Insights into Aflatoxin B1 Toxicity in Cattle: An In Vitro Whole-Transcriptomic Approach
Abstract
1. Introduction
2. Results
2.1. Preliminary Evaluations of BFH12 Cell Responsiveness to PCB126
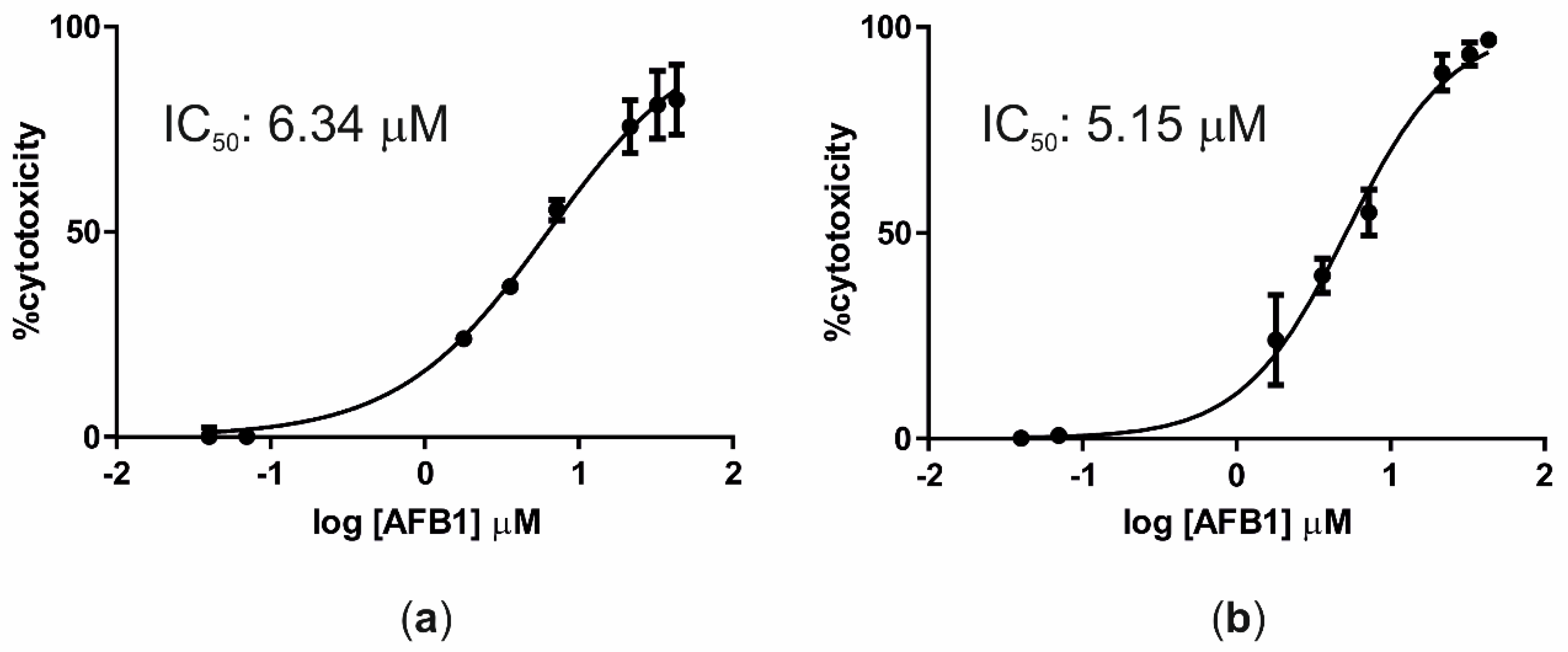
2.2. Aflatoxin B1 Cytotoxicity
2.3. Aflatoxin B1 Biotransformation in BFH12 Cells
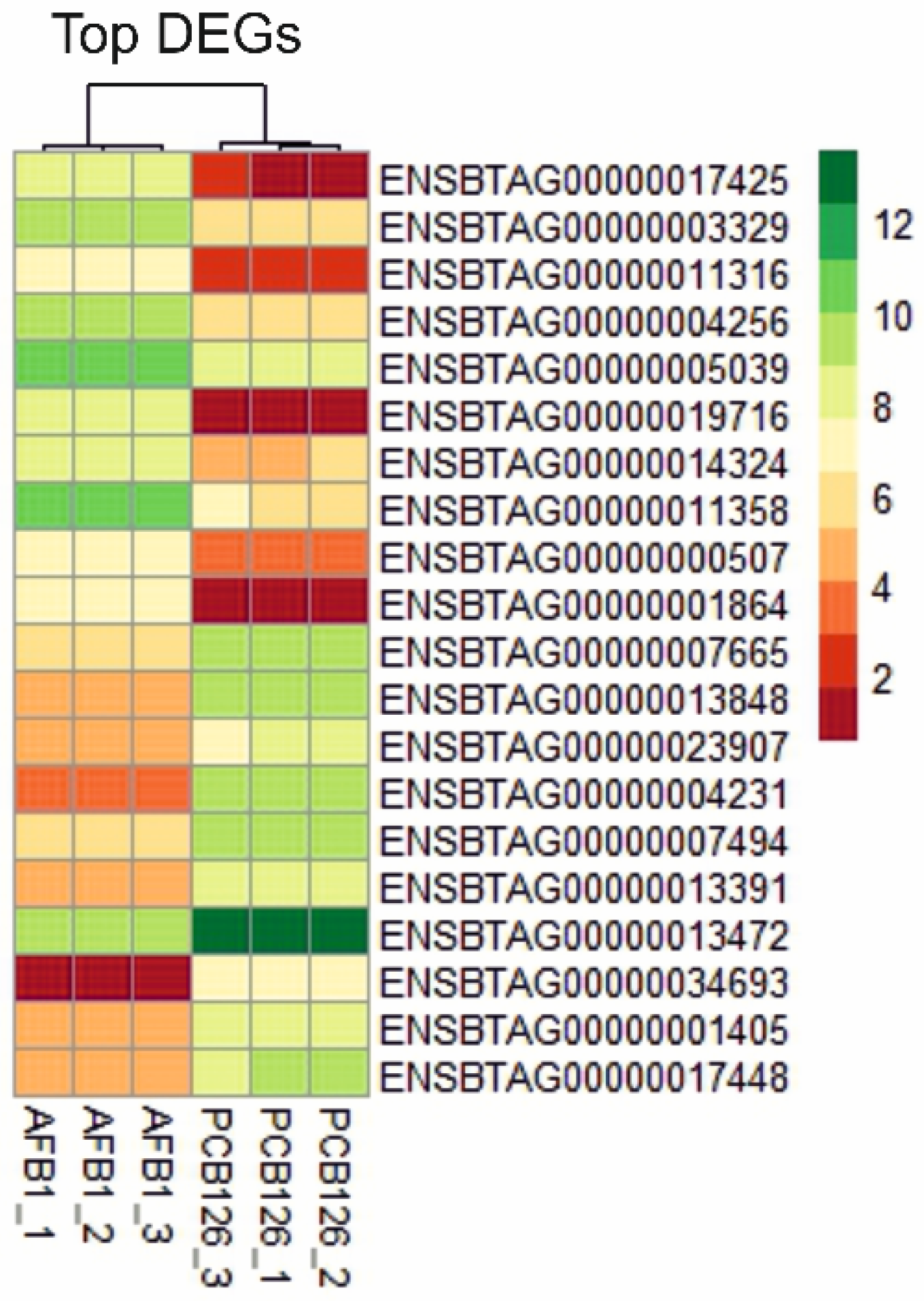
2.4. Whole-Transcriptome Differential Expression Analysis
2.5. Gene Set Enrichment Analysis
2.6. Functional Enrichment Analysis
2.7. Targeted Gene Expression (qPCR)
3. Discussion
3.1. Aflatoxin B1 Cytotoxicity
3.2. Aflatoxin B1 Biotransformation
3.3. RNA-Seq Analysis of PCB126-Exposed Cells
3.4. RNA-Seq Analysis of AFB1-Exposed Cells
3.4.1. Cancer
3.4.2. Cellular Damage—Apoptosis
3.4.3. Inflammation
3.4.4. Bioactivation, Detoxification, and Transport across Cell Membranes: CYPs and GSTs and ATP Binding Cassette (ABC) Efflux Transporters
3.4.5. Comparison with Data from Human In Vitro Models
4. Conclusions
5. Materials and Methods
5.1. Materials
5.2. Cell Culture
5.3. Preliminary Evaluations of BFH12 Responsiveness to PCB126
5.4. Aflatoxin B1 Cytotoxicity
5.5. Cell Incubation for Gene Expression Analyses
5.6. RNA-Seq Library Preparation and Sequencing
5.7. Differential Expression Analysis
5.8. Enrichment Analysis
5.9. Quantitative Real-Time PCR
5.10. Analytical Investigations
5.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Koehler, P.E.; Hanlin, R.T.; Beraha, L. Production of Aflatoxins B1 and G1 by Aspergillus flavus and Aspergillus parasiticus Isolated from Market Pecans. Appl. Microbiol. 1975, 30, 581–583. [Google Scholar] [CrossRef] [PubMed]
- IARC IARC monographs on the evaluation of carcinogenic risks to humans. Iarc Monogr. Eval. Carcinog. Risks Hum. 2010, 93, 9–38. [CrossRef]
- Wu, H.C.; Wang, Q.; Yang, H.I.; Ahsan, H.; Tsai, W.Y.; Wang, L.Y.; Chen, S.Y.; Chen, C.J.; Santella, R.M. Aflatoxin B1 exposure, hepatitis B virus infection, and hepatocellular carcinoma in Taiwan. Cancer Epidemiol. Biomark. Prev. 2009, 18, 846–853. [Google Scholar] [CrossRef] [PubMed]
- Yarru, L.P.; Settivari, R.S.; Antoniou, E.; Ledoux, D.R.; Rottinghaus, G.E. Toxicological and gene expression analysis of the impact of aflatoxin B1 on hepatic function of male broiler chicks. Poult. Sci. 2009, 88, 360–371. [Google Scholar] [CrossRef] [PubMed]
- Hussein, H.S.; Brasel, J.M. Toxicity, metabolism, and impact of mycotoxins on humans and animals. Toxicology 2001, 167, 101–134. [Google Scholar] [CrossRef]
- Monson, M.S.; Settlage, R.E.; McMahon, K.W.; Mendoza, K.M.; Rawal, S.; El-Nezami, H.S.; Coulombe, R.A.; Reed, K.M. Response of the hepatic transcriptome to aflatoxin b1in domestic turkey (Meleagris gallopavo). PLoS ONE 2014, 9, e100930. [Google Scholar] [CrossRef]
- Rustemeyer, S.M.; Lamberson, W.R.; Ledoux, D.R.; Wells, K.; Austin, K.J.; Cammack, K.M. Effects of dietary aflatoxin on the hepatic expression of apoptosis genes in growing barrows. J. Anim. Sci. 2011, 89, 916–925. [Google Scholar] [CrossRef]
- Shi, F.; Seng, X.; Tang, H.; Zhao, S.; Deng, Y.; Jin, R.; Li, Y. Effect of low levels of aflatoxin B1 on performance, serum biochemistry, hepatocyte apoptosis and liver histopathological changes of cherry valley ducks. J. Anim. Vet. Adv. 2013, 12, 1126–1130. [Google Scholar] [CrossRef]
- Rawal, S.; Kim, J.E.; Coulombe, R. Aflatoxin B1 in poultry: Toxicology, metabolism and prevention. Res. Vet. Sci. 2010, 89, 325–331. [Google Scholar] [CrossRef]
