The New Genetic Landscape of Cushing’s Disease: Deubiquitinases in the Spotlight
Abstract
1. Introduction
2. Results and Discussions
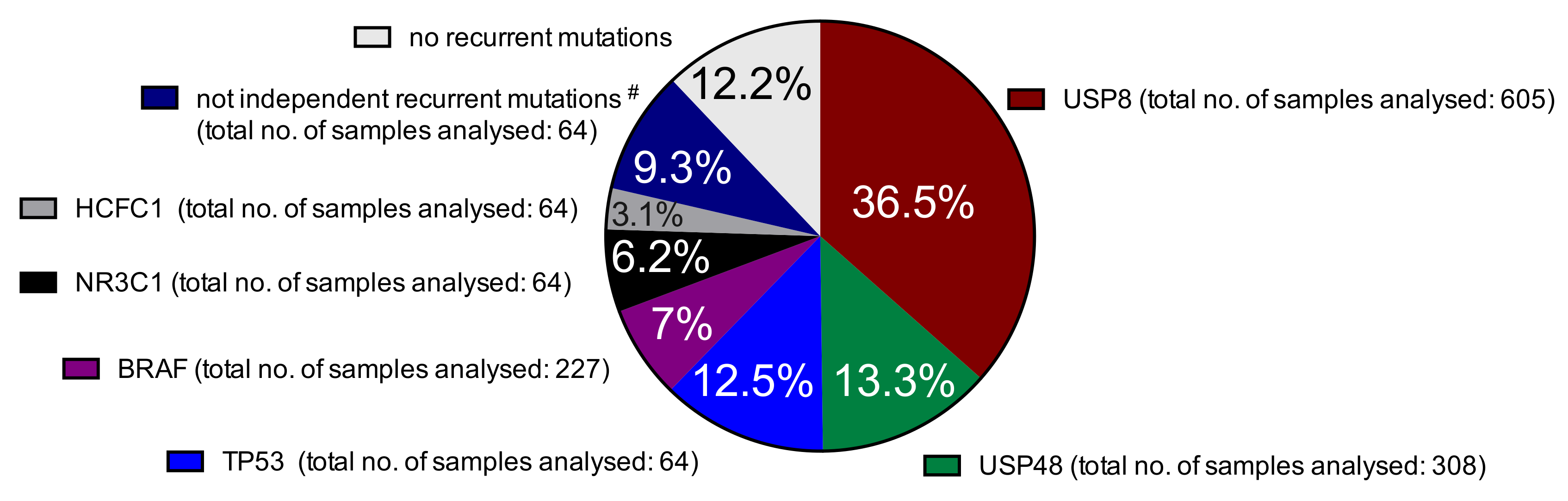
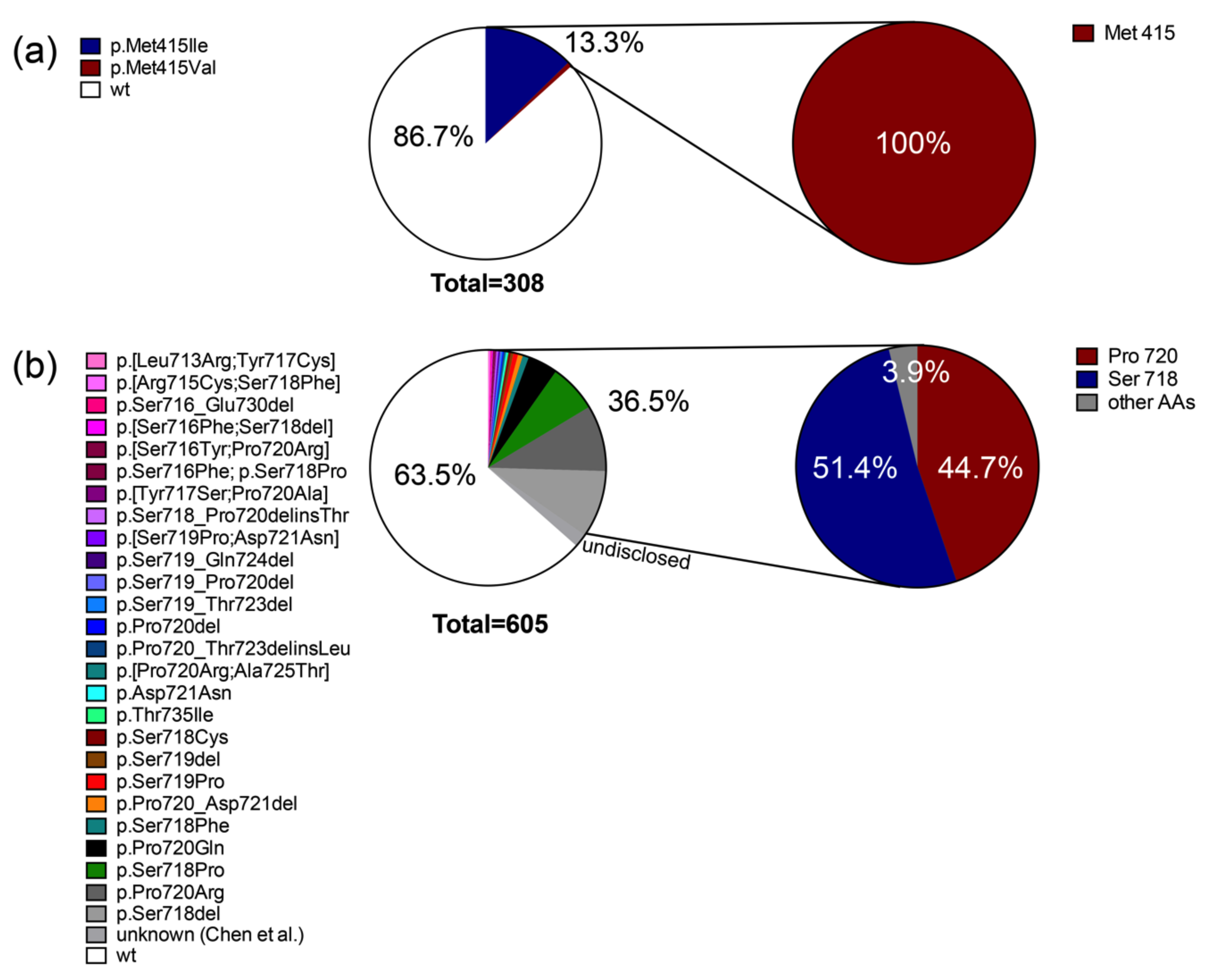
2.1. Somatic Mutations in Deubiquitinases: The Most Frequent Recurrent Mutations in Cushing’s Disease
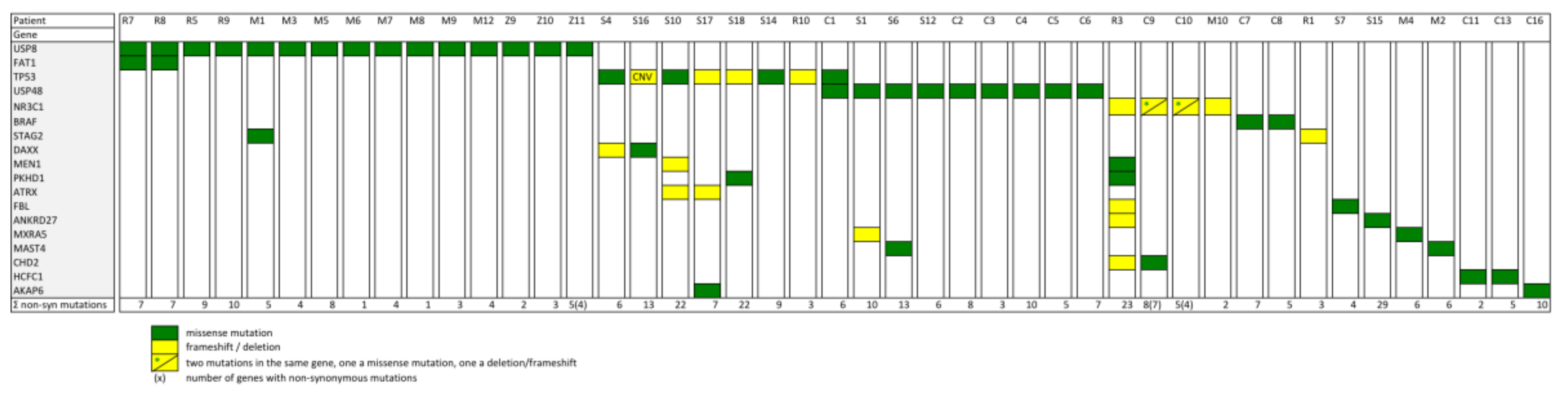
2.2. Prevalence of Different Other Recurrent Mutations in Cushing’s Disease
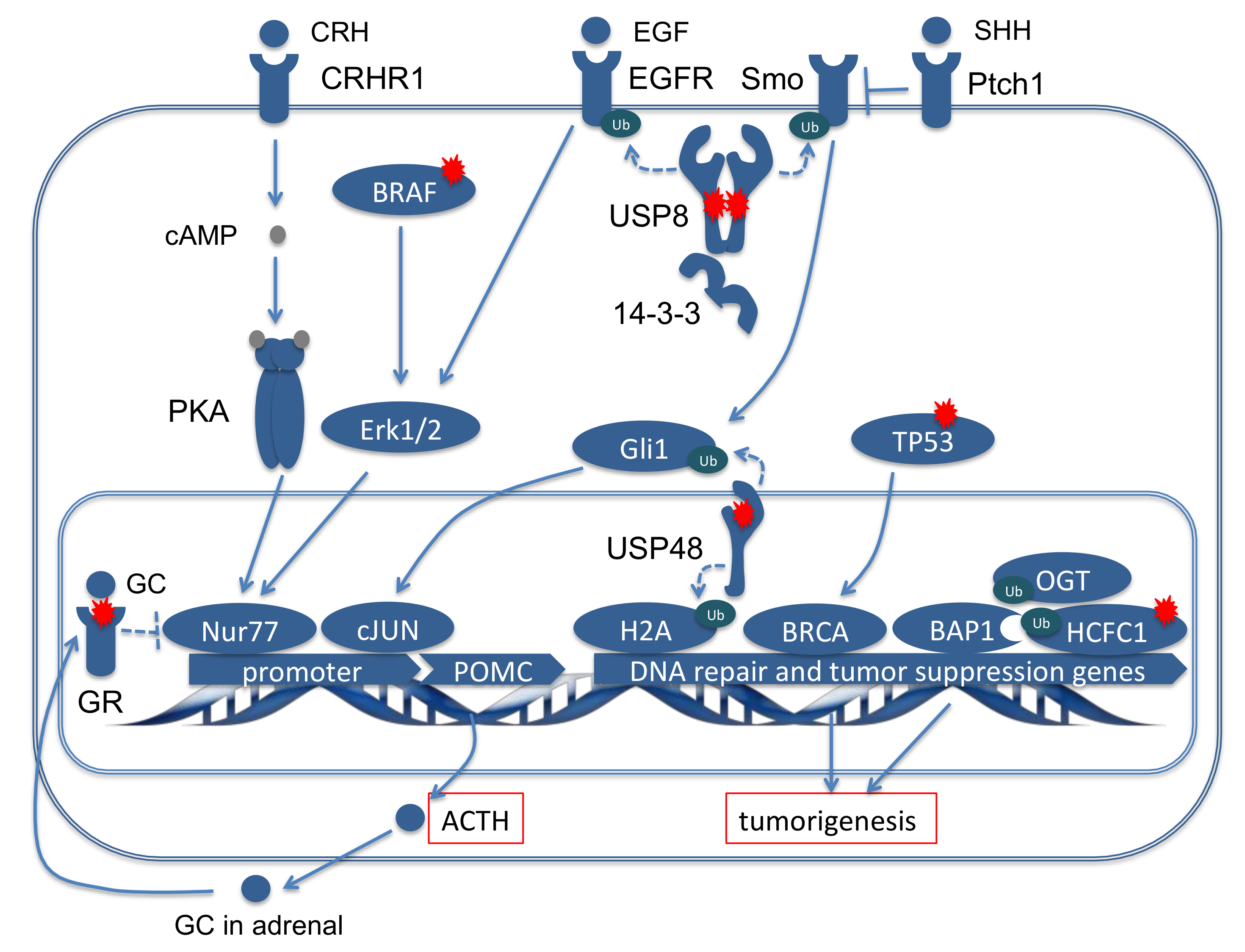
2.3. Pathogenetic Relevance of the Most Frequent Mutations in Cushing’s Disease
2.3.1. Deubiquitinases, the Key Driver Genes in CD
2.3.2. Classical Oncogenes and Tumor Suppressor Genes
2.3.3. Alterations in the Glucocorticoid Pathway
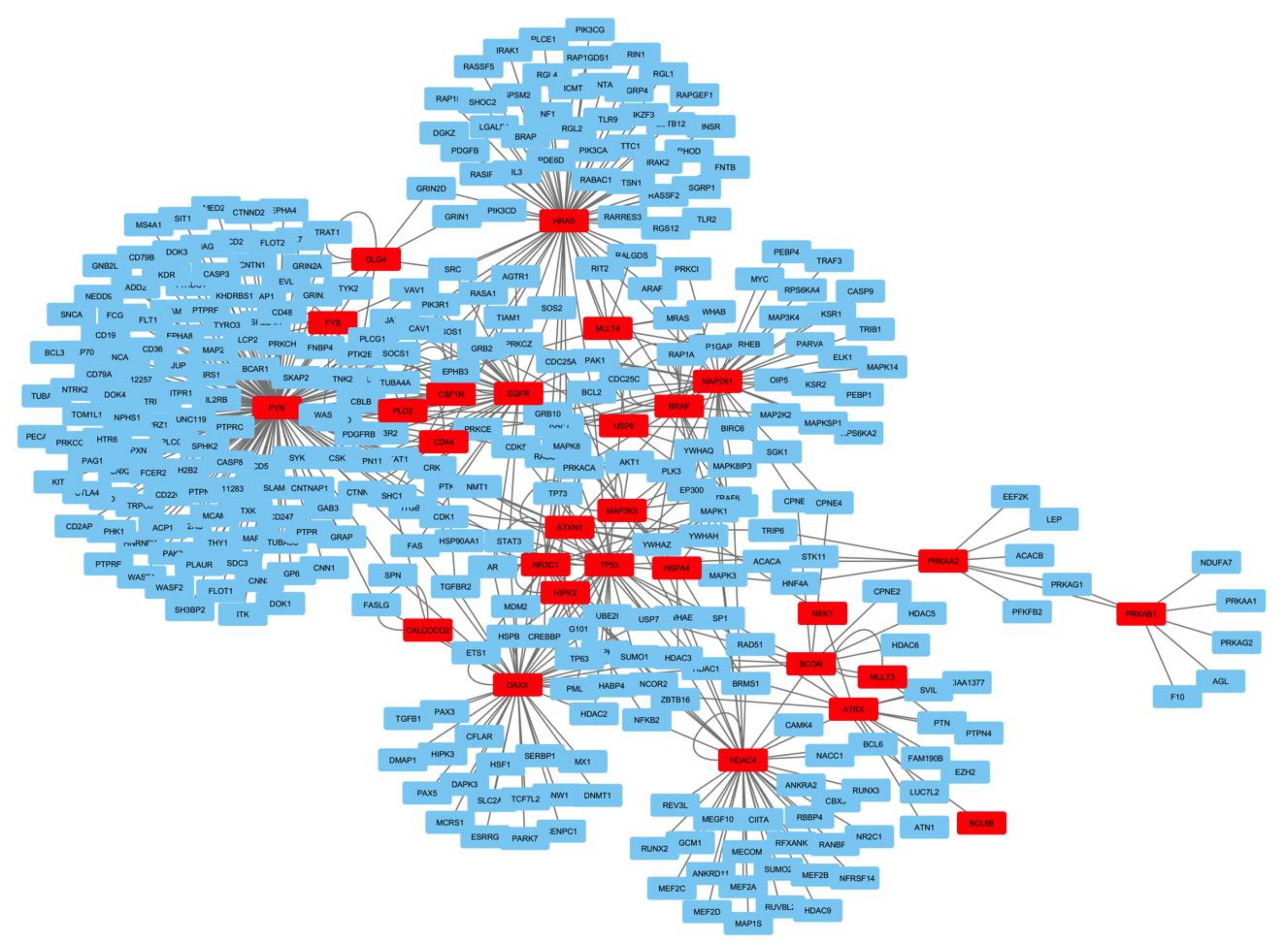
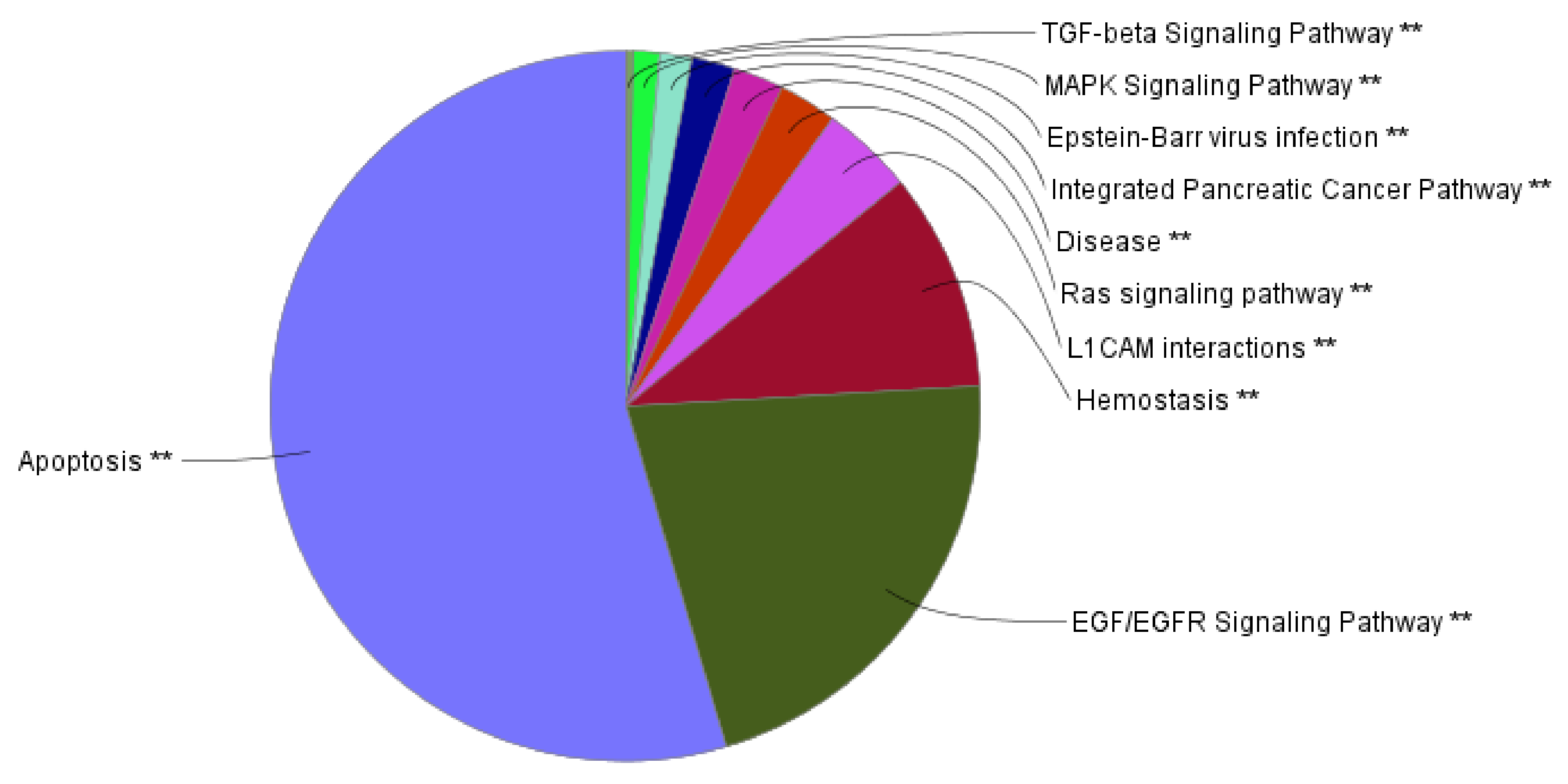
2.4. Functional Network Analysis of the Genes Mutated in ACTH Producing Pituitary Adenomas
3. Materials and Methods
3.1. Patient Cohorts
3.2. Bioinformatics Analyses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lambert, J.K.; Goldberg, L.; Fayngold, S.; Kostadinov, J.; Post, K.D.; Geer, E.B. Predictors of mortality and long-term outcomes in treated Cushing’s disease: A study of 346 patients. J. Clin. Endocrinol. Metab. 2013, 98, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Lacroix, A.; Feelders, R.A.; Stratakis, C.A.; Nieman, L.K. Cushing’s syndrome. Lancet 2015, 386, 913–927. [Google Scholar] [CrossRef]
- Cushing, H., III. Partial Hypophysectomy for Acromegaly: With Remarks on the Function of the Hypophysis. Ann. Surg. 1909, 50, 1002–1017. [Google Scholar] [CrossRef] [PubMed]
- Biller, B.M.; Grossman, A.B.; Stewart, P.M.; Melmed, S.; Bertagna, X.; Bertherat, J.; Buchfelder, M.; Colao, A.; Hermus, A.R.; Hofland, L.J.; et al. Treatment of adrenocorticotropin-dependent Cushing’s syndrome: A consensus statement. J. Clin. Endocrinol. Metab. 2008, 93, 2454–2462. [Google Scholar] [CrossRef] [PubMed]
