Deciphering the Mechanism of Action Involved in Enhanced Suicide Gene Colon Cancer Cell Killer Effect Mediated by Gef and Apoptin
Abstract
1. Introduction
2. Results
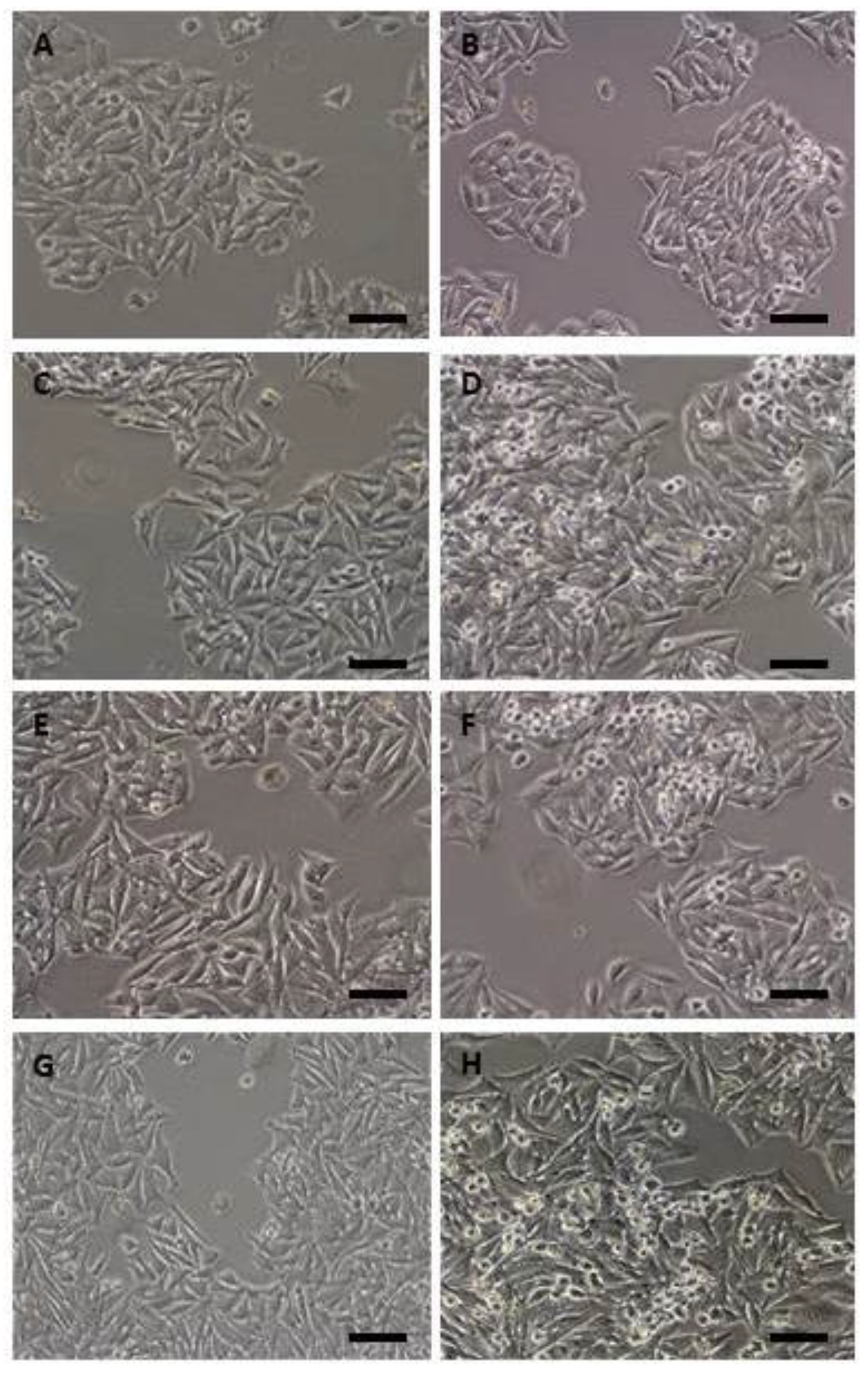
2.1. Morphological Findings
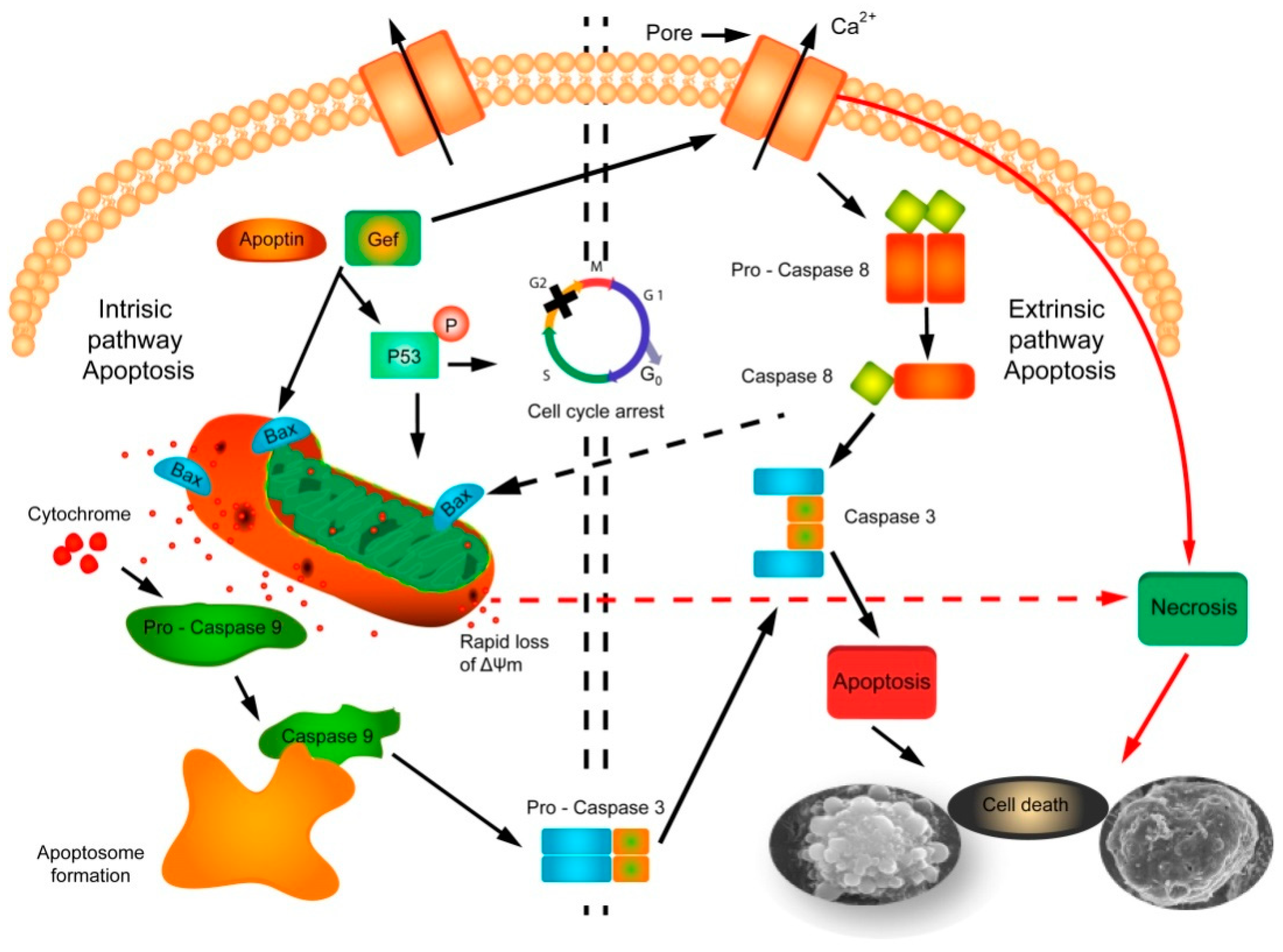
2.2. Mechanism of Action of Gef and/or Apoptin
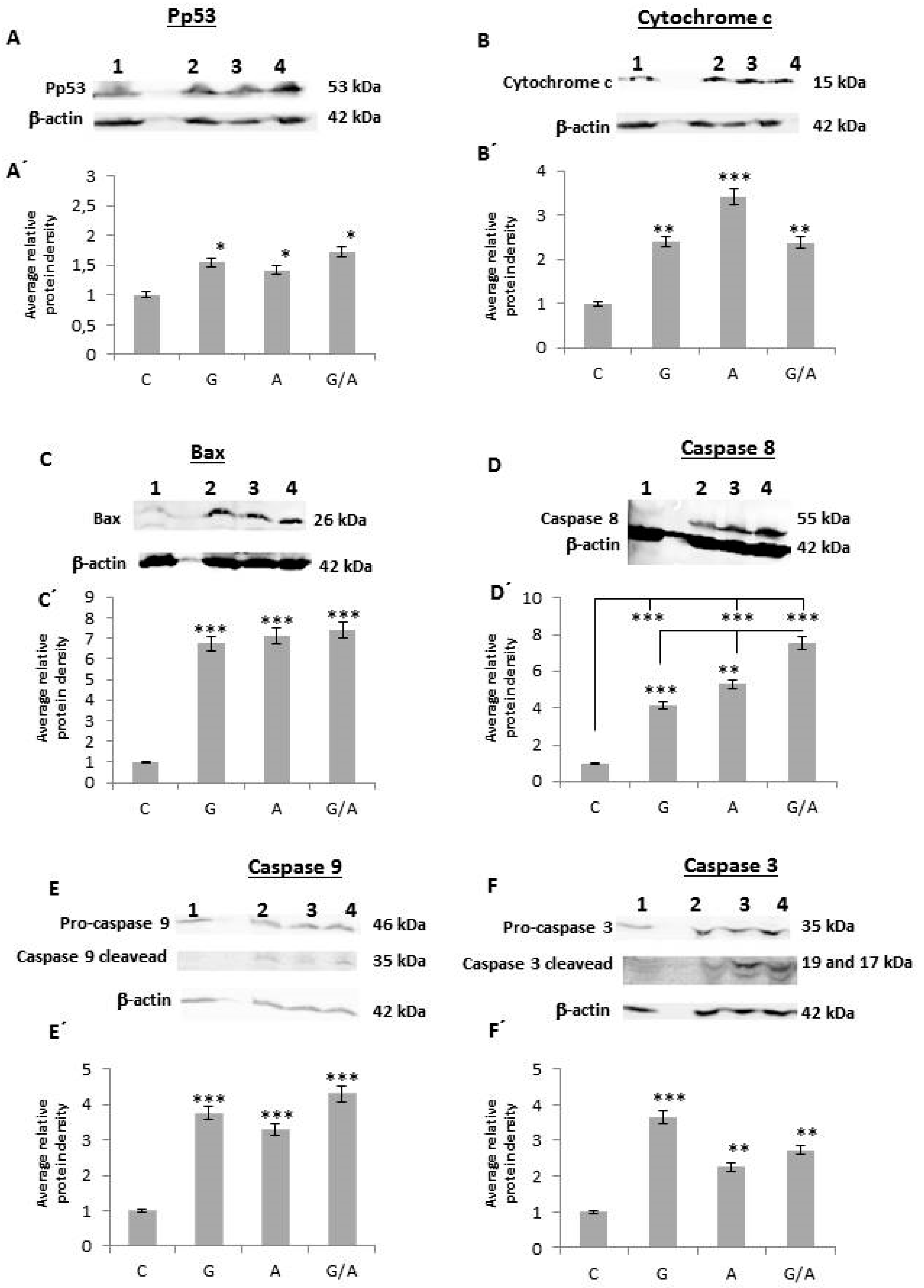
2.2.1. Pp53
2.2.2. Cytochrome c
2.2.3. Bax
2.2.4. Caspase 8
2.2.5. Caspase 9
2.2.6. Caspase 3
3. Discussion
4. Material and Methods
4.1. Cell Lines
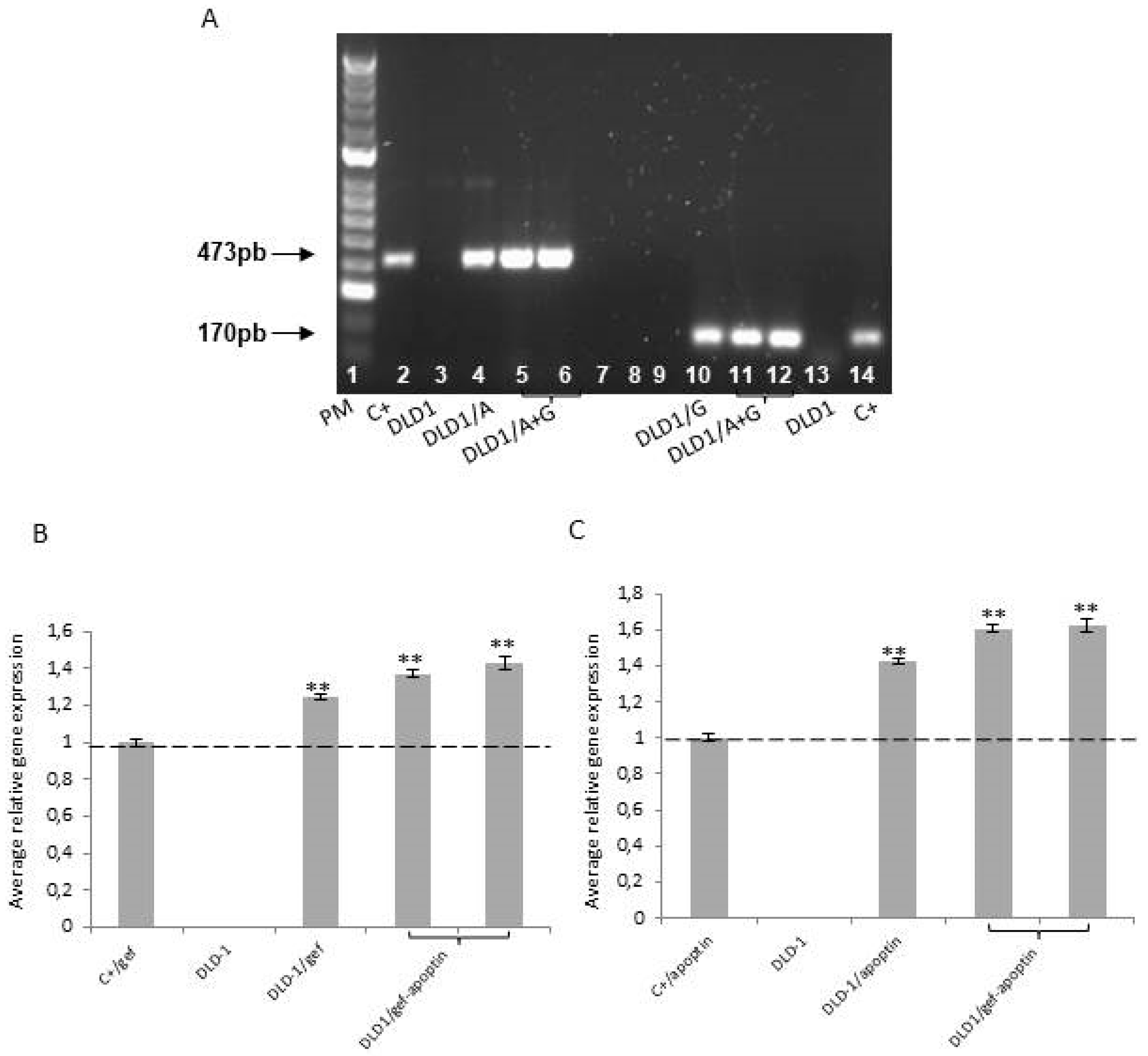
4.2. RNA Extraction Protocol and RT-PCR Analysis
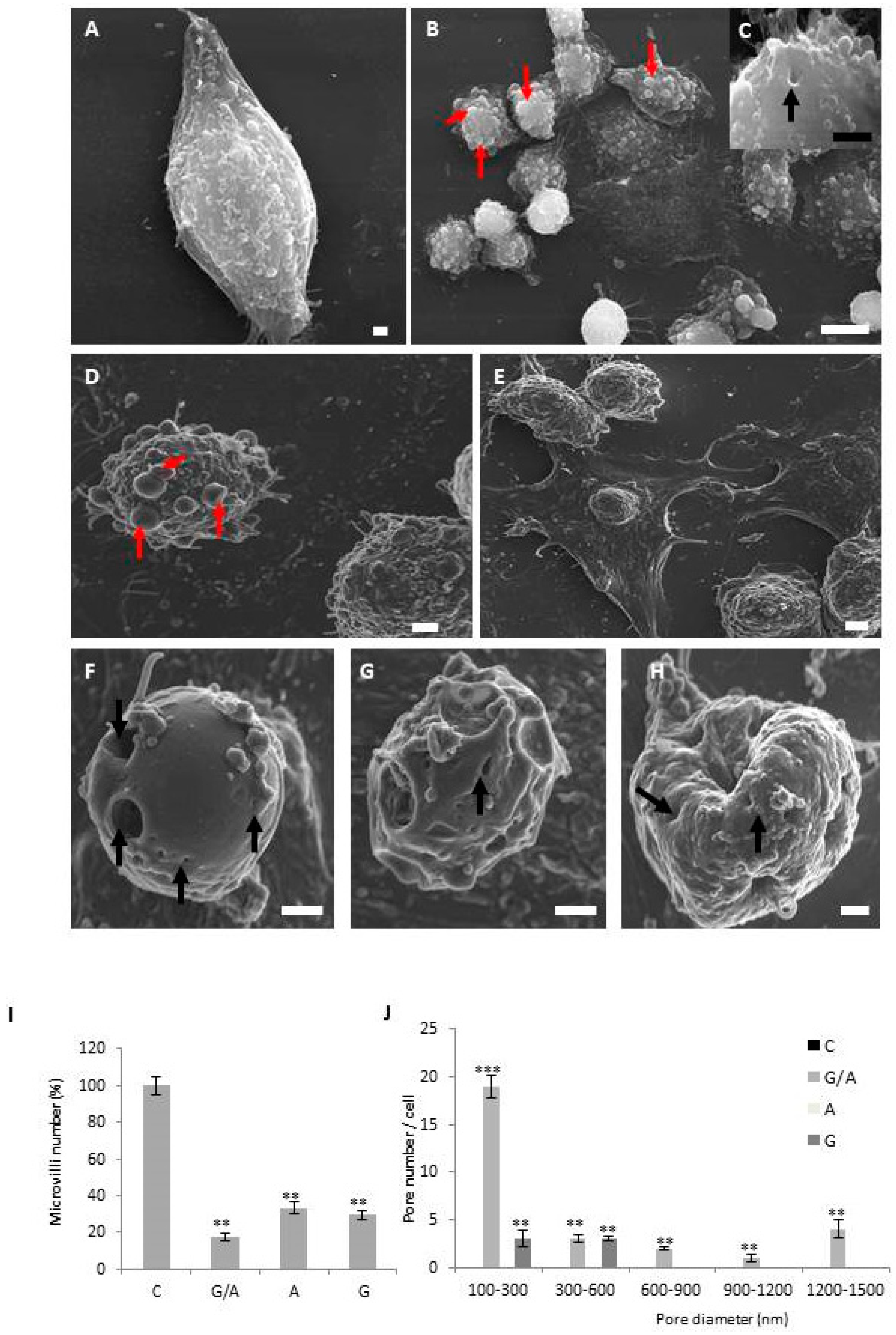
4.3. Scanning Electron Microscopy
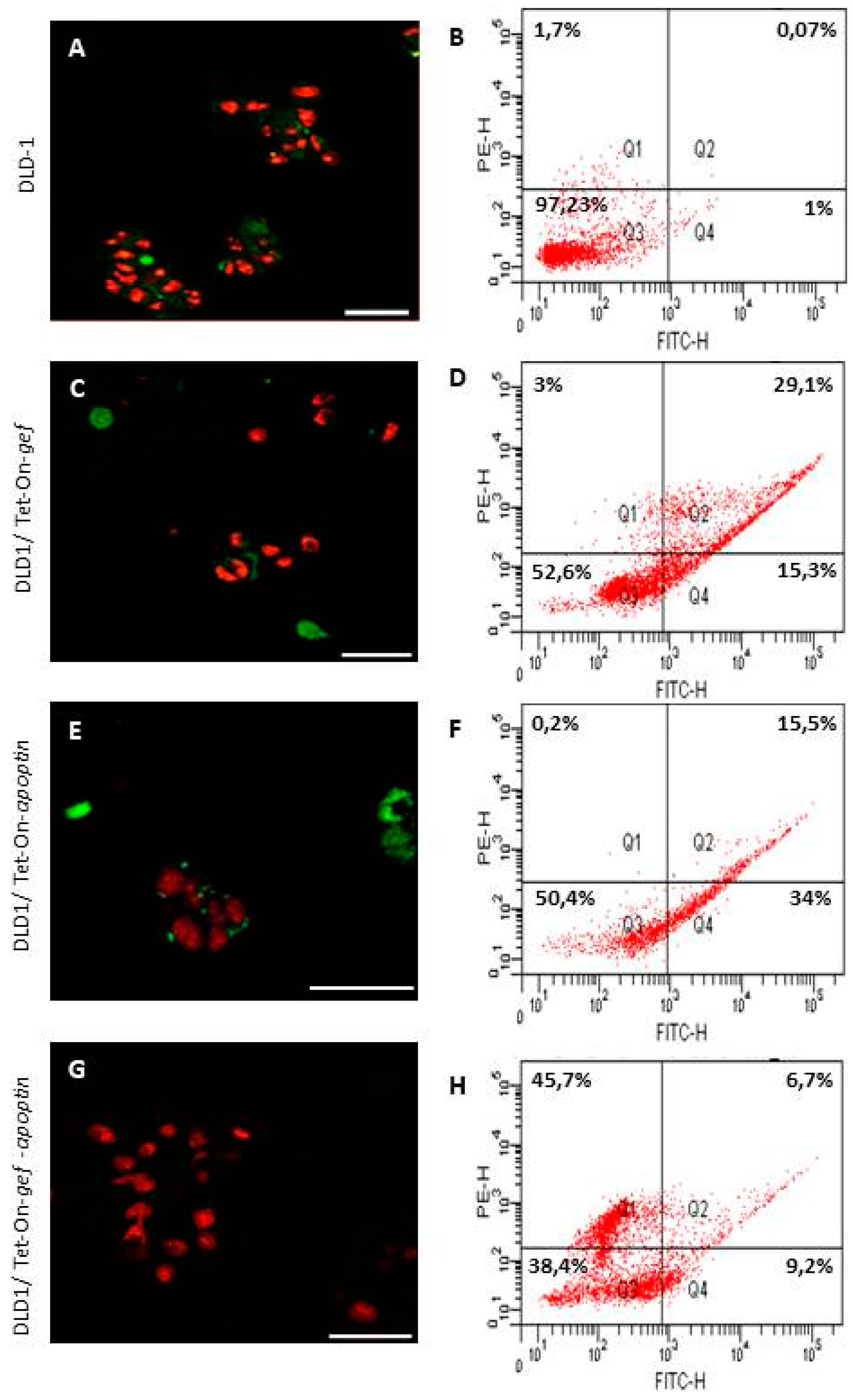
4.4. Confocal Microscopy and Flow Cytometry for Apoptosis Detection
4.5. Protein Analysis
4.6. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, D.; Atkin, W.; Lenz, H.J.; Lynch, H.T.; Minsky, B.; Nordlinger, B.; Starling, N. Colorectal cancer. Lancet 2010, 375, 1030–1047. [Google Scholar] [CrossRef]
- Hu, T.; Li, Z.; Gao, C.Y.; Cho, C.H. Mechanisms of drug resistance in colon cancer and its therapeutic strategies. World J. Gastroenterol. 2016, 22, 6876–6889. [Google Scholar] [CrossRef] [PubMed]
- Navarro, S.A.; Carrillo, E.; Grinan-Lison, C.; Martin, A.; Peran, M.; Marchal, J.A.; Boulaiz, H. Cancer suicide gene therapy: A patent review. Expert Opin. Ther. Pat. 2016, 26, 1095–1104. [Google Scholar] [CrossRef] [PubMed]
- Pahle, J.; Walther, W. Bacterial Toxins for Oncoleaking Suicidal Cancer Gene Therapy. Recent Results Cancer Res. 2016, 209, 95–110. [Google Scholar] [CrossRef] [PubMed]
- Shapira, A.; Benhar, I. Toxin-based therapeutic approaches. Toxins 2010, 2, 2519–2583. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.S.; Park, S.O.; Yoon, A.R.; Yoo, J.Y.; Kim, M.K.; Yun, C.O.; Kim, C.W. Suicide cancer gene therapy using pore-forming toxin, streptolysin O. Mol. Cancer Ther. 2006, 5, 1610–1619. [Google Scholar] [CrossRef] [PubMed]
- Fiorentini, G.; Banfi, R.; Dentico, P.; Moriconi, S.; Turrisi, G.; Pelagotti, F.; Rossi, S.; Montagnani, F. Clinical experience of treatment of metastatic melanoma and solid tumours adopting a derivative of diphtheria toxin: Cross-reacting material 197. In Vivo 2013, 27, 197–202. [Google Scholar] [PubMed]