- Wu, F.; Guclu, H. Aflatoxin Regulations in a Network of Global Maize Trade. PLoS ONE 2012, 7, e45151. [Google Scholar] [CrossRef]
- Peles, F.; Sipos, P.; Győri, Z.; Pfliegler, W.P.; Giacometti, F.; Serraino, A.; Pagliuca, G.; Gazzotti, T.; Pócsi, I. Adverse Effects, Transformation and Channeling of Aflatoxins Into Food Raw Materials in Livestock. Front. Microbiol. 2019, 10, 2861. [Google Scholar] [CrossRef] [PubMed]
- Radostits, O.M.; Gay, C.C.; Hinchcliff, K.W.; Constable, P.D. A Textbook of the Disease of Cattle, Horses, Sheep, Pigs and Goats. Vet. Med. 2007, 1452–1461. [Google Scholar]
- Monson, M.; Coulombe, R.; Reed, K. Aflatoxicosis: Lessons from Toxicity and Responses to Aflatoxin B1 in Poultry. Agriculture 2015, 5, 742–777. [Google Scholar] [CrossRef]
- Santacroce, M.P.; Conversano, M.C.; Casalino, E.; Lai, O.; Zizzadoro, C.; Centoducati, G.; Crescenzo, G. Aflatoxins in aquatic species: Metabolism, toxicity and perspectives. Rev. Fish Biol. Fish. 2008, 18, 99–130. [Google Scholar] [CrossRef]
- Dohnal, V.; Wu, Q.; Kuča, K. Metabolism of aflatoxins: Key enzymes and interindividual as well as interspecies differences. Arch. Toxicol. 2014, 88, 1635–1644. [Google Scholar] [CrossRef]
- O’Brien, K.; Moss, E.; Judah, D.; Neal, G. Metabolic basis of the species difference to aflatoxin B1 induced hepatotoxicity. Biochem. Biophys. Res. Commun. 1983, 114, 813–821. [Google Scholar] [CrossRef]
- Klein, P.J.; Buckner, R.; Kelly, J.; Coulombe, R.A. Biochemical basis for the extreme sensitivity of Turkeys to aflatoxin B1. Toxicol. Appl. Pharm. 2000, 165, 45–52. [Google Scholar] [CrossRef]
- Reed, K.M.; Mendoza, K.M.; Coulombe, R.A. Altered gene response to aflatoxin b1 in the spleens of susceptible and resistant turkeys. Toxins 2019, 11, 242. [Google Scholar] [CrossRef]
- Reed, K.M.; Mendoza, K.M.; Coulombe, R.A. Differential transcriptome responses to aflatoxin B 1 in the cecal tonsil of susceptible and resistant Turkeys. Toxins 2019, 11, 55. [Google Scholar] [CrossRef]
- Kuilman, M.E.M.; Maas, R.F.M.; Judah, D.J.; Fink-Gremmels, J. Bovine Hepatic Metabolism of Aflatoxin B1. J. Agric. Food Chem. 1998, 46, 2707–2713. [Google Scholar] [CrossRef]
- Wogan, G.N.; Kensler, T.W.; Groopman, J.D. Present and future directions of translational research on aflatoxin and hepatocellular carcinoma. A review. Food Addit. Contam. Part A. 2012, 29, 249–257. [Google Scholar] [CrossRef]
- Kuilman, M.E.M.; Maas, R.F.M.; Fink-Gremmels, J. Cytochrome P450-mediated metabolism and cytotoxicity of aflatoxin B1 in bovine hepatocytes. Toxicol. In Vitro 2000, 14, 321–327. [Google Scholar] [CrossRef]
- McLean, M.; Dutton, M.F. Cellular interactions and metabolism of aflatoxin: An update. Pharm. Ther. 1995, 65, 163–192. [Google Scholar] [CrossRef]
- Rushing, B.R.; Selim, M.I. Aflatoxin B1: A review on metabolism, toxicity, occurrence in food, occupational exposure, and detoxification methods. Food Chem. Toxicol. 2019, 124, 81–100. [Google Scholar] [CrossRef]
- Marchese, S.; Sorice, A.; Ariano, A.; Florio, S.; Budillon, A.; Costantini, S.; Severino, L. Evaluation of Aflatoxin M1 Effects on the Metabolomic and Cytokinomic Profiling of a Hepatoblastoma Cell Line. Toxins 2018, 10, 436. [Google Scholar] [CrossRef]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans Chemical agents and related occupations. Iarc Monogr. Eval. Carcinog. Risks Hum. 2012, 100, 9–562.