- Petersenn, S.; Beckers, A.; Ferone, D.; van der Lely, A.; Bollerslev, J.; Boscaro, M.; Brue, T.; Bruzzi, P.; Casanueva, F.F.; Chanson, P.; et al. Therapy of endocrine disease: Outcomes in patients with Cushing’s disease undergoing transsphenoidal surgery: Systematic review assessing criteria used to define remission and recurrence. Eur. J. Endocrinol. 2015, 172, R227–R239. [Google Scholar] [CrossRef]
- Feelders, R.A.; Hofland, L.J. Medical treatment of Cushing’s disease. J. Clin. Endocrinol. Metab. 2013, 98, 425–438. [Google Scholar] [CrossRef]
- Hamrahian, A.H.; Yuen, K.C.; Hoffman, A.R.; Neuroendocrine, A.; Pituitary Scientific, C. AACE/ACE Disease State Clinical Review: Medical Management of Cushing Disease. Endocr. Pract. 2014, 20, 746–757. [Google Scholar] [CrossRef]
- Stratakis, C.A.; Tichomirowa, M.A.; Boikos, S.; Azevedo, M.F.; Lodish, M.; Martari, M.; Verma, S.; Daly, A.F.; Raygada, M.; Keil, M.F.; et al. The role of germline AIP, MEN1, PRKAR1A, CDKN1B and CDKN2C mutations in causing pituitary adenomas in a large cohort of children, adolescents, and patients with genetic syndromes. Clin. Genet. 2010, 78, 457–463. [Google Scholar] [CrossRef]
- Dinesen, P.T.; Dal, J.; Gabrovska, P.; Gaustadnes, M.; Gravholt, C.H.; Stals, K.; Denes, J.; Asa, S.L.; Korbonits, M.; Jorgensen, J.O. An unusual case of an ACTH-secreting macroadenoma with a germline variant in the aryl hydrocarbon receptor-interacting protein (AIP) gene. Endocrinol. Diabetes Metab. Case Rep. 2015, 2015, 140105. [Google Scholar] [CrossRef]
- De Kock, L.; Sabbaghian, N.; Plourde, F.; Srivastava, A.; Weber, E.; Bouron-Dal Soglio, D.; Hamel, N.; Choi, J.H.; Park, S.H.; Deal, C.L.; et al. Pituitary blastoma: A pathognomonic feature of germ-line DICER1 mutations. Acta Neuropathol 2014, 128, 111–122. [Google Scholar] [CrossRef]
- Sahakitrungruang, T.; Srichomthong, C.; Pornkunwilai, S.; Amornfa, J.; Shuangshoti, S.; Kulawonganunchai, S.; Suphapeetiporn, K.; Shotelersuk, V. Germline and somatic DICER1 mutations in a pituitary blastoma causing infantile-onset Cushing’s disease. J. Clin. Endocrinol. Metab. 2014, 99, E1487–E1492. [Google Scholar] [CrossRef] [PubMed]
