The Finding of a Group IIE Phospholipase A2 Gene in a Specified Segment of Protobothrops flavoviridis Genome and Its Possible Evolutionary Relationship to Group IIA Phospholipase A2 Genes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Cloning and Sequencing of the Genome Segments Harboring IIE PLA2 Gene and Its 5' and 3' Flanking Regions of Crotalinae Snakes and of Their IIE PLA2 cDNAs
2.3. Acquisition of the Genome Segment Harboring IIA PLA2 Gene, IIE PLA2 Gene, and OTUD3 of P. flavoviridis
2.4. Expression Analysis by Semi-Quantitative RT-PCR of Crotalinae IIE PLA2 mRNA
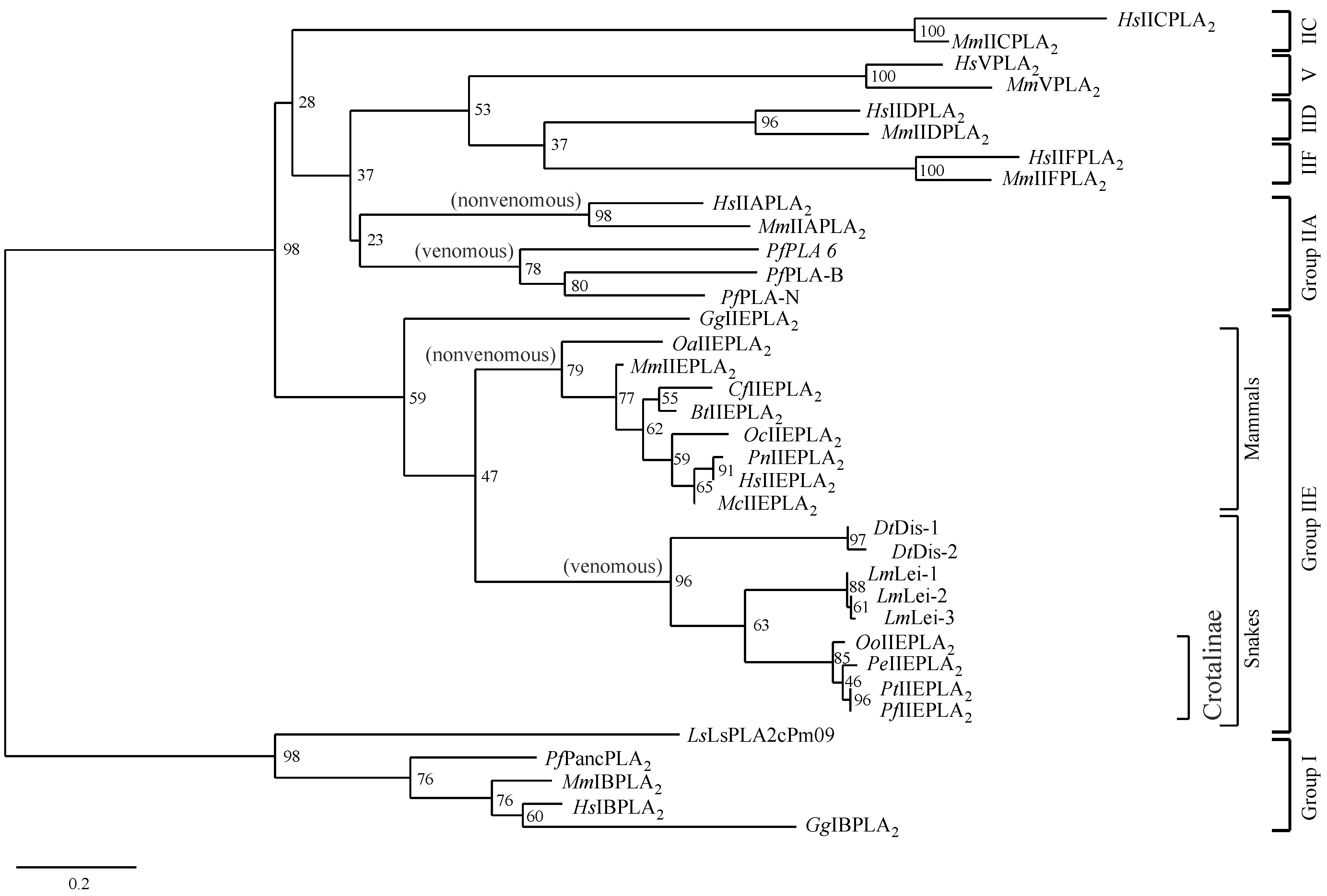
2.5. Phylogenetic Analysis of Secretory PLA2s
2.6. Comparative Structural Analysis of the Cluster Domains of Secretory PLA2 Genes in the Genomes
3. Results and Discussion
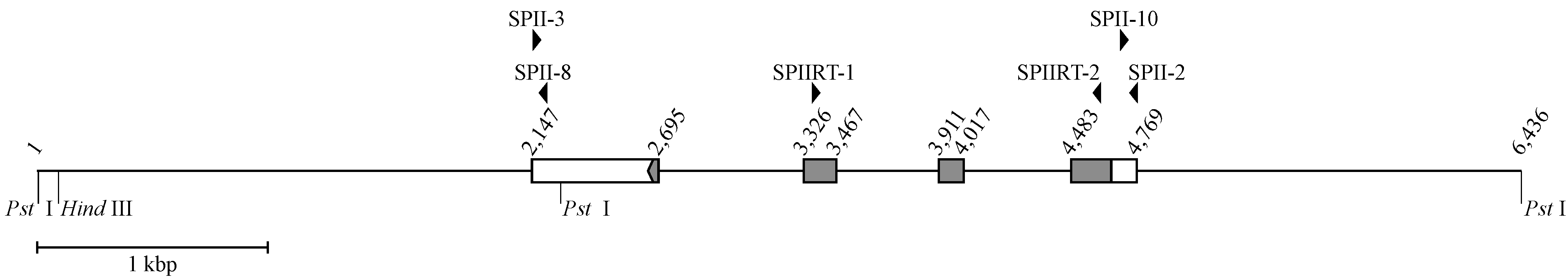
3.1. The Structure of a 6436 bp P. flavoviridis Genome Segment Containing the IIE PLA2 Gene
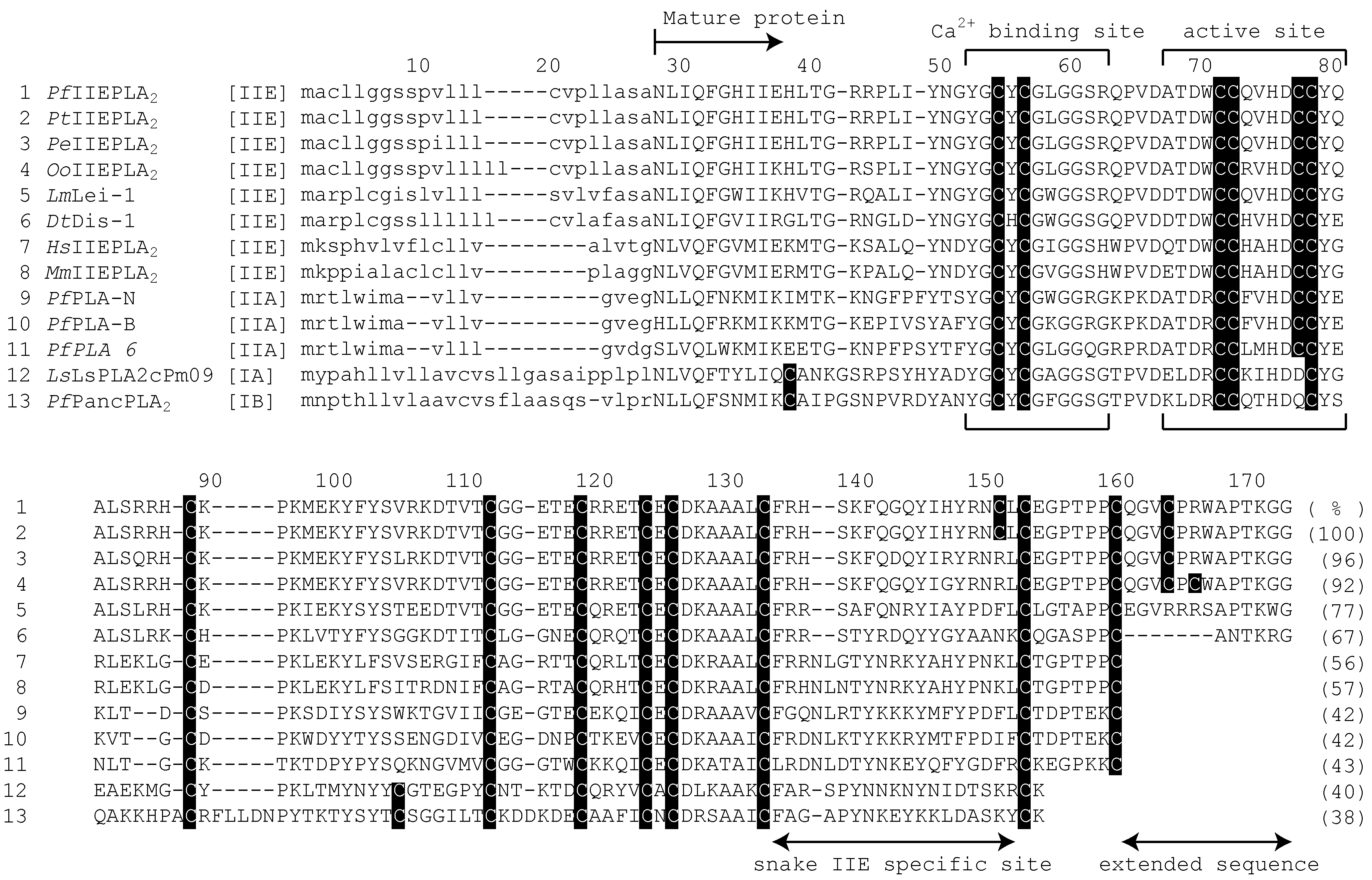
3.2. The Characteristic Primary Structures of Snake IIE PLA2 Proteins
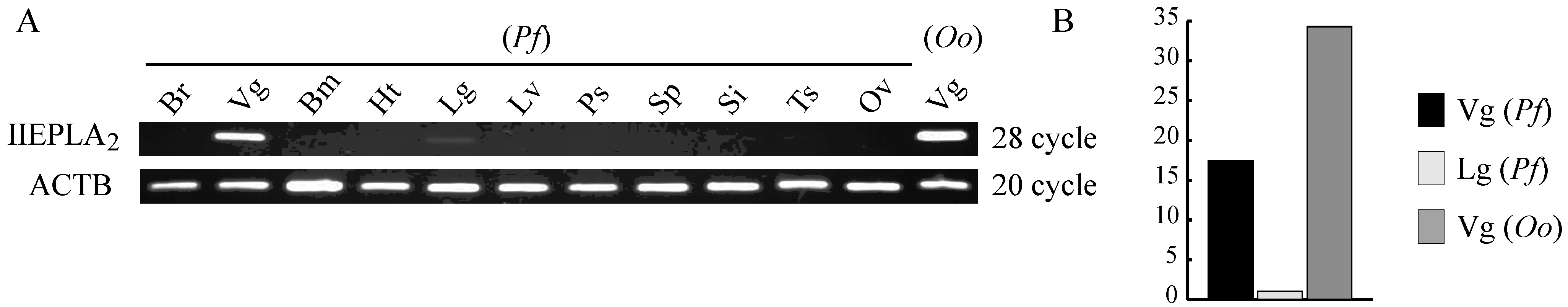
3.3. Venom Gland-Specific Expression of IIE PLA2s in Crotalinae Snakes
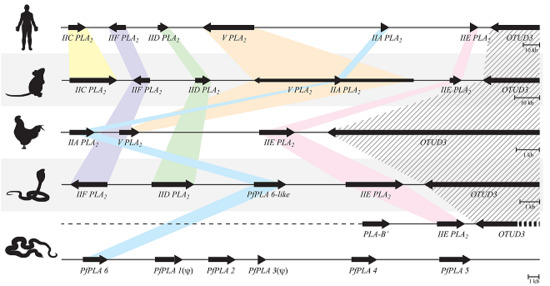
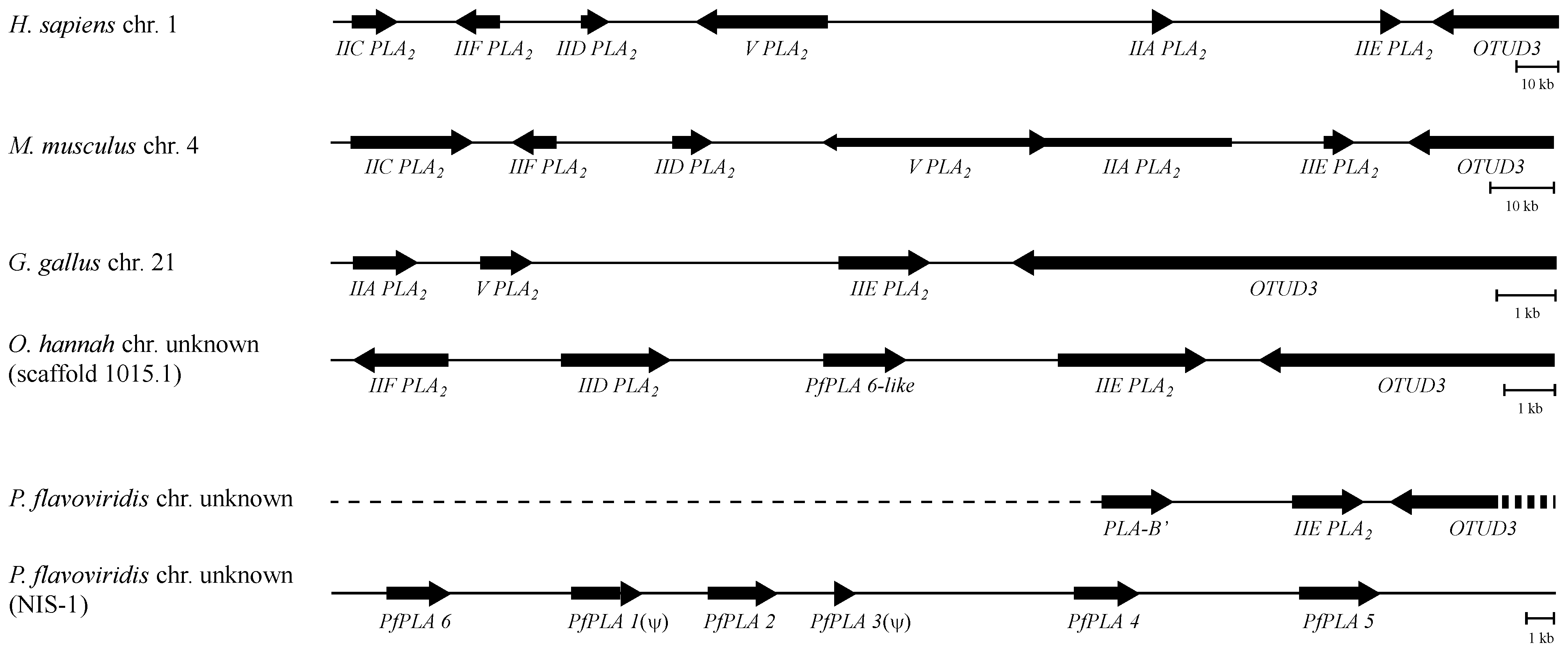
3.4. The Genome Structures Harboring Secretory PLA2 Genes Are Conserved from Human to Snake
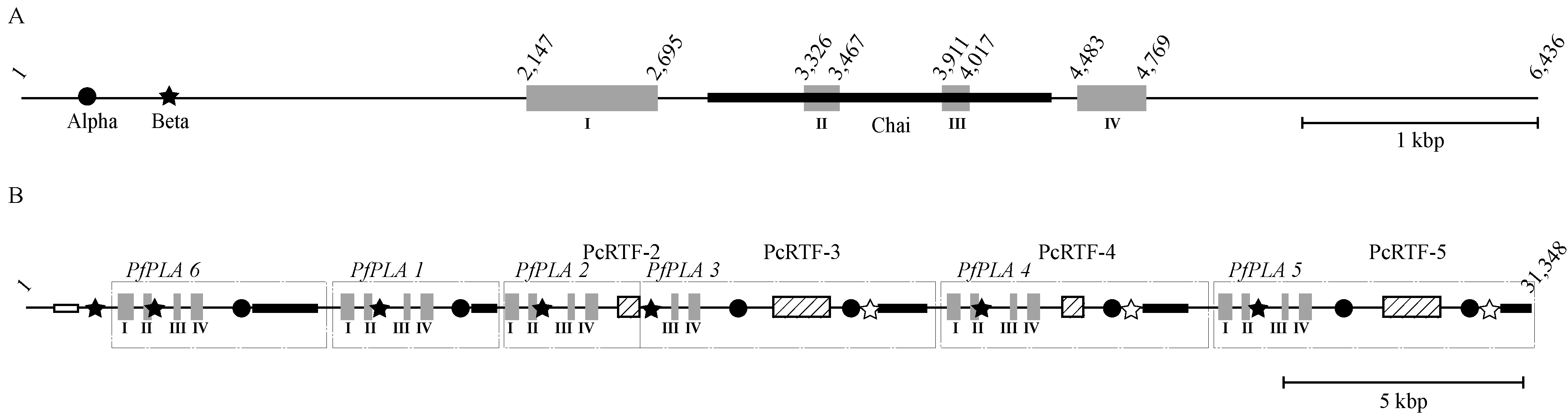
3.5. The Structural Relationship between the IIE PLA2 Gene and IIA PLA2 Genes in the P. flavoviridis Genome
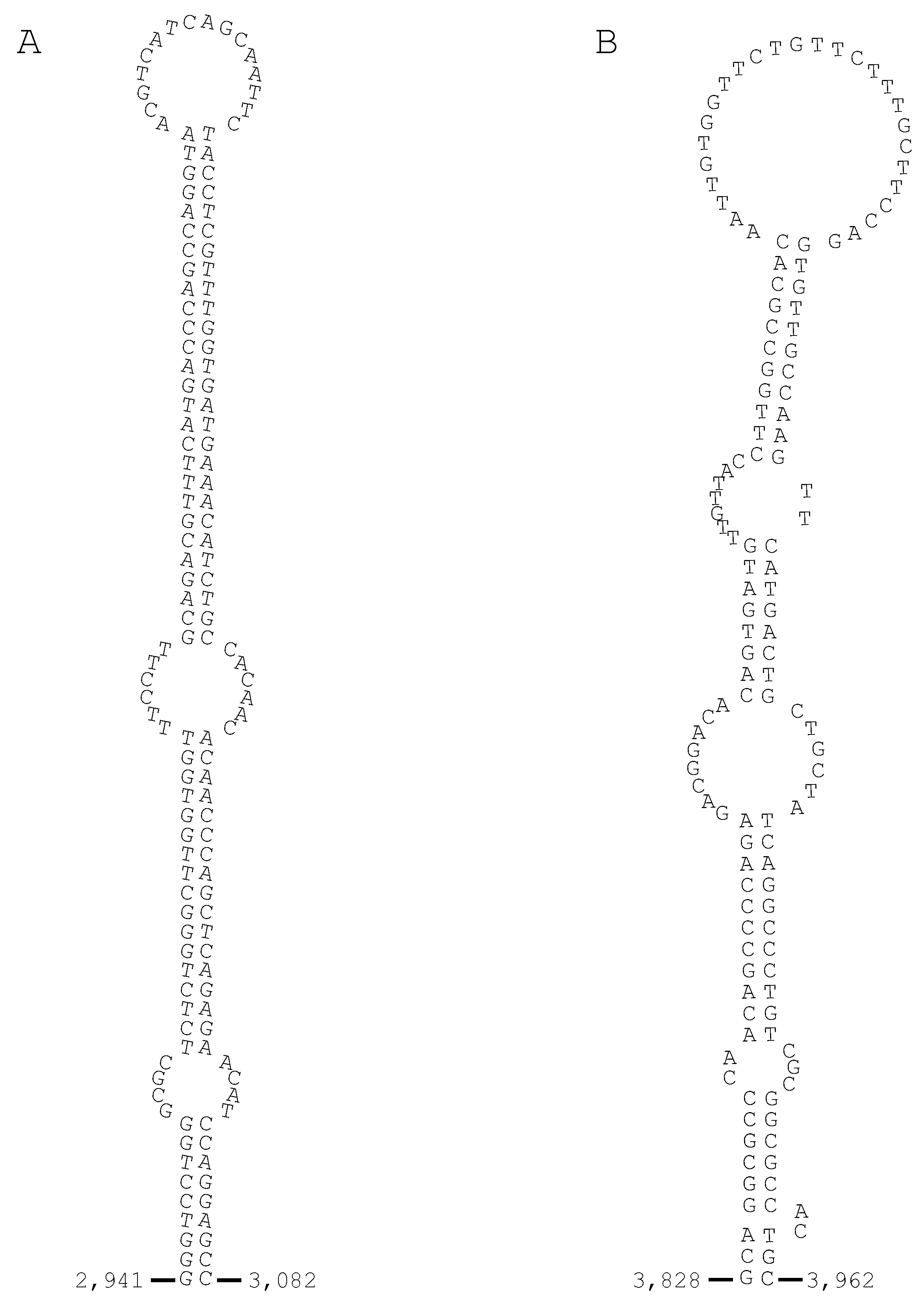

3.6. Different Multiplication Processes between Non-Venomous Secretory PLA2 Genes and P. flavoviridis Venom IIA PLA2 Genes
Acknowledgments
Author Contributions
Abbreviations
| CR 1 | chicken repeat 1 |
| EST | expressed sequence tag |
| LINE | long interspersed nuclear element |
| OTUD3 | ovarian tumor domain-containing protein 3 |
| Oo | Ovophis okinavensis |
| ORF | open reading frame |
| P | Protobothrops |
| PLA2 | phospholipase A2 |
| SINE | short interspersed nuclear element |
| UTR | untranslated region |
Conflicts of Interest
References