- Yang, X.; Kessler, E.; Su, L.J.; Thorburn, A.; Frankel, A.E.; Li, Y.; La Rosa, F.G.; Shen, J.; Li, C.Y.; Varella-Garcia, M.; et al. Diphtheria toxin-epidermal growth factor fusion protein DAB389EGF for the treatment of bladder cancer. Clin. Cancer Res. 2013, 19, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Amit, D.; Hochberg, A. Development of targeted therapy for a broad spectrum of cancers (pancreatic cancer, ovarian cancer, glioblastoma and HCC) mediated by a double promoter plasmid expressing diphtheria toxin under the control of H19 and IGF2-P4 regulatory sequences. Int. J. Clin. Exp. Med. 2012, 5, 296–305. [Google Scholar] [PubMed]
- Martin, V.; Cortes, M.L.; de Felipe, P.; Farsetti, A.; Calcaterra, N.B.; Izquierdo, M. Cancer gene therapy by thyroid hormone-mediated expression of toxin genes. Cancer Res. 2000, 60, 3218–3224. [Google Scholar] [PubMed]
- Zhou, S.; Zhang, M.; Zhang, J.; Shen, H.; Tangsakar, E.; Wang, J. Mechanisms of Apoptin-induced cell death. Med. Oncol. 2012, 29, 2985–2991. [Google Scholar] [CrossRef] [PubMed]
- Boulaiz, H.; Alvarez, P.J.; Prados, J.; Marchal, J.; Melguizo, C.; Carrillo, E.; Peran, M.; Rodriguez, F.; Ramirez, A.; Ortiz, R.; et al. gef gene expression in MCF-7 breast cancer cells is associated with a better prognosis and induction of apoptosis by p53-mediated signaling pathway. Int. J. Mol. Sci. 2011, 12, 7445–7458. [Google Scholar] [CrossRef] [PubMed]
- Boulaiz, H.; Prados, J.; Melguizo, C.; Garcia, A.M.; Marchal, J.A.; Ramos, J.L.; Carrillo, E.; Velez, C.; Aranega, A. Inhibition of growth and induction of apoptosis in human breast cancer by transfection of gef gene. Br. J. Cancer 2003, 89, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Boulaiz, H.; Prados, J.; Marchal, J.A.; Garcia, A.M.; Alvarez, L.; Melguizo, C.; Carrillo, E.; Ramos, J.L.; Aranega, A. Transfection of MS-36 melanoma cells with gef gene inhibits proliferation and induces modulation of the cell cycle. Cancer Sci. 2003, 94, 564–568. [Google Scholar] [CrossRef] [PubMed]
