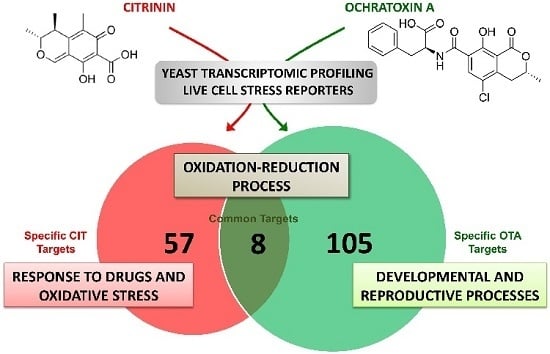
Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast
Abstract
:1. Introduction
2. Results
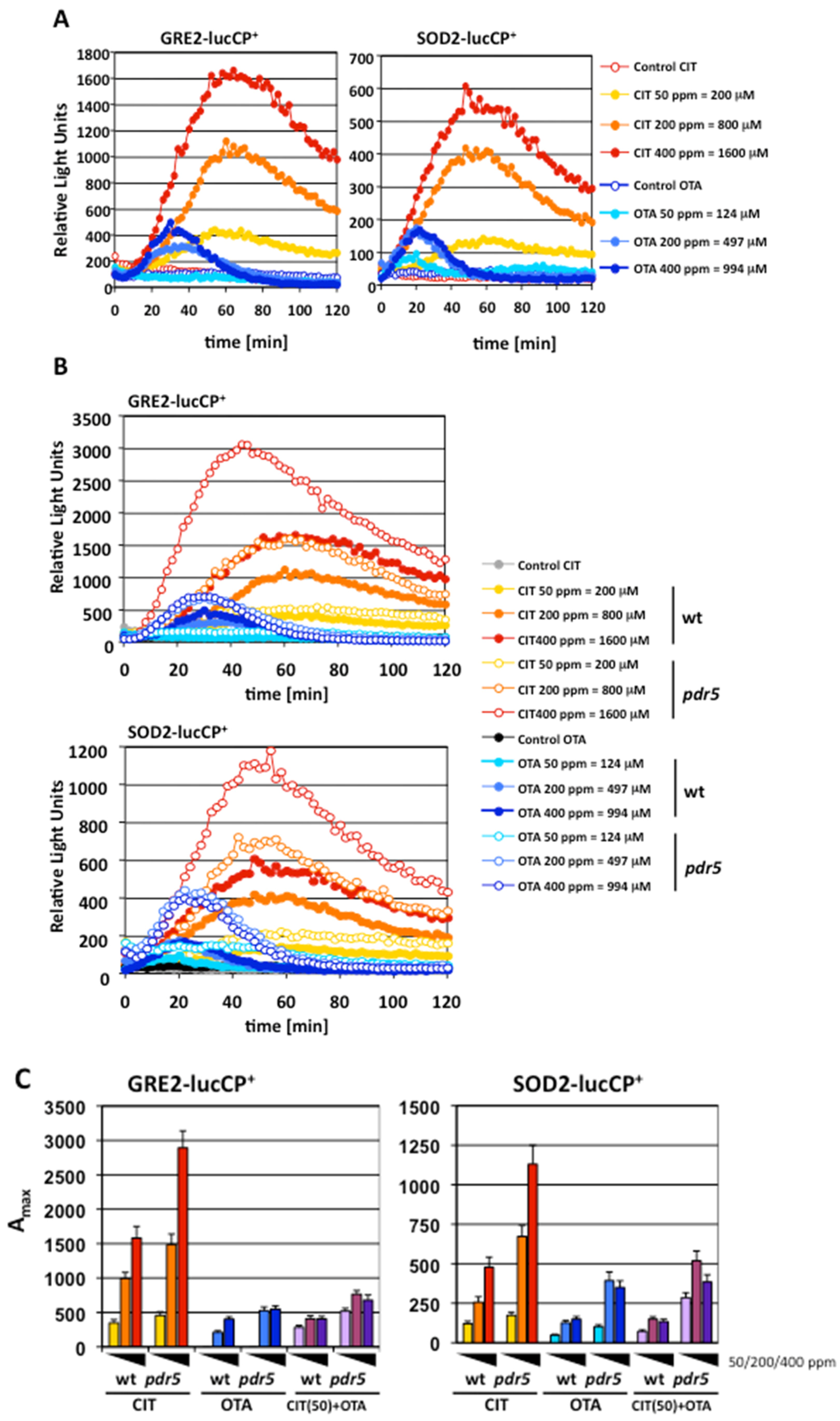
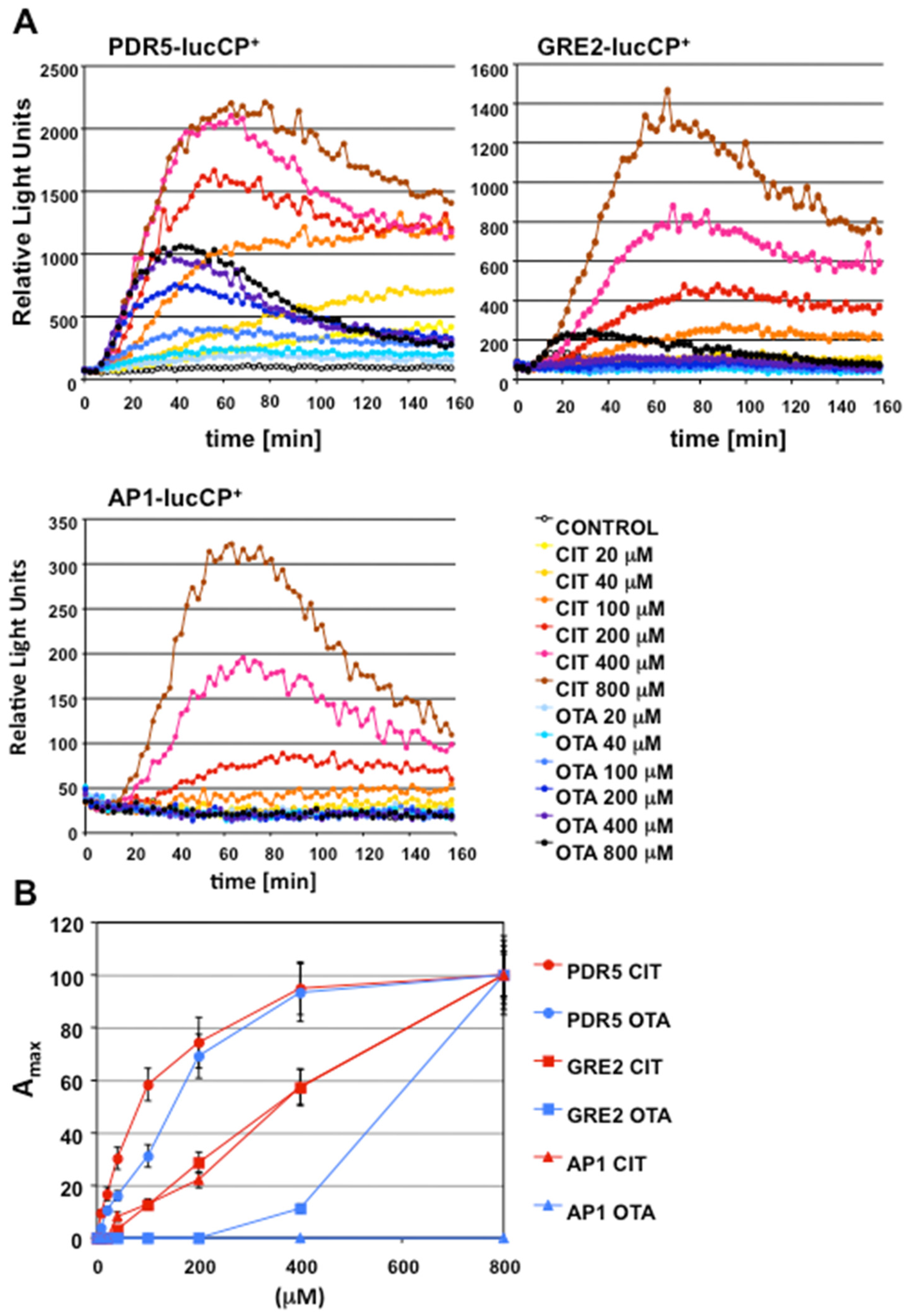
2.1. Gene Expression Profiles of Stress Response Genes upon CIT and OTA Exposure
2.2. Genomic Expression Profiles upon Separated and Combined Exposure to CIT and OTA
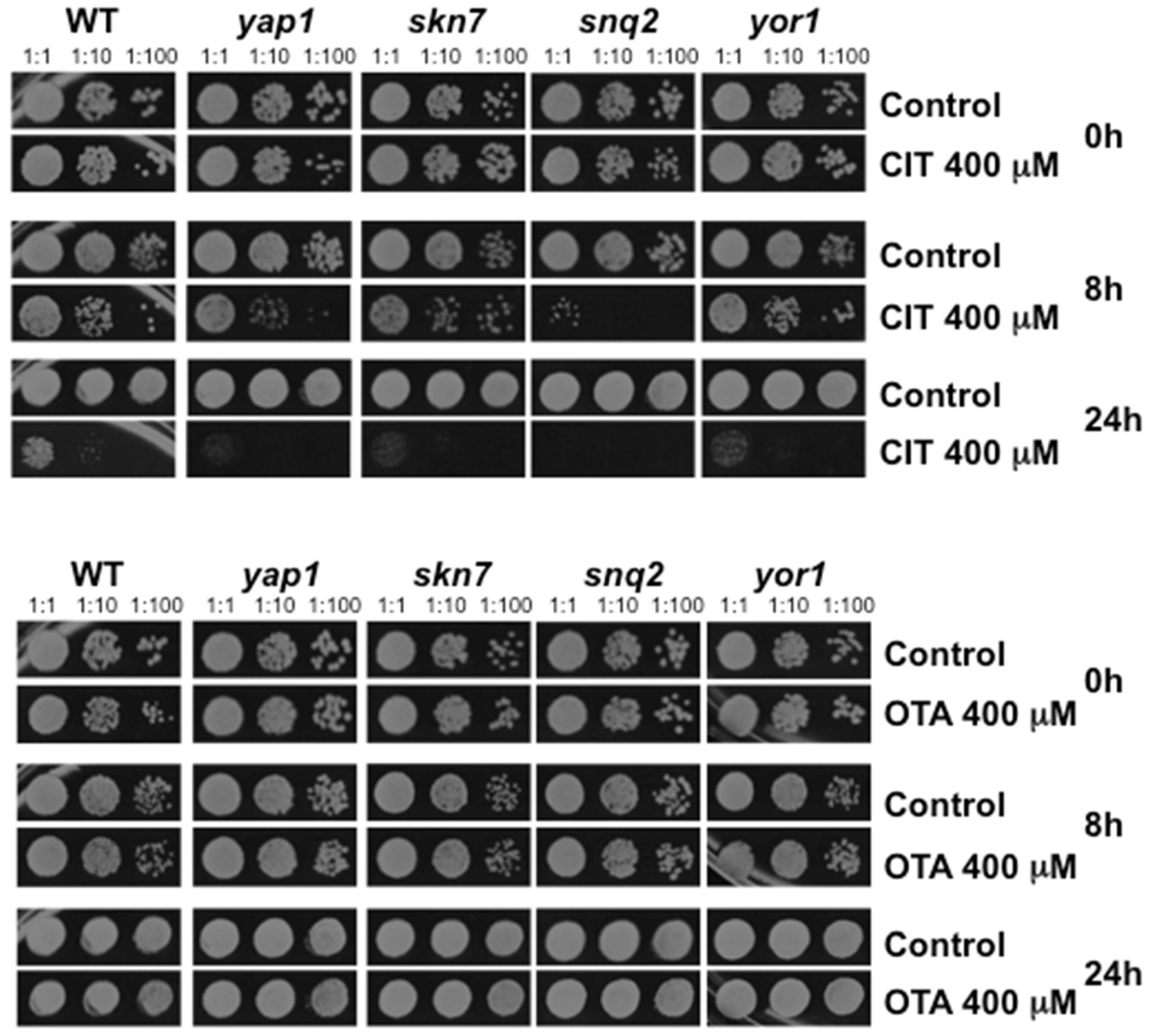
2.3. Oxidative Stress is a Hallmark for CIT, but not OTA, Toxicity
3. Discussion
4. Materials and Methods
4.1. Yeast Strains and Growth Conditions
4.2. Plasmid Constructions
4.3. Live Cell Luciferase Assays
4.4. Yeast Sensitivity Assays
4.5. Microarray Experiments and Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [PubMed]
- Marroquin-Cardona, A.G.; Johnson, N.M.; Phillips, T.D.; Hayes, A.W. Mycotoxins in a changing global environment—A review. Food Chem. Toxicol. 2014, 69, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Moretti, A.; Susca, A.; Mule, G.; Logrieco, A.F.; Proctor, R.H. Molecular biodiversity of mycotoxigenic fungi that threaten food safety. Int. J. Food Microbiol. 2013, 167, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Groopman, J.D.; Pestka, J.J. Public health impacts of foodborne mycotoxins. Annu. Rev. Food Sci. Technol. 2014, 5, 351–372. [Google Scholar] [CrossRef] [PubMed]
- Mobius, N.; Hertweck, C. Fungal phytotoxins as mediators of virulence. Curr. Opin. Plant. Biol. 2009, 12, 390–398. [Google Scholar] [CrossRef] [PubMed]
- Doi, K.; Uetsuka, K. Mechanisms of Mycotoxin-induced Dermal Toxicity and Tumorigenesis through Oxidative Stress-related Pathways. J. Toxicol. Pathol. 2014, 27, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Escriva, L.; Font, G.; Manyes, L. In vivo toxicity studies of fusarium mycotoxins in the last decade: A review. Food. Chem. Toxicol. 2015, 78, 185–206. [Google Scholar] [CrossRef] [PubMed]
- Vettorazzi, A.; Gonzalez-Penas, E.; de Cerain, A.L. Ochratoxin A kinetics: A review of analytical methods and studies in rat model. Food Chem. Toxicol. 2014, 72, 273–288. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, L.; Liu, F.; Wang, Q.; Selvaraj, J.N.; Xing, F.; Zhao, Y.; Liu, Y. Ochratoxin A Producing Fungi, Biosynthetic Pathway and Regulatory Mechanisms. Toxins 2016, 8. [Google Scholar] [CrossRef] [PubMed]
- Koszegi, T.; Poor, M. Ochratoxin A: Molecular Interactions, Mechanisms of Toxicity and Prevention at the Molecular Level. Toxins 2016, 8. [Google Scholar] [CrossRef] [PubMed]
- Faucet, V.; Pfohl-Leszkowicz, A.; Dai, J.; Castegnaro, M.; Manderville, R.A. Evidence for covalent DNA adduction by Ochratoxin A following chronic exposure to rat and subacute exposure to pig. Chem. Res. Toxicol. 2004, 17, 1289–1296. [Google Scholar] [CrossRef] [PubMed]
- Mantle, P.G.; Faucet-Marquis, V.; Manderville, R.A.; Squillaci, B.; Pfohl-Leszkowicz, A. Structures of covalent adducts between DNA and Ochratoxin A: A new factor in debate about genotoxicity and human risk assessment. Chem. Res. Toxicol. 2010, 23, 89–98. [Google Scholar] [CrossRef] [PubMed]
- Pfohl-Leszkowicz, A.; Manderville, R.A. An update on direct genotoxicity as a molecular mechanism of ochratoxin a carcinogenicity. Chem. Res. Toxicol. 2012, 25, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Rahimtula, A.D.; Bereziat, J.C.; Bussacchini-Griot, V.; Bartsch, H. Lipid peroxidation as a possible cause of ochratoxin A toxicity. Biochem. Pharmacol. 1988, 37, 4469–4477. [Google Scholar] [CrossRef]
- Sorrenti, V.; di Giacomo, C.; Acquaviva, R.; Barbagallo, I.; Bognanno, M.; Galvano, F. Toxicity of ochratoxin a and its modulation by antioxidants: A review. Toxins 2013, 5, 1742–1766. [Google Scholar] [CrossRef] [PubMed]
- Bragulat, M.R.; Martinez, E.; Castella, G.; Cabanes, F.J. Ochratoxin A and citrinin producing species of the genus Penicillium from feedstuffs. Int. J. Food. Microbiol. 2008, 126, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Vrabcheva, T.; Usleber, E.; Dietrich, R.; Martlbauer, E. Co-occurrence of ochratoxin A and citrinin in cereals from Bulgarian villages with a history of Balkan endemic nephropathy. J. Agric. Food. Chem. 2000, 48, 2483–2488. [Google Scholar] [CrossRef] [PubMed]
- Ostry, V.; Malir, F.; Ruprich, J. Producers and important dietary sources of ochratoxin A and citrinin. Toxins 2013, 5, 1574–1586. [Google Scholar] [CrossRef] [PubMed]