- Jafari, T.; Fallah, A.A.; Kheiri, S.; Fadaei, A.; Amini, S.A. Aflatoxin M1 in human breast milk in Shahrekord, Iran and association with dietary factors. Food Addit. Contam. Part B Surveill. 2017, 10, 128–136. [Google Scholar] [CrossRef]
- Veldman, A.; Meijs, J.A.C.; Borggreve, G.J.; Heeres-Van Der Tol, J.J. Carry-over of aflatoxin from cows’ food to milk. Anim. Prod. 1992, 55, 163–168. [Google Scholar] [CrossRef]
- Diaz, G.J.; Sánchez, M.P. Determination of aflatoxin M1 in breast milk as a biomarker of maternal and infant exposure in Colombia. Food Addit. Contam. Part A 2015, 32, 1192–1198. [Google Scholar] [CrossRef]
- Shuib, N.S.; Makahleh, A.; Salhimi, S.M.; Saad, B. Natural occurrence of aflatoxin M1 in fresh cow milk and human milk in Penang, Malaysia. Food Control 2017, 73, 966–970. [Google Scholar] [CrossRef]
- Tahoun, A.; Ahmed, M.; Abou Elez, R.; AbdEllatif, S. Aflatoxin M1 in Milk and some Dairy Products: Level, Effect of Manufature and Public Health Concerns. Zagazig Vet. J. 2017, 45, 188–196. [Google Scholar] [CrossRef]
- Engin, A.B.; Engin, A. DNA damage checkpoint response to aflatoxin B1. Environ. Toxicol. Pharm. 2019, 65, 90–96. [Google Scholar] [CrossRef]
- Macé, K.; Aguilar, F.; Wang, J.S.; Vautravers, P.; Gómez-Lechön, M.; Gonzalez, F.J.; Groopman, J.; Harris, C.C.; Pfeifer, A.M.A. Aflatoxin B1-induced DNA adduct formation and p53 mutations in CYP450-expressing human liver cell lines. Carcinogenesis 1997, 18, 1291–1297. [Google Scholar] [CrossRef]
- Riley, J.; Mandel, H.G.; Sinha, S.; Judah, D.J.; Neal, G.E. In vitro activation of the human Harvey-ras proto-oncogene by aflatoxin B1. Carcinogenesis 1997, 18, 905–910. [Google Scholar] [CrossRef]
- Soman, N.R.; Wogan, G.N. Activation of the c-Ki-ras oncogene in aflatoxin B1-induced hepatocellular carcinoma and adenoma in the rat: Detection by denaturing gradient gel electrophoresis. Proc. Natl. Acad. Sci. USA 1993, 90, 2045–2049. [Google Scholar] [CrossRef]
- Zhang, J.; Zheng, N.; Liu, J.; Li, F.D.; Li, S.L.; Wang, J.Q. Aflatoxin B1 and aflatoxin M1 induced cytotoxicity and DNA damage in differentiated and undifferentiated Caco-2 cells. Food Chem. Toxicol. 2015, 83, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, W. Aflatoxin B1 impairs mitochondrial functions, activates ROS generation, induces apoptosis and involves Nrf2 signal pathway in primary broiler hepatocytes. Anim. Sci. J. 2016, 87, 1490–1500. [Google Scholar] [CrossRef] [PubMed]
- Weng, M.W.; Lee, H.W.; Choi, B.; Wang, H.T.; Hu, Y.; Mehta, M.; Desai, D.; Amin, S.; Zheng, Y.; Tang, M.S. AFB1 hepatocarcinogenesis is via lipid peroxidation that inhibits DNA repair, sensitizes mutation susceptibility and induces aldehyde-DNA adducts at p53 mutational hotspot codon 249. Oncotarget 2017, 8, 18213. [Google Scholar] [CrossRef]
- Eshelli, M.; Qader, M.M.; Jambi, E.J.; Hursthouse, A.S.; Rateb, M.E. Current status and future opportunities of omics tools in mycotoxin research. Toxins 2018, 10, 433. [Google Scholar] [CrossRef]
- Tryndyak, V.; Kindrat, I.; Dreval, K.; Churchwell, M.I.; Beland, F.A.; Pogribny, I.P. Effect of aflatoxin B1, benzo[a]pyrene, and methapyrilene on transcriptomic and epigenetic alterations in human liver HepaRG cells. Food Chem. Toxicol. 2018, 121, 214–223. [Google Scholar] [CrossRef]
- Jetten, M.J.A.; Kleinjans, J.C.S.; Claessen, S.M.; Chesné, C.; van Delft, J.H.M. Baseline and genotoxic compound induced gene expression profiles in HepG2 and HepaRG compared to primary human hepatocytes. Toxicol. In Vitro 2013, 27, 2031–2040. [Google Scholar] [CrossRef] [PubMed]
- Smit, E.; Souza, T.; Jennen, D.G.J.; Kleinjans, J.C.S.; van den Beucken, T. Identification of essential transcription factors for adequate DNA damage response after benzo(a)pyrene and aflatoxin B1 exposure by combining transcriptomics with functional genomics. Toxicology 2017, 390, 74–82. [Google Scholar] [CrossRef]
- Zhang, N.Y.; Qi, M.; Gao, X.; Zhao, L.; Liu, J.; Gu, C.Q.; Song, W.J.; Krumm, C.S.; Sun, L.H.; Qi, D.S. Response of the hepatic transcriptome to aflatoxin B1 in ducklings. Toxicon 2016, 111, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Reed, K.M.; Mendoza, K.M.; Abrahante, J.E.; Coulombe, R.A. Comparative response of the hepatic transcriptomes of domesticated and wild turkey to aflatoxin B1. Toxins 2018, 10, 42. [Google Scholar] [CrossRef]
- Gleich, A.; Kaiser, B.; Honscha, W.; Fuhrmann, H.; Schoeniger, A. Evaluation of the hepatocyte-derived cell line BFH12 as an in vitro model for bovine biotransformation. Cytotechnology 2019, 71, 231–244. [Google Scholar] [CrossRef]
- Gleich, A.; Kaiser, B.; Schumann, J.; Fuhrmann, H. Establishment and characterisation of a novel bovine SV40 large T-antigen-transduced foetal hepatocyte-derived cell line. Vitr. Cell. Dev. Biol.-Anim. 2016, 52, 662–672. [Google Scholar] [CrossRef]
- Bandiera, S.; Safe, S.; Okey, A.B. Binding of polychlorinated biphenyls classified as either phenobarbitone-, 3-methylcholanthrene- or mixed-type inducers to cytosolic Ah receptor. Chem. Biol. Interact. 1982, 39, 259–277. [Google Scholar] [CrossRef]
- Nebert, D.W.; Roe, A.L.; Dieter, M.Z.; Solis, W.A.; Yang, Y.; Dalton, T.P. Role of the aromatic hydrocarbon receptor and (Ah) gene battery in the oxidative stress response, cell cycle control, and apoptosis. Biochem. Pharmacol. 2000, 59, 65–85. [Google Scholar] [CrossRef]
- Wu, X.; Wu, J.; Xin, Z.; Wang, H.; Zhu, X.; Pan, L.; Li, Z.; Li, H.; Liu, Y. A 3′ UTR SNP in COL18A1 is associated with susceptibility to HBV related hepatocellular carcinoma in chinese: Three independent case-control studies. PLoS ONE 2012, 7, e33855. [Google Scholar] [CrossRef]
- Yu, Y.; Liu, D.; Liu, Z.; Li, S.; Ge, Y.; Sun, W.; Liu, B. The inhibitory effects of COL1A2 on colorectal cancer cell proliferation, migration, and invasion. J. Cancer 2018, 9, 2953–2962. [Google Scholar] [CrossRef]
