Zearalenone-Induced Interaction between PXR and Sp1 Increases Binding of Sp1 to a Promoter Site of the eNOS, Decreasing Its Transcription and NO Production in BAECs
Abstract
1. Introduction
2. Results
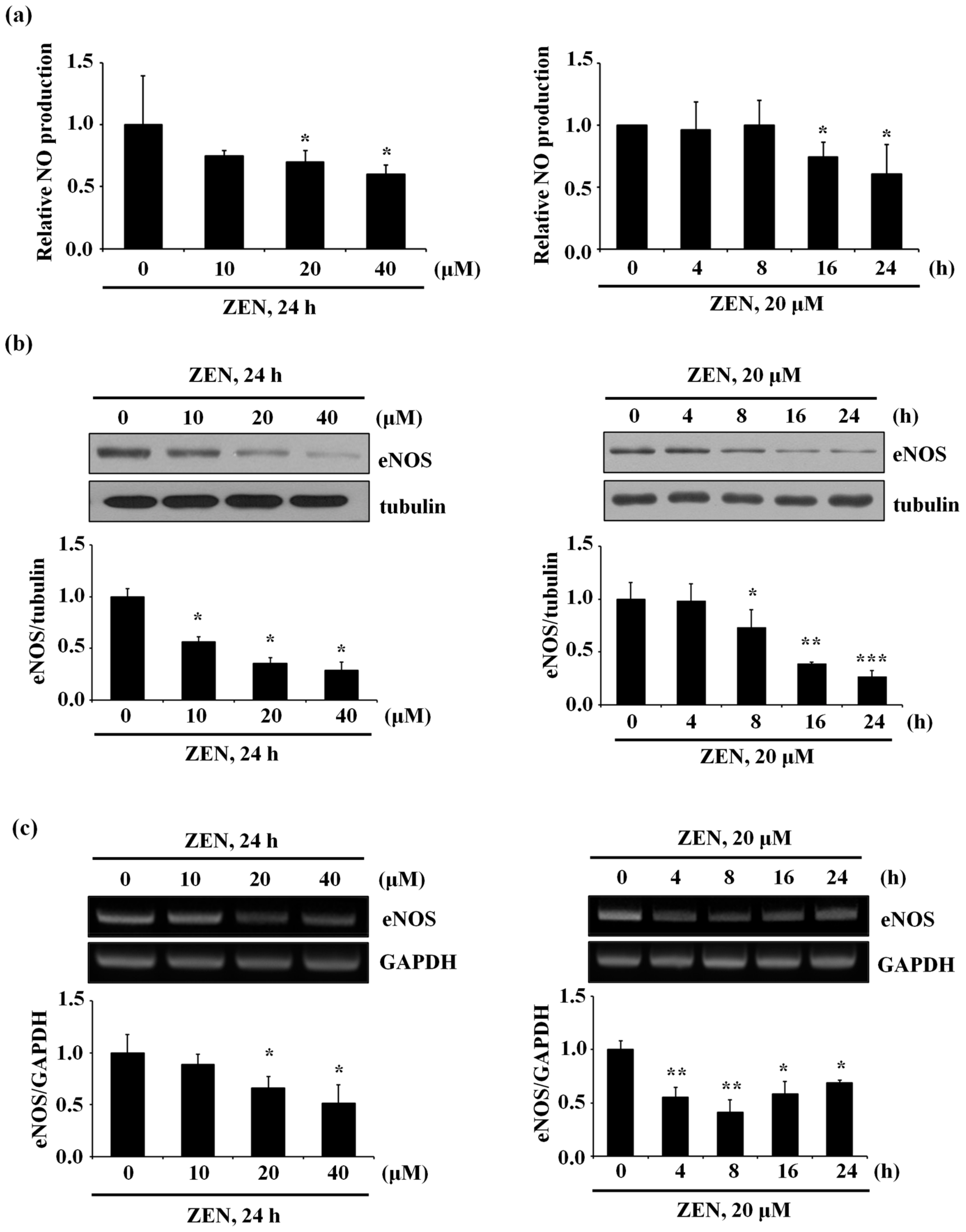
2.1. Zearalenone Decreases Nitric Oxide (NO) Production, Which is Accompanied by Decreased Protein and mRNA Expressions of eNOS
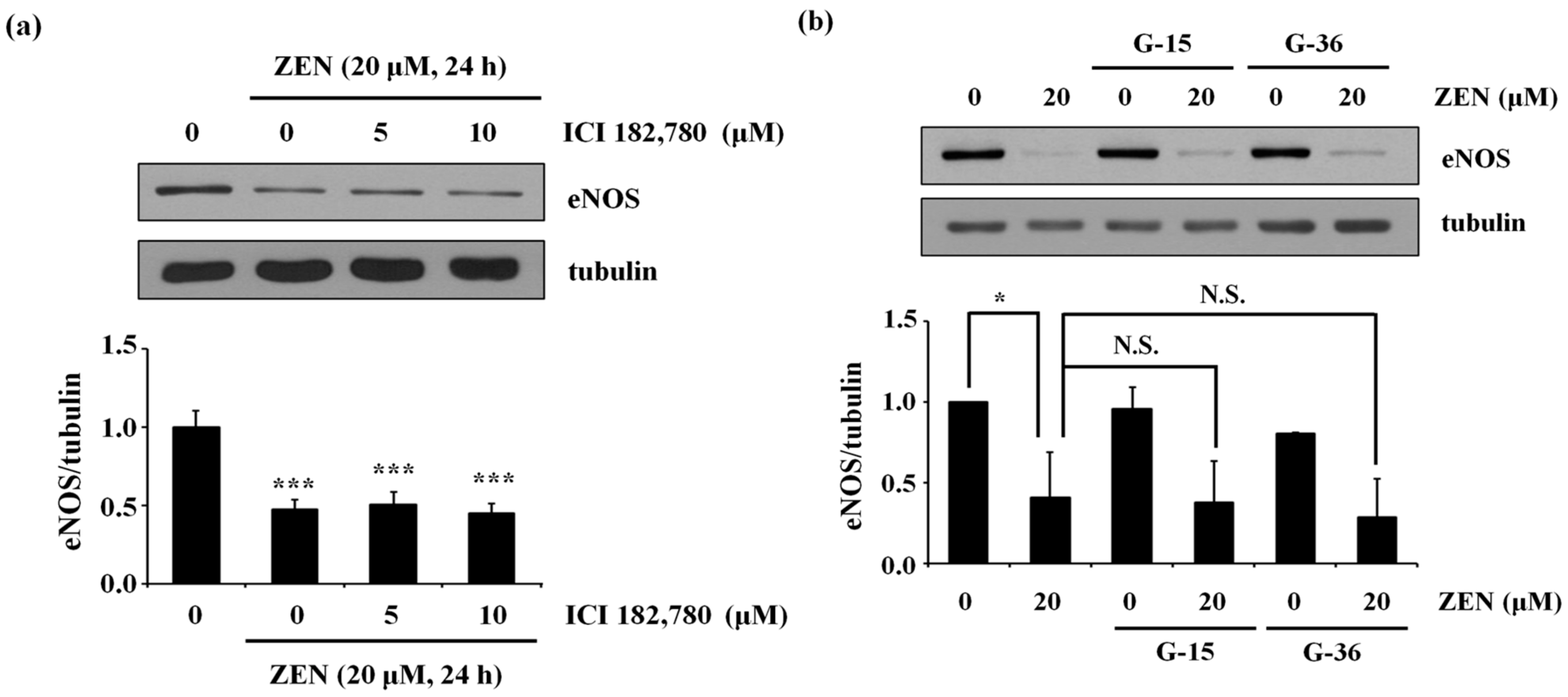
2.2. Estrogen Receptors (ERs) Are Not Involved in the ZEN-Mediated Decrease in eNOS Protein Expression
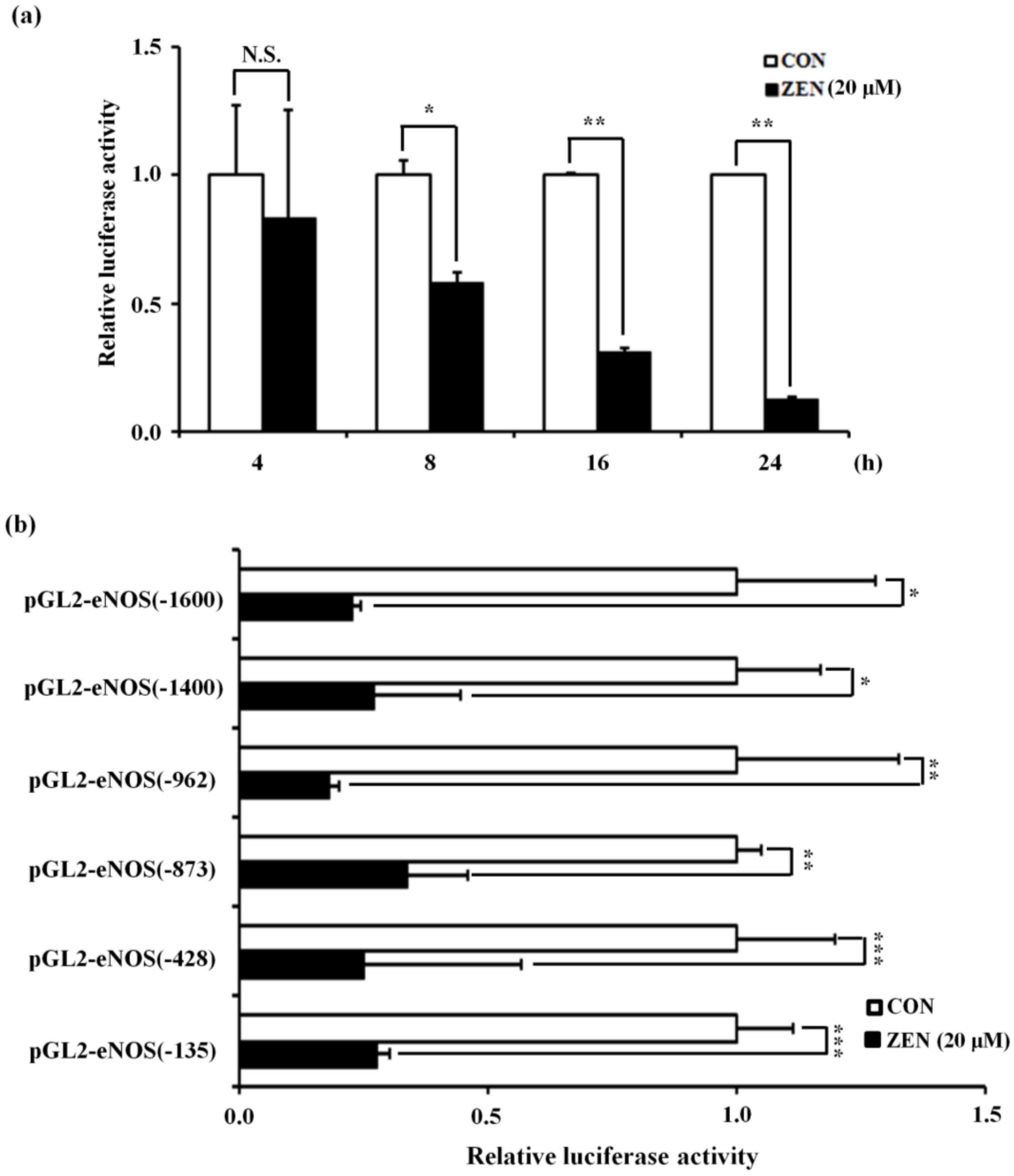
2.3. The Proximal Promoter Region (−135 to +22) of the Human eNOS Gene is Involved in the ZEN-Mediated Decrease in eNOS Gene Transcription
2.4. ZEN Decreases the Transcriptional Activation of eNOS by Increasing Sp1 Binding to −135 to +22 of the Human eNOS Promoter Region
2.5. Mithramycin A, a Selective Sp1 Inhibitor, and Sp1 siRNA Can Reverse the ZEN-Mediated Decrease in eNOS Promoter Activity, Protein Expression, and NO Production
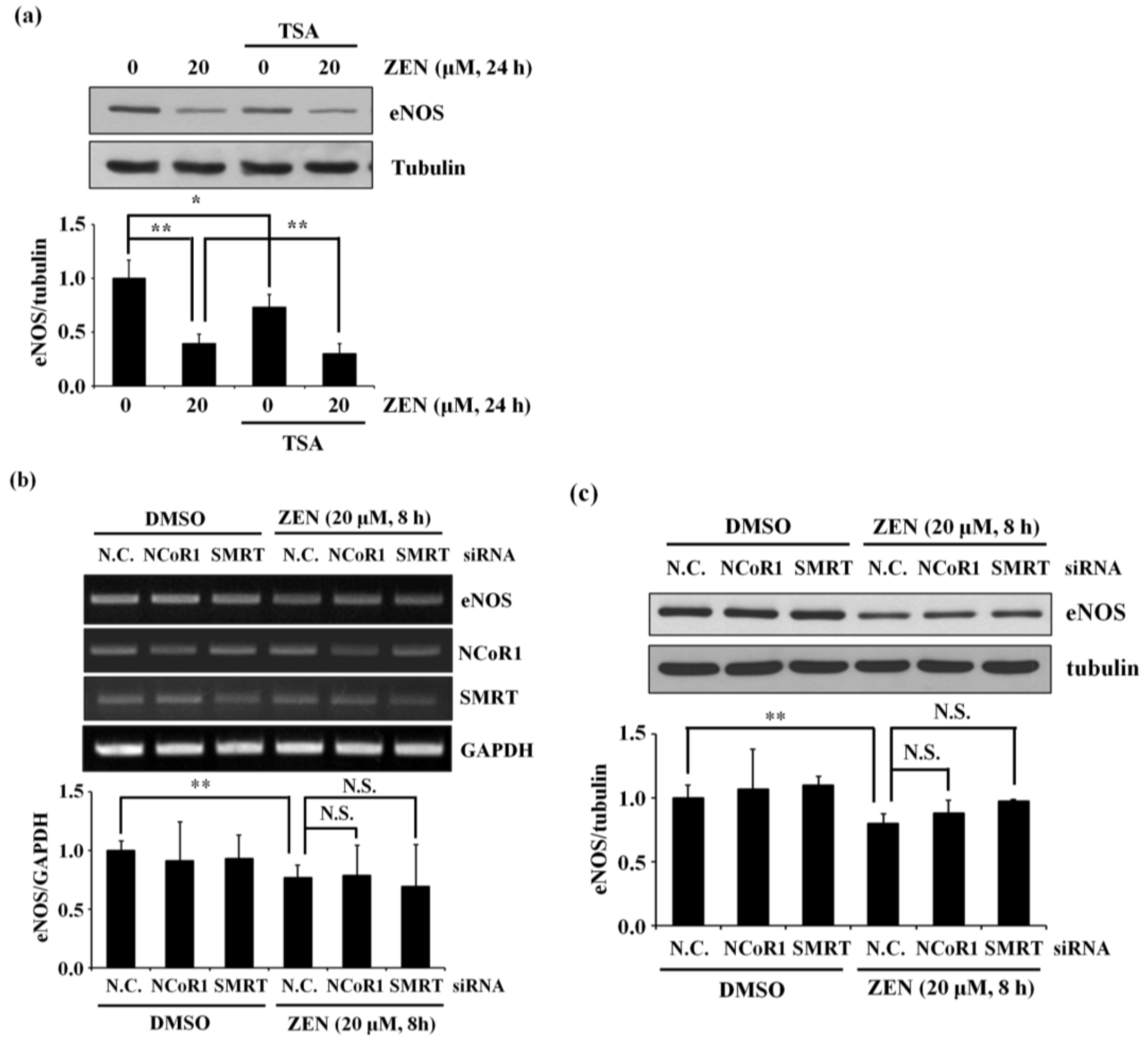
2.6. Histone Deacetylases (HDACs), NCoR1, or SMRT Do Not Reverse the ZEN-Stimulated Decrease in the Expression of eNOS Protein and mRNA
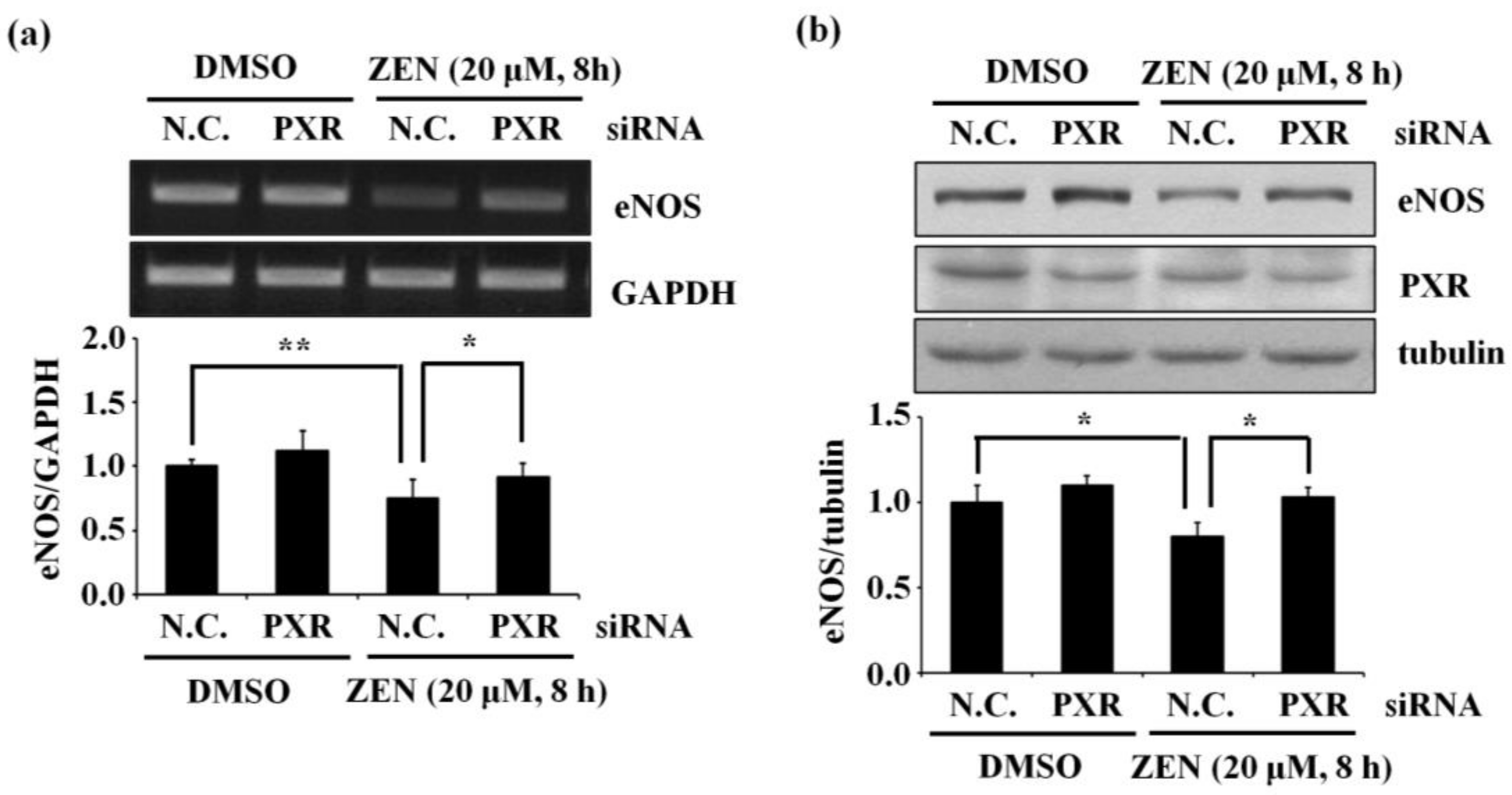
2.7. Pregnane X Receptor (PXR) Induces the ZEN-Mediated Decrease in eNOS mRNA and Protein Expression
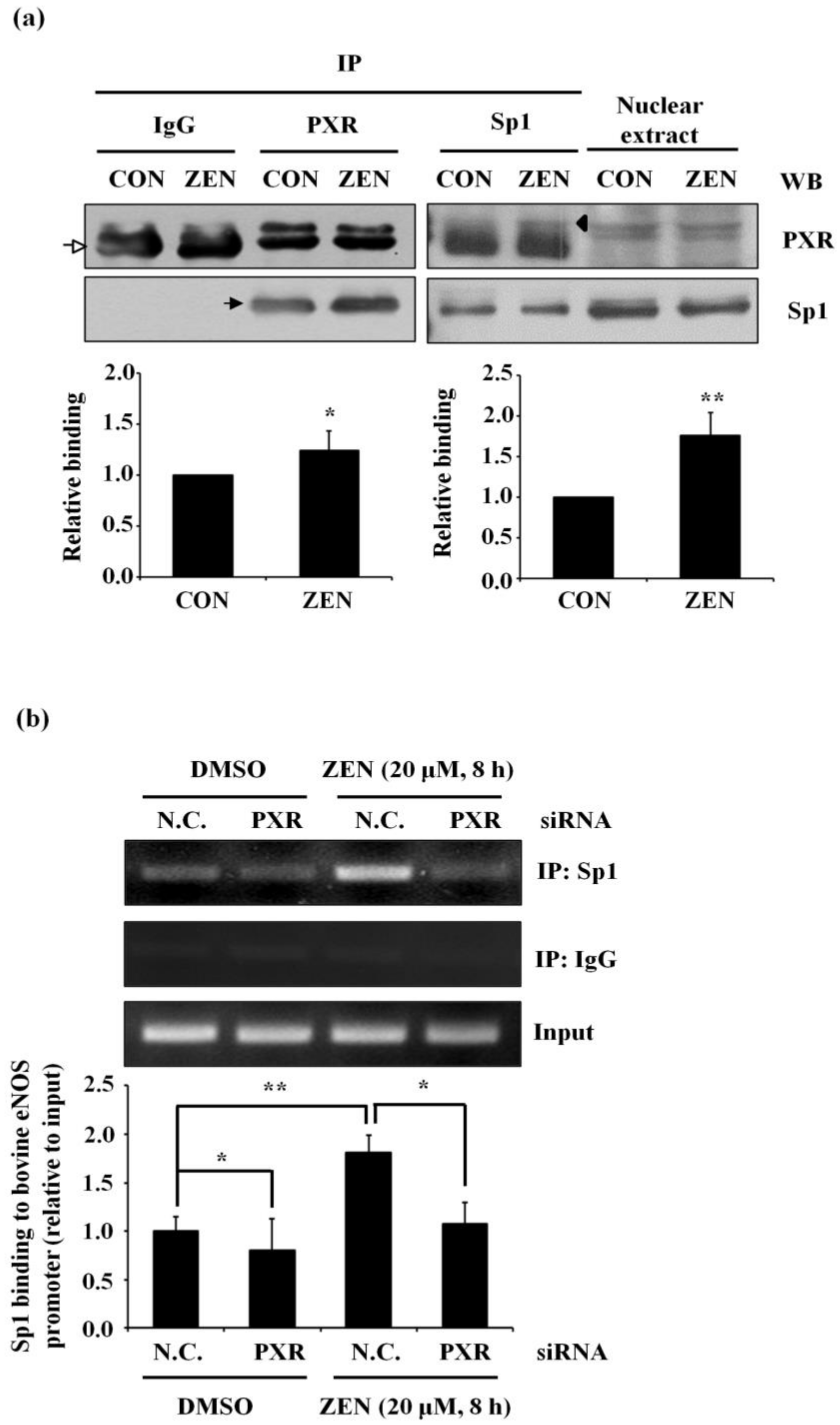
2.8. ZEN-Stimulated PXR Binding to Sp1 Is Required for Binding of Sp1 to the eNOS Promoter of BAECs
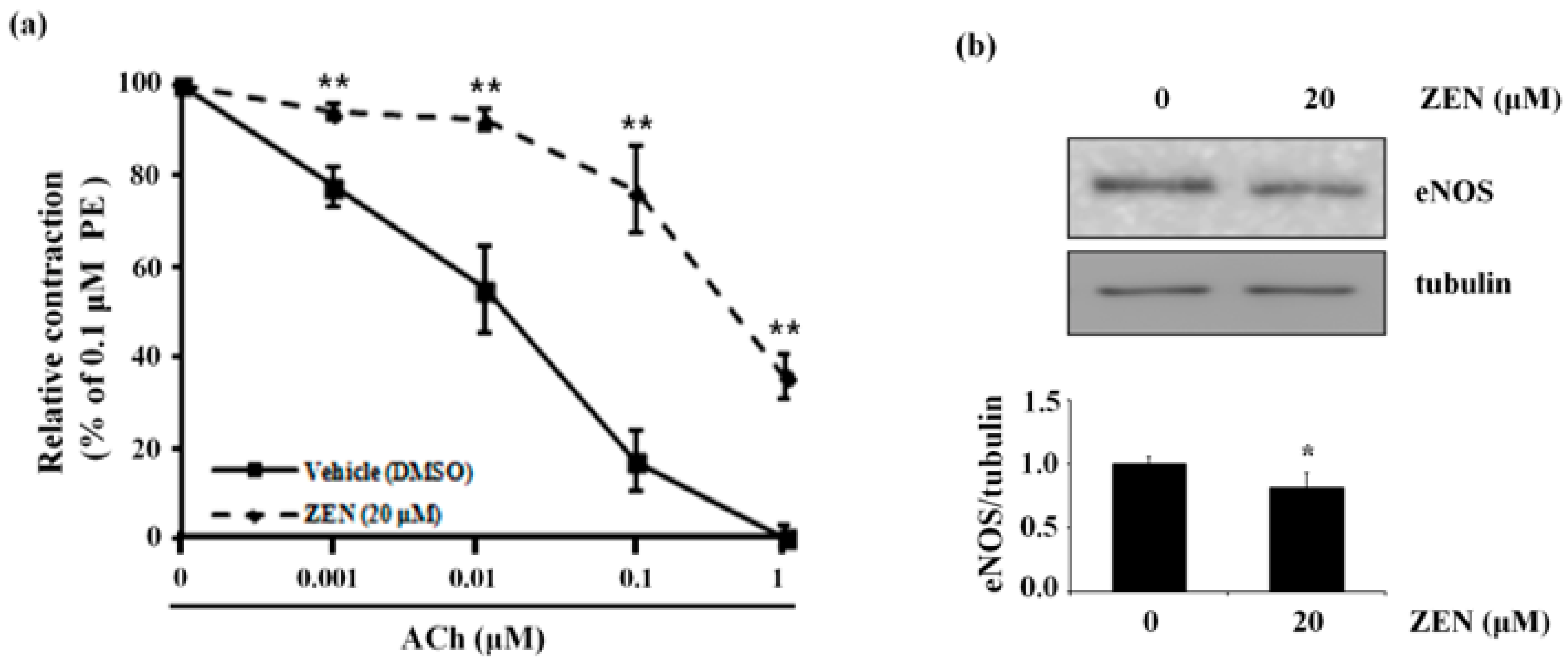
2.9. ZEN Decreases Acetylcholine (ACh)-Induced Vessel Relaxation Accompanied by Decreased eNOS Protein Expression in Mouse Aortas
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Materials
5.2. Full-Length and 5′-Deleted Human eNOS Promoter
5.3. Site-Directed Mutagenesis
5.4. Cell Culture and Drug Treatments
5.5. Transfection
5.6. Luciferase Assay
5.7. Reverse Transcription (RT)-PCR
5.8. Western Blot Analysis
5.9. Assessment of Nitric Oxide (NO) Production
5.10. In Silico Analysis
5.11. Chromatin Immunoprecipitation (ChIP) Assay
5.12. Preparation of Nuclear and Non-Nuclear Fractions
5.13. Co-Immunoprecipitation (co-IP)
5.14. Animals
5.15. Measurement of Endothelium-Dependent Vessel Relaxation
5.16. Statistical Analysis
5.17. Ethical Approval
Author Contributions
Funding
Conflicts of Interest
References