- Schmidt-Heydt, M.; Graf, E.; Stoll, D.; Geisen, R. The biosynthesis of ochratoxin A by Penicillium as one mechanism for adaptation to NaCl rich foods. Food Microbiol. 2012, 29, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Schmidt-Heydt, M.; Stoll, D.; Schutz, P.; Geisen, R. Oxidative stress induces the biosynthesis of citrinin by Penicillium verrucosum at the expense of ochratoxin. Int. J. Food Microbiol. 2015, 192, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Stoll, D.; Schmidt-Heydt, M.; Geisen, R. Differences in the regulation of ochratoxin A by the HOG pathway in Penicillium and Aspergillus in response to high osmolar environments. Toxins 2013, 5, 1282–1298. [Google Scholar] [CrossRef] [PubMed]
- Flajs, D.; Peraica, M. Toxicological properties of citrinin. Arh. Hig. Rada Toksikol. 2009, 60, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Bouslimi, A.; Ouannes, Z.; Golli, E.E.; Bouaziz, C.; Hassen, W.; Bacha, H. Cytotoxicity and oxidative damage in kidney cells exposed to the mycotoxins ochratoxin a and citrinin: Individual and combined effects. Toxicol. Mech. Methods 2008, 18, 341–349. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.H. Citrinin induces apoptosis via a mitochondria-dependent pathway and inhibition of survival signals in embryonic stem cells, and causes developmental injury in blastocysts. Biochem. J. 2007, 404, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Dwivedi, P.; Sharma, A.K.; Sankar, M.; Patil, R.D.; Singh, N.D. Apoptosis and lipid peroxidation in ochratoxin A- and citrinin-induced nephrotoxicity in rabbits. Toxicol. Ind. Health 2014, 30, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Dwivedi, P.D.; Dhawan, A.; Das, M.; Ansari, K.M. Citrinin-generated reactive oxygen species cause cell cycle arrest leading to apoptosis via the intrinsic mitochondrial pathway in mouse skin. Toxicol. Sci. 2011, 122, 557–566. [Google Scholar] [CrossRef] [PubMed]
- Mate, G.; Gazdag, Z.; Mike, N.; Papp, G.; Pocsi, I.; Pesti, M. Regulation of oxidative stress-induced cytotoxic processes of citrinin in the fission yeast Schizosaccharomyces pombe. Toxicon 2014, 90, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Pascual-Ahuir, A.; Vanacloig-Pedros, E.; Proft, M. Toxicity mechanisms of the food contaminant citrinin: Application of a quantitative yeast model. Nutrients 2014, 6, 2077–2087. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, S.M.; Chagas, G.M.; Campello, A.P.; Kluppel, M.L. Mechanism of citrinin-induced dysfunction of mitochondria. V. Effect on the homeostasis of the reactive oxygen species. Cell. Biochem. Funct. 1997, 15, 203–209. [Google Scholar] [CrossRef]
- Singh, N.D.; Sharma, A.K.; Dwivedi, P.; Leishangthem, G.D.; Rahman, S.; Reddy, J.; Kumar, M. Effect of feeding graded doses of citrinin on apoptosis and oxidative stress in male Wistar rats through the F1 generation. Toxicol. Ind. Health 2016, 32, 385–397. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.Y.; Liao, Y.C.; Chang, C.H.; Liu, B.H. Citrinin induces apoptosis in HL-60 cells via activation of the mitochondrial pathway. Toxicol. Lett. 2006, 161, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Follmann, W.; Behm, C.; Degen, G.H. Toxicity of the mycotoxin citrinin and its metabolite dihydrocitrinone and of mixtures of citrinin and ochratoxin A in vitro. Arch. Toxicol. 2014, 88, 1097–1107. [Google Scholar] [CrossRef] [PubMed]
- Klaric, M.S.; Rasic, D.; Peraica, M. Deleterious effects of mycotoxin combinations involving ochratoxin A. Toxins 2013, 5, 1965–1987. [Google Scholar] [CrossRef] [PubMed]
- Afshari, C.A.; Hamadeh, H.K.; Bushel, P.R. The evolution of bioinformatics in toxicology: Advancing toxicogenomics. Toxicol. Sci. 2011, 120 (Suppl. 1), S225–S237. [Google Scholar] [CrossRef] [PubMed]
- Yasokawa, D.; Iwahashi, H. Toxicogenomics using yeast DNA microarrays. J. Biosci. Bioeng. 2010, 110, 511–522. [Google Scholar] [CrossRef] [PubMed]
- Arbillaga, L.; Azqueta, A.; van Delft, J.H.; Lopez de Cerain, A. In vitro gene expression data supporting a DNA non-reactive genotoxic mechanism for ochratoxin A. Toxicol. Appl. Pharmacol. 2007, 220, 216–224. [Google Scholar] [CrossRef] [PubMed]
- Hibi, D.; Kijima, A.; Kuroda, K.; Suzuki, Y.; Ishii, Y.; Jin, M.; Nakajima, M.; Sugita-Konishi, Y.; Yanai, T.; Nohmi, T.; et al. Molecular mechanisms underlying ochratoxin A-induced genotoxicity: Global gene expression analysis suggests induction of DNA double-strand breaks and cell cycle progression. J. Toxicol. Sci. 2013, 38, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Hundhausen, C.; Boesch-Saadatmandi, C.; Matzner, N.; Lang, F.; Blank, R.; Wolffram, S.; Blaschek, W.; Rimbach, G. Ochratoxin a lowers mRNA levels of genes encoding for key proteins of liver cell metabolism. Cancer Genom. Proteom. 2008, 5, 319–332. [Google Scholar]
