Putative Nonribosomal Peptide Synthetase and Cytochrome P450 Genes Responsible for Tentoxin Biosynthesis in Alternaria alternata ZJ33
Abstract
:1. Introduction
2. Results
2.1. Identification of Isolate ZJ33
2.2. PCR Amplification of NRPS Gene Fragments
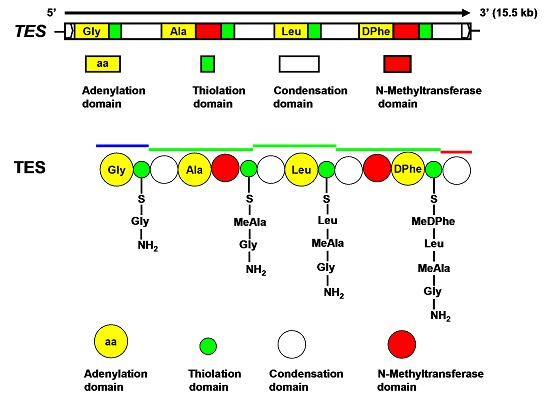
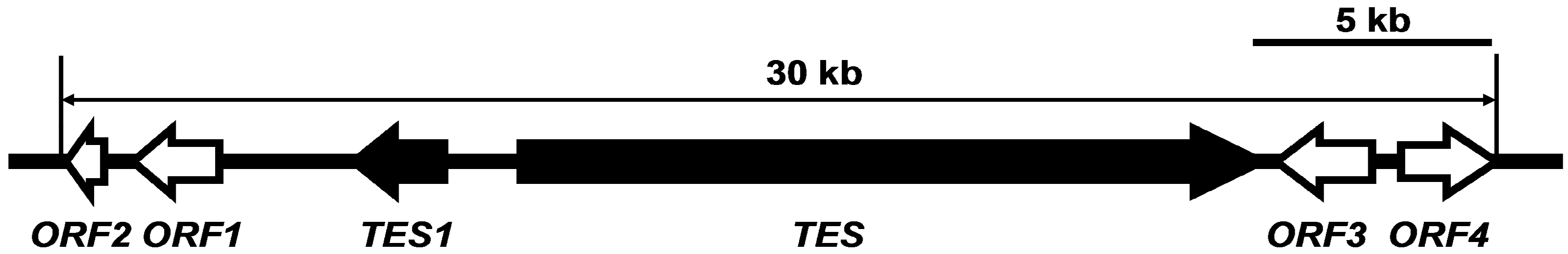
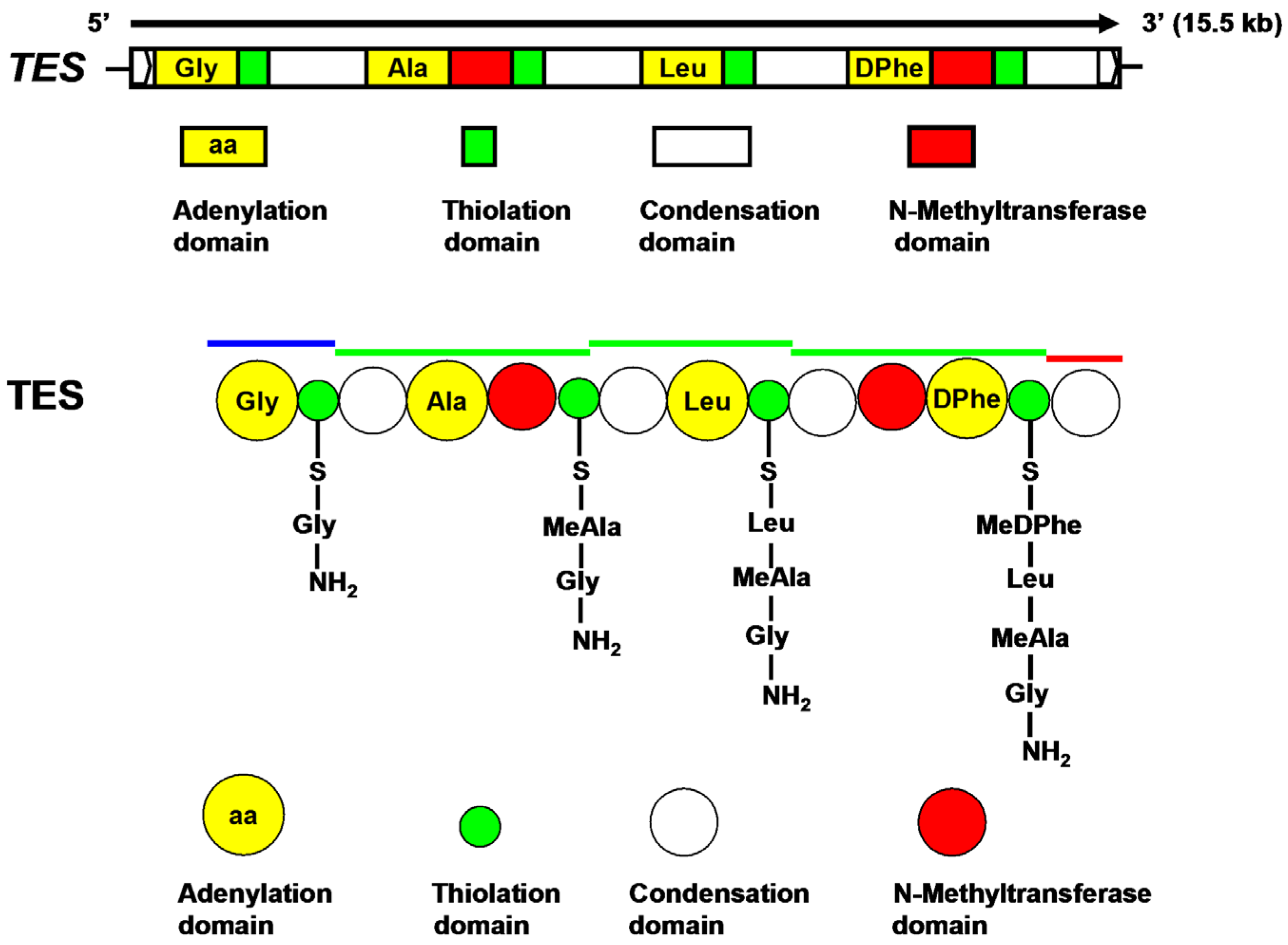
2.3. Characterization of TES and TES1
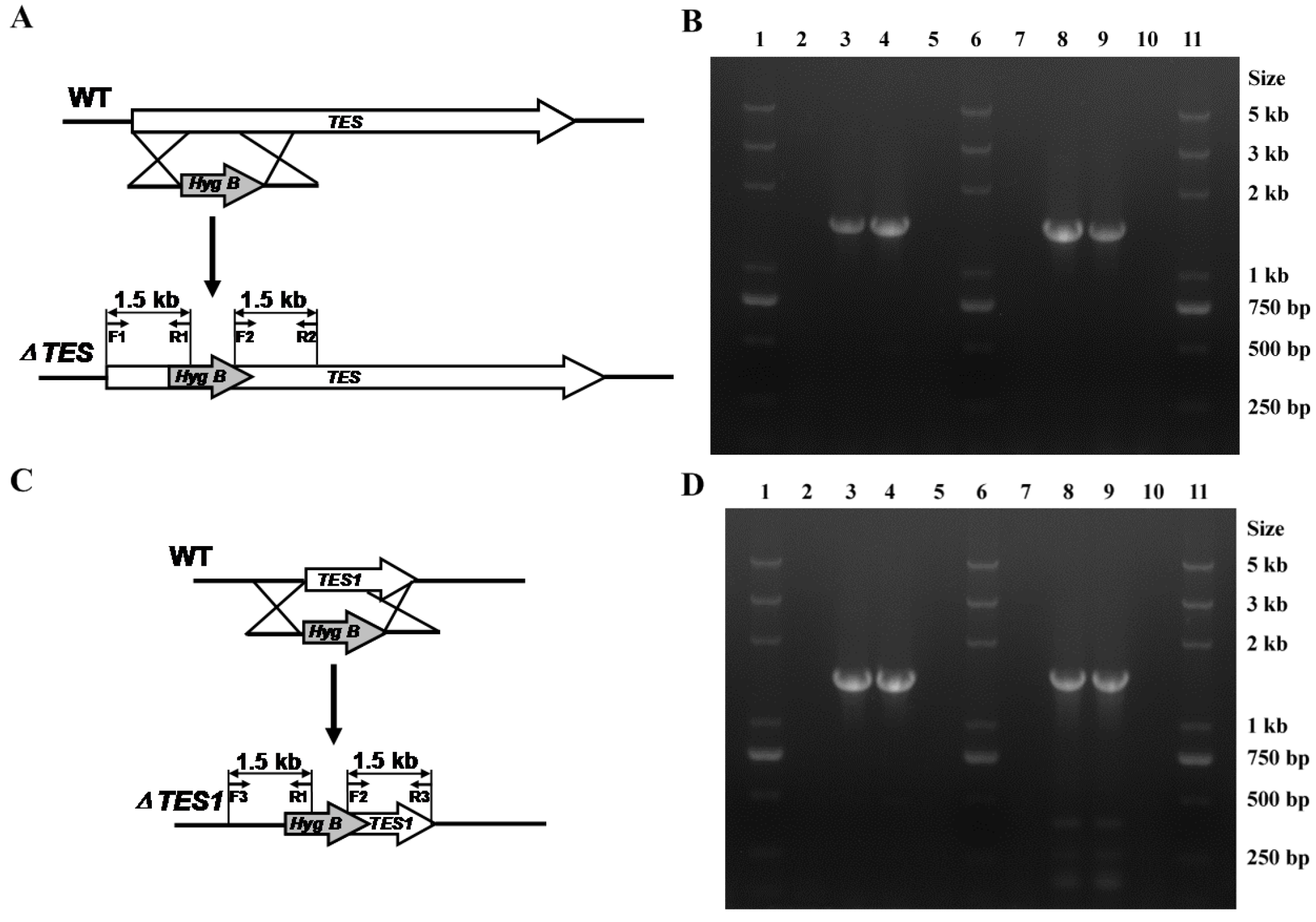
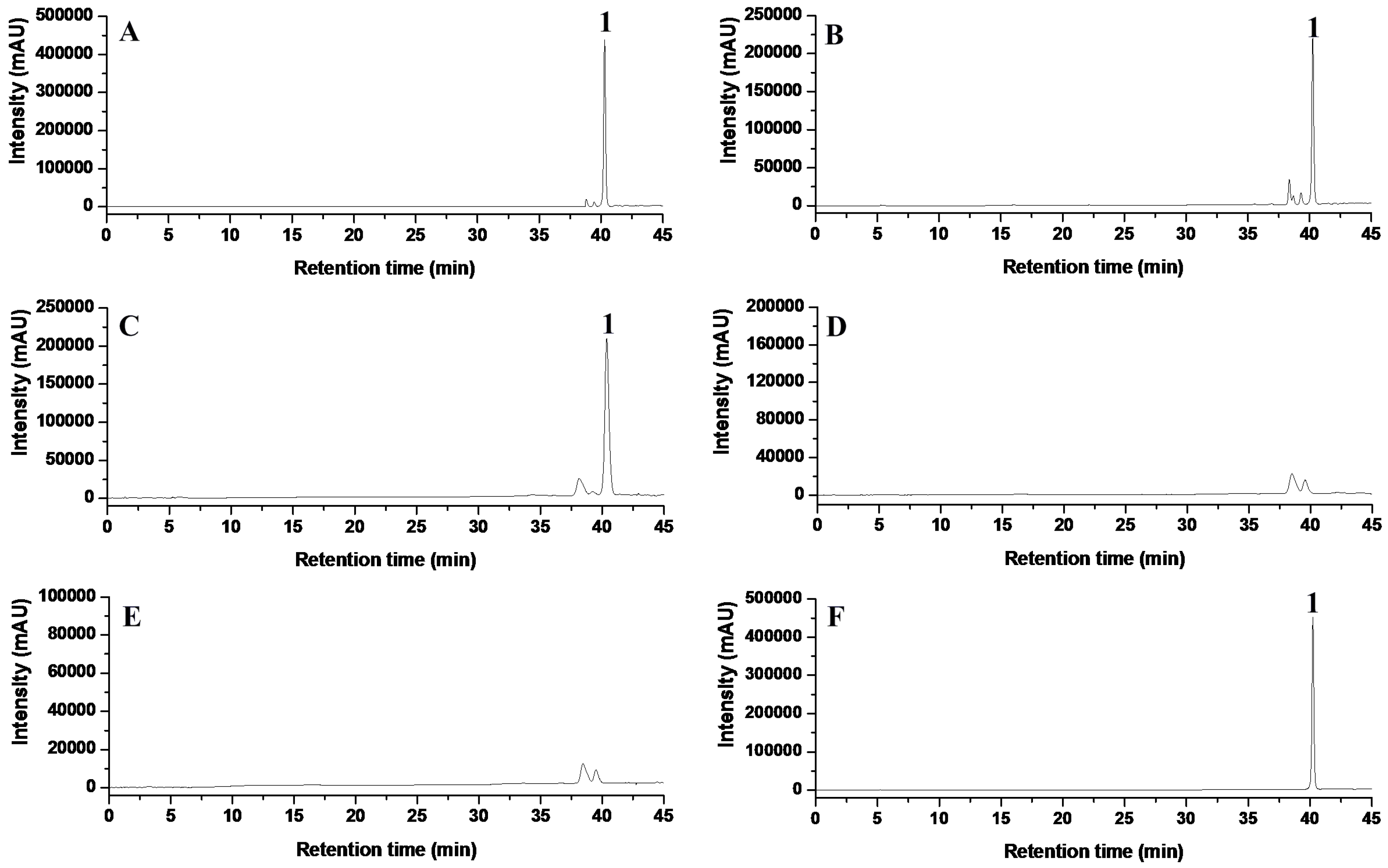
2.4. Targeted Disruption of TES and TES1 in A. alternata ZJ33
3. Discussion
4. Experimental Section
4.1. Strains, Media, and Culture Conditions
4.2. Nucleic Acid Manipulations
4.3. Identification of Fungi
4.4. Cloning of NRPS Genes from A. alternata ZJ33
4.5. Accession Numbers
4.6. Double-Joint PCR (DJ-PCR) for Constructing Mutant Alleles
4.7. Targeted Mutagenesis
4.8. High-Performance Liquid Chromatography (HPLC) Analysis of Tentoxin
Supplementary Materials
Acknowledgments
Author Contributions
Conflict of Interest
References
- Barkai-Golan, R. Alternaria Mycotoxins. In Mycotoxins in Fruits and Vegetables; Elsevier Inc.: Amsterdam, The Netherlands, 2008; pp. 185–203. [Google Scholar]
- Saad, S.M.; Halloin, J.M.; Hagedorn, D.J. Production, Purification, and bioassay of tentoxin. Phytopathology 1970, 60, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Meyer, W.L.; Templeton, G.E.; Grable, C.I.; Sigel, C.W.; Jones, R.; Woodhead, S.H.; Sauer, C. The structure of tentoxin. Tetrahedron Lett. 1971, 12, 2357–2360. [Google Scholar] [CrossRef]