- Tateno, I.; Suzuki, S.; Kitamoto, O.; Chiku, N.; Sawai, Y. Anticoagulant activity of habu snake venom (Trimeresurus flavoviridis) I. Anticoagulant activity of crude habu venom. Jpn. J. Exp. Med. 1960, 30, 409–419. [Google Scholar] [PubMed]
- Tateno, I.; Suzuki, S.; Chiku, N.; Kitamoto, O. Anticoagulant activity of habu snake (Trimeresurus flavoviridis) venom. II. Effect of crude venom on blood vessels and fibrin clots. Jpn. J. Exp. Med. 1960, 30, 421–426. [Google Scholar] [PubMed]
- Mitsuhashi, S.; Maeno, H.; Sato, I.; Tanaka, T.; Kawakami, M.; Yagi, S.; Okonogi, T.; Sawai, Y.; Ono, T.; Matsushita, J. Studies on Trimeresurus venom. 1b) Comparison of the toxicological action of the venoms of Trimeresurus flavoviridis Hallowell, Trimeresurus elegans Gray and Trimeresurus okinavensis Boulenger. Nihon Saikingaku Zasshi 1961, 16, 904–908. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, B.W.; Kalk, K.H.; Hol, W.G.; Drenth, J. Structure of bovine pancreatic phospholipase A2 at 1.7 Å resolution. J. Mol. Biol. 1981, 147, 97–123. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Taketomi, Y.; Miki, T.; Sato, H.; Hirabayashi, T.; Yamamoto, K. Recent progress in phospholipase A2 research: From cells to animals to humans. Prog. Lipid Res. 2011, 50, 152–192. [Google Scholar] [CrossRef] [PubMed]
- Balsinde, J.; Winstead, M.V.; Dennis, E.A. Phospholipase A2 regulation of arachidonic acid mobilization. FEBS Lett. 2002, 531, 2–6. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Taketomi, Y.; Girard, C.; Yamamoto, K.; Lambeau, G. Emerging roles of secreted phospholipase A2 enzymes: Lessons from transgenic and knockout mice. Biochimie 2010, 92, 561–582. [Google Scholar] [CrossRef] [PubMed]
- Dufton, M.J.; Hider, R.C. Classification of phospholipases A2 according to sequence. Evolutionary and pharmacological implications. Eur. J. Biochem. 1983, 137, 545–551. [Google Scholar] [CrossRef] [PubMed]
- Kini, R.M. Venom Phospholipase A2 Enzymes: Structure, Function and Mechanism; Kini, R.M., Ed.; John Wiley & Sons: Hoboken, NJ, USA, 1997. [Google Scholar]
