q.; Zhao, H. Long non-coding RNA HNF1A-AS1 promotes cell proliferation and invasion via regulating miR-17-5p in non-small cell lung cancer. Biomed. Pharmacother. 2018, 98, 594–599. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Wu, X.; Wang, Z.; Li, B.; Zhu, X. Effect of miR-18a overexpression on the radiosensitivity of non-small cell lung cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 643–648. [Google Scholar] [PubMed]
- Lin, Q.; Chen, T.; Lin, Q.; Lin, G.; Lin, J.; Chen, G.; Guo, L. Serum miR-19a expression correlates with worse prognosis of patients with non-small cell lung cancer. J. Surg. Oncol. 2013, 107, 767–771. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Cao, Y.; He, Z.; He, J.; Hu, C.; Duan, H.; Jiang, J. Serum levels of miR-19b and miR-146a as prognostic biomarkers for non-small cell lung cancer. Tohoku J. Exp. Med. 2014, 232, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.G.; Zhu, W.Y.; Huang, Y.Y.; Ma, L.N.; Zhou, S.Q.; Wang, Y.K.; Zeng, F.; Zhou, J.H.; Zhang, Y.K. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med. Oncol. 2012, 29, 618–626. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.X.; Bian, H.B.; Wang, J.R.; Cheng, Z.X.; Wang, K.M.; De, W. Prognostic significance of serum miRNA-21 expression in human non-small cell lung cancer. J. Surg. Oncol. 2011, 104, 847–851. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.L.; Wang, H.; Liu, J.; Wang, Z.X. MicroRNA-21 (miR-21) expression promotes growth, metastasis, and chemo- or radioresistance in non-small cell lung cancer cells by targeting PTEN. Mol. Cell. Biochem. 2013, 372, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.C.; Wang, W.; Zhang, Z.B.; Zhao, J.; Tan, X.G.; Luo, J.C. Overexpression of miRNA-21 promotes radiation-resistance of non-small cell lung cancer. Radiat. Oncol. 2013, 8, 146. [Google Scholar] [CrossRef] [PubMed]
- Tian, L.; Shan, W.; Zhang, Y.; Lv, X.; Li, X.; Wei, C. Up-Regulation of miR-21 Expression Predicate Advanced Clinicopathological Features and Poor Prognosis in Patients with Non-Small Cell Lung Cancer. Pathol. Oncol. Res. 2016, 22, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.X.; Su, Y.L.; Zhang, H.; Kong, J.Y.; Yu, H.; Qian, B.Y. Prognostic implications for high expression of MiR-25 in lung adenocarcinomas of female non-smokers. Asian Pac. J. Cancer Prev. 2014, 15, 1197–1203. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Tu, Y.S.; Long, J.; Zhang, H.Q.; Qi, C.L.; Xie, X.B.; Li, S.H.; Zhang, Y.J. SRF-miR29b-MMP2 axis inhibits NSCLC invasion and metastasis. Int. J. Oncol. 2015, 47, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.Y.; Zhang, X.C.; Su, J.; Meng, W.; Yang, X.N.; Yang, J.J.; Zhou, Q.; Chen, Z.Y.; Chen, Z.H.; Xie, Z.; et al. BCL11A overexpression predicts survival and relapse in non-small cell lung cancer and is modulated by microRNA-30a and gene amplification. Mol. Cancer 2013, 12, 61. [Google Scholar] [CrossRef] [PubMed]
- Qi, Z.; Zhang, B.; Zhang, J.; Hu, Q.; Xu, F.; Chen, B.; Zhu, C. MicroRNA-30b inhibits non-small cell lung cancer cell growth by targeting the epidermal growth factor receptor. Neoplasma 2018, 65, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Li, P.; Yang, R.; Cheng, R.; Zhang, F.; Wang, Y.; Chen, X.; Sun, Q.; Zang, W.; Du, Y.; et al. microRNA-30b inhibits cell invasion and migration through targeting collagen triple helix repeat containing 1 in non-small cell lung cancer. Cancer Cell Int. 2015, 15, 85. [Google Scholar] [CrossRef] [PubMed]
- Meng, W.; Ye, Z.; Cui, R.; Perry, J.; Dedousi-Huebner, V.; Huebner, A.; Wang, Y.; Li, B.; Volinia, S.; Nakanishi, H.; et al. MicroRNA-31 predicts the presence of lymph node metastases and survival in patients with lung adenocarcinoma. Clin. Cancer Res. 2013, 19, 5423–5433. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Wang, Y.L.; Yao, W.J.; Guo, L.; Xi, H.F.; Li, S.Y.; Zhao, B.S. Expression of miR-32 in human non-small cell lung cancer and its correlation with tumor progression and patient survival. Int. J. Clin. Exp. Pathol. 2015, 8, 824–829. [Google Scholar] [PubMed]
- Zhao, K.; Cheng, J.; Chen, B.; Liu, Q.; Xu, D.; Zhang, Y. Circulating microRNA-34 family low expression correlates with poor prognosis in patients with non-small cell lung cancer. J. Thorac. Dis. 2017, 9, 3735–3746. [Google Scholar] [CrossRef] [PubMed]
- Ren, P.; Gong, F.; Zhang, Y.; Jiang, J.; Zhang, H. MicroRNA-92a promotes growth, metastasis, and chemoresistance in non-small cell lung cancer cells by targeting PTEN. Tumour Biol. 2016, 37, 3215–3225. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, P.; Chen, T.; Gao, G.; Chen, X.; Du, Y.; Zhang, R.; Yang, R.; Zhao, W.; Dun, S.; et al. Expression of microRNA-96 and its potential functions by targeting FOXO3 in non-small cell lung cancer. Tumour Biol. 2015, 36, 685–692. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Dong, L.; Fang, Q.; Xia, H.; Hou, X. Low serum miR-98 as an unfavorable prognostic biomarker in patients with non-small cell lung cancer. Cancer Biomark. 2017, 20, 283–288. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Fang, S.; Gao, L.; Tan, Y.; Yang, Z. Clinic significance of microRNA-99a expression in human lung adenocarcinoma. J. Surg. Oncol. 2013, 108, 248–255. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lu, K.H.; Liu, Z.L.; Sun, M.; De, W.; Wang, Z.X. MicroRNA-100 is a potential molecular marker of non-small cell lung cancer and functions as a tumor suppressor by targeting polo-like kinase 1. BMC Cancer 2012, 12, 519. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Zhang, T.; Liu, H.; Lv, T.; Yuan, D.; Yao, Y.; Lv, Y.; Song, Y. MiR-101 and Mcl-1 in non-small-cell lung cancer: Expression profile and clinical significance. Med. Oncol. 2012, 29, 1681–1686. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.M.; Yi, W.W.; Ma, Y.S.; Wu, W.; Yu, F.; Fan, H.W.; Lv, Z.W.; Yang, H.Q.; Chang, Z.Y.; Zhang, C.; et al. Prognostic implications of decreased microRNA-101-3p expression in patients with non-small cell lung cancer. Oncol. Lett. 2018, 16, 7048–7056. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Zhang, F.; Zhu, C.; Geng, L.; Tian, T.; Liu, H. Upregulated lncRNA SNHG1 contributes to progression of non-small cell lung cancer through inhibition of miR-101-3p and activation of Wnt/beta-catenin signaling pathway. Oncotarget 2017, 8, 17785–17794. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Tian, J.; Guo, Z.J.; Zhang, Z.B.; Xiao, C.Y.; Wang, X.C. Expression of microRNAs-106b in nonsmall cell lung cancer. J. Cancer Res. Ther. 2018, 14, S295–S298. [Google Scholar] [CrossRef] [PubMed]
- Zhong, K.Z.; Chen, W.W.; Hu, X.Y.; Jiang, A.L.; Zhao, J. Clinicopathological and prognostic significance of microRNA-107 in human non small cell lung cancer. Int. J. Clin. Exp. Pathol. 2014, 7, 4545–4551. [Google Scholar] [PubMed]
- Xia, H.; Li, Y.; Lv, X. MicroRNA-107 inhibits tumor growth and metastasis by targeting the BDNF-mediated PI3K/AKT pathway in human non-small lung cancer. Int. J. Oncol. 2016, 49, 1325–1333. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Han, Y.; Liu, Y.; Han, L.; Liu, J. miRNA-124 down-regulates SOX8 expression and suppresses cell proliferation in non-small cell lung cancer. Int. J. Clin. Exp. Pathol. 2014, 7, 6534–6542. [Google Scholar] [PubMed]
- Yu, W.; Peng, W.; Jiang, H.; Sha, H.; Li, J. LncRNA HOXA11-AS promotes proliferation and invasion by targeting miR-124 in human non-small cell lung cancer cells. Tumour Biol. 2017, 39, 1010428317721440. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Huang, Q.; Zhang, S.; Zhang, Q.; Chang, J.; Qiu, X.; Wang, E. Hsa-miR-125a-3p and hsa-miR-125a-5p are downregulated in non-small cell lung cancer and have inverse effects on invasion and migration of lung cancer cells. BMC Cancer 2010, 10, 318. [Google Scholar] [CrossRef] [PubMed]
- Hou, L.; Luo, P.; Ma, Y.; Jia, C.; Yu, F.; Lv, Z.; Wu, C.; Fu, D. MicroRNA-125a-3p downregulation correlates with tumorigenesis and poor prognosis in patients with non-small cell lung cancer. Oncol. Lett. 2017, 14, 4441–4448. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Wu, J.; Shi, J.; Lu, C.; Wang, Y.; Sun, Q.; Zhang, G.; Zhao, G. miR-125a-5p upregulation suppresses the proliferation and induces the cell apoptosis of lung adenocarcinoma by targeting NEDD9. Oncol. Rep. 2017, 38, 1790–1796. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Wei, F.; Yu, J.; Zhao, H.; Jia, L.; Ye, Y.; Du, R.; Ren, X.; Li, H. Matrix metalloproteinase 13: A potential intermediate between low expression of microRNA-125b and increasing metastatic potential of non-small cell lung cancer. Cancer Genet. 2015, 208, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Gu, Y.Y.; Ma, F.C.; He, R.Q.; Li, Z.Y.; Zhai, G.Q.; Lin, X.; Hu, X.H.; Pan, L.J.; Chen, G. Expression levels and cotargets of miRNA1263p and miRNA1265p in lung adenocarcinoma tissues: Alphan exploration with RTqPCR, microarray and bioinformatic analyses. Oncol. Rep. 2019, 41, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.W.; Wang, T.B.; Tian, Y.H.; Zheng, Y.G. Down-regulation of microRNA-126 and microRNA-133b acts as novel predictor biomarkers in progression and metastasis of non small cell lung cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 14983–14988. [Google Scholar] [PubMed]