- Zinedine, A.; Soriano, J.M.; Molto, J.C.; Manes, J. Review on the toxicity, occurrence, metabolism, detoxification, regulations and intake of zearalenone: An oestrogenic mycotoxin. Food Chem. Toxicol. 2007, 45, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Mauro, T.; Hao, L.; Pop, L.C.; Buckley, B.; Schneider, S.H.; Bandera, E.V.; Shapses, S.A. Circulating zearalenone and its metabolites differ in women due to body mass index and food intake. Food Chem. Toxicol. 2018, 116, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Adibnia, E.; Razi, M.; Malekinejad, H. Zearalenone and 17 beta-estradiol induced damages in male rats reproduction potential; evidence for ERalpha and ERbeta receptors expression and steroidogenesis. Toxicon 2016, 120, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Zatecka, E.; Ded, L.; Elzeinova, F.; Kubatova, A.; Dorosh, A.; Margaryan, H.; Dostalova, P.; Korenkova, V.; Hoskova, K.; Peknicova, J. Effect of zearalenone on reproductive parameters and expression of selected testicular genes in mice. Reprod. Toxicol. 2014, 45, 20–30. [Google Scholar] [CrossRef]
- Wang, Y.; Zheng, W.; Bian, X.; Yuan, Y.; Gu, J.; Liu, X.; Liu, Z.; Bian, J. Zearalenone induces apoptosis and cytoprotective autophagy in primary Leydig cells. Toxicol. Lett. 2014, 226, 182–191. [Google Scholar] [CrossRef]
- Prouillac, C.; Koraichi, F.; Videmann, B.; Mazallon, M.; Rodriguez, F.; Baltas, M.; Lecoeur, S. In vitro toxicological effects of estrogenic mycotoxins on human placental cells: Structure activity relationships. Toxicol. Appl. Pharmacol. 2012, 259, 366–375. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, B.; He, X.; Cheng, W.H.; Xu, W.; Luo, Y.; Liang, R.; Luo, H.; Huang, K. Analysis of individual and combined effects of ochratoxin A and zearalenone on HepG2 and KK-1 cells with mathematical models. Toxins (Basel) 2014, 6, 1177–1192. [Google Scholar] [CrossRef]
- Gao, F.; Jiang, L.P.; Chen, M.; Geng, C.Y.; Yang, G.; Ji, F.; Zhong, L.F.; Liu, X.F. Genotoxic effects induced by zearalenone in a human embryonic kidney cell line. Mutat. Res. 2013, 755, 6–10. [Google Scholar] [CrossRef]
- Cao, H.; Zhi, Y.; Xu, H.; Fang, H.; Jia, X. Zearalenone causes embryotoxicity and induces oxidative stress and apoptosis in differentiated human embryonic stem cells. Toxicol. In Vitro 2019, 54, 243–250. [Google Scholar] [CrossRef]
- Mokoena, M.P.; Chelule, P.K.; Gqaleni, N. Reduction of fumonisin B1 and zearalenone by lactic acid bacteria in fermented maize meal. J. Food Prot. 2005, 68, 2095–2099. [Google Scholar] [CrossRef]
- Bae, S.W.; Kim, H.S.; Cha, Y.N.; Park, Y.S.; Jo, S.A.; Jo, I. Rapid increase in endothelial nitric oxide production by bradykinin is mediated by protein kinase A signaling pathway. Biochem. Biophys. Res. Commun. 2003, 306, 981–987. [Google Scholar] [CrossRef]