- Marin-Kuan, M.; Nestler, S.; Verguet, C.; Bezencon, C.; Piguet, D.; Mansourian, R.; Holzwarth, J.; Grigorov, M.; Delatour, T.; Mantle, P.; et al. A toxicogenomics approach to identify new plausible epigenetic mechanisms of ochratoxin a carcinogenicity in rat. Toxicol. Sci. 2006, 89, 120–134. [Google Scholar] [CrossRef] [PubMed]
- Vettorazzi, A.; van Delft, J.; Lopez de Cerain, A. A review on ochratoxin A transcriptomic studies. Food Chem. Toxicol. 2013, 59, 766–783. [Google Scholar] [CrossRef] [PubMed]
- Iwahashi, H.; Kitagawa, E.; Suzuki, Y.; Ueda, Y.; Ishizawa, Y.H.; Nobumasa, H.; Kuboki, Y.; Hosoda, H.; Iwahashi, Y. Evaluation of toxicity of the mycotoxin citrinin using yeast ORF DNA microarray and Oligo DNA microarray. BMC Genomics 2007, 8, 95. [Google Scholar] [CrossRef] [PubMed]
- Toone, W.M.; Morgan, B.A.; Jones, N. Redox control of AP-1-like factors in yeast and beyond. Oncogene 2001, 20, 2336–2346. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Wang, J.; Liu, B.; Wang, Z.; Yuan, Y.; Yue, T. Effect of yeast cell morphology, cell wall physical structure and chemical composition on patulin adsorption. PLoS ONE 2015, 10, e0136045. [Google Scholar] [CrossRef] [PubMed]
- Piotrowska, M.; Masek, A. Saccharomyces cerevisiae cell wall components as tools for ochratoxin A decontamination. Toxins 2015, 7, 1151–1162. [Google Scholar] [CrossRef] [PubMed]
- Jungwirth, H.; Kuchler, K. Yeast ABC transporters—A tale of sex, stress, drugs and aging. FEBS Lett. 2006, 580, 1131–1138. [Google Scholar] [CrossRef] [PubMed]
- Prasad, R.; Goffeau, A. Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu. Rev. Microbiol. 2012, 66, 39–63. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.K.; Arthanari, H.; Yang, F.; Pan, S.J.; Fan, X.; Breger, J.; Frueh, D.P.; Gulshan, K.; Li, D.K.; Mylonakis, E.; et al. A nuclear receptor-like pathway regulating multidrug resistance in fungi. Nature 2008, 452, 604–609. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.C.; Chan, W.H. Inhibition of citrinin-induced apoptotic biochemical signaling in human hepatoma G2 cells by resveratrol. Int. J. Mol. Sci. 2009, 10, 3338–3357. [Google Scholar] [CrossRef] [PubMed]
- Gayathri, L.; Dhivya, R.; Dhanasekaran, D.; Periasamy, V.S.; Alshatwi, A.A.; Akbarsha, M.A. Hepatotoxic effect of ochratoxin A and citrinin, alone and in combination, and protective effect of vitamin E: In vitro study in HepG2 cell. Food Chem. Toxicol. 2015, 83, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Aleo, M.D.; Wyatt, R.D.; Schnellmann, R.G. The role of altered mitochondrial function in citrinin-induced toxicity to rat renal proximal tubule suspensions. Toxicol. Appl. Pharmacol. 1991, 109, 455–463. [Google Scholar] [CrossRef]
- Qi, X.; Yu, T.; Zhu, L.; Gao, J.; He, X.; Huang, K.; Luo, Y.; Xu, W. Ochratoxin A induces rat renal carcinogenicity with limited induction of oxidative stress responses. Toxicol. Appl. Pharmacol. 2014, 280, 543–549. [Google Scholar] [CrossRef] [PubMed]
- Taniai, E.; Yafune, A.; Nakajima, M.; Hayashi, S.M.; Nakane, F.; Itahashi, M.; Shibutani, M. Ochratoxin A induces karyomegaly and cell cycle aberrations in renal tubular cells without relation to induction of oxidative stress responses in rats. Toxicol. Lett. 2014, 224, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Govin, J.; Berger, S.L. Genome reprogramming during sporulation. Int. J. Dev. Biol. 2009, 53, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Winter, E. The Sum1/Ndt80 transcriptional switch and commitment to meiosis in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 2012, 76, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Grunstein, M.; Gasser, S.M. Epigenetics in Saccharomyces cerevisiae. Cold Spring Harb. Perspect. Biol. 2013, 5. [Google Scholar] [CrossRef] [PubMed]
- Pijnappel, W.W.; Schaft, D.; Roguev, A.; Shevchenko, A.; Tekotte, H.; Wilm, M.; Rigaut, G.; Seraphin, B.; Aasland, R.; Stewart, A.F. The S. cerevisiae SET3 complex includes two histone deacetylases, Hos2 and Hst1, and is a meiotic-specific repressor of the sporulation gene program. Genes Dev. 2001, 15, 2991–3004. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Pierce, M.; Gailus-Durner, V.; Wagner, M.; Winter, E.; Vershon, A.K. Sum1 and Hst1 repress middle sporulation-specific gene expression during mitosis in Saccharomyces cerevisiae. EMBO J. 1999, 18, 6448–6454. [Google Scholar] [CrossRef] [PubMed]
- Chalkiadaki, A.; Guarente, L. The multifaceted functions of sirtuins in cancer. Nat. Rev. Cancer 2015, 15, 608–624. [Google Scholar] [CrossRef] [PubMed]
- Roth, M.; Chen, W.Y. Sorting out functions of sirtuins in cancer. Oncogene 2014, 33, 1609–1620. [Google Scholar] [CrossRef] [PubMed]