- Hu, J.; Cheng, Y.; Li, Y.; Jin, Z.; Pan, Y.; Liu, G.; Fu, S.; Zhang, Y.; Feng, K.; Feng, Y. microRNA-128 plays a critical role in human non-small cell lung cancer tumourigenesis, angiogenesis and lymphangiogenesis by directly targeting vascular endothelial growth factor-C. Eur. J. Cancer 2014, 50, 2336–2350. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Jiang, Y. LncRNA NNT-AS1 promotes the proliferation, and invasion of lung cancer cells via regulating miR-129-5p expression. Biomed. Pharmacother. 2018, 105, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Wang, Y.; Nie, L.; Qian, S.; Xu, M. MiR-130 exerts tumor suppressive function on the tumorigenesis of human non-small cell lung cancer by targeting PTEN. Am. J. Transl. Res. 2017, 9, 1856–1865. [Google Scholar] [PubMed]
- Wang, X.C.; Tian, L.L.; Wu, H.L.; Jiang, X.Y.; Du, L.Q.; Zhang, H.; Wang, Y.Y.; Wu, H.Y.; Li, D.G.; She, Y.; et al. Expression of miRNA-130a in nonsmall cell lung cancer. Am. J. Med. Sci. 2010, 340, 385–388. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.Q.; Wu, C.A.; Cheng, Y.X. Prognostic Value of microRNA-133a Expression and Its Clinicopathologic Significance in Non-Small Cell Lung Cancer: A Comprehensive Study Based on Meta-Analysis and the TCGA Database. Oncol. Res. Treat. 2018, 41, 762–768. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Shao, X.; Gao, W.; Zhang, Z.; Liu, P.; Wang, R.; Huang, P.; Yin, Y.; Shu, Y. MicroRNA-133b inhibits the growth of non-small-cell lung cancer by targeting the epidermal growth factor receptor. FEBS J. 2012, 279, 3800–3812. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.; Jing, H.; Li, Y.; Lv, X. Long noncoding RNA HOXD-AS1 promotes non-small cell lung cancer migration and invasion through regulating miR-133b/MMP9 axis. Biomed. Pharmacother. 2018, 106, 156–162. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Zhang, G.; Zhang, N.; Li, H.; Liu, Y.; Fu, A.; Zheng, Y. Prognostic potential of microRNA-138 and its target mRNA PDK1 in sera for patients with non-small cell lung cancer. Med. Oncol. 2014, 31, 129. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.; Fang, B.; Pan, J.; Zhang, N.; Huang, J.; Xie, C.; Lou, T.; Cao, Z. miR-138 suppresses the proliferation, metastasis and autophagy of non-small cell lung cancer by targeting Sirt1. Oncol. Rep. 2017, 37, 3244–3252. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L.; Zhou, H.; Li, X.P.; Chen, J.; Fang, C.; Mao, C.X.; Cui, J.J.; Zhang, W.; Zhou, H.H.; Yin, J.Y.; et al. MicroRNA-138 acts as a tumor suppressor in non small cell lung cancer via targeting YAP1. Oncotarget 2016, 7, 40038–40046. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Amano, Y.; Ishikawa, R.; Sunohara, M.; Kage, H.; Ichinose, J.; Sano, A.; Nakajima, J.; Fukayama, M.; Yatomi, Y.; et al. Histone methylation-mediated silencing of miR-139 enhances invasion of non-small-cell lung cancer. Cancer Med. 2015, 4, 1573–1582. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, P.; Rong, M.; He, R.; Hou, X.; Xie, Y.; Chen, G. MicroRNA-141 is a biomarker for progression of squamous cell carcinoma and adenocarcinoma of the lung: Clinical analysis of 125 patients. Tohoku J. Exp. Med. 2015, 235, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Wang, J.; Deng, J.; Li, X.; Long, W.; Chang, Y. MiR-145 acts as a metastasis suppressor by targeting metadherin in lung cancer. Med. Oncol. 2015, 32, 344. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Chen, H.; You, Q.; Ye, Q.; Wang, F.; Wang, S.; Zhang, S.; Yu, K.; Li, W.; Gu, M. miR145 inhibits human nonsmall-cell lung cancer growth by dual-targeting RIOK2 and NOB1. Int. J. Oncol. 2018, 53, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Gan, T.Q.; Xie, Z.C.; Tang, R.X.; Zhang, T.T.; Li, D.Y.; Li, Z.Y.; Chen, G. Clinical value of miR-145-5p in NSCLC and potential molecular mechanism exploration: A retrospective study based on GEO, qRT-PCR, and TCGA data. Tumour Biol. 2017, 39, 1010428317691683. [Google Scholar] [CrossRef] [PubMed]
- Chu, G.; Zhang, J.; Chen, X. Serum level of microRNA-147 as diagnostic biomarker in human non-small cell lung cancer. J. Drug Target. 2016, 24, 613–617. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Min, L.; Zhang, X.; Hu, S.; Wang, B.; Liu, W.; Wang, R.; Gu, X.; Shen, W.; Lv, H.; et al. Decreased miRNA-148a is associated with lymph node metastasis and poor clinical outcomes and functions as a suppressor of tumor metastasis in non-small cell lung cancer. Oncol. Rep. 2013, 30, 1832–1840. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Min, L.; Ren, C.; Xu, X.; Yang, J.; Sun, X.; Wang, T.; Wang, F.; Sun, C.; Zhang, X. miRNA-148a serves as a prognostic factor and suppresses migration and invasion through Wnt1 in non-small cell lung cancer. PLoS ONE 2017, 12, e0171751. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yu, T.; Cao, J.; Liu, L.; Liu, Y.; Kong, H.W.; Zhu, M.X.; Lin, H.C.; Chu, D.D.; Yao, M.; et al. MicroRNA-148a Suppresses Invasion and Metastasis of Human Non-Small-Cell Lung Cancer. Cell. Physiol. Biochem. 2015, 37, 1847–1856. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Li, B.; Hu, W.X.; Li, R.J.; Jin, H.; Gao, M.M.; Ding, C.M. MicroRNA-148b is down-regulated in non-small cell lung cancer and associated with poor survival. Int. J. Clin. Exp. Pathol. 2015, 8, 800–805. [Google Scholar] [PubMed]
- Yin, Q.W.; Sun, X.F.; Yang, G.T.; Li, X.B.; Wu, M.S.; Zhao, J. Increased expression of microRNA-150 is associated with poor prognosis in non-small cell lung cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 842–846. [Google Scholar] [PubMed]
- Shan, N.; Shen, L.; Wang, J.; He, D.; Duan, C. MiR-153 inhibits migration and invasion of human non-small-cell lung cancer by targeting ADAM19. Biochem. Biophys. Res. Commun. 2015, 456, 385–391. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.J.; Zhang, E.N.; Zhong, Z.K.; Jiang, M.Z.; Yang, X.F.; Zhou, D.M.; Wang, X.W. MicroRNA-153 expression and prognosis in non-small cell lung cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 8671–8675. [Google Scholar] [PubMed]
- Li, S.; Yang, J.; Xia, Y.; Fan, Q.; Yang, K.P. Long Noncoding RNA NEAT1 Promotes Proliferation and Invasion via Targeting miR-181a-5p in Non-Small Cell Lung Cancer. Oncol. Res. 2018, 26, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, H.; Wang, H.; Sun, Y. Down-regulation of microRNA-181b is a potential prognostic marker of non-small cell lung cancer. Pathol. Res. Pract. 2013, 209, 490–494. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Zhang, H.; Su, Y.; Kong, J.; Yu, H.; Qian, B. Up-regulation of microRNA-183-3p is a potent prognostic marker for lung adenocarcinoma of female non-smokers. Clin. Transl. Oncol. 2014, 16, 980–985. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhang, Y.; Liu, J.; Yin, W.; Jin, D.; Wang, D.; Zhang, W. MiR-185 inhibits cell proliferation and invasion of non-small cell lung cancer by targeting KLF7. Oncol. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Jin, X.; Sun, Z.; Zhao, Y.; Song, X. MiR-186 Inhibited Migration of NSCLC via Targeting cdc42 and Effecting EMT Process. Mol. Cells 2017, 40, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yin, C.; Zhang, B.; Sun, Y.; Shi, L.; Liu, N.; Liang, S.; Lu, S.; Liu, Y.; Zhang, J.; et al. PTTG1 promotes migration and invasion of human non-small cell lung cancer cells and is modulated by miR-186. Carcinogenesis 2013, 34, 2145–2155. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Li, J.; Yan, M.; Liu, L.; Lin, H.; Zhao, F.; Sun, L.; Zhang, Y.; Cui, Y.; Zhang, F.; et al. MicroRNA-193a-3p and -5p suppress the metastasis of human non-small-cell lung cancer by downregulating the ERBB4/PIK3R3/mTOR/S6K2 signaling pathway. Oncogene 2015, 34, 413–423. [Google Scholar] [CrossRef] [PubMed]
- Ren, F.; Ding, H.; Huang, S.; Wang, H.; Wu, M.; Luo, D.; Dang, Y.; Yang, L.; Chen, G. Expression and clinicopathological significance of miR-193a-3p and its potential target astrocyte elevated gene-1 in non-small lung cancer tissues. Cancer Cell Int. 2015, 15, 80. [Google Scholar] [CrossRef] [PubMed]