- Liebermann, B.; Oertel, B. Bildung und isolierung des phytotoxins tentoxin aus Alternaria alternata. J. Basic Microbiol. 1983, 23, 503–511. [Google Scholar] [CrossRef]
- Kono, Y.; Gardner, J.M.; Takeuchi, S. Nonselective phytotoxins simultaneously produced with hostselective ACTG-toxins by a pathotype of Alternaria citri causing brown spot. Agric. Biol. Chem. 1986, 50, 2401–2403. [Google Scholar]
- Suemitsu, R.; Ohnishi, K.; Nobuhara, T.; Horiuchi, M.; Horiuchi, K. Isolation and identification of tentoxin from Alternaria porri (Ellis) Ciferri. Agric. Biol. Chem. 1990, 54, 2449–2450. [Google Scholar] [CrossRef]
- Steele, J.A.; Uchytil, T.F.; Durbin, R.D.; Bhatnagar, P.K.; Rich, D.H. Chloroplast coupling factor 1: A species-specific receptor for tentoxin. Proc. Natl. Acad. Sci. USA 1976, 73, 2245–2248. [Google Scholar] [CrossRef] [PubMed]
- Steele, J.A.; Uchytil, T.F.; Durbin, R.D. The stimulation of coupling factor 1 ATPase by tentoxin. Biochim. Biophys. Acta 1978, 504, 136–141. [Google Scholar] [CrossRef]
- Meiss, E.; Konno, H.; Groth, G.; Hisabori, T. Molecular processes of inhibition and stimulation of ATP synthase caused by the phytotoxin tentoxin. J. Biol. Chem. 2008, 283, 24548–24553. [Google Scholar] [CrossRef] [PubMed]
- Finking, R.; Marahiel, M.A. Biosynthesis of nonribosomal peptides. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef] [PubMed]
- Caboche, S.; Pupin, M.; Leclère, V.; Fontaine, A.; Jacques, P.; Kucherov, G. NORINE: A database of nonribosomal peptides. Nucleic Acids Res. 2008, 36, D326–D331. [Google Scholar] [CrossRef] [PubMed]
