Necessities in the Processing of DNA Double Strand Breaks and Their Effects on Genomic Instability and Cancer
Abstract
:1. Introduction: General Considerations for DSB Repair Pathway Utilization
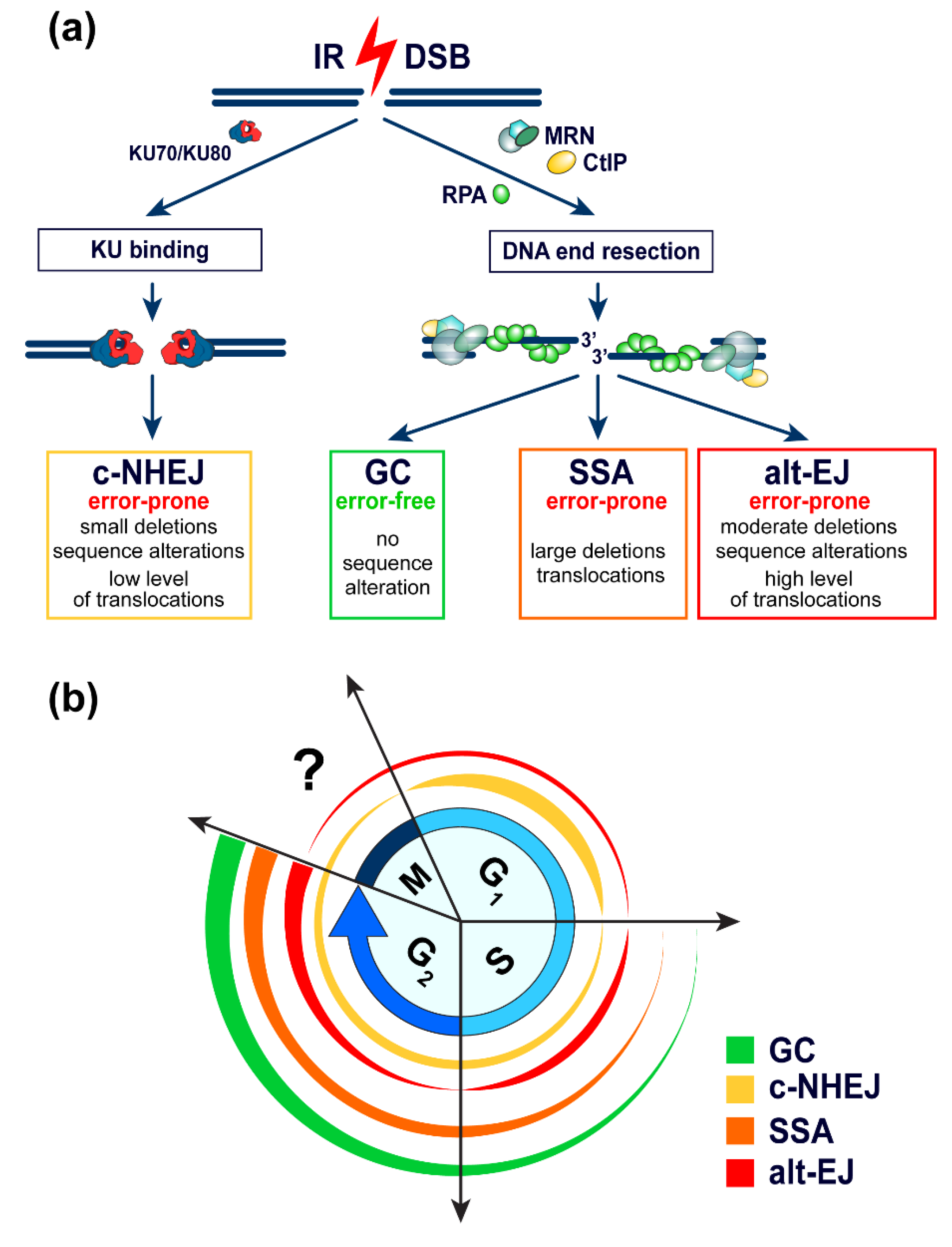
2. Characteristics of Pathways Processing DSBs
2.1. General Considerations
2.2. Rapid Removal of DNA Ends by c-NHEJ
2.3. DNA-End-Resection-Dependent Rejoining of DSBs by GC, SSA, and Alt-EJ
2.3.1. Gene Conversion (GC)
2.3.2. Single Strand Annealing (SSA)
2.3.3. Alternative End-Joining (Alt-EJ)
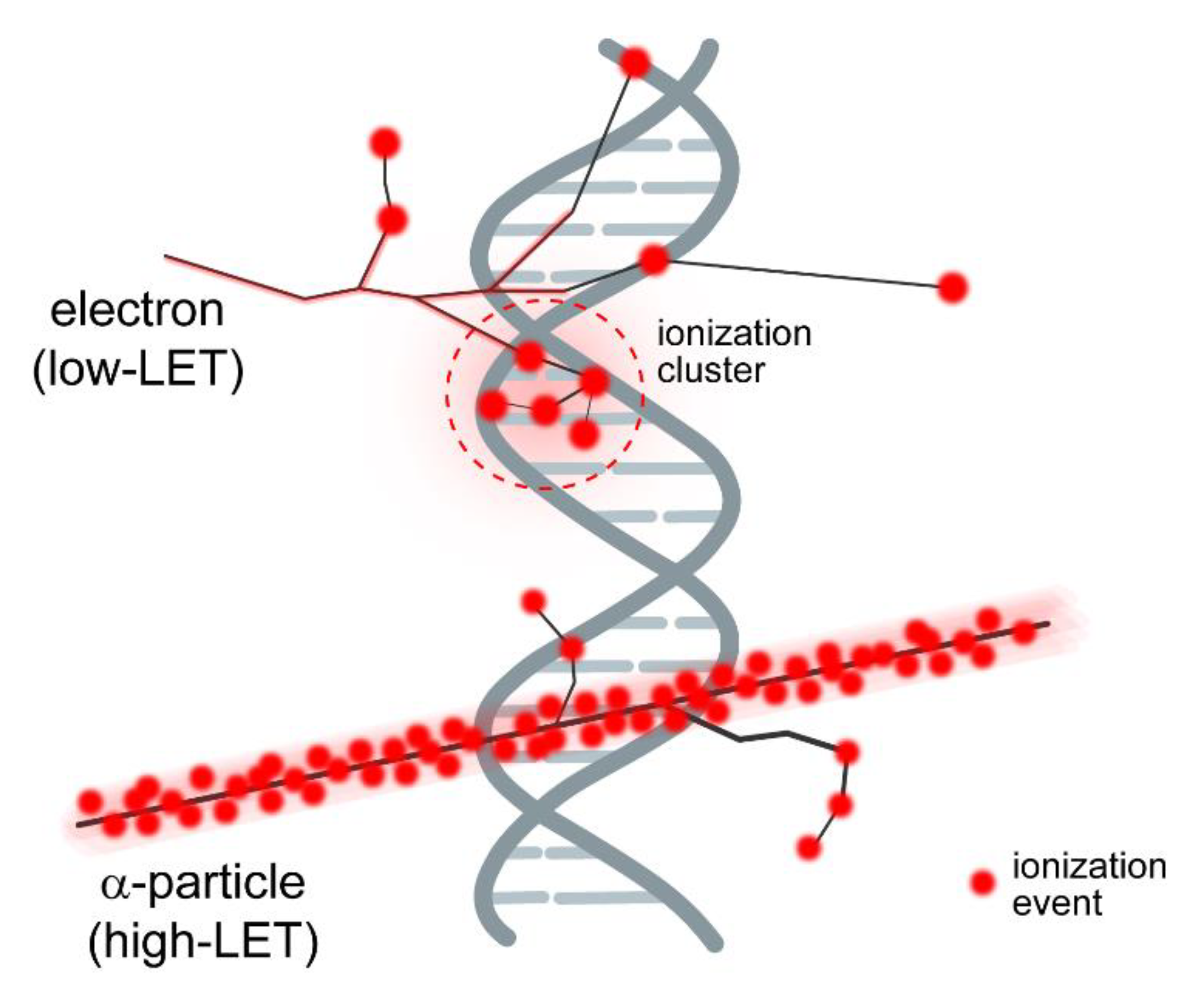
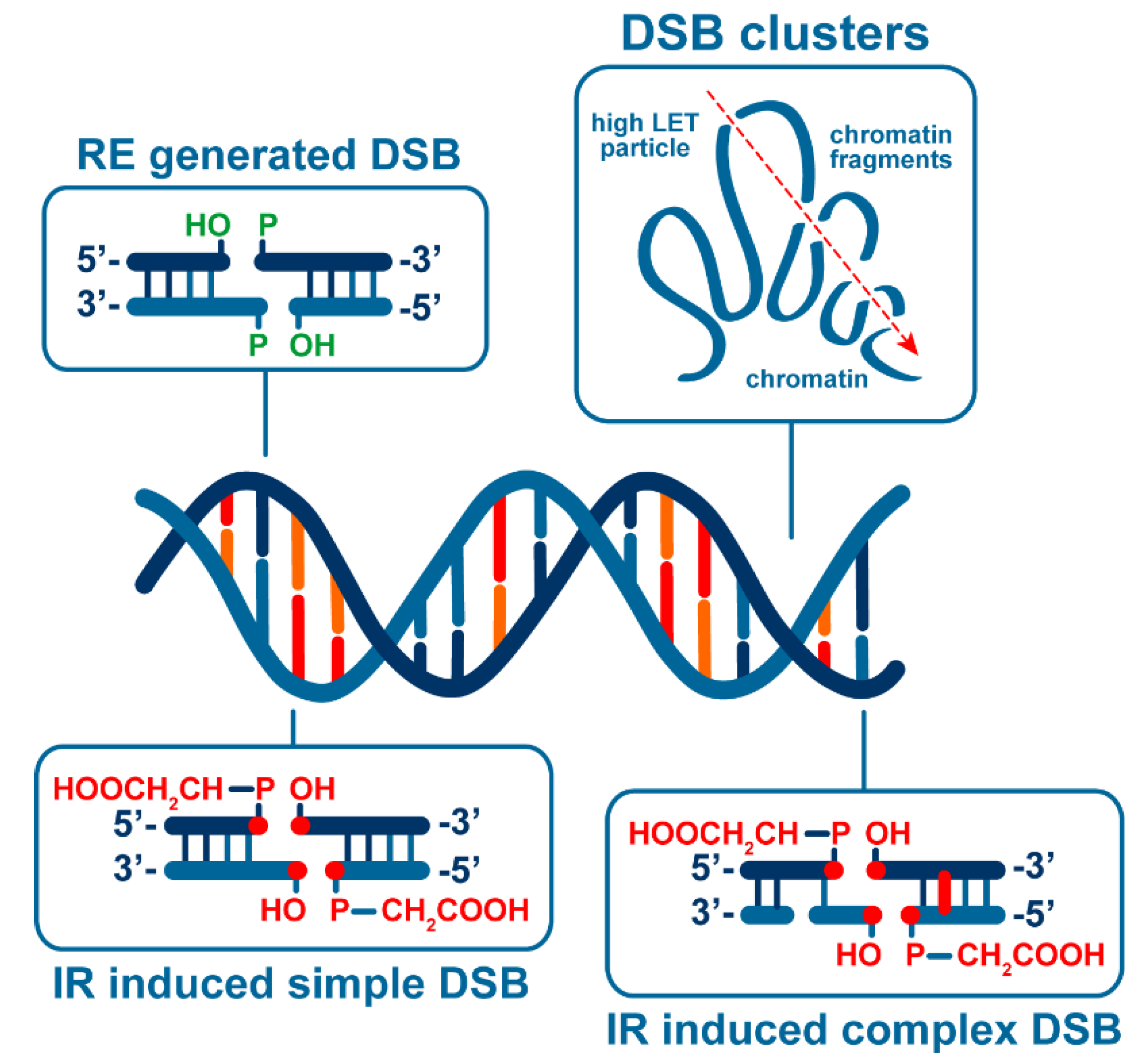
3. IR Induces DSBs of Different Complexity
4. A Subset of DSBs Induced by High LET IR Compromises c-NHEJ
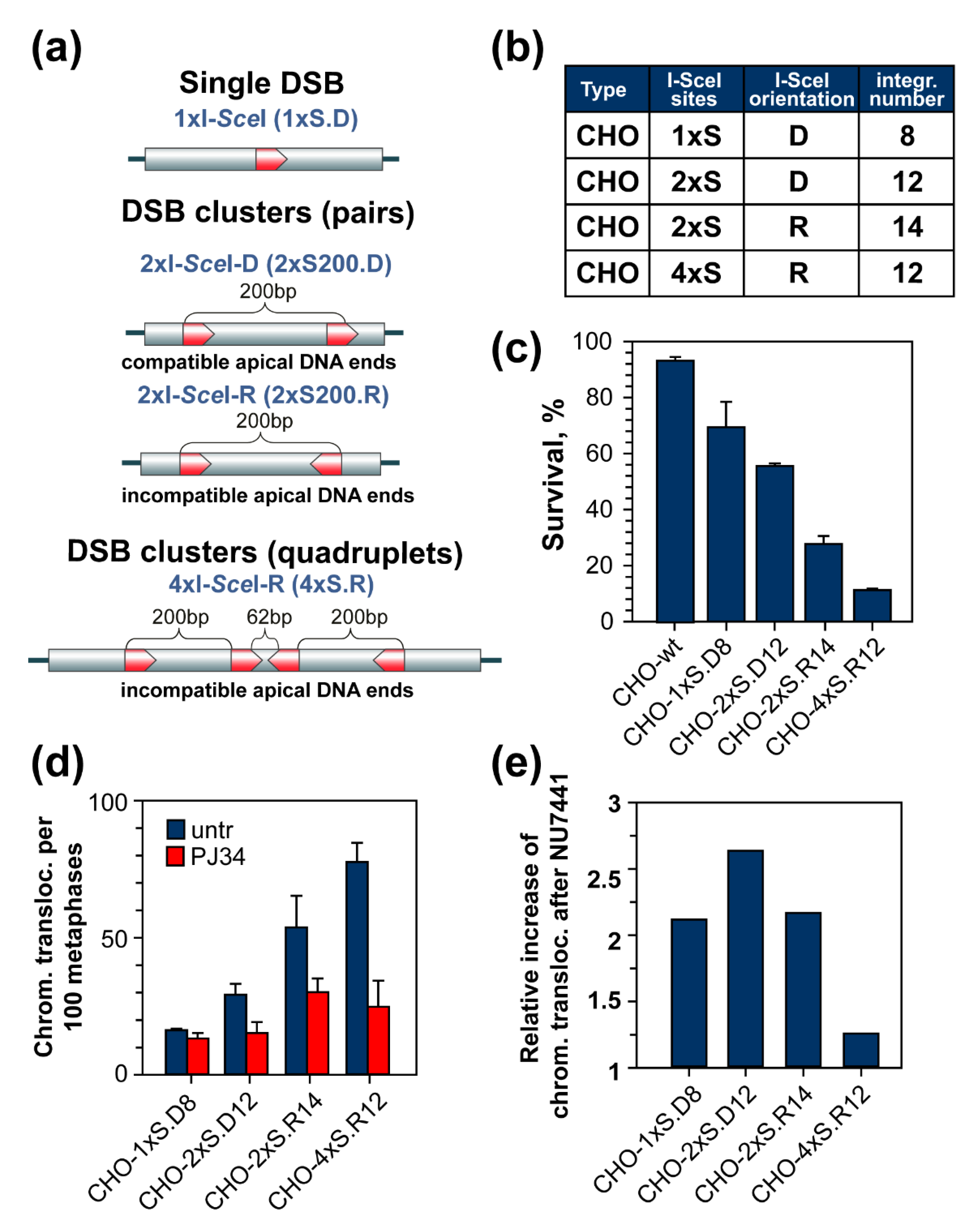
5. DSB Clusters Suppress c-NHEJ and Cause Translocations
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Kuppers, R. Mechanisms of B-cell lymphoma pathogenesis. Nat. Rev. Cancer 2005, 5, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Lieber, M.R. Mechanisms of human lymphoid chromosomal translocations. Nat. Rev. Cancer 2016, 16, 387–398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bunting, S.F.; Nussenzweig, A. End-joining, translocations and cancer. Nat. Rev. Cancer 2013, 13, 443–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schipler, A.; Iliakis, G. DNA double-strand-break complexity levels and their possible contributions to the probability for error-prone processing and repair pathway choice. Nucleic Acids Res. 2013, 41, 7589–7605. [Google Scholar] [CrossRef] [Green Version]
- Mladenova, V.; Mladenov, E.; Iliakis, G. Novel Biological Approaches for Testing the Contributions of Single DSBs and DSB Clusters to the Biological Effects of High LET Radiation. Front. Oncol. 2016, 6, 163. [Google Scholar] [CrossRef] [Green Version]
- Mladenov, E.; Magin, S.; Soni, A.; Iliakis, G. DNA double-strand break repair as determinant of cellular radiosensitivity to killing and target in radiation therapy. Front. Oncol. 2013, 3, 113. [Google Scholar] [CrossRef]
- Mladenov, E.; Iliakis, G. Induction and repair of DNA double strand breaks: The increasing spectrum of non-homologous end joining pathways. Mutat. Res. 2011, 711, 61–72. [Google Scholar] [CrossRef]