- Trang, P.; Wiggins, J.F.; Daige, C.L.; Cho, C.; Omotola, M.; Brown, D.; Weidhaas, J.B.; Bader, A.G.; Slack, F.J. Systemic delivery of tumor suppressor microRNA mimics using a neutral lipid emulsion inhibits lung tumors in mice. Mol. Ther. 2011, 19, 1116–1122. [Google Scholar] [CrossRef] [PubMed]
- Su, K.; Zhang, T.; Wang, Y.; Hao, G. Diagnostic and prognostic value of plasma microRNA-195 in patients with non-small cell lung cancer. World J. Surg. Oncol. 2016, 14, 224. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zhang, G.; Li, P.; Chen, S.; Zhang, F.; Li, J.; Jiang, C.; Chen, X.; Wang, Y.; Du, Y.; et al. miR-198 targets SHMT1 to inhibit cell proliferation and enhance cell apoptosis in lung adenocarcinoma. Tumour Biol. 2016, 37, 5193–5202. [Google Scholar] [CrossRef] [PubMed]
- Ceppi, P.; Mudduluru, G.; Kumarswamy, R.; Rapa, I.; Scagliotti, G.V.; Papotti, M.; Allgayer, H. Loss of miR-200c expression induces an aggressive, invasive, and chemoresistant phenotype in non-small cell lung cancer. Mol. Cancer Res. 2010, 8, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- Si, L.; Tian, H.; Yue, W.; Li, L.; Li, S.; Gao, C.; Qi, L. Potential use of microRNA-200c as a prognostic marker in non-small cell lung cancer. Oncol. Lett. 2017, 14, 4325–4330. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Lv, B.; Zhang, L.; Zhao, N.; Lv, Y. miR-202 functions as a tumor suppressor in non-small cell lung cancer by targeting STAT3. Mol. Med. Rep. 2017, 16, 2281–2289. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Wang, F.; Lu, S.; Wang, X. LASP-1, regulated by miR-203, promotes tumor proliferation and aggressiveness in human non-small cell lung cancer. Exp. Mol. Pathol. 2016, 100, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Osugi, J.; Kimura, Y.; Owada, Y.; Inoue, T.; Watanabe, Y.; Yamaura, T.; Fukuhara, M.; Muto, S.; Okabe, N.; Matsumura, Y.; et al. Prognostic Impact of Hypoxia-Inducible miRNA-210 in Patients with Lung Adenocarcinoma. J. Oncol. 2015, 2015, 316745. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.H.; Zhang, H.; Yang, Z.G.; Wen, G.Q.; Cui, Y.B.; Shao, G.G. Prognostic significance of serum microRNA-210 levels in nonsmall-cell lung cancer. J. Int. Med. Res. 2013, 41, 1437–1444. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.X.; Zhang, M.Y.; Liu, K.; Liu, J.; Zhang, Z.L.; Fu, L. LncRNA SNHG15 promotes proliferation and migration of lung cancer via targeting microRNA-211-3p. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 6838–6844. [Google Scholar] [CrossRef] [PubMed]
- Tang, T.; Huan, L.; Zhang, S.; Zhou, H.; Gu, L.; Chen, X.; Zhang, L. MicroRNA-212 functions as a tumor-suppressor in human non-small cell lung cancer by targeting SOX4. Oncol. Rep. 2017, 38, 2243–2250. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhao, Y.; Sun, S.; Liu, Z.; Zhang, Y.; Jiao, S. Overexpression of MicroRNA-221 is associated with poor prognosis in non-small cell lung cancer patients. Tumour Biol. 2016, 37, 10155–10160. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Chen, H.; Yang, X.; Chen, W.; Wang, L.; Xu, J.; Yu, L. Decreased microRNA-224 and its clinical significance in non-small cell lung cancer patients. Diagn. Pathol. 2014, 9, 198. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.K.; Zang, Q.L.; Li, G.X.; Huang, Y.; Wang, S.Z. Increased expression of microRNA-301a in nonsmall-cell lung cancer and its clinical significance. J. Cancer Res. Ther. 2016, 12, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Ma, C.N.; Zhou, N.N.; Li, X.D.; Zhang, Y.J. Up- regulation of miR-328-3p sensitizes non-small cell lung cancer to radiotherapy. Sci. Rep. 2016, 6, 31651. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Li, M.; Zhang, R.; Wang, Y.; Zang, W.; Ma, Y.; Zhao, G.; Zhang, G. Effect of miR-335 upregulation on the apoptosis and invasion of lung cancer cell A549 and H1299. Tumour Biol. 2013, 34, 3101–3109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Shao, G.; Lin, X.; Liu, Y.; Yang, Z. MiR-338-3p inhibits the growth and invasion of non-small cell lung cancer cells by targeting IRS2. Am. J. Cancer Res. 2017, 7, 53–63. [Google Scholar] [PubMed]
- Li, Y.; Zhao, W.; Bao, P.; Li, C.; Ma, X.Q.; Li, Y.; Chen, L.A. miR-339-5p inhibits cell migration and invasion in vitro and may be associated with the tumor-node-metastasis staging and lymph node metastasis of non-small cell lung cancer. Oncol. Lett. 2014, 8, 719–725. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, X.; Yang, Z.; Li, Y.; Han, B.; Chen, L.A. miR-339-5p inhibits metastasis of non-small cell lung cancer by regulating the epithelial-to-mesenchymal transition. Oncol. Lett. 2018, 15, 2508–2514. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Zhou, X.; Huang, C.; Li, L.; Liu, H.; Liang, N.; Chen, Y.; Ma, D.; Han, Z.; Xu, X.; et al. Lower miR-340 expression predicts poor prognosis of non-small cell lung cancer and promotes cell proliferation by targeting CDK4. Gene 2018, 675, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Z.L.; Tian, F.M.; Sun, C.L. Downregulation of miR-361-5p associates with aggressive clinicopathological features and unfavorable prognosis in non-small cell lung cancer. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 5132–5136. [Google Scholar] [PubMed]
- Liu, Y.; Zhang, G.; Li, H.; Han, L.; Fu, A.; Zhang, N.; Zheng, Y. Serum microRNA-365 in combination with its target gene TTF-1 as a non-invasive prognostic marker for non-small cell lung cancer. Biomed. Pharmacother. 2015, 75, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Sun, R.; Liu, Z.; Ma, G.; Lv, W.; Zhao, X.; Lei, G.; Xu, C. Associations of deregulation of mir-365 and its target mRNA TTF-1 and survival in patients with NSCLC. Int. J. Clin. Exp. Pathol. 2015, 8, 2392–2399. [Google Scholar] [PubMed]
- Skrzypek, K.; Tertil, M.; Golda, S.; Ciesla, M.; Weglarczyk, K.; Collet, G.; Guichard, A.; Kozakowska, M.; Boczkowski, J.; Was, H.; et al. Interplay between heme oxygenase-1 and miR-378 affects non-small cell lung carcinoma growth, vascularization, and metastasis. Antioxid. Redox Signal. 2013, 19, 644–660. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Nie, L.; Feng, D.; Guo, S.; Luo, R. MicroRNA-379 acts as a tumor suppressor in non-small cell lung cancer by targeting the IGF1R-mediated AKT and ERK pathways. Oncol. Rep. 2017, 38, 1857–1866. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Li, J.; Ren, L.; Peng, R.; Chen, B.; Lin, Y. MicroRNA-381 serves as a prognostic factor and inhibits migration and invasion in non-small cell lung cancer by targeting LRH-1. Oncol. Rep. 2017, 38, 3071–3077. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Zang, A.; Li, J.; Jia, Y.; Li, X.; Zhang, L.; Huo, R.; Yang, J.; Feng, J.; Ge, K.; et al. MicroRNA-383 is a tumor suppressor and potential prognostic biomarker in human non-small cell lung caner. Biomed. Pharmacother. 2016, 83, 1175–1181. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.; Zhu, L.; Xu, J.; Lu, B.; Yang, Y.; Liu, F.; Wang, Z. MicroRNA-409-3p functions as a tumor suppressor in human lung adenocarcinoma by targeting c-Met. Cell. Physiol. Biochem. 2014, 34, 1273–1290. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.Y.; Li, Y.; Jiang, Y.S.; Li, R.Z. Investigation of serum miR-411 as a diagnosis and prognosis biomarker for non-small cell lung cancer. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 4092–4097. [Google Scholar] [PubMed]
- Li, Y.; Cui, X.; Li, Y.; Zhang, T.; Li, S. Upregulated expression of miR-421 is associated with poor prognosis in non-small-cell lung cancer. Cancer Manag. Res. 2018, 10, 2627–2633. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Hu, B.; Zhao, B.; Liu, Y.; Yang, Y.; Zhang, L.; Chen, J. Circulating microRNA-422a is associated with lymphatic metastasis in lung cancer. Oncotarget 2017, 8, 42173–42188. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Liu, Z.; Zhang, W.; Li, Y.; Cao, J.; Yang, H.; Li, X. MicroRNA433 reduces cell proliferation and invasion in nonsmall cell lung cancer via directly targeting E2F transcription factor 3. Mol. Med. Rep. 2018, 18, 1155–1164. [Google Scholar] [CrossRef] [PubMed]