- Maraganore, J.M.; Merutka, G.; Cho, W.; Welches, W.; Kezdy, F.J.; Heinrikson, R.L. A new class of phospholipase A2 with lysine in place of aspartate 49. Functional consequences for calcium and substrate binding. J. Biol. Chem. 1984, 259, 13839–13843. [Google Scholar] [PubMed]
- Maraganore, J.M.; Heinrikson, R.L. The lysine-49 phopholipase A2 from the venom of Agkistrodon piscivorus piscivorus. Relation of structure and function to other phospholipase A2. J. Biol. Chem. 1986, 261, 4797–4804. [Google Scholar] [PubMed]
- Nakashima, K.; Ogawa, T.; Oda-Ueda, N.; Hattori, S.; Sakaki, Y.; Kihara, H.; Ohno, M. Accelerated evolution of Trimeresurus flavoviridis venom gland phospholipase A2 isozymes. Proc. Natl. Acad. Sci. USA 1993, 90, 5964–5968. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, N.; Chijiwa, T.; Matsubara, K.; Oda-Ueda, N.; Hattori, S.; Matsuda, Y.; Ohno, M. Unique structural characteristics and evolution of a cluster of venom phospholipase A2 isozyme genes of Protobothrops flavoviridis snake. Gene 2010, 461, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Ohno, M.; Ménez, R.; Ogawa, T.; Danse, J.M.; Shimohigashi, Y.; Fromen, C.; Ducancel, F.; Zinn-Justin, S.; le Du, M.H.; Boulain, J.C.; et al. Molecular evolution of snake toxins: Is the functional diversity of snake toxins associated with a mechanism of accelerated evolution? Prog. Nucleic Acid Res. Mol. Biol. 1998, 59, 307–364. [Google Scholar] [PubMed]
- Ohno, M.; Ogawa, T.; Oda-Ueda, N.; Chijiwa, T.; Hattori, S. Accelerated and regional evolution of snake venom gland isozymes. In Perspectives in Molecular Toxinology; John Wiley & Sons: Hoboken, NJ, USA, 2002; pp. 387–400. [Google Scholar]
- Ohno, M.; Chijiwa, T.; Oda-Ueda, N.; Ogawa, T.; Hattori, S. Molecular evolution of myotoxic phospholipases A2 from snake venom. Toxicon 2003, 42, 841–854. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Scheib, H.; de, L.M.; de Azevedo, I.J.; Silva, D.A.; Casewell, N.R. Novel transcripts in the maxillary venom glands of advanced snakes. Toxicon 2012, 59, 696–708. [Google Scholar] [CrossRef] [PubMed]