- Friedberg, E.C.; Walker, G.C.; Siede, W.; Wood, R.D.; Schultz, R.A.; Ellenberger, T. DNA Repair and Mutagenesis, 2nd ed.; ASM Press: Washington, DC, USA, 2006. [Google Scholar]
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature 2009, 461, 1071–1078. [Google Scholar] [CrossRef] [Green Version]
- Zhou, B.B.; Elledge, S.J. The DNA damage response: Putting checkpoints in perspective. Nature 2000, 408, 433–439. [Google Scholar] [CrossRef]
- Terasima, T.; Tolmach, L.J. X-ray sensitivity and DNA synthesis in synchronous populations of HeLa cells. Science 1963, 140, 490–492. [Google Scholar] [CrossRef]
- Chang, H.H.Y.; Pannunzio, N.R.; Adachi, N.; Lieber, M.R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 2017, 18, 495. [Google Scholar] [CrossRef] [PubMed]
- Ceccaldi, R.; Rondinelli, B.; D’Andrea, A.D. Repair Pathway Choices and Consequences at the Double-Strand Break. Trends Cell Biol. 2016, 26, 52–64. [Google Scholar] [CrossRef] [PubMed]
- Nick McElhinny, S.A.; Havener, J.M.; Garcia-Diaz, M.; Juarez, R.; Bebenek, K.; Kee, B.L.; Blanco, L.; Kunkel, T.A.; Ramsden, D.A. A Gradient of Template Dependence Defines Distinct Biological Roles for Family X Polymerases in Nonhomologous End Joining. Mol. Cell 2005, 19, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Soni, A.; Siemann, M.; Pantelias, G.E.; Iliakis, G. Marked cell cycle-dependent contribution of alternative end joining to formation of chromosome translocations by stochastically induced DNA double strand breaks in human cells. Mutat. Res. 2015, 793, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Ghezraoui, H.; Piganeau, M.; Renouf, B.; Renaud, J.-B.; Sallmyr, A.; Ruis, B.; Oh, S.; Tomkinson, A.E.; Hendrickson, E.A.; Giovannangeli, C.; et al. Chromosomal Translocations in Human Cells Are Generated by Canonical Nonhomologous End-Joining. Mol. Cell 2014, 55, 829–842. [Google Scholar] [CrossRef] [Green Version]
- Symington, L.S.; Gautier, J. Double-Strand Break End Resection and Repair Pathway Choice. Annu. Rev. Genet. 2011, 45, 247–271. [Google Scholar] [CrossRef]
