Microfluidic System for Observation of Bacterial Culture and Effects on Biofilm Formation at Microscale
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials and Reagents
2.2. Sample Preparation
2.3. Bacteria Culture and Biofilm Formation in Microfluidic Device
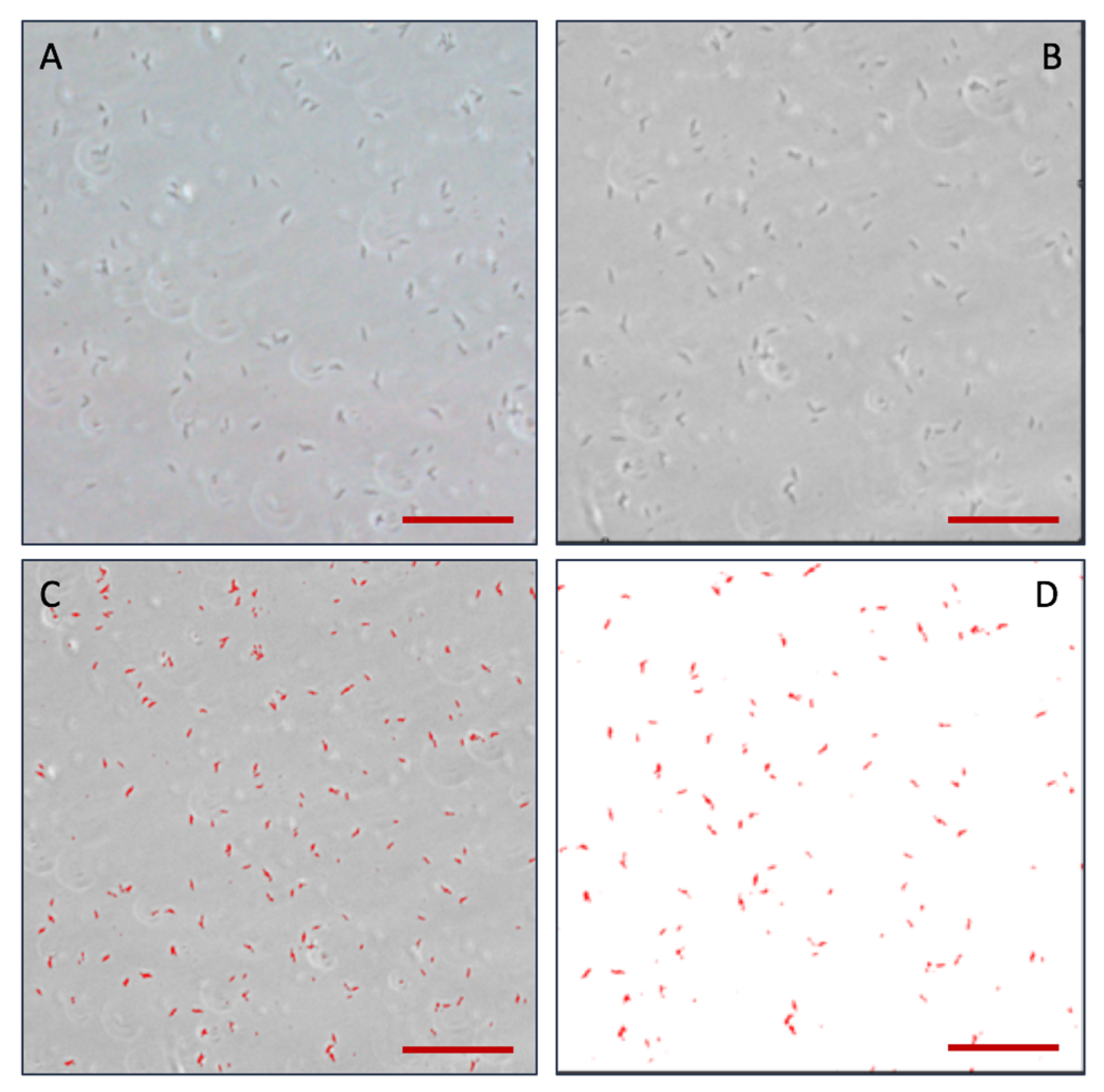
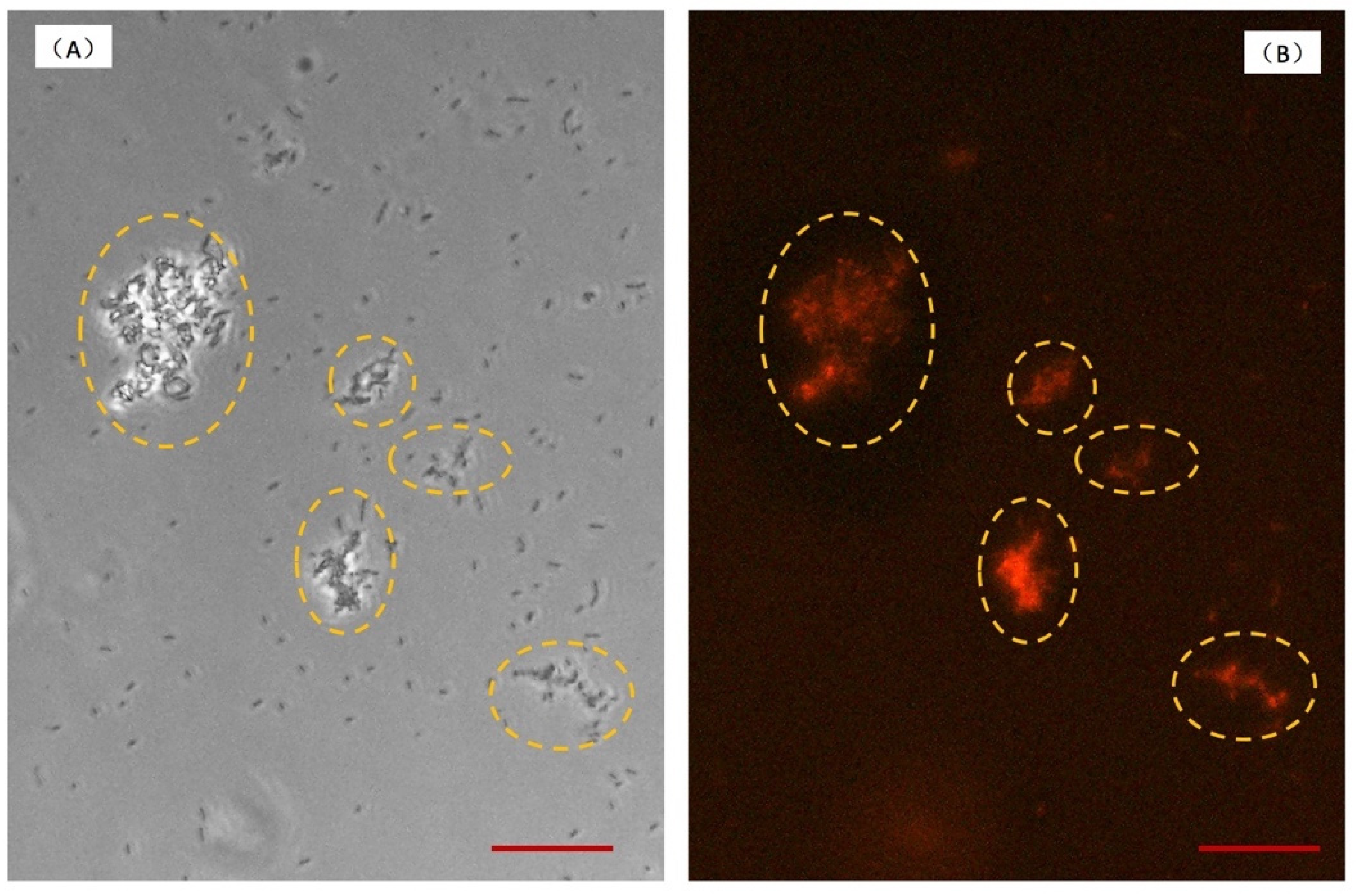
2.4. The Process of Biofilm Stain on Chip
2.5. Microfluidic Device Fabrication
2.6. Microfluidic System and Optical Image Analysis
3. Results and Discussion
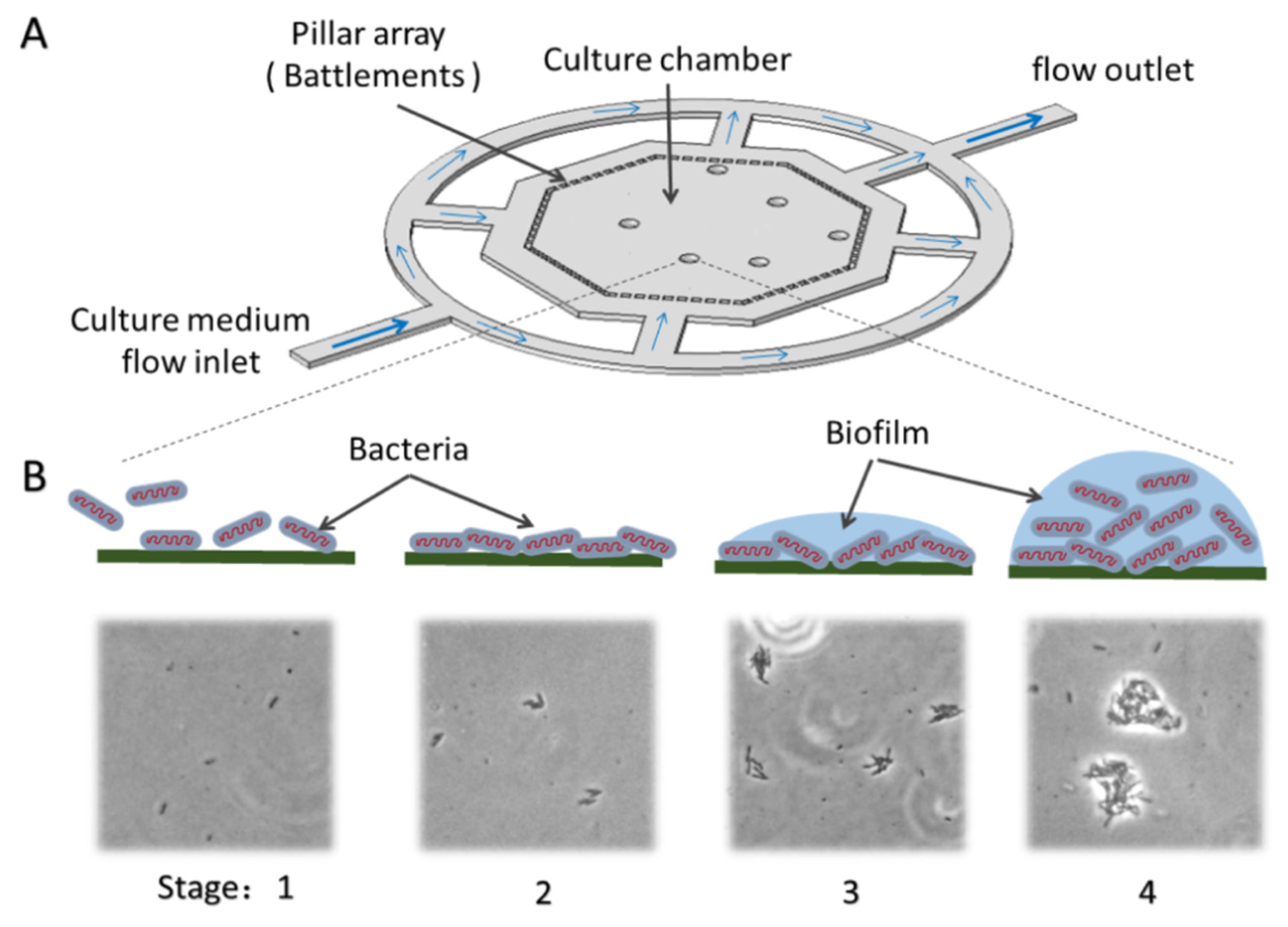
3.1. Chip Design and Operation
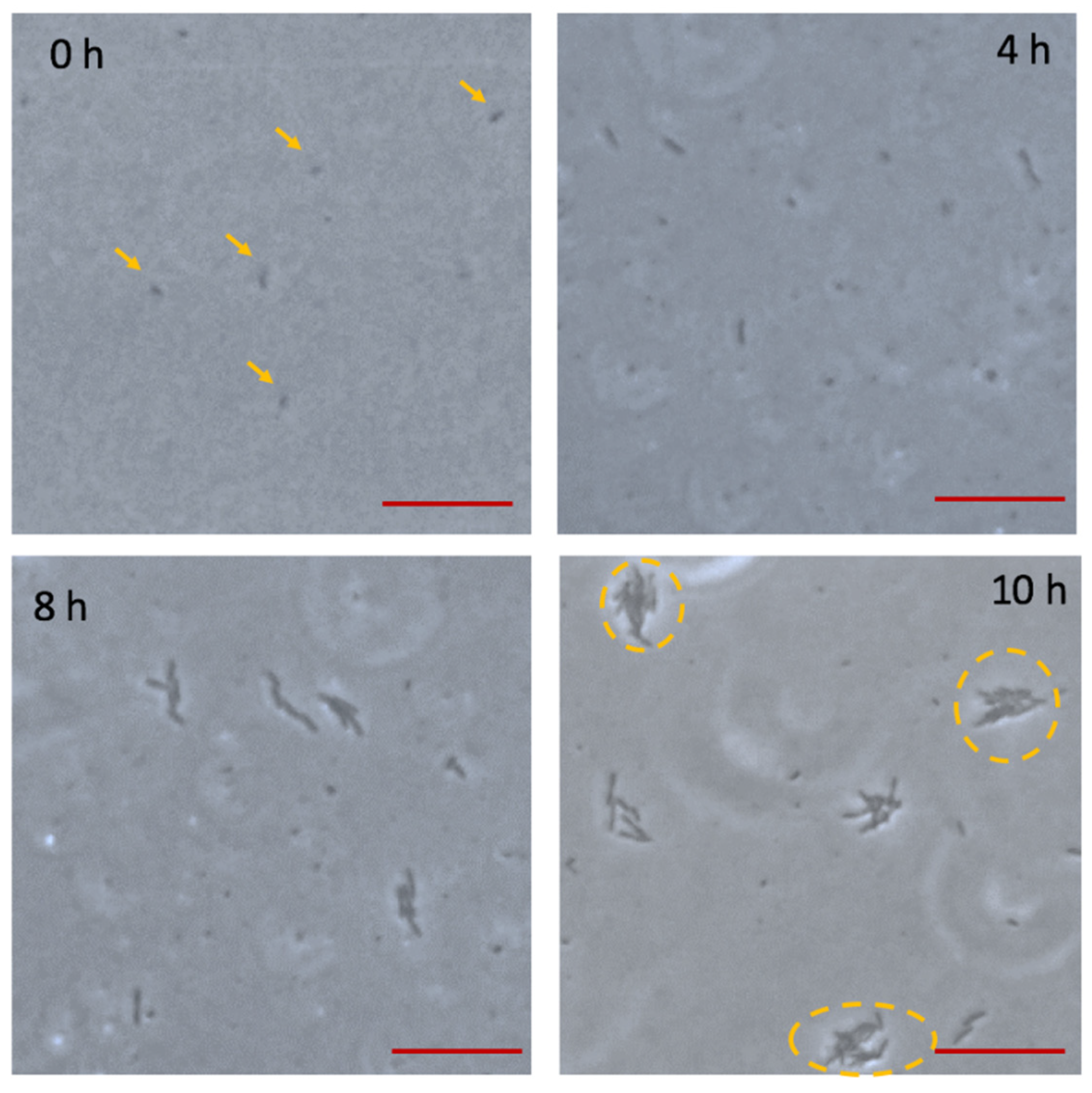
3.2. Bacterial Adhesion and Biofilm Formation on the Chip
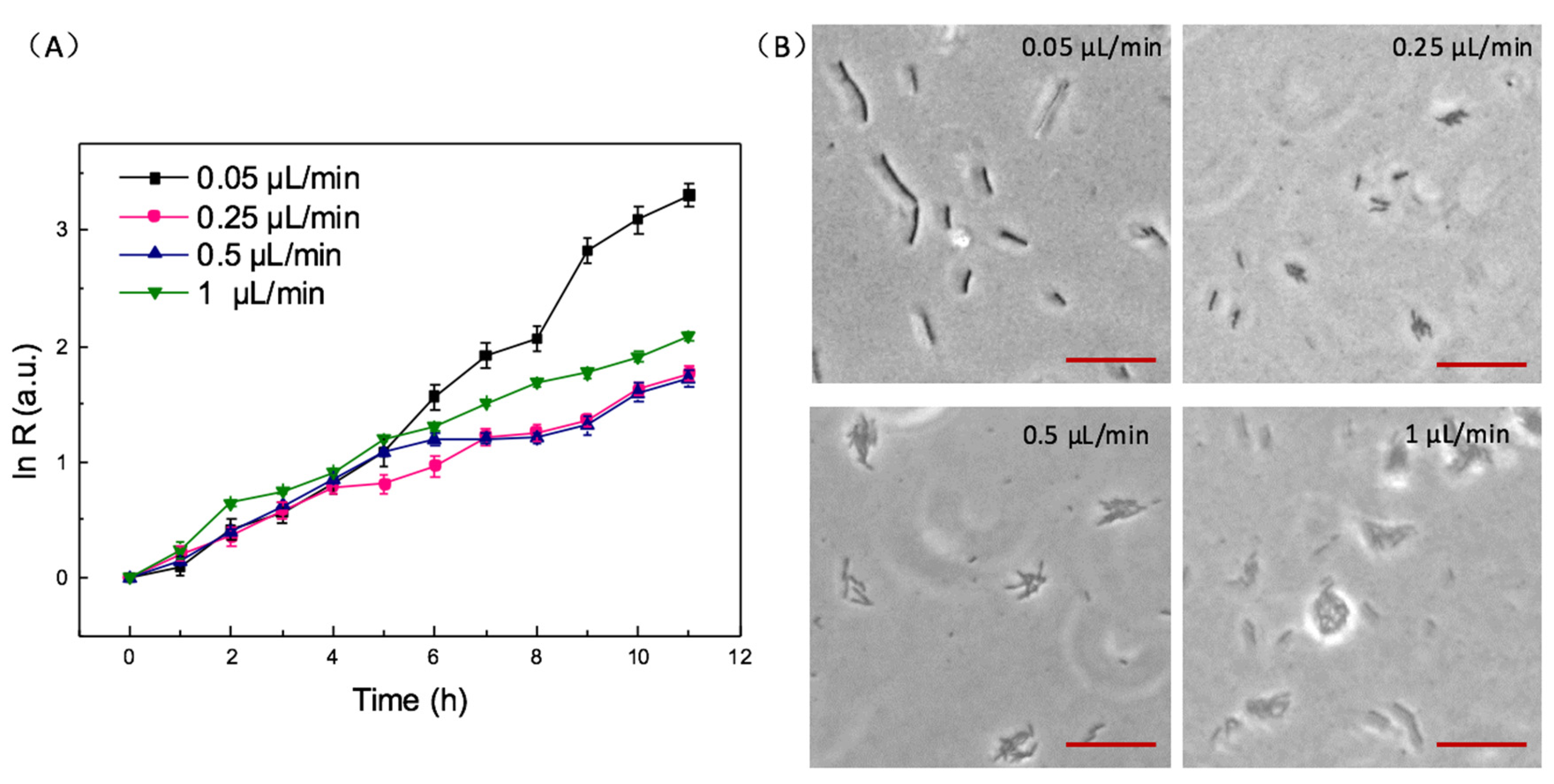
3.3. The Effect of Velocity on Bacteria Growth and Biofilms Formation
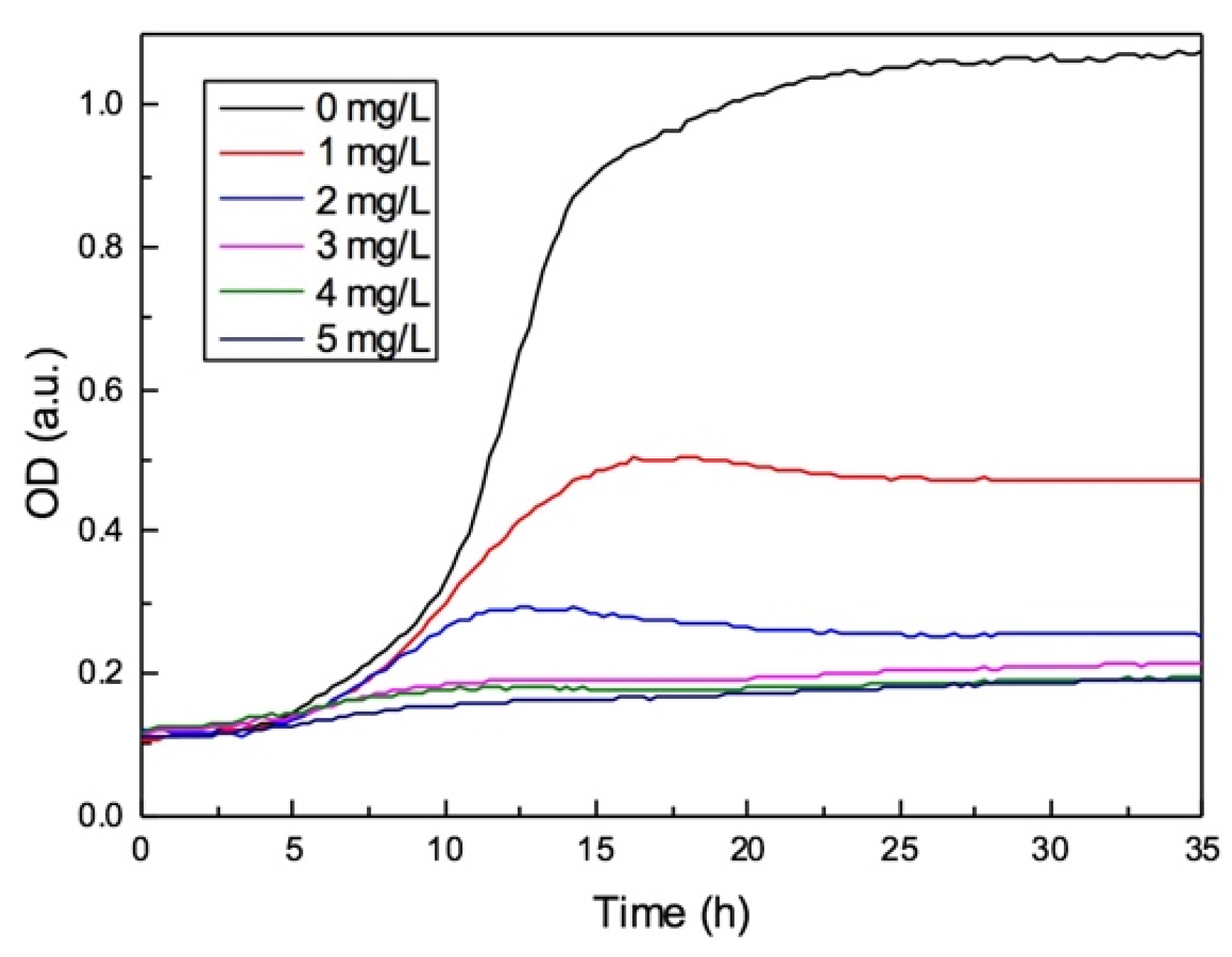
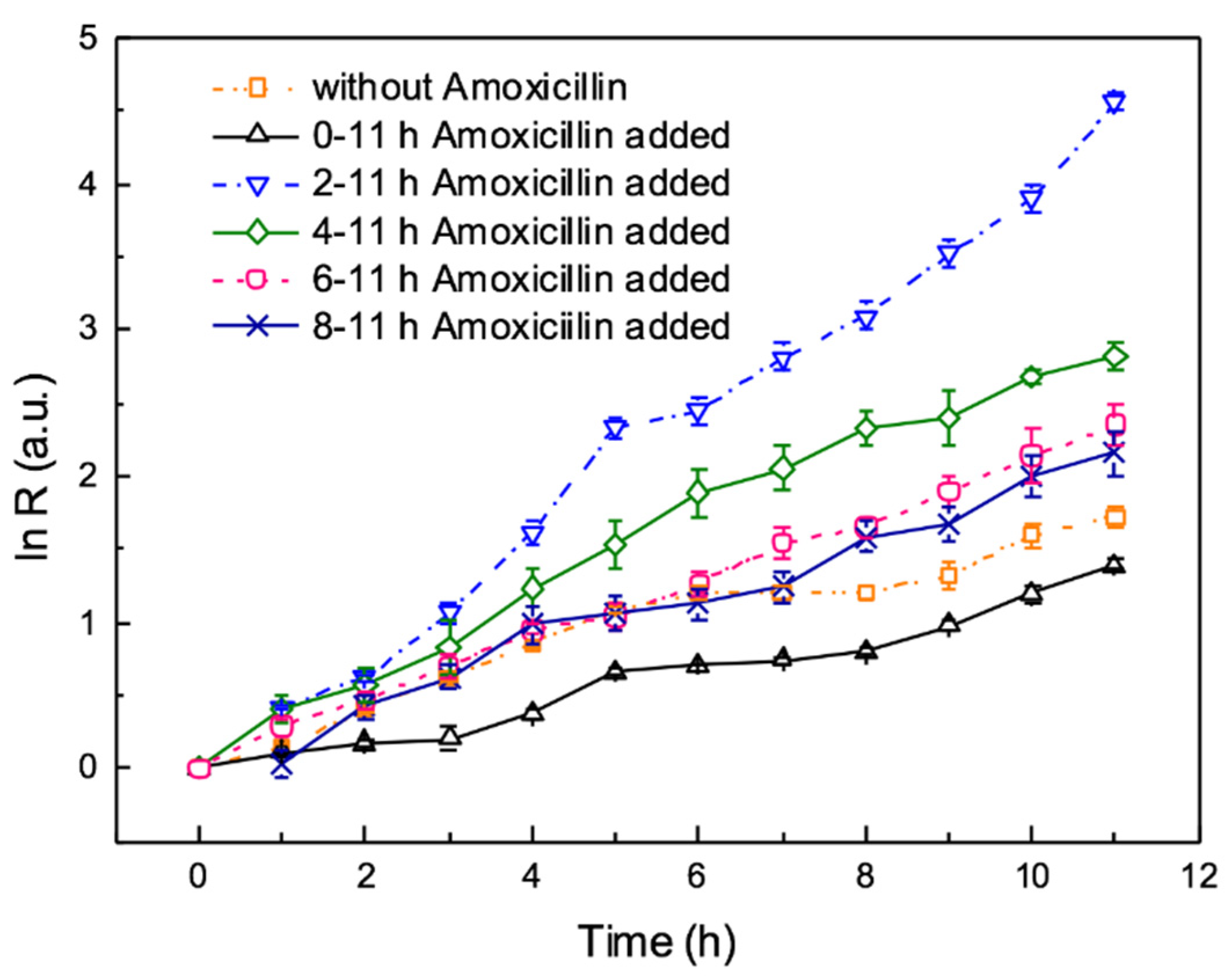
3.4. The Effects of Antibiotic on Biofilm Formation during the On-Chip Growth
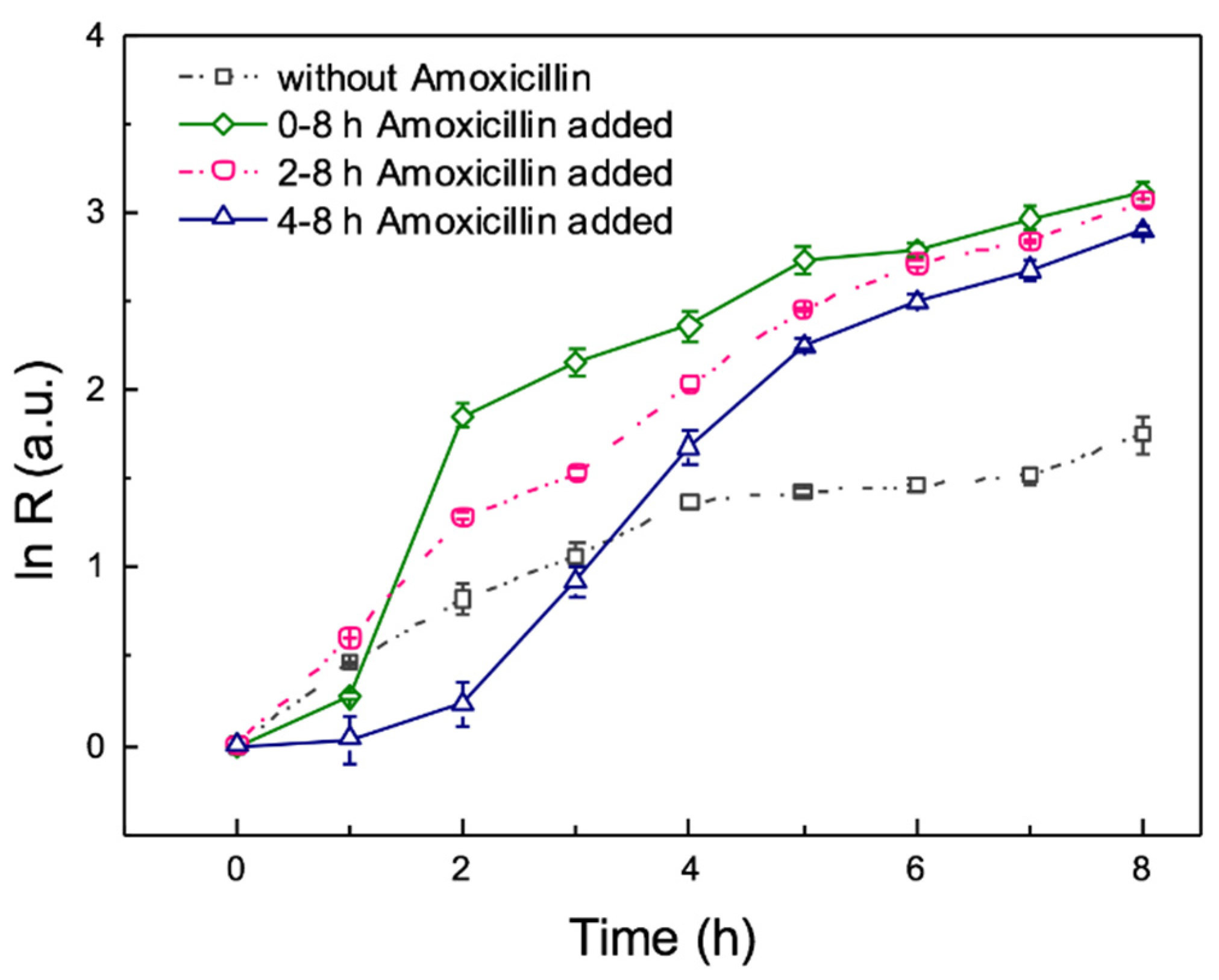
3.5. Effect of Antibiotic on Mixed Microorganism by On-Chip Culture
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Miura, Y.; Watanabe, Y.; Okabe, S. Membrane biofouling in pilot-scale membrane bioreactors (MBRs) treating municipal wastewater: Impact of biofilm formation. Environ. Sci. Technol. 2007, 41, 632–638. [Google Scholar] [CrossRef] [PubMed]