- Blin, N.; Stafford, D.W. A general method for isolation of high molecular weight DNA from eukaryotes. Nucleic Acids Res. 1976, 3, 2303–2308. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Ishizaki, J.; Yokota, Y.; Higashino, K.; Ono, T.; Ikeda, M.; Fujii, N.; Kawamoto, K.; Hanasaki, K. Structures, enzymatic properties, and expression of novel human and mouse secretory phospholipase A2s. J. Biol. Chem. 2000, 275, 5785–5793. [Google Scholar] [CrossRef] [PubMed]
- Valentin, E.; Ghomashchi, F.; Gelb, M.H.; Lazdunski, M.; Lambeau, G. On the diversity of secreted phospholipase A2. Cloning, tissue distribution, and functional expression of two novel mouse group II enzymes. J. Biol. Chem. 1999, 274, 31195–31202. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Deshimaru, M.; Nobuhisa, I.; Nakai, M.; Ogawa, T.; Oda-Ueda, N.; Nakashima, K.; Fukumaki, Y.; Shimohigashi, Y.; Hattori, S.; et al. Regional evolution of venom-gland phospholipase A2 isoenzymes of Trimeresurus flavoviridis snakes in the southwestern islands of Japan. Biochem. J. 2000, 347, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Nakasone, H.; Irie, S.; Ikeda, N.; Tomoda, K.; Oda-Ueda, N.; Hattori, S.; Ohno, M. Structural characteristics and evolution of the Protobothrops elegans pancreatic phospholipase A2 gene in contrast with those of Protobothrops genus venom phospholipase A2 genes. Biosci. Biotechnol. Biochem. 2013, 77, 97–102. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Kimura, M. The rate of molecular evolution considered from the standpoint of population genetics. Proc. Natl. Acad. Sci. USA 1969, 63, 1181–1188. [Google Scholar] [CrossRef] [PubMed]
- Vonk, F.J.; Casewell, N.R.; Henkel, C.V.; Heimberg, A.M.; Jansen, H.J.; McCleary, R.J.; Kerkkamp, H.M.; Vos, R.A.; Guerreiro, I.; Calvete, J.J.; et al. The king cobra genome reveals dynamic gene evolution and adaptation in the snake venom system. Proc. Natl. Acad. Sci. USA 2013, 110, 20651–20656. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Hamai, S.; Tsubouchi, S.; Ogawa, T.; Deshimaru, M.; Oda-Ueda, N.; Hattori, S.; Kihara, H.; Tsunasawa, S.; Ohno, M. Interisland mutation of a novel phospholipase A2 from Trimeresurus flavoviridis venom and evolution of Crotalinae group II phospholipases A2. J. Mol. Evol. 2003, 57, 546–554. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Abe, K.; Ogawa, T.; Nikandrov, N.N.; Hattori, S.; Oda-Ueda, N.; Ohno, M. Amino acid sequence of a basic aspartate-49-phospholipase A2 from Trimeresurus flavoviridis venom and phylogenetic analysis of Crotalinae venom phospholipases A2. Toxicon 2005, 46, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Ikeda, N.; Masuda, H.; Hara, H.; Oda-Ueda, N.; Hattori, S.; Ohno, M. Structural characteristics and evolution of a novel venom phospholipase A2 gene from Protobothrops flavoviridis. Biosci. Biotechnol. Biochem. 2012, 76, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Fujimi, T.J.; Tsuchiya, T.; Tamiya, T. A comparative analysis of invaded sequences from group IA phospholipase A2 genes provides evidence about the divergence period of genes groups and snaked families. Toxicon 2002, 40, 873–884. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Nakasone, H.; Ikeda, N.; Irie, S.; Oda-Ueda, N.; Hattori, S.; Ohno, M. Structural characterization and evolution of pancreatic phospholipase A2 from Crotalinae snakes. Submitted to the EMBL/GenBank/DDBJ databases, Bethesda, MD, USA, BAN08536.
- Tamiya, T.; Fujimi, T.J. Laticauda semifasciata phospholipase A2 cDNA clone LsPLA2cPm09. Submitted to the EMBL/GenBank/DDBJ databases, Bethesda, MD, USA, BAB03302.
- Zimin, A.V.; Delcher, A.L.; Florea, L.; Kelley, D.R.; Schatz, M.C.; Puiu, D.; Hanrahan, F.; Pertea, G.; van Tassell, C.P.; Sonstegard, T.S.; et al. A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol. 2009, 10, R42. [Google Scholar] [CrossRef] [PubMed]