- Hustedt, N.; Durocher, D. The control of DNA repair by the cell cycle. Nat. Cell Biol. 2017, 19, 1–9. [Google Scholar] [CrossRef]
- McVey, M.; Khodaverdian, V.Y.; Meyer, D.; Cerqueira, P.G.; Heyer, W.D. Eukaryotic DNA Polymerases in Homologous Recombination. Annu. Rev. Genet. 2016, 50, 393–421. [Google Scholar] [CrossRef] [Green Version]
- Heyer, W.-D.; Ehmsen, K.T.; Liu, J. Regulation of Homologous Recombination in Eukaryotes. Annu. Rev. Genet. 2010, 44, 113–139. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.-C.; Feng, W.; Lim, P.X.; Kass, E.M.; Jasin, M. Homology-Directed Repair and the Role of BRCA1, BRCA2, and Related Proteins in Genome Integrity and Cancer. Annu. Rev. Cancer Biol. 2018, 2, 313–336. [Google Scholar] [CrossRef]
- Haber, J. Genome Stability; Garland Science, Taylor & Francis Group, LLC: Abingdon, UK, 2014. [Google Scholar]
- Ivanov, E.L.; Sugawara, N.; Fishman-Lobell, J.; Haber, J.E. Genetic requirements for the single-strand annealing pathway of double-strand break repair in Saccharomyces cerevisiae. Genetics 1996, 142, 693–704. [Google Scholar] [PubMed]
- Ochs, F.; Somyajit, K.; Altmeyer, M.; Rask, M.-B.; Lukas, J.; Lukas, C. 53BP1 fosters fidelity of homology-directed DNA repair. Nat. Struct. Mol. Biol. 2016, 23, 714–721. [Google Scholar] [CrossRef] [PubMed]
- Rothenberg, E.; Grimme, J.M.; Spies, M.; Ha, T. Human Rad52-mediated homology search and annealing occurs by continuous interactions between overlapping nucleoprotein complexes. Proc. Natl. Acad. Sci. USA 2008, 105, 20274–20279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sallmyr, A.; Tomkinson, A.E. Repair of DNA double-strand breaks by mammalian alternative end-joining pathways. J. Biol. Chem. 2018, 293, 10536–10546. [Google Scholar] [CrossRef] [Green Version]
- Iliakis, G. Backup pathways of NHEJ in cells of higher eukaryotes: Cell cycle dependence. Radiother. Oncol. 2009, 92, 310–315. [Google Scholar] [CrossRef]
- McVey, M.; Lee, S.E. MMEJ repair of double-strand breaks (director’s cut): Deleted sequences and alternative endings. Trends Genet. 2008, 24, 529–538. [Google Scholar] [CrossRef]