- Strieker, M.; Tanović, A.; Marahiel, M.A. Nonribosomal peptide synthetases: Structures and dynamics. Curr. Opin. Struct. Biol. 2010, 20, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Doekel, S.; Marahiel, M.A. Biosynthesis of natural products on modular peptide synthetases. Metab. Eng. 2001, 3, 64–77. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R.D.; Johnson, L.; Itoh, Y.; Kodama, M.; Otani, H.; Kohmoto, K. Cloning and characterization of a cyclic peptide synthetase gene from Alternaria alernata apple pathotype whose product is involved in AM-toxin synthesis and pathogenicity. Mol. Plant Microbe Interact. 2000, 13, 742–753. [Google Scholar] [CrossRef] [PubMed]
- Wight, W.D.; Labuda, R.; Walton, J.D. Conservation of the genes for HC-toxin biosynthesis in Alternaria jesenskae. BMC Microbiol. 2013, 13, 165. [Google Scholar] [CrossRef] [PubMed]
- Guillemette, T.; Sellam, A.; Simoneau, P. Analysis of a nonribosomal peptide synthetase gene from Alternaria brassicae and flanking genomic sequences. Curr. Genet. 2004, 45, 214–224. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.Y.; Cho, Y.; La Tota, M.; Cramer, R.A.; Lawrence, C.B. Functional analysis of the Alternaria brassicicola non-ribosomal peptide synthetase gene AbNPS2 reveals a role in conidial cell wall construction. Mol. Plant Pathol. 2007, 8, 23–39. [Google Scholar] [CrossRef] [PubMed]