- Karray, A.; Ben, A.Y.; Boujelben, J.; Amara, S.; Carriere, F.; Gargouri, Y.; Bezzine, S. Drastic changes in the tissue-specific expression of secreted phospholipase A2 in chicken pulmonary disease. Biochimie 2012, 94, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Karray, A.; Frikha, F.; Ben, A.Y.; Gargouri, Y.; Bezzine, S. Purification and biochemical characterization of a secreted group IIA chicken intestinal phospholipase A2. Lipids Health Dis. 2011, 10, 27. [Google Scholar] [CrossRef] [PubMed]
- Seilhamer, J.J.; Randall, T.L.; Yamanaka, M.; Johnson, L.K. Pancreatic phospholipase A2: Isolation of the human gene and cDNAs from porcine pancreas and human lung. DNA 1986, 5, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Seilhamer, J.J.; Randall, T.L.; Johnson, J.K.; Heinzmann, C.; Klisak, I.; Sparkes, R.S.; Lusis, A.J. Novel gene exon homologous to pancreatic phospholipase A2: Sequence and chromosomal mapping of both human genes. J. Cell. Biochem. 1989, 39, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Tischfield, J.A.; Xia, Y.R.; Shih, D.M.; Klisak, I.; Chen, J.; Engle, S.J.; Siakotos, A.N.; Winstead, M.V.; Seilhamer, J.J.; Allamand, V.; et al. Low-molecular-weight, calcium-dependent phospholipase A2 genes are linked and map to homologous chromosome regions in mouse and human. Genomics 1996, 32, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Ishizaki, J.; Suzuki, N.; Higashino, K.; Yokota, Y.; Ono, T.; Kawamoto, K.; Fujii, N.; Arita, H.; Hanasaki, K. Cloning and characterization of novel mouse and human secretory phospholipase A2s. J. Biol. Chem. 1999, 274, 24973–24979. [Google Scholar] [CrossRef] [PubMed]
- Valentin, E.; Singer, A.G.; Ghomashchi, F.; Lazdunski, M.; Gelb, M.H.; Lambeau, G. Cloning and recombinant expression of human group IIF-secreted phospholipase A2. Biochem. Biophys. Res. Commun. 2000, 279, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Dennis, E.A. Diversity of group types, regulation, and function of phospholipase A2. J. Biol. Chem. 1994, 269, 13057–13060. [Google Scholar] [PubMed]
- Moser, A.R.; Dove, W.F.; Roth, K.A.; Gordon, J.I. The Min (multiple intestinal neoplasia) mutation: Its effect on gut epithelial cell differentiation and interaction with a modifier system. J. Cell Biol. 1992, 116, 1517–1526. [Google Scholar] [CrossRef] [PubMed]
