Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow
Abstract
1. Introduction
2. Materials and Methods
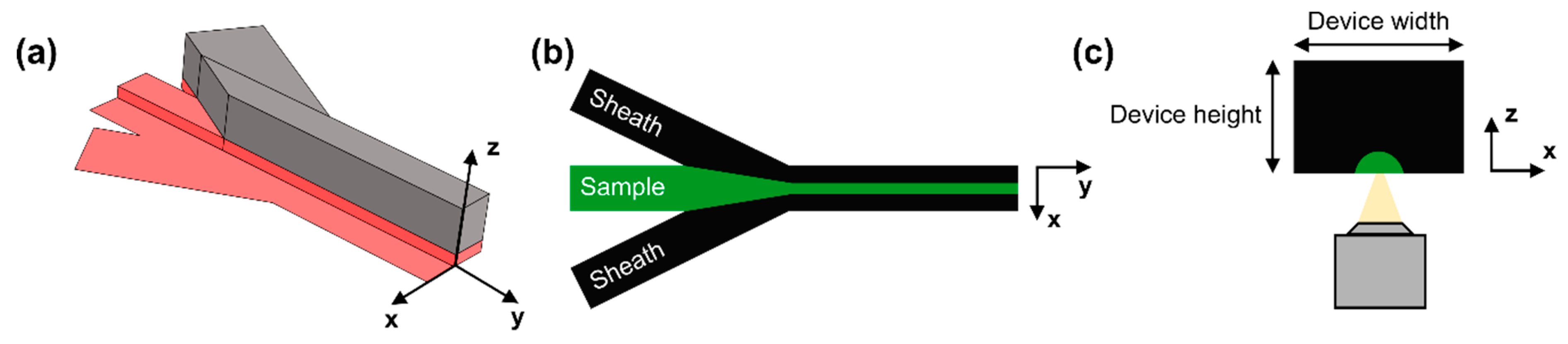
2.1. Device Design and Fabrication
2.2. Flow Focusing Principle
2.3. Device Simulations
2.4. Microscopy
2.5. Microsphere Z-Displacement Regression
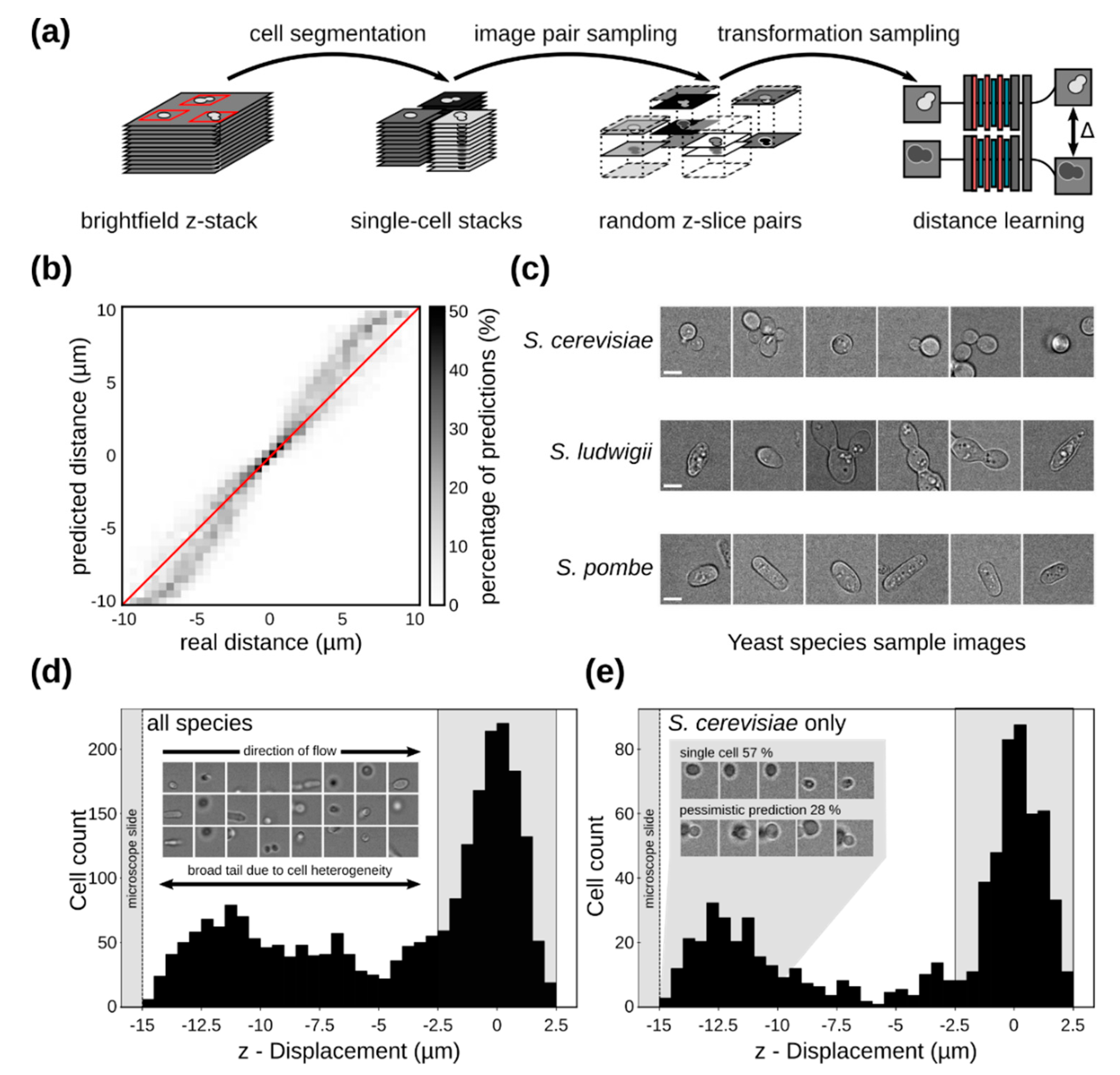
2.6. Yeast Cell Z-Distance Regression
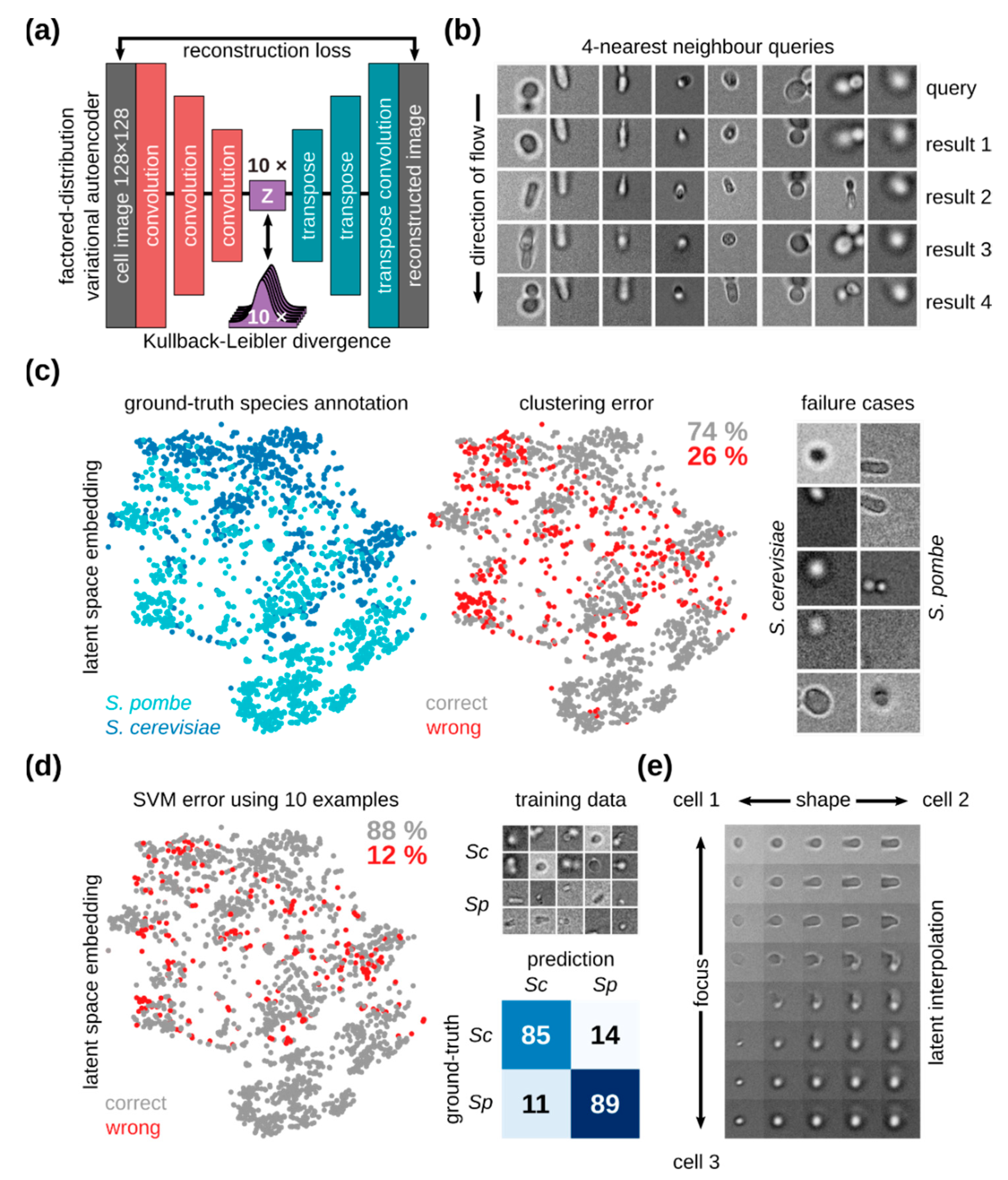
2.7. Unsupervised Learning For Cellular Mixtures
3. Results and Discussion
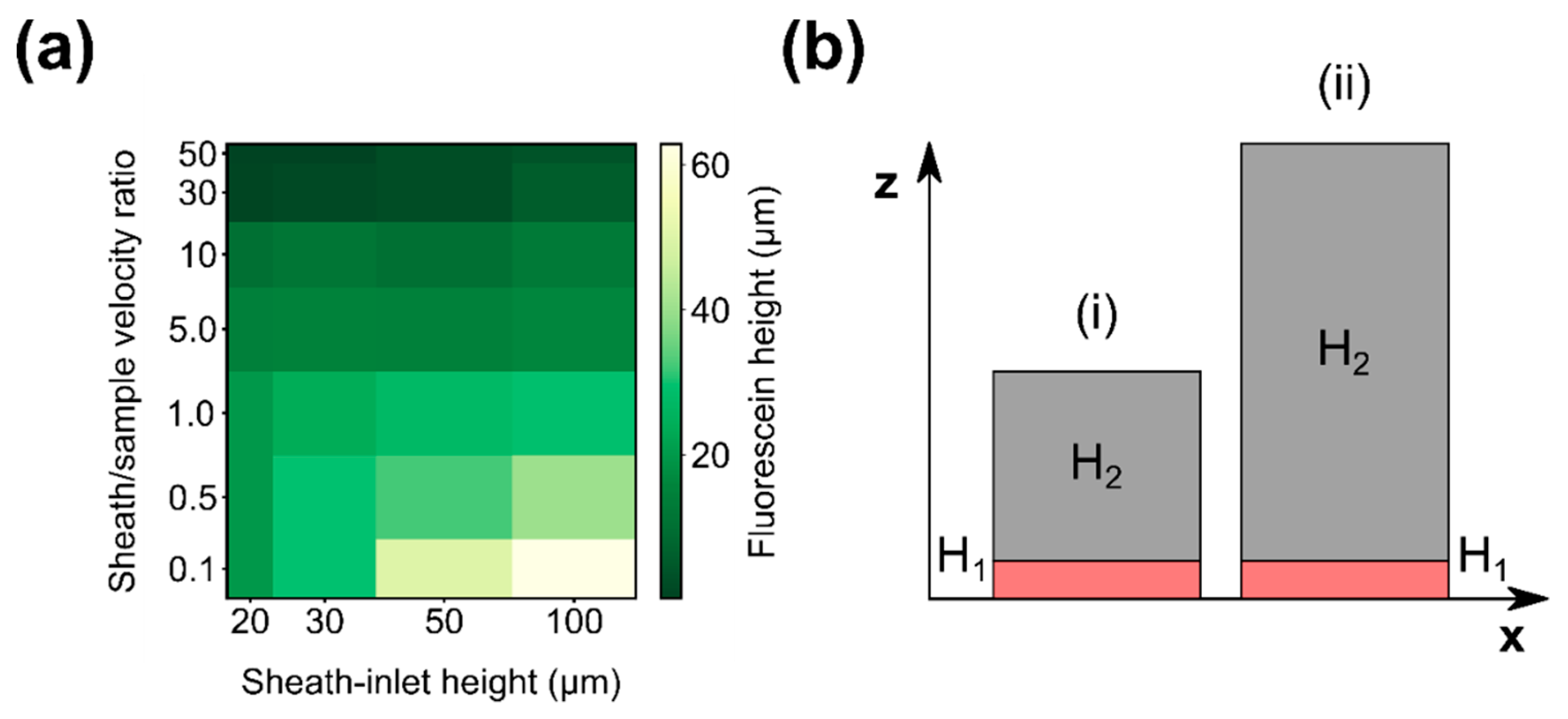
3.1. Simulation Results
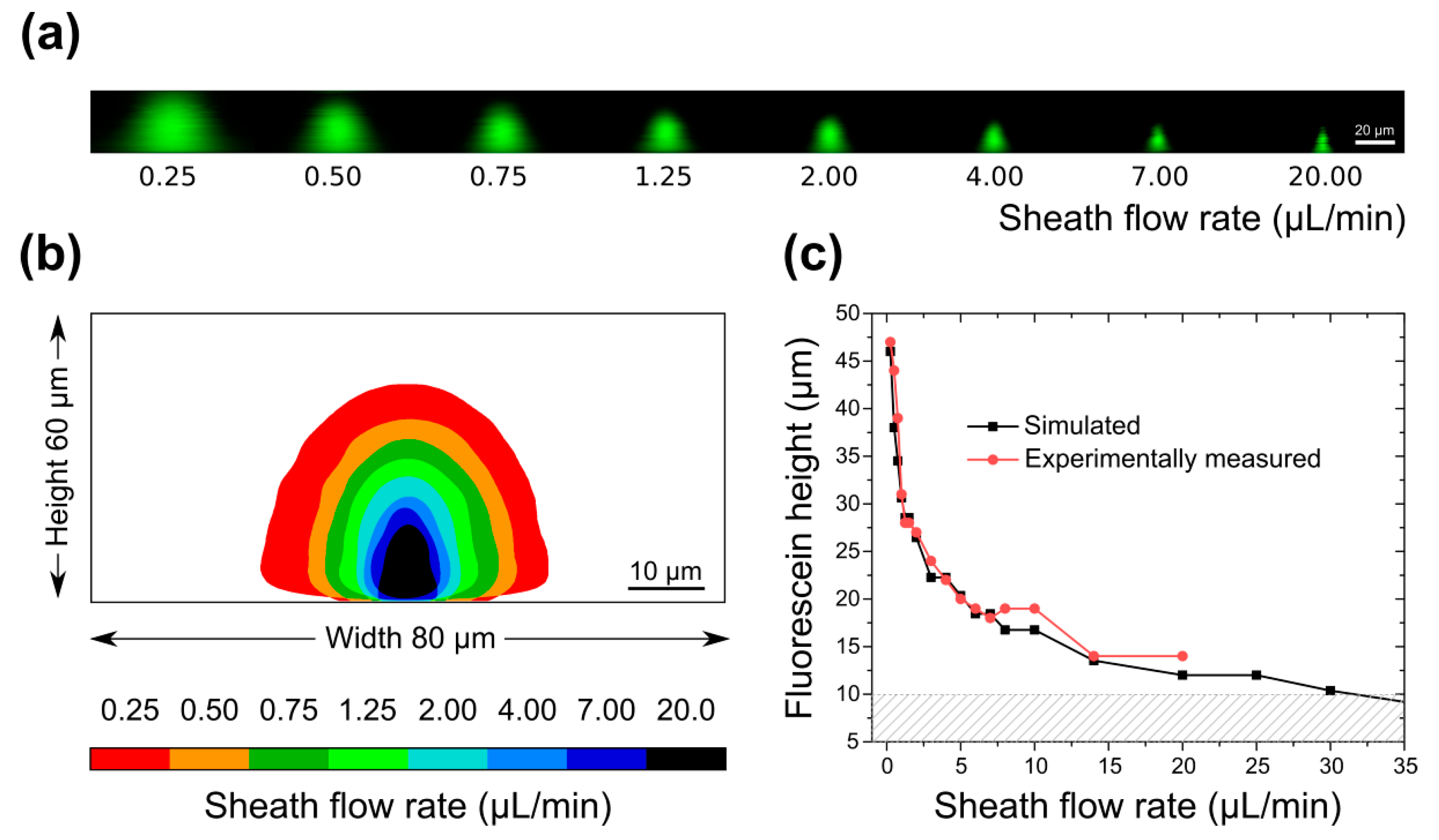
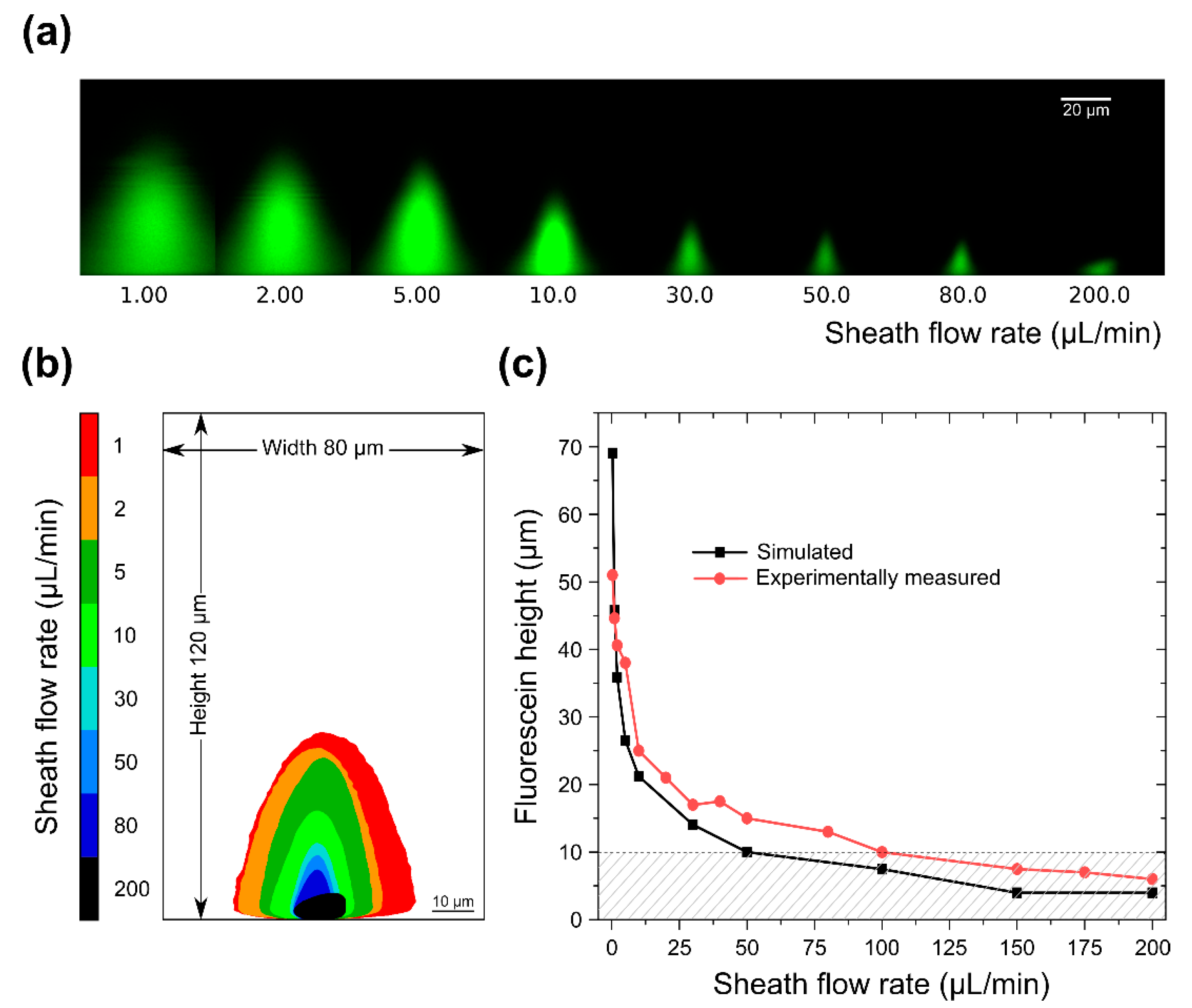
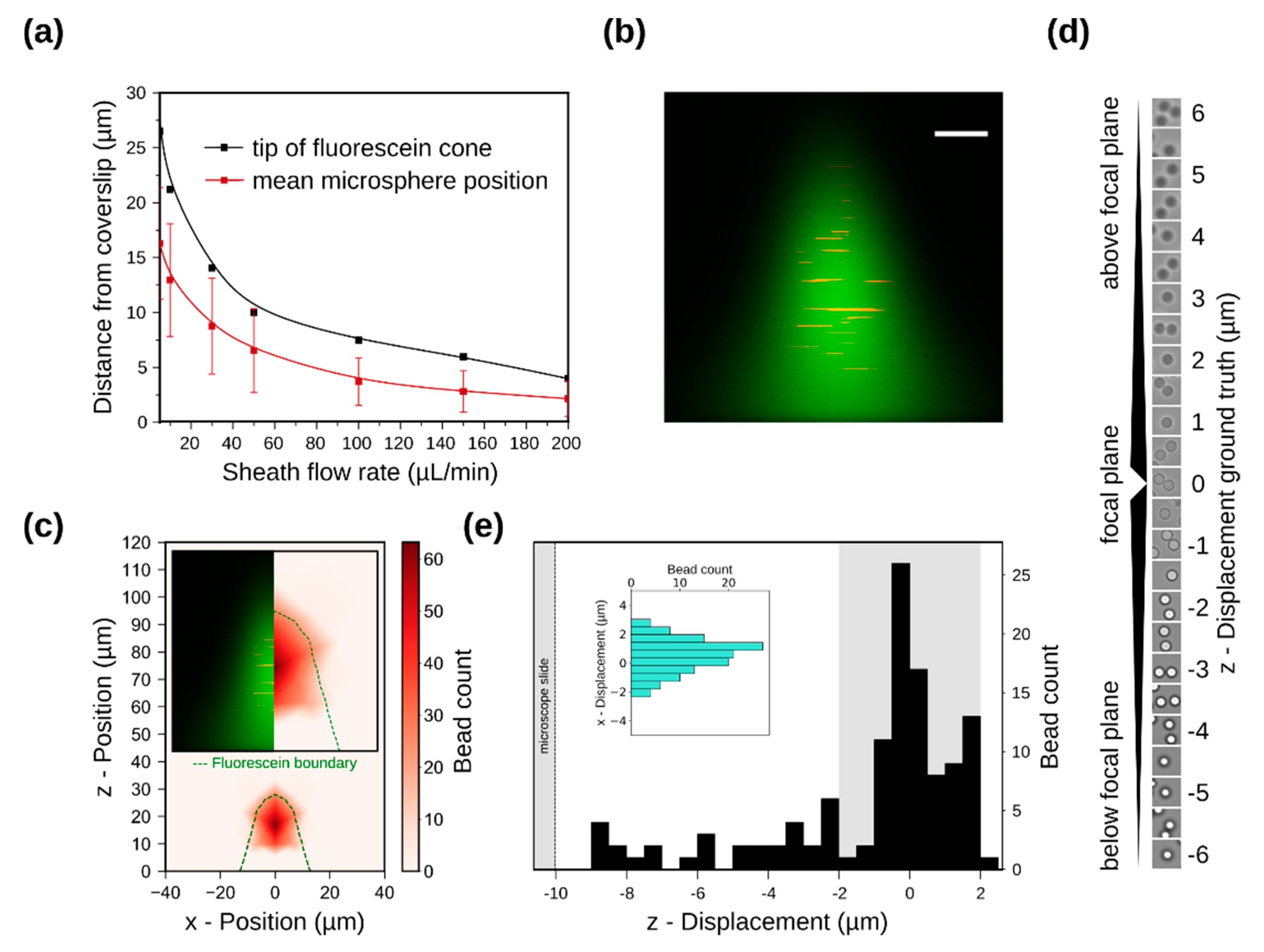
3.2. Sample Confinement Testing Using Fluorescein
3.3. Simulation of Particle Positioning and Validation Using Microspheres
3.4. Single-Cell Imaging in Flow
3.5. In-Flow Cell Imaging and Cell Classification
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gross, H.-J.; Verwer, B.; Houck, D.; Recktenwald, D. Detection of rare cells at a frequency of one per million by flow cytometry. Cytometry 1993, 14, 519–526. [Google Scholar] [CrossRef] [PubMed]
- De Rosa, S.C.; Herzenberg, L.A.; Herzenberg, L.A.; Roederer, M. 11-color, 13-parameter flow cytometry: Identification of human naive T cells by phenotype, function, and T-cell receptor diversity. Nat. Med. 2001, 7, 245–248. [Google Scholar] [CrossRef]
- Sandberg, J.; Werne, B.; Dessing, M.; Lundeberg, J. Rapid flow-sorting to simultaneously resolve multiplex massively parallel sequencing products. Sci. Rep. 2011, 1, 1–7. [Google Scholar] [CrossRef]
- Schonbrun, E.; Gorthi, S.S.; Schaak, D. Microfabricated multiple field of view imaging flow cytometry. Lab Chip 2012, 12, 268–273. [Google Scholar] [CrossRef]
- Barteneva, N.S.; Fasler-kan, E.; Vorobjev, I.A. Imaging Flow Cytometry: Coping with Heterogeneity in Biological Systems. J. Histochem. Cytochem. 2012, 60, 723–733. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Gu, Y.; Zhang, A.C.; Lo, Y. Review: Imaging technologies for flow cytometry. Lab Chip 2016, 16, 4639–4647. [Google Scholar] [CrossRef] [PubMed]