- Shan, C.; Fei, F.; Li, F.; Zhuang, B.; Zheng, Y.; Wan, Y.; Chen, J. miR-448 is a novel prognostic factor of lung squamous cell carcinoma and regulates cells growth and metastasis by targeting DCLK1. Biomed. Pharmacother. 2017, 89, 1227–1234. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Huang, B.; Li, Z.; Li, H.; Sun, L.; Zhang, Q.; Qiu, X.; Wang, E. MicroRNA-449a is downregulated in non-small cell lung cancer and inhibits migration and invasion by targeting c-Met. PLoS ONE 2013, 8, e64759. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, Z.X.; Yang, J.S.; Pan, X.; De, W.; Chen, L.B. MicroRNA-451 functions as a tumor suppressor in human non-small cell lung cancer by targeting ras-related protein 14 (RAB14). Oncogene 2011, 30, 2644–2658. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.C.; Tian, L.L.; Jiang, X.Y.; Wang, Y.Y.; Li, D.G.; She, Y.; Chang, J.H.; Meng, A.M. The expression and function of miRNA-451 in non-small cell lung cancer. Cancer Lett. 2011, 311, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Kanaoka, R.; Iinuma, H.; Dejima, H.; Sakai, T.; Uehara, H.; Matsutani, N.; Kawamura, M. Usefulness of Plasma Exosomal MicroRNA-451a as a Noninvasive Biomarker for Early Prediction of Recurrence and Prognosis of Non-Small Cell Lung Cancer. Oncology 2018, 94, 311–323. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Xia, Y.; Pan, C.; Ma, T.; Liu, B.; Wang, J.; Chen, L.; Chen, Y. Up-Regulation of MiR-452 Inhibits Metastasis of Non-Small Cell Lung Cancer by Regulating BMI1. Cell. Physiol. Biochem. 2015, 37, 387–398. [Google Scholar] [CrossRef] [PubMed]
- Gan, X.N.; Luo, J.; Tang, R.X.; Wang, H.L.; Zhou, H.; Qin, H.; Gan, T.Q.; Chen, G. Clinical value of miR-452-5p expression in lung adenocarcinoma: A retrospective quantitative real-time polymerase chain reaction study and verification based on The Cancer Genome Atlas and Gene Expression Omnibus databases. Tumour Biol. 2017, 39, 1010428317705755. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.Y.; Li, X.N.; Qi, Y.; Liu, D.L.; Yang, Y.; Zhao, J.; Zhang, C.Y.; Wu, K.; Zhao, S. MiR-454 promotes the progression of human non-small cell lung cancer and directly targets PTEN. Biomed. Pharmacother. 2016, 81, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.S.; Zheng, Y.L.; Li, C.; Ding, C.; Xu, C.; Zhao, J. MicroRNA-485-5p suppresses growth and metastasis in non-small cell lung cancer cells by targeting IGF2BP2. Life Sci. 2018, 199, 104–111. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Liu, Z.; Todd, N.W.; Zhang, H.; Liao, J.; Yu, L.; Guarnera, M.A.; Li, R.; Cai, L.; Zhan, M.; et al. Diagnosis of lung cancer in individuals with solitary pulmonary nodules by plasma microRNA biomarkers. BMC Cancer 2011, 11, 374. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tian, X.; Han, R.; Zhang, X.; Wang, X.; Shen, H.; Xue, L.; Liu, Y.; Yan, X.; Shen, J.; et al. Downregulation of miR-486-5p contributes to tumor progression and metastasis by targeting protumorigenic ARHGAP5 in lung cancer. Oncogene 2014, 33, 1181–1189. [Google Scholar] [CrossRef] [PubMed]
- Pang, W.; Tian, X.; Bai, F.; Han, R.; Wang, J.; Shen, H.; Zhang, X.; Liu, Y.; Yan, X.; Jiang, F.; et al. Pim-1 kinase is a target of miR-486-5p and eukaryotic translation initiation factor 4E, and plays a critical role in lung cancer. Mol. Cancer 2014, 13, 240. [Google Scholar] [CrossRef] [PubMed]
- Gong, F.; Ren, P.; Zhang, Y.; Jiang, J.; Zhang, H. MicroRNAs-491-5p suppresses cell proliferation and invasion by inhibiting IGF2BP1 in non-small cell lung cancer. Am. J. Transl. Res. 2016, 8, 485–495. [Google Scholar] [PubMed]
- Zhang, J.; Wang, T.; Zhang, Y.; Wang, H.; Wu, Y.; Liu, K.; Pei, C. Upregulation of serum miR-494 predicts poor prognosis in non-small cell lung cancer patients. Cancer Biomark. 2018, 21, 763–768. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, H.; Liao, Y.; Chen, N.; Liu, T.; Zhang, H.; Zhang, H. Expression and clinical evidence of miR-494 and PTEN in non-small cell lung cancer. Tumour Biol. 2015, 36, 6965–6972. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.F.; Zhang, J.G.; Guo, T.X.; Pan, X.J. MiR-504 inhibits cell proliferation and invasion by targeting LOXL2 in non small cell lung cancer. Biomed. Pharmacother. 2018, 97, 1289–1295. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Yang, P.; Jiang, X.; Li, X.; Wang, Y.; Zhang, X.; Sun, B.; Zhang, Y.; Jia, Y. Genetic and epigenetic silencing of mircoRNA-506-3p enhances COTL1 oncogene expression to foster non-small lung cancer progression. Oncotarget 2017, 8, 644–657. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yu, S. Role of miR-520b in non-small cell lung cancer. Exp. Ther. Med. 2018, 16, 3987–3995. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Yang, H.; Xu, Z.; Li, D.; Zhou, M.; Xiao, K.; Shi, Z.; Zhu, L.; Yang, L.; Zhou, R. microRNA-548l is involved in the migration and invasion of non-small cell lung cancer by targeting the AKT1 signaling pathway. J. Cancer Res. Clin. Oncol. 2015, 141, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.L.; Zhang, M. miR-582-5p is a potential prognostic marker in human non-small cell lung cancer and functions as a tumor suppressor by targeting MAP3K2. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 7760–7767. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.; Lin, L.; Du, Y.; Hao, X.; Zhao, Y.; Liu, X. MicroRNA-588 suppresses tumor cell migration and invasion by targeting GRN in lung squamous cell carcinoma. Mol. Med. Rep. 2016, 14, 3021–3028. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, B.; Niu, L.; Ge, L. miR-592 functions as a tumor suppressor in human non-small cell lung cancer by targeting SOX9. Oncol. Rep. 2017, 37, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Lou, J.F.; Cao, Y.; Shi, X.H.; Wang, P.; Xu, J.; Xie, E.F.; Xu, T.; Sun, R.H.; Rao, J.Y.; et al. miR-638 is a new biomarker for outcome prediction of non-small cell lung cancer patients receiving chemotherapy. Exp. Mol. Med. 2015, 47, e162. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.Y.; Cui, S.Y.; Chen, Y.T.; Song, H.Z.; Huang, G.C.; Feng, B.; Sun, M.; De, W.; Wang, R.; Chen, L.B. MicroRNA-650 was a prognostic factor in human lung adenocarcinoma and confers the docetaxel chemoresistance of lung adenocarcinoma cells via regulating Bcl-2/Bax expression. PLoS ONE 2013, 8, e72615. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Zhou, C.; Luo, M.; Shi, X.; Li, Y.; Sun, Z.; Zhou, F.; Chen, Z.; He, J. MiR-652-3p is up-regulated in non-small cell lung cancer and promotes proliferation and metastasis by directly targeting Lgl1. Oncotarget 2016, 7, 16703–16715. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, Y.; Wu, B.; Shi, C.; Li, C. MicroRNA-661 promotes non-small cell lung cancer progression by directly targeting RUNX3. Mol. Med. Rep. 2017, 16, 2113–2120. [Google Scholar] [CrossRef] [PubMed]
- He, D.; Wang, J.; Zhang, C.; Shan, B.; Deng, X.; Li, B.; Zhou, Y.; Chen, W.; Hong, J.; Gao, Y.; et al. Down-regulation of miR-675-5p contributes to tumor progression and development by targeting pro-tumorigenic GPR55 in non-small cell lung cancer. Mol. Cancer 2015, 14, 73. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Li, P.; Zhang, G.; Lu, C.; Wang, H.; Zhao, G. Long Non-Coding RNA XLOC_008466 Functions as an Oncogene in Human Non-Small Cell Lung Cancer by Targeting miR-874. Cell. Physiol. Biochem. 2017, 42, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, S.; Xu, J.; Niu, W.; Li, S. microRNA-935 is reduced in non-small cell lung cancer tissue, is linked to poor outcome, and acts on signal transduction mediator E2F7 and the AKT pathway. Br. J. Biomed. Sci. 2019, 76, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; He, Y.; Shi, Y.; Guo, Z.; Yang, S.; Wei, K.; Pan, C.; Xia, Y.; Chen, Y. MicroRNA-1204 promotes cell proliferation by regulating PITX1 in non-small cell lung cancer. Cell Biol. Int. 2018. [Google Scholar] [CrossRef] [PubMed]
- Bao, M.; Song, Y.; Xia, J.; Li, P.; Liu, Q.; Wan, Z. miR-1269 promotes cell survival and proliferation by targeting tp53 and caspase-9 in lung cancer. Onco-Targets Ther. 2018, 11, 1721–1732. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.J.; Liu, Y.H.; Si, J.M.; Ni, R.; Wang, J. Overexpression of miR-1290 contributes to cell proliferation and invasion of non small cell lung cancer by targeting interferon regulatory factor 2. Int. J. Biochem. Cell Biol. 2018, 95, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Mo, D.; Gu, B.; Gong, X.; Wu, L.; Wang, H.; Jiang, Y.; Zhang, B.; Zhang, M.; Zhang, Y.; Xu, J.; et al. miR-1290 is a potential prognostic biomarker in non-small cell lung cancer. J. Thorac. Dis. 2015, 7, 1570–1579. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Ren, Y.; Zuo, T. Long noncoding RNA LINC00978 promotes cell proliferation and invasion in nonsmall cell lung cancer by inhibiting miR67545p. Mol. Med. Rep. 2018, 18, 4725–4732. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Gorski, D.H. Regulation of angiogenesis through a microRNA (miR-130a) that down-regulates antiangiogenic homeobox genes GAX and HOXA5. Blood 2008, 111, 1217–1226. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Egranov, S.D.; Yang, L.; Lin, C. Molecular Mechanisms of Long Noncoding RNAs-mediated Cancer Metastasis. Genes Chromosomes Cancer 2018. [Google Scholar] [CrossRef] [PubMed]
