Towards Multiplex Molecular Diagnosis—A Review of Microfluidic Genomics Technologies
Abstract
1. Introduction
2. Lysis Techniques
2.1. Chemical Lysis
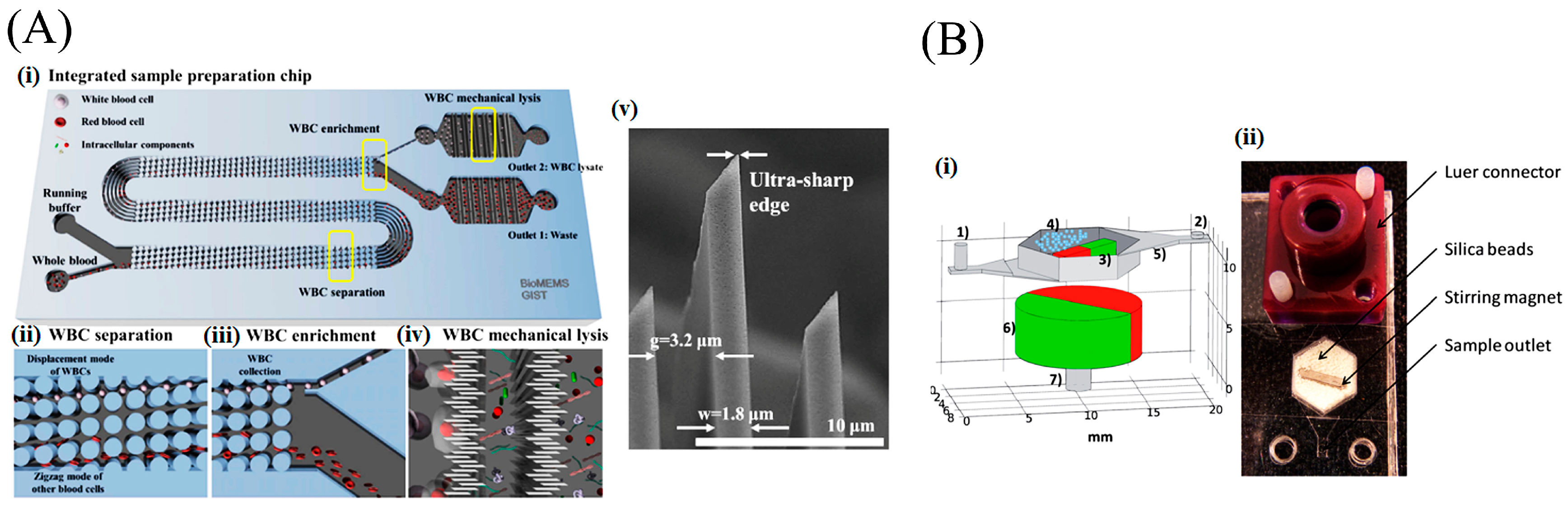
2.2. Mechanical Lysis
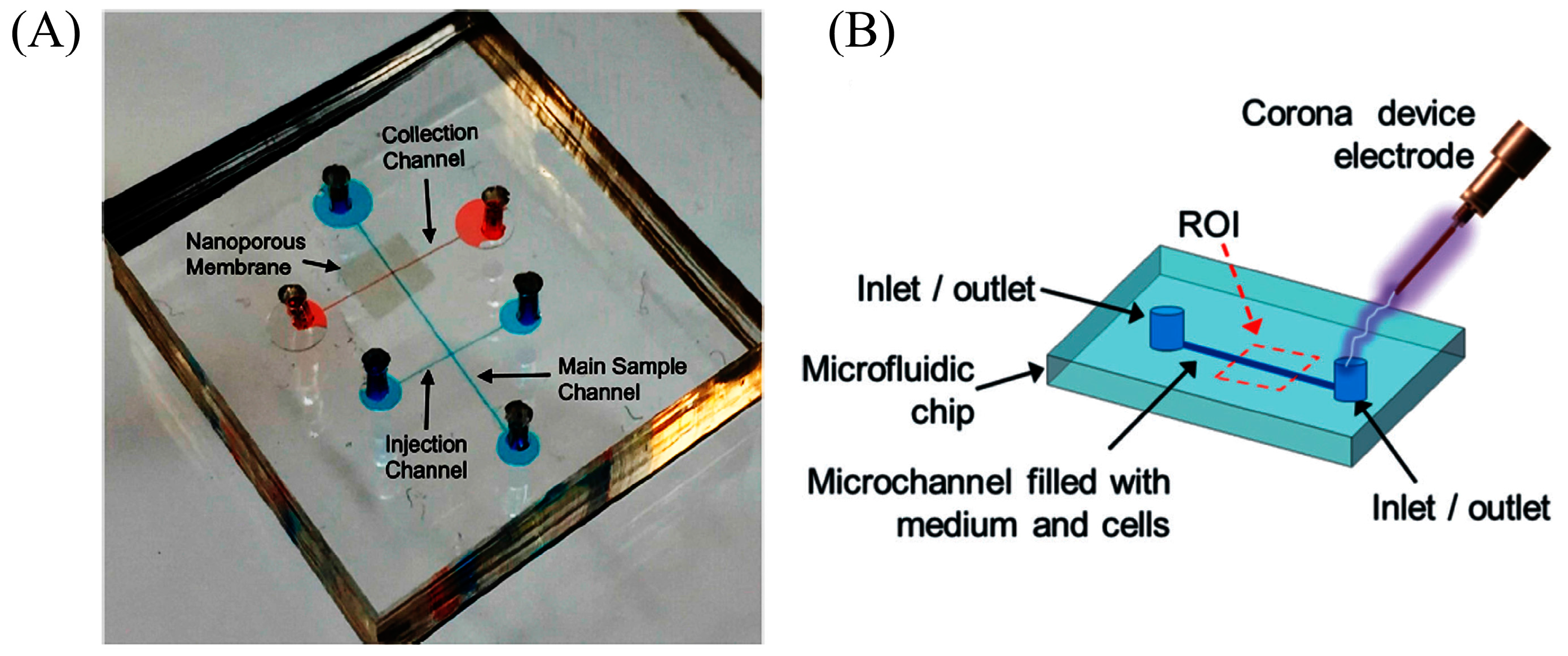
2.3. Electrical Lysis
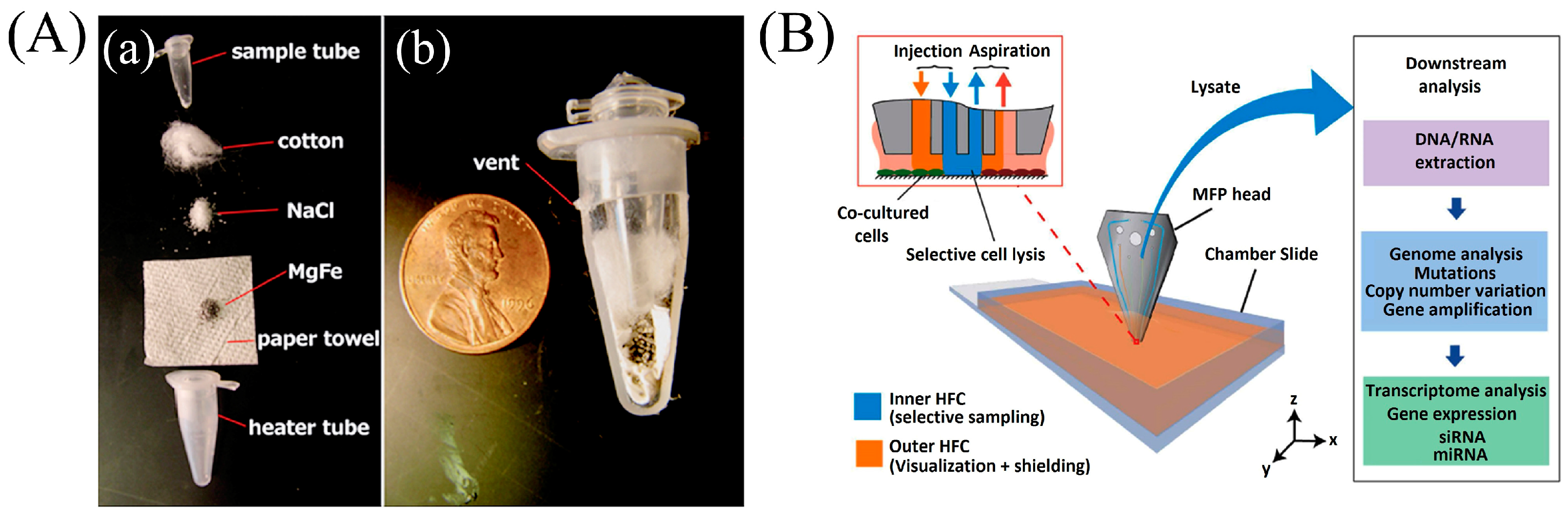
2.4. Thermal Lysis Techniques
3. Nucleic Acid Extraction
3.1. Solid Phase Extraction
3.2. Isotachophoresis
4. Amplification
4.1. Cyclo-Thermal Amplification
4.1.1. Stationary Chamber Polymerase Chain Reaction (PCR)
4.1.2. Continuous-Flow PCR
4.1.3. Multiple Annealing and Looping-Based Amplification Cycles (MALBAC)
4.2. Isothermal Amplification
4.2.1. Nucleic Acid Sequence-Based Amplification (NASBA)
4.2.2. Loop-Mediated Isothermal Amplification (LAMP)
4.2.3. Recombinase Polymerase Amplification (RPA)
4.2.4. Helicase-Dependent Amplification (HDA)
4.2.5. Rolling Circle Amplification (RCA)
4.3. Digital Amplification
5. Detection
6. Novel Microfluidic Actuators
6.1. Paper-Based Microfluidics
6.2. Centrifugal-Based Microfluidics
6.3. Digital Microfluidics
7. Multiplex Integrated Devices
8. Commercial Microfluidic Diagnosis
9. Summary and Conclusions
10. Future Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lee, W.G.; Kim, Y.-G.; Chung, B.G.; Demirci, U.; Khademhosseini, A. Nano/Microfluidics for diagnosis of infectious diseases in developing countries. Adv. Drug Deliv. Rev. 2010, 62, 449–457. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.F.; Goldberg, M.; Rosenthal, S.; Carlson, L.; Chen, J.; Chen, C.; Ramachandran, S. Global rise in human infectious disease outbreaks. J. R. Soc. Interface 2014, 11, 20140950. [Google Scholar] [CrossRef] [PubMed]
- Dye, C. After 2015: Infectious diseases in a new era of health and development. Philos. Trans. R. Soc. B 2015, 369. [Google Scholar] [CrossRef] [PubMed]
- Heesterbeek, H.; Anderson, R.M.; Andreasen, V.; Bansal, S.; De Angelis, D.; Dye, C.; Eames, K.T.D.; Edmunds, W.J.; Frost, S.D.W.; Funk, S.; et al. Modeling infectious disease dynamics in the complex landscape of global health. Science 2015, 347, aaa4339. [Google Scholar] [CrossRef] [PubMed]
- Fauci, A.; Morens, D. Zika Virus in the Americas—Yet Another Arbovirus Threat. N. Engl. J. Med. 2016, 374, 601–604. [Google Scholar] [CrossRef] [PubMed]
- To, K.K.W.; Chan, J.F.W.; Tsang, A.K.L.; Cheng, V.C.C.; Yuen, K.-Y. Ebola virus disease: A highly fatal infectious disease reemerging in West Africa. Microbes Infect. 2015, 17, 84–97. [Google Scholar] [CrossRef] [PubMed]
- Johnson, B.W.; Goodman, C.H.; Jee, Y.; Featherstone, D.A. Differential diagnosis of Japanese encephalitis virus infections with the inbios JE DetectTM and DEN DetectTM MAC-ELISA kits. Am. J. Trop. Med. Hyg. 2016, 94, 820–828. [Google Scholar] [CrossRef] [PubMed]
- Johnson, B.W.; Russell, B.J.; Goodman, C.H. Laboratory diagnosis of chikungunya virus infections and commercial sources for diagnostic assays. J. Infect. Dis. 2016, 214, S471–S474. [Google Scholar] [CrossRef] [PubMed]
- Golding, C.G.; Lamboo, L.L.; Beniac, D.R.; Booth, T.F. The scanning electron microscope in microbiology and diagnosis of infectious disease. Sci. Rep. 2016, 6, 26516. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Hung, T.; Song, J.D.; He, J.S. Electron microscopy: Essentials for viral structure, morphogenesis and rapid diagnosis. Sci. China Life Sci. 2013, 56, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Bouguelia, S.; Roupioz, Y.; Slimani, S.; Mondani, L.; Casabona, M.G.; Durmort, C.; Vernet, T.; Calemczuk, R.; Livache, T. On-chip microbial culture for the specific detection of very low levels of bacteria. Lab Chip 2013, 13, 4024. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Zhang, S.; Xiang, F.; Wu, D.; Guo, M.; Ge, S.; Li, K.; Ye, X.; Xia, N.; Qian, S. Instrument-free point-of-care molecular diagnosis of H1N1 based on microfluidic convective PCR. Sens. Actuators B Chem. 2017, 243, 738–744. [Google Scholar] [CrossRef]
- Liu, N.; Zou, D.; Dong, D.; Yang, Z.; Ao, D.; Liu, W.; Huang, L. Development of a multiplex loop-mediated isothermal amplification method for the simultaneous detection of Salmonella spp. and Vibrio parahaemolyticus. Sci. Rep. 2017, 7, 45601. [Google Scholar] [CrossRef] [PubMed]
- Bissonnette, L.; Bergeron, M.G. Infectious Disease Management through Point-of-Care Personalized Medicine Molecular Diagnostic Technologies. J. Pers. Med. 2012, 2, 50–70. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Wang, S.; Wang, L.; Li, F.; Pingguan-murphy, B. Advances in paper-based point-of-care diagnostics. Biosens. Bioelectron. 2014, 54, 585–597. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, N.M.; Linnes, J.C.; Fan, A.; Ellenson, C.K.; Pollock, N.R.; Klapperich, C.M. Paper-Based RNA Extraction, in Situ Isothermal Amplification, and Lateral Flow Detection for Low-Cost, Rapid Diagnosis of Influenza A (H1N1) from Clinical Specimens. Anal. Chem. 2015, 87, 7872–7879. [Google Scholar] [CrossRef] [PubMed]
- Su, W.; Gao, X.; Jiang, L.; Qin, J. Microfluidic platform towards point-of-care diagnostics in infectious. J. Chromatogr. A 2015, 1377, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Tay, A.; Pavesi, A.; Yazdi, S.R.; Lim, C.T.; Warkiani, M.E. Advances in microfluidics in combating infectious diseases. Biotechnol. Adv. 2016, 34, 404–421. [Google Scholar] [CrossRef] [PubMed]
- Buonora, S.N.; Passos, S.R.L.; do Carmo, C.N.; Quintela, F.M.; de Oliveira, D.N.R.; dos Santos, F.B.; Hökerberg, Y.H.M.; Nogueira, R.M.R.; Daumas, R.P. Accuracy of clinical criteria and an immunochromatographic strip test for dengue diagnosis in a DENV-4 epidemic. BMC Infect. Dis. 2016, 16, 37. [Google Scholar] [CrossRef] [PubMed]
- Eggerbauer, E.; de Benedictis, P.; Hoffmann, B.; Mettenleiter, T.C.; Schlottau, K.; Ngoepe, E.C.; Sabeta, C.T.; Freuling, C.M.; Müller, T. Evaluation of Six Commercially Available Rapid Immunochromatographic Tests for the Diagnosis of Rabies in Brain Material. PLoS Negl. Trop. Dis. 2016, 10, e0004776. [Google Scholar] [CrossRef] [PubMed]
- Screaton, G.; Mongkolsapaya, J.; Yacoub, S.; Roberts, C. New insights into the immunopathology and control of dengue virus infection. Nat. Rev. Immunol. 2015, 15, 745–759. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Supasa, P.; Wongwiwat, W.; Rouvinski, A.; Barba-Spaeth, G.; Duangchinda, T.; Sakuntabhai, A.; Cao-Lormeau, V.-M.; Malasit, P.; Rey, F.A.; et al. Dengue virus sero-cross-reactivity drives antibody-dependent enhancement of infection with zika virus. Nat. Immunol. 2016, 17, 1102–1108. [Google Scholar] [CrossRef] [PubMed]
- Nan, L.; Jiang, Z.; Wei, X. Emerging microfluidic devices for cell lysis: A review. Lab Chip 2014, 14, 1060–1073. [Google Scholar] [CrossRef] [PubMed]
- Werner, M.; Palankar, R.; Arm, L.; Hovius, R.; Vogel, H. Microfluidic Single-Cell Analysis with Affinity Beads. Small 2015, 1–7. [Google Scholar] [CrossRef]
- Berasaluce, A.; Matthys, L.; Mujika, J.; Anto, M. Bead beating-based continuous flow cell lysis in a microfluidic device. RSC Adv. 2015, 5, 22350–22355. [Google Scholar] [CrossRef]
- Jiang, F.; Chen, J.; Yu, J. Design and application of a microfluidic cell lysis microelectrode chip. Instrum. Sci. Technol. 2015, 44, 223–232. [Google Scholar] [CrossRef]
- Packard, M.; Wheeler, E.; Alocilja, E.; Shusteff, M. Performance Evaluation of Fast Microfluidic Thermal Lysis of Bacteria for Diagnostic Sample Preparation. Diagnostics 2013, 3, 105–116. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cui, D.; Liu, C.; Cai, H. Microfluidic Biochip for Blood Cell Lysis. Chin. J. Anal. Chem. 2006, 34, 1656–1660. [Google Scholar] [CrossRef]
- Cooper, G.M.; Hausman, R.E. The Cell: A Molecular Approach; ASM Press: Washington, DC, USA, 2007. [Google Scholar]
- Yu, Z.; Lu, S.; Huang, Y. Microfluidic Whole Genome Amplification Device for Single Cell Sequencing. Anal. Chem. 2014, 86, 9386–9390. [Google Scholar] [CrossRef] [PubMed]