- Sfeir, A.; Symington, L.S. Microhomology-Mediated End Joining: A Back-up Survival Mechanism or Dedicated Pathway? Trends Biochem. Sci. 2015, 40, 701–714. [Google Scholar] [CrossRef]
- Mateos-Gomez, P.A.; Gong, F.; Nair, N.; Miller, K.M.; Lazzerini-Denchi, E.; Sfeir, A. Mammalian polymerase [θ] promotes alternative NHEJ and suppresses recombination. Nature 2015, 518, 254–257. [Google Scholar] [CrossRef]
- Wood, R.D.; Doublie, S. DNA polymerase theta (POLQ), double-strand break repair, and cancer. DNA Repair 2016, 44, 22–32. [Google Scholar] [CrossRef]
- Ceccaldi, R.; Liu, J.C.; Amunugama, R.; Hajdu, I.; Primack, B.; Petalcorin, M.I.R.; O’Connor, K.W.; Konstantinopoulos, P.A.; Elledge, S.J.; Boulton, S.J.; et al. Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair. Nature 2015, 518, 258–262. [Google Scholar] [CrossRef]
- Windhofer, F.; Wu, W.; Wang, M.; Singh, S.K.; Saha, J.; Rosidi, B.; Iliakis, G. Marked dependence on growth state of backup pathways of NHEJ. Int. J. Radiat. Oncol. Biol. Phys. 2007, 68, 1462–1470. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Wang, M.; Wu, W.; Singh, S.K.; Mussfeldt, T.; Iliakis, G. Repair of radiation induced DNA double strand breaks by backup NHEJ is enhanced in G2. DNA Repair 2008, 7, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Bindra, R.S.; Goglia, A.G.; Jasin, M.; Powell, S.N. Development of an assay to measure mutagenic non-homologous end-joining repair activity in mammalian cells. Nucleic Acids Res. 2013, 41, e115. [Google Scholar] [CrossRef]
- Ward, J.F. Biochemistry of DNA lesions. Radiat. Res. 1985, 104, S103–S111. [Google Scholar] [CrossRef]
- Ward, J.F. The yield of DNA double-strand breaks produced intracellularly by ionizing radiation: A review. Int. J. Radiat. Biol. 1990, 57, 1141–1150. [Google Scholar] [CrossRef] [PubMed]
- Allen, C.; Borak, T.B.; Tsujii, H.; Nickoloff, J.A. Heavy charged particle radiobiology: Using enhanced biological effectiveness and improved beam focusing to advance cancer therapy. Mutat. Res. 2011, 711, 150–157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, J.F.; Evans, J.W.; Limoli, C.L.; Calabro-Jones, P.M. Radiation and hydrogen peroxide induced free radical damage to DNA. Br. J. Cancer 1987, 55, 105–112. [Google Scholar]