- Wood, L.D.; Parsons, D.W.; Jones, S.; Lin, J.; Sjoblom, T.; Leary, R.J.; Shen, D.; Boca, S.M.; Barber, T.; Ptak, J.; et al. The genomic landscapes of human breast and colorectal cancers. Science 2007, 318, 1108–1113. [Google Scholar] [CrossRef] [PubMed]
- Barrett, I.P. Cancer genome analysis informatics. Methods Mol. Biol. 2010, 628, 75–102. [Google Scholar] [CrossRef] [PubMed]
- Amado, R.G.; Wolf, M.; Peeters, M.; Van Cutsem, E.; Siena, S.; Freeman, D.J.; Juan, T.; Sikorski, R.; Suggs, S.; Radinsky, R.; et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J. Clin. Oncol. 2008, 26, 1626–1634. [Google Scholar] [CrossRef]
- Reincke, M.; Sbiera, S.; Hayakawa, A.; Theodoropoulou, M.; Osswald, A.; Beuschlein, F.; Meitinger, T.; Mizuno-Yamasaki, E.; Kawaguchi, K.; Saeki, Y.; et al. Mutations in the deubiquitinase gene USP8 cause Cushing’s disease. Nat. Genet. 2015, 47, 31–38. [Google Scholar] [CrossRef]
- Ma, Z.Y.; Song, Z.J.; Chen, J.H.; Wang, Y.F.; Li, S.Q.; Zhou, L.F.; Mao, Y.; Li, Y.M.; Hu, R.G.; Zhang, Z.Y.; et al. Recurrent gain-of-function USP8 mutations in Cushing’s disease. Cell Res. 2015, 25, 306–317. [Google Scholar] [CrossRef]
- Chen, J.; Jian, X.; Deng, S.; Ma, Z.; Shou, X.; Shen, Y.; Zhang, Q.; Song, Z.; Li, Z.; Peng, H.; et al. Identification of recurrent USP48 and BRAF mutations in Cushing’s disease. Nat. Commun. 2018, 9, 3171. [Google Scholar] [CrossRef]
- Sbiera, S.; Perez-Rivas, L.G.; Taranets, L.; Weigand, I.; Flitsch, J.; Graf, E.; Monoranu, C.M.; Saeger, W.; Hagel, C.; Honegger, J.; et al. Driver mutations in USP8 wild type Cushing’s disease. Neuro-Oncology 2019. [Google Scholar] [CrossRef]
- Faucz, F.R.; Tirosh, A.; Tatsi, C.; Berthon, A.; Hernandez-Ramirez, L.C.; Settas, N.; Angelousi, A.; Correa, R.; Papadakis, G.Z.; Chittiboina, P.; et al. Somatic USP8 Gene Mutations Are a Common Cause of Pediatric Cushing Disease. J. Clin. Endocrinol. Metab. 2017, 102, 2836–2843. [Google Scholar] [CrossRef]
- Hayashi, K.; Inoshita, N.; Kawaguchi, K.; Ibrahim Ardisasmita, A.; Suzuki, H.; Fukuhara, N.; Okada, M.; Nishioka, H.; Takeuchi, Y.; Komada, M.; et al. The USP8 mutational status may predict drug susceptibility in corticotroph adenomas of Cushing’s disease. Eur. J. Endocrinol. 2016, 174, 213–226. [Google Scholar] [CrossRef]