- Jen, C.P.; Hsiao, J.H.; Maslov, N.A. Single-cell chemical lysis on microfluidic chips with arrays of microwells. Sensors 2012, 12, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Houston, K.; Tucker, M.R.; Chowdhury, J.; Shirley, N.; Little, A. The Plant Cell Wall: A Complex and Dynamic Structure as Revealed by the Responses of Genes under Stress Conditions. Front. Plant Sci. 2016, 7, 984. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.A. Gene Cloning and DNA Analysis: An Introduction; Wiley-Blackwell: Hoboken, NJ, USA, 2011. [Google Scholar]
- Cichova, M.; Proksova, M.; Tothova, L.; Santha, H.; Mayer, V. On-line cell lysis of bacteria and its spores using a microfluidic biochip. Cent. Eur. J. Biol. 2012, 7, 230–240. [Google Scholar]
- Buser, J.; Zhang, X.; Byrnes, S.; Ladd, P.D.; Heiniger, E.K.; Wheeler, M.D.; Bishop, J.D.; Englund, J.A.; Lutz, B.R.; Weigl, B.H.; et al. A disposable chemical heater and dry enzyme preparation for lysis and extraction of DNA and RNA from microorganisms. Anal. Methods 2016, 8, 2880–2886. [Google Scholar] [CrossRef]
- Strohmeier, O.; Keil, S.; Kanat, B.; Patel, P.; Niedrig, M.; Weidmann, M.; Hufert, F.; Drexler, J.; Zengerle, R.; von Stetten, F. Automated nucleic acid extraction from whole blood, B. subtilis, E. coli, and Rift Valley fever virus on a centrifugal microfluidic LabDisk. RSC Adv. 2015, 5, 32144–32150. [Google Scholar] [CrossRef]
- Lee, H.J.; Kim, J.-H.; Lim, H.K.; Cho, E.C.; Huh, N.; Ko, C.; Park, J.C.; Choi, J.-W.; Lee, S.S. Electrochemical cell lysis device for DNA extraction. Lab Chip 2010, 10, 626–633. [Google Scholar] [CrossRef] [PubMed]
- Jha, S.K.; Ra, G.-S.S.; Joo, G.-S.S.; Kim, Y.-S.S. Electrochemical cell lysis on a miniaturized flow-through device. Curr. Appl. Phys. 2009, 9, e301–e303. [Google Scholar] [CrossRef]
- Zelenin, S.; Hansson, J.; Ardabili, S.; Ramachandraiah, H.; Brismar, H.; Russom, A. Microfluidic-based isolation of bacteria from whole blood for sepsis diagnostics. Biotechnol. Lett. 2015, 37, 825–830. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, A.; Autebert, J.; Delamarche, E.; Kaigala, G. V Selective local lysis and sampling of live cells for nucleic acid analysis using a microfluidic probe. Sci. Rep. 2016, 6, 29579. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cui, D.F.; Liu, C.C. On-line cell lysis and DNA extraction on a microfluidic biochip fabricated by microelectromechanical system technology. Electrophoresis 2008, 29, 1844–1851. [Google Scholar] [CrossRef] [PubMed]
- Karle, M.; Miwa, J.; Czilwik, G.; Auwärter, V.; Roth, G.; Zengerle, R.; von Stetten, F. Continuous microfluidic DNA extraction using phase-transfer magnetophoresis. Lab Chip 2010, 10, 3284–3290. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.; Yoon, Y.; Song, M.; Im, S.; Kim, S.; Lee, J.; Yang, S. Handheld mechanical cell lysis chip with ultra-sharp silicon nano-blade arrays for rapid intracellular protein extraction. Lab Chip 2010, 10, 1442–1446. [Google Scholar] [CrossRef] [PubMed]
- Caffiyar, M.Y.; Ho, E.T.W.; Hussain, I.; Hamid, N.H.B. Microfluidic Platform for Cell Isolation and Manipulation Based on Cell Properties. Micromachines 2017, 8, 15. [Google Scholar]
- Choi, J.; Hyun, J.; Yang, S. On-Chip Extraction of Intracellular Molecules in White Blood Cells from Whole Blood. Sci. Rep. 2015, 5, 15167. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Hong, J.W.; Kim, D.P.; Shin, J.H.; Park, I. Nanowire-integrated microfluidic devices for facile and reagent-free mechanical cell lysis. Lab Chip 2012, 12, 2914–2921. [Google Scholar] [CrossRef] [PubMed]
- Mahalanabis, M.; Al-Muayad, H.; Kulinski, M.D.; Altman, D.; Klapperich, C.M. Cell lysis and DNA extraction of gram-positive and gram-negative bacteria from whole blood in a disposable microfluidic chip. Lab Chip 2009, 9, 2811–2817. [Google Scholar] [CrossRef] [PubMed]
- Burke, J.M.; Smela, E. A novel surface modification technique for forming porous polymer monoliths in Poly(dimethylsiloxane). Biomicrofluidics 2012, 6, 016506. [Google Scholar] [CrossRef] [PubMed]
- Saad Aly, M.A.; Nguon, O.; Gauthier, M.; Yeow, J.T.W. Antibacterial porous polymeric monolith columns with amphiphilic and polycationic character on cross-linked PMMA substrates for cell lysis applications. RSC Adv. 2013, 3, 24177. [Google Scholar] [CrossRef]
- Aly, M.A.S.; Aly, S.; Gauthier, M.; Yeow, J. Lysis of gram-positive and gram-negative bacteria by antibacterial porous polymeric monolith formed in microfluidic biochips for sample preparation. Anal. Bioanal. Chem. 2014, 406, 5977–5987. [Google Scholar] [CrossRef] [PubMed]
- Geissler, M.; Beauregard, J.A.; Charlebois, I.; Isabel, S.; Normandin, F.; Voisin, B.; Boissinot, M.; Bergeron, M.G.; Veres, T. Extraction of nucleic acids from bacterial spores using bead-based mechanical lysis on a plastic chip. Eng. Life Sci. 2011, 11, 174–181. [Google Scholar] [CrossRef]
- Cheng, Y.; Wang, Y.; Wang, Z.; Huang, L.; Bi, M.; Xu, W. A mechanical cell disruption microfluidic platform based an on-chip micropump. Biomicrofluidics 2017, 24112, 11–14. [Google Scholar] [CrossRef] [PubMed]
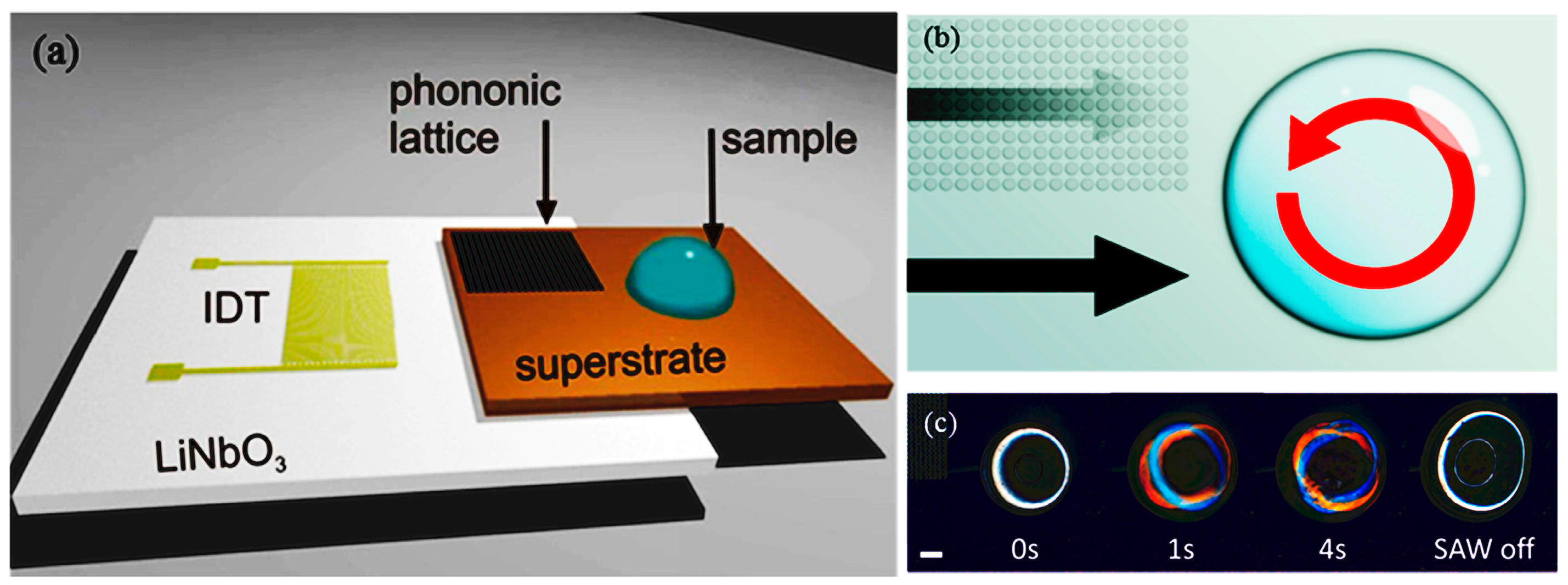
- Salehi-Reyhani, A.; Gesellchen, F.; Mampallil, D.; Wilson, R.; Reboud, J.; Ces, O.; Willison, K.R.; Cooper, J.M.; Klug, D.R. Chemical-free lysis and fractionation of cells by use of surface acoustic waves for sensitive protein assays. Anal. Chem. 2015, 87, 2161–2169. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Chen, Y.; Farooq, U.; Xuan, W.; Jin, H.; Dong, S.; Luo, J. Ultrafast chemical-free cell lysis by high speed stream collision induced by surface acoustic waves. Appl. Phys. Lett. 2017, 110, 143504. [Google Scholar] [CrossRef]
- Taller, D.; Richards, K.; Slouka, Z.; Senapati, S.; Hill, R.; Go, D.B.; Chang, H.-C. On-chip surface acoustic wave lysis and ion-exchange nanomembrane detection of exosomal RNA for pancreatic cancer study and diagnosis. Lab Chip 2015, 15, 1656–1666. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.-H.; Hung, L.-Y.; Lee, G.-B. Continuous nucleus extraction by optically-induced cell lysis on a batch-type microfluidic platform. Lab Chip 2016, 16, 1447–1456. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.H.; Bhunia, A.K.; Lu, C. A microfluidic flow-through device for high throughput electrical lysis of bacterial cells based on continuous dc voltage. Biosens. Bioelectron. 2006, 22, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.W.; Cho, Y. A continuous electrical cell lysis device using a low dc voltage for a cell transport and rupture. Sens. Actuators B Chem. 2007, 124, 84–89. [Google Scholar] [CrossRef]
- Gabardo, C.M.; Kwong, A.M.; Soleymani, L. Rapidly prototyped multi-scale electrodes to minimize the voltage requirements for bacterial cell lysis. Analyst 2015, 140, 1599–1608. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.; Shahid, A.; Kuryllo, K.; Li, Y.; Deen, M.; Selvaganapathy, P. Electrophoretic Concentration and Electrical Lysis of Bacteria in a Microfluidic Device Using a Nanoporous Membrane. Micromachines 2017, 8, 45. [Google Scholar] [CrossRef]
- Escobedo, C.; Bürgel, S.C.; Kemmerling, S.; Sauter, N.; Braun, T.; Hierlemann, A. On-chip lysis of mammalian cells through a handheld corona device. Lab Chip 2015, 15, 2990–2997. [Google Scholar] [CrossRef] [PubMed]
- Talebpour, A.; Maaskant, R.; Khine, A.A.; Alavie, T. Use of surface enhanced blocking (SEB) electrodes for microbial cell lysis in flow-through devices. PLoS ONE 2014, 9, e102707. [Google Scholar] [CrossRef] [PubMed]
- Ameri, S.K.; Singh, P.K.; Sonkusale, S. Utilization of graphene electrode in transparent microwell arrays for high throughput cell trapping and lysis. Biosens. Bioelectron. 2014, 61, 625–630. [Google Scholar] [CrossRef] [PubMed]
- Witte, C.; Kremer, C.; Chanasakulniyom, M.; Reboud, J.; Wilson, R.; Cooper, J.M.; Neale, S.L. Spatially selecting a single cell for lysis using light-induced electric fields. Small 2014, 10, 3026–3031. [Google Scholar] [CrossRef] [PubMed]