- Boulaiz, H.; Aranega, A.; Caceres, B.; Alvarez, P.; Serrano-Rodriguez, F.; Carrillo, E.; Melguizo, C.; Prados, J. A novel double-enhanced suicide gene therapy in a colon cancer cell line mediated by gef and apoptin. BioDrugs 2014, 28, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Ortiz, R.; Prados, J.; Melguizo, C.; Rama, A.R.; Alvarez, P.J.; Rodriguez-Serrano, F.; Caba, O.; Boulaiz, H.; Aranega, A. Gef gene therapy enhances the therapeutic efficacy of cytotoxics in colon cancer cells. Biomed. Pharmacother. 2012, 66, 563–567. [Google Scholar] [CrossRef] [PubMed]
- Wajant, H. The Fas signaling pathway: More than a paradigm. Science 2002, 296, 1635–1636. [Google Scholar] [CrossRef] [PubMed]
- Wickman, G.; Julian, L.; Olson, M.F. How apoptotic cells aid in the removal of their own cold dead bodies. Cell Death Differ. 2012, 19, 735–742. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.J.; Chao, Y.; Hung, Y.M.; Lin, W.C.; Yang, D.M.; Shih, Y.L.; Ch’ang, L.Y.; Whang-Peng, J.; Yang, W.K. S- and G2-phase cell cycle arrests and apoptosis induced by ganciclovir in murine melanoma cells transduced with herpes simplex virus thymidine kinase. Exp. Cell Res. 1998, 241, 66–75. [Google Scholar] [CrossRef] [PubMed]
- Kerr, J.F.; Wyllie, A.H.; Currie, A.R. Apoptosis: A basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 1972, 26, 239–257. [Google Scholar] [CrossRef] [PubMed]
- Inbal, B.; Bialik, S.; Sabanay, I.; Shani, G.; Kimchi, A. DAP kinase and DRP-1 mediate membrane blebbing and the formation of autophagic vesicles during programmed cell death. J. Cell Biol. 2002, 157, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Coleman, M.L.; Sahai, E.A.; Yeo, M.; Bosch, M.; Dewar, A.; Olson, M.F. Membrane blebbing during apoptosis results from caspase-mediated activation of ROCK I. Nature Cell Biol. 2001, 3, 339–345. [Google Scholar] [CrossRef] [PubMed]
- Khajah, M.A.; Luqmani, Y.A. Involvement of Membrane Blebbing in Immunological Disorders and Cancer. Med. Prin. Pract. 2016, 25 (Suppl. 2), 18–27. [Google Scholar] [CrossRef]