- Sutherland, B.M.; Bennett, P.V.; Sidorkina, O.; Laval, J. Clustered DNA damages induced in isolated DNA and in human cells by low doses of ionizing radiation. Proc. Natl. Acad. Sci. USA 2000, 97, 103–108. [Google Scholar] [CrossRef] [Green Version]
- Sutherland, R.M.; Bennett, P.V.; Sutherland, J.C.; Laval, J. Clustered DNA Damages Induced by X rays in Human Cells. Radiat. Res. 2002, 157, 611–616. [Google Scholar] [CrossRef]
- Prise, K.M.; Davies, S.; Michael, B.D. The relationship between radiation-induced DNA double-strand breaks and cell kill in hamster V79 fibroblasts irradiated with 250 kVp X-rays, 2.3 MeV neutrons or 238 Pu a-particles. Int. J. Radiat. Biol. 1987, 52, 893–902. [Google Scholar] [CrossRef]
- Peak, M.J.; Wang, L.; Hill, C.K.; Peak, J.G. Comparison of repair of DNA double-strand breaks caused by neutron or gamma radiation in cultured human cells. Int. J. Radiat. Biol. 1991, 60, 891–898. [Google Scholar] [CrossRef] [PubMed]
- Prise, K.M.; Folkard, M.; Davies, S.; Michael, B.D. The irradiation of V79 mammalian cells by protons with energies below 2 MeV. Part II. Measurement of oxygen enhancement ratios and DNA damage. Int. J. Radiat. Biol. 1990, 58, 261–277. [Google Scholar] [CrossRef] [PubMed]
- Jenner, T.J.; Belli, M.; Goodhead, D.T.; Ianzini, F.; Simone, G.; Tabocchini, M.A. Direct comparison of biological effectiveness of protons and alpha-particles of the same LET. III. Initial yield of DNA double-strand breaks in V79 cells. Int. J. Radiat. Biol. 1992, 61, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Jenner, T.J.; de Lara, C.M.; O’Neill, P.; Stevens, D.L. Induction and rejoining of DNA double-strand breaks in V79-4 mammalian cells following gamma- and alpha-irradiation. Int. J. Radiat. Biol. 1993, 64, 265–273. [Google Scholar] [CrossRef]
- Coquerelle, T.M.; Weibezahn, K.F.; Luecke-Huhle, C. Rejoining of double strand breaks in normal human and ataxia-telangiectasia fibroblasts after exposure to 60Co g-rays, 241Am a-particles or bleomycin. Int. J. Radiat. Biol. 1987, 51, 209–218. [Google Scholar]