- De Bruyne, L.; Van Poucke, C.; Di Mavungu, D.J.; Zainudin, N.A.I.I.M.; Vanhaecke, L.; De Vleesschauwer, D.; Turgeon, B.G.; De Saeger, S.; HÖfte, M. Comparative chemical screening and genetic analysis reveal tentoxin as a new virulence factor in Cochliobolus miyabeanus, the causal agent of brown spot disease on rice. Mol. Plant Pathol. 2015. [Google Scholar] [CrossRef]
- Woudenberg, J.H.C.; Groenewald, J.Z.; Binder, M.; Crous, P.W. Alternaria redefined. Stud. Mycol. 2013, 75, 171–212. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.N.; Kroken, S.; Chou, D.Y.T.; Robbertse, B.; Yodder, O.C.; Turgeon, B.G. Functional analysis of all nonribosomal peptide synthetases in Cochliobolus heterostrophus reveals a factor, NPS6, involved in virulence and resistance to oxidative stress. Eukaryot. Cell 2005, 4, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Condon, B.J.; Leng, Y.; Wu, D.; Bushley, K.E.; Ohm, R.A.; Otillar, R.; Martin, J.; Schackwitz, W.; Grimwood, J.; MohdZainudin, N.; et al. Comparative genome structure, secondary metabolite, and effector coding capacity across Cochliobolus pathogens. PLoS Genet. 2013, 9, e1003233. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Dai, X.; Qiang, S.; Tang, Y. Effect of a nonhost-selective toxin from Alternaria alternata on chloroplast-electron transfer activity in Eupatorium adenophorum. Plant Pathol. 2005, 54, 671–677. [Google Scholar] [CrossRef]
- Qiang, S.; Chang, Y.; Wan, Z.X.; Li, Y.H. Comparison on pathogenicity and other characteristics of five isolates of Alternaria alternata from Eupatorium adenophorum. J. Nanjing Agric. Univ. 2002, 25, 23–27. [Google Scholar]