- Cheng, D.; Ngo, H.H.; Guo, W.; Liu, Y.; Chang, S.W.; Nguyen, D.D.; Nghiem, L.D.; Zhou, J.; Ni, B. Anaerobic membrane bioreactors for antibiotic wastewater treatment: Performance and membrane fouling issues. Bioresour. Technol. 2018, 267, 714–724. [Google Scholar] [CrossRef] [PubMed]
- Seviour, T.; Derlon, N.; Dueholm, M.S.; Flemming, H.C.; Girbal-Neuhauser, E.; Horn, H.; Kjelleberg, S.; van Loosdrecht, M.C.M.; Lotti, T.; Malpei, M.F.; et al. Extracellular polymeric substances of biofilms: Suffering from an identity crisis. Water Res. 2019, 151, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Karimi, A.; Karig, D.; Kumar, A.; Ardekani, A.M. Interplay of physical mechanisms and biofilm processes: Review of microfluidic methods. Lab Chip 2015, 15, 23–42. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Lawrence, J.R.; Korber, D.R.; Hoyle, B.D.; Costerton, J.W.; Caldwell, D.E. Optical sectioning of microbial biofilms. J. Bacteriol. 1991, 173, 6558–6567. [Google Scholar] [CrossRef]
- de Beer, D.; Stoodley, P.; Roe, F.; Lewandowski, Z. Effects of biofilms structures on oxygen distribution and mass transport. Biotechnol. Bioeng. 1994, 43, 1131–1138. [Google Scholar] [CrossRef]
- Huang, C.; Xu, D.K.; McFeters, A.G.; Stewart, S.P. Spatial patterns of alkaline phosphatase expression within bacterial colonies and biofilms in response to phosphate starvation. Appl. Environ. Microbiol. 1998, 64, 1526–1531. [Google Scholar]
- Hogan, S.; Kasotakis, E.; Maher, S.; Cavanagh, B.; O’Gara, P.E.; Pandit, A.; Keyes, E.T.; Devocelle, M.; O’Neill, E. A novel medical device coating prevents Staphylococcus aureusbiofilm formation on medical device surfaces. FEMS Microbiol. Lett. 2019, 366. [Google Scholar] [CrossRef]
- Almeida, M.I.G.S.; Jayawardane, B.M.; Kolev, S.D.; McKelvie, I.D. Developments of microfluidic paper-based analytical devices (μPADs) for water analysis: A review. Talanta 2018, 177, 176–190. [Google Scholar] [CrossRef]
- Lee, W.B.; Fu, C.Y.; Chang, W.H.; You, H.L.; Wang, C.H.; Lee, M.S.; Lee, G.B. A microfluidic device for antimicrobial susceptibility testing based on a broth dilution method. Biosens. Bioelectron. 2017, 87, 669–678. [Google Scholar] [CrossRef]
- Tang, Y.; Gan, M.; Xie, Y.; Li, X.; Chen, L. Fast screening of bacterial suspension culture conditions on chips. Lab Chip 2014, 14, 1162–1167. [Google Scholar] [CrossRef]