- Dolz-Edo, L.; Rienzo, A.; Poveda-Huertes, D.; Pascual-Ahuir, A.; Proft, M. Deciphering dynamic dose responses of natural promoters and single cis elements upon osmotic and oxidative stress in yeast. Mol. Cell. Biol. 2013, 33, 2228–2240. [Google Scholar] [CrossRef] [PubMed]
- Rienzo, A.; Pascual-Ahuir, A.; Proft, M. The use of a real-time luciferase assay to quantify gene expression dynamics in the living yeast cell. Yeast 2012, 29, 219–231. [Google Scholar] [CrossRef] [PubMed]

| Gene | Standard Name | FC * | p-Value | Description |
|---|---|---|---|---|
| YPL171C | OYE3 | 473.1 | 3.00 × 10−8 | Conserved NADPH oxidoreductase containing flavin mononucleotide (FMN) |
| YFL056C | AAD6 | 252.4 | 9.60 × 10−7 | Putative aryl-alcohol dehydrogenase |
| YDL243C | AAD4 | 252.1 | 1.77 × 10−9 | Putative aryl-alcohol dehydrogenase |
| YCL026C-A | FRM2 | 177.2 | 1.77 × 10−5 | Type II nitroreductase |
| YLL060C | GTT2 | 142.4 | 4.50 × 10−5 | Glutathione S-transferase |
| YBR008C | FLR1 | 120.6 | 1.83 × 10−7 | Plasma membrane multidrug transporter of the major facilitator superfamily |
| YCL026C-B | HBN1 | 61.7 | 1.81 × 10−6 | Protein of unknown function |
| YGR213C | RTA1 | 57.8 | 1.77 × 10−8 | Protein involved in 7-aminocholesterol resistance |
| YML116W | ATR1 | 54.8 | 1.45 × 10−7 | Multidrug efflux pump of the major facilitator superfamily |
| YKR076W | ECM4 | 51.6 | 6.10 × 10−6 | Omega class glutathione transferase |
| YML131W | - | 41.2 | 9.44 × 10−3 | Protein of unknown function |
| YHR139C | SPS100 | 35.2 | 4.78 × 10−7 | Protein required for spore wall maturation |
| YFL057C | AAD16 | 33.5 | 2.12 × 10−2 | Putative aryl-alcohol dehydrogenase |
| YDR011W | SNQ2 | 31.1 | 6.78 × 10−7 | Plasma membrane ATP-binding cassette (ABC) transporter |
| YOL151W | GRE2 | 25.8 | 2.28 × 10−6 | 3-methylbutanal reductase and NADPH-dependent methylglyoxal reductase |
| YKL086W | SRX1 | 25.6 | 5.22 × 10−7 | Sulfiredoxin |
| YDR406W | PDR15 | 18.1 | 1.12 × 10−6 | Plasma membrane ATP binding cassette (ABC) transporter |
| YLR108C | - | 16.6 | 5.77 × 10−8 | Protein of unknown function |
| YDL020C | RPN4 | 15.8 | 9.41 × 10−8 | Transcription factor that stimulates expression of proteasome genes |
| YNL117W | MLS1 | 15.1 | 3.95 × 10−6 | Malate synthase |
| YOR328W | PDR10 | 14.5 | 2.41 × 10−8 | ATP-binding cassette (ABC) transporter |
| YHR199C | AIM46 | 13.3 | 1.58 × 10−7 | Putative protein of unknown function |
| YHR029C | YHI9 | 12.9 | 3.86 × 10−7 | Protein of unknown function |
| YGR256W | GND2 | 10.9 | 5.58 × 10−7 | 6-phosphogluconate dehydrogenase |
| YBR244W | GPX2 | 10.5 | 3.56 × 10−6 | Phospholipid hydroperoxide glutathione peroxidase |
| YFL030W | AGX1 | 10.3 | 4.28 × 10−8 | Alanine:glyoxylate aminotransferase (AGT) |
| YDR453C | TSA2 | 9.6 | 2.01 × 10−7 | Stress inducible cytoplasmic thioredoxin peroxidase |
| YER143W | DDI1 | 9.5 | 8.66 × 10−5 | DNA damage-inducible v-SNARE binding protein |
| YNR074C | AIF1 | 9.1 | 4.46 × 10−7 | Mitochondrial cell death effector |
| YER042W | MXR1 | 9.0 | 2.11 × 10−6 | Methionine-S-sulfoxide reductase |
| YJL101C | GSH1 | 8.9 | 1.30 × 10−7 | Gamma glutamylcysteine synthetase |
| YHR138C | - | 8.8 | 1.42 × 10−3 | Protein of unknown function |
| YHL036W | MUP3 | 8.6 | 1.13 × 10−5 | Low affinity methionine permease |
| YNL129W | NRK1 | 8.5 | 1.61 × 10−5 | Nicotinamide riboside kinase |
| YPR200C | ARR2 | 8.1 | 1.57 × 10−4 | Arsenate reductase |
| YER103W | SSA4 | 7.8 | 2.65 × 10−5 | Heat shock protein |
| YJL045W | - | 7.7 | 3.32 × 10−7 | Minor succinate dehydrogenase isozyme |
| YPL027W | SMA1 | 7.7 | 9.86 × 10−7 | Protein of unknown function involved in prospore membrane assembly |
| YGR010W | NMA2 | 7.5 | 1.02 × 10−7 | Nicotinic acid mononucleotide adenylyltransferase |
| YMR169C | ALD3 | 7.4 | 6.10 × 10−4 | Cytoplasmic aldehyde dehydrogenase |
| YDR132C | - | 7.3 | 1.74 × 10−6 | Protein of unknown function |
| YOR162C | YRR1 | 7.2 | 1.24 × 10−7 | Zn2-Cys6 zinc-finger transcription factor |
| YMR038C | CCS1 | 6.9 | 6.96 × 10−5 | Copper chaperone for superoxide dismutase Sod1p |
| YJL219W | HXT9 | 6.9 | 1.67 × 10−7 | Putative hexose transporter |
| YER142C | MAG1 | 6.8 | 5.46 × 10−7 | 3-methyl-adenine DNA glycosylase |
| YBR046C | ZTA1 | 6.7 | 1.13 × 10−5 | NADPH-dependent quinone reductase |
| YNL231C | PDR16 | 6.6 | 7.41 × 10−3 | Phosphatidylinositol transfer protein (PITP) |
| YPL091W | GLR1 | 6.5 | 1.49 × 10−5 | Cytosolic and mitochondrial glutathione oxidoreductase |
| YGR281W | YOR1 | 6.4 | 2.16 × 10−3 | Plasma membrane ATP-binding cassette (ABC) transporter |
| YGR197C | SNG1 | 6.3 | 3.47 × 10−7 | Protein involved in resistance to nitrosoguanidine and 6-azauracil |
| YNL155W | CUZ1 | 6.1 | 5.38 × 10−3 | Protein with a role in the ubiquitin-proteasome pathway |