- Perez-Rivas, L.G.; Osswald, A.; Knosel, T.; Lucia, K.; Schaaf, C.; Hristov, M.; Fazel, J.; Kirchner, T.; Beuschlein, F.; Reincke, M.; et al. Expression and mutational status of USP8 in tumors causing ectopic ACTH secretion syndrome. Endocr. Relat. Cancer 2017, 24, L73–L77. [Google Scholar] [CrossRef] [PubMed]
- Perez-Rivas, L.G.; Theodoropoulou, M.; Ferrau, F.; Nusser, C.; Kawaguchi, K.; Stratakis, C.A.; Faucz, F.R.; Wildemberg, L.E.; Assie, G.; Beschorner, R.; et al. The Gene of the Ubiquitin-Specific Protease 8 Is Frequently Mutated in Adenomas Causing Cushing’s Disease. J. Clin. Endocrinol. Metab. 2015, 100, E997–E1004. [Google Scholar] [CrossRef] [PubMed]
- Perez-Rivas, L.G.; Theodoropoulou, M.; Puar, T.H.; Fazel, J.; Stieg, M.R.; Ferrau, F.; Assie, G.; Gadelha, M.R.; Deutschbein, T.; Fragoso, M.C.; et al. Somatic USP8 mutations are frequent events in corticotroph tumor progression causing Nelson’s tumor. Eur. J. Endocrinol. 2018, 178, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Yaffe, M.B.; Rittinger, K.; Volinia, S.; Caron, P.R.; Aitken, A.; Leffers, H.; Gamblin, S.J.; Smerdon, S.J.; Cantley, L.C. The structural basis for 14-3-3:phosphopeptide binding specificity. Cell 1997, 91, 961–971. [Google Scholar] [CrossRef]
- Mizuno, E.; Kitamura, N.; Komada, M. 14-3-3-dependent inhibition of the deubiquitinating activity of UBPY and its cancellation in the M phase. Exp. Cell Res. 2007, 313, 3624–3634. [Google Scholar] [CrossRef]
- Ronchi, C.L.; Peverelli, E.; Herterich, S.; Weigand, I.; Mantovani, G.; Schwarzmayr, T.; Sbiera, S.; Allolio, B.; Honegger, J.; Appenzeller, S.; et al. Landscape of somatic mutations in sporadic GH-secreting pituitary adenomas. Eur. J. Endocrinol. 2016, 174, 363–372. [Google Scholar] [CrossRef]
- Song, Z.J.; Reitman, Z.J.; Ma, Z.Y.; Chen, J.H.; Zhang, Q.L.; Shou, X.F.; Huang, C.X.; Wang, Y.F.; Li, S.Q.; Mao, Y.; et al. The genome-wide mutational landscape of pituitary adenomas. Cell Res. 2016, 26, 1255–1259. [Google Scholar] [CrossRef]