- Rosenauer, M.; Buchegger, W.; Finoulst, I.; Verhaert, P.; Vellekoop, M. Miniaturized flow cytometer with 3D hydrodynamic particle focusing and integrated optical elements applying silicon photodiodes. Microfluid. Nanofluid. 2011, 10, 761–771. [Google Scholar] [CrossRef]
- Simonnet, C.; Groisman, A. High-throughput and high-resolution flow cytometry in molded microfluidic devices. Anal. Chem. 2006, 78, 5653–5663. [Google Scholar] [CrossRef]
- Sundararajan, N.; Pio, M.S.; Lee, L.P.; Berlin, A.A. Three-Dimensional Hydrodynamic Focusing in Polydimethylsiloxane (PDMS) Microchannels. J. Microelectromech. Syst. 2004, 13, 559–567. [Google Scholar] [CrossRef]
- Chang, C.-C.; Huang, Z.-X.; Yang, R.-J. Three-dimensional hydrodynamic focusing in two-layer polydimethylsiloxane (PDMS) microchannels. J. Micromech. Microeng. 2007, 17, 1479–1486. [Google Scholar] [CrossRef]
- Wu, T.; Chen, Y.; Park, S.; Hong, J.; Teslaa, T.; Zhong, J.F.; Carlo, D.; Teitell, M.A.; Chiou, P. Pulsed laser triggered high speed microfluidic fluorescence activated cell sorter. Lab Chip 2012, 12, 1378–1383. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, S.; Kasai, Y.; Hayakawa, T.; Arai, F. On-chip cell sorting by high-speed local-flow control using dual membrane pumps. Lab Chip 2017, 17, 2760–2767. [Google Scholar] [CrossRef]
- Mao, X.; Lin, S.C.S.; Dong, C.; Huang, T.J. Single-layer planar on-chip flow cytometer using microfluidic drifting based three-dimensional (3D) hydrodynamic focusing. Lab Chip 2009, 9, 1583–1589. [Google Scholar] [CrossRef]
- Eluru, G.; Julius, L.A.N.; Gorthi, S.S. Single-layer microfluidic device to realize hydrodynamic 3D flow focusing. Lab Chip 2016, 16, 4133–4141. [Google Scholar] [CrossRef] [PubMed]
- Gualda, E.J.; Pereira, H.; Martins, G.G.; Gardner, R.; Moreno, N. Three-dimensional imaging flow cytometry through light-sheet fluorescence microscopy. Cytom. Part A 2017, 91, 144–151. [Google Scholar] [CrossRef]