- Cho, D.H.; Choi, Y.J.; Jo, S.A.; Jo, I. Nitric oxide production and regulation of endothelial nitric-oxide synthase phosphorylation by prolonged treatment with troglitazone: Evidence for involvement of peroxisome proliferator-activated receptor (PPAR) gamma-dependent and PPARgamma-independent signaling pathways. J. Biol. Chem. 2004, 279, 2499–2506. [Google Scholar] [PubMed]
- Cho, D.H.; Choi, Y.J.; Jo, S.A.; Nam, J.H.; Jung, S.C.; Jo, I. Retinoic acid decreases nitric oxide production in endothelial cells: A role of phosphorylation of endothelial nitric oxide synthase at Ser(1179). Biochem. Biophys. Res. Commun. 2005, 326, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Kim, W.S.; Kim, J.Y.; Park, M.H.; Nam, J.H.; Yun, C.W.; Kwon, Y.G.; Jo, I. Chk1 and Hsp90 cooperatively regulate phosphorylation of endothelial nitric oxide synthase at serine 1179. Free Radic. Biol. Med. 2011, 51, 2217–2226. [Google Scholar] [CrossRef]
- Cho, D.H.; Park, J.H.; Lee, E.J.; Won, K.J.; Lee, S.H.; Kim, Y.H.; Hwang, S.; Kwon, K.J.; Shin, C.Y.; Song, K.H.; et al. Valproic acid increases NO production via the SH-PTP1-CDK5-eNOS-Ser(116) signaling cascade in endothelial cells and mice. Free Radic. Biol. Med. 2014, 76, 96–106. [Google Scholar] [CrossRef]
- Seo, J.; Lee, J.Y.; Sung, M.S.; Byun, C.J.; Cho, D.H.; Lee, H.J.; Park, J.H.; Cho, H.S.; Cho, S.J.; Jo, I. Arsenite Acutely Decreases Nitric Oxide Production via the ROS-Protein Phosphatase 1-Endothelial Nitric Oxide Synthase-Thr(497) Signaling Cascade. Biomol. Ther. (Seoul) 2014, 22, 510–518. [Google Scholar] [CrossRef][Green Version]
- Seo, J.; Cho, D.H.; Lee, H.J.; Sung, M.S.; Lee, J.Y.; Won, K.J.; Park, J.H.; Jo, I. Citron Rho-interacting kinase mediates arsenite-induced decrease in endothelial nitric oxide synthase activity by increasing phosphorylation at threonine 497: Mechanism underlying arsenite-induced vascular dysfunction. Free Radic. Biol. Med. 2016, 90, 133–144. [Google Scholar] [CrossRef]
- MacRitchie, A.N.; Jun, S.S.; Chen, Z.; German, Z.; Yuhanna, I.S.; Sherman, T.S.; Shaul, P.W. Estrogen upregulates endothelial nitric oxide synthase gene expression in fetal pulmonary artery endothelium. Circ. Res. 1997, 81, 355–362. [Google Scholar] [CrossRef]
- He, J.; Wei, C.; Li, Y.; Liu, Y.; Wang, Y.; Pan, J.; Liu, J.; Wu, Y.; Cui, S. Zearalenone and alpha-zearalenol inhibit the synthesis and secretion of pig follicle stimulating hormone via the non-classical estrogen membrane receptor GPR30. Mol. Cell. Endocrinol. 2018, 461, 43–54. [Google Scholar] [CrossRef]