- Guerrero-Martínez, J.A.; Reyes, J.C. High expression of SMARCA4 or SMARCA2 is frequently associated with an opposite prognosis in cancer. Sci. Rep. 2018, 8, 2043. [Google Scholar] [CrossRef]
- Nomoto, S.; Kanda, M.; Okamura, Y.; Nishikawa, Y.; Qiyong, L.; Fujii, T.; Sugimoto, H.; Takeda, S.; Nakao, A. Epidermal growth factor-containing fibulin-like extracellular matrix protein 1, EFEMP1, a novel tumor-suppressor gene detected in hepatocellular carcinoma using double combination array analysis. Ann. Surg. Oncol. 2010, 17, 923–932. [Google Scholar] [CrossRef]
- Prudnikova, T.Y.; Mostovich, L.A.; Domanitskaya, N.V.; Pavlova, T.V.; Kashuba, V.I.; Zabarovsky, E.R.; Grigorieva, E.V. Antiproliferative effect of D-glucuronyl C5-epimerase in human breast cancer cells. Cancer Cell Int. 2010, 10, 27. [Google Scholar] [CrossRef]
- Zancanella, V.; Giantin, M.; Dacasto, M. Absolute quantification and modulation of cytochrome P450 3A isoforms in cattle liver. Vet. J. 2014, 202, 106–111. [Google Scholar] [CrossRef]
- Fink-Gremmels, J. Mycotoxins in cattle feeds and carry-over to dairy milk: A review. Food Addit. Contam. Part A 2008, 25, 172–180. [Google Scholar] [CrossRef]
- Zhang, L.; Ye, Y.; An, Y.; Tian, Y.; Wang, Y.; Tang, H. Systems responses of rats to aflatoxin B1 exposure revealed with metabonomic changes in multiple biological matrices. J. Proteome Res. 2011, 10, 614–623. [Google Scholar] [CrossRef]
- Ratajewski, M.; Walczak-Drzewiecka, A.; Sałkowska, A.; Dastych, J. Aflatoxins upregulate CYP3A4 mRNA expression in a process that involves the PXR transcription factor. Toxicol. Lett. 2011, 205, 146–153. [Google Scholar] [CrossRef]
- Kim, J.; Park, S.-H.; Do, K.H.; Kim, D.; Moon, Y. Interference with mutagenic aflatoxin B1-induced checkpoints through antagonistic action of ochratoxin A in intestinal cancer cells: A molecular explanation on potential risk of crosstalk between carcinogens. Oncotarget 2016, 7, 39627–39639. [Google Scholar] [CrossRef]
- Jiang, J.; Pieterman, C.D.; Ertaylan, G.; Peeters, R.L.M.; de Kok, T.M.C.M. The application of omics-based human liver platforms for investigating the mechanism of drug-induced hepatotoxicity in vitro. Arch. Toxicol. 2019, 93, 3067–3098. [Google Scholar] [CrossRef]
- Aranguren-Abadía, L.; Lille-Langøy, R.; Madsen, A.K.; Karchner, S.I.; Franks, D.G.; Yadetie, F.; Hahn, M.E.; Goksøyr, A.; Karlsen, O.A. Molecular and Functional Properties of the Atlantic Cod (Gadus morhua) Aryl Hydrocarbon Receptors Ahr1a and Ahr2a. Env. Sci. Technol. 2020, 54, 1033–1044. [Google Scholar] [CrossRef]
- Liu, Y.; Du, M.; Zhang, G. Proapoptotic activity of aflatoxin B1 and sterigmatocystin in HepG2 cells. Toxicol. Rep. 2014, 1, 1076–1086. [Google Scholar] [CrossRef]
- McKean, C.; Tang, L.; Tang, M.; Billam, M.; Wang, Z.; Theodorakis, C.W.; Kendall, R.J.; Wang, J.S. Comparative acute and combinative toxicity of aflatoxin B1 and fumonisin B1 in animals and human cells. Food Chem. Toxicol. 2006, 44, 868–876. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, Y. Non-coplanar and coplanar polychlorinated biphenyls potentiate genotoxicity of aflatoxin B1 in a human hepatocyte line by enhancing CYP1A2 and CYP3A4 expression. Environ. Pollut. 2019, 246, 945–954. [Google Scholar] [CrossRef]
- Arenas-Huertero, F.; Zaragoza-Ojeda, M.; Sánchez-Alarcón, J.; Milić, M.; Šegvić Klarić, M.; Montiel-González, J.M.; Valencia-Quintana, R. Involvement of Ahr Pathway in Toxicity of Aflatoxins and Other Mycotoxins. Front. Microbiol. 2019, 10, 2347. [Google Scholar] [CrossRef]
- Ghadiri, S.; Spalenza, V.; Dellafiora, L.; Badino, P.; Barbarossa, A.; Dall’Asta, C.; Nebbia, C.; Girolami, F. Modulation of aflatoxin B1 cytotoxicity and aflatoxin M1 synthesis by natural antioxidants in a bovine mammary epithelial cell line. Toxicol. In Vitro 2019, 57, 174–183. [Google Scholar] [CrossRef]
- van Herwaarden, A.E.; Wagenaar, E.; Karnekamp, B.; Merino, G.; Jonker, J.W.; Schinkel, A.H. Breast cancer resistance protein (Bcrp1/Abcg2) reduces systemic exposure of the dietary carcinogens aflatoxin B1, IQ and Trp-P-1 but also mediates their secretion into breast milk. Carcinogenesis 2006, 27, 123–130. [Google Scholar] [CrossRef]
- Manzini, L.; Halwachs, S.; Girolami, F.; Badino, P.; Honscha, W.; Nebbia, C. Interaction of mammary bovine ABCG2 with AFB1 and its metabolites and regulation by PCB 126 in a MDCKII in vitro model. J. Vet. Pharm. 2017, 40, 591–598. [Google Scholar] [CrossRef]
- Huuskonen, P.; Myllynen, P.; Storvik, M.; Pasanen, M. The effects of aflatoxin B1 on transporters and steroid metabolizing enzymes in JEG-3 cells. Toxicol. Lett. 2013, 218, 200–206. [Google Scholar] [CrossRef]
- Nault, R.; Forgacs, A.L.; Dere, E.; Zacharewski, T.R. Comparisons of differential gene expression elicited by TCDD, PCB126, βNF, or ICZ in mouse hepatoma Hepa1c1c7 cells and C57BL/6 mouse liver. Toxicol. Lett. 2013, 223, 52–59. [Google Scholar] [CrossRef]
- Denison, M.S.; Heath-Pagliuso, S. The Ah receptor: A regulator of the biochemical and toxicological actions of structurally diverse chemicals. Bull. Environ. Contam. Toxicol. 1998, 61, 557–568. [Google Scholar] [CrossRef]
- Denison, M.S.; Soshilov, A.A.; He, G.; Degroot, D.E.; Zhao, B. Exactly the same but different: Promiscuity and diversity in the molecular mechanisms of action of the aryl hydrocarbon (dioxin) receptor. Toxicol. Sci. 2011, 124, 1–22. [Google Scholar] [CrossRef]
- Du, F.; Zhao, T.; Ji, H.C.; Luo, Y.B.; Wang, F.; Mao, G.H.; Feng, W.W.; Chen, Y.; Wu, X.Y.; Yang, L.Q. Dioxin-like (DL-) polychlorinated biphenyls induced immunotoxicity through apoptosis in mice splenocytes via the AhR mediated mitochondria dependent signaling pathways. Food Chem. Toxicol. 2019, 134, 110803. [Google Scholar] [CrossRef]