- Nawaz, A.A.; Zhang, X.; Mao, X.; Rufo, J.; Lin, S.C.S.; Guo, F.; Zhao, Y.; Lapsley, M.; Li, P.; McCoy, J.P.; et al. Sub-micrometer-precision, three-dimensional (3D) hydrodynamic focusing via “microfluidic drifting”. Lab Chip Miniaturisation Chem. Biol. 2014, 14, 415–423. [Google Scholar] [CrossRef]
- Paiè, P.; Bragheri, F.; Di Carlo, D.; Osellame, R. Particle focusing by 3D inertial microfluidics. Microsyste. Nanoeng. 2017, 3, 17027. [Google Scholar] [CrossRef]
- Rane, A.S.; Rutkauskaite, J.; deMello, A.; Stavrakis, S. High-Throughput Multi-parametric Imaging Flow Cytometry. Chem 2017, 3, 588–602. [Google Scholar] [CrossRef]
- Nitta, N.; Sugimura, T.; Isozaki, A.; Mikami, H.; Hiraki, K.; Sakuma, S.; Iino, T.; Arai, F.; Endo, T.; Fujiwaki, Y.; et al. Intelligent Image-Activated Cell Sorting. Cell 2018, 175, 266–276.e13. [Google Scholar] [CrossRef]
- Normolle, D.P.; Donnenberg, V.S.; Donnenberg, A.D. Statistical classification of multivariate flow cytometry data analyzed by manual gating: Stem, progenitor, and epithelial marker expression in nonsmall cell lung cancer and normal lung. Cytom. Part A 2013, 83A, 150–160. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Ho, J.W.K. Ultrafast clustering of single-cell flow cytometry data using FlowGrid. BMC Syst. Biol. 2019, 13 (Suppl. 2), 35. [Google Scholar] [CrossRef]
- Pouyan, M.B.; Jindal, V.; Birjandtalab, J.; Nourani, M. Single and multi-subject clustering of flow cytometry data for cell-type identification and anomaly detection. BMC Med. Genom. 2016, 9, 41. [Google Scholar] [CrossRef]
- Kraus, O.Z.; Grys, B.T.; Ba, J.; Chong, Y.T.; Frey, B.J.; Boone, C.; Andrews, B.J.J.; Pärnamaa, T.; Parts, L.; Humeau-Heurtier, A.; et al. The curse of dimensionality. Mach. Learn. 2018, 7, 18. [Google Scholar]
- Köppen, M. The curse of dimensionality. In Proceedings of the 5th Online World Conference on Soft Computing in Industrial Applications, 4–18 September 2000; pp. 4–8. [Google Scholar]