- Sanchez Calle, A.; Kawamura, Y.; Yamamoto, Y.; Takeshita, F.; Ochiya, T. Emerging roles of long non-coding RNA in cancer. Cancer Sci. 2018, 109, 2093–2100. [Google Scholar] [CrossRef] [PubMed]
- Berghmans, T.; Ameye, L.; Willems, L.; Paesmans, M.; Mascaux, C.; Lafitte, J.J.; Meert, A.P.; Scherpereel, A.; Cortot, A.B.; Cstoth, I.; et al. Identification of microRNA-based signatures for response and survival for non-small cell lung cancer treated with cisplatin-vinorelbine A ELCWP prospective study. Lung Cancer 2013, 82, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Boeri, M.; Milione, M.; Proto, C.; Signorelli, D.; Lo Russo, G.; Galeone, C.; Verri, C.; Mensah, M.; Centonze, G.; Martinetti, A.; et al. Circulating microRNAs and PD-L1 tumor expression are associated with survival in advanced NSCLC patients treated with immunotherapy: A prospective study. Clin. Cancer Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Portnoy, V.; Lin, S.H.; Li, K.H.; Burlingame, A.; Hu, Z.H.; Li, H.; Li, L.C. saRNA-guided Ago2 targets the RITA complex to promoters to stimulate transcription. Cell Res. 2016, 26, 320–335. [Google Scholar] [CrossRef] [PubMed]
- Krutzfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Elmen, J.; Lindow, M.; Silahtaroglu, A.; Bak, M.; Christensen, M.; Lind-Thomsen, A.; Hedtjarn, M.; Hansen, J.B.; Hansen, H.F.; Straarup, E.M.; et al. Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Res. 2008, 36, 1153–1162. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002, 297, 2056–2060. [Google Scholar] [CrossRef] [PubMed]
- Kota, J.; Chivukula, R.R.; O’Donnell, K.A.; Wentzel, E.A.; Montgomery, C.L.; Hwang, H.W.; Chang, T.C.; Vivekanandan, P.; Torbenson, M.; Clark, K.R.; et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 2009, 137, 1005–1017. [Google Scholar] [CrossRef] [PubMed]
- Gumireddy, K.; Young, D.D.; Xiong, X.; Hogenesch, J.B.; Huang, Q.; Deiters, A. Small-molecule inhibitors of microrna miR-21 function. Angew. Chem. Int. Ed. Engl. 2008, 47, 7482–7484. [Google Scholar] [CrossRef] [PubMed]
- Bakhshandeh, B.; Soleimani, M.; Hafizi, M.; Ghaemi, N. A comparative study on nonviral genetic modifications in cord blood and bone marrow mesenchymal stem cells. Cytotechnology 2012, 64, 523–540. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Kanasty, R.L.; Eltoukhy, A.A.; Vegas, A.J.; Dorkin, J.R.; Anderson, D.G. Non-viral vectors for gene-based therapy. Nat. Rev. Genet. 2014, 15, 541–555. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Reinhardt, F.; Pan, E.; Soutschek, J.; Bhat, B.; Marcusson, E.G.; Teruya-Feldstein, J.; Bell, G.W.; Weinberg, R.A. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 2010, 28, 341–347. [Google Scholar] [CrossRef] [PubMed]
- Tazawa, H.; Tsuchiya, N.; Izumiya, M.; Nakagama, H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc. Natl. Acad. Sci. USA 2007, 104, 15472–15477. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhu, X.; Zhang, X.; Liu, B.; Huang, L. Nanoparticles modified with tumor-targeting scFv deliver siRNA and miRNA for cancer therapy. Mol. Ther. 2010, 18, 1650–1656. [Google Scholar] [CrossRef] [PubMed]
- Trang, P.; Medina, P.P.; Wiggins, J.F.; Ruffino, L.; Kelnar, K.; Omotola, M.; Homer, R.; Brown, D.; Bader, A.G.; Weidhaas, J.B.; et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 2010, 29, 1580–1587. [Google Scholar] [CrossRef] [PubMed]
- Wiggins, J.F.; Ruffino, L.; Kelnar, K.; Omotola, M.; Patrawala, L.; Brown, D.; Bader, A.G. Development of a lung cancer therapeutic based on the tumor suppressor microRNA-34. Cancer Res. 2010, 70, 5923–5930. [Google Scholar] [CrossRef] [PubMed]
- Stahlhut, C.; Slack, F.J. Combinatorial Action of MicroRNAs let-7 and miR-34 Effectively Synergizes with Erlotinib to Suppress Non-small Cell Lung Cancer Cell Proliferation. Cell Cycle 2015, 14, 2171–2180. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Bai, Y.; Zhang, F.; Wang, Y.; Guo, Y.; Guo, L. miR-126 inhibits non-small cell lung cancer cells proliferation by targeting EGFL7. Biochem. Biophys. Res. Commun. 2010, 391, 1483–1489. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zeng, H.; Guo, Y.; Liu, P.; Pan, H.; Deng, A.; Hu, J. miRNA-145 inhibits non-small cell lung cancer cell proliferation by targeting c-Myc. J. Exp. Clin. Cancer Res. 2010, 29, 151. [Google Scholar] [CrossRef] [PubMed]
- Chiou, G.Y.; Cherng, J.Y.; Hsu, H.S.; Wang, M.L.; Tsai, C.M.; Lu, K.H.; Chien, Y.; Hung, S.C.; Chen, Y.W.; Wong, C.I.; et al. Cationic polyurethanes-short branch PEI-mediated delivery of Mir145 inhibited epithelial-mesenchymal transdifferentiation and cancer stem-like properties and in lung adenocarcinoma. J. Control. Release 2012, 159, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Crawford, M.; Yu, B.; Mao, Y.; Nana-Sinkam, S.P.; Lee, L.J. MicroRNA delivery by cationic lipoplexes for lung cancer therapy. Mol. Pharm. 2011, 8, 1381–1389. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Crawford, M.; Mao, Y.; Lee, R.J.; Davis, I.C.; Elton, T.S.; Lee, L.J.; Nana-Sinkam, S.P. Therapeutic Delivery of MicroRNA-29b by Cationic Lipoplexes for Lung Cancer. Mol. Ther. Nucleic Acids 2013, 2, e84. [Google Scholar] [CrossRef] [PubMed]
- Bouchie, A. First microRNA mimic enters clinic. Nat. Biotechnol. 2013, 31, 577. [Google Scholar] [CrossRef] [PubMed]
- Beg, M.S.; Brenner, A.J.; Sachdev, J.; Borad, M.; Kang, Y.K.; Stoudemire, J.; Smith, S.; Bader, A.G.; Kim, S.; Hong, D.S. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Investig. New Drugs 2017, 35, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Bader, A.G. Evaluating Synergistic Effects of miR-34a Mimics in Combination with Other Therapeutic Agents in Cultured Non-Small Cell Lung Cancer Cells. Methods Mol. Biol. 2017, 1517, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Van Zandwijk, N.; Pavlakis, N.; Kao, S.C.; Linton, A.; Boyer, M.J.; Clarke, S.; Huynh, Y.; Chrzanowska, A.; Fulham, M.J.; Bailey, D.L.; et al. Safety and activity of microRNA-loaded minicells in patients with recurrent malignant pleural mesothelioma: A first-in-man, phase 1, open-label, dose-escalation study. Lancet Oncol 2017, 18, 1386–1396. [Google Scholar] [CrossRef]
- Reid, G.; Kao, S.C.; Pavlakis, N.; Brahmbhatt, H.; MacDiarmid, J.; Clarke, S.; Boyer, M.; van Zandwijk, N. Clinical development of TargomiRs, a miRNA mimic-based treatment for patients with recurrent thoracic cancer. Epigenomics 2016, 8, 1079–1085. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N.; Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 2010, 79, 351–379. [Google Scholar] [CrossRef] [PubMed]
- Levin, A.A. Treating Disease at the RNA Level with Oligonucleotides. N. Engl. J. Med. 2019, 380, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.P.; Wang, Z.; Liu, X.Y.; Sun, Z.G.; Cheng, G.; Yu, Y. Inhibition effect of up-regulation of p21~(WAF1/CIP1) by small activating RNA on cell proliferation and promotes apoptosis in lung cancer cells. Chin. J. Cancer Prev. Treat. 2011, 14, 1080–1083. [Google Scholar]
- Wei, J.; Zhao, J.; Long, M.; Han, Y.; Wang, X.; Lin, F.; Ren, J.; He, T.; Zhang, H. p21WAF1/CIP1 gene transcriptional activation exerts cell growth inhibition and enhances chemosensitivity to cisplatin in lung carcinoma cell. BMC Cancer 2010, 10, 632. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Wang, L.; Gan, J.; Zhang, H. RNA activation: Promise as a new weapon against cancer. Cancer Lett. 2014, 355, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.J.; Chau, J.; Ebert, P.J.; Sylvester, G.; Min, H.; Liu, G.; Braich, R.; Manoharan, M.; Soutschek, J.; Skare, P.; et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 2007, 129, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Aagaard, L.; Rossi, J.J. RNAi therapeutics: Principles, prospects and challenges. Adv. Drug Deliv. Rev. 2007, 59, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Pan, F.; Holt, C.M.; Lewis, A.L.; Lu, J.R. Controlled delivery of antisense oligonucleotides: A brief review of current strategies. Expert Opin. Drug Deliv. 2009, 6, 673–686. [Google Scholar] [CrossRef] [PubMed]
- Grimm, D.; Streetz, K.L.; Jopling, C.L.; Storm, T.A.; Pandey, K.; Davis, C.R.; Marion, P.; Salazar, F.; Kay, M.A. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 2006, 441, 537–541. [Google Scholar] [CrossRef] [PubMed]
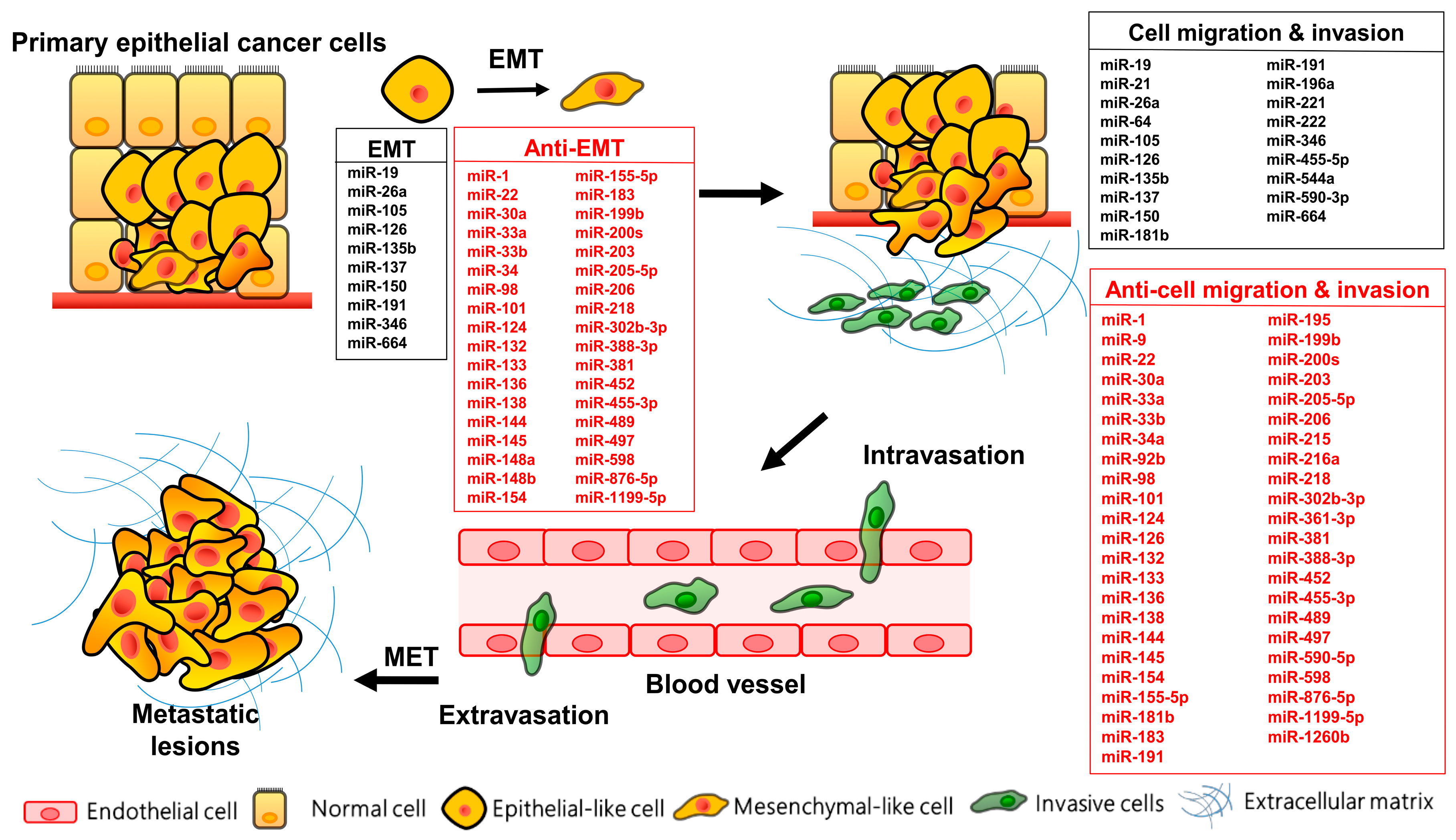
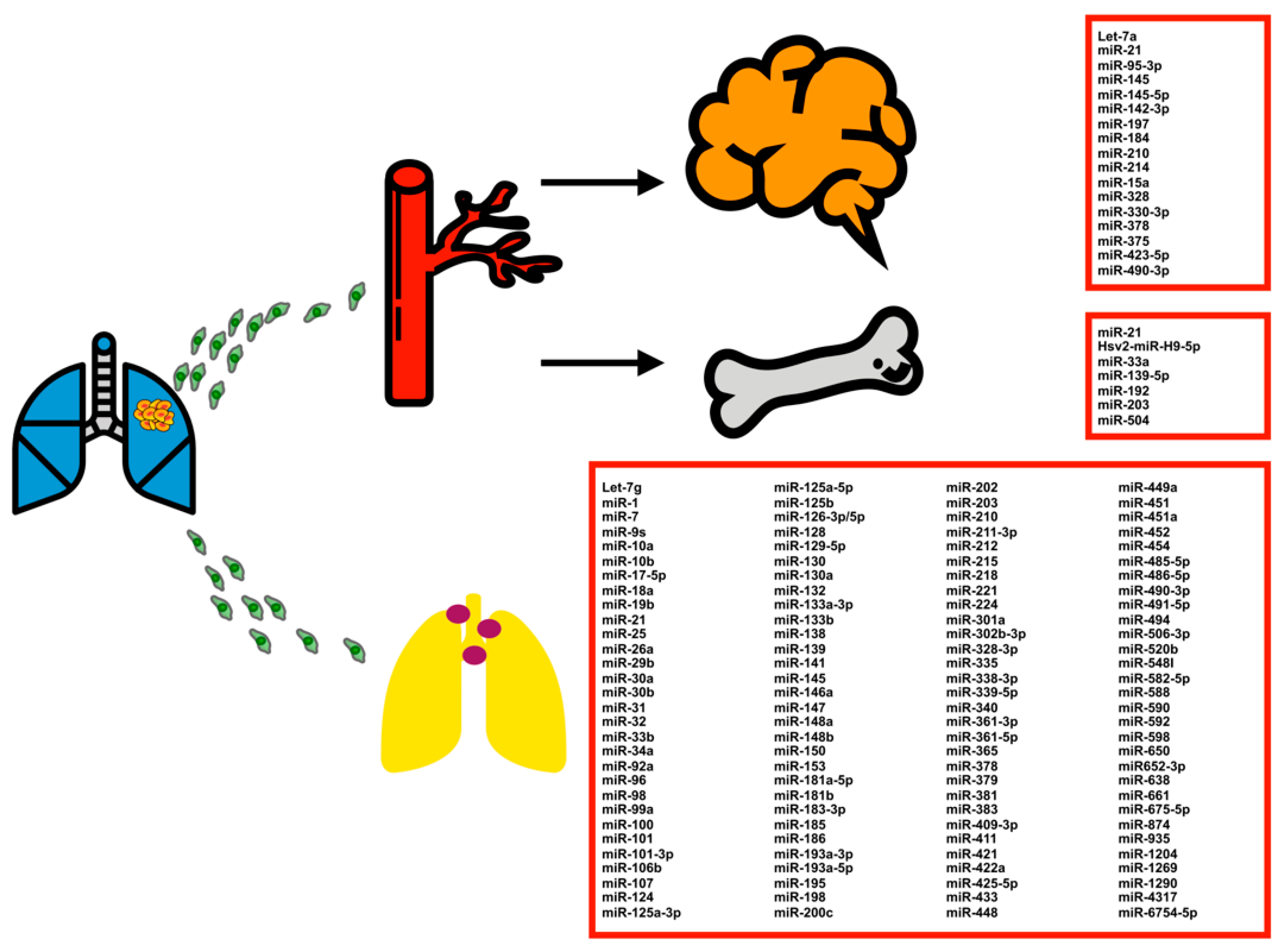