| YAL054C | ACS1 | 6.1 | 3.74 × 10−7 | Acetyl-coA synthetase isoform |
| YOL119C | MCH4 | 6.1 | 1.27 × 10−5 | Protein with similarity to mammalian monocarboxylate permeases |
| YDL168W | SFA1 | 6.0 | 1.21 × 10−5 | Bifunctional alcohol dehydrogenase and formaldehyde dehydrogenase |
| YCR021C | HSP30 | 6.0 | 5.37 × 10−3 | Negative regulator of the H(+)-ATPase Pma1p |
| YBR256C | RIB5 | 5.9 | 1.15 × 10−3 | Riboflavin synthase |
| YOR052C | TMC1 | 5.8 | 9.56 × 10−3 | AN1-type zinc finger protein of unknown function |
| YOL155C | HPF1 | 5.8 | 6.09 × 10−5 | Haze-protective mannoprotein |
| YMR318C | ADH6 | 5.8 | 7.64 × 10−3 | NADPH-dependent medium chain alcohol dehydrogenase |
| YJL082W | IML2 | 5.8 | 4.56 × 10−4 | Protein of unknown function |
| YKL051W | SFK1 | 5.6 | 6.62 × 10−6 | Plasma membrane protein that may act to generate normal levels of PI4P |
| YER185W | PUG1 | 5.6 | 3.14 × 10−5 | Plasma membrane protein involved in protoprophyrin and heme transport |
| YIR017C | MET28 | 5.6 | 3.48 × 10−6 | Basic leucine zipper (bZIP) transcriptional activator in the Cbf1p-Met4p-Met28p complex |
| YHL024W | RIM4 | 5.5 | 4.66 × 10−6 | Putative RNA-binding protein |
| YGR243W | MPC3 | 5.4 | 7.07 × 10−5 | Highly conserved subunit of mitochondrial pyruvate carrier |
| YGL010W | MPO1 | 5.3 | 7.58 × 10−6 | Protein involved in metabolism of phytosphingosine |
| YDR513W | GRX2 | 5.1 | 6.09 × 10−3 | Cytoplasmic glutaredoxin |
| YHR179W | OYE2 | 5.1 | 1.04 × 10−2 | Conserved NADPH oxidoreductase containing flavin mononucleotide (FMN) |
| YDR059C | UBC5 | 5.1 | 2.39 × 10−4 | Ubiquitin-conjugating enzyme |
| YMR276W | DSK2 | 5.0 | 5.01 × 10−3 | Nuclear-enriched ubiquitin-like polyubiquitin-binding protein |
| Gene | Standard Name | FC * | p-Value | Description |
|---|---|---|---|---|
| YER106W | MAM1 | 60.2 | 2.77 × 10−8 | Monopolin |
| YGR225W | AMA1 | 57.4 | 9.19 × 10−10 | Activator of meiotic anaphase promoting complex (APC/C) |
| YER179W | DMC1 | 40.5 | 5.34 × 10−7 | Meiosis-specific recombinase |
| YOR298W | MUM3 | 33.5 | 9.62 × 10−4 | Protein of unknown function |
| YFL011W | HXT10 | 33.2 | 1.38 × 10−7 | Putative hexose transporter |
| YLL046C | RNP1 | 27.3 | 1.08 × 10−7 | Ribonucleoprotein |
| YER104W | RTT105 | 26.0 | 3.22 × 10−8 | Protein with a role in regulation of Ty1 transposition |
| YLR377C | FBP1 | 23.3 | 1.62 × 10−7 | Fructose-1,6-bisphosphatase |
| YDR523C | SPS1 | 22.7 | 6.27 × 10−6 | Putative protein serine/threonine kinase |
| YHR176W | FMO1 | 20.2 | 1.11 × 10−5 | Flavin-containing monooxygenase |
| YBR040W | FIG1 | 19.7 | 1.16 × 10−7 | Integral membrane protein |
| YGR059W | SPR3 | 18.6 | 4.74 × 10−5 | septin protein involved in sporulation |
| YEL039C | CYC7 | 16.9 | 6.54 × 10−7 | Cytochrome c isoform 2 |
| YMR101C | SRT1 | 16.7 | 3.73 × 10−7 | Forms the dehydrodolichyl diphosphate syntase (DDS) complex with NUS1 |
| YDR218C | SPR28 | 14.1 | 1.11 × 10−6 | Meiotic septin |
| YDR256C | CTA1 | 13.5 | 7.51 × 10−8 | Catalase A |
| YIL113W | SDP1 | 13.3 | 2.62 × 10−7 | Stress-inducible dual-specificity MAP kinase phosphatase |
| YOL123W | HRP1 | 12.9 | 1.98 × 10−6 | Subunit of cleavage factor I complex |
| YGL254W | FZF1 | 12.6 | 2.03 × 10−7 | Transcription factor involved in sulfite metabolism |
| YPL201C | YIG1 | 12.4 | 3.23 × 10−5 | Protein that interacts with glycerol 3-phosphatase |
| Q0275 | COX3 | 12.3 | 1.01 × 10−4 | Subunit III of cytochrome c oxidase (Complex IV) |
| YFL055W | AGP3 | 12.3 | 2.34 × 10−6 | Low-affinity amino acid permease |
| YDR259C | YAP6 | 11.4 | 1.88 × 10−5 | Basic leucine zipper (bZIP) transcription factor |
| YPR193C | HPA2 | 11.3 | 2.74 × 10−5 | Tetrameric histone acetyltransferase |
| YOR378W | AMF1 | 11.3 | 2.33 × 10−6 | Low affinity NH4+ transporter |
| YLL042C | ATG10 | 11.3 | 3.47 × 10−6 | Conserved E2-like conjugating enzyme |
| YIL101C | XBP1 | 11.1 | 3.43 × 10−4 | Transcriptional repressor |
| YBR018C | GAL7 | 11.0 | 2.12 × 10−5 | Galactose-1-phosphate uridyl transferase |
| YEL019C | MMS21 | 10.9 | 6.11 × 10−6 | SUMO ligase and component of the SMC5-SMC6 complex |
| YPR040W | TIP41 | 10.9 | 3.19 × 10−5 | Protein that interacts with Tap42p |
| YPL033C | SRL4 | 10.7 | 1.75 × 10−6 | Protein of unknown function |
| YLL057C | JLP1 | 10.5 | 1.82 × 10−6 | Fe(II)-dependent sulfonate/alpha-ketoglutarate dioxygenase |
| YGR142W | BTN2 | 10.3 | 2.34 × 10−5 | v-SNARE binding protein |
| YPL279C | FEX2 | 10.3 | 2.64 × 10−7 | Protein involved in fluoride export |
| YHL022C | SPO11 | 10.2 | 2.70 × 10−7 | Meiosis-specific protein |
| YKL055C | OAR1 | 10.0 | 2.10 × 10−6 | Mitochondrial 3-oxoacyl-[acyl-carrier-protein] reductase |