- Sun, P.; Liu, Y.; Sha, J.; Zhang, Z.; Tu, Q.; Chen, P.; Wang, J. High-throughput microfluidic system for long-term bacterial colony monitoring and antibiotic testing in zero-flow environments. Biosens. Bioelectron. 2011, 26, 1993–1999. [Google Scholar] [CrossRef]
- Xu, B.Y.; Hu, S.W.; Qian, G.S.; Xu, J.J.; Chen, H.Y. A novel microfluidic platform with stable concentration gradient for on chip cell culture and screening assays. Lab Chip 2013, 13, 3714–3720. [Google Scholar] [CrossRef] [PubMed]
- Skolimowski, M.; Nielsen, M.W.; Emneus, J.; Molin, S.; Taboryski, R.; Sternberg, C.; Dufva, M.; Geschke, O. Microfluidic dissolved oxygen gradient generator biochip as a useful tool in bacterial biofilm studies. Lab Chip 2010, 10, 2162–2169. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.; Hall-Stoodley, L.; Allan, N.R.; Faust, N.S. New approaches to the treatment of biofilm—Related infections. J. Infect. 2014, 69, S47–S52. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Keatch, R.; Zhao, Q.; Wright, J.A.; Bryant, C.E.; Redmann, A.L.; Terentjev, E.M. Influence of type I fimbriae and fluid shear stress on bacterial behavior and multicellular architecture of early escherichia coli biofilms at single-cell resolution. Appl. Environ. Microbiol. 2018, 84, e02343–e02417. [Google Scholar] [CrossRef]