- Boulaiz, H.; Prados, J.; Marchal, J.A.; Melguizo, C.; Concha, A.; Carrillo, E.; Velez, C.; Martinez, A.; Aranega, A. Modulation of Ki-67 expression and morphological changes induced by gef gene in MCF-7 human breast cancer cells. Cell. Mol. Biol. 2005, 51, 87–92. [Google Scholar] [PubMed]
- Mohiyuddin, S.; Naqvi, S.; Packirisamy, G. Enhanced antineoplastic/therapeutic efficacy using 5-fluorouracil-loaded calcium phosphate nanoparticles. Beilstein J. Nanotechnol. 2018, 9, 2499–2515. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, K.; Bech, F.W.; Jorgensen, S.T.; Lobner-Olesen, A.; Rasmussen, P.B.; Atlung, T.; Boe, L.; Karlstrom, O.; Molin, S.; von Meyenburg, K. Mechanism of postsegregational killing by the hok gene product of the parB system of plasmid R1 and its homology with the relF gene product of the E. coli relB operon. EMBO J. 1986, 5, 2023–2029. [Google Scholar] [CrossRef] [PubMed]
- Ono, K.; Akimoto, S.; Ohnishi, Y. Nucleotide sequence of the pnd gene in plasmid R483 and role of the pnd gene product in plasmolysis. Microbiol. Immunol. 1987, 31, 1071–1083. [Google Scholar] [CrossRef] [PubMed]
- Burattini, S.; Falcieri, E. Analysis of cell death by electron microscopy. Methods Mol. Biol. 2013, 1004, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Kroemer, G.; Galluzzi, L.; Vandenabeele, P.; Abrams, J.; Alnemri, E.S.; Baehrecke, E.H.; Blagosklonny, M.V.; El-Deiry, W.S.; Golstein, P.; Green, D.R.; et al. Classification of cell death: Recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ. 2009, 16, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Proskuryakov, S.Y.; Konoplyannikov, A.G.; Gabai, V.L. Necrosis: A specific form of programmed cell death? Exp. Cell Res. 2003, 283, 1–16. [Google Scholar] [CrossRef]
- Golstein, P.; Kroemer, G. Cell death by necrosis: Towards a molecular definition. Trends Biochem. Sci. 2007, 32, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Backes, C.S.; Friedmann, K.S.; Mang, S.; Knorck, A.; Hoth, M.; Kummerow, C. Natural killer cells induce distinct modes of cancer cell death: Discrimination, quantification, and modulation of apoptosis, necrosis, and mixed forms. J. Biol. Chem. 2018, 293, 16348–16363. [Google Scholar] [CrossRef] [PubMed]