- Fox, J.C.; McNally, N.H. The rejoining of DNA double-strand breaks following irradiation with 238Pu a-particles: Evidence for a fast component of repair as measured by neutral filter elution. Int. J. Radiat. Biol. 1990, 57, 513–521. [Google Scholar] [CrossRef]
- Rydberg, B.; Heilbronn, L.; Holley, W.R.; Löbrich, M.; Zeitlin, C.; Chatterjee, A.; Cooper, P.K. Spatial Distribution and Yield of DNA Double-Strand Breaks Induced by 3–7 MeV Helium Ions in Human Fibroblasts. Radiat. Res. 2002, 158, 32–42. [Google Scholar] [CrossRef]
- Stewart, R.D.; Yu, V.K.; Georgakilas, A.G.; Koumenis, C.; Park, J.H.; Carlson, D.J. Effects of Radiation Quality and Oxygen on Clustered DNA Lesions and Cell Death. Radiat. Res. 2011, 176, 587–602. [Google Scholar] [CrossRef]
- Georgakilas, A.G.; O’Neill, P.; Stewart, R.D. Induction and Repair of Clustered DNA Lesions: What do we know so far? Radiat. Res. 2013, 180, 100–109. [Google Scholar] [CrossRef]
- Hada, M.; Georgakilas, A.G. Formation of Clustered DNA Damage after High-LET Irradiation: A Review. J. Radiat. Res. 2008, 49, 203–210. [Google Scholar] [CrossRef]
- Dianov, G.L.; O’Neill, P.; Goodhead, D.T. Securing genome stability by orchestrating DNA repair: Removal of radiation-induced clustered lesions in DNA. BioEssays 2001, 23, 745–749. [Google Scholar] [CrossRef] [PubMed]
- Henner, W.D.; Grunberg, S.M.; Haseltine, W.A. Sites and structure of g radiation-induced DNA strand breaks. J. Biol. Chem. 1982, 257, 11750–11754. [Google Scholar] [PubMed]
- Henner, W.D.; Rodriguez, L.O.; Hecht, S.M.; Haseltine, W.A. Gamma-ray Induced deoxyribonucleic acid strand breaks. J. Biol. Chem. 1983, 258, 711–713. [Google Scholar] [PubMed]
- Friedland, W.; Dingfelder, M.; Kundrát, P.; Jacob, P. Track structures, DNA targets and radiation effects in the biophysical Monte Carlo simulation code PARTRAC. Mutat. Res. Mol. Mech. Mutagen. 2011, 711, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Schipler, A.; Mladenova, V.; Soni, A.; Nikolov, V.; Saha, J.; Mladenov, E.; Iliakis, G. Chromosome thripsis by DNA double strand break clusters causes enhanced cell lethality, chromosomal translocations and 53BP1-recruitment. Nucleic Acids Res. 2016, 44, 7673–7690. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, S.K.; Bencsik-Theilen, A.; Mladenov, E.; Jakob, B.; Taucher-Scholz, G.; Iliakis, G. Reduced contribution of thermally labile sugar lesions to DNA double strand break formation after exposure to heavy ions. Radiat. Oncol. 2013, 8, 77. [Google Scholar] [CrossRef]
- Shadley, J.D.; Whitlock, J.L.; Rotmensch, J.; Atcher, R.W.; Tang, J.; Schwartz, J.L. The effects of radon daughter alpha-particle irradiation in K1 and xrs-5 CHO cell lines. Mutat. Res. 1991, 248, 73–83. [Google Scholar] [CrossRef]
- Zyuzikov, N.A.; Prise, K.M.; Zdzienicka, M.Z.; Newman, H.C.; Michael, B.D.; Trott, K.R. The relationship between the RBE of alpha particles and the radiosensitivity of different mutations of Chinese hamster cells. Radiat. Environ. Biophys. 2001, 40, 243–248. [Google Scholar] [CrossRef]
- Thacker, J.; Stretch, A. Responses of 4 X-ray-sensitive CHO cell mutants to different radiations and to irradiation conditions promoting cellular recovery. Mutat. Res. 1985, 146, 99–108. [Google Scholar] [CrossRef]
- Hill, M.A.; Herdman, M.T.; Stevens, D.L.; Jones, N.J.; Thacker, J.; Goodhead, D.T. Relative Sensitivities of Repair-Deficient Mammalian Cells for Clonogenic Survival after a-Particle Irradiation. Radiat. Res. 2004, 162, 667–676. [Google Scholar] [CrossRef]
- Weyrather, W.K.; Ritter, S.; Scholz, M.; Kraft, G. RBE for carbon track-segment irradiation in cell lines of differing repair capacity. Int. J. Radiat. Biol. 1999, 75, 1357–1364. [Google Scholar] [PubMed]
- Wang, H.; Wang, X.; Zhang, P.; Wang, Y. The Ku-dependent non-homologous end-joining but not other repair pathway is inhibited by high linear energy transfer ionizing radiation. DNA Repair 2008, 7, 725–733. [Google Scholar] [CrossRef] [PubMed]
- Grosse, N.; Fontana, A.O.; Hug, E.B.; Lomax, A.; Coray, A.; Augsburger, M.; Paganetti, H.; Sartori, A.A.; Pruschy, M. Deficiency in Homologous Recombination Renders Mammalian Cells More Sensitive to Proton Versus Photon Irradiation. Int. J. Radiat. Oncol. Biol. Phys. 2014, 88, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Lord, C.J.; Ashworth, A. BRCAness revisited. Nat. Rev. Cancer 2016, 16, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Fenina, M.; Simon-Chazottes, D.; Vandormael-Pournin, S.; Soueid, J.; Langa, F.; Cohen-Tannoudji, M.; Bernard, B.A.; Panthier, J.