- Cohen, M.; Persky, R.; Stegemann, R.; Hernandez-Ramirez, L.C.; Zeltser, D.; Lodish, M.B.; Chen, A.; Keil, M.F.; Tatsi, C.; Faucz, F.R.; et al. Germline USP8 Mutation Associated With Pediatric Cushing Disease and Other Clinical Features: A New Syndrome. J. Clin. Endocrinol. Metab. 2019, 104, 4676–4682. [Google Scholar] [CrossRef]
- Wilkinson, K.D. Ubiquitination and deubiquitination: Targeting of proteins for degradation by the proteasome. Semin. Cell Dev. Biol. 2000, 11, 141–148. [Google Scholar] [CrossRef]
- Suresh, B.; Lee, J.; Kim, K.S.; Ramakrishna, S. The Importance of Ubiquitination and Deubiquitination in Cellular Reprogramming. Stem Cells Int. 2016, 2016, 6705927. [Google Scholar] [CrossRef]
- Wing, S.S. Deubiquitinating enzymes—The importance of driving in reverse along the ubiquitin-proteasome pathway. Int. J. Biochem. Cell Biol. 2003, 35, 590–605. [Google Scholar] [CrossRef]
- Avvakumov, G.V.; Walker, J.R.; Xue, S.; Finerty, P.J., Jr.; Mackenzie, F.; Newman, E.M.; Dhe-Paganon, S. Amino-terminal dimerization, NRDP1-rhodanese interaction, and inhibited catalytic domain conformation of the ubiquitin-specific protease 8 (USP8). J. Biol. Chem. 2006, 281, 38061–38070. [Google Scholar] [CrossRef] [PubMed]
- Roberts, P.J.; Der, C.J. Targeting the Raf-MEK-ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene 2007, 26, 3291–3310. [Google Scholar] [CrossRef] [PubMed]
- Weigand, I.; Knobloch, L.; Flitsch, J.; Saeger, W.; Monoranu, C.M.; Hofner, K.; Herterich, S.; Rotermund, R.; Ronchi, C.L.; Buchfelder, M.; et al. Impact of USP8 Gene Mutations on Protein Deregulation in Cushing Disease. J. Clin. Endocrinol. Metab. 2019, 104, 2535–2546. [Google Scholar] [CrossRef]
- Sbiera, S.; Deutschbein, T.; Weigand, I.; Reincke, M.; Fassnacht, M.; Allolio, B. The New Molecular Landscape of Cushing’s Disease. Trends Endocrinol. Metab. 2015, 26, 573–583. [Google Scholar] [CrossRef]
- Xia, R.; Jia, H.; Fan, J.; Liu, Y.; Jia, J. USP8 promotes smoothened signaling by preventing its ubiquitination and changing its subcellular localization. PLoS Biol. 2012, 10, e1001238. [Google Scholar] [CrossRef]