- Murakami, M.; Yoshihara, K.; Shimabara, S.; Lambeau, G.; Singer, A.; Gelb, MH.; Sawada, M.; Inagaki, N.; Nagai, H.; Kudo, I. Arachidonate release and eicosanoid generation by group IIE phospholipase A2. Biochem. Biophys. Res. Commun. 2002, 292, 689–696. [Google Scholar] [CrossRef] [PubMed]
- Touqui, L.; Wu, Y.Z. Interaction of secreted phospholipase A2 and pulmonary surfactant and its pathophysiological relevance in acute respiratory distress syndrome. Acta Pharmacol. Sin. 2003, 24, 1292–1296. [Google Scholar] [PubMed]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Tsai, I.H.; Wang, Y.M.; Chen, Y.H.; Tsai, T.S.; Tu, M.C. Venom phospholipase A2 of bamboo viper (Trimeresurus stejnegeri): Molecular characterization, geographic variations and evidence of multiple ancestries. Biochem. J. 2004, 377, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Chijiwa, T.; Yamaguchi, Y.; Ogawa, T.; Deshimaru, M.; Nobuhisa, I.; Nakashima, K.; Oda-Ueda, N.; Fukumaki, Y.; Hattori, S.; Ohno, M. Interisland evolution of Trimeresurus flavoviridis venom phospholipase A2 isozymes. J. Mol. Evol. 2003, 56, 286–293. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, Y.; Shimohigashi, Y.; Chijiwa, T.; Nakai, M.; Ogawa, T.; Hattori, S.; Ohno, M. Characterization, amino acid sequence and evolution of edema-inducing, basic phospholipase A2 from Trimeresurus flavoviridis venom. Toxicon 2001, 39, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Lemoine, F.J.; Degtyareva, N.P.; Lobachev, K.; Petes, T.D. Chromosomal translocations in yeast induced by low levels of DNA polymerase a model for chromosome fragile sites. Cell 2005, 120, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Koszul, R.; Fischer, G. A prominent role for segmental duplications in modeling eukaryotic genomes. C. R. Biol. 2009, 332, 254–266. [Google Scholar] [CrossRef] [PubMed]
- Castoe, T.A.; Hall, K.T.; Guibotsy Mboulas, M.L.; Gu, W.; de Koning, A.P.; Fox, S.E.; Poole, A.W.; Vemulapalli, V.; Daza, J.M.; Mockler, T.; et al. Discovery of highly divergent repeat landscapes in snake genomes using high-throughput sequencing. Genome Biol. Evol. 2011, 3, 641–653. [Google Scholar] [CrossRef] [PubMed]