- Qiang, S.; Zhu, Y.; Summerell, B.A.; Li, Y. Mycelium of Alternaria alternata as a potential biological control agent for Eupatorium adenophorum. Biocontrol Sci. Technol. 2006, 16, 653–668. [Google Scholar] [CrossRef]
- Liebermann, B.; Ramm, K. N-Methylation in the biosynthesis of the phytotoxin tentoxin. Phytochemistry 1991, 30, 1815–1817. [Google Scholar] [CrossRef]
- Ramm, K.; Ramm, M.; Liebermann, B.; Reuter, G. Studies of the biosynthesis of tentoxin by Alternaria alternata. Microbiology 1994, 140, 3257–3266. [Google Scholar] [CrossRef] [PubMed]
- Stachelhaus, T.; Mootz, H.D.; Marahiel, M.A. The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem. Biol. 1999, 6, 493–505. [Google Scholar] [CrossRef]
- Črešnar, B.; Petrič, Š. Cytochrome P450 enzymes in the fungal kingdom. Biochim. Biophys. Acta 2011, 1814, 29–35. [Google Scholar]
- Tohge, T.; Watanabe, M.; Hoefgen, R.; Fernie, A.R. Shikimate and phenylalanine biosynthesis in the green lineage. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.M.; Lee, S.; Lee, J.; Baek, S.R.; Kim, J.C.; Yun, S.H.; Park, S.Y.; Kang, S.; Lee, Y.W. Functional characterization and manipulation of the apicidin biosynthetic pathway in Fusarium semitectum. Mol. Microbiol. 2010, 76, 456–466. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: London, UK, 1990; pp. 315–322. [Google Scholar]