- Vila, G.; Papazoglou, M.; Stalla, J.; Theodoropoulou, M.; Stalla, G.K.; Holsboer, F.; Paez-Pereda, M. Sonic hedgehog regulates CRH signal transduction in the adult pituitary. FASEB J. 2005, 19, 281–283. [Google Scholar] [CrossRef]
- Du, H.N. Transcription, DNA damage and beyond: The roles of histone ubiquitination and deubiquitination. Curr. Protein Pept. Sci. 2012, 13, 447–466. [Google Scholar] [CrossRef]
- Jeong, C.H. Inhibition of Ubiquitin-specific Peptidase 8 Suppresses Growth of Gefitinib-resistant Non-small Cell Lung Cancer Cells by Inducing Apoptosis. J. Cancer Prev. 2015, 20, 57–63. [Google Scholar] [CrossRef]
- Ben-Shlomo, A.; Cooper, O. Role of tyrosine kinase inhibitors in the treatment of pituitary tumours: From bench to bedside. Curr. Opin. Endocrinol. Diabetes Obes. 2017, 24, 301–305. [Google Scholar] [CrossRef]
- Gonnissen, A.; Isebaert, S.; Haustermans, K. Targeting the Hedgehog signaling pathway in cancer: Beyond Smoothened. Oncotarget 2015, 6, 13899–13913. [Google Scholar] [CrossRef] [PubMed]
- Li, F.P.; Fraumeni, J.F., Jr. Soft-tissue sarcomas, breast cancer, and other neoplasms. A familial syndrome? Ann. Intern. Med. 1969, 71, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Varley, J.M. Germline TP53 mutations and Li-Fraumeni syndrome. Hum. Mutat. 2003, 21, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Hollstein, M.; Sidransky, D.; Vogelstein, B.; Harris, C.C. p53 mutations in human cancers. Science 1991, 253, 49–53. [Google Scholar] [CrossRef]
- Schmitt, C.A.; Fridman, J.S.; Yang, M.; Baranov, E.; Hoffman, R.M.; Lowe, S.W. Dissecting p53 tumor suppressor functions in vivo. Cancer Cell 2002, 1, 289–298. [Google Scholar] [CrossRef]
- Lambrughi, M.; De Gioia, L.; Gervasio, F.L.; Lindorff-Larsen, K.; Nussinov, R.; Urani, C.; Bruschi, M.; Papaleo, E. DNA-binding protects p53 from interactions with cofactors involved in transcription-independent functions. Nucl. Acids Res. 2016, 44, 9096–9109. [Google Scholar] [CrossRef]
