Molecular Docking and Dynamics Simulation Studies Predict Munc18b as a Target of Mycolactone: A Plausible Mechanism for Granule Exocytosis Impairment in Buruli Ulcer Pathogenesis
Abstract
1. Introduction
2. Results
2.1. Molecular Docking
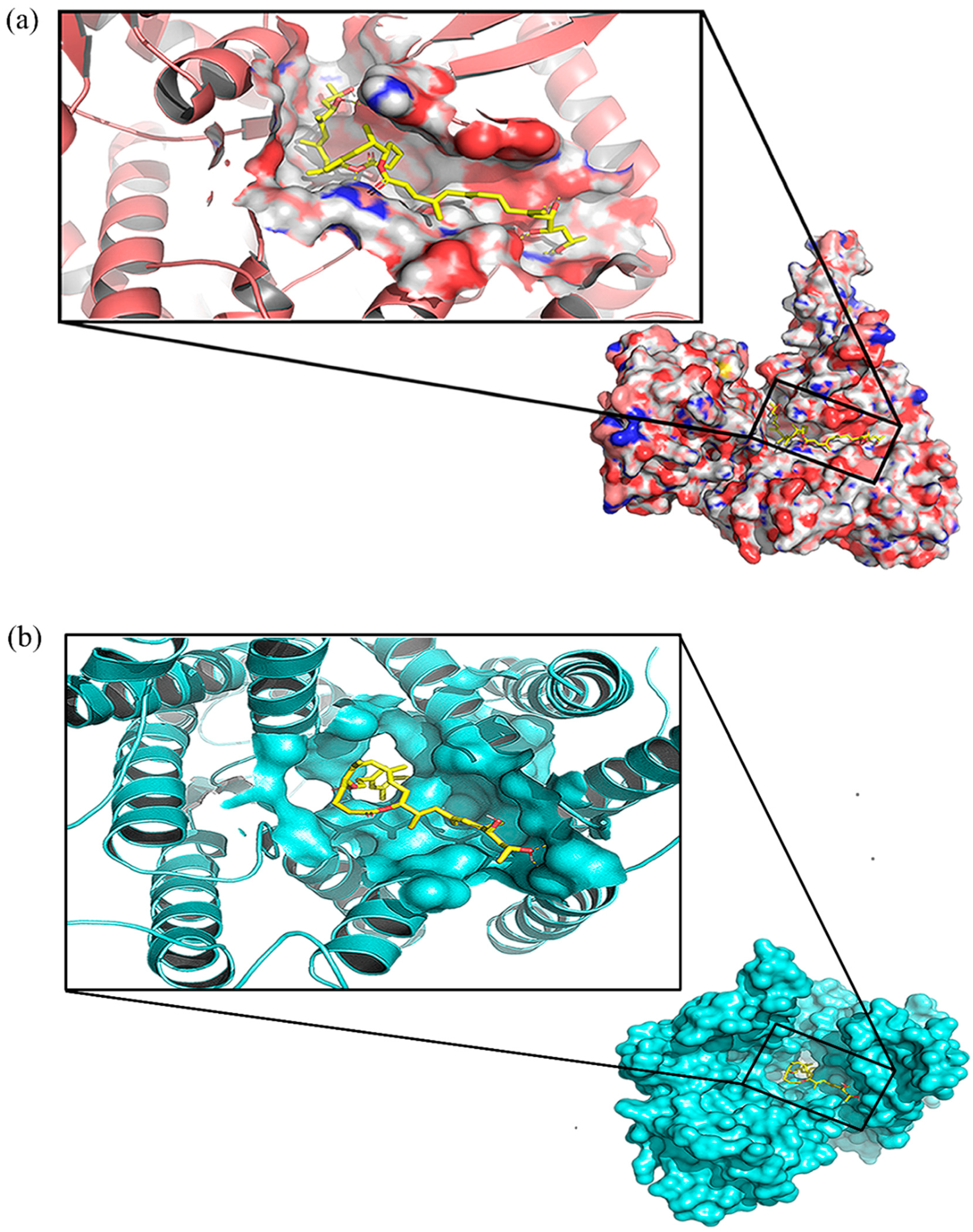