- Pryor, B.M.; Bigelow, D.M. Molecular characterization of Embellisia and Nimbya species and their relationship to Alternaria, Ulocladium and Stemphylium. Mycologia 2003, 95, 1141–1154. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Wang, K.; Yu, X.; Liu, J.; Zhang, H.; Zhou, F.; Xie, B.; Li, S. Transcription factor CCG-8 as a new regulator in the adaptation to antifungal azole stress. Antimicrob. Agents Chemother. 2014, 58, 1434–1442. [Google Scholar] [CrossRef] [PubMed]
- Shabana, Y.M.; Charudattan, R. Preparation and regeneration of mycelial protoplasts of Alternaria eichhorniae. J. Phytopathol. 1997, 145, 335–338. [Google Scholar] [CrossRef]

| Domain | Position | Substrate | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 235 | 236 | 239 | 278 | 299 | 301 | 322 | 330 | 331 | ||
| A1 | D | I | A | Q | V | G | V | I | W | Gly |
| A2 | D | V | W | F | C | G | G | T | F | Ala |
| A3 | D | A | L | L | V | G | A | V | S | Leu |
| A4 | D | G | W | F | L | A | A | V | M | DPhe |
| Primer a | Sequence b (5′→3′) |
|---|---|
| ITS4 | TCCTCCGCTTATTGATATGC |
| ITS5 | GGAAGTAAAAGTCGTAACAAGG |
| cps1 | AATCTAGATAYGGNCCNACNGA |
| cps2 | CCTCTAGANAGRTCNCCNGTYTTR |
| NRPS-for | GAGGCAAGGCAACCGCAACGATGA |
| NRPS-rev | CCCTTCATGTCGGGACTTGCGACA |
| HygB-for | CCGGGCTGCAGGAATTCGAT |
| HygB-rev | GGATCCCGGTCGGCATCTAC |
| TES-5for | CGGGATCGCTACTGTTTGACGTCA |
| TES-5rev | ATCGAATTCCTGCAGCCCGGTTGCCTTGCCTCAGCCAGA |
| TES-3for | GTAGATGCCGACCGGGATCCGCTAAAGTGGACCGGCAGA |
| TES-3rev | CATCGATCCCGATTGGCGTTCACA |
| TES-5nest | GAAGATATGGGAGAGAAACCGCGG |
| TES-3nest | AATGGTGGCATCGTTCTGGCCAGT |