- Dickschat, S.J. Quorum sensing and bacterial biofilms. Nat. Prod. Rep. 2010, 27, 343–369. [Google Scholar] [CrossRef]
- Vertes, A.; Hitchins, V.; Phillips, K.S. Analytical challenges of microbial biofilms on medical devices. Anal. Chem. 2012, 84, 3858–3866. [Google Scholar] [CrossRef]
- Kim, J.; Park, H.D.; Chung, S. Microfluidic approaches to bacterial biofilm formation. Molecules 2012, 17, 9818–9834. [Google Scholar] [CrossRef]
- Funari, R.; Bhalla, N.; Chu, K.Y.; Soderstrom, B.; Shen, A.Q. Nanoplasmonics for real-time and label-free monitoring of microbial biofilm formation. ACS Sens. 2018, 3, 1499–1509. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Skauge, T.; Landa-Marbán, D.; Hovland, B.; Thorbjørnsen, B.; Radu, F.A.; Vik, B.F.; Baumann, T.; Bødtker, G. Microfuidic study of effects of flow velocity and nutrient concentration on biofilm accumulation and adhesive strength in the flowing and no-flowing microchannels. J. Ind. Microbiol. Biotechnol. 2019, 46, 855–868. [Google Scholar] [CrossRef]
- Li, B.; Qiu, Y.; Glidle, A.; McIlvenna, D.; Luo, Q.; Cooper, J.; Shi, H.C.; Yin, H. Gradient microfluidics enables rapid bacterial growth inhibition testing. Anal. Chem. 2014, 86, 3131–3137. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.P.; Kim, Y.G.; Choi, C.H.; Kim, H.E.; Lee, S.H.; Chang, W.S.; Lee, C.S. In situ monitoring of antibiotic susceptibility of bacterial biofilms in a microfluidic device. Lab Chip 2010, 10, 3296–3299. [Google Scholar] [CrossRef] [PubMed]
- Blair, D.F. How bacteria sense and swim. Annu. Rev. Microbiol. 1995, 49, 489–522. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; William Costerton, J. Antibiotic resistance of bacteria in biofilms. Lancet 2011, 358, 135–138. [Google Scholar] [CrossRef]
- Kim, J.; Kim, H.S.; Han, S.; Lee, J.Y.; Oh, J.E.; Chung, S.; Park, H.D. Hydrodynamic effects on bacterial biofilm development in a microfluidic environment. Lab Chip 2013, 13, 1846–1849. [Google Scholar] [CrossRef]
- Kalashnikov, M.; Lee, C.J.; Campbell, J.; Sharon, A.; Sauer-Budge, F.A. A microfluidic platform for rapid, stress-induced antibiotic susceptibility testing of Staphylococcus aureus. Lab Chip 2012, 12, 4523–4532. [Google Scholar] [CrossRef]