- Humeau-Heurtier, A. Texture Feature Extraction Methods: A Survey. IEEE Access 2019, 7, 8975–9000. [Google Scholar] [CrossRef]
- Carpenter, A.E.; Jones, T.R.; Lamprecht, M.R.; Clarke, C.; Kang, I.H.; Friman, O.; Guertin, D.A.; Chang, J.H.; Lindquist, R.A.; Moffat, J.; et al. CellProfiler: Image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006, 7, R100. [Google Scholar] [CrossRef]
- Kraus, O.Z.; Grys, B.T.; Ba, J.; Chong, Y.; Frey, B.J.; Boone, C.; Andrews, B.J. Automated analysis of high-content microscopy data with deep learning. Mol. Syst. Biol. 2017, 13, 924. [Google Scholar] [CrossRef]
- Rumetshofer, E.; Hofmarcher, M.; Röhrl, C.; Hochreiter, S.; Klambauer, G. Human-level Protein Localization with Convolutional Neural Networks. In Proceedings of the International Conference on Learning Representations, New Orleans, LA, USA, 6–9 May 2019. [Google Scholar]
- Pärnamaa, T.; Parts, L. Accurate Classification of Protein Subcellular Localization from High-Throughput Microscopy Images Using Deep Learning. G3 (Bethesda) 2017, 7, 1385–1392. [Google Scholar] [CrossRef]
- Platt, J. Sequential Minimal Optimization: A Fast Algorithm for Training Support Vector Machines; Technical Report MSR-TR-98-14; Microsoft Research: Redmond, WA, USA, 1998. [Google Scholar]
- Chong, Y.T.; Koh, J.L.Y.; Friesen, H.; Kaluarachchi Duffy, S.; Cox, M.J.; Moses, A.; Moffat, J.; Boone, C.; Andrews, B.J. Yeast Proteome Dynamics from Single Cell Imaging and Automated Analysis. Cell 2015, 161, 1413–1424. [Google Scholar] [CrossRef] [PubMed]
- Bengio, Y. Deep Learning of Representations: Looking Forward. In Statistical Language and Speech Processing, Lecture Notes in Computer Science; Dediu, A.-H., Martín-Vide, C., Mitkov, R., Truthe, B., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 1–27. [Google Scholar]
- Gidaris, S.; Singh, P.; Komodakis, N. Unsupervised Representation Learning by Predicting Image Rotations. arXiv 2018, arXiv:1803.07728. [Google Scholar]