-J. I-SceI-Mediated Double-Strand Break Does Not Increase the Frequency of Homologous Recombination at the Dct Locus in Mouse Embryonic Stem Cells. PLoS ONE 2012, 7, e39895. [Google Scholar] [CrossRef]
- Iacovoni, J.S.; Caron, P.; Lassadi, I.; Nicolas, E.; Massip, L.; Trouche, D.; Legube, G. High-resolution profiling of gH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 2010, 29, 1446–1457. [Google Scholar] [CrossRef]
- Jasin, M. Genetic manipulation of genomes with rare-cutting endonucleases. Trends Genet. 1996, 12, 224–228. [Google Scholar] [CrossRef]
- Gunn, A.; Stark, J.M. I-SceI-based assays to examine distinct repair outcomes of mammalian chromosomal double strand-breaks. Methods Mol. Biol. 2012, 920, 379–391. [Google Scholar]
- Soutoglou, E.; Misteli, T. Activation of the cellular DNA damage response in the absence of DNA lesions. Science 2008, 320, 1507–1510. [Google Scholar] [CrossRef]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iliakis, G.; Mladenov, E.; Mladenova, V. Necessities in the Processing of DNA Double Strand Breaks and Their Effects on Genomic Instability and Cancer. Cancers 2019, 11, 1671. https://doi.org/10.3390/cancers11111671
Iliakis G, Mladenov E, Mladenova V. Necessities in the Processing of DNA Double Strand Breaks and Their Effects on Genomic Instability and Cancer. Cancers. 2019; 11(11):1671. https://doi.org/10.3390/cancers11111671
Chicago/Turabian StyleIliakis, George, Emil Mladenov, and Veronika Mladenova. 2019. "Necessities in the Processing of DNA Double Strand Breaks and Their Effects on Genomic Instability and Cancer" Cancers 11, no. 11: 1671. https://doi.org/10.3390/cancers11111671
APA StyleIliakis, G., Mladenov, E., & Mladenova, V. (2019). Necessities in the Processing of DNA Double Strand Breaks and Their Effects on Genomic Instability and Cancer. Cancers, 11(11), 1671. https://doi.org/10.3390/cancers11111671