- Geng, T.; Bao, N.; Sriranganathanw, N.; Li, L.; Lu, C. Genomic DNA extraction from cells by electroporation on an integrated microfluidic platform. Anal. Chem. 2013, 84, 9632–9639. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Bryson, B.D.; Sun, C.; Fortune, S.M.; Lu, C. RNA Extraction from a Mycobacterium under Ultrahigh Electric Field Intensity in a Microfluidic Device. Anal. Chem. 2016, 88, 5053–5057. [Google Scholar] [CrossRef] [PubMed]
- Baek, S.; Min, J.; Park, J.-H. Wireless induction heating in a microfluidic device for cell lysis. Lab Chip 2010, 10, 909–917. [Google Scholar] [CrossRef] [PubMed]
- Tsougeni, K.; Papadakis, G.; Gianneli, M.; Grammoustianou, A.; Constantoudis, V.; Dupuy, B.; Petrou, P.S.; Kakabakos, S.E.; Tserepi, A.; Gizeli, E.; et al. Plasma nanotextured polymeric lab-on-a-chip for highly efficient bacteria capture and lysis. Lab Chip 2015, 1, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Reinholt, S.J.; Baeumner, A.J. Microfluidic Isolation of Nucleic Acids. Angew. Chem. Int. Ed. 2014, 53, 13988–14001. [Google Scholar] [CrossRef] [PubMed]
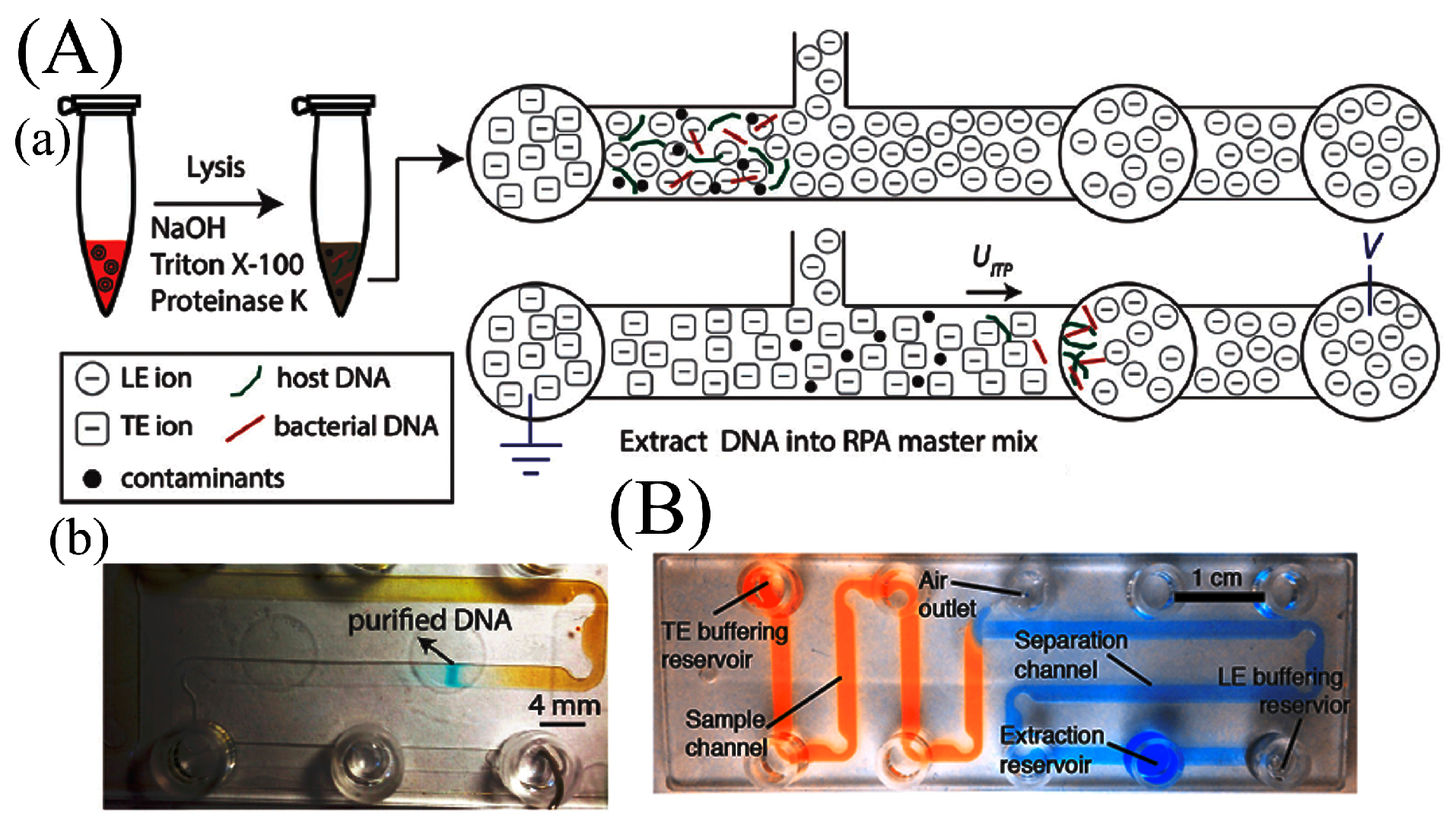
- Eid, C.; Santiago, J.G. Assay for Listeria monocytogenes cells in whole blood using isotachophoresis and recombinase polymerase amplification. Analyst 2017, 142, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cui, D.; Sun, J.; Zhang, L.; Li, H. Microdevice-based DNA Extraction Method Using Green Reagent. Micro-Nano Technol. XIV 2013, 565, 1111–1115. [Google Scholar] [CrossRef]
- Hagan, K.A.; Reedy, C.R.; Uchimoto, M.L.; Basu, D.; Engel, A.; Landers, J.P. An integrated, valveless system for microfluidic purification and reverse transcription-PCR amplification of RNA for detection of infectious agents. Lab Chip 2011, 11, 957–961. [Google Scholar] [CrossRef] [PubMed]
- Hwang, K.; Kwon, S.H.; Jung, S.; Namkoong, K.; Jung, W.; Kim, J.; Suh, K.; Huh, N. Solid Phase DNA Extraction with a Flexible Bead-Packed Microfluidic Device to Detect Methicillin-Resistant Staphylococcus aureus in Nasal Swabs. Anal. Chim. Acta 2012, 84, 7912–7918. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.-J.; Lien, K.-Y.; Weng, C.-Y.; Shin, J.-W.; Chang, T.-Y.; Lee, G.-B. Magnetic-bead-based microfluidic system for ribonucleic acid extraction and reverse transcription processes. Biomed. Microdevices 2009, 11, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.-H.; Cheng, L.; Wang, C.; Ling, W.; Wang, S.; Lee, G.-B. An integrated chip capable of performing sample pretreatment and nucleic acid amplification for HIV-1 detection. Biosens. Bioelectron. 2013, 41, 484–491. [Google Scholar] [CrossRef] [PubMed]
- Han, N.; Shin, J.H.; Han, K.-H. An on-chip RT-PCR microfluidic device, that integrates mRNA extraction, cDNA synthesis, and gene amplification. RSC Adv. 2014, 4, 9160. [Google Scholar] [CrossRef]
- Adams, N.M.; Bordelon, H.; Wang, K.-K.A.; Albert, L.E.; Wright, D.W.; Haselton, F.R. Comparison of three magnetic bead surface functionalities for RNA extraction and detection. ACS Appl. Mater. Interfaces 2015, 7, 6062–6069. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Mauk, M.G. An integrated, cellulose membrane-based PCR chamber. Microsyst. Technol. 2014, 21, 841–850. [Google Scholar] [CrossRef]
- Liu, C.; Geva, E.; Mauk, M.; Qiu, X.; Abrams, W. An isothermal amplification reactor with an integrated isolation membrane for point-of-care detection of infectious diseases. Analyst 2011, 136, 2069–2076. [Google Scholar] [CrossRef] [PubMed]
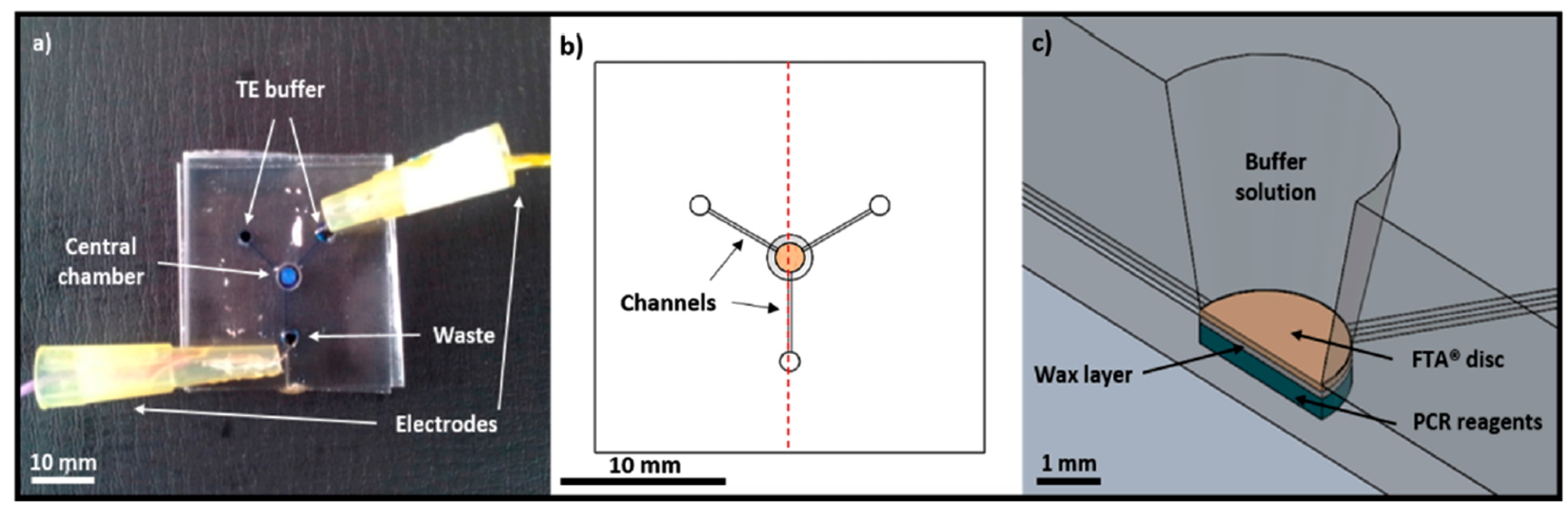
- Wimbles, R.; Melling, L.; Shaw, K. Combining Electro-Osmotic Flow and FTA® Paper for DNA Analysis on Microfluidic Devices. Micromachines 2016, 7, 119. [Google Scholar] [CrossRef]
- Rogacs, A.; Marshall, L.A.; Santiago, J.G. Purification of nucleic acids using isotachophoresis. J. Chromatogr. A 2014, 1335, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Smejkal, P.; Bottenus, D.; Breadmore, M.C.; Guijt, R.M.; Ivory, C.F.; Foret, F.; Macka, M. Microfluidic isotachophoresis: A review. Electrophoresis 2013, 34, 1493–1509. [Google Scholar] [CrossRef] [PubMed]
- Boček, P.; Deml, M.; Janák, J. Instrumentation for high-speed isotachophoresis. J. Chromatogr. A 1975, 106, 283–290. [Google Scholar] [CrossRef]
- Kondratova, V.; Serd’uk, O.; Shelepov, V.; Lichtenstein, A. Concentration and isolation of DNA from biological fluids by agarose gel isotachophoresis. Biotechniques 2005, 39, 695–699. [Google Scholar] [CrossRef] [PubMed]
- Kondratova, V.; Serdyuk, O. Counterflow Isotachophoresis as a Method of Concentration and Isolation of DNA from Biological Fluids. Dokl. Biochem. Biophys. 2005, 402, 268–271. [Google Scholar] [CrossRef]
- Marshall, L.; Han, C.; Santiago, J. Extraction of DNA from malaria-infected erythrocytes using isotachophoresis. Anal. Chem. 2011, 9715–9718. [Google Scholar] [CrossRef] [PubMed]
- Schoch, R.B.; Ronaghi, M.; Santiago, J.G. Rapid and selective extraction, isolation, preconcentration, and quantitation of small RNAs from cell lysate using on-chip isotachophoresis. Lab Chip 2009, 9, 2145–2152. [Google Scholar] [CrossRef] [PubMed]
- Bercovici, M.; Kaigala, G.V.; Mach, K.E.; Han, C.M.; Liao, J.C.; Santiago, J.G. Rapid Detection of Urinary Tract Infections Using Isotachophoresis and Molecular Beacons. Anal. Chem. 2011, 83, 4110–4117. [Google Scholar] [CrossRef] [PubMed]
- Rogacs, A.; Qu, Y.; Santiago, J.G. Bacterial RNA Extraction and Purification from Whole Human Blood Using Isotachophoresis. Anal. Chem. 2012, 84, 5858–5863. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.L.; Marshall, L.A.; Babikian, S.; Han, C.M.; Santiago, J.G.; Bachman, M. A Printed Circuit Board Based Microfluidic System for Point-of-Care Diagnostics Applications. In Proceedings of the 15th International Conference on Miniaturized Systems for Chemistry and Life Sciences, Seattle, WA, USA, 2–6 October 2011; pp. 1819–1821. [Google Scholar]
- Marshall, L.A.; Rogacs, A.; Meinhart, C.D.; Santiago, J.G. An injection molded microchip for nucleic acid purification from 25 microliter samples using isotachophoresis. J. Chromatogr. A 2014, 1331, 139–142. [Google Scholar] [CrossRef] [PubMed]
- Shintaku, H.; Nishikii, H.; Marshall, L.L.A.; Kotera, H.; Santiago, J.G. On-chip separation and analysis of RNA and DNA from single cells. Anal. Chem. 2014, 86, 1953–1957. [Google Scholar] [CrossRef] [PubMed]
- Kuriyama, K.; Shintaku, H.; Santiago, J.G. Isotachophoresis for fractionation and recovery of cytoplasmic RNA and nucleus from single cells. Electrophoresis 2015, 36, 1658–1662. [Google Scholar] [CrossRef] [PubMed]
- LIopis, S.P.; Brugarolas, J. Simultaneous isolation of high-quality DNA, RNA, miRNA and proteins from tissues for genomic applications. Nat. Protoc. 2013, 8, 2240–2255. [Google Scholar] [CrossRef] [PubMed]
- Dey, S.S.; Kester, L.; Spanjaard, B.; Bienko, M.; van Oudenaarden, A. Integrated genome and transcriptome sequencing of the same cell. Nat. Biotechnol. 2015, 33, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Jackman, J.A.; Linardy, E.; Yoo, D.; Seo, J.; Ng, W.B.; Klemme, D.J.; Wittenberg, N.J.; Oh, S.H.; Cho, N.J. Plasmonic Nanohole Sensor for Capturing Single Virus-Like Particles toward Virucidal Drug Evaluation. Small 2016, 12, 1159–1166. [Google Scholar] [CrossRef] [PubMed]
- Henihan, G.; Schulze, H.; Corrigan, D.K.; Giraud, G.; Terry, J.G.; Hardie, A.; Campbell, C.J.; Walton, A.J.; Crain, J.; Pethig, R.; et al. Label- and amplification-free electrochemical detection of bacterial ribosomal RNA. Biosens. Bioelectron. 2016, 81, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-M.; Chang, W.-H.; Wang, C.-H.; Wang, J.-H.; Mai, J.D.; Lee, G.-B. Nucleic acid amplification using microfluidic systems. Lab Chip 2013, 13, 1225–1242. [Google Scholar] [CrossRef] [PubMed]