- Nebert, D.W. Aryl hydrocarbon receptor (AHR): “pioneer member” of the basic-helix/loop/helix per-Arnt-sim (bHLH/PAS) family of “sensors” of foreign and endogenous signals. Prog. Lipid Res. 2017, 67, 38–57. [Google Scholar] [CrossRef]
- Taguchi, Y.; Saeki, K.; Komano, T. Functional analysis of MRP1 cloned from bovine. FEBS Lett. 2002, 521, 211–213. [Google Scholar] [CrossRef]
- Fujise, H.; Sasawatari, S.; Annoura, T.; Ikeda, T.; Ueda, K. 3,3′,4,4′,5-Pentachlorobiphenyl inhibits drug efflux through P-glycoprotein in KB-3 cells expressing mutant human P-glycoprotein. J. Biomed. Biotechnol. 2004, 2004, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Gräns, J.; Wassmur, B.; Fernández-Santoscoy, M.; Zanette, J.; Woodin, B.R.; Karchner, S.I.; Nacci, D.E.; Champlin, D.; Jayaraman, S.; Hahn, M.E.; et al. Regulation of pregnane-X-receptor, CYP3A and P-glycoprotein genes in the PCB-resistant killifish (Fundulus heteroclitus) population from New Bedford Harbor. Aquat. Toxicol. 2015, 159, 198–207. [Google Scholar] [CrossRef] [PubMed]
- Faust, D.; Vondráček, J.; Krčmář, P.; Šmerdová, L.; Procházková, J.; Hrubá, E.; Hulinková, P.; Kaina, B.; Dietrich, C.; MacHala, M. AhR-mediated changes in global gene expression in rat liver progenitor cells. Arch. Toxicol. 2013, 87, 681–698. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q.; Wang, C.Q.; Yang, L.Y.; Gao, X.M.; Sun, H.T.; Zhang, Y.; Zhang, K.L.; Zhu, Y.; Zheng, Y.; Sheng, Y.Y.; et al. FOXQ1/NDRG1 axis exacerbates hepatocellular carcinoma initiation via enhancing crosstalk between fibroblasts and tumor cells. Cancer Lett. 2018, 417, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Rieswijk, L.; Claessen, S.M.H.; Bekers, O.; van Herwijnen, M.; Theunissen, D.H.J.; Jennen, D.G.J.; de Kok, T.M.C.M.; Kleinjans, J.C.S.; van Breda, S.G.J. Aflatoxin B1 induces persistent epigenomic effects in primary human hepatocytes associated with hepatocellular carcinoma. Toxicology 2016, 350–352, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Merrick, B.A.; Phadke, D.P.; Auerbach, S.S.; Mav, D.; Stiegelmeyer, S.M.; Shah, R.R.; Tice, R.R. RNA-Seq Profiling Reveals Novel Hepatic Gene Expression Pattern in Aflatoxin B1 Treated Rats. PLoS ONE 2013, 8, e61768. [Google Scholar] [CrossRef]
- Grusch, M.; Drucker, C.; Peter-Vörösmarty, B.; Erlach, N.; Lackner, A.; Losert, A.; Macheiner, D.; Schneider, W.J.; Hermann, M.; Groome, N.P.; et al. Deregulation of the activin/follistatin system in hepatocarcinogenesis. J. Hepatol. 2006, 45, 673–680. [Google Scholar] [CrossRef] [PubMed]
- Ooe, H.; Chen, Q.; Kon, J.; Sasaki, K.; Miyoshi, H.; Ichinohe, N.; Tanimizu, N.; Mitaka, T. Proliferation of rat small hepatocytes requires follistatin expression. J. Cell. Physiol. 2012, 227, 2363–2370. [Google Scholar] [CrossRef] [PubMed]
- Omran, G.; Abo El-Maali, N.; Ismail, M.; Ahmed, N.; Mostafa, M.; Nasser, N. Differential Hepatic Gene Expression and Antioxidant Activity in Male and Female Rats Induced by Subchronic Aflatoxicosis B1. Mansoura J. Forensic Med. Clin. Toxicol. 2019, 27, 13–28. [Google Scholar] [CrossRef]
- Tomasi, M.L.; Ryoo, M.; Skay, A.; Tomasi, I.; Giordano, P.; Mato, J.M.; Lu, S.C. Polyamine and methionine adenosyltransferase 2A crosstalk in human colon and liver cancer. Exp. Cell Res. 2013, 319, 1902–1911. [Google Scholar] [CrossRef]
- Xu, L.; Long, J.; Wang, P.; Liu, K.; Mai, L.; Guo, Y. Association between the ornithine decarboxylase G316A polymorphism and breast cancer survival. Oncol. Lett. 2015, 10, 485–491. [Google Scholar] [CrossRef]
- Maurer, G.; Tarkowski, B.; Baccarini, M. Raf kinases in cancer-roles and therapeutic opportunities. Oncogene 2011, 30, 3477–3488. [Google Scholar] [CrossRef]
- Li, J.K.; Chen, C.; Liu, J.Y.; Shi, J.Z.; Liu, S.P.; Liu, B.; Wu, D.S.; Fang, Z.Y.; Bao, Y.; Jiang, M.M.; et al. Long noncoding RNA MRCCAT1 promotes metastasis of clear cell renal cell carcinoma via inhibiting NPR3 and activating p38-MAPK signaling. Mol. Cancer 2017, 16, 111. [Google Scholar] [CrossRef]
- Lee, S.M.; Kim-Ha, J.; Choi, W.Y.; Lee, J.; Kim, D.; Lee, J.; Choi, E.; Kim, Y.J. Interplay of genetic and epigenetic alterations in hepatocellular carcinoma. Epigenomics 2016, 8, 993–1005. [Google Scholar] [CrossRef]
- Schulze, K.; Imbeaud, S.; Letouzé, E.; Alexandrov, L.B.; Calderaro, J.; Rebouissou, S.; Couchy, G.; Meiller, C.; Shinde, J.; Soysouvanh, F.; et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 2015, 47, 505–511. [Google Scholar] [CrossRef]
- Land, H.; Parada, L.F.; Weinberg, R.A. Tumorigenic conversion of primary embryo fibroblasts requires at least two cooperating oncogenes. Nature 1983, 304, 596–602. [Google Scholar] [CrossRef]
- Kranenburg, O. The KRAS oncogene: Past, present, and future. Biochim. Biophys. Acta 2005, 1756, 81–82. [Google Scholar] [CrossRef] [PubMed]
- Nesbit, C.E.; Tersak, J.M.; Prochownik, E.V. MYC oncogenes and human neoplastic disease. Oncogene 1999, 18, 3004–3016. [Google Scholar] [CrossRef] [PubMed]
- Dang, C.V. MYC on the path to cancer. Cell 2012, 149, 22–35. [Google Scholar] [CrossRef]
- Larson, P.S.; Mcmahon, G.; Wogan, G.N. Modulation of c-myc gene expression in rat livers by aflatoxin B1 exposure and age. Toxicol. Sci. 1993, 20, 316–324. [Google Scholar] [CrossRef]
- Verma, R.J. Aflatoxin Cause DNA Damage. Int. J. Hum. Genet. 2004, 4, 231–236. [Google Scholar] [CrossRef]
- Zhou, H.; Wang, J.; Ma, L.; Chen, L.; Guo, T.; Zhang, Y.; Dai, H.; Yu, Y. Oxidative DNA damage and multi-organ pathologies in male mice subchronically treated with aflatoxin B1. Ecotoxicol. Environ. Saf. 2019, 186, 109697. [Google Scholar] [CrossRef]