- Lecuyer, S.; Rusconi, R.; Shen, Y.; Forsyth, A.; Vlamakis, H.; Kolter, R.; Stone, H.A. Shear stress increases the residence time of adhesion of Pseudomonas aeruginosa. Biophys. J. 2011, 100, 341–350. [Google Scholar] [CrossRef]
- Picioreanu, C.; van Loosdrecht, M.C.M.; Heijnen, J.J. Effect of diffusive and convective substrate transport on biofilm structure formation: A two-dimensional modeling study. Biotechnol. Bioeng. 2000, 68, 504–515. [Google Scholar] [CrossRef]
- Zheng, D.; Chang, Q.; Li, Z.; Gao, M.; She, Z.; Wang, X.; Guo, L.; Zhao, Y.; Jin, C.; Gao, F. Performance and microbial community of a sequencing batch biofilm reactor treating synthetic mariculture wastewater under long-term exposure to norfloxacin. Bioresour. Technol. 2016, 222, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Wang, Y.; Zhou, S.; Jiang, X.; Ma, X.; Liu, C. Robust performance of a membrane bioreactor for removing antibiotic resistance genes exposed to antibiotics: Role of membrane foulants. Water Res. 2018, 130, 139–150. [Google Scholar] [CrossRef] [PubMed]
- Bollinger, N.; Hassett, D.J.; Iglewski, B.H.; Costerton, J.W.; McDermott, T.R. Gene expression in Pseudomonas aeruginosa: Evidence of iron override effects on quorum sensing and biofilm-specific gene regulation. J. Bacteriol. 2001, 183, 1990–1996. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef] [PubMed]