- Nakamura, U.; Kadokawa, H. The nonsteroidal mycoestrogen zearalenone and its five metabolites suppress LH secretion from the bovine anterior pituitary cells via the estradiol receptor GPR30 in vitro. Theriogenology 2015, 84, 1342–1349. [Google Scholar] [CrossRef]
- Yang, Y.; Liu, S.; Fan, Z.; Li, Z.; Liu, J.; Xing, F. Sp1 modification of human endothelial nitric oxide synthase promoter increases the hypoxia-stimulated activity. Microvasc. Res. 2014, 93, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Padala, C.; Tupurani, M.A.; Puranam, K.; Gantala, S.; Shyamala, N.; Kondapalli, M.S.; Gundapaneni, K.K.; Mudigonda, S.; Galimudi, R.K.; Kupsal, K.; et al. Synergistic effect of collagenase-1 (MMP1), stromelysin-1 (MMP3) and gelatinase-B (MMP9) gene polymorphisms in breast cancer. PLoS ONE 2017, 12, e0184448. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Zaman, K.; Ryu, H.; Conforto, A.; Ratan, R.R. Sequence-selective DNA binding drugs mithramycin A and chromomycin A3 are potent inhibitors of neuronal apoptosis induced by oxidative stress and DNA damage in cortical neurons. Ann. Neurol. 2001, 49, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Karlseder, J.; Rotheneder, H.; Wintersberger, E. Interaction of Sp1 with the growth- and cell cycle-regulated transcription factor E2F. Mol. Cell. Biol. 1996, 16, 1659–1667. [Google Scholar] [CrossRef] [PubMed]
- Doetzlhofer, A.; Rotheneder, H.; Lagger, G.; Koranda, M.; Kurtev, V.; Brosch, G.; Wintersberger, E.; Seiser, C. Histone deacetylase 1 can repress transcription by binding to Sp1. Mol. Cell. Biol. 1999, 19, 5504–5511. [Google Scholar] [CrossRef]
- Lee, J.A.; Suh, D.C.; Kang, J.E.; Kim, M.H.; Park, H.; Lee, M.N.; Kim, J.M.; Jeon, B.N.; Roh, H.E.; Yu, M.Y.; et al. Transcriptional activity of Sp1 is regulated by molecular interactions between the zinc finger DNA binding domain and the inhibitory domain with corepressors, and this interaction is modulated by MEK. J. Biol. Chem. 2005, 280, 28061–28071. [Google Scholar] [CrossRef]
- Higgins, K.J.; Liu, S.; Abdelrahim, M.; Vanderlaag, K.; Liu, X.; Porter, W.; Metz, R.; Safe, S. Vascular endothelial growth factor receptor-2 expression is down-regulated by 17beta-estradiol in MCF-7 breast cancer cells by estrogen receptor alpha/Sp proteins. Mol. Endocrinol. 2008, 22, 388–402. [Google Scholar] [CrossRef]
- Ding, X.; Lichti, K.; Staudinger, J.L. The mycoestrogen zearalenone induces CYP3A through activation of the pregnane X receptor. Toxicol. Sci. 2006, 91, 448–455. [Google Scholar] [CrossRef]