- Haeusser, P.; Plapp, J.; Golkov, V.; Aljalbout, E.; Cremers, D. Associative Deep Clustering: Training a Classification Network with no Labels. In Proceedings of the German Conference on Pattern Recognition (GCPR), Stuttgart, Germany, 9–12 October 2018. [Google Scholar]
- Caron, M.; Bojanowski, P.; Joulin, A.; Douze, M. Deep Clustering for Unsupervised Learning of Visual Features. arXiv 2018, arXiv:1807.05520. [Google Scholar]
- Kingma, D.P.; Welling, M. Auto-Encoding Variational Bayes. arXiv 2013, arXiv:1312.6114. [Google Scholar]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Networks. arXiv 2014, arXiv:1406.2661. [Google Scholar]
- Vincent, P.; Larochelle, H.; Lajoie, I.; Bengio, Y.; Manzagol, P.-A. Stacked Denoising Autoencoders: Learning Useful Representations in a Deep Network with a Local Denoising Criterion. J. Mach. Learn. Res. 2010, 11, 3371–3408. [Google Scholar]
- Higgins, I.; Amos, D.; Pfau, D.; Racaniere, S.; Matthey, L.; Rezende, D.; Lerchner, A. Towards a Definition of Disentangled Representations. arXiv 2018, arXiv:1812.02230. [Google Scholar]
- Kim, H.; Mnih, A. Disentangling by Factorising. arXiv 2018, arXiv:1802.05983. [Google Scholar]
- Kim, M.; Wang, Y.; Sahu, P.; Pavlovic, V. Relevance Factor VAE: Learning and Identifying Disentangled Factors. arXiv 2019, arXiv:1902.01568. [Google Scholar]
- Higgins, I.; Matthey, L.; Pal, A.; Burgess, C.; Glorot, X.; Botvinick, M.; Mohamed, S.; Lerchner, A. beta-VAE: Learning Basic Visual Concepts with a Constrained Variational Framework. In Proceedings of the International Conference on Learning Representations, Toulon, France, 24–26 April 2017. [Google Scholar]
- Burgess, C.P.; Higgins, I.; Pal, A.; Matthey, L.; Watters, N.; Desjardins, G.; Lerchner, A. Understanding disentangling in β-VAE. arXiv 2018, arXiv:1804.03599. [Google Scholar]
- Chen, R.T.Q.; Li, X.; Grosse, R.; Duvenaud, D. Isolating Sources of Disentanglement in Variational Autoencoders. arXiv 2018, arXiv:1802.04942. [Google Scholar]