| Epithelial-to-Mesenchymal Transition-Related Transcription Factors | ||||||
|---|---|---|---|---|---|---|
| Snail | Slug | ZEB1 | ZEB2 | Twist | Other (miRNA/Related Gene or Target) | |
| miR-22 [36] | miR-1 [40] | miR-33b [41] | miR-132 [42] | miR-33a [43] | miR-19/PTEN [44] | miR-204/SIX1 [45] |
| miR-30 [37,38] | miR-124 [46,47] | miR-34a [48] | miR-138 [49] | miR-92b [50] | miR-21/Pdcd4 [51] | miR-205-5p/Integrin α5 [52] |
| miR-34 [33] | miR-137 [53] | miR-101 [54] | miR-145 [55] | miR-98 [56] | miR-26a/PTEN [57] | miR-205-5p/Smad4 [58] |
| miR-126 [34] | miR-218 [59] | miR-124 [60] | miR-154 [61] | miR-381 [39] | miR-105/Mcl-1 [62] | miR-206/MET [63] |
| miR-346 [35] | miR-452 [64] | miR-144 [65,66] | miR-155-5p [67] | miR-124/STAT3 [68] | miR-221&222/PTEN, TIMP3 [69] | |
| miR-381 [39] | miR-155-5p [67] | miR-200c [70] | miR-133/FOXQ1 [71] | miR-302b-3p/GCNT3 [72] | ||
| miR-181 [73] | miR-205-5p [74] | miR-135b/LZTS1, Hippo pathway [75] | miR-361-3p/SH2B1 [76] | |||
| miR-199b [77] | miR-203 [78] | miR-136/Smad2/3 [79] | miR-455-5p/SOCO3 [80] | |||
| miR-200s [81,82] | miR-215 [83] | miR-145/N-cadherin [84] | miR-489/SUL12 [85] | |||
| miR-205-5p [74] | miR-218 [59] | miR-145/MTDH [86] | miR-497/MTDH [86] | |||
| miR-216a [87] | miR-338-3p [88] | miR-148a/ROCK1 [89] | miR-544a/cadherina 1 [90] | |||
| miR-455-3p [91] | miR-598 [92] | miR-148b/ROCK1 [93] | miR-590-3p/OLFM4 [94] | |||
| miR-1199-5p [95] | miR-150/FOXO4 [96] | miR-590-5p/ADAM9 [97] | ||||
| miR-155-5p/Smad2/3 [67] | miR-590-5p/GAB1 [98] | |||||
| miR-183/MTA1 [99] | miR-598/Derlin 1 [100] | |||||
| miR-191/HIF-2α [101] | miR-664/AKT [102] | |||||
| miR-195/MYB [103] | miR-876-5p/BMP-4 [104] | |||||
| miR-196a/HOXA5 [105] | miR-1260b/PTPRK [106] | |||||
| miRNA | Direct or Related Target | Tumor Suppressor/Oncogene | Tissue | Effect | Author/Reference |
|---|---|---|---|---|---|
| miR-21 | COX19 | Oncogene | Metastatic bone tissue | Promoted cell proliferation, inhibited apoptosis | Guo et al. [149] |
| miR-21 | PDCD4 | Oncogene | TCGA database | Promoted osteoclastogenesis and tumorigenesis. High miR-21 correlated with poor prognosis according to TCGA database. | Xu et al. [152] |
| Hsv2-miR-H9-5p | SOCS2 | Oncogene | Lung tumors | Increased cell survival, migration, and invasion | Wang et al. [155] |
| miR-33a | PTHrP | Suppressor | Lung cancer cell lines (A549, H1299, and BEAS-2B) | Reduced the stimulatory effect of A549 on the production of osteoclastogenesis activator RANKL and M-CSF on osteoblasts, and increased the production of osteoprotegerin. | Kuo et al. [158] |
| miR-139-5p | Notch1 | Suppressor | Serum | MiR-139-5p expression was increased during MSC differentiation toward osteoblasts and positively regulated osteogenic differentiation. | Xu et al. [159] |
| miR-192 | ICAM-1 and PTPRJ | Suppressor | In vivo (mice) | Decreased tumor-induced angiogenesis | Valencia et al. [160] |
| miR-203 | TGF-β/SMAD2 | Suppressor | Tumor tissues | Suppressed cell proliferation and migration, induced apoptosis; repressed TGF-β/Smad2 | Wei et al. [161] |
| miRNA | Direct or Related Target | Tumor Suppressor/Oncogene | Tissue | Effect | Author/Reference |
|---|---|---|---|---|---|
| miR-21 | SPRY2, TIMP3, CDKN1A, SERPINB5 and PTEN. | Oncogene | In vivo | Promoted brain metastasis-initiating cell (BMIC) self-renewal and proliferation. | Singh et al. [166] |
| miR-95-3p | Cyclin D1 | Suppressor | In vivo | Inhibited cell invasion, proliferation, and colony formation. | Hwang et al. [167] |
| miR-145 | Suppressor | Brain and lung tumors | Inhibited cell proliferation | Zhao et al. [163] | |
| miR-145-5p | TPD52 | Suppressor | Brain and lung tumors | Inhibited cell invasion and migration. Restrained brain orthotopic tumor engraftment in vivo. | Donzelli et al. [165] |
| miR-142-3p | TRPA1 | Suppressor | TCGA data | Suppressed NSCLC progression | Berrout et al. [168] |
| miR-184, miR-197 | EGFR-mutant lung tumors | Remon et al. [169] | |||
| miR-15a, miR-210, miR-214 | Lung tumor | The Forest model of the three-miRNA signature could be used to predict brain metastasis of lung adenocarcinoma patients | Zhao et al. [170] | ||
| miR-328 | PRKCA | Oncogene | Brain and lung tumors | Increased cell migration. Up-regulated PRKCA | Arora et al. [171] |
| miR-330-3p | GRIA3 | Oncogene | Lung tumors | Promoted cell growth, invasion, and migration. Up-regulated total DNA methylation. Radiation-resistance | Wei et al. [172] Jiang et al. [173] |
| miR-375 | VEGF and MMP-9 | Suppressor | Brain and lung tumors | Chen et al. [174] | |
| miR-378 | MMP-2, MMP-9 and VEGF | Oncogene | Brain and lung tumors | Promoted cell migration, invasion, and tumor angiogenesis. | Chen et al. [175] |
| miR-423-5p | MTSS1 | Oncogene | Lung tumors | Promoted NSCLC cell colony formation, cell motility, invasion, and migration. | Sun et al. [176] |
| miR-490-3p | PCBP1 | Oncogene | Brain tissues | Promoted cell proliferation, invasion, and migration. | Li et al. [177] |
| miR-590 | ADAM9 | Suppressor | Lung tumors | Suppressed tumorigenesis and invasion. | Wang et al. [97] |
| miR-4317 | FGF9 and CCND2 | Suppressor | Lung tumors | Inhibited proliferation, colony formation, migration, and invasion, and hampered cycling | He et al. [178] |
| miRNA | Direct or Related Target | Tumor Suppressor/Oncogene | Tissue | Effect | Author/Reference |
|---|---|---|---|---|---|
| Let-7g | HMGA2, ERCC6 and MAP3K3 * | Suppressor | Lung tumor | The combination of Let-7g and miR-21 profiling and KRAS mutational status may be considered a useful biomarker for clinical management of NSCLC patients. | Capodanno et al. [194] |
| miR-1 | PIK3CA | Suppressor | Lung tumors | Low expression of miR-1 and overexpression of PIK3CA in NSCLC tissues may be useful predictors of lymph node metastasis and postoperative recurrence in patients with NSCLC. | Zhao et al. [195] |
| miR-7 | Bcl-2 | Suppressor | Lung tumor | Overexpressed CDR1as in NSCLC functioned to promote tumor progression via miR-7 signals. Up-regulated miR-7 increased the sensitivity of lung adenocarcinoma cells to CDDP by inducing apoptosis. | Zhang et al. [196], Cheng et al. [197] |
| miR-9s | Oncogene | Lung tumors | Involved in NSCLC progression and could serve as a promising biomarker. | Muraoka et al. [198], Xu et al. [199] | |
| miR-10a | PTEN | Oncogene | Lung tumors | Promoted NSCLC cell proliferation, migration, and invasion. | Yu et al. [200] |
| miR-10b | E-cadherin | Oncogene | Lunt tumors PBMC | E-cadherin mRNA and protein were overexpressed in miR-10b-suppressed cells compared with controls. MiR-10b expression in PBMCs had predictive value for tumor response to chemotherapy and prognosis for advanced NSCLC patients. | Zhang et al. [201], Yang et al. [202], Yang et al. [203], Li et al. [204] |
| miR-17-5p | Oncogene | Lung tumors | Increased cell proliferation. LncRNA HNF1A-AS1 promoted cell proliferation and invasion by directly targeting miR-17-5p in NSCLC. | Zhang et al. [205] | |
| miR-18a | Oncogene | Lung tumors | Correlated with stage, lymph node metastasis, and radio-resistance. | Shen et al. [206] | |
| miR-19a/b | Oncogene | Lunt tumors, Serum | MiR-19b is a potential biomarker for the prediction of survival and response to chemotherapy in NSCLC. | Lin et al. [207], Wu et al. [208] | |
| miR-21 | PTEN | Oncogene | Lung tumors, Serum | Reduced radio-sensitivity in vitro. Promoted cell proliferation and cell cycle progression. High serum level was associated with poor prognosis. | Liu et al. [209], Wang et al. [210], Liu et al. [211], Wang et al. [212], Tian et al. [213] |
| miR-25 | Oncogene | Lung adenocarcinoma tissues | Positive correlation with lymph node metastasis, stage, and EGFR mutations. | Xu et al. [214] | |
| miR-26a | PTEN | Oncogene | Lung tumors | Enhanced lung cancer cell migration and invasion abilities. Up-regulated β-catenin, MMP-2, Twist, and VEGF. | Liu et al. [57] |
| miR-29b | MMP2 | Suppressor | Lung tumor | Suppressed migration and invasion. | Wang et al. [215] |
| miR-30a | BCL11A | Suppressor | Lung tumors | A potential diagnostic and prognostic biomarker. | Jiang et al. [216] |
| miR-30b | EGFR Cthrc1 | Suppressor | Lung tumors | Inhibited proliferation, migration, and invasion, induced apoptosis, and enhanced sensitivity of NSCLC cells to EGFR-TKIs. | Qi et al. [217], Chen et al. [218] |
| miR-31 | CDK5, PTEN, p70S6K, ERK/MAPK, and PI3K/AKT # | Oncogene | Lung tumors | Promoted cell proliferation, invasion, and migration. | Meng et al. [219] |
| miR-32 | Suppressor | Lung tumors | Inversely correlated with tumor stage, lymph node metastasis, and OS. | Bai et al. [220] | |