| YNL009W | IDP3 | 10.0 | 1.42 × 10−2 | Peroxisomal NADP-dependent isocitrate dehydrogenase |
| YOR297C | TIM18 | 9.9 | 3.75 × 10−5 | Component of the mitochondrial TIM22 complex |
| YER053C-A | - | 9.8 | 7.45 × 10−6 | Protein of unknown function |
| YPL027W | SMA1 | 9.7 | 1.50 × 10−7 | Protein of unknown function |
| YBR074W | PFF1 | 9.6 | 5.70 × 10−6 | Multi-spanning vacuolar membrane protease |
| YEL048C | TCA17 | 9.6 | 2.14 × 10−7 | Component of transport protein particle (TRAPP) complex II |
| YGR197C | SNG1 | 9.2 | 7.32 × 10−8 | Protein involved in resistance to nitrosoguanidine and 6-azauracil |
| YJR047C | ANB1 | 9.2 | 1.29 × 10−6 | Translation elongation factor eIF-5A |
| YKL093W | MBR1 | 9.0 | 3.41 × 10−5 | Protein involved in mitochondrial functions and stress response |
| YGR212W | SLI1 | 9.0 | 2.03 × 10−5 | N-acetyltransferase |
| YCL026C-A | FRM2 | 8.8 | 1.66 × 10−6 | Type II nitroreductase |
| YEL072W | RMD6 | 8.7 | 6.39 × 10−7 | Protein required for sporulation |
| YML054C | CYB2 | 8.5 | 2.74 × 10−6 | Cytochrome b2 (l-lactate cytochrome-c oxidoreductase) |
| YNL187W | SWT21 | 8.5 | 6.08 × 10−6 | Protein involved in mRNA splicing |
| YNR064C | - | 8.5 | 1.99 × 10−5 | Epoxide hydrolase |
| YBR065C | ECM2 | 8.4 | 9.49 × 10−6 | Pre-mRNA splicing factor |
| YPL171C | OYE3 | 8.4 | 6.43 × 10−6 | Conserved NADPH oxidoreductase containing flavin mononucleotide (FMN) |
| YGL212W | VAM7 | 8.4 | 1.02 × 10−4 | Vacuolar SNARE protein |
| YOR390W | FEX1 | 8.2 | 3.59 × 10−6 | Protein involved in fluoride export |
| YMR069W | NAT4 | 8.1 | 1.76 × 10−4 | N-alpha-acetyl-transferase |
| YDL020C | RPN4 | 8.0 | 3.51 × 10−7 | Transcription factor that stimulates expression of proteasome genes |
| YDR171W | HSP42 | 8.0 | 6.87 × 10−6 | Small heat shock protein (sHSP) with chaperone activity |
| YER054C | GIP2 | 7.9 | 2.59 × 10−6 | Putative regulatory subunit of protein phosphatase Glc7p |
| YPR151C | SUE1 | 7.9 | 9.84 × 10−7 | Protein required for degradation of unstable forms of cytochrome c |
| YGR131W | FHN1 | 7.7 | 1.62 × 10−6 | Protein of unknown function |
| YEL061C | CIN8 | 7.6 | 1.15 × 10−5 | Kinesin motor protein |
| YDR079W | PET100 | 7.6 | 4.29 × 10−6 | Chaperone that specifically facilitates the assembly of cytochrome c oxidase |
| YKL051W | SFK1 | 7.6 | 1.38 × 10−4 | Plasma membrane protein |
| YMR017W | SPO20 | 7.5 | 1.72 × 10−3 | Meiosis-specific subunit of the t-SNARE complex |
| YDR011W | SNQ2 | 7.5 | 4.53 × 10−7 | Plasma membrane ATP-binding cassette (ABC) transporter |
| YOR152C | ATG40 | 7.4 | 4.01 × 10−5 | Autophagy receptor |
| YLR312C | ATG39 | 7.4 | 2.53 × 10−7 | Autophagy receptor |
| YBL078C | ATG8 | 7.3 | 7.40 × 10−7 | Component of autophagosomes and Cvt vesicles |
| YPL186C | UIP4 | 7.2 | 4.47 × 10−4 | Protein that interacts with Ulp1p |
| YLR142W | PUT1 | 7.1 | 2.11 × 10−6 | Proline oxidase |
| YOR065W | CYT1 | 7.0 | 4.71 × 10−5 | Cytochrome c1 |
| YOL149W | DCP1 | 7.0 | 1.35 × 10−3 | Subunit of the Dcp1p-Dcp2p decapping enzyme complex |
| Q0250 | COX2 | 6.7 | 3.78 × 10−2 | Subunit II of cytochrome c oxidase (Complex IV) |
| YDR402C | DIT2 | 6.6 | 1.08 × 10−3 | N-formyltyrosine oxidase |
| YGR243W | MPC3 | 6.6 | 1.70 × 10−5 | Highly conserved subunit of the mitochondrial pyruvate carrier (MPC) |
| YOR005C | DNL4 | 6.6 | 5.57 × 10−6 | DNA ligase |
| YJR010W | MET3 | 6.6 | 9.83 × 10−7 | ATP sulfurylase |
| YLR151C | PCD1 | 6.5 | 2.79 × 10−6 | 8-oxo-dGTP diphosphatase |
| YNL158W | PGA1 | 6.3 | 4.04 × 10−4 | Essential component of GPI-mannosyltransferase II |
| YDR524C | AGE1 | 6.3 | 8.02 × 10−7 | ADP-ribosylation factor (ARF) GTPase activating protein (GAP) effector |
| YNL012W | SPO1 | 6.3 | 4.68 × 10−6 | Meiosis-specific prospore protein |
| YGL240W | DOC1 | 6.3 | 6.44 × 10−5 | Processivity factor |
| YDR076W | RAD55 | 6.3 | 1.32 × 10−4 | Protein that stimulates strand exchange |
| YOR192C | THI72 | 6.3 | 7.85 × 10−6 | Transporter of thiamine or related compound |
| YMR251W | GTO3 | 6.3 | 2.35 × 10−5 | Omega class glutathione transferase |
| YDR185C | UPS3 | 6.2 | 4.77 × 10−6 | Mitochondrial protein of unknown function |
| YNL014W | HEF3 | 6.2 | 1.32 × 10−4 | Translational elongation factor EF-3 |
| YML087C | AIM33 | 6.2 | 1.01 × 10−4 | Putative protein of unknown function |
| YNR034W | SOL1 | 6.2 | 7.19 × 10−7 | Protein with a possible role in tRNA export |
| YDR070C | FMP16 | 6.1 | 3.24 × 10−4 | Protein of unknown function |
| YJR129C | EFM3 | 6.1 | 4.06 × 10−2 | S-adenosylmethionine-dependent methyltransferase |
| Q0045 | COX1 | 6.0 | 1.76 × 10−2 | Subunit I of cytochrome c oxidase (Complex IV) |
| YNL036W | NCE103 | 5.9 | 4.88 × 10−5 | Carbonic anhydrase |