- Pulakazhi Venu, V.K.; Saifeddine, M.; Mihara, K.; Tsai, Y.C.; Nieves, K.; Alston, L.; Mani, S.; McCoy, K.D.; Hollenberg, M.D.; Hirota, S.A. The pregnane X receptor and its microbiota-derived ligand indole 3-propionic acid regulate endothelium-dependent vasodilation. Am. J. Physiol. Endocrinol. Metab. 2019, 317, E350–E361. [Google Scholar] [CrossRef]
- Furchgott, R.F.; Zawadzki, J.V. The obligatory role of endothelial cells in the relaxation of arterial smooth muscle by acetylcholine. Nature 1980, 288, 373–376. [Google Scholar] [CrossRef]
- Bauer, V.; Sotnikova, R. Nitric oxide--the endothelium-derived relaxing factor and its role in endothelial functions. Gen. Physiol. Biophys. 2010, 29, 319–340. [Google Scholar] [CrossRef] [PubMed]
- Drummond, G.R.; Cai, H.; Davis, M.E.; Ramasamy, S.; Harrison, D.G. Transcriptional and posttranscriptional regulation of endothelial nitric oxide synthase expression by hydrogen peroxide. Circ. Res. 2000, 86, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Yan, G.J.; You, B.; Chen, S.P.; Liao, J.K.; Sun, J.X. Tumor necrosis factor-alpha downregulates endothelial nitric oxide synthase mRNA stability via translation elongation factor 1-alpha 1. Circ. Res. 2008, 103, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.S.; Kim, J.; Kwak, S.N.; Lee, D.K.; Ha, K.S.; Won, M.H.; Jeoung, D.; Lee, H.; Kwon, Y.G.; Kim, Y.M. Functional role of NF-kappaB in expression of human endothelial nitric oxide synthase. Biochem. Biophys. Res. Commun. 2014, 448, 101–107. [Google Scholar] [CrossRef]
- Gonzalez-Rubio, S.; Lopez-Sanchez, L.; Munoz-Castaneda, J.; Linares, C.I.; Aguilar-Melero, P.; Rodriguez-Peralvarez, M.; Sanchez-Sanchez, R.; Fernandez-Alvarez, A.; Casado, M.; Montero-Alvarez, J.L.; et al. GCDCA down-regulates gene expression by increasing Sp1 binding to the NOS-3 promoter in an oxidative stress dependent manner. Biochem. Pharmacol. 2015, 96, 39–51. [Google Scholar] [CrossRef]
- Ma, X.; Idle, J.R.; Gonzalez, F.J. The pregnane X receptor: From bench to bedside. Expert Opin. Drug Metab. Toxicol. 2008, 4, 895–908. [Google Scholar] [CrossRef]
- Kliewer, S.A.; Goodwin, B.; Willson, T.M. The nuclear pregnane X receptor: A key regulator of xenobiotic metabolism. Endocr. Rev. 2002, 23, 687–702. [Google Scholar] [CrossRef]
- Pavek, P. Pregnane X Receptor (PXR)-Mediated Gene Repression and Cross-Talk of PXR with Other Nuclear Receptors via Coactivator Interactions. Front. Pharmacol. 2016, 7, 456. [Google Scholar] [CrossRef]