- Mullis, K.; Faloona, F.; Scharf, S.; Saiki, R.; Horn, G.; Erlich, H. Specific enzymatic amplification of DNA in vitro: The polymerase chain reaction. Cold Spring Harb. Symp. Quant. Biol. 1986, 51, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, V.E.; Kokkoris, G.; Kefala, I.N.; Tserepi, A. Comparison of continuous-flow and static chamber μPCR devices through a computational study: The potential of flexible polymeric substrates. Microfluid. Nanofluid. 2015, 19, 867–882. [Google Scholar] [CrossRef]
- Northrup, M.; Ching, M.; White, R.; Watson, R. DNA amplification in a microfabricated reaction chamber. In Proceedings of the 7th International Conference on Solid-State Sensors and Actuators (Transducers ’93), Yokohama, Japan, 7–10 June 1993; pp. 924–926. [Google Scholar]
- Houssin, T.; Cramer, J.; Grojsman, R.; Bellahsene, L.; Colas, G.; Moulet, H.; Minnella, W.; Pannetier, C.; Leberre, M.; Plecis, A.; et al. Ultrafast, sensitive and large-volume on-chip real-time PCR for the molecular diagnosis of bacterial and viral infections. Lab Chip 2016, 16, 1401–1411. [Google Scholar] [CrossRef] [PubMed]
- Farrar, J.S.; Wittwer, C.T. Extreme PCR: Efficient and specific DNA amplification in 15–60 s. Clin. Chem. 2015, 61, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Le Roux, D.; Root, B.E.; Reedy, C.R.; Hickey, A.; Scott, O.N.; Bienvenue, J.M.; Landers, J.P.; Chassagne, L.; Mazancourt, P. De DNA Analysis Using an Integrated Microchip for Multiplex PCR Amplification and Electrophoresis for Reference Samples. Anal. Chem. 2014, 86, 8192–8199. [Google Scholar] [CrossRef] [PubMed]
- Pak, N.; Saunders, D.C.; Phaneuf, C.R.; Forest, C.R. Plug-and-play, infrared, laser-mediated PCR in a microfluidic chip. Biomed. Microdevices 2012, 14, 427–433. [Google Scholar] [CrossRef] [PubMed]
- Lounsbury, J.A.; Karlsson, A.; Miranian, D.C.; Cronk, S.M.; Nelson, D.A.; Li, J.; Haverstick, D.M.; Kinnon, P.; Saul, J.; Landers, J.P. From sample to PCR product in under 45 min: A polymeric integrated microdevice for clinical and forensic DNA analysis. Lab Chip 2013, 13, 1384–1393. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Zhang, M.; Liu, X.; Sharma, A.; Ding, X. A Point-of-Need infrared mediated PCR platform with compatible lateral flow strip for HPV detection. Biosens. Bioelectron. 2017, 96, 213–219. [Google Scholar] [CrossRef] [PubMed]
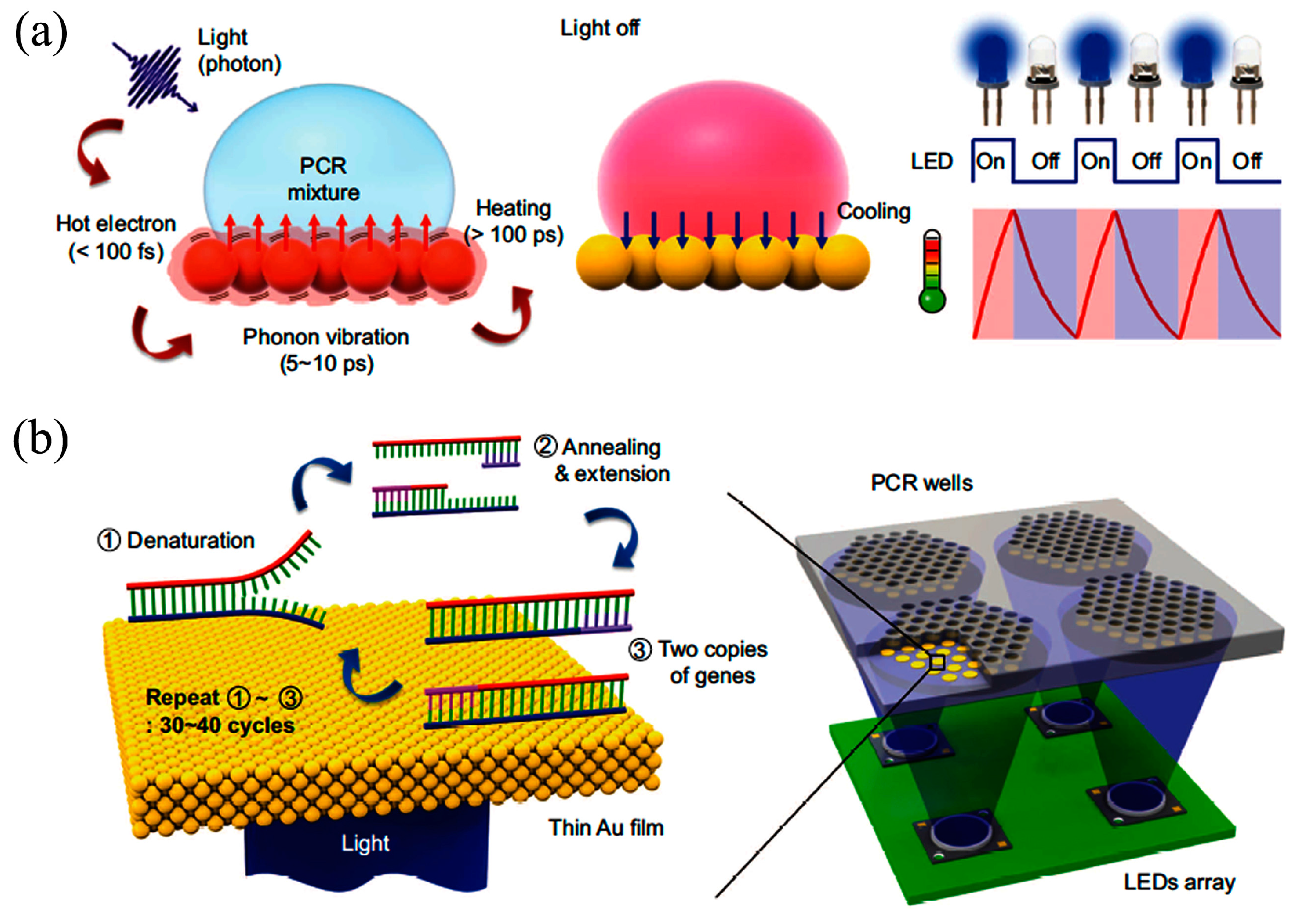
- Son, J.H.; Cho, B.; Hong, S.; Lee, S.H.; Hoxha, O.; Haack, A.J.; Lee, L.P. Ultrafast photonic PCR. Light Sci. Appl. 2015, 4, e280. [Google Scholar] [CrossRef]
- Phaneuf, C.R.; Pak, N.; Saunders, D.C.; Holst, G.L.; Birjiniuk, J.; Nagpal, N.; Culpepper, S.; Popler, E.; Shane, A.L.; Jerris, R.; et al. Thermally multiplexed polymerase chain reaction. Biomicrofluidics 2015, 9, 044117. [Google Scholar] [CrossRef] [PubMed]
- Manage, D.P.; Morrissey, Y.C.; Stickel, A.J.; Lauzon, J.; Atrazhev, A.; Acker, J.P.; Pilarski, L.M. On-chip PCR amplification of genomic and viral templates in unprocessed whole blood. Microfluid. Nanofluid. 2011, 10, 697–702. [Google Scholar] [CrossRef]
- Hilton, J.P.; Nguyen, T.; Barbu, M.; Pei, R.; Stojanovic, M.; Lin, Q. Bead-based polymerase chain reaction on a microchip. Microfluid. Nanofluid. 2012, 13, 749–760. [Google Scholar] [CrossRef]
- Sun, L.; Dhumpa, R.; Bang, D.D.; Hogberg, J.; Handberg, K.; Wolff, A. A lab-on-a-chip device for rapid identification of avian influenza viral RNA by soild-phase PCR. Lab Chip 2011, 11, 1457–1463. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ostlund, E.N.; Jun, Y.; Nie, F.P.; Li, Y.G.; Johnson, D.J.; Lin, R.; Li, Z.G. Combining reverse-transcription multiplex PCR and microfluidic electrophoresis to simultaneously detect seven mosquito-transmitted zoonotic encephalomyelitis viruses. Vet. J. 2016, 212, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Dean, D.; Turingan, R.S.; Thomann, H.; Zolotova, A.; Rothschild, J.; Joseph, S.J.; Read, T.D.; Tan, E.; Selden, R.F. A Multiplexed Microfluidic PCR Assay for Sensitive and Specific Point-of-Care Detection of Chlamydia trachomatis. PLoS ONE 2012, 7, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Nakano, H.; Matsuda, K.; Yohda, M.; Nagamune, T.; Endo, I.; Yamane, T. High speed polymerase chain reaction in constant flow. Biosci. Biotechnol. Biochem. 1994, 58, 349–352. [Google Scholar] [CrossRef] [PubMed]
- Kopp, M.U.; Mello, A.J.; Manz, A. Chemical Amplification: Continuous-Flow PCR on a Chip. Science 1998, 280, 1046–1048. [Google Scholar] [CrossRef] [PubMed]
- Aboud, M.; Oh, H.H.; Mccord, B. Rapid direct PCR for forensic genotyping in under 25 min. Electrophoresis 2013, 34, 1539–1547. [Google Scholar] [CrossRef] [PubMed]
- Ha, B.H.; Lee, K.S.; Destgeer, G.; Park, J.; Choung, J.S.; Jung, J.H.; Shin, J.H.; Sung, H.J. Acoustothermal heating of polydimethylsiloxane microfluidic system. Sci. Rep. 2015, 5, 11851. [Google Scholar] [CrossRef] [PubMed]
- Brunklaus, S.; Hansen-hagge, T.E.; Erwes, J.; Jung, M.; Latta, D.; Strobach, X.; Winkler, C.; Ritzi-lehnert, M.; Drese, K.S. Fast nucleic acid amplification for integration in point-of-care applications. Electrophoresis 2012, 33, 3222–3228. [Google Scholar] [CrossRef] [PubMed]
- Nie, J.; Zhao, Y.; Peng, N. Multichannel oscillatory-flow PCR micro-fluidic chip with controllable temperature gradient. Microsyst. Technol. 2014, 21, 41–48. [Google Scholar] [CrossRef]
- Wu, W.; Loan, T.; Yoon, N.L. Flow-through PCR on a 3D qiandu-shaped polydimethylsiloxane (PDMS) microdevice employing a single heater: Toward microscale multiplex PCR. Analyst 2012, 137, 2069–2076. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, H.; Saito, M.; Shibuya, S.; Tsuji, K.; Miyagawa, N.; Yamanaka, K.; Tamiya, E. On-chip quantitative detection of pathogen genes by autonomous microfluidic PCR platform. Biosens. Bioelectron. 2015, 74, 725–730. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, H.; Saito, M.; Tsuji, K.; Yamanaka, K.; Hoa, L.Q.; Tamiya, E. Self-propelled continuous-flow PCR in capillary-driven microfluidic device: Microfluidic behavior and DNA amplification. Sens. Actuators B Chem. 2015, 206, 303–310. [Google Scholar] [CrossRef]
- Wu, W.; Kang, K.-T.; Lee, Y.N. Bubble-free on-chip continuous-flow polymerase chain reaction: Concept and application. Analyst 2011, 136, 2287–2293. [Google Scholar] [CrossRef] [PubMed]
- Shu, B.; Zhang, C.; Xing, D. Highly sensitive identification of foodborne pathogenic Listeria monocytogenes using single-phase continuous-flow nested PCR microfluidics with on-line fluorescence detection. Microfluid. Nanofluid. 2013, 15, 161–172. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, H.; Xing, D. Multichannel oscillatory-flow multiplex PCR microfluidics for high-throughput and fast detection of foodborne bacterial pathogens. Biomed. Microdevices 2011, 13, 885–897. [Google Scholar] [CrossRef] [PubMed]
- Compton, J. Nucleic acid sequence-based amplification. Nature 1991, 354, 56–58. [Google Scholar] [CrossRef] [PubMed]
- Hataoka, Y.; Zhang, L.; Mori, Y.; Tomita, N.; Notomi, T.; Baba, Y. Analysis of Specific Gene by Integration of Isothermal Amplification and Electrophoresis on Poly(methyl methacrylate) Microchips. Anal. Chem. 2004, 76, 3689–3693. [Google Scholar] [CrossRef] [PubMed]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N. A DNA detection using recombination proteins. PLoS Biol. 2006, 4, e204. [Google Scholar] [CrossRef] [PubMed]
- Vincent, M.; Xu, Y.; Kong, H. Helicase-dependent isothermal DNA amplification. EMBO Rep. 2004, 5, 795–800. [Google Scholar] [CrossRef] [PubMed]
- Uroda, A.K.; Shigaki, Y.I.; Ilsson, M.N.; Ato, K.S.; Ato, K.S. Microfluidics-based in situ Padloc /Rolling Circle Amplification System for Counting Single DNA Molecules in a Cell. Anal. Sci. 2014, 30, 1107–1112. [Google Scholar]
- Murakami, T.; Sumaoka, J.; Komiyama, M. Sensitive isothermal detection of nucleic-acid sequence by primer generation-rolling circle amplification. Nucleic Acids Res. 2009, 37, e19. [Google Scholar] [CrossRef] [PubMed]