- Bischofberger, M.; Iacovache, I.; van der Goot, F.G. Pathogenic pore-forming proteins: Function and host response. Cell Host Microbe 2012, 12, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Peng, D.; Cheng, C.; Zhou, W.; Ju, S.; Wan, D.; Yu, Z.; Shi, J.; Deng, Y.; Wang, F.; et al. Bacillus thuringiensis Crystal Protein Cry6Aa Triggers Caenorhabditis elegans Necrosis Pathway Mediated by Aspartic Protease (ASP-1). PLoS Pathog. 2016, 12, e1005389. [Google Scholar] [CrossRef] [PubMed]
- Seike, S.; Miyamoto, K.; Kobayashi, K.; Takehara, M.; Nagahama, M. Clostridium perfringens Delta-Toxin Induces Rapid Cell Necrosis. PLoS ONE 2016, 11, e0147957. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Lee, Y.L.; Lee, Y.J.; Park, S.Y.; Kim, I.S.; Choi, T.H.; Ha, J.H.; Ahn, B.C.; Lee, J. Enhanced antitumor effects by combination gene therapy using MDR1 gene shRNA and HSV1-tk in a xenograft mouse model. Cancer Lett. 2010, 291, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Prewett, M.C.; Hooper, A.T.; Bassi, R.; Ellis, L.M.; Waksal, H.W.; Hicklin, D.J. Enhanced antitumor activity of anti-epidermal growth factor receptor monoclonal antibody IMC-C225 in combination with irinotecan (CPT-11) against human colorectal tumor xenografts. Clin. Cancer Res. 2002, 8, 994–1003. [Google Scholar] [PubMed]
- Salehi, M.; Sharifi, M. Induction of apoptosis and necrosis in human acute erythroleukemia cells by inhibition of long non-coding RNA PVT1. Mol. Biol. Res. Commun. 2018, 7, 89–96. [Google Scholar] [CrossRef] [PubMed]
- McIlwain, D.R.; Berger, T.; Mak, T.W. Caspase functions in cell death and disease. Cold Spring Harb. Perspect. Biol. 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Pfeffer, C.M.; Singh, A.T.K. Apoptosis: A Target for Anticancer Therapy. Int. J. Mol. Sci. 2018, 19, 448. [Google Scholar] [CrossRef] [PubMed]
- Abraha, A.M.; Ketema, E.B. Apoptotic pathways as a therapeutic target for colorectal cancer treatment. World J. Gastro. Oncol. 2016, 8, 583–591. [Google Scholar] [CrossRef] [PubMed]