- Kodama, S.; Moore, R.; Yamamoto, Y.; Negishi, M. Human nuclear pregnane X receptor cross-talk with CREB to repress cAMP activation of the glucose-6-phosphatase gene. Biochem. J. 2007, 407, 373–381. [Google Scholar] [CrossRef]
- Husmann, M.; Dragneva, Y.; Romahn, E.; Jehnichen, P. Nuclear receptors modulate the interaction of Sp1 and GC-rich DNA via ternary complex formation. Biochem. J. 2000, 352 Pt 3, 763–772. [Google Scholar] [CrossRef]
- Oladimeji, P.O.; Chen, T. PXR: More Than Just a Master Xenobiotic Receptor. Mol. Pharmacol. 2018, 93, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Olsen, M.; Malmlof, K.; Pettersson, H.; Sandholm, K.; Kiessling, K.H. Plasma and urinary levels of zearalenone and alpha-zearalenol in a prepubertal gilt fed zearalenone. Acta Pharmacol. Toxicol. (Copenh) 1985, 56, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Wang, B.; Li, X.; Wang, T.; Zou, H.; Gu, J.; Yuan, Y.; Liu, X.; Bai, J.; Bian, J.; et al. Zearalenone Promotes Cell Proliferation or Causes Cell Death? Toxins (Basel) 2018, 10, 184. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Wang, Y.; Ma, L.; Chen, X.; Lu, Y.; Ge, F.; Chen, Y.; Lian, Q.; Jin, X.D.; Ge, R.S. Zearalenone Delays Rat Leydig Cell Regeneration. Toxicol. Sci. 2018, 164, 60–71. [Google Scholar] [CrossRef]
- Zollner, P.; Jodlbauer, J.; Kleinova, M.; Kahlbacher, H.; Kuhn, T.; Hochsteiner, W.; Lindner, W. Concentration levels of zearalenone and its metabolites in urine, muscle tissue, and liver samples of pigs fed with mycotoxin-contaminated oats. J. Agric. Food Chem. 2002, 50, 2494–2501. [Google Scholar] [CrossRef]
- Koraichi, F.; Videmann, B.; Mazallon, M.; Benahmed, M.; Prouillac, C.; Lecoeur, S. Zearalenone exposure modulates the expression of ABC transporters and nuclear receptors in pregnant rats and fetal liver. Toxicol. Lett. 2012, 211, 246–256. [Google Scholar] [CrossRef]
- Kowalska, K.; Habrowska-Gorczynska, D.E.; Urbanek, K.A.; Dominska, K.; Piastowska-Ciesielska, A.W. Estrogen Receptor alpha Is Crucial in Zearalenone-Induced Invasion and Migration of Prostate Cancer Cells. Toxins (Basel) 2018, 10, 98. [Google Scholar] [CrossRef]
- Kowalska, K.; Habrowska-Gorczynska, D.E.; Urbanek, K.A.; Dominska, K.; Sakowicz, A.; Piastowska-Ciesielska, A.W. Estrogen receptor beta plays a protective role in zearalenone-induced oxidative stress in normal prostate epithelial cells. Ecotoxicol. Environ. Saf. 2019, 172, 504–513. [Google Scholar] [CrossRef]