2.1.1. Mycolactone has a Higher Binding Affinity for Munc18b than other SNARE Proteins
2.1.2. Predicted Binding Energies for Known Mycolactone Receptors (Sec61 and AT2R) and Munc18b Are High
2.2. Molecular Dynamics Simulation
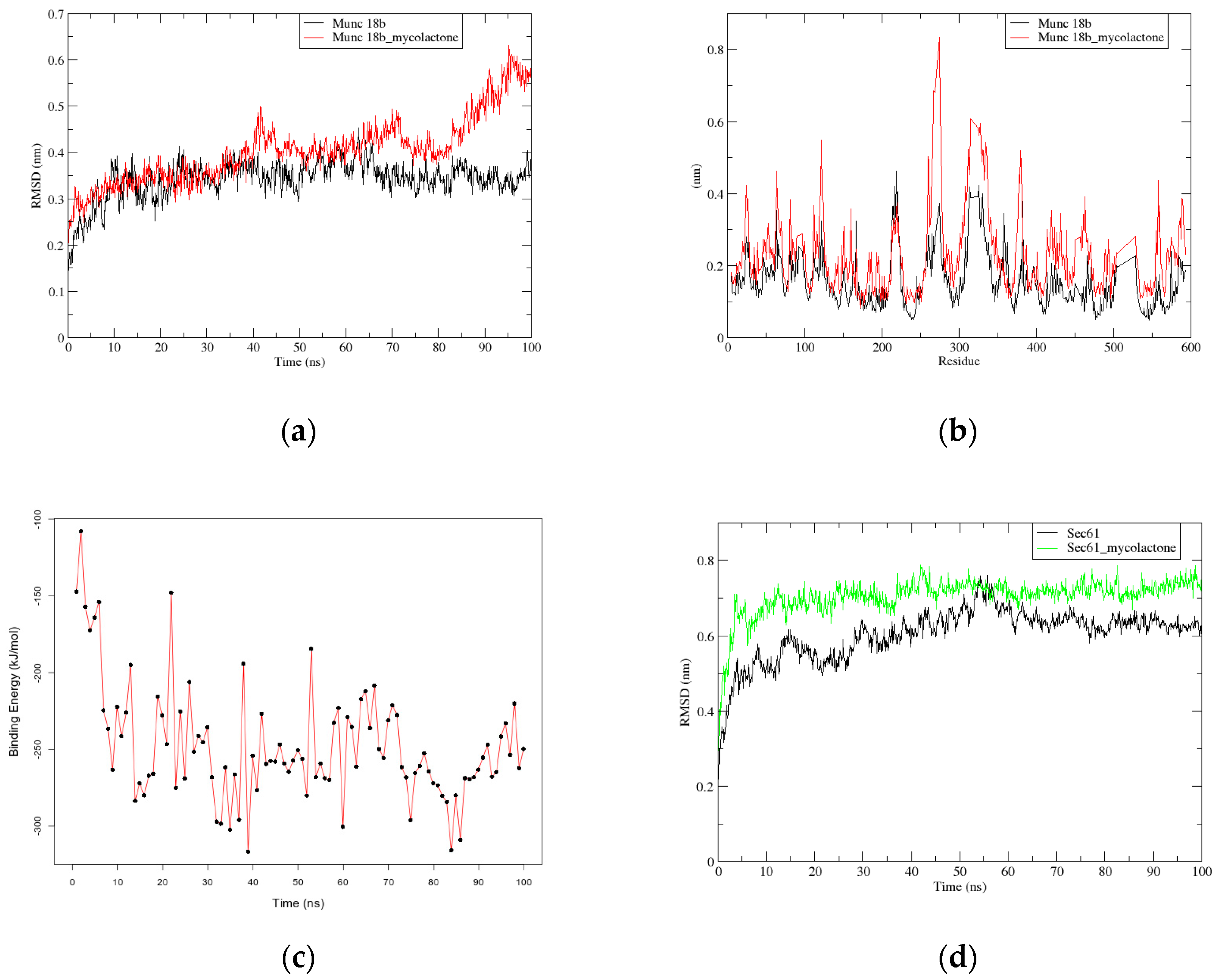