- Mughal, M.J.; Peng, X.; Zhou, Y.; Fang, J. Aflatoxin B1 invokes apoptosis via death receptor pathway in hepatocytes. Oncotarget 2017, 8, 8239. [Google Scholar] [CrossRef]
- Wu, B.; Mughal, M.J.; Fang, J.; Peng, X. The Protective Role of Selenium Against AFB1-Induced Liver Apoptosis by Death Receptor Pathway in Broilers. Biol. Trace Elem. Res. 2019, 191, 453–463. [Google Scholar] [CrossRef]
- Fu, P.Y.; Hu, B.; Ma, X.L.; Yang, Z.F.; Yu, M.C.; Sun, H.X.; Huang, A.; Zhang, X.; Wang, J.; Hu, Z.Q.; et al. New insight into BIRC3: A novel prognostic indicator and a potential therapeutic target for liver cancer. J. Cell. Biochem. 2019, 120, 6035–6045. [Google Scholar] [CrossRef]
- Hinton, D.M.; Myers, M.J.; Raybourne, R.A.; Francke-Carroll, S.; Sotomayor, R.E.; Shaddock, J.; Warbritton, A.; Chou, M.W. Immunotoxicity of aflatoxin B1 in rats: Effects on lymphocytes and the inflammatory response in a chronic intermittent dosing study. Toxicol. Sci. 2003, 73, 362–377. [Google Scholar] [CrossRef]
- Luedde, T.; Schwabe, R.F. NF-κB in the liver-linking injury, fibrosis and hepatocellular carcinoma. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.; Li, Y.; Fan, Y.; Zhao, L.; Wei, H.; Ji, C.; Zhang, J. Molecular mechanisms of lipoic acid protection against aflatoxin b1-induced liver oxidative damage and inflammatory responses in broilers. Toxins 2015, 7, 5435–5447. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Li, H.; Zhou, X.; Peng, C.; Tan, H.; Wang, M. Effect of superoxide and inflammatory factor on aflatoxin B1 triggered hepatocellular carcinoma. Am. J. Transl. Res. 2016, 8, 4003–4008. [Google Scholar] [PubMed]
- Prieto, J. Inflammation, HCC and sex: IL-6 in the centre of the triangle. J. Hepatol. 2008, 48, 380–381. [Google Scholar] [CrossRef] [PubMed]
- Hsia, C.Y.; Huo, T.I.; Chiang, S.Y.; Lu, M.F.; Sun, C.L.; Wu, J.C.; Lee, P.C.; Chi, C.W.; Lui, W.Y.; Lee, S.D. Evaluation of interleukin-6, interleukin-10 and human hepatocyte growth factor as tumor markers for hepatocellular carcinoma. Eur. J. Surg. Oncol. 2007, 33, 208–212. [Google Scholar] [CrossRef]
- Jiang, R.; Deng, L.; Zhao, L.; Li, X.; Zhang, F.; Xia, Y.; Gao, Y.; Wang, X.; Sun, B. miR-22 promotes HBV-related hepatocellular carcinoma development in males. Clin. Cancer Res. 2011, 17, 5593–5603. [Google Scholar] [CrossRef]
- Karin, M.; Greten, F.R. NF-κB: Linking inflammation and immunity to cancer development and progression. Nat. Rev. Immunol. 2005, 5, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Feng, Y.; Lin, H.; Ma, R.; Cai, X. Role of estrogen in hepatocellular carcinoma: Is inflammation the key? J. Transl. Med. 2014, 12, 93. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Zhao, L.; Zhang, N.Y.; Karrow, N.A.; Krumm, C.S.; Qi, D.S.; Sun, L.H. Aflatoxin B 1 metabolism: Regulation by phase I and II metabolizing enzymes and chemoprotective agents. Mutat. Res./Rev. Mutat. Res. 2018, 778, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Ayed-Boussema, I.; Pascussi, J.-M.; Maurel, P.; Bacha, H.; Hassen, W. Effect of Aflatoxin B1 on Nuclear Receptors PXR, CAR, and AhR and Their Target Cytochromes P450 mRNA Expression in Primary Cultures of Human Hepatocytes. Int. J. Toxicol. 2012, 31, 86–93. [Google Scholar] [CrossRef]
- Chen, X.; Horn, N.; Applegate, T.J. Efficiency of hydrated sodium calcium aluminosilicate to ameliorate the adverse effects of graded levels of aflatoxin B1 in broiler chicks. Poult. Sci. 2014, 93, 2037–2047. [Google Scholar] [CrossRef] [PubMed]
- Guerre, P.; Pineau, T.; Costet, P.; Burgat, V.; Galtier, P. Effects of AFB1 on CYP 1A1, 1A2 and 3A6 mRNA, and P450 expression in primary culture of rabbit hepatocytes. Toxicol. Lett. 2000, 111, 243–251. [Google Scholar] [CrossRef]
- Mary, V.S.; Valdehita, A.; Navas, J.M.; Rubinstein, H.R.; Fernández-Cruz, M.L. Effects of aflatoxin B1, fumonisin B1 and their mixture on the aryl hydrocarbon receptor and cytochrome P450 1A induction. Food Chem. Toxicol. 2015, 75, 104–111. [Google Scholar] [CrossRef]
- Go, R.E.; Hwang, K.A.; Choi, K.C. Cytochrome P450 1 family and cancers. J. Steroid Biochem. Mol. Biol. 2015, 147, 24–30. [Google Scholar] [CrossRef] [PubMed]
- Crespi, C.L.; Penman, B.W.; Steimel, D.T.; Smith, T.; Yang, C.S.; Sutter, T.R. Development of a human lymphoblastoid cell line constitutively expressing human CYP1B1 cDNA: Substrate specificity with model substrates and promutagens. Mutagenesis 1997, 12, 83–89. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bahari, A.; Mehrzad, J.; Mahmoudi, M.; Bassami, M.R.; Dehghani, H. Cytochrome P450 isoforms are differently up-regulated in aflatoxin B 1 -exposed human lymphocytes and monocytes. Immunopharmacol. Immunotoxicol. 2014, 36, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Andersson, E.R.; Sandberg, R.; Lendahl, U. Notch signaling: Simplicity in design, versatility in function. Development 2011, 138, 3593–3612. [Google Scholar] [CrossRef]
- Chiang, K.C.; Yen, C.L.; Yeh, C.N.; Hsu, J.T.; Chen, L.W.; Kuo, S.F.; Wang, S.Y.; Sun, C.C.; Kittaka, A.; Chen, T.C.; et al. Hepatocellular carcinoma cells express 25(OH)D-1α-hydroxylase and are able to convert 25(OH)D to 1α,25(OH)2D, leading to the 25(OH)D-induced growth inhibition. J. Steroid Biochem. Mol. Biol. 2015, 154, 47–52. [Google Scholar] [CrossRef]
- Storvik, M.; Huuskonen, P.; Kyllönen, T.; Lehtonen, S.; El-Nezami, H.; Auriola, S.; Pasanen, M. Aflatoxin B1–A potential endocrine disruptor–Up-regulates CYP19A1 in JEG-3 cells. Toxicol. Lett. 2011, 202, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wong, T.Y.; Chan, F.L.; Chen, S.; Leung, L.K. Assessing the effect of food mycotoxins on aromatase by using a cell-based system. Toxicol. In Vitro 2014, 28, 640–646. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Hu, B.; Shao, L.; Tian, Y.; Jin, T.; Jin, Y.; Ji, S.; Fan, X. Integrated analysis of transcriptomics and metabonomics profiles in aflatoxin B1-induced hepatotoxicity in rat. Food Chem. Toxicol. 2013, 55, 444–455. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.E.; Bunderson, B.R.; Croasdell, A.; Reed, K.M.; Coulombe, R.A. Alpha-Class Glutathione S-Transferases in Wild Turkeys (Meleagris gallopavo): Characterization and Role in Resistance to the Carcinogenic Mycotoxin Aflatoxin B1. PLoS ONE 2013, 8, e60662. [Google Scholar] [CrossRef]