- Wang, J.J.; Wei, Z.K.; Han, Z.; Liu, Z.Y.; Zhu, X.Y.; Li, X.W.; Wang, K.; Yang, Z.T. Zearalenone Induces Estrogen-Receptor-Independent Neutrophil Extracellular Trap Release in Vitro. J. Agric. Food Chem. 2019, 67, 4588–4594. [Google Scholar] [CrossRef]
- Min, J.; Jin, Y.M.; Moon, J.S.; Sung, M.S.; Jo, S.A.; Jo, I. Hypoxia-induced endothelial NO synthase gene transcriptional activation is mediated through the tax-responsive element in endothelial cells. Hypertension 2006, 47, 1189–1196. [Google Scholar] [CrossRef]
- Kim, H.P.; Lee, J.Y.; Jeong, J.K.; Bae, S.W.; Lee, H.K.; Jo, I. Nongenomic stimulation of nitric oxide release by estrogen is mediated by estrogen receptor alpha localized in caveolae. Biochem. Biophys. Res. Commun. 1999, 263, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.Y.; Lee, S.; Hwang, S.; Jo, S.A.; Kim, M.; Kim, Y.J.; Pang, M.G.; Jo, I. Histone H3 lysine 27 and 9 hypermethylation within the Bad promoter region mediates 5-Aza-2’-deoxycytidine-induced Leydig cell apoptosis: Implications of 5-Aza-2’-deoxycytidine toxicity to male reproduction. Apoptosis 2013, 18, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Kizuka, Y.; Kitazume, S.; Yoshida, M.; Taniguchi, N. Brain-specific expression of N-acetylglucosaminyltransferase IX (GnT-IX) is regulated by epigenetic histone modifications. J. Biol. Chem. 2011, 286, 31875–31884. [Google Scholar] [CrossRef] [PubMed]
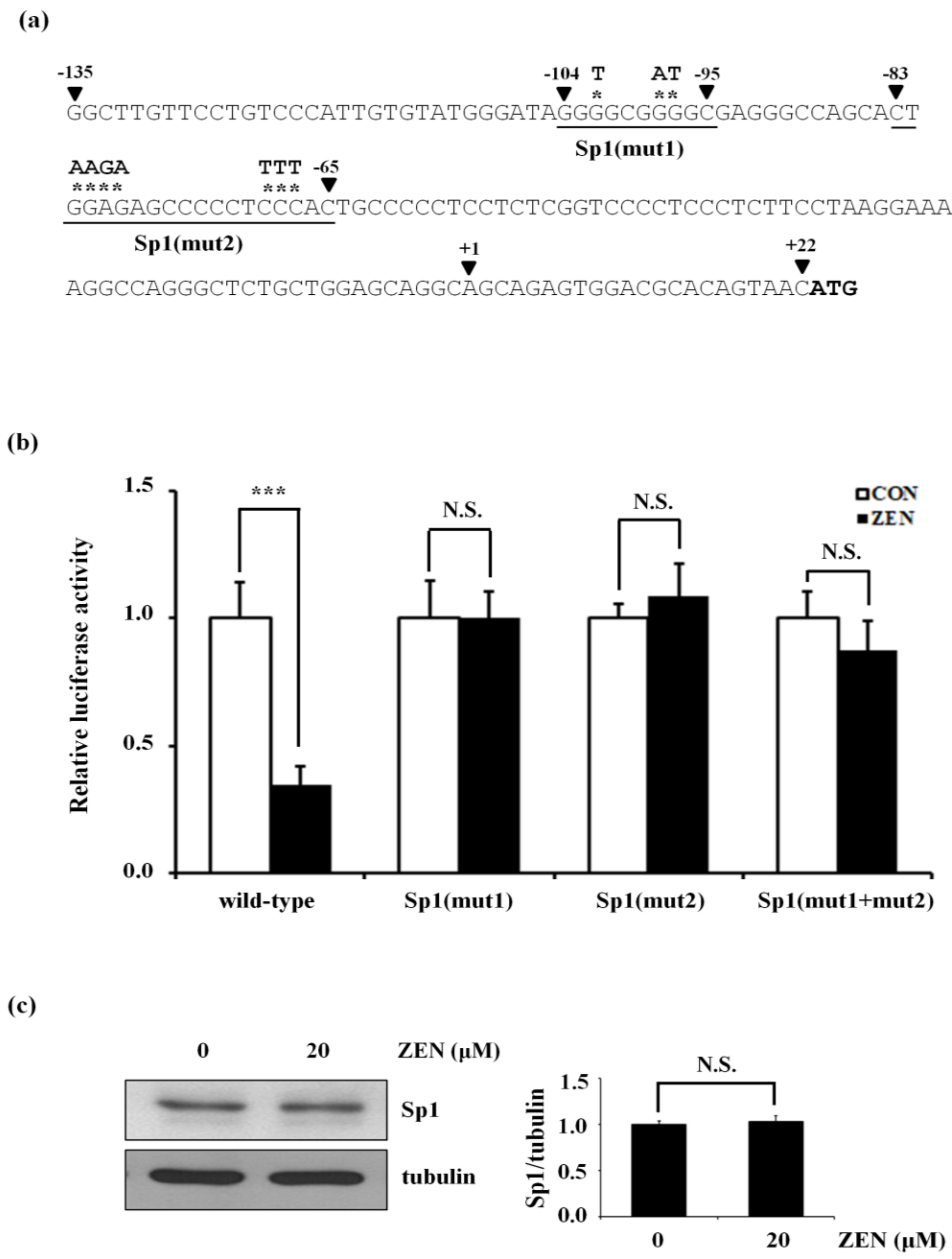
 ) are indicated in bold. (b) After pretreatment with vehicle or 20 nM of the Sp1 inhibitor mithramycin A for 1 h, BAECs were further co-treated with or without 20 μM of ZEN for 8 h. Binding of Sp1 to the bovine eNOS promoter gene relative to the total chromatin extract (Input) was quantified using a ChIP assay, as described in Materials and Methods. (c) Luciferase activities of the bovine eNOS(−135) vector and CMV promoter (Renilla) in BAECs treated with vehicle or 20 nM mithramycin A for 1 h, followed by further co-treatment with 20 µM of ZEN for 8 h. (d) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses, and (e) NO production was quantified using the Griess method in BAECs treated with vehicle or 20 nM mithramycin A followed by further co-treatment with 20 µM ZEN for 4, 8, 16, and 24 h. (f) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses of BAECs transfected with siRNA targeting Sp1 for 24 h and then exposed to 20 µM of ZEN for 8 h. N.C., negative control. Plots depict mean fold alterations from the control (± S.D.) from at least four independent trials. Statistical significance is denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, and N.S. (not statistically significant).