- Zhao, L.Y.; Liu, J.; Sidhu, G.S.; Niu, Y.; Liu, Y.; Wang, R.; Liao, D. Negative regulation of p53 functions by Daxx and the involvement of MDM2. J. Biol. Chem. 2004, 279, 50566–50579. [Google Scholar] [CrossRef]
- Dyer, M.A.; Qadeer, Z.A.; Valle-Garcia, D.; Bernstein, E. ATRX and DAXX: Mechanisms and Mutations. Cold Spring Harb. Perspect. Med. 2017, 7. [Google Scholar] [CrossRef]
- Wang, Y.; Suh, Y.A.; Fuller, M.Y.; Jackson, J.G.; Xiong, S.; Terzian, T.; Quintas-Cardama, A.; Bankson, J.A.; El-Naggar, A.K.; Lozano, G. Restoring expression of wild-type p53 suppresses tumor growth but does not cause tumor regression in mice with a p53 missense mutation. J. Clin. Investig. 2011, 121, 893–904. [Google Scholar] [CrossRef]
- Synnott, N.C.; Madden, S.F.; Bykov, V.J.N.; Crown, J.; Wiman, K.G.; Duffy, M.J. The Mutant p53-Targeting Compound APR-246 Induces ROS-Modulating Genes in Breast Cancer Cells. Transl. Oncol. 2018, 11, 1343–1349. [Google Scholar] [CrossRef]
- Hugdahl, E.; Kalvenes, M.B.; Puntervoll, H.E.; Ladstein, R.G.; Akslen, L.A. BRAF-V600E expression in primary nodular melanoma is associated with aggressive tumour features and reduced survival. Br. J. Cancer 2016, 114, 801–808. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Lee, K.E.; Myong, J.P.; Park, J.H.; Jeon, Y.K.; Min, H.S.; Park, S.Y.; Jung, K.C.; Koo do, H.; Youn, Y.K. BRAF V600E mutation is associated with tumor aggressiveness in papillary thyroid cancer. World J. Surg. 2012, 36, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Hillhouse, E.W.; Grammatopoulos, D.K. The molecular mechanisms underlying the regulation of the biological activity of corticotropin-releasing hormone receptors: Implications for physiology and pathophysiology. Endocr. Rev. 2006, 27, 260–286. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, L.L.S.; Franco, O.L.; Alencar, S.A.; Porto, W.F. Deciphering the structural basis for glucocorticoid resistance caused by missense mutations in the ligand binding domain of glucocorticoid receptor. J. Mol. Graph. Model. 2019, 92, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Wagner, K.V.; Wang, X.D.; Liebl, C.; Scharf, S.H.; Muller, M.B.; Schmidt, M.V. Pituitary glucocorticoid receptor deletion reduces vulnerability to chronic stress. Psychoneuroendocrinology 2011, 36, 579–587. [Google Scholar] [CrossRef]