- Walker, G.T.; Fraiser, M.S.; Schram, J.L.; Little, M.C.; Nadeau, J.G.; Malinowski, D.P. Strand displacement amplification-an isothermal, in vitro DNA amplification technique. Nucleic Acids Res. 1992, 20, 1691–1696. [Google Scholar] [CrossRef] [PubMed]
- Jonas, V.; Alden, M.J.; Curry, J.I.; Kamisango, K.; Knott, C.A.; Lankford, R.; Wolfe, J.M.; Moore, D.F. Detection and identification of Mycobacterium tuberculosis directly from sputum sediments by amplification of rRNA. J. Clin. Microbiol. 1993, 31, 2410–2416. [Google Scholar] [PubMed]
- Stary, A.; Schuh, E.; Kerschbaumer, M. Performance of Transcription-Mediated Amplification and Ligase Chain Reaction Assays for Detection of Chlamydial Infection in Urogenital Samples Obtaine. J. Clin. Microbiol. 1998, 36, 2666–2670. [Google Scholar] [PubMed]
- Urdea, M.S. Branched DNA Signal Amplification. BioTechnology 1994, 12, 926–928. [Google Scholar] [CrossRef] [PubMed]
- Collins, M.L.; Irvine, B.; Tyner, D.; Fine, E.; Zayati, C.; Chang, C.A.; Horn, T.; Ahle, D.; Detmer, J.; Shen, L.P.; et al. A branched DNA signal amplification assay for quantification of nucleic acid targets below 100 molecules/mL. Nucleic Acids Res. 1997, 25, 2979–2984. [Google Scholar] [CrossRef] [PubMed]
- Kurn, N.; Chen, P.; Heath, J.D.; Kopf-Sill, A.; Stephens, K.M.; Wang, S. Novel isothermal, linear nucleic acid amplification systems for highly multiplexed applications. Clin. Chem. 2005, 51, 1973–1981. [Google Scholar] [CrossRef] [PubMed]
- García-Elorriaga, G.; del Rey-Pineda, G. Practical and Laboratory Diagnosis of Tuberculosis; Springer International Publishing: Cham, Switzerland, 2015. [Google Scholar]
- Tsaloglou, M.-N.; Bahi, M.M.; Waugh, E.M.; Morgan, H.; Mowlem, M. On-chip real-time nucleic acid sequence-based amplification for RNA detection and amplification. Anal. Methods 2011, 3, 2127. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Rafati, A.; Gill, P. Microfluidic method for rapid turbidimetric detection of the DNA of Mycobacterium tuberculosis using loop-mediated isothermal amplification in capillary tubes. Microchim. Acta 2014, 182, 523–530. [Google Scholar] [CrossRef]
- Fang, X.; Liu, Y.; Kong, J.; Jiang, X. Loop-mediated isothermal amplification integrated on microfluidic chips for point-of-care quantitative detection of pathogens. Anal. Chem. 2010, 82, 3002–3006. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Giri, H.; Hewlett, I.K. Point of Care Technologies for HIV. AIDS Res. Treat. 2014, 2014, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Zhang, T.; Chen, F.; Wang, L.; Bai, X. Advances in loop-mediated isothermal ampli fi cation: Integrated with several point-of-care. Anal. Methods 2014, 6, 7585–7589. [Google Scholar] [CrossRef]
- Liu, C.; Mauk, M.G.; Bau, H.H. A disposable, integrated loop-mediated isothermal amplification cassette with thermally actuated valves. Microfluid. Nanofluid. 2011, 11, 209–220. [Google Scholar] [CrossRef]
- Tourlousse, D.M.; Ahmad, F.; Stedtfeld, R.D.; Seyrig, G.; Tiedje, J.M.; Hashsham, S.A. A polymer microfluidic chip for quantitative detection of multiple water- and foodborne pathogens using real-time fluorogenic loop-mediated isothermal amplification. Biomed. Microdevices 2012, 14, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Lien, K.; Wang, T.; Chen, T.; Lee, G.-B. An integrated microfluidic loop-mediated-isothermal-amplification system for rapid sample pre-treatment and detection of viruses. Biosens. Bioelectron. 2011, 26, 2045–2052. [Google Scholar] [CrossRef] [PubMed]
- Borysiak, M.; Kimura, K.; Posner, J. NAIL: Nucleic Acid detection using Isotachophoresis and Loop-mediated isothermal amplification. Lab Chip 2015, 15, 1697–1707. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, N.M.; Wong, W.S.; Liu, L.; Dewar, R.; Klapperich, C.M. A fully integrated paperfluidic molecular diagnostic chip for the extraction, amplification, and detection of nucleic acids from clinical samples. Lab Chip 2016, 16, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Michelle Wong Tzeling, J.; Yean Yean, C. A shelf-stable fluorogenic isothermal amplification assay for the detection of Burkholderia pseudomallei. Analyst 2016, 141, 1246–1249. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Liu, P.; Zhao, X.; Du, W.; Feng, X.; Liu, B.F. A self-contained microfluidic in-gel loop-mediated isothermal amplification for multiplexed pathogen detection. Sens. Actuators B Chem. 2017, 239, 1–8. [Google Scholar] [CrossRef]
- Lee, D.; Kim, Y.T.; Lee, J.W.; Kim, D.H.; Seo, T.S. An integrated direct loop-mediated isothermal amplification microdevice incorporated with an immunochromatographic strip for bacteria detection in human whole blood and milk without a sample preparation step. Biosens. Bioelectron. 2016, 79, 273–279. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, F.; Stedtfeld, R.D.; Waseem, H.; Williams, M.R.; Cupples, A.M.; Tiedje, J.M.; Hashsham, S.A. Most probable number—Loop mediated isothermal amplification (MPN-LAMP) for quantifying waterborne pathogens in <25 min. J. Microbiol. Methods 2017, 132, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.-J.; Wang, L.; Chen, J.; Wang, R.; Shi, Y.; Li, C.; Zhang, D.; Yan, X.; Zhang, Y. Development and evaluation of a real-time fluorogenic loop-mediated isothermal amplification assay integrated on a microfluidic disc chip (on-chip LAMP) for rapid and simultaneous detection of ten pathogenic bacteria in aquatic animals. J. Microbiol. Methods 2014, 104, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Kong, J. A multiplexed nucleic acid microsystem for point-of-care detection of HIV co-infection with MTB and PCP. Talanta 2013, 117, 532–535. [Google Scholar] [CrossRef] [PubMed]
- Tsaloglou, M.-N.; Watson, R.J.; Rushworth, C.M.; Zhao, Y.; Niu, X.; Sutton, J.M.; Morgan, H. Real-time microfluidic recombinase polymerase amplification for the toxin B gene of Clostridium difficile on a SlipChip platform. Analyst 2015, 140, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Lillis, L.; Siverson, J.; Lee, A.; Cantera, J.; Parker, M.; Piepenburg, O.; Lehman, D.A.; Boyle, D.S. Factors influencing Recombinase polymerase amplification (RPA) assay outcomes at point of care. Mol. Cell. Probes 2016, 30, 74–78. [Google Scholar] [CrossRef] [PubMed]
- Rohrman, B.A.; Richards-kortum, R.R. A Paper and Plastic Device for Performing Recombinase Polymerase Amplification of HIV DNA. Lab Chip 2012, 12, 3082–3088. [Google Scholar] [CrossRef] [PubMed]
- Kersting, S.; Rausch, V.; Bier, F.F.; von Nickisch-Rosenegk, M. Multiplex isothermal solid-phase recombinase polymerase amplification for the specific and fast DNA-based detection of three bacterial pathogens. Mikrochim. Acta 2014, 181, 1715–1723. [Google Scholar] [CrossRef] [PubMed]
- Shen, F.; Davydova, E.K.; Du, W.; Kreutz, J.E.; Piepenburg, O.; Ismagilov, R.F. Digital Isothermal Quantification of Nucleic Acids via Simultaneous Chemical Initiation of Recombinase Polymerase Amplification Reactions on SlipChip. Anal. Chem. 2011, 83, 3533–3540. [Google Scholar] [CrossRef] [PubMed]
- Zaghloul, H.; El-Shahat, M. Recombinase polymerase amplification as a promising tool in hepatitis C virus diagnosis. World J. Hepatol. 2014, 6, 916–922. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Hoshika, S.; Hutter, D.; Bradley, K.M.; Benner, S. A Recombinase-based isothermal amplification of nucleic acids with self-avoiding molecular recognition systems (SAMRS). Chembiochem 2014, 15, 2268–2274. [Google Scholar] [CrossRef] [PubMed]
- Kersting, S.; Rausch, V.; Bier, F.F.; von Nickisch-Rosenegk, M. Rapid detection of Plasmodium falciparum with isothermal recombinase polymerase amplification and lateral flow analysis. Malar. J. 2014, 13, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Rohrman, B.; Richards-Kortum, R. Inhibition of recombinase polymerase amplification by background DNA: A lateral flow-based method for enriching target DNA. Anal. Chem. 2015, 87, 1963–1967. [Google Scholar] [CrossRef] [PubMed]