- Prados, J.; Melguizo, C.; Ortiz, R.; Boulaiz, H.; Carrillo, E.; Segura, A.; Rodriguez-Herva, J.J.; Ramos, J.L.; Aranega, A. Regression of established subcutaneous B16-F10 murine melanoma tumors after gef gene therapy associated with the mitochondrial apoptotic pathway. Exp. Dermatol. 2010, 19, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Solt, L.A.; Conkright, J.; Wang, Y.; Istrate, M.A.; Busby, S.A.; Garcia-Ordonez, R.D.; Nuhant, P.; Burris, T.; Mercer, B.A.; et al. Campaign to identify novel modulators of the Retinoic acid receptor-related Orphan Receptors (ROR). In Probe Reports from the NIH Molecular Libraries Program; National Center for Biotechnology Information: Bethesda, MD, USA, 2010. [Google Scholar]
- Lavrik, I.N.; Golks, A.; Krammer, P.H. Caspases: Pharmacological manipulation of cell death. J. Clin. Investig. 2005, 115, 2665–2672. [Google Scholar] [CrossRef] [PubMed]
- Porter, A.G.; Janicke, R.U. Emerging roles of caspase-3 in apoptosis. Cell Death Differ. 1999, 6, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.Y.; Choi, Y.; Jeon, Y.K.; Aung, K.C.; Kim, C.W. Over-expression of adenine nucleotide translocase 1 (ANT1) induces apoptosis and tumor regression in vivo. BMC Cancer 2008, 8, 160. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Candas, M.; Griko, N.B.; Taussig, R.; Bulla, L.A., Jr. A mechanism of cell death involving an adenylyl cyclase/PKA signaling pathway is induced by the Cry1Ab toxin of Bacillus thuringiensis. Proc. Natl. Acad. Sci. USA 2006, 103, 9897–9902. [Google Scholar] [CrossRef] [PubMed]
- Prados, J.; Melguizo, C.; Rama, A.; Ortiz, R.; Boulaiz, H.; Rodriguez-Serrano, F.; Caba, O.; Rodriguez-Herva, J.J.; Ramos, J.L.; Aranega, A. Combined therapy using suicide gef gene and paclitaxel enhances growth inhibition of multicellular tumour spheroids of A-549 human lung cancer cells. Int. J. Oncol. 2008, 33, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Slee, E.A.; Adrain, C.; Martin, S.J. Executioner caspase-3,-6, and-7 perform distinct, non-redundant roles during the demolition phase of apoptosis. J. Biol. Chem. 2001, 276, 7320–7326. [Google Scholar] [CrossRef] [PubMed]