- Moran, J.V.; DeBerardinis, R.J.; Kazazia, H.H., Jr. Exon shuffling by L1 retrotransposition. Science 1999, 283, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Xing, J.; Wang, H.; Belancio, V.P.; Cordaux, R.; Deininger, P.L.; Batzer, M.A. Emergence of primate genes by retrotransposon-mediated sequence transduction. Proc. Natl. Acad. Sci. USA 2006, 103, 17608–17613. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamaguchi, K.; Chijiwa, T.; Ikeda, N.; Shibata, H.; Fukumaki, Y.; Oda-Ueda, N.; Hattori, S.; Ohno, M. The Finding of a Group IIE Phospholipase A2 Gene in a Specified Segment of Protobothrops flavoviridis Genome and Its Possible Evolutionary Relationship to Group IIA Phospholipase A2 Genes. Toxins 2014, 6, 3471-3487. https://doi.org/10.3390/toxins6123471
Yamaguchi K, Chijiwa T, Ikeda N, Shibata H, Fukumaki Y, Oda-Ueda N, Hattori S, Ohno M. The Finding of a Group IIE Phospholipase A2 Gene in a Specified Segment of Protobothrops flavoviridis Genome and Its Possible Evolutionary Relationship to Group IIA Phospholipase A2 Genes. Toxins. 2014; 6(12):3471-3487. https://doi.org/10.3390/toxins6123471
Chicago/Turabian StyleYamaguchi, Kazuaki, Takahito Chijiwa, Naoki Ikeda, Hiroki Shibata, Yasuyuki Fukumaki, Naoko Oda-Ueda, Shosaku Hattori, and Motonori Ohno. 2014. "The Finding of a Group IIE Phospholipase A2 Gene in a Specified Segment of Protobothrops flavoviridis Genome and Its Possible Evolutionary Relationship to Group IIA Phospholipase A2 Genes" Toxins 6, no. 12: 3471-3487. https://doi.org/10.3390/toxins6123471
APA StyleYamaguchi, K., Chijiwa, T., Ikeda, N., Shibata, H., Fukumaki, Y., Oda-Ueda, N., Hattori, S., & Ohno, M. (2014). The Finding of a Group IIE Phospholipase A2 Gene in a Specified Segment of Protobothrops flavoviridis Genome and Its Possible Evolutionary Relationship to Group IIA Phospholipase A2 Genes. Toxins, 6(12), 3471-3487. https://doi.org/10.3390/toxins6123471