- Kaprou, G.D.; Papadakis, G.; Papageorgiou, D.P.; Kokkoris, G.; Papadopoulos, V.; Kefala, I.; Gizeli, E.; Tserepi, A. Miniaturized devices for isothermal DNA amplification addressing DNA diagnostics. Microsyst. Technol. 2015, 22, 1–6. [Google Scholar] [CrossRef]
- Mahalanabis, M.; Do, J.; AlMuayad, H.; Zhang, J.Y.; Klapperich, C.M. An integrated disposable device for DNA extraction and helicase dependent amplification. Biomed. Microdevices 2011, 12, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Do, J.; Mahalanabis, M.; Fan, A.; Zhao, L.; Jepeal, L.; Singh, S.K.; Klapperich, C.M. Low Cost Extraction and Isothermal Amplification of DNA for Infectious Diarrhea Diagnosis. PLoS ONE 2013, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Sadoshima, J. Helicase Dependent Isothermal Amplification of DNA and RNA using Self-Avoiding Molecular Recognition Systems. Med. Image Anal. 2015, 116, 1477–1490. [Google Scholar]
- Denys, G.A. Portrait Toxigenic Clostridium difficile assay, an isothermal amplification assay detects toxigenic C. difficile in clinical stool specimens. Expert Rev. Mol. Diagn. 2014, 14, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Chen, Q.; Liu, W.; Li, H.; Lin, J.M. A portable microchip for ultrasensitive and high-throughput assay of thrombin by rolling circle amplification and hemin/G-quadruplex system. Biosens. Bioelectron. 2014, 56, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Nixon, G.; Garson, J.; Grant, P. Comparative study of sensitivity, linearity, and resistance to inhibition of digital and nondigital polymerase chain reaction and loop mediated isothermal amplification. Anal. Chem. 2014, 86, 4387–4394. [Google Scholar] [CrossRef] [PubMed]
- Hindson, B.J.; Ness, K.D.; Masquelier, D.A.; Belgrader, P.; Heredia, N.J.; Makarewicz, A.J.; Bright, I.J.; Lucero, M.Y.; Hiddessen, A.L.; Legler, T.C.; et al. High-Throughput Droplet Digital PCR System for Absolute Quantitation of DNA Copy Number. Anal. Chem. 2011, 83, 8604–8610. [Google Scholar] [CrossRef] [PubMed]
- Rane, T.D.; Chen, L.; Zec, H.C.; Wang, T.-H. Microfluidic continuous flow digital loop-mediated isothermal amplification (LAMP). Lab Chip 2015, 15, 776–782. [Google Scholar] [CrossRef] [PubMed]
- Dobnik, D.; Štebih, D.; Blejec, A.; Morisset, D.; Žel, J. Multiplex quantification of four DNA targets in one reaction with Bio-Rad droplet digital PCR system for GMO detection. Sci. Rep. 2016, 6, 35451. [Google Scholar] [CrossRef] [PubMed]
- Lagus, T.P.; Edd, J.F. A review of the theory, methods and recent applications of high-throughput single-cell droplet microfluidics. J. Phys. D Appl. Phys. 2013, 46, 114005. [Google Scholar] [CrossRef]
- Hatch, A.C.; Fisher, J.S.; Tovar, A.R.; Hsieh, A.T.; Lin, R.; Pentoney, S.L.; Yang, D.L.; Lee, A.P. 1-Million droplet array with wide-field fluorescence imaging for digital PCR. Lab Chip 2011, 11, 3838. [Google Scholar] [CrossRef] [PubMed]
- Yukl, S.A.; Kaiser, P.; Kim, P.; Li, P.; Wong, J.K. Advantages of using the QIAshredder instead of restriction digestion to prepare DNA for droplet digital PCR. Biotechniques 2014, 56, 194–196. [Google Scholar] [CrossRef] [PubMed]
- Bauer, W.-A.C.; Fischlechner, M.; Abell, C.; Huck, W.T.S. Hydrophilic PDMS microchannels for high-throughput formation of oil-in-water microdroplets and water-in-oil-in-water double emulsions. Lab Chip 2010, 10, 1814. [Google Scholar] [CrossRef] [PubMed]
- Sang, Y.Y.C.; Lorenceau, E.; Wahl, S.; Stoffel, M.; Angelescu, D.E.; Höhler, R. A microfluidic technique for generating monodisperse submicron-sized drops. RSC Adv. 2013, 3, 2330. [Google Scholar] [CrossRef]
- Bian, X.; Jing, F.; Li, G.; Fan, X.; Jia, C.; Zhou, H.; Jin, Q.; Zhao, J. A microfluidic droplet digital PCR for simultaneous detection of pathogenic Escherichia coli O157 and Listeria monocytogenes. Biosens. Bioelectron. 2015, 74, 770–777. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Jing, F.; Li, G.; Wu, Z.; Cheng, Z.; Zhang, J.; Zhang, H.; Jia, C.; Jin, Q.; Mao, H.; Zhao, J. Absolute quantification of lung cancer related microRNA by droplet digital PCR. Biosens. Bioelectron. 2015, 74, 836–842. [Google Scholar] [CrossRef] [PubMed]
- Schuler, F.; Siber, C.; Hin, S.; Wadle, S.; Paust, N.; Zengerle, R.; von Stetten, F. Digital droplet LAMP as a microfluidic app on standard laboratory devices. Anal. Methods 2016, 8, 2750–2755. [Google Scholar] [CrossRef]
- Schuler, F.; Schwemmer, F.; Trotter, M. Centrifugal step emulsification applied for absolute quantification of nucleic acids by digital droplet RPA. Lab Chip 2015, 15, 2759–2766. [Google Scholar] [CrossRef] [PubMed]
- Schneider, T.; Yen, G.S.; Thompson, A.M.; Burnham, D.R.; Chiu, D.T. Self-digitization of samples into a high-density microfluidic bottom-well array. Anal. Chem. 2013, 85, 10417–10423. [Google Scholar] [CrossRef] [PubMed]
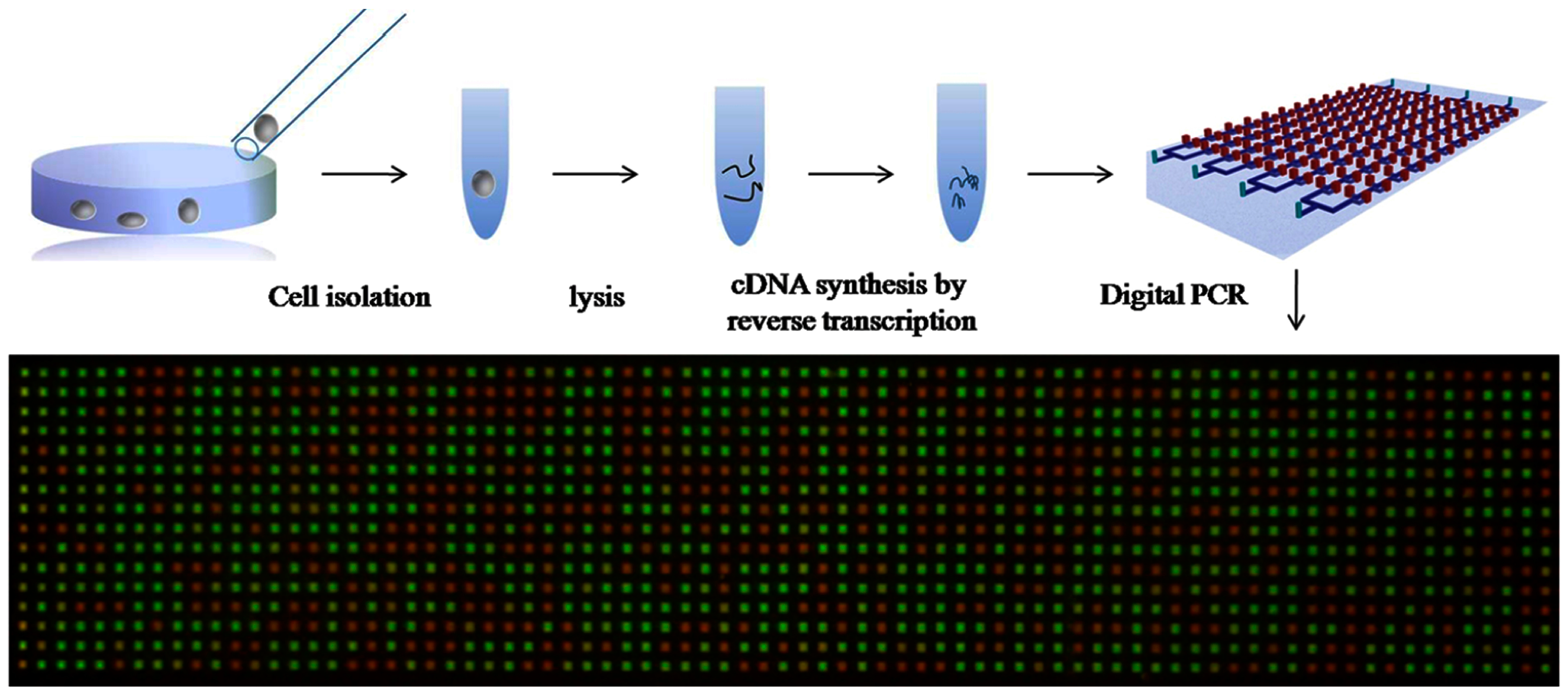
- Thompson, A.M.; Gansen, A.; Paguirigan, A.L.; Kreutz, J.E.; Radich, J.P.; Chiu, D.T. Self-Digitization Microfluidic Chip for Absolute Quantification of mRNA in Single Cells. Anal. Chem. 2014, 86, 12308–12314. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Li, L.; Nichols, K.P.; Ismagilov, R.F. SlipChip. Lab Chip 2009, 9, 2286. [Google Scholar] [CrossRef] [PubMed]
- Shen, F.; Du, W.; Kreutz, J.E.; Fok, A.; Ismagilov, R.F. Digital PCR on a SlipChip. Lab Chip 2010, 10, 2666. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Gao, Y.; Yu, B.; Ren, H.; Qiu, L.; Han, S.; Jin, W.; Jin, Q.; Mu, Y. Self-priming compartmentalization digital LAMP for point-of-care. Lab Chip 2012, 12, 4755–4763. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Qiu, L.; Yu, B.; Xu, Y.; Gao, Y.; Pan, T.; Tian, Q.; Song, Q.; Jin, W.; Jin, Q.; Mu, Y. Digital PCR on an integrated self-priming compartmentalization chip. Lab Chip 2014, 14, 1176–1185. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Qiu, L.; Xu, Y.; Li, G.; Mu, Y. Single cell digital polymerase chain reaction on self-priming compartmentalization chip. Biomicrofluidics 2017, 11, 14109. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Zhou, H.; Jia, C.; Jing, F.; Jin, Q.; Zhao, J.; Li, G. A microfluidic chip based on surfactant-doped polydimethylsiloxane (PDMS) in a sandwich configuration for low-cost and robust digital PCR. Sens. Actuators B Chem. 2017, 245, 414–422. [Google Scholar] [CrossRef]
- Yeh, E.-C.; Fu, C.-C.; Hu, L.; Thakur, R.; Feng, J.; Lee, L.P. Self-powered integrated microfluidic point-of-care low-cost enabling (SIMPLE) chip. Sci. Adv. 2017, 3, e1501645. [Google Scholar] [CrossRef] [PubMed]
- Zonta, E.; Garlan, F.; Pécuchet, N.; Perez-Toralla, K.; Caen, O.; Milbury, C.; Didelot, A.; Fabre, E.; Blons, H.; Laurent-Puig, P.; et al. Multiplex Detection of Rare Mutations by Picoliter Droplet Based Digital PCR: Sensitivity and Specificity Considerations. PLoS ONE 2016, 11, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Rajeswari, P.K.P.; Soderberg, L.M.; Yacoub, A.; Leijon, M.; Svahn, H.A.; Joensson, H.N. Multiple pathogen biomarker detection using an encoded bead array in droplet PCR. J. Microbiol. Methods 2017, 139, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Liang, G.; Zhang, Q.; Chen, B. Detection of Mycobacterium tuberculosis Using a Capillary-Array Microsystem with Integrated DNA Extraction, Loop-Mediated Isothermal Amplification, and Fluorescence Detection. Anal. Chem. 2013, 85, 4698–4704. [Google Scholar] [CrossRef] [PubMed]
- Yao, M.; Shah, G.; Fang, J. Highly Sensitive and Miniaturized Fluorescence Detection System with an Autonomous Capillary Fluid Manipulation Chip. Micromachines 2012, 3, 462–479. [Google Scholar] [CrossRef]
- Gravel, J.-F.; Geissler, M.; Chapdelaine, S.; Boissinot, K.; Voisin, B.; Charlebois, I.; Poirier-Richard, H.-P.; Grégoire, A.; Boissinot, M.; Bergeron, M.G.; et al. Portable bead-based fluorescence detection system for multiplex nucleic acid testing: A case study with Bacillus anthracis. Microfluid. Nanofluid. 2013, 16, 1075–1087. [Google Scholar] [CrossRef]