- Mescheder, L.; Geiger, A.; Nowozin, S. Which Training Methods for GANs do actually Converge? arXiv 2018, arXiv:1801.04406. [Google Scholar]
- Scott, R.; Sethu, P.; Harnett, C.K. Three-dimensional hydrodynamic focusing in a microfluidic Coulter counter. Rev. Sci. Instrum. 2008, 79, 46104. [Google Scholar] [CrossRef]
- Hairer, G.; Pärr, G.S.; Svasek, P.; Jachimowicz, A.; Vellekoop, M.J. Investigations of micrometer sample stream profiles in a three-dimensional hydrodynamic focusing device. Sens. Actuators B Chem. 2008, 132, 518–524. [Google Scholar] [CrossRef]
- Chung, S.; Park, S.J.; Kim, J.K.; Chung, C.; Han, D.C.; Chang, J.K. Plastic microchip flow cytometer based on 2- and 3-dimensional hydrodynamic flow focusing. Microsyst. Technol. 2003, 9, 525–533. [Google Scholar] [CrossRef]
- Lake, M.; Narciso, C.; Cowdrick, K.; Storey, T.; Zhang, S.; Zartman, J.; Hoelzle, D. Microfluidic device design, fabrication, and testing protocols. Protoc. Exch. 2015. [Google Scholar] [CrossRef]
- Qin, D.; Xia, Y.; Whitesides, G.M. Soft lithography for micro- and nanoscale patterning. Nat. Protoc. 2010, 5, 491. [Google Scholar] [CrossRef]
- Ward, K.; Fan, Z.H. Mixing in microfluidic devices and enhancement methods. J. Micromech. Microeng. 2015, 25, 094001. [Google Scholar] [CrossRef]
- Tan, J.N.; Neild, A. Microfluidic mixing in a Y-junction open channel. AIP Adv. 2012, 2, 032160. [Google Scholar] [CrossRef]
- Ushikubo, F.Y.; Birribilli, F.S.; Oliveira, D.R.B.; Cunha, R.L. Y- and T-junction microfluidic devices: Effect of fluids and interface properties and operating conditions. Microfluid. Nanofluid. 2014, 17, 711–720. [Google Scholar] [CrossRef]
- Watkins, N.; Venkatesan, B.M.; Toner, M.; Rodriguez, W.; Bashir, R. A robust electrical microcytometer with 3-dimensional hydrofocusing. Lab Chip 2009, 9, 3177–3184. [Google Scholar] [CrossRef] [PubMed]
- Bryan, A.K.; Goranov, A.; Amon, A.; Manalis, S.R. Measurement of mass, density, and volume during the cell cycle of yeast. Proc. Natl. Acad. Sci. USA 2010, 107, 999–1004. [Google Scholar] [CrossRef]