- Gotovdorj, T.; Lee, E.; Lim, Y.; Cha, E.J.; Kwon, D.; Hong, E.; Kim, Y.J.; Oh, M.Y. 2,3,7,8-tetrachlorodibenzo-p-dioxin induced Cell-Specific drug transporters with acquired cisplatin resistance in cisplatin sensitive cancer cells. J. Korean Med. Sci. 2014, 29, 1188–1198. [Google Scholar] [CrossRef]
- Liao, X.; Song, G.; Xu, Z.; Bu, Y.; Chang, F.; Jia, F.; Xiao, X.; Ren, X.; Zhang, M.; Jia, Q. Oxaliplatin resistance is enhanced by saracatinib via upregulation Wnt-ABCG1 signaling in hepatocellular carcinoma. BMC Cancer 2020, 20, 31. [Google Scholar] [CrossRef]
- Kreuzer, K.; Frenzel, F.; Lampen, A.; Braeuning, A.; Böhmert, L. Transcriptomic effect marker patterns of genotoxins—A comparative study with literature data. J. Appl. Toxicol. 2020, 40, 448–457. [Google Scholar] [CrossRef] [PubMed]
- Kreuzer, K.; Böhmert, L.; Alhalabi, D.; Buhrke, T.; Lampen, A.; Braeuning, A. Identification of a transcriptomic signature of food-relevant genotoxins in human HepaRG hepatocarcinoma cells: Effect marker patterns of food genotoxins. Food Chem. Toxicol. 2020, 140, 111297. [Google Scholar] [CrossRef] [PubMed]
- Tryndyak, V.; Borowa-Mazgaj, B.; Beland, F.A.; Pogribny, I.P. Gene expression and cytosine DNA methylation alterations in induced pluripotent stem-cell-derived human hepatocytes treated with low doses of chemical carcinogens. Arch. Toxicol. 2019, 93, 3335–3344. [Google Scholar] [CrossRef]
- Caruso, M.; Mariotti, A.; Zizzadoro, C.; Zaghini, A.; Ormas, P.; Altafini, A.; Belloli, C. A clonal cell line (BME-UV1) as a possible model to study bovine mammary epithelial metabolism: Metabolism and cytotoxicity of aflatoxin B1. Toxicon 2009, 53, 400–408. [Google Scholar] [CrossRef]
- Andrews, S.; Krueger, F.; Seconds-Pichon, A.; Biggins, F.; Wingett, S. FastQC. A quality control tool for high throughput sequence data. Babraham Bioinformatics. Babraham Inst. 2015, 1, 1. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Kopylova, E.; Noé, L.; Touzet, H. SortMeRNA: Fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 2012, 28, 3211–3217. [Google Scholar] [CrossRef]
- Dobin, A.; Gingeras, T.R. Mapping RNA-seq Reads with STAR. Curr. Protoc. Bioinform. 2015, 51, 11.14.1–11.14.19. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

| AFB1 (ng/mL) | AFM1 (ng/mL) | AFL (ng/mL) | |
|---|---|---|---|
| Medium | 974.24 ± 186.09 | 45.6 ± 3.78 | 92.8 ± 10.50 |
| Cellular pellet | 1.43 ± 0.99 | 0 | 0 |
| Ensemble Gene ID | logFC | FDR | Gene Description | Gene Acronym |
|---|---|---|---|---|
| ENSBTAG00000001021 | 4.00 | 4.61 × 10−5 | Cytochrome P450 family 1 subfamily A member 1 | CYP1A1 |
| ENSBTAG00000010531 | 1.79 | 4.93 × 10−5 | Cytochrome P450, family 1, subfamily B, polypeptide 1 | CYP1B1 |
| ENSBTAG00000026527 | 4.56 | 0.00014 | NA | NA |
| ENSBTAG00000005997 | 0.97 | 0.00611 | Multidrug resistance protein 1 | ABCB1 |
| ENSBTAG00000052132 | 1.84 | 0.00618 | Forkhead box Q1 | FOXQ1 |
| ENSBTAG00000048991 | 1.18 | 0.00618 | REC114 meiotic recombination protein | REC114 |
| ENSBTAG00000000494 | 1.29 | 0.01107 | Phosphodiesterase 4D | PDE4D |
| ENSBTAG00000017466 | 1.29 | 0.01110 | Hyperpolarization activated cyclic nucleotide gated potassium channel 4 | HCN4 |
| Ensembl Gene ID | logFC | FDR | Gene Description | Gene Acronym | Pattern of Expression |
|---|---|---|---|---|---|
| ENSBTAG00000017425 | 6.38 | 1.74 × 10−17 | Gap junction protein beta 2 | GJB2 | ↑ |
| ENSBTAG00000003329 | 3.40 | 1.98 × 10−16 | Follistatin | FST | ↑ |
| ENSBTAG00000011316 | 4.09 | 7.33 × 10−16 | Zinc finger CCCH-type containing 12A | ZC3H12A | ↑ |
| ENSBTAG00000004256 | 2.91 | 1.06 × 10−15 | Ornithine decarboxylase 1 | ODC1 | ↑ |
| ENSBTAG00000005039 | 2.65 | 1.24 × 10−15 | A-Raf proto-oncogene, serine/threonine kinase | ARAF | ↑ |
| ENSBTAG00000019716 | 7.54 | 1.24 × 10−15 | C-X-C motif chemokine ligand 8 | CXCL8 | ↑ |
| ENSBTAG00000014324 | 2.97 | 1.50 × 10−15 | ANTXR cell adhesion molecule 2 | ANTXR2 | ↑ |
| ENSBTAG00000011358 | 3.74 | 1.63 × 10−15 | Immediate early response 3 | IER3 | ↑ |
| ENSBTAG00000000507 | 3.62 | 2.80 × 10−15 | Nuclear receptor subfamily 4 group A member 1 | NR4A1 | ↑ |
| ENSBTAG00000001864 | 5.47 | 3.68 × 10−15 | Nuclear receptor subfamily 4 group A member 3 | NR4A3 | ↑ |
| ENSBTAG00000007665 | −3.35 | 1.98 × 10−16 | Natriuretic peptide receptor 3 | NPR3 | ↓ |
| ENSBTAG00000013848 | −4.20 | 1.98 × 10−16 | Adhesion G protein-coupled receptor D1 | ADGRD1 | ↓ |
| ENSBTAG00000023907 | −3.52 | 1.98 × 10−16 | Collagen type XVIII alpha 1 chain | COL18A1 | ↓ |
| ENSBTAG00000004231 | −5.70 | 1.98 × 10−16 | Glycoprotein M6A | GPM6A | ↓ |
| ENSBTAG00000007494 | −3.34 | 1.98 × 10−16 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | SMARCA2 | ↓ |
| ENSBTAG00000013391 | −3.66 | 2.19 × 10−16 | ANKH inorganic pyrophosphate transport regulator | ANKH | ↓ |
| ENSBTAG00000013472 | −3.83 | 2.24 × 10−16 | Collagen type I alpha 2 chain | COL1A2 | ↓ |
| ENSBTAG00000034693 | −5.91 | 2.79 × 10−16 | Synaptotagmin 1 | SYT1 | ↓ |
| ENSBTAG00000001405 | −3.78 | 2.99 × 10−16 | Glucuronic acid epimerase | GLCE | ↓ |
| ENSBTAG00000017448 | −4.08 | 3.46 × 10−16 | EGF containing fibulin extracellular matrix protein 1 | EFEMP1 | ↓ |
| Ensembl Gene ID | logFC | FDR | Gene Description | Gene Acronym | Pattern of Expression |