- Hsu, Y.M.; Chang, C.C. The portable fluorescence detection system matched with PDMS microfluidic biochip for DNA hybridization detection. Opt. Int. J. Light Electron Opt. 2015, 126, 2600–2605. [Google Scholar] [CrossRef]
- Craw, P.; Mackay, R.E.; Naveenathayalan, A.; Hudson, C.; Branavan, M.; Sadiq, S.T.; Balachandran, W. A Simple, Low-Cost Platform for Real-Time Isothermal Nucleic Acid Amplification. Sensors 2015, 15, 23418–23430. [Google Scholar] [CrossRef] [PubMed]
- Ryu, G.; Huang, J.; Hofmann, O.; Walshe, C.A.; Sze, J.Y.Y.; McClean, G.D.; Mosley, A.; Rattle, S.J.; deMello, J.C.; deMello, A.J.; et al. Highly sensitive fluorescence detection system for microfluidic lab-on-a-chip. Lab Chip 2011, 11, 1664–1670. [Google Scholar] [CrossRef] [PubMed]
- Hung, T.Q.; Chin, W.H.; Sun, Y.; Wolff, A.; Bang, D.D. A novel lab-on-chip platform with integrated solid phase PCR and supercritical angle fluorescence (SAF) microlens array for highly sensitive and multiplexed pathogen detection. Biosens. Bioelectron. 2016, 90, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Tae, Y.; Chen, Y.; Young, J.; Kim, W.; Dae, H.; Jung, J.; Seok, T. Integrated microdevice of reverse transcription-polymerase chain reaction with colorimetric immunochromatographic detection for rapid gene expression analysis of influenza A H1N1 virus. Biosens. Bioelectron. 2012, 33, 88–94. [Google Scholar]
- Safavieh, M.; Ahmed, M.M.U.; Sokullu, E.; Ng, A.; Braescu, L.; Zourob, M. A simple cassette as point-of-care diagnostic device for naked-eye colorimetric bacteria detection. Analyst 2014, 139, 482–487. [Google Scholar] [CrossRef] [PubMed]
- Chang, W.-H.; Yang, S.-Y.; Lin, C.-L.; Wang, C.-H.; Li, P.-C.; Chen, T.-Y.; Jan, F.-J.; Lee, G.-B. Detection of viruses directly from the fresh leaves of a Phalaenopsis orchid using a microfluidic system. Nanomedicine 2013, 9, 1274–1282. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.C.-L.; Chang, W.-H.W.; Wang, C.; Lee, C.-H.; Chen, T.-Y.; Jan, F.-J.; Lee, G.-B. A microfluidic system integrated with buried optical fibers for detection of Phalaenopsis orchid pathogens. Biosens. Bioelectron. 2015, 63, 572–579. [Google Scholar] [CrossRef] [PubMed]
- Comina, G.; Suska, A.; Filippini, D. Towards autonomous lab-on-a-chip devices for cell phone biosensing. Biosens. Bioelectron. 2016, 77, 1153–1167. [Google Scholar] [CrossRef] [PubMed]
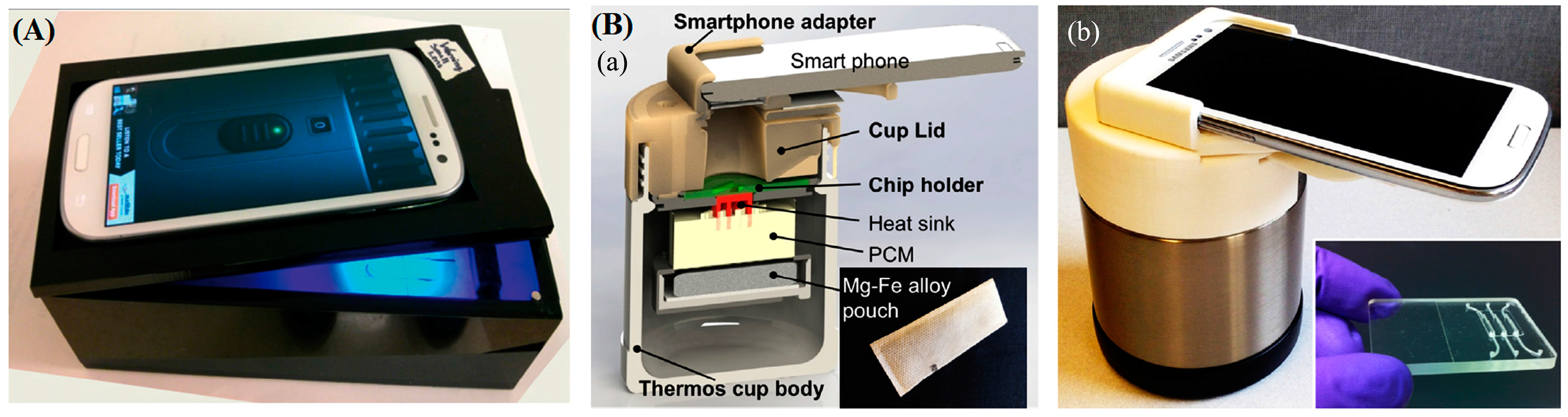
- Damhorst, G.; Duarte-Guevara, C. Smartphone-Imaged HIV-1 Reverse-Transcription Loop-Mediated Isothermal Amplification (RT-LAMP) on a Chip from Whole Blood. Engineering 2015, 1, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Liao, S.; Peng, J.; Mauk, M.; Awasthi, S. Smart cup: A minimally-instrumented, smartphone-based point-of-care molecular diagnostic device. Sens. Actuators B Chem. 2016, 229, 232–238. [Google Scholar] [CrossRef] [PubMed]
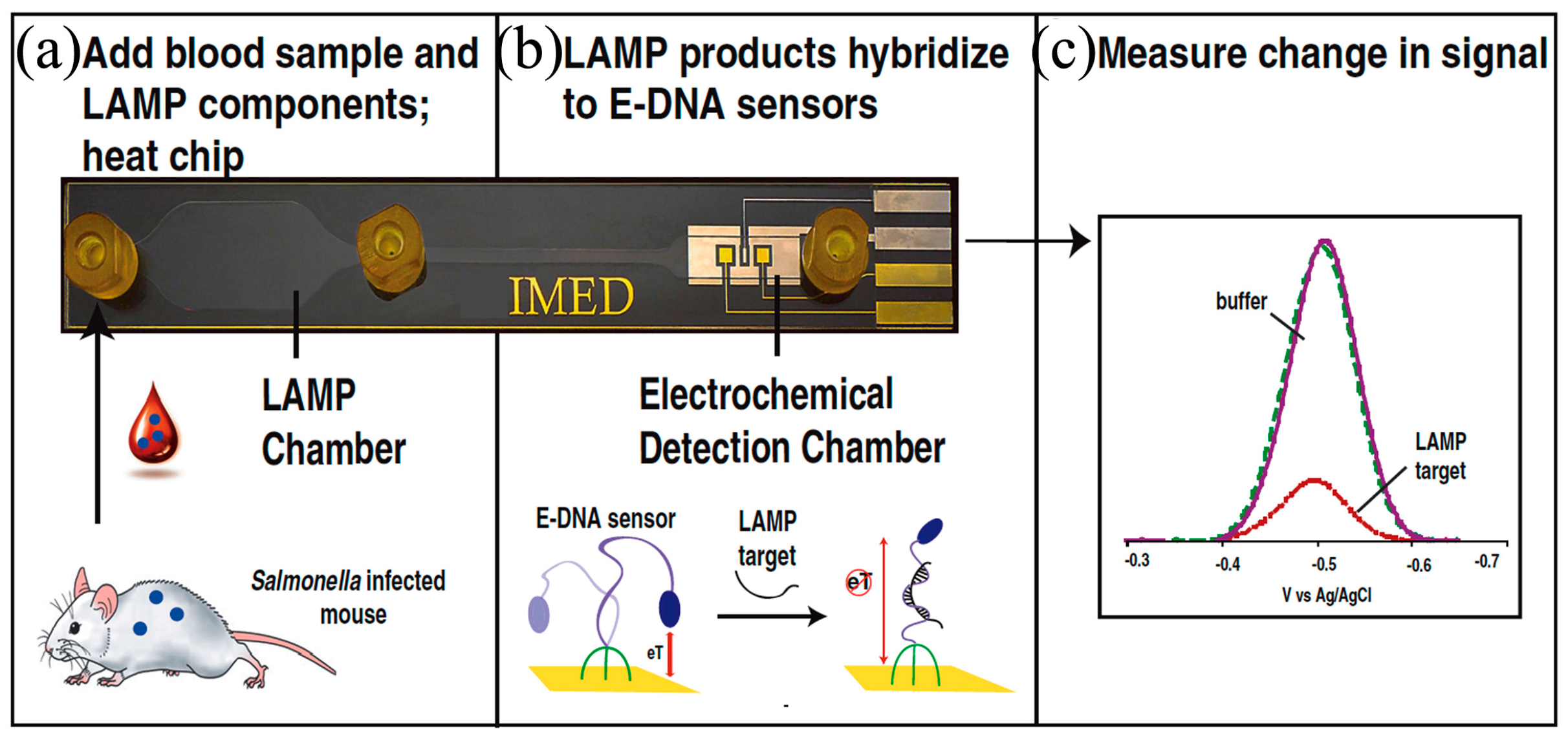
- Patterson, A.S.; Heithoff, D.M.; Ferguson, B.S.; Soh, H.T.; Mahan, M.J.; Plaxco, K.W. Microfluidic chip-based detection and intraspecies strain discrimination of Salmonella serovars derived from whole blood of septic mice. Appl. Environ. Microbiol. 2013, 79, 2302–2311. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, K.; Patterson, A.S.; Ferguson, B.S.; Plaxco, K.W.; Soh, H.T. Rapid, Sensitive, and Quantitative Detection of Pathogenic DNA at the Point of Care through Microfluidic Electrochemical Quantitative Loop-Mediated Isothermal Amplification. Angew. Chem. 2012, 124, 4980–4984. [Google Scholar] [CrossRef]
- Lee, D.C.; Yip, S.P.; Lee, T.M.H. Simple and Sensitive Electrochemical DNA Detection of Primer Generation-Rolling Circle Amplification. Electroanalysis 2013, 25, 1310–1315. [Google Scholar] [CrossRef]
- Kivlehan, F.; Mavré, F.; Talini, L.; Limoges, B.; Marchal, D. Real-time electrochemical monitoring of isothermal helicase-dependent amplification of nucleic acids. Analyst 2011, 136, 3635–3642. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.U.; Nahar, S.; Safavieh, M.; Zourob, M. Real-time electrochemical detection of pathogen DNA using electrostatic interaction of a redox probe. Analyst 2013, 138, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Safavieh, M.; Ahmed, M.U.; Ng, A.; Zourob, M. High-throughput real-time electrochemical monitoring of LAMP for pathogenic bacteria detection. Biosens. Bioelectron. 2014, 58, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Fang, X.; Ye, D.; Li, H.; Chen, H.; Zhang, S.; Kong, J. A real-time microfluidic multiplex electrochemical loop-mediated isothermal amplification chip for differentiating bacteria. Biosens. Bioelectron. 2014, 60, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.W.; Phillips, S.T.; Butte, M.J.; Whitesides, G.M. Patterned paper as a platform for inexpensive, low-volume, portable bioassays. Angew. Chem. Int. Ed. 2007, 46, 1318–1320. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.W.; Phillips, S.T.; Whitesides, G.M. Three-dimensional microfluidic devices fabricated in layered paper and tape. Proc. Natl. Acad. Sci. USA 2008, 105, 19606–19611. [Google Scholar] [CrossRef] [PubMed]
- Fronczek, C.F.; Park, T.S.; Harshman, D.K.; Nicolini, A.M.; Yoon, J.-Y. Paper microfluidic extraction and direct smartphone-based identification of pathogenic nucleic acids from field and clinical samples. RSC Adv. 2014, 4, 11103–11110. [Google Scholar] [CrossRef]
- Li, X.; Luo, L.; Crooks, R. Low-voltage paper isotachophoresis device for DNA focusing. Lab Chip 2015, 15, 4090–4098. [Google Scholar] [CrossRef] [PubMed]
- Moghadam, B.Y.; Connelly, K.T.; Posner, J.D. Isotachophoretic Preconcenetration on Paper-Based Microfluidic Devices. Anal. Chem. 2014, 86, 5829–5837. [Google Scholar] [CrossRef] [PubMed]
- Connelly, J.T.; Rolland, J.P.; Whitesides, G.M. “Paper Machine” for Molecular Diagnostics. Anal. Chem. 2015, 87, 7595–7601. [Google Scholar] [CrossRef] [PubMed]
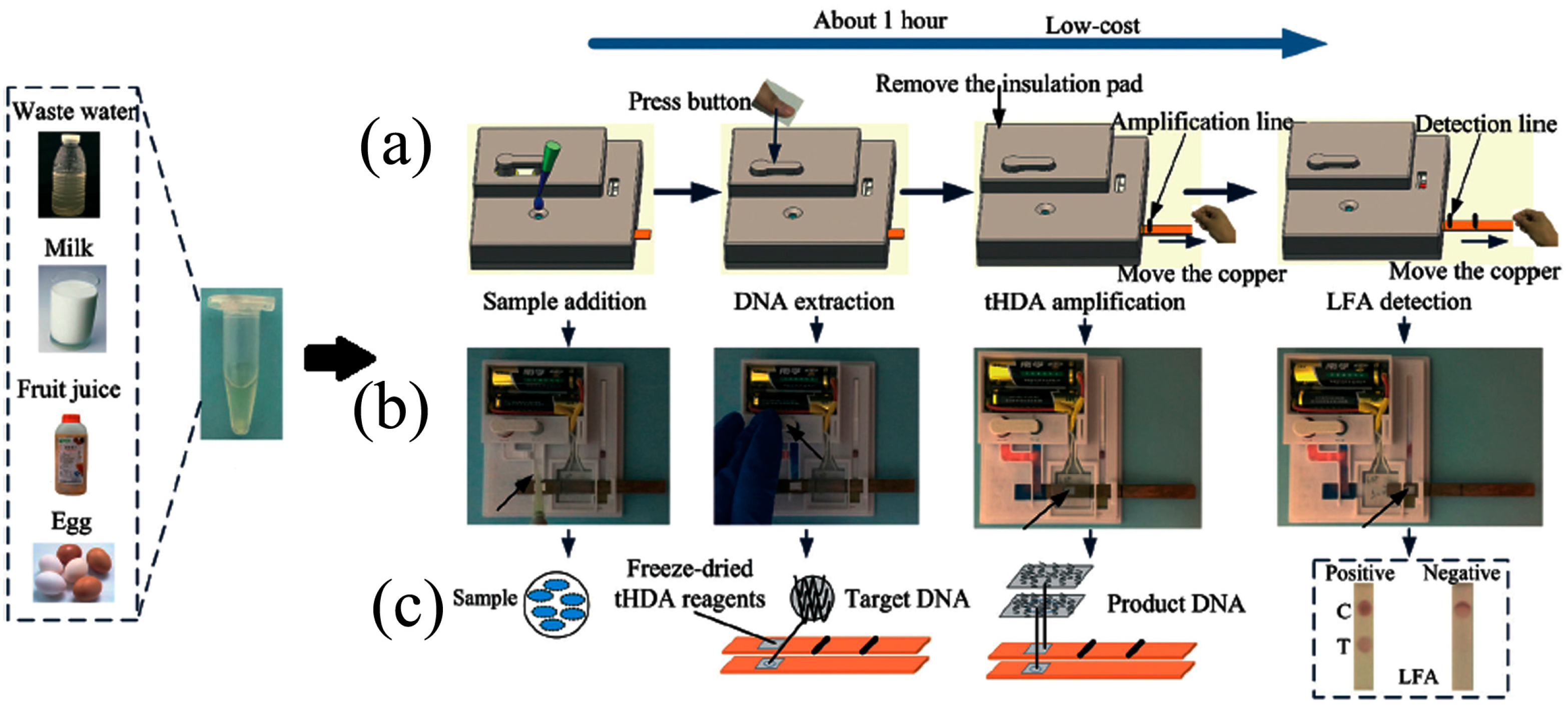
- Tang, R.; Yang, H.; Gong, Y.; You, M.; Liu, Z.; Choi, J.R.; Wen, T.; Qu, Z.; Mei, Q.; Xu, F. A fully disposable and integrated paper-based device for nucleic acid extraction, amplification and detection. Lab Chip 2017, 17, 1270–1279. [Google Scholar] [CrossRef] [PubMed]
- Miao, B.; Peng, N.; Li, L.; Li, Z.; Hu, F.; Zhang, Z.; Wang, C. Centrifugal Microfluidic System for Nucleic Acid Amplification and Detection. Sensors 2015, 15, 27954–27968. [Google Scholar] [CrossRef] [PubMed]
- Duvall, J.A.; Le Roux, D.; Tsuei, A.-C.; Thompson, B.L.; Birch, C.; Li, J.; Nelson, D.A.; Mills, D.L.; Ewing, M.M.; Mclaren, R.S.; et al. A rotationally-driven polyethylene terephthalate microdevice with integrated reagent mixing for multiplexed PCR amplification of DNA. Anal. Methods 2016, 8, 7331–7340. [Google Scholar] [CrossRef]
- Jung, J.H.; Park, B.H.; Oh, S.J.; Choi, G.; Seo, T.S. Integrated centrifugal reverse transcriptase loop-mediated isothermal amplification microdevice for influenza A virus detection. Biosens. Bioelectron. 2015, 68, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Felipe, S.; Tortajada-Genaro, L.A.; Carrascosa, J.; Puchades, R.; Maquieira, Á. Real-time loop-mediated isothermal DNA amplification in compact disc micro-reactors. Biosens. Bioelectron. 2016, 79, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Tortajada-Genaro, L.A.; Santiago-Felipe, S.; Amasia, M.; Russom, A.; Maquieira, Á. Isothermal solid-phase recombinase polymerase amplification on microfluidic digital versatile discs (DVDs). RSC Adv. 2015, 5, 29987–29995. [Google Scholar] [CrossRef]
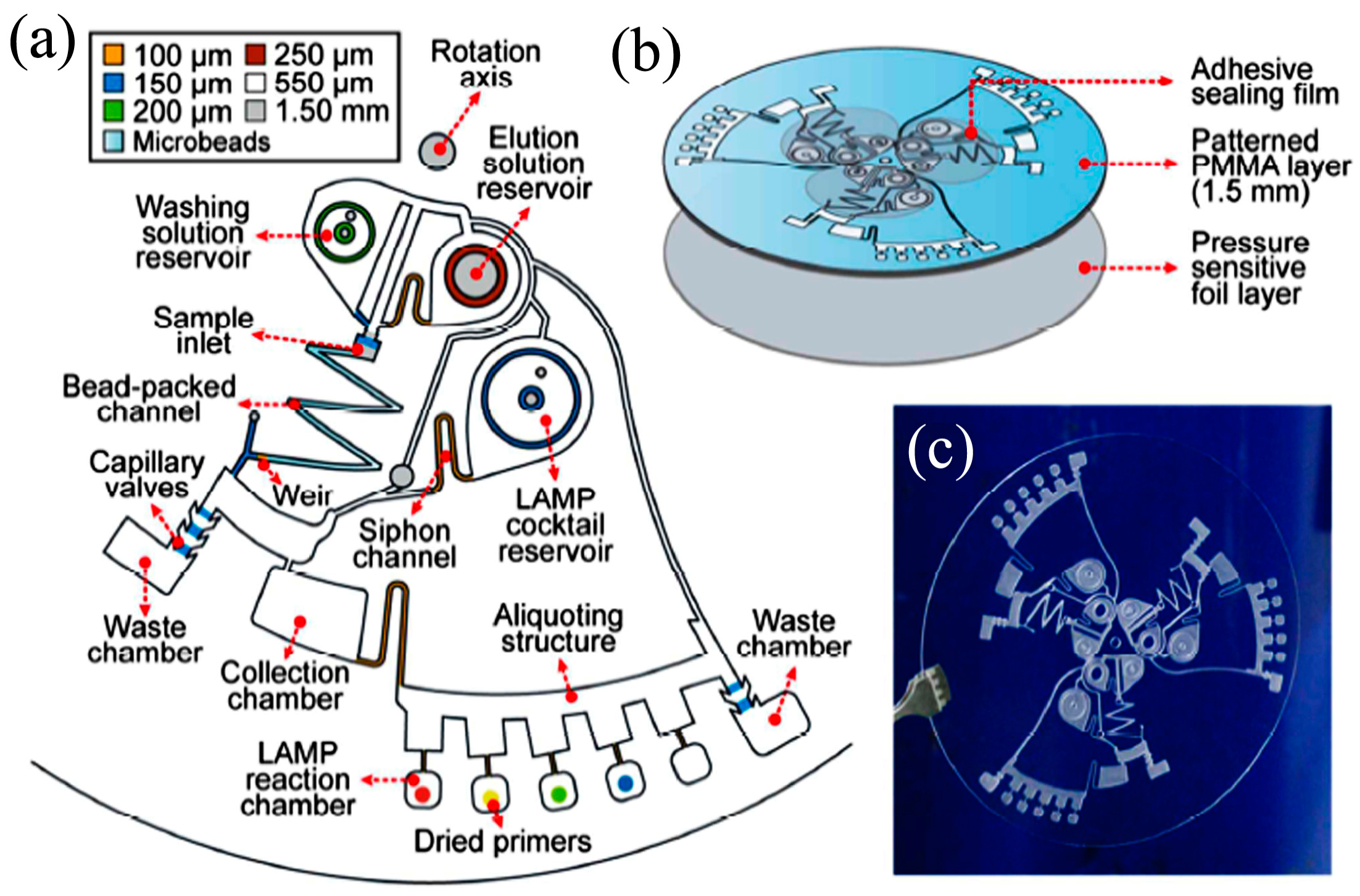
- Sayad, A.A.; Ibrahim, F.; Uddin, S.M.; Pei, K.X.; Mohktar, M.S.; Madou, M.; Thong, K.L. A microfluidic lab-on-a-disc integrated loop mediated isothermal amplification for foodborne pathogen detection. Sens. Actuators B Chem. 2016, 227, 600–609. [Google Scholar] [CrossRef]
- Oh, S.J.; Park, B.H.; Jung, J.H.; Choi, G.; Lee, D.C.; Kim, D.H.; Seo, T.S. Centrifugal loop-mediated isothermal amplification microdevice for rapid, multiplex and colorimetric foodborne pathogen detection. Biosens. Bioelectron. 2016, 75, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.; Chao, Y.; Yen, J.; Yu, Y.; Lee, C.; Ho, B.; Liu, K.; Fang, J.; Lin, C.; Lee, J. A Turbidity Test Based Centrifugal Microfluidics Diagnostic System for Simultaneous Detection of HBV, HCV, and CMV. Adv. Mater. Sci. Eng. 2015, 2015, 1–8. [Google Scholar] [CrossRef]
- Oh, S.J.; Park, B.H.; Choi, G.; Seo, J.H.; Jung, J.H.; Choi, J.S.; Kim, D.H.; Seo, T.S. Fully automated and colorimetric foodborne pathogen detection on an integrated centrifugal microfluidic device. Lab Chip 2016, 16, 1917–1926. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.S.; Kim, D.H.; Seo, T.S. A centrifugal direct recombinase polymerase amplification (direct-RPA) microdevice for multiplex and real-time identification of food poisoning bacteria. Lab Chip 2016, 16, 2309–2316. [Google Scholar] [CrossRef] [PubMed]
- Hyun, J.; Hyun, B.; Jun, S.; Choi, G.; Hyun, D.; Yeol, E.; Seok, T. Development of a high-throughput centrifugal loop-mediated isothermal amplification microdevice for multiplex foodborne pathogenic bacteria detection. Sens. Actuators B Chem. 2017, 246, 146–153. [Google Scholar]
- Park, B.H.; Oh, S.J.; Jung, J.H.; Choi, G.; Seo, J.H.; Kim, D.H.; Lee, E.Y.; Seo, T.S. An integrated rotary microfluidic system with DNA extraction, loop-mediated isothermal amplification, and lateral flow strip based detection for point-of-care pathogen diagnostics. Biosens. Bioelectron. 2016, 91, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Park, J.; Kim, C.; Cho, Y. Fully integrated lab-on-a-disc for nucleic acid analysis of food-borne pathogens. Anal. Chem. 2014, 86, 3841–3848. [Google Scholar] [CrossRef] [PubMed]
- Hung, P.-Y.; Jiang, P.-S.; Lee, E.-F.; Fan, S.-K.; Lu, Y.-W. Genomic DNA extraction from whole blood using a digital microfluidic (DMF) platform with magnetic beads. Microsyst. Technol. 2015, 23, 313–320. [Google Scholar] [CrossRef]
- Rival, A.; Jary, D.; Delattre, C.; Fouillet, Y. An EWOD-based microfluidic chip for single-cell isolation, mRNA purification and subsequent multiplex qPCR. Lab Chip 2014, 14, 3739–3749. [Google Scholar] [CrossRef] [PubMed]
- Kalsi, S.; Sellars, S.; Turner, C.; Sutton, J.; Morgan, H. A Programmable Digital Microfluidic Assay for the Simultaneous Detection of Multiple Anti-Microbial Resistance Genes. Micromachines 2017, 8, 12. [Google Scholar] [CrossRef]
- Agarwal, A. Digital Microfluidics: Techniques, Their Applications and Advantages. J. Bioeng. Biomed. Sci. 2013, 3, 1–6. [Google Scholar] [CrossRef]
- Zhang, Y.; Park, S.; Liu, K.; Tsuan, J.; Yang, S.; Wang, T.-H. A surface topography assisted droplet manipulation platform for biomarker detection and pathogen identification. Lab Chip 2011, 11, 398–406. [Google Scholar] [CrossRef] [PubMed]
- Chiou, C.H.; Jin Shin, D.; Zhang, Y.; Wang, T.H. Topography-assisted electromagnetic platform for blood-to-PCR in a droplet. Biosens. Bioelectron. 2013, 50, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.J.; Zhang, Y.; Wang, T.-H. A droplet microfluidic approach to single-stream nucleic acid isolation and mutation detection. Microfluid. Nanofluid. 2014, 17, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.-H.; Lien, K.-Y.; Hung, L.; Lei, H.; Lee, G.-B. Integrated microfluidic system for the identification and multiple subtyping of influenza viruses by using a molecular diagnostic approach. Microfluid. Nanofluid. 2012, 13, 113–123. [Google Scholar] [CrossRef]
- Tai, C.-H.; Tsai, Y.-C.; Wang, C.-H.; Ho, T.-S.; Chang, C.-P.; Lee, G.-B. An integrated microfluidic platform for rapid detection and subtyping of influenza viruses from clinical samples. Microfluid. Nanofluid. 2013, 16, 501–512. [Google Scholar] [CrossRef]
- Wolff, A.; Sun, Y.; Quyen, T.; Hung, T.; Chin, W. A lab-on-a-chip system with integrated sample preparation and loop-mediated isothermal amplification for rapid and quantitative detection of Salmonella spp. in food samples Lab on a Chip. Lab Chip 2015, 15, 1898–1904. [Google Scholar]
- Branavan, M.; Mackay, R.E.; Craw, P.; Naveenathayalan, A.; Ahern, J.C.; Sivanesan, T.; Hudson, C.; Stead, T.; Kremer, J.; Garg, N.; et al. Modular development of a prototype point of care molecular diagnostic platform for sexually transmitted infections. Med. Eng. Phys. 2016, 38, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Dou, M.; Sanjay, S.T.; Liu, P.; Xu, F.; Li, X. Multiplexed instrument-free meningitis diagnosis on a polymer/paper hybrid microfluidic biochip. Biosens. Bioelectron. 2017, 87, 865–873. [Google Scholar] [CrossRef] [PubMed]
- Czilwik, G.; Messinger, T.; Strohmeier, O.; Wadle, S.; von Stetten, F.; Paust, N.; Roth, G.; Zengerle, R.; Saarinen, P.; Niittymäki, J.; et al. Rapid and fully automated bacterial pathogen detection on a centrifugal-microfluidic LabDisk using highly sensitive nested PCR with integrated sample preparation. Lab Chip 2015, 15, 3749–3759. [Google Scholar] [CrossRef] [PubMed]
- Stumpf, F.; Schwemmer, F.; Hutzenlaub, T.; Baumann, D.; Strohmeier, O.; Dingemanns, G.; Simons, G.; Sager, C.; Plobner, L.; von Stetten, F.; et al. LabDisk with complete reagent prestorage for sample-to-answer nucleic acid based detection of respiratory pathogens verified with influenza A H3N2 virus. Lab Chip 2016, 16, 199–207. [Google Scholar] [CrossRef] [PubMed]
- Roy, E.; Stewart, G.; Mounier, M.; Malic, L.; Peytavi, R.; Clime, L.; Madou, M.; Bossinot, M.; Bergeron, M.G.; Veres, T. From cellular lysis to microarray detection, an integrated thermoplastic elastomer (TPE) point of care Lab on a Disc. Lab Chip 2015, 15, 406–416. [Google Scholar] [CrossRef] [PubMed]
- Micronics, I. PanNAT® Molecular Diagnostic System. Available online: https://www.micronics.net/products/diagnostic-products/PanNAT (accessed on 12 August 2016).
- QuantuMDx Q-POC. Available online: http://quantumdx.com/systems/q-poc/ (accessed on 12 August 2016).
- SRI International. Sentinel Nucleic Acid Analysis System. Available online: https://www.sri.com/work/projects/sentinel-portable-bioanalysis-system (accessed on 12 August 2016).
- Fluidigm Biomark HD System. Available online: https://www.fluidigm.com/products/biomark-hd-system#workflow (accessed on 12 August 2016).
- Technologies, L. OpenArray. Available online: https://www.thermofisher.com/my/en/home/life-science/pcr/real-time-pcr/real-time-openarray.html (accessed on 12 August 2016).
- Wafergen SmartChip System. Available online: http://www.wafergen.com/products/smartchip-real-time-pcr (accessed on 12 August 2016).
- Technologies, R. RainDropTM Digital PCR System. Available online: http://raindancetech.com/digital-pcr-tech/raindrop-digital-pcr-system/ (accessed on 12 August 2016).
- BIO-RAD QX200 Droplet Digital PCR. Available online: http://www.bio-rad.com/en-us/product/qx200-droplet-digital-pcr-system (accessed on 12 August 2016).


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Basha, I.H.K.; Ho, E.T.W.; Yousuff, C.M.; Hamid, N.H.B. Towards Multiplex Molecular Diagnosis—A Review of Microfluidic Genomics Technologies. Micromachines 2017, 8, 266. https://doi.org/10.3390/mi8090266
Basha IHK, Ho ETW, Yousuff CM, Hamid NHB. Towards Multiplex Molecular Diagnosis—A Review of Microfluidic Genomics Technologies. Micromachines. 2017; 8(9):266. https://doi.org/10.3390/mi8090266
Chicago/Turabian StyleBasha, Ismail Hussain Kamal, Eric Tatt Wei Ho, Caffiyar Mohamed Yousuff, and Nor Hisham Bin Hamid. 2017. "Towards Multiplex Molecular Diagnosis—A Review of Microfluidic Genomics Technologies" Micromachines 8, no. 9: 266. https://doi.org/10.3390/mi8090266
APA StyleBasha, I. H. K., Ho, E. T. W., Yousuff, C. M., & Hamid, N. H. B. (2017). Towards Multiplex Molecular Diagnosis—A Review of Microfluidic Genomics Technologies. Micromachines, 8(9), 266. https://doi.org/10.3390/mi8090266