- Ridler, T.W.; Calvard, S. Picture Thresholding Using an Iterative Selection Method. IEEE Trans. Syst. Man Cybern. 1978, 8, 630–632. [Google Scholar]
- Salvi, M.; Molinari, F. Multi-tissue and multi-scale approach for nuclei segmentation in H&E stained images. Biomed. Eng. Online 2018, 17, 89. [Google Scholar] [PubMed]
- Paszke, A.; Chanan, G.; Lin, Z.; Gross, S.; Yang, E.; Antiga, L.; Devito, Z. Automatic differentiation in PyTorch. In Proceedings of the 31st Conference Neural Information Processing, Long Beach, CA, USA, 4–9 December 2017; pp. 1–4. [Google Scholar]
- Nair, V.; Hinton, G.E. Rectified Linear Units Improve Restricted Boltzmann Machines. In Proceedings of the 27th International Conference on Machine Learning, Haifa, Israel, 21–24 June 2010; p. 8. [Google Scholar]
- Xu, B.; Wang, N.; Chen, T.; Li, M. Empirical Evaluation of Rectified Activations in Convolutional Network. arXiv 2015, arXiv:1505.00853. [Google Scholar]
- Kingma, D.P.; Ba, J.L. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference for Learning Representations, San Diego, CA, USA, 7–9 May 2015; pp. 1–15. [Google Scholar]
- Koch, G.; Zemel, R.; Salakhutdinov, R. Siamese Neural Networks for One-shot Image Recognition. In Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; p. 8. [Google Scholar]
- Watanabe, S. Information Theoretical Analysis of Multivariate Correlation. IBM J. Res. Dev. 1960, 4, 66–82. [Google Scholar] [CrossRef]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. arXiv 2015, arXiv:1502.03167. [Google Scholar]
- Rezende, D.J.; Mohamed, S.; Wierstra, D. Stochastic Backpropagation and Approximate Inference in Deep Generative Models. arXiv 2014, arXiv:1401.4082. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Van der Maaten, L.; Hinton, G. Visualizing Data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Wyatt Shields, C., IV; Reyes, C.D.; López, G.P. Microfluidic cell sorting: A review of the advances in the separation of cells from debulking to rare cell isolation. Lab Chip 2015, 15, 1230–1249. [Google Scholar] [CrossRef]