|---|---|---|---|---|---|
| ENSBTAG00000012212 | 5.16 | 1.3 × 10−14 | cytochrome P450, family 26, subfamily B, polypeptide | CYP26B1 | ↑ |
| ENSBTAG00000052665 | 4.01 | 9.7 × 10−14 | cytochrome P450, subfamily IIIA, polypeptide 4 | CYP3A4 CYP3A28 | ↑ |
| ENSBTAG00000016906 | 5.43 | 4.3 × 10−13 | cytochrome P450, family 27, subfamily B, polypeptide 1 | CYP27B1 | ↑ |
| ENSBTAG00000010531 | −1.87 | 4.4 × 10−8 | cytochrome P450, family 1, subfamily B, polypeptide 1 | CYP1B1 | ↓ |
| ENSBTAG00000012972 | −1.39 | 1.3 × 10−6 | cytochrome P450, family 2, subfamily U, polypeptide 1 | CYP2U1 | ↓ |
| ENSBTAG00000014890 | −5.61 | 1.8 × 10−6 | cytochrome P450, family 19, subfamily A, polypeptide 1 | CYP19A1 | ↓ |
| ENSBTAG00000003632 | −1.91 | 3.0 × 10−6 | cytochrome P450, family 39, subfamily A, polypeptide 1 | CYP39A1 | ↓ |
| ENSBTAG00000039319 | −2.09 | 7.3 × 10−6 | cytochrome P450, family 4, subfamily F, polypeptide 2 | CYP4F2 | ↓ |
| ENSBTAG00000011976 | −5.65 | 1.0 × 10−5 | cytochrome P450, family 4, subfamily B, polypeptide 1 | CYP4B1 | ↓ |
| ENSBTAG00000053766 | −3.24 | 7.7 × 10−5 | cytochrome P450 2J2-like | CYP2J2L | ↓ |
| ENSBTAG00000003871 | −3.31 | 0.00031 | cytochrome P450 subfamily 2B | CYP2B6 | ↓ |
| ENSBTAG00000001021 | −1.28 | 0.00064 | cytochrome P450 family 1 subfamily A member 1 | CYP1A1 | ↓ |
| Ensembl Gene ID | logFC | FDR | Gene Description | Gene Acronym | Pattern of Expression |
|---|---|---|---|---|---|
| ENSBTAG00000008541 | −3.49 | 8.4 × 10−15 | microsomal glutathione S-transferase 1 | MGST1 | ↓ |
| ENSBTAG00000000170 | −2.67 | 4.9 × 10−11 | glutathione S-transferase, theta 4 | GSTT4 | ↓ |
| ENSBTAG00000017765 | −1.76 | 1.1 × 10−10 | Bos taurus glutathione S-transferase M1 | GSTM1 | ↓ |
| ENSBTAG00000006546 | −2.85 | 6.4 × 10−8 | glutathione S-transferase alpha 2 | GSTA2 | ↓ |
| ENSBTAG00000002706 | −1.33 | 2.3 × 10−6 | glutathione S-transferase zeta 1 | GSTZ1 | ↓ |
| ENSBTAG00000040298 | −1.11 | 1.0 × 10−5 | glutathione S-transferase theta 1 | GSTT1 | ↓ |
| ENSBTAG00000003989 | 1.07 | 4.6 × 10−5 | glutathione S-transferase omega 1 | GSTO1 | ↑ |
| ENSBTAG00000010265 | −0.83 | 0.00255 | microsomal glutathione S-transferase 3 | MGST3 | ↓ |
| ENSBTAG00000038540 | −1.54 | 0.00308 | glutathione S-transferase omega-1 | GSTO1 | ↓ |
| ENSBTAG00000051210 | 0.72 | 0.00626 | glutathione S-transferase pi 1 | GSTP1 | ↑ |
| ENSBTAG00000003548 | 1.13 | 0.00796 | glutathione S-transferase pi 1 | GSTP1 | ↑ |
| ENSBTAG00000037673 | −0.99 | 0.00799 | glutathione S-transferase mu 1 | GSTM1 | ↓ |
| ENSBTAG00000021779 | −0.77 | 0.01075 | microsomal glutathione S-transferase 2 | MGST2 | ↓ |
| Hallmark GS Upregulated by AFB1 | GS Size | ES | NES | NOM p-val | FDR q-val |
| MYC_targets_v1 | 196 | 0.39 | 6.20 | 0.000 | 0.000 |
| TNFα_signaling_via_NF-kB | 174 | 0.34 | 5.23 | 0.000 | 0.000 |
| Oxidative_phosphorylation | 194 | 0.32 | 5.18 | 0.000 | 0.000 |
| MYC_targets_v2 | 55 | 0.56 | 4.83 | 0.000 | 0.000 |
| E2F_targets | 195 | 0.23 | 3.79 | 0.000 | 0.000 |
| DNA_repair | 143 | 0.25 | 3.58 | 0.000 | 0.000 |
| G2M_checkpoint | 192 | 0.22 | 3.53 | 0.000 | 0.000 |
| Inflammatory_response | 121 | 0.24 | 3.04 | 0.000 | 0.000 |
| UV_response_up | 133 | 0.19 | 2.58 | 0.000 | 0.000 |
| KRAS_signaling_up | 141 | 0.18 | 2.50 | 0.000 | 0.000 |
| Unfolded_protein_response | 108 | 0.19 | 2.27 | 0.000 | 0.003 |
| Allograft_rejection | 102 | 0.17 | 1.97 | 0.010 | 0.013 |
| MTORC1_signaling | 192 | 0.12 | 1.96 | 0.002 | 0.013 |
| P53_pathway | 184 | 0.12 | 1.84 | 0.015 | 0.022 |
| Interferon gamma_response | 139 | 0.13 | 1.73 | 0.024 | 0.038 |
| IL2_STAT5_signaling | 158 | 0.11 | 1.61 | 0.030 | 0.063 |
| PI3K_AKT_mTOR_signaling | 92 | 0.14 | 1.60 | 0.035 | 0.063 |
| Apoptosis | 135 | 0.12 | 1.59 | 0.047 | 0.062 |
| Hypoxia | 176 | 0.10 | 1.58 | 0.047 | 0.064 |
| Hallmark Gene Sets Downregulated by AFB1 | Size | ES | NES | NOM p-val | FDR q-val |
| Epithelial_mesenchymal_transition | 168 | −0.25 | −3.74 | 0.000 | 0.000 |
| UV_response_dn | 128 | −0.23 | −3.06 | 0.000 | 0.000 |
| Protein_secretion | 91 | −0.25 | −2.77 | 0.000 | 0.001 |
| Apical_junction | 156 | −0.17 | −2.59 | 0.000 | 0.001 |
| Myogenesis | 141 | −0.18 | −2.59 | 0.000 | 0.001 |
| KRAS_signaling_dn | 93 | −0.20 | −2.31 | 0.000 | 0.003 |
| Coagulation | 90 | −0.19 | −2.16 | 0.002 | 0.007 |
| Apical_surface | 31 | −0.29 | −1.95 | 0.010 | 0.020 |
| Bile_acid_metabolism | 76 | −0.19 | −1.92 | 0.004 | 0.022 |
| Estrogen_response_late | 145 | −0.13 | −1.82 | 0.022 | 0.035 |
| Estrogen_response_early | 154 | −0.12 | −1.81 | 0.014 | 0.034 |
| Angiogenesis | 26 | −0.28 | −1.70 | 0.032 | 0.053 |
| Heme_metabolism | 154 | −0.11 | −1.60 | 0.042 | 0.083 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pauletto, M.; Tolosi, R.; Giantin, M.; Guerra, G.; Barbarossa, A.; Zaghini, A.; Dacasto, M. Insights into Aflatoxin B1 Toxicity in Cattle: An In Vitro Whole-Transcriptomic Approach. Toxins 2020, 12, 429. https://doi.org/10.3390/toxins12070429
Pauletto M, Tolosi R, Giantin M, Guerra G, Barbarossa A, Zaghini A, Dacasto M. Insights into Aflatoxin B1 Toxicity in Cattle: An In Vitro Whole-Transcriptomic Approach. Toxins. 2020; 12(7):429. https://doi.org/10.3390/toxins12070429
Chicago/Turabian StylePauletto, Marianna, Roberta Tolosi, Mery Giantin, Giorgia Guerra, Andrea Barbarossa, Anna Zaghini, and Mauro Dacasto. 2020. "Insights into Aflatoxin B1 Toxicity in Cattle: An In Vitro Whole-Transcriptomic Approach" Toxins 12, no. 7: 429. https://doi.org/10.3390/toxins12070429
APA StylePauletto, M., Tolosi, R., Giantin, M., Guerra, G., Barbarossa, A., Zaghini, A., & Dacasto, M. (2020). Insights into Aflatoxin B1 Toxicity in Cattle: An In Vitro Whole-Transcriptomic Approach. Toxins, 12(7), 429. https://doi.org/10.3390/toxins12070429