| YOR178C | GAC1 | 5.9 | 6.08 × 10−4 | Regulatory subunit for Glc7p type-1 protein phosphatase (PP1) |
| YGR088W | CTT1 | 5.9 | 8.13 × 10−5 | Cytosolic catalase T |
| YDL247W | MPH2 | 5.8 | 2.28 × 10−5 | Alpha-glucoside permease |
| YCL066W | HMLALPHA1 | 5.7 | 6.90 × 10−4 | Silenced copy of ALPHA1 at HML |
| YNL077W | APJ1 | 5.6 | 3.33 × 10−6 | Chaperone with a role in SUMO-mediated protein degradation |
| YKL095W | YJU2 | 5.6 | 1.29 × 10−3 | Essential protein required for pre-mRNA splicing |
| YJL030W | MAD2 | 5.6 | 1.64 × 10−4 | Component of the spindle-assembly checkpoint complex |
| YHL016C | DUR3 | 5.6 | 9.87 × 10−7 | Plasma membrane transporter for urea and polyamines |
| YNL188W | KAR1 | 5.6 | 1.64 × 10−4 | Protein involved in karyogamy and spindle pole body duplication |
| YGR234W | YHB1 | 5.6 | 1.02 × 10−5 | Nitric oxide oxidoreductase |
| YCR040W | MATALPHA1 | 5.5 | 6.76 × 10−4 | Transcriptional co-activator that regulates mating-type-specific genes |
| YFL016C | MDJ1 | 5.5 | 2.05 × 10−4 | Co-chaperone that stimulates HSP70 protein Ssc1p ATPase activity |
| YNL194C | - | 5.4 | 4.89 × 10−4 | Integral membrane protein |
| YDR475C | JIP4 | 5.3 | 2.01 × 10−3 | Protein of unknown function |
| YJR160C | MPH3 | 5.3 | 8.87 × 10−5 | Alpha-glucoside permease |
| YCR104W | PAU3 | 5.3 | 1.92 × 10−3 | Member of the seripauperin multigene family |
| YIL084C | SDS3 | 5.3 | 6.30 × 10−6 | Component of the Rpd3L histone deacetylase complex |
| YIL056W | VHR1 | 5.1 | 3.53 × 10−3 | Transcriptional activator |
| YAR020C | PAU7 | 5.0 | 1.56 × 10−4 | Member of the seripauperin multigene family |
| YDR227W | SIR4 | 5.0 | 1.71 × 10−5 | Silent information regulator |
| YLR376C | PSY3 | 5.0 | 6.70 × 10−6 | Component of Shu complex (aka PCSS complex) |
| Gene | Standard Name | FC * | p-Value | Description |
|---|---|---|---|---|
| YPL171C | OYE3 | 199.6 | 1.29 × 10−4 | Conserved NADPH oxidoreductase containing flavin mononucleotide (FMN) |
| YDL243C | AAD4 | 46.5 | 1.49 × 10−9 | Putative aryl-alcohol dehydrogenase |
| YFL056C | AAD6 | 41.2 | 1.16 × 10−7 | Putative aryl-alcohol dehydrogenase |
| YLL060C | GTT2 | 34.6 | 1.44 × 10−9 | Glutathione S-transferase capable of homodimerization |
| YBR008C | FLR1 | 28.0 | 1.21 × 10−7 | Plasma membrane transporter of the major facilitator superfamily |
| YML131W | - | 24.2 | 2.41 × 10−6 | Protein of unknown function |
| YOL151W | GRE2 | 21.9 | 1.17 × 10−4 | 3-methylbutanal reductase and NADPH-dependent methylglyoxal reductase |
| YCL026C-A | FRM2 | 21.5 | 1.53 × 10−8 | Type II nitroreductase |
| YMR101C | SRT1 | 21.3 | 2.64 × 10−8 | Forms the dehydrodolichyl diphosphate syntase (DDS) complex with NUS1 |
| YGR225W | AMA1 | 20.5 | 3.87 × 10−7 | Activator of meiotic anaphase promoting complex (APC/C) |
| YDL020C | RPN4 | 19.5 | 4.24 × 10−4 | Transcription factor that stimulates expression of proteasome genes |
| YDR256C | CTA1 | 18.8 | 4.53 × 10−9 | Catalase A |
| YGR197C | SNG1 | 18.7 | 5.35 × 10−9 | Protein involved in resistance to nitrosoguanidine and 6-azauracil |
| YKL051W | SFK1 | 18.7 | 6.62 × 10−8 | Plasma membrane protein that may act to generate normal levels of PI4P |
| YML116W | ATR1 | 16.3 | 9.44 × 10−6 | Multidrug efflux pump of the major facilitator superfamily |
| YGR142W | BTN2 | 15.2 | 7.12 × 10−6 | v-SNARE binding protein |
| YHR087W | RTC3 | 15.0 | 1.01 × 10−6 | Protein of unknown function involved in RNA metabolism |
| YDR406W | PDR15 | 14.2 | 3.31 × 10−6 | Plasma membrane ATP binding cassette (ABC) transporter |
| YFL057C | AAD16 | 13.7 | 1.51 × 10−5 | Putative aryl-alcohol dehydrogenase |
| YOL149W | DCP1 | 13.5 | 3.61 × 10−5 | Subunit of the Dcp1p-Dcp2p decapping enzyme complex |
| YDR171W | HSP42 | 13.5 | 1.19 × 10−3 | Small heat shock protein (sHSP) with chaperone activity |
| YIL101C | XBP1 | 12.3 | 3.01 × 10−5 | Transcriptional repressor |
| YHR139C | SPS100 | 12.3 | 1.95 × 10−7 | Protein required for spore wall maturation |
| YGR213C | RTA1 | 12.1 | 1.04 × 10−8 | Protein involved in 7-aminocholesterol resistance |
| YEL039C | CYC7 | 11.8 | 4.14 × 10−8 | Cytochrome c isoform 2 |
| YIL056W | VHR1 | 10.5 | 4.95 × 10−7 | Transcriptional activator |
| YCL026C-B | HBN1 | 10.5 | 8.33 × 10−6 | Protein of unknown function |
| YOL123W | HRP1 | 10.4 | 2.81 × 10−6 | Subunit of cleavage factor I |
| YHL036W | MUP3 | 9.5 | 6.44 × 10−7 | Low affinity methionine permease |
| YKR076W | ECM4 | 9.4 | 4.12 × 10−7 | Omega class glutathione transferase |
| YLR108C | - | 9.1 | 3.50 × 10−7 | Protein of unknown function |
| YER054C | GIP2 | 8.9 | 1.55 × 10−7 | Putative regulatory subunit of protein phosphatase Glc7p |