- Kuo, J.S.; Chiu, D.T. Controlling Mass Transport in Microfluidic Devices. Annu. Rev. Anal. Chem. 2011, 4, 275–296. [Google Scholar] [CrossRef]
- Salmon, J.B.; Ajdari, A. Transverse transport of solutes between co-flowing pressure-driven streams for microfluidic studies of diffusion/reaction processes. J. Appl. Phys. 2007, 101, 074902. [Google Scholar] [CrossRef]
- Kuntaegowdanahalli, S.S.; Bhagat, A.A.; Kumar, G.; Papautsky, I. Inertial microfluidics for continuous particle separation in spiral microchannels. Lab Chip 2009, 9, 2973–2980. [Google Scholar] [CrossRef]
- Carlo, D. Di Inertial microfluidics. Lab Chip 2009, 9, 3038–3046. [Google Scholar] [CrossRef]
- Yu, C.; Qian, X.; Chen, Y.; Yu, Q.; Ni, K.; Wang, X. Three-Dimensional Electro-Sonic Flow Focusing Ionization Microfluidic Chip for Mass Spectrometry. Micromachines 2015, 6, 1890–1902. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Constantinou, I.; Jendrusch, M.; Aspert, T.; Görlitz, F.; Schulze, A.; Charvin, G.; Knop, M. Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow. Micromachines 2019, 10, 311. https://doi.org/10.3390/mi10050311
Constantinou I, Jendrusch M, Aspert T, Görlitz F, Schulze A, Charvin G, Knop M. Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow. Micromachines. 2019; 10(5):311. https://doi.org/10.3390/mi10050311
Chicago/Turabian StyleConstantinou, Iordania, Michael Jendrusch, Théo Aspert, Frederik Görlitz, André Schulze, Gilles Charvin, and Michael Knop. 2019. "Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow" Micromachines 10, no. 5: 311. https://doi.org/10.3390/mi10050311
APA StyleConstantinou, I., Jendrusch, M., Aspert, T., Görlitz, F., Schulze, A., Charvin, G., & Knop, M. (2019). Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow. Micromachines, 10(5), 311. https://doi.org/10.3390/mi10050311