| miR-33b | ZEB1 | Suppressor | Lung tumors | Inhibited cell growth, invasion, and EMT by suppressing Wnt/β-catenin/ZEB1 signaling. | Qu et al. [41] |
| miR-34a | Suppressor | Lung tumors and plasma | Plasma miR-34a negatively predicted lymph node metastasis. Lower miR-34a was correlated with longer survival. | Zhao et al. [221] | |
| miR-92a | PTEN | Oncogene | Lung tumors | Promoted cell growth, metastasis, and chemo-resistance. | Ren et al. [222] |
| miR-96 | FOXO3 | Oncogene | Lung tumors | Promoted cell invasion and inhibited apoptosis. | Li et al. [223] |
| miR-98 | Suppressor | Serum | Low serum miR-98 was positively correlated with advanced TNM stage, lymph node metastasis, and unfavorable OS. | Wang et al. [224] | |
| miR-99a | mTOR | Suppressor | Lung tumors | Inversely correlated with lymph node metastasis. | Gu et al. [225] |
| miR-100 | PLK1 | Suppressor | Lung tumors | Inhibited cell proliferation and caused G2/M cell cycle arrest | Liu et al. [226] |
| miR-101 | Mcl-1 ZEB1 | Suppressor | Lung tumors | Inhibited cell proliferation, invasion, and migration. | Luo et al. [227], Han et al. [54] |
| miR-101-3p | SOX9 | Suppressor | Lung tumors | LncRNA SNHG1 contributed to the progression of NSCLC through inhibition of miR-101-3p and activation of the Wnt/β-catenin signaling pathway. | Lu et al. [228], Cui et al. [229] |
| miR-106b | Oncogene | Lung tumor | Overexpression of miRNA-106b was strongly associated with lymph node metastasis and poor prognosis. | Li et al. [230] | |
| miR-107 | BNDF | Suppressor | Lung tumors | miR-107 significantly inversely correlated with tumor progression and decreased survival in patients with NSCLC. | Zhong et al. [231], Xia et al. [232] |
| miR-124 | SOX8, STAT3 | Suppressor | Lung tumors | Inhibited cell proliferation and induced apoptosis. HOXA11-AS acted as a competing endogenous RNA to regulate transcriptional factor Sp1 expression by sponging miR-124. | Xie et al. [233], Li et al. [68], Yu et al. [234] |
| miR-125a-3p | IGF2, CCL4 | Suppressor | Lung tumors | Suppressed cell invasion and migration. Inversely correlated with lymph node metastasis. | Jiang et al. [235], Hou et al. [236] |
| miR-125a-5p | NEDD9 | Uncertain | Lung tumors | The effects on cell invasion and migration and the relationship between miR-125a-5p and lymph node metastasis were controversial in lung cancers. | Jiang et al. [235] Zheng et al. [237] |
| miR-125b | MMP13 | Suppressor | Lung tumors | Inhibited cell invasion in vitro and in vivo. | Yu et al. [238] |
| miR-126-3p/5p | 44 co-targets ¶ | Suppressor | Lung tumors, | Lower expression of miRNA-126-3p and -5p was indicative of vascular invasion, lymph node spread, and an advanced TNM stage of lung adenocarcinoma. | Chen et al. [239], Chen et al. [240] |
| miR-128 | VEGF-C | Suppressor | Lung tumors | Inhibited cell proliferation, colony formation, invasion, and migration. | Hu et al. [241] |
| miR-129-5p | Suppressor | Lung tumor | LncRNA NNT-AS1 exerted functions in NSCLC by altering NNT-AS1/miR-129-5p axis | Shen et al. [242] | |
| miR-130 | PTEN | Suppressor | Lung tumor | Inhibited NSCLC cell growth and increased cell apoptosis. | Ye et al. [243] |
| miR-130a | Oncogene | Lung tumor | Overexpressed in NSCLC tissue and correlated with lymph node spreading. | Wang et al. [244] | |
| miR-132 | ZEB2 | Suppressor | Lung tumor | Inhibited cell proliferation, invasion, and migration, and decreased apoptosis | You et al. [42] |
| miR-133a-3p | Suppressor | Lung tumor (TCGA) | Associated with longer survival time and negative lymph node metastasis. | Yang et al. [245] | |
| miR-133b | EGFR | Suppressor | Lung tumor | Inhibited cell invasion, induced apoptosis, and enhanced sensitivity to gefitinib. HOXD-AS1 directly targeted miR-133b to promote cell migration and invasion. | Liu et al. [246], Xia et al. [247], Chen et al. [240] |
| miR-138 | PDK1, Sirt1, YAP1 | Suppressor | Lung tumor | Inversely correlated with lymph node metastasis | Han et al. [248], Ye et al. [249], Xiao et al. [250] |
| miR-139 | PDE2A | Suppressor | Lung tumor | H3K27me3-mediated down-regulation of miR-139. Enhanced invasive and metastasis ability of NSCLC cells. | Watanabe et al. [251] |
| miR-141 | Oncogene | Lung tumor | Positively associated with tumor size and, lymph node metastasis. | Zhang et al. [252] | |
| miR-145 | AEG-1/MTDH RIOK2, NOB1 N-cadherin | Suppressor | Lung tumors | Inhibited cell invasion and migration | Wang et al. [253], Liu et al. [254], Gan et al. [255], Mo et al. [84], Li et al. [204] |
| miR-146a | Suppressor | Serum | Lower serum level in NSCLC patients. | Wu et al. [208] | |
| miR-147 | Suppressor | Lung tumor, Serum | Low serum miR-147 expression level was correlated with lymph node metastasis and worse OS. | Chu et al. [256] | |
| miR-148a | ROCK1 Wnt1 | Suppressor | Lung tumors | Reduced cell invasion and inhibited EMT | Li et al. [89], Chen et al. [257], Chen et al. [258], Li et al. [259] |
| miR-148b | Suppressor | Lung tumors | High miR-148b expression had a favorable prognosis. | Ge et al. [260] | |
| miR-150 | Oncogene | Lung tumors | High miR-150 expression had a poor prognosis. | Yin et al. [261] | |
| miR-153 | ADAM19 | Suppressor | Lung tumors | Inhibited cell proliferation, migration, and invasion. | Shan et al. [262], Chen et al. [263] |
| miR-181a-5p | HMGB2 | Suppressor | Lung tumors | LncRNA NEAT1 promoted proliferation and invasion by targeting miR-181a-5p. | Li et al. [264] |
| miR-181b | Suppressor | Lung tumors | LncRNA NEAT1 up-regulated the miR-181a-5p-targeted gene HMGB2 through acting as a competitive "sponge" of miR-181a-5p. | Yang et al. [265] | |
| miR-183-3p | Oncogene | Lung adenocarcinoma tissues | Involved in lung cancer pathogenesis and progression, and could be used as a potential prognostic biomarker of female lung adenocarcinoma. | Xu et al. [266] | |
| miR-185 | KLF7 | Suppressor | Lung tumors | Inhibited the cell propagation, cell colony formation, and incursion capacities in vitro. | Zhao et al. [267] |
| miR-186 | Cdc42 | Suppressor | Lung tumors | Inhibited cell invasion and metastasis | Dong et al. [268], Li et al. [269] |
| miR-193a-3p | ERBB4, S6K2 AEG-1 | Suppressor | Lung tumors | Inhibited NSCLC cell migration, invasion, and EMT in vitro and lung metastasis formation in vivo. | Yu et al. [270], Ren et al. [271] |
| miR-193a-5p | PIK3R3 mTOR | Suppressor | Lung tumors | Inhibited NSCLC cell migration, invasion, and EMT in vitro and lung metastasis formation in vivo. | Yu et al. [272] |
| miR-195 | Suppressor | Plasma | Decreased plasma miRNA-195 expression was associated with lymph node metastasis and advanced clinical stage | Su et al. [273] | |
| miR-198 | SHMT1 | Suppressor | Lung adenocarcinoma | Inhibited cell proliferation, enhanced cell apoptosis, and led to cell-cycle arrest | Wu et al. [274] |
| miR-200c | USP25 | Suppressor | Lung tumors | Inhibited cell invasion and migration. Negatively correlated with lymph node metastasis. | Li et al. [115], Ceppi et al. [275] |
| miR-200c | Oncogene | Lung tumor | Higher expression of miR-200c was associated with poor prognosis. | Si et al. [276], Liu et al. [209] | |
| miR-202 | STAT3 | Suppressor | Lung tumors | Inhibited cell proliferation, migration, and invasion. | Zhao et al. [277] |
| miR-203 | LASP-1 | Suppressor | Lung tumors | LASP-1, regulated by miR-203, promoted tumor proliferation and aggressiveness in human NSCLC. | Zheng et al. [278] |
| miR-210 | Oncogene | Lung tumor Serum | MiR-210 expression levels might be a novel diagnostic and prognostic marker of NSCLC | Osugi et al. [279], Li et al. [280] | |
| miR-211-3p | Suppressor | Lung tumors | LncRNA SNHG15 promoted cell proliferation and migration by targeting miR-211-3p | Cui et al. [281] | |
| miR-212 | SOX4 | Suppressor | Lung tumors | Suppressed cell migration and invasion, and EMT in NSCLC cells | Tang et al. [282] |
| miR-215 | ZEB2 | Suppressor | Lung tumors | Inhibited cell migration and invasion. | Hou et al. [83] |
| miR-218 | Slug/ZEB2 | Suppressor | Lung tumors | Inhibited cell migration and invasion. | Shi et al. [59] |
| miR-221 | Oncogene | Lung tumors | Correlated with lymph node metastasis and disease progression. | Zhang et al. [283] | |
| miR-224 | Suppressor | Lung tumors | Inhibited cell proliferation, invasion, and migration, and promoted cell apoptosis. | Zhu et al. [284] | |
| miR-301a | Oncogene | Lung tumors | miR-301a overexpression was correlated with lymph node metastasis and poor prognosis. | Shi et al. [285] | |
| miR-302b-3p | GCNT3 | Suppressor | Lung tumors | Inhibited proliferation, migration, and invasion | Li et al. [72] |