- de Kievit, T.R. Quorum sensing in Pseudomonas aeruginosa biofilm. Environ. Microbiol. 2009, 11, 278–288. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.-Y.; Sun, K.; Abulimiti, A.; Xu, P.-P.; Li, Z.-Y. Microfluidic System for Observation of Bacterial Culture and Effects on Biofilm Formation at Microscale. Micromachines 2019, 10, 606. https://doi.org/10.3390/mi10090606
Zhang X-Y, Sun K, Abulimiti A, Xu P-P, Li Z-Y. Microfluidic System for Observation of Bacterial Culture and Effects on Biofilm Formation at Microscale. Micromachines. 2019; 10(9):606. https://doi.org/10.3390/mi10090606
Chicago/Turabian StyleZhang, Xiao-Yan, Kai Sun, Aliya Abulimiti, Pian-Pian Xu, and Zhe-Yu Li. 2019. "Microfluidic System for Observation of Bacterial Culture and Effects on Biofilm Formation at Microscale" Micromachines 10, no. 9: 606. https://doi.org/10.3390/mi10090606
APA StyleZhang, X.-Y., Sun, K., Abulimiti, A., Xu, P.-P., & Li, Z.-Y. (2019). Microfluidic System for Observation of Bacterial Culture and Effects on Biofilm Formation at Microscale. Micromachines, 10(9), 606. https://doi.org/10.3390/mi10090606