- Melmed, S. Mechanisms for pituitary tumorigenesis: The plastic pituitary. J. Clin. Investig. 2003, 112, 1603–1618. [Google Scholar] [CrossRef]
- Al-Gahtany, M.; Horvath, E.; Kovacs, K. Pituitary hyperplasia. Hormones 2003, 2, 149–158. [Google Scholar] [CrossRef]
- Miesfeld, R.; Rusconi, S.; Godowski, P.J.; Maler, B.A.; Okret, S.; Wikstrom, A.C.; Gustafsson, J.A.; Yamamoto, K.R. Genetic complementation of a glucocorticoid receptor deficiency by expression of cloned receptor cDNA. Cell 1986, 46, 389–399. [Google Scholar] [CrossRef]
- Meijer, O.C.; Koorneef, L.L.; Kroon, J. Glucocorticoid receptor modulators. Ann. Endocrinol. 2018, 79, 107–111. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Bader, G.D.; Hogue, C.W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4. [Google Scholar] [CrossRef] [PubMed]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pages, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sbiera, S.; Kunz, M.; Weigand, I.; Deutschbein, T.; Dandekar, T.; Fassnacht, M. The New Genetic Landscape of Cushing’s Disease: Deubiquitinases in the Spotlight. Cancers 2019, 11, 1761. https://doi.org/10.3390/cancers11111761
Sbiera S, Kunz M, Weigand I, Deutschbein T, Dandekar T, Fassnacht M. The New Genetic Landscape of Cushing’s Disease: Deubiquitinases in the Spotlight. Cancers. 2019; 11(11):1761. https://doi.org/10.3390/cancers11111761
Chicago/Turabian StyleSbiera, Silviu, Meik Kunz, Isabel Weigand, Timo Deutschbein, Thomas Dandekar, and Martin Fassnacht. 2019. "The New Genetic Landscape of Cushing’s Disease: Deubiquitinases in the Spotlight" Cancers 11, no. 11: 1761. https://doi.org/10.3390/cancers11111761
APA StyleSbiera, S., Kunz, M., Weigand, I., Deutschbein, T., Dandekar, T., & Fassnacht, M. (2019). The New Genetic Landscape of Cushing’s Disease: Deubiquitinases in the Spotlight. Cancers, 11(11), 1761. https://doi.org/10.3390/cancers11111761