- Danial, N.N.; Korsmeyer, S.J. Cell death: Critical control points. Cell 2004, 116, 205–219. [Google Scholar] [CrossRef]
- Sakahira, H.; Enari, M.; Nagata, S. Cleavage of CAD inhibitor in CAD activation and DNA degradation during apoptosis. Nature 1998, 391, 96–99. [Google Scholar] [CrossRef] [PubMed]
- Elmore, S. Apoptosis: A review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef] [PubMed]
- Boland, K.; Flanagan, L.; Prehn, J.H. Paracrine control of tissue regeneration and cell proliferation by Caspase-3. Cell Death Dis. 2013, 4, e725. [Google Scholar] [CrossRef] [PubMed]
- Park, C.; Hong, S.H.; Kim, G.Y.; Choi, Y.H. So-Cheong-Ryong-Tang induces apoptosis through activation of the intrinsic and extrinsic apoptosis pathways, and inhibition of the PI3K/Akt signaling pathway in non-small-cell lung cancer A549 cells. BMC Complement. Altern. Med. 2015, 15, 113. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.T. Secondary necrosis: The natural outcome of the complete apoptotic program. FEBS Lett. 2010, 584, 4491–4499. [Google Scholar] [CrossRef] [PubMed]
- Sachet, M.; Liang, Y.Y.; Oehler, R. The immune response to secondary necrotic cells. Apoptosis 2017, 22, 1189–1204. [Google Scholar] [CrossRef] [PubMed]
- Rogers, C.; Fernandes-Alnemri, T.; Mayes, L.; Alnemri, D.; Cingolani, G.; Alnemri, E.S. Cleavage of DFNA5 by caspase-3 during apoptosis mediates progression to secondary necrotic/pyroptotic cell death. Nat. Commun. 2017, 8, 14128. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, A.; Conejo-García, A.; Griñán-Lisón, C.; López-Cara, L.C.; Jiménez, G.; Campos, J.M.; Marchal, J.A.; Boulaiz, H. Enhancement of Tumor Cell Death by Combining gef Gene Mediated Therapy and New 1,4-Benzoxazepin-2,6-Dichloropurine Derivatives in Breast Cancer Cells. Front. Pharmacol. 2018, 9, 798. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.C.; Wu, J.L.; Her, G.M.; Hong, J.R. Aquatic birnavirus induces necrotic cell death via the mitochondria-mediated caspase pathway. Fish Shellfish Immunol. 