| P450-5for | ACCTGACGAAACTCACTGCCTCCT |
| P450-5rev | ATCGAATTCCTGCAGCCCGGCTCAGGCTCCATCTTTGGAG |
| P450-3for | GTAGATGCCGACCGGGATCCAGGGCCTGGAGGAAACCTACAC |
| P450-3rev | GCTCGACCATTGGATACCTAGCCT |
| P450-5nest | ACAATGGCCTAAGTCTGCCGCTCA |
| P450-3nest | CCCTCAACATTCCCTGGCATCTTG |
| Hyg-5rev | CGCACAAGTTATCGTGCACCAAGC |
| Hyg-3for | GGCGTATATGCTCCGCATTGGTCT |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.-H.; Han, W.-J.; Gui, X.-W.; Wei, T.; Tang, S.-Y.; Jin, J.-M. Putative Nonribosomal Peptide Synthetase and Cytochrome P450 Genes Responsible for Tentoxin Biosynthesis in Alternaria alternata ZJ33. Toxins 2016, 8, 234. https://doi.org/10.3390/toxins8080234
Li Y-H, Han W-J, Gui X-W, Wei T, Tang S-Y, Jin J-M. Putative Nonribosomal Peptide Synthetase and Cytochrome P450 Genes Responsible for Tentoxin Biosynthesis in Alternaria alternata ZJ33. Toxins. 2016; 8(8):234. https://doi.org/10.3390/toxins8080234
Chicago/Turabian StyleLi, You-Hai, Wen-Jin Han, Xi-Wu Gui, Tao Wei, Shuang-Yan Tang, and Jian-Ming Jin. 2016. "Putative Nonribosomal Peptide Synthetase and Cytochrome P450 Genes Responsible for Tentoxin Biosynthesis in Alternaria alternata ZJ33" Toxins 8, no. 8: 234. https://doi.org/10.3390/toxins8080234
APA StyleLi, Y.-H., Han, W.-J., Gui, X.-W., Wei, T., Tang, S.-Y., & Jin, J.-M. (2016). Putative Nonribosomal Peptide Synthetase and Cytochrome P450 Genes Responsible for Tentoxin Biosynthesis in Alternaria alternata ZJ33. Toxins, 8(8), 234. https://doi.org/10.3390/toxins8080234