) are indicated in bold. (b) After pretreatment with vehicle or 20 nM of the Sp1 inhibitor mithramycin A for 1 h, BAECs were further co-treated with or without 20 μM of ZEN for 8 h. Binding of Sp1 to the bovine eNOS promoter gene relative to the total chromatin extract (Input) was quantified using a ChIP assay, as described in Materials and Methods. (c) Luciferase activities of the bovine eNOS(−135) vector and CMV promoter (Renilla) in BAECs treated with vehicle or 20 nM mithramycin A for 1 h, followed by further co-treatment with 20 µM of ZEN for 8 h. (d) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses, and (e) NO production was quantified using the Griess method in BAECs treated with vehicle or 20 nM mithramycin A followed by further co-treatment with 20 µM ZEN for 4, 8, 16, and 24 h. (f) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses of BAECs transfected with siRNA targeting Sp1 for 24 h and then exposed to 20 µM of ZEN for 8 h. N.C., negative control. Plots depict mean fold alterations from the control (± S.D.) from at least four independent trials. Statistical significance is denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, and N.S. (not statistically significant).
 ) are indicated in bold. (b) After pretreatment with vehicle or 20 nM of the Sp1 inhibitor mithramycin A for 1 h, BAECs were further co-treated with or without 20 μM of ZEN for 8 h. Binding of Sp1 to the bovine eNOS promoter gene relative to the total chromatin extract (Input) was quantified using a ChIP assay, as described in Materials and Methods. (c) Luciferase activities of the bovine eNOS(−135) vector and CMV promoter (Renilla) in BAECs treated with vehicle or 20 nM mithramycin A for 1 h, followed by further co-treatment with 20 µM of ZEN for 8 h. (d) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses, and (e) NO production was quantified using the Griess method in BAECs treated with vehicle or 20 nM mithramycin A followed by further co-treatment with 20 µM ZEN for 4, 8, 16, and 24 h. (f) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses of BAECs transfected with siRNA targeting Sp1 for 24 h and then exposed to 20 µM of ZEN for 8 h. N.C., negative control. Plots depict mean fold alterations from the control (± S.D.) from at least four independent trials. Statistical significance is denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, and N.S. (not statistically significant).
) are indicated in bold. (b) After pretreatment with vehicle or 20 nM of the Sp1 inhibitor mithramycin A for 1 h, BAECs were further co-treated with or without 20 μM of ZEN for 8 h. Binding of Sp1 to the bovine eNOS promoter gene relative to the total chromatin extract (Input) was quantified using a ChIP assay, as described in Materials and Methods. (c) Luciferase activities of the bovine eNOS(−135) vector and CMV promoter (Renilla) in BAECs treated with vehicle or 20 nM mithramycin A for 1 h, followed by further co-treatment with 20 µM of ZEN for 8 h. (d) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses, and (e) NO production was quantified using the Griess method in BAECs treated with vehicle or 20 nM mithramycin A followed by further co-treatment with 20 µM ZEN for 4, 8, 16, and 24 h. (f) Protein expression of eNOS relative to tubulin was quantified using Western blot analyses of BAECs transfected with siRNA targeting Sp1 for 24 h and then exposed to 20 µM of ZEN for 8 h. N.C., negative control. Plots depict mean fold alterations from the control (± S.D.) from at least four independent trials. Statistical significance is denoted as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, and N.S. (not statistically significant).
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, H.-J.; Park, J.-H.; Oh, S.-Y.; Cho, D.-H.; Kim, S.; Jo, I. Zearalenone-Induced Interaction between PXR and Sp1 Increases Binding of Sp1 to a Promoter Site of the eNOS, Decreasing Its Transcription and NO Production in BAECs. Toxins 2020, 12, 421. https://doi.org/10.3390/toxins12060421
Lee H-J, Park J-H, Oh S-Y, Cho D-H, Kim S, Jo I. Zearalenone-Induced Interaction between PXR and Sp1 Increases Binding of Sp1 to a Promoter Site of the eNOS, Decreasing Its Transcription and NO Production in BAECs. Toxins. 2020; 12(6):421. https://doi.org/10.3390/toxins12060421
Chicago/Turabian StyleLee, Hyeon-Ju, Jung-Hyun Park, Se-Young Oh, Du-Hyong Cho, Suji Kim, and Inho Jo. 2020. "Zearalenone-Induced Interaction between PXR and Sp1 Increases Binding of Sp1 to a Promoter Site of the eNOS, Decreasing Its Transcription and NO Production in BAECs" Toxins 12, no. 6: 421. https://doi.org/10.3390/toxins12060421
APA StyleLee, H.-J., Park, J.-H., Oh, S.-Y., Cho, D.-H., Kim, S., & Jo, I. (2020). Zearalenone-Induced Interaction between PXR and Sp1 Increases Binding of Sp1 to a Promoter Site of the eNOS, Decreasing Its Transcription and NO Production in BAECs. Toxins, 12(6), 421. https://doi.org/10.3390/toxins12060421