2010, 28, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Hitomi, J.; Christofferson, D.E.; Ng, A.; Yao, J.; Degterev, A.; Xavier, R.J.; Yuan, J. Identification of a molecular signaling network that regulates a cellular necrotic cell death pathway. Cell 2008, 135, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Stoll, G.; Ma, Y.; Yang, H.; Kepp, O.; Zitvogel, L.; Kroemer, G. Pro-necrotic molecules impact local immunosurveillance in human breast cancer. Oncoimmunology 2017, 6, e1299302. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Kroemer, G. Secondary necrosis: Accidental no more. Trends Cancer 2017, 3, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Wolmark, N.; Fisher, B.; Wieand, H.S. The prognostic value of the modifications of the Dukes’ C class of colorectal cancer. An analysis of the NSABP clinical trials. Ann. Surg. 1986, 203, 115–122. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cáceres, B.; Ramirez, A.; Carrillo, E.; Jimenez, G.; Griñán-Lisón, C.; López-Ruiz, E.; Jiménez-Martínez, Y.; Marchal, J.A.; Boulaiz, H. Deciphering the Mechanism of Action Involved in Enhanced Suicide Gene Colon Cancer Cell Killer Effect Mediated by Gef and Apoptin. Cancers 2019, 11, 264. https://doi.org/10.3390/cancers11020264
Cáceres B, Ramirez A, Carrillo E, Jimenez G, Griñán-Lisón C, López-Ruiz E, Jiménez-Martínez Y, Marchal JA, Boulaiz H. Deciphering the Mechanism of Action Involved in Enhanced Suicide Gene Colon Cancer Cell Killer Effect Mediated by Gef and Apoptin. Cancers. 2019; 11(2):264. https://doi.org/10.3390/cancers11020264
Chicago/Turabian StyleCáceres, Blanca, Alberto Ramirez, Esmeralda Carrillo, Gema Jimenez, Carmen Griñán-Lisón, Elena López-Ruiz, Yaiza Jiménez-Martínez, Juan A. Marchal, and Houria Boulaiz. 2019. "Deciphering the Mechanism of Action Involved in Enhanced Suicide Gene Colon Cancer Cell Killer Effect Mediated by Gef and Apoptin" Cancers 11, no. 2: 264. https://doi.org/10.3390/cancers11020264
APA StyleCáceres, B., Ramirez, A., Carrillo, E., Jimenez, G., Griñán-Lisón, C., López-Ruiz, E., Jiménez-Martínez, Y., Marchal, J. A., & Boulaiz, H. (2019). Deciphering the Mechanism of Action Involved in Enhanced Suicide Gene Colon Cancer Cell Killer Effect Mediated by Gef and Apoptin. Cancers, 11(2), 264. https://doi.org/10.3390/cancers11020264