| miR-328-3p | γ-H2AX | Suppressor | Lung tumors | Up-regulated miR-328-3p demonstrated a survival inhibition effect in A549 and restored NSCLC cell sensitivity to radiotherapy. | Ma et al. [286] |
| miR-335 | Bcl-w, SP1 | Suppressor | Lung tumors | Inhibited cell proliferation, migration, Increased apoptosis. | Wang et al. [287] |
| miR-338-3p | IRS2 | Suppressor | Lung tumors | Inhibited growth and invasion. | Zhang et al. [288] |
| miR-339-5p | BCL6 | Suppressor | Lung tumors, Peripheral blood | Inhibited cell migration and invasion | Li et al. [289], Li et al. [290] |
| miR-340 | CDK4 | Suppressor | Lung tumors | Suppressed cell proliferation. | Qin et al. [291] |
| miR-361-3p | SH2B1 | Suppressor | Lung tumors | Inhibited cell growth, proliferation, colony formation, invasion, and migration | Chen et al. [76] |
| miR-361-5p | Suppressor | Lung tumors | Lower miR-361-5p expression was found in NSCLC and associated lymph node metastasis. | Zhuang et al. [292] | |
| miR-365 | TTF-1 | Suppressor | Serum | High miR-365 serum level had less lymph node metastasis and longer OS. | Liu et al. [293], Sun et al. [294] |
| miR-378 | HMOX1 | Oncogene | Lung tumors | Modulated NSCLC progression and angiogenesis | Skrzypek et al. [295] |
| Mir-379 | IGF-1R | Suppressor | Lung tumors | Inhibited cell proliferation, migration, and invasion. | Zhou et al. [296] |
| miR-381 | LRH-1 | Suppressor | Lung tumors | Inhibited cell migration and invasion in vitro and in vivo. | Tian et al. [297] |
| miR-383 | Suppressor | Lung tumors | Reduced proliferation, invasion, and migration. | Shang et al. [298] | |
| miR-409-3p | c-MET | Suppressor | Lung adenocarcinoma tumors | Inhibited cell proliferation, induced apoptosis, and reduced invasion and migration by silencing of AKT signaling. | Wan et al. [299] |
| miR-411 | Oncogene | Serum | Elevated serum miR-411 expression was correlated with lymph node metastasis and poor prognosis. | Wang et al. [300] | |
| miR-421 | Oncogene | Lung tumors, Serum | Promoted cell proliferation, invasion, and migration. | Li et al. [301] | |
| miR-422a | 61 potential target genes § | Oncogene | Lymph nodes and plasma | High expression in NSCLC metastatic lymph nodes and validated by fresh blood of patients. | Wu et al. [302] |
| miR-433 | E2F3 | Suppressor | Lung tumors | Reduced cell proliferation and invasion | Liu et al. [303] |
| miR-448 | DCLK1 | Suppressor | Lung squamous cell carcinoma | Inhibited cell proliferation, colony formation, migration, and invasion | Shan et al. [304] |
| miR-449a | c-MET | Suppressor | Lung tumors | Inhibited cell migration and invasion | Luo et al. [305] |
| miR-451 | RAB14 | Suppressor | Lung tumors, | Inhibited cell proliferation and triggered apoptosis | Wang et al. [306], Wang et al. [307] |
| miR-451a | Oncogene | Plasma | Exosomal miR-451a showed a significant association with lymph node metastasis, vascular invasion, and stage | Kanaoka et al. [308] | |
| miR-452 | BMI1 | Suppressor | Lung tumors | Inhibited cell invasion, but not cell proliferation or apoptosis. | He et al. [309] |
| miR-452-5p | 10 hub genes@ | Suppressor | Lung tumors, TCGA | Low miR-452-5p expression level played an essential role in lung adenocarcinoma. | Gan et al. [310] |
| miR-454 | PTEN | Oncogene | Lung tumors | Promoted cell proliferation, invasion, and migration and inhibited apoptosis. | Zhu et al. [311] |
| miR-485-5p | IGF2BP2 | Suppressor | Lung tumors | Inhibited cell growth, invasion, and caused G0/G1 arrest | Huang et al. [312] |
| miR-486-5p | ARHGAP5, Pim-1 | Suppressor | Lung tumors, Sputum, plasma | Inhibited tumor progression and metastasis | Shen et al. [313], Wang et al. [314], Pang et al. [315] |
| miR-490-3p | PCBP1 | Oncogene | Lung tumors | Promoted cell proliferation, invasion, and migration. | Li et al. [177] |
| miR-491-5p | IGF2BP1 | Suppressor | Lung tumors | Reduced cell proliferation, colony formation, migration, and invasion | Gong et al. [316] |
| miR-494 | Oncogene | Lung tumor, Serum | High miR-494 level was correlated with lymph node metastasis and poor prognosis. | Zhang et al. [317], Wang et al. [318] | |
| miR-504 | LOXL2 | Suppressor | Lung tumors | Inhibited cell proliferation, cell invasion, and EMT process of NSCLC | Ye et al. [319] |
| miR-506-3p | COTL1 | Suppressor | Lung tumors | Reduced cell growth, migration, and invasion in vitro and in vivo. | Guo et al. [320] |
| miR-520b | Rad22A | Suppressor | Lung tumors | Inhibited cell proliferation, invasion, and metastasis abilities | Zhang et al. [321] |
| miR-548I | AKT1 | Suppressor | Lung tumors | Inhibited NSCLC cell migration and invasion. | Liu et al. [322] |
| miR-582-5p | MAP3K2 | Suppressor | Lung tumors | Suppressed the proliferation, migration, and invasion of NSCLC cells | Wang et al. [323] |
| miR-588 | GRN | Suppressor | Lung squamous cell carcinoma | Suppressed tumor cell migration and invasion. | Qian et al. [324] |
| miR-590 | OLFM4 | Oncogene | Lung adenocarcinoma | Promoted cell migration and invasion. | Liu et al. [94] |
| miR-592 | SOX9 | Suppressor | Lung tumors | Reduced cell proliferation, colony formation, migration, and invasion. | Li et al. [325] |
| miR-598 | ZEB2 | Suppressor | Lung tumors | Reduced NSCLC cell proliferation and invasion. | Tong et al. [92] |
| miR-638 | Suppressor | Serum | Serum miR-638 expression levels in NSCLC patients after chemotherapy were associated with disease prognosis. | Wang et al. [326] | |
| miR-650 | ING4 | Oncogene | Lung tumors | Inhibited caspase-3-dependent apoptosis | Huang et al. [327] |
| miR652-3p | Lgl1 | Oncogene | Lung tumors | Promoted cell proliferation, invasion, and migration. | Yang et al. [328] |
| miR-661 | RUNX3 | Oncogene | Lung tumors | Down-regulation of miR-661 suppressed NSCLC proliferation and invasion. | Wang et al. [329] |
| miR-675-5p | GPR55 | Suppressor | Lung tumors | Inhibited cell proliferation, colony formation, invasion, and migration, and attenuated the tumorigenicity in vivo. | He et al. [330] |
| miR-874 | MMP2, XIAP | Suppressor | Lung tumors | LncRNA XLOC_008466 functioned as an oncogene in NSCLC by regulating the miR-874-MMP2/XIAP axis | Yang et al. [331] |
| miR-935 | E2F7 | Suppressor | Lung tumors | Suppressed cell proliferation, migration, and invasion | Wang et al. [332] |
| miR-1204 | PITX1 | Oncogene | Lung tumors | Promoted cell proliferation and reduced cell cycle arrest. | Jiang et al. [333] |
| miR-1269 | TP53, Caspase-9 | Oncogene | Lung tumors, TCGA | Promoted cell survival and proliferation. | Bao et al. [334] |
| miR-1290 | IRF2 | Oncogene | Lung tumors Serum | Promoted cell growth | Jin et al. [335], Mo. Et al. [336] |
| miR-4317 | FGF9 CCND2 | Suppressor | Lung tumors | Inhibited proliferation, colony formation, migration, and invasion, and hampered cycles | He et al. [178] |
| miR-6754-5p | Suppressor | Lung tumors | LncRNA LINC00978 promoted cell proliferation and invasion in NSCLC by inhibiting miR-6754-5p. | Li et al. [337] |
| Phase of Drug Development | Year | miRNA | Target | Delivery System | Author/Reference |
|---|---|---|---|---|---|
| In vitro | 2010 | miR-145 | C-MYC | Lentivirus | Chen et al. [359] |
| In vitro, In vivo | 2012 | miR-145 | OCT4, SOX2 | Polyethylenimines | Chiou et al. [360] |
| In vitro, In vivo | 2010 | miR-126 | EGFL7 | Lipid | Sun et al. [358] |
| In vitro, In vivo | 2010 | Let-7 | KRAS | Lentivirus | Trang et al. [355] |
| In vitro, In vivo | 2010, 2011 | miR-34a | BCL-2 | Neutral lipid | Wiggins et al. [356], Trang et al. [272] |
| In vitro, In vivo | 2011, 2015 | Let-7, miR-34a | KRAS, TP53 | Lentivirus, Lipid | Trang et al. [272], Stahlhut et al. [357] |
| In vitro, In vivo | 2011 | miR-133b | MCL-1 | Cationic lipoplex | Wu et al. [361] |
| In vitro, In vivo | 2013 | miR-29b | MCL-1, CDK6, DNMT3 | Lipid | Wu et al. [362] |
| Phase I (NCT01829971) | Start: 2013 Termination: 2016 | miR-34a | BCL-2 | Lipid | |
| Phase I (NCT02369198) | 2017 | miR-16 | EGFR | Bacterial minicells | van Zandwijk et al. [366] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, S.-G.; Chang, T.-H.; Liu, Y.-N.; Shih, J.-Y. MicroRNA in Lung Cancer Metastasis. Cancers 2019, 11, 265. https://doi.org/10.3390/cancers11020265
Wu S-G, Chang T-H, Liu Y-N, Shih J-Y. MicroRNA in Lung Cancer Metastasis. Cancers. 2019; 11(2):265. https://doi.org/10.3390/cancers11020265
Chicago/Turabian StyleWu, Shang-Gin, Tzu-Hua Chang, Yi-Nan Liu, and Jin-Yuan Shih. 2019. "MicroRNA in Lung Cancer Metastasis" Cancers 11, no. 2: 265. https://doi.org/10.3390/cancers11020265
APA StyleWu, S.-G., Chang, T.-H., Liu, Y.-N., & Shih, J.-Y. (2019). MicroRNA in Lung Cancer Metastasis. Cancers, 11(2), 265. https://doi.org/10.3390/cancers11020265