| YOR298W | MUM3 | 8.9 | 3.49 × 10−6 | Protein of unknown function involved in outer spore wall organization |
| YHL024W | RIM4 | 8.6 | 4.31 × 10−8 | Putative RNA-binding protein |
| YMR169C | ALD3 | 8.3 | 6.99 × 10−3 | Cytoplasmic aldehyde dehydrogenase |
| YOR028C | CIN5 | 8.2 | 2.47 × 10−7 | Basic leucine zipper (bZIP) transcription factor of the yAP-1 family |
| YGR088W | CTT1 | 8.1 | 3.34 × 10−6 | Cytosolic catalase T |
| YER103W | SSA4 | 8.0 | 1.02 × 10−5 | Heat shock protein member of the HSP70 family |
| YER185W | PUG1 | 7.5 | 1.07 × 10−5 | Plasma membrane protein involved in protoprophyrin and heme transport |
| YER053C-A | - | 7.2 | 1.56 × 10−4 | Protein of unknown function |
| YOR152C | ATG40 | 7.2 | 3.92 × 10−5 | Autophagy receptor |
| YDL204W | RTN2 | 6.7 | 1.74 × 10−6 | Reticulon protein |
| YOR065W | CYT1 | 6.6 | 4.43 × 10−6 | Cytochrome c1 |
| YJL051W | IRC8 | 6.6 | 5.68 × 10−5 | Bud tip localized protein of unknown function |
| YLR329W | REC102 | 6.5 | 4.83 × 10−6 | Protein involved in early stages of meiotic recombination |
| YKR077W | MSA2 | 6.4 | 6.97 × 10−6 | Putative transcriptional activator |
| YHR138C | - | 6.1 | 7.19 × 10−3 | Protein of unknown function |
| YPL201C | YIG1 | 6.0 | 4.46 × 10−7 | Protein that interacts with glycerol 3-phosphatase |
| YDL025C | RTK1 | 6.0 | 3.61 × 10−2 | Putative protein kinase |
| YOR178C | GAC1 | 5.9 | 9.69 × 10−4 | Regulatory subunit for Glc7p type-1 protein phosphatase (PP1) |
| YFL016C | MDJ1 | 5.8 | 4.14 × 10−5 | Co-chaperone member of the HSP40 (DnaJ) family of chaperones |
| YFL030W | AGX1 | 5.8 | 1.51 × 10−6 | Alanine:glyoxylate aminotransferase (AGT) |
| YKL086W | SRX1 | 5.8 | 5.28 × 10−5 | Sulfiredoxin |
| YOR328W | PDR10 | 5.8 | 1.96 × 10−6 | ATP-binding cassette (ABC) transporter |
| YPR151C | SUE1 | 5.6 | 1.71 × 10−7 | Protein required for degradation of unstable forms of cytochrome c |
| YLL026W | HSP104 | 5.5 | 4.85 × 10−2 | Disaggregase |
| YGR243W | MPC3 | 5.5 | 5.30 × 10−5 | Highly conserved subunit of the mitochondrial pyruvate carrier (MPC) |
| YKL093W | MBR1 | 5.5 | 2.19 × 10−5 | Protein involved in mitochondrial functions and stress response |
| YNL036W | NCE103 | 5.5 | 5.13 × 10−5 | Carbonic anhydrase |
| YNL008C | ASI3 | 5.5 | 1.69 × 10−5 | Subunit of the nuclear inner membrane Asi ubiquitin ligase complex |
| YLR343W | GAS2 | 5.5 | 4.37 × 10−6 | 1,3-beta-glucanosyltransferase |
| YGR223C | HSV2 | 5.4 | 1.69 × 10−6 | Phosphatidylinositol 3,5-bisphosphate-binding protein |
| YER060W-A | FCY22 | 5.2 | 1.17 × 10−5 | Putative purine-cytosine permease |
| YNL155W | CUZ1 | 5.2 | 1.90 × 10−3 | Protein with a role in the ubiquitin-proteasome pathway |
| YHL021C | AIM17 | 5.2 | 1.36 × 10−4 | Putative protein of unknown function |
| YHR199C | AIM46 | 5.2 | 1.08 × 10−5 | Putative protein of unknown function |
| YGR281W | YOR1 | 5.1 | 2.18 × 10−3 | Plasma membrane ATP-binding cassette (ABC) transporter |
| YGL010W | MPO1 | 5.1 | 3.53 × 10−6 | Protein involved in metabolism of phytosphingosine |
| CIT | p-value |
| Gene Ontology Group | |
| Oxidation-reduction process | 1.8 × 10−13 |
| Cell response to oxidative stress | 2.2 × 10−9 |
| Glutathione metabolic process | 1.8 × 10−6 |
| Drug transport | 1.3 × 10−5 |
| Response to reactive oxygen species | 1.3 × 10−4 |
| OTA | p-value |
| Gene Ontology Group | |
| Single organism developmental process | 2.2 × 10−8 |
| Oxidation-reduction process | 2.0 × 10−7 |
| Cell differentiation | 3.0 × 10−6 |
| Developmental process involved in reproduction | 5.4 × 10−6 |
| Sporulation | 1.6 × 10−5 |
| Cell response to oxidative stress | 5.4 × 10−3 |
| CIT + OTA | p-value |
| Gene Ontology Group | |
| Oxidation-reduction process | 1.7 × 10−7 |
| Drug transport | 1.3 × 10−5 |
| Cell response to oxidative stress | 3.1 × 10−4 |
| Spore wall assembly | 1.4 × 10−3 |
| Single organism developmental process | 4.2 × 10−3 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vanacloig-Pedros, E.; Proft, M.; Pascual-Ahuir, A. Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast. Toxins 2016, 8, 273. https://doi.org/10.3390/toxins8100273
Vanacloig-Pedros E, Proft M, Pascual-Ahuir A. Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast. Toxins. 2016; 8(10):273. https://doi.org/10.3390/toxins8100273
Chicago/Turabian StyleVanacloig-Pedros, Elena, Markus Proft, and Amparo Pascual-Ahuir. 2016. "Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast" Toxins 8, no. 10: 273. https://doi.org/10.3390/toxins8100273
APA StyleVanacloig-Pedros, E., Proft, M., & Pascual-Ahuir, A. (2016). Different Toxicity Mechanisms for Citrinin and Ochratoxin A Revealed by Transcriptomic Analysis in Yeast. Toxins, 8(10), 273. https://doi.org/10.3390/toxins8100273