Microfluidic Approaches and Methods Enabling Extracellular Vesicle Isolation for Cancer Diagnostics
Abstract
1. Introduction
2. Importance of Extracellular Vesicles
3. Conventional Isolation Techniques
3.1. Centrifugation-Based Techniques
3.2. Size Exclusion
3.3. Polymer-Based Precipitation Methods
4. Microfluidic-Based Devices for Extracellular Vesicle Isolation
4.1. Isolation Based on Immunoaffinity
4.2. Isolation Based on Size
4.2.1. Filtration
4.2.2. Acoustofluidics
4.2.3. Inertial Microfluidics
4.3. Isolation Based on Electrokinetics
5. Summary and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Diamantis, A.; Magiorkinis, E.; Koutselini, H. Fine-needle aspiration (FNA) biopsy: Historical aspects. Folia Histochem. Cytobiol 2009, 47, 191–197. [Google Scholar] [CrossRef]
- Robertson, E.G.; Baxter, G. Tumour seeding following percutaneous needle biopsy: The real story! Clin. Radiol. 2011, 66, 1007–1014. [Google Scholar] [CrossRef]
- Lu, J.; Pang, J.; Chen, Y.; Dong, Q.; Sheng, J.; Luo, Y.; Lu, Y.; Lin, B.; Liu, T. Application of Microfluidic Chips in Separation and Analysis of Extracellular Vesicles in Liquid Biopsy for Cancer. Micromachines 2019, 10, 390. [Google Scholar] [CrossRef]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A., Jr.; Kinzler, K.W. Cancer genome landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; Tarpey, P.; et al. Intratumor Heterogeneity and Branched Evolution Revealed by Multiregion Sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Pasha, N.; Turner, N.C. Understanding and overcoming tumor heterogeneity in metastatic breast cancer treatment. Nat. Cancer 2021, 2, 680–692. [Google Scholar] [CrossRef]
- Hu, Z.; Li, Z.; Ma, Z.; Curtis, C. Multi-cancer analysis of clonality and the timing of systemic spread in paired primary tumors and metastases. Nat. Genet. 2020, 52, 701–708. [Google Scholar] [CrossRef] [PubMed]
- Tellez-Gabriel, M.; Ory, B.; Lamoureux, F.; Heymann, M.-F.; Heymann, D. Tumour Heterogeneity: The Key Advantages of Single-Cell Analysis. Int. J. Mol. Sci. 2016, 17, 2142. [Google Scholar] [CrossRef]
- Winkler, J.; Abisoye-Ogunniyan, A.; Metcalf, K.J.; Werb, Z. Concepts of extracellular matrix remodelling in tumour progression and metastasis. Nat. Commun. 2020, 11, 5120. [Google Scholar] [CrossRef]
- Prakash, S.; Karnes, M.P.; Sequin, E.K.; West, J.D.; Hitchcock, C.L.; Nichols, S.D.; Bloomston, M.; Abdel-Misih, S.R.; Schmidt, C.R.; Martin, E.W., Jr.; et al. Ex vivo electrical impedance measurements on excised hepatic tissue from human patients with metastatic colorectal cancer. Physiol Meas 2015, 36, 315–328. [Google Scholar] [CrossRef] [PubMed]
- Lochab, V.; Jones, T.H.; Alkandry, E.; West, J.D.; Abdel-Rahman, M.H.; Subramaniam, V.V.; Prakash, S. Evaluation of electrical properties of ex vivo human hepatic tissue with metastatic colorectal cancer. Physiol. Meas. 2020, 41, 085005. [Google Scholar] [CrossRef]
- 40 Years of the War on Cancer. Science 2011, 331, 1540–1544. [CrossRef]
- Mino-Kenudson, M. Cons: Can liquid biopsy replace tissue biopsy?—the US experience. Transl. Lung Cancer Res. 2016, 5, 424–427. [Google Scholar] [CrossRef]
- Špilak, A.; Brachner, A.; Kegler, U.; Neuhaus, W.; Noehammer, C. Implications and pitfalls for cancer diagnostics exploiting extracellular vesicles. Adv. Drug Deliv. Rev. 2021, 175, 113819. [Google Scholar] [CrossRef]
- Tadimety, A.; Closson, A.; Li, C.; Yi, S.; Shen, T.; Zhang, J.X.J. Advances in liquid biopsy on-chip for cancer management: Technologies, biomarkers, and clinical analysis. Crit. Rev. Clin. Lab. Sci. 2018, 55, 140–162. [Google Scholar] [CrossRef] [PubMed]
- Vaidyanathan, R.; Soon, R.H.; Zhang, P.; Jiang, K.; Lim, C.T. Cancer diagnosis: From tumor to liquid biopsy and beyond. Lab A Chip 2019, 19, 11–34. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Calderón, D.; Pedraza, A.; Mancera Urrego, C.; Mejía-Mejía, A.; Montealegre-Páez, A.L.; Perdomo, S. Analysis of the Cost-Effectiveness of Liquid Biopsy to Determine Treatment Change in Patients with Her2-Positive Advanced Breast Cancer in Colombia. Clin. Outcomes Res. 2020, 12, 115–122. [Google Scholar] [CrossRef]
- Chen, M.; Zhao, H. Next-generation sequencing in liquid biopsy: Cancer screening and early detection. Hum. Genom. 2019, 13, 34. [Google Scholar] [CrossRef] [PubMed]
- Bieńkowski, M.; Pęksa, R.; Popęda, M.; Kołaczkowska, M.; Frankiewicz, A.; Żaczek, A.J.; Gruchała, M.; Biernat, W.; Siondalski, P. Liquid biopsy for minimally invasive heart transplant monitoring: A pilot study. J. Clin. Pathol. 2020, 73, 507–510. [Google Scholar] [CrossRef]
- Calandri, M.; Siravegna, G.; Yevich, S.M.; Stranieri, G.; Gazzera, C.; Kopetz, S.; Fonio, P.; Gupta, S.; Bardelli, A.; Veltri, A.; et al. Liquid biopsy, a paradigm shift in oncology: What interventional radiologists should know. Eur. Radiol. 2020, 30, 4496–4503. [Google Scholar] [CrossRef]
- Liu, C.; Xiang, X.; Han, S.; Lim, H.Y.; Li, L.; Zhang, X.; Ma, Z.; Yang, L.; Guo, S.; Soo, R.; et al. Blood-based liquid biopsy: Insights into early detection and clinical management of lung cancer. Cancer Lett. 2022, 524, 91–102. [Google Scholar] [CrossRef]
- Li, G.; Zhao, A.; Péoch, M.; Cottier, M.; Mottet, N. Detection of urinary cell-free miR-210 as a potential tool of liquid biopsy for clear cell renal cell carcinoma. Urol. Oncol. Semin. Orig. Investig. 2017, 35, 294–299. [Google Scholar] [CrossRef]
- Tu, M.; Cheng, J.; Chen, Y.-L.; Jea, W.-C.; Chen, W.-L.; Chen, C.-J.; Ho, C.-L.; Huang, W.-L.; Lin, C.-C.; Su, W.-C.; et al. Electric Field–Induced Release and Measurement (EFIRM): Characterization and Technical Validation of a Novel Liquid Biopsy Platform in Plasma and Saliva. J. Mol. Diagn. 2020, 22, 1050–1062. [Google Scholar] [CrossRef]
- Le Rhun, E.; Seoane, J.; Salzet, M.; Soffietti, R.; Weller, M. Liquid biopsies for diagnosing and monitoring primary tumors of the central nervous system. Cancer Lett. 2020, 480, 24–28. [Google Scholar] [CrossRef] [PubMed]
- von Baumgarten, L.; Kumbrink, J.; Jung, A.; Reischer, A.; Flach, M.; Liebmann, S.; Metzeler, K.H.; Holch, J.W.; Niyazi, M.; Thon, N.; et al. Therapeutic management of neuro-oncologic patients—Potential relevance of CSF liquid biopsy. Theranostics 2020, 10, 856–866. [Google Scholar] [CrossRef] [PubMed]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Prakash, S.; Yeom, J. Chapter 2—Fundamentals for Microscale and Nanoscale Flows. In Nanofluidics and Microfluidics, Prakash, S., Yeom, J., Eds.; William Andrew Publishing: Norwich, NY, USA, 2014; pp. 9–38. [Google Scholar]
- Yamamoto, K.; Ota, N.; Tanaka, Y. Nanofluidic Devices and Applications for Biological Analyses. Anal. Chem. 2021, 93, 332–349. [Google Scholar] [CrossRef] [PubMed]
- Prakash, S.; Piruska, A.; Gatimu, E.N.; Bohn, P.W.; Sweedler, J.V.; Shannon, M.A. Nanofluidics: Systems and Applications. IEEE Sens. J. 2008, 8, 441–450. [Google Scholar] [CrossRef]
- Convery, N.; Gadegaard, N. 30 years of microfluidics. Micro Nano Eng. 2019, 2, 76–91. [Google Scholar] [CrossRef]
- Breadmore, M.C.; Wolfe, K.A.; Arcibal, I.G.; Leung, W.K.; Dickson, D.; Giordano, B.C.; Power, M.E.; Ferrance, J.P.; Feldman, S.H.; Norris, P.M.; et al. Microchip-Based Purification of DNA from Biological Samples. Anal. Chem. 2003, 75, 1880–1886. [Google Scholar] [CrossRef]
- He, M.; Crow, J.; Roth, M.; Zeng, Y.; Godwin, A.K. Integrated immunoisolation and protein analysis of circulating exosomes using microfluidic technology. Lab A Chip 2014, 14, 3773–3780. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Yang, Y.; Zeng, Y.; He, M. A microfluidic ExoSearch chip for multiplexed exosome detection towards blood-based ovarian cancer diagnosis. Lab A Chip 2016, 16, 489–496. [Google Scholar] [CrossRef]
- Kang, Y.-T.; Purcell, E.; Palacios-Rolston, C.; Lo, T.-W.; Ramnath, N.; Jolly, S.; Nagrath, S. Isolation and Profiling of Circulating Tumor-Associated Exosomes Using Extracellular Vesicular Lipid–Protein Binding Affinity Based Microfluidic Device. Small 2019, 15, 1903600. [Google Scholar] [CrossRef] [PubMed]
- Lamanda, A.; Lu, Y.; Gill, N.; Wong, P.K. An electrokinetic microdevice for isolation and quantification of circulating cell-free DNA from physiological samples. In Proceedings of the 2015 Transducers—2015 18th International Conference on Solid-State Sensors, Actuators and Microsystems (TRANSDUCERS), Anchorage, AK, USA, 21–25 June 2015; pp. 544–547. [Google Scholar]
- Reyes, D.R.; Iossifidis, D.; Auroux, P.-A.; Manz, A. Micro Total Analysis Systems. 1. Introduction, Theory, and Technology. Anal. Chem. 2002, 74, 2623–2636. [Google Scholar] [CrossRef] [PubMed]
- Auroux, P.-A.; Iossifidis, D.; Reyes, D.R.; Manz, A. Micro Total Analysis Systems. 2. Analytical Standard Operations and Applications. Anal. Chem. 2002, 74, 2637–2652. [Google Scholar] [CrossRef]
- Nayak, S.; Blumenfeld, N.R.; Laksanasopin, T.; Sia, S.K. Point-of-Care Diagnostics: Recent Developments in a Connected Age. Anal. Chem. 2017, 89, 102–123. [Google Scholar] [CrossRef] [PubMed]
- Dincer, C.; Bruch, R.; Kling, A.; Dittrich, P.S.; Urban, G.A. Multiplexed Point-of-Care Testing—xPOCT. Trends Biotechnol. 2017, 35, 728–742. [Google Scholar] [CrossRef]
- Oey, O.; Ghaffari, M.; Li, J.J.; Hosseini-Beheshti, E. Application of extracellular vesicles in the diagnosis and treatment of prostate cancer: Implications for clinical practice. Crit. Rev. Oncol. Hematol. 2021, 167, 103495. [Google Scholar] [CrossRef]
- Iliescu, F.S.; Vrtačnik, D.; Neuzil, P.; Iliescu, C. Microfluidic Technology for Clinical Applications of Exosomes. Micromachines 2019, 10, 392. [Google Scholar] [CrossRef] [PubMed]
- Maas, S.L.N.; Breakefield, X.O.; Weaver, A.M. Extracellular Vesicles: Unique Intercellular Delivery Vehicles. Membr. Biol. 2017, 27, 172–188. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Skog, J.; Würdinger, T.; Van Rijn, S.; Meijer, D.H.; Gainche, L.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.S.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- van Niel, G.; D′Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Simons, M.; Raposo, G. Exosomes—vesicular carriers for intercellular communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Xu, M.; Jing, C.; Wu, X.; Chen, X.; Zhang, W. Extracellular vesicle-derived miR-320a targets ZC3H12B to inhibit tumorigenesis, invasion, and angiogenesis in ovarian cancer. Discov. Oncol. 2021, 12, 51. [Google Scholar] [CrossRef]
- Melo, S.A.; Sugimoto, H.; O′Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef]
- Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Investig. 2016, 126, 1208–1215. [Google Scholar] [CrossRef] [PubMed]
- Azadi, S.; Aboulkheyr Es, H.; Kulasinghe, A.; Bordhan, P.; Ebrahimi Warkiani, M. Chapter Five—Application of microfluidic technology in cancer research and therapy. In Advances in Clinical Chemistry; Makowski, G.S., Ed.; Elsevier: Amsterdam, The Netherlands, 2020; Volume 99, pp. 193–235. [Google Scholar]
- Xu, Y.; Zhang, Y.; Wang, L.; Zhao, R.; Qiao, Y.; Han, D.; Sun, Q.; Dong, N.; Liu, Y.; Wu, D.; et al. miR-200a targets Gelsolin: A novel mechanism regulating secretion of microvesicles in hepatocellular carcinoma cells. Oncol. Rep. 2017, 37, 2711–2719. [Google Scholar] [CrossRef] [PubMed]
- Bebelman, M.P.; Smit, M.J.; Pegtel, D.M.; Baglio, S.R. Biogenesis and function of extracellular vesicles in cancer. Pharmacol. Ther. 2018, 188, 1–11. [Google Scholar] [CrossRef]
- Maacha, S.; Bhat, A.A.; Jimenez, L.; Raza, A.; Haris, M.; Uddin, S.; Grivel, J.-C. Extracellular vesicles-mediated intercellular communication: Roles in the tumor microenvironment and anti-cancer drug resistance. Mol. Cancer 2019, 18, 55. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, M.; Martin-Jaular, L.; Lavieu, G.; Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 2019, 21, 9–17. [Google Scholar] [CrossRef]
- Gonda, A.; Kabagwira, J.; Senthil, G.N.; Wall, N.R. Internalization of Exosomes through Receptor-Mediated Endocytosis. Mol. Cancer Res. 2019, 17, 337. [Google Scholar] [CrossRef]
- Giusti, I.; Di Francesco, M.; D’Ascenzo, S.; Palmerini, M.G.; Macchiarelli, G.; Carta, G.; Dolo, V. Ovarian cancer-derived extracellular vesicles affect normal human fibroblast behavior. Cancer Biol. Ther. 2018, 19, 722–734. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.P.; Spary, L.K.; Sanders, A.J.; Chowdhury, R.; Jiang, W.G.; Steadman, R.; Wymant, J.; Jones, A.T.; Kynaston, H.; Mason, M.D.; et al. Differentiation of tumour-promoting stromal myofibroblasts by cancer exosomes. Oncogene 2015, 34, 290–302. [Google Scholar] [CrossRef] [PubMed]
- Hosseini-Beheshti, E.; Choi, W.; Weiswald, L.-B.; Kharmate, G.; Ghaffari, M.; Roshan-Moniri, M.; Hassona, M.D.; Chan, L.; Chin, M.Y.; Tai, I.T.; et al. Exosomes confer pro-survival signals to alter the phenotype of prostate cells in their surrounding environment. Oncotarget 2016, 7, 14639. [Google Scholar] [CrossRef]
- van Niel, G.; Porto-Carreiro, I.; Simoes, S.; Raposo, G. Exosomes: A common pathway for a specialized function. J. Biochem. 2006, 140, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Głuszko, A.; Szczepański, M.J.; Ludwig, N.; Mirza, S.M.; Olejarz, W. Exosomes in Cancer: Circulating Immune-Related Biomarkers. BioMed Res. Int. 2019, 2019, 1628029. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Tan, Z.; Lubman, D. Exosome enrichment of human serum using multiple cycles of centrifugation. Electrophoresis 2015, 36, 2017–2026. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Gong, M.; Hu, Y.; Liu, H.; Zhang, W.; Zhang, M.; Hu, X.; Aubert, D.; Zhu, S.; Wu, L.; et al. Quality and efficiency assessment of six extracellular vesicle isolation methods by nano-flow cytometry. J. Extracell Vesicles 2019, 9, 1697028. [Google Scholar] [CrossRef]
- Tang, Y.-T.; Huang, Y.-Y.; Zheng, L.; Qin, S.-H.; Xu, X.-P.; An, T.-X.; Xu, Y.; Qu, Y.-S.; Hu, X.-M.; Ping, B.-H.; et al. Comparison of isolation methods of exosomes and exosomal RNA from cell culture medium and serum. Int. J. Mol. Med 2017, 40, 834–844. [Google Scholar] [CrossRef] [PubMed]
- Cvjetkovic, A.; Lötvall, J.; Lässer, C. The influence of rotor type and centrifugation time on the yield and purity of extracellular vesicles. J. Extracell Vesicles 2014, 3, 23111. [Google Scholar] [CrossRef] [PubMed]
- Brennan, K.; Martin, K.; FitzGerald, S.P.; O’ Sullivan, J.; Wu, Y.; Blanco, A.; Richardson, C.; McGee, M.M. A comparison of methods for the isolation and separation of extracellular vesicles from protein and lipid particles in human serum. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef]
- Webber, J.; Clayton, A. How pure are your vesicles? J. Extracell Vesicles 2013, 2, 19861. [Google Scholar] [CrossRef] [PubMed]
- Monguió-Tortajada, M.; Gálvez-Montón, C.; Bayes-Genis, A.; Roura, S.; Borrás, F. Extracellular vesicle isolation methods: Rising impact of size-exclusion chromatography. Cell. Mol. Life Sci. 2019, 76, 2369–2382. [Google Scholar] [CrossRef] [PubMed]
- Sidhom, K.; Obi, P.; Saleem, A. A Review of Exosomal Isolation Methods: Is Size Exclusion Chromatography the Best Option? Int. J. Mol. Sci. 2020, 21, 6466. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-‘t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell Vesicles 2013, 2, 20360. [Google Scholar] [CrossRef]
- Momen-Heravi, F.; Balaj, L.; Alian, S.; Trachtenberg, A.; Hochberg, F.; Skog, J.; Kuo, W. Impact of Biofluid Viscosity on Size and Sedimentation Efficiency of the Isolated Microvesicles. Front. Physiol. 2012, 3, 162. [Google Scholar] [CrossRef] [PubMed]
- Liangsupree, T.; Multia, E.; Riekkola, M.-L. Modern isolation and separation techniques for extracellular vesicles. J. Chromatogr. A 2021, 1636, 461773. [Google Scholar] [CrossRef]
- Gámez-Valero, A.; Monguió-Tortajada, M.; Carreras-Planella, L.; Franquesa, M.l.; Beyer, K.; Borràs, F.E. Size-Exclusion Chromatography-based isolation minimally alters Extracellular Vesicles’ characteristics compared to precipitating agents. Sci. Rep. 2016, 6, 33641. [Google Scholar] [CrossRef] [PubMed]
- Stranska, R.; Gysbrechts, L.; Wouters, J.; Vermeersch, P.; Bloch, K.; Dierickx, D.; Andrei, G.; Snoeck, R. Comparison of membrane affinity-based method with size-exclusion chromatography for isolation of exosome-like vesicles from human plasma. J. Transl. Med. 2018, 16, 1. [Google Scholar] [CrossRef] [PubMed]
- Askeland, A.; Borup, A.; Østergaard, O.; Olsen, J.V.; Lund, S.M.; Christiansen, G.; Kristensen, S.R.; Heegaard, N.H.H.; Pedersen, S. Mass-Spectrometry Based Proteome Comparison of Extracellular Vesicle Isolation Methods: Comparison of ME-kit, Size-Exclusion Chromatography, and High-Speed Centrifugation. Biomedicines 2020, 8, 246. [Google Scholar] [CrossRef] [PubMed]
- Niu, Z.; Pang, R.; Liu, W.; Li, Q.; Cheng, R.; Yeung, W. Polymer-based precipitation preserves biological activities of extracellular vesicles from an endometrial cell line. PLoS ONE 2017, 12, e0186534. [Google Scholar] [CrossRef] [PubMed]
- Konoshenko, M.Y.; Lekchnov, E.A.; Vlassov, A.V.; Laktionov, P.P. Isolation of Extracellular Vesicles: General Methodologies and Latest Trends. BioMed Res. Int. 2018, 2018, 8545347. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.; Yin, H. Polymer-Based Purification of Extracellular Vesicles. Methods Mol. Biol. 2017, 1660, 91–103. [Google Scholar]
- Taylor, D.D.; Shah, S. Methods of isolating extracellular vesicles impact down-stream analyses of their cargoes. Methods 2015, 87, 3–10. [Google Scholar] [CrossRef]
- Taylor, D.D.; Zacharias, W.; Gercel-Taylor, C. Exosome isolation for proteomic analyses and RNA profiling. Methods Mol. Biol. 2011, 728, 235–246. [Google Scholar] [CrossRef] [PubMed]
- Lobb, R.J.; Becker, M.; Wen, S.W.; Wong, C.S.; Wiegmans, A.P.; Leimgruber, A.; Möller, A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J. Extracell Vesicles 2015, 4, 27031. [Google Scholar] [CrossRef] [PubMed]
- Van Deun, J.; Mestdagh, P.; Sormunen, R.; Cocquyt, V.; Vermaelen, K.; Vandesompele, J.; Bracke, M.; De Wever, O.; Hendrix, A. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J. Extracell Vesicles 2014, 3, 24858. [Google Scholar] [CrossRef] [PubMed]
- Andreu, Z.; Rivas, E.; Sanguino-Pascual, A.; Lamana, A.; Marazuela, M.; González-Alvaro, I.; Sánchez-Madrid, F.; de la Fuente, H.; Yáñez-Mó, M. Comparative analysis of EV isolation procedures for miRNAs detection in serum samples. J. Extracell Vesicles 2016, 5, 31655. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Inoshima, Y.; Matsuda, T.; Ishiguro, N. Comparison of Methods for Isolating Exosomes from Bovine Milk. J. Vet. Med. Sci. 2012, 74, 1523–1525. [Google Scholar] [CrossRef] [PubMed]
- Kahlert, C. Liquid Biopsy: Is There an Advantage to Analyzing Circulating Exosomal DNA Compared to cfDNA or Are They the Same? Cancer Res. 2019, 79, 2462. [Google Scholar] [CrossRef] [PubMed]
- Corcoran, R.B.; Chabner, B.A. Application of Cell-free DNA Analysis to Cancer Treatment. N. Engl. J. Med. 2018, 379, 1754–1765. [Google Scholar] [CrossRef] [PubMed]
- Mouliere, F.; Chandrananda, D.; Piskorz Anna, M.; Moore Elizabeth, K.; Morris, J.; Ahlborn Lise, B.; Mair, R.; Goranova, T.; Marass, F.; Heider, K.; et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci. Transl. Med. 2018, 10, eaat4921. [Google Scholar] [CrossRef] [PubMed]
- van den Berg, M.M.J.; Krauskopf, J.; Ramaekers, J.G.; Kleinjans, J.C.S.; Prickaerts, J.; Briedé, J.J. Circulating microRNAs as potential biomarkers for psychiatric and neurodegenerative disorders. Prog. Neurobiol. 2020, 185, 101732. [Google Scholar] [CrossRef]
- Lee, J.-S.; Park, S.S.; Lee, Y.K.; Norton, J.A.; Jeffrey, S.S. Liquid biopsy in pancreatic ductal adenocarcinoma: Current status of circulating tumor cells and circulating tumor DNA. Mol. Oncol. 2019, 13, 1623–1650. [Google Scholar] [CrossRef]
- Zhang, Y.; Tong, X.; Yang, L.; Yin, R.; Li, Y.; Zeng, D.; Wang, X.; Deng, K. A herringbone mixer based microfluidic device HBEXO-chip for purifying tumor-derived exosomes and establishing miRNA signature in pancreatic cancer. Sens. Actuators B Chem. 2021, 332, 129511. [Google Scholar] [CrossRef]
- Song, Z.; Mao, J.; Barrero, R.A.; Wang, P.; Zhang, F.; Wang, T. Development of a CD63 Aptamer for Efficient Cancer Immunochemistry and Immunoaffinity-Based Exosome Isolation. Molecules 2020, 25, 5585. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.-T.; Hadlock, T.; Lo, T.-W.; Purcell, E.; Mutukuri, A.; Fouladdel, S.; Raguera, M.D.S.; Fairbairn, H.; Murlidhar, V.; Durham, A.; et al. Dual-Isolation and Profiling of Circulating Tumor Cells and Cancer Exosomes from Blood Samples with Melanoma Using Immunoaffinity-Based Microfluidic Interfaces. Adv. Sci. 2020, 7, 2001581. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Ludwig, S.; Muller, L.; Hong, C.S.; Kirkwood, J.M.; Ferrone, S.; Whiteside, T.L. Immunoaffinity-based isolation of melanoma cell-derived exosomes from plasma of patients with melanoma. J. Extracell Vesicles 2018, 7, 1435138. [Google Scholar] [CrossRef]
- Liang, L.-G.; Sheng, Y.-F.; Zhou, S.; Inci, F.; Li, L.; Demirci, U.; Wang, S. An Integrated Double-Filtration Microfluidic Device for Detection of Extracellular Vesicles from Urine for Bladder Cancer Diagnosis. In Extracellular Vesicles: Methods and Protocols; Kuo, W.P., Jia, S., Eds.; Springer: New York, NY, USA, 2017; pp. 355–364. [Google Scholar]
- Wang, Z.; Wu, H.-j.; Fine, D.; Schmulen, J.; Hu, Y.; Godin, B.; Zhang, J.X.J.; Liu, X. Ciliated micropillars for the microfluidic-based isolation of nanoscale lipid vesicles. Lab A Chip 2013, 13, 2879–2882. [Google Scholar] [CrossRef]
- Casadei, L.; Choudhury, A.; Sarchet, P.; Mohana Sundaram, P.; Lopez, G.; Braggio, D.; Balakirsky, G.; Pollock, R.; Prakash, S. Cross-flow microfiltration for isolation, selective capture and release of liposarcoma extracellular vesicles. J. Extracell Vesicles 2021, 10, e12062. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, F.; Rufo, J.; Chen, C.; Yang, S.; Li, L.; Zhang, J.; Cheng, J.; Kim, Y.; Wu, M.; et al. Acoustofluidic Salivary Exosome Isolation: A Liquid Biopsy Compatible Approach for Human Papillomavirus-Associated Oropharyngeal Cancer Detection. J. Mol. Diagn. 2020, 22, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Ouyang, Y.; Wang, Z.; Zhang, R.; Huang, P.H.; Chen, C.; Li, H.; Li, P.; Quinn, D.; Dao, M.; et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc. Natl. Acad. Sci. USA 2017, 114, 10584–10589. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Chen, C.; Mao, Z.; Bachman, H.; Becker, R.; Rufo, J.; Wang, Z.; Zhang, P.; Mai, J.; Yang, S.; et al. Acoustofluidic centrifuge for nanoparticle enrichment and separation. Sci. Adv. 2021, 7, eabc0467. [Google Scholar] [CrossRef] [PubMed]
- Ku, A.; Lim, H.C.; Evander, M.; Lilja, H.; Laurell, T.; Scheding, S.; Ceder, Y. Acoustic Enrichment of Extracellular Vesicles from Biological Fluids. Anal. Chem. 2018, 90, 8011–8019. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Shao, H.; Weissleder, R.; Lee, H. Acoustic Purification of Extracellular Microvesicles. ACS Nano 2015, 9, 2321–2327. [Google Scholar] [CrossRef]
- Liu, C.; Guo, J.; Tian, F.; Yang, N.; Yan, F.; Ding, Y.; Wei, J.; Hu, G.; Nie, G.; Sun, J. Field-Free Isolation of Exosomes from Extracellular Vesicles by Microfluidic Viscoelastic Flows. ACS Nano 2017, 11, 6968–6976. [Google Scholar] [CrossRef] [PubMed]
- Akagi, T.; Kato, K.; Kobayashi, M.; Kosaka, N.; Ochiya, T.; Ichiki, T. On-Chip Immunoelectrophoresis of Extracellular Vesicles Released from Human Breast Cancer Cells. PLoS ONE 2015, 10, e0123603. [Google Scholar] [CrossRef] [PubMed]
- Peoples, M.C.; Karnes, H.T. Microfluidic immunoaffinity separations for bioanalysis. J. Chromatogr. B 2008, 866, 14–25. [Google Scholar] [CrossRef]
- Moser, A.C.; Hage, D.S. Immunoaffinity chromatography: An introduction to applications and recent developments. Bioanalysis 2010, 2, 769–790. [Google Scholar] [CrossRef]
- Prakash, S.; Karacor, M.B.; Banerjee, S. Surface modification in microsystems and nanosystems. Surf. Sci. Rep. 2009, 64, 233–254. [Google Scholar] [CrossRef]
- Wu, Y.; Misra, S.; Karacor, M.B.; Prakash, S.; Shannon, M.A. Dynamic Response of AFM Cantilevers to Dissimilar Functionalized Silica Surfaces in Aqueous Electrolyte Solutions. Langmuir 2010, 26, 16963–16972. [Google Scholar] [CrossRef] [PubMed]
- Lo, T.-W.; Zhu, Z.; Purcell, E.; Watza, D.; Wang, J.; Kang, Y.-T.; Jolly, S.; Nagrath, D.; Nagrath, S. Microfluidic device for high-throughput affinity-based isolation of extracellular vesicles. Lab A Chip 2020, 20, 1762–1770. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.; Kim, J.; Park, J. Methods to isolate extracellular vesicles for diagnosis. Micro Nano Syst. Lett. 2017, 5, 15. [Google Scholar] [CrossRef]
- Sunkara, V.; Woo, H.-K.; Cho, Y.-K. Emerging techniques in the isolation and characterization of extracellular vesicles and their roles in cancer diagnostics and prognostics. Analyst 2016, 141, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro-Samy, S.; Oliveira, M.I.; Pereira-Veiga, T.; Muinelo-Romay, L.; Carvalho, S.; Gaspar, J.; Freitas, P.P.; López-López, R.; Costa, C.; Diéquez, L. Fast and efficient microfluidic cell filter for isolation of circulating tumor cells from unprocessed whole blood of colorectal cancer patients. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Doh, I.; Yoo, H.-i.; Cho, Y.-H.; Lee, J.; Kim, H.K.; Kim, J. Viable capture and release of cancer cells in human whole blood. Appl. Phys. Lett. 2012, 101, 043701. [Google Scholar] [CrossRef]
- Zhang, P.; Bachman, H.; Ozcelik, A.; Huang, T.J. Acoustic Microfluidics. Annu. Rev. Anal. Chem. 2020, 13, 17–43. [Google Scholar] [CrossRef]
- Li, P.; Huang, T.J. Applications of Acoustofluidics in Bioanalytical Chemistry. Anal. Chem. 2019, 91, 757–767. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Li, P.; Lin, S.-C.S.; Stratton, Z.S.; Nama, N.; Guo, F.; Slotcavage, D.; Mao, X.; Shi, J.; Costanzo, F.; et al. Surface acoustic wave microfluidics. Lab A Chip 2013, 13, 3626–3649. [Google Scholar] [CrossRef]
- Hahn, P.; Leibacher, I.; Baasch, T.; Dual, J. Numerical simulation of acoustofluidic manipulation by radiation forces and acoustic streaming for complex particles. Lab A Chip 2015, 15, 4302–4313. [Google Scholar] [CrossRef] [PubMed]
- Wiklund, M.; Green, R.; Ohlin, M. Acoustofluidics 14: Applications of acoustic streaming in microfluidic devices. Lab A Chip 2012, 12, 2438–2451. [Google Scholar] [CrossRef] [PubMed]
- Connacher, W.; Zhang, N.; Huang, A.; Mei, J.; Zhang, S.; Gopesh, T.; Friend, J. Micro/nano acoustofluidics: Materials, phenomena, design, devices, and applications. Lab A Chip 2018, 18, 1952–1996. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Papautsky, I. Viscoelastic microfluidics: Progress and challenges. Microsyst. Nanoeng. 2020, 6, 113. [Google Scholar] [CrossRef] [PubMed]
- Asghari, M.; Cao, X.; Mateescu, B.; van Leeuwen, D.; Aslan, M.K.; Stavrakis, S.; deMello, A.J. Oscillatory Viscoelastic Microfluidics for Efficient Focusing and Separation of Nanoscale Species. ACS Nano 2020, 14, 422–433. [Google Scholar] [CrossRef]
- Zhou, J.; Kulasinghe, A.; Bogseth, A.; O’Byrne, K.; Punyadeera, C.; Papautsky, I. Isolation of circulating tumor cells in non-small-cell-lung-cancer patients using a multi-flow microfluidic channel. Microsyst. Nanoeng. 2019, 5, 8. [Google Scholar] [CrossRef] [PubMed]
- Gossett, D.R.; Weaver, W.M.; Mach, A.J.; Hur, S.C.; Tse, H.T.K.; Lee, W.; Amini, H.; Di Carlo, D. Label-free cell separation and sorting in microfluidic systems. Anal. Bioanal. Chem. 2010, 397, 3249–3267. [Google Scholar] [CrossRef] [PubMed]
- Lochab, V.; Prakash, S. Combined electrokinetic and shear flows control colloidal particle distribution across microchannel cross-sections. Soft Matter 2021, 17, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Lochab, V.; Yee, A.; Yoda, M.; Conlisk, A.T.; Prakash, S. Dynamics of colloidal particles in microchannels under combined pressure and electric potential gradients. Microfluid. Nanofluidics 2019, 23, 134. [Google Scholar] [CrossRef]
- Dabros, T. Electrokinetic and Colloid Transport Phenomena: Jacob, H. Masliyah and Subir Bhattacharjee Publisher: Wiley-Interscience, 2006 ISBN: 0471799734. Can. J. Chem. Eng. 2006, 84, 729. [Google Scholar] [CrossRef]
- Zhao, C.; Yang, C. Advances in electrokinetics and their applications in micro/nano fluidics. Microfluid. Nanofluidics 2012, 13, 179–203. [Google Scholar] [CrossRef]
- Voldman, J. Electrical forces for microscale cell manipulation. Annu. Rev. Biomed. Eng. 2006, 8, 425–454. [Google Scholar] [CrossRef] [PubMed]
- Ghosal, S.; Chen, Z. Electromigration dispersion in a capillary in the presence of electro-osmotic flow. J. Fluid Mech. 2012, 697, 436–454. [Google Scholar] [CrossRef]
- Zhao, W.; Zhang, L.; Ye, Y.; Li, Y.; Luan, X.; Liu, J.; Cheng, J.; Zhao, Y.; Li, M.; Huang, C. Microsphere mediated exosome isolation and ultra-sensitive detection on a dielectrophoresis integrated microfluidic device. Analyst 2021, 146, 5962–5972. [Google Scholar] [CrossRef]
- Pak Kin, W.; Tza-Huei, W.; Deval, J.H.; Chih-Ming, H. Electrokinetics in micro devices for biotechnology applications. IEEE/ASME Trans. Mechatron. 2004, 9, 366–376. [Google Scholar] [CrossRef]
- Krishnan, R.; Sullivan, B.D.; Mifflin, R.L.; Esener, S.C.; Heller, M.J. Alternating current electrokinetic separation and detection of DNA nanoparticles in high-conductance solutions. Electrophoresis 2008, 29, 1765–1774. [Google Scholar] [CrossRef] [PubMed]
- Cheng, I.F.; Chen, T.Y.; Chang, H.C. Electrokinetics-Based Microfluidic Technology for the Rapid Separation and Concentration of Bacteria/Cells/Biomolecules. Adv. Mater. Res. 2014, 911, 347–351. [Google Scholar] [CrossRef]
- Diaz-Armas, G.G.; Cervantes-Gonzalez, A.P.; Martinez-Duarte, R.; Perez-Gonzalez, V.H. Electrically driven microfluidic platforms for exosome manipulation and characterization. Electrophoresis 2021, 43, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Romero-Soto, F.O.; Polanco-Oliva, M.I.; Gallo-Villanueva, R.C.; Martinez-Chapa, S.O.; Perez-Gonzalez, V.H. A survey of electrokinetically-driven microfluidics for cancer cells manipulation. Electrophoresis 2021, 42, 605–625. [Google Scholar] [CrossRef] [PubMed]
- Aïzel, K.; Fouillet, Y.; Pudda, C. Electropreconcentration of nanoparticles using a radial micro-nanofluidic device. J. Nanoparticle Res. 2014, 16, 2731. [Google Scholar] [CrossRef]
- Lu, Y.; Liu, T.; Lamanda, A.C.; Sin, M.L.Y.; Gau, V.; Liao, J.C.; Wong, P.K. AC Electrokinetics of Physiological Fluids for Biomedical Applications. J. Lab. Autom. 2014, 20, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Dey, R.; Shaik, V.A.; Chakraborty, D.; Ghosal, S.; Chakraborty, S. AC Electric Field-Induced Trapping of Microparticles in Pinched Microconfinements. Langmuir 2015, 31, 5952–5961. [Google Scholar] [CrossRef] [PubMed]
- Kato, K.; Kobayashi, M.; Hanamura, N.; Akagi, T.; Kosaka, N.; Ochiya, T.; Ichiki, T. Electrokinetic Evaluation of Individual Exosomes by On-Chip Microcapillary Electrophoresis with Laser Dark-Field Microscopy. Jpn. J. Appl. Phys. 2013, 52, 06GK10. [Google Scholar] [CrossRef]
- Akagi, T.; Kato, K.; Hanamura, N.; Kobayashi, M.; Ichiki, T. Evaluation of desialylation effect on zeta potential of extracellular vesicles secreted from human prostate cancer cells by on-chip microcapillary electrophoresis. Jpn. J. Appl. Phys. 2014, 53, 06JL01. [Google Scholar] [CrossRef]
| Isolation Method | Operating Principle | Advantages | Study | Isolation Efficiency | Throughput |
|---|---|---|---|---|---|
| Centrifugation | Spinning results in separation and pellet formation | High purity | Use of multiple centrifugation cycles for exosome enrichment from human serum [62] | - | - |
| Size exclusion | Particles separated based on size | Minimal impact on size and features | Quality and efficiency assessment of qEV using nano-flow cytometry [63] | 67.7 ± 13.1% [63] | - |
| Polymer-based precipitation | Precipitation of lower solubility components of sample out of solution | Time efficient; requires minimal equipment | Quality and efficiency assessment of ExoQuick isolation kit using nano-flow cytometry [63] | ~82% [63] | - |
| Isolation Method | Operating Principle | Advantages | Study | Isolation Efficiency | Throughput |
|---|---|---|---|---|---|
| Immunoaffinity | Interactions between antibodies and antigens | High specificity | HBEXO-chip for purifying tumor-derived exosomes and establishing miRNA signature in pancreatic cancer with GPC1+exosomes as biomarkers [90] | ~75% [90] | - |
| Using CD-63-1 aptamer for the isolation of EVs (50–150 nm) from CD-63 positive tumor samples [91] | - | - | |||
| OncoBean (DUO) using melanoma-specific antibodies MCAM and MCSP for exosome isolation [92] | - | - | |||
| Immunoaffinity-based isolation of melanoma cell-derived exosomes from plasma of patients with melanoma with CSPG4-specific mAb 763.74 as biomarker [93] | - | - | |||
| Filtration | Difference in particle size population | No need for external actuation; easy to use | Isolation of bladder cancer EVs from urine samples using integrated double-filtration device [94] | 74.2% [94] | - |
| Isolation of exosome-like lipid vesicles via a ciliated micropillar device [95] | 60% (83 nm lipid vesicles), 45% (120 nm lipid vesicles) [95] | 3 μL/min [95] | |||
| Isolation and capture of EVs from liposarcoma cell-conditioned media (LCCM) and dedifferentiated liposarcoma patient serum, with MDM2 and CD-63 as biomarkers [96] | 76% (LCCM), 36% (dedifferentiated liposarcoma patient serum) [96] | 10 μL/min [96] | |||
| Acoustofluidics | Acoustic waves | Biocompatibility, versatility, precision, flexibility | Isolation of salivary exosomes from Human papilloma viral (HPV)- associated oropharyngeal cancer patients with HPV DNA as biomarker [97] | - | - |
| Isolating exosomes directly from undiluted human blood [98] | 82% [98] | 4 μL/min [98] | |||
| Nanoparticle enrichment and separation using acoustic centrifugation [99] | - | - | |||
| Acoustic trapping for the enrichment of EVs from cell culture conditioned media, urine, and blood plasma from healthy volunteers [100] | - | 10 μL/min [100] | |||
| Separation of exosomes using acoustic nanofilter system [101] | >90% [101] | - | |||
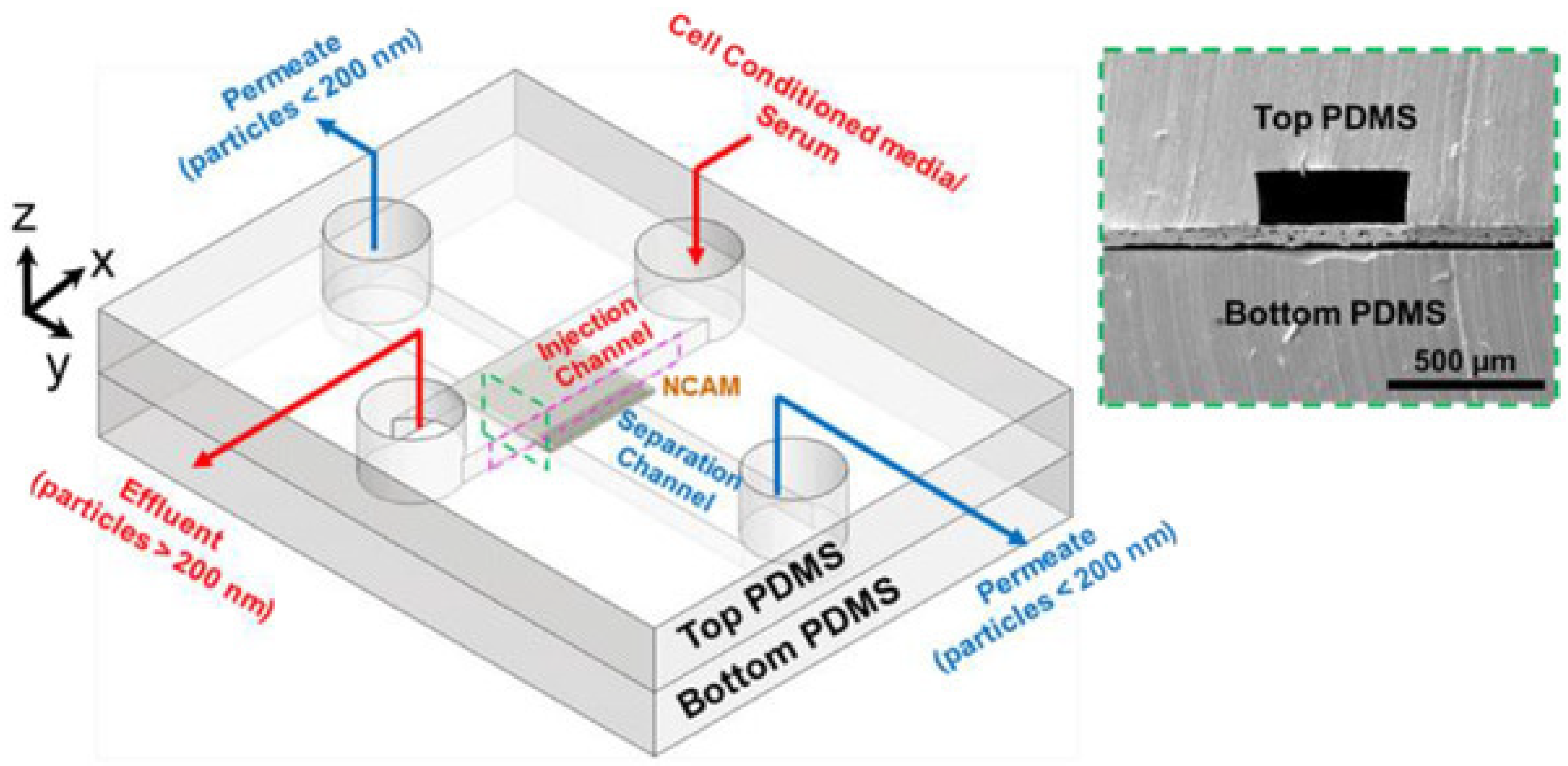
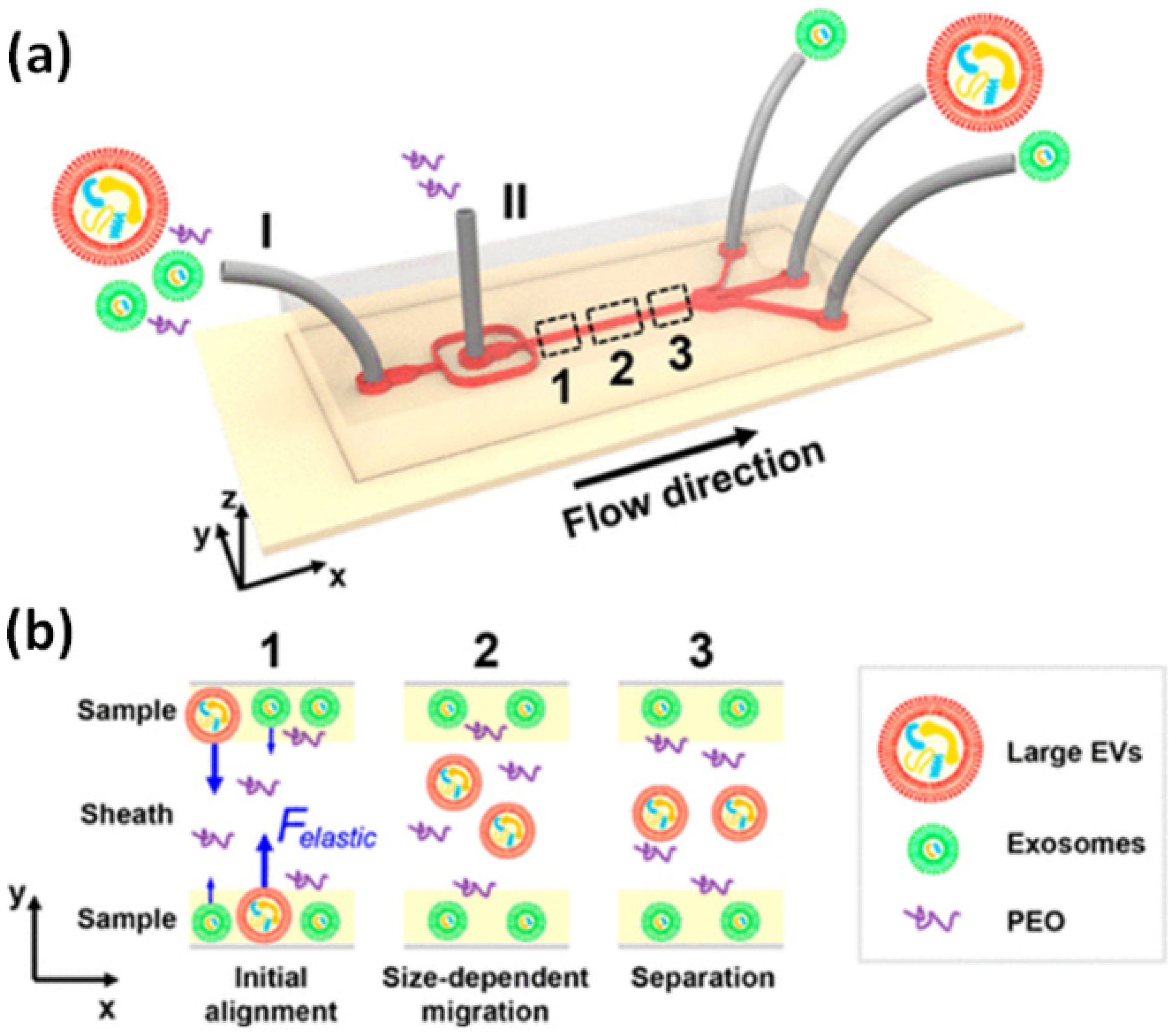
| Viscoelastic flow | Imbalance of normal forces in a non-Newtonian medium | Ease of use, no requirements for external actuation, robust performance once operational parameters are optimized | Separation of exosomes from cell culture media and serum of adenocarcinomic human alveolar basal epithelial cells [102] | >80% [102] | 200 μL/h [102] |
| Electrokinetics | Charge of the particle and electrolyte | Strong actuation force due to linear scaling law | On-chip microcapillary electrophoresis for separation of human breast cancer derived exosomes [103] | - | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singh, P.K.; Patel, A.; Kaffenes, A.; Hord, C.; Kesterson, D.; Prakash, S. Microfluidic Approaches and Methods Enabling Extracellular Vesicle Isolation for Cancer Diagnostics. Micromachines 2022, 13, 139. https://doi.org/10.3390/mi13010139
Singh PK, Patel A, Kaffenes A, Hord C, Kesterson D, Prakash S. Microfluidic Approaches and Methods Enabling Extracellular Vesicle Isolation for Cancer Diagnostics. Micromachines. 2022; 13(1):139. https://doi.org/10.3390/mi13010139
Chicago/Turabian StyleSingh, Premanshu Kumar, Aarti Patel, Anastasia Kaffenes, Catherine Hord, Delaney Kesterson, and Shaurya Prakash. 2022. "Microfluidic Approaches and Methods Enabling Extracellular Vesicle Isolation for Cancer Diagnostics" Micromachines 13, no. 1: 139. https://doi.org/10.3390/mi13010139
APA StyleSingh, P. K., Patel, A., Kaffenes, A., Hord, C., Kesterson, D., & Prakash, S. (2022). Microfluidic Approaches and Methods Enabling Extracellular Vesicle Isolation for Cancer Diagnostics. Micromachines, 13(1), 139. https://doi.org/10.3390/mi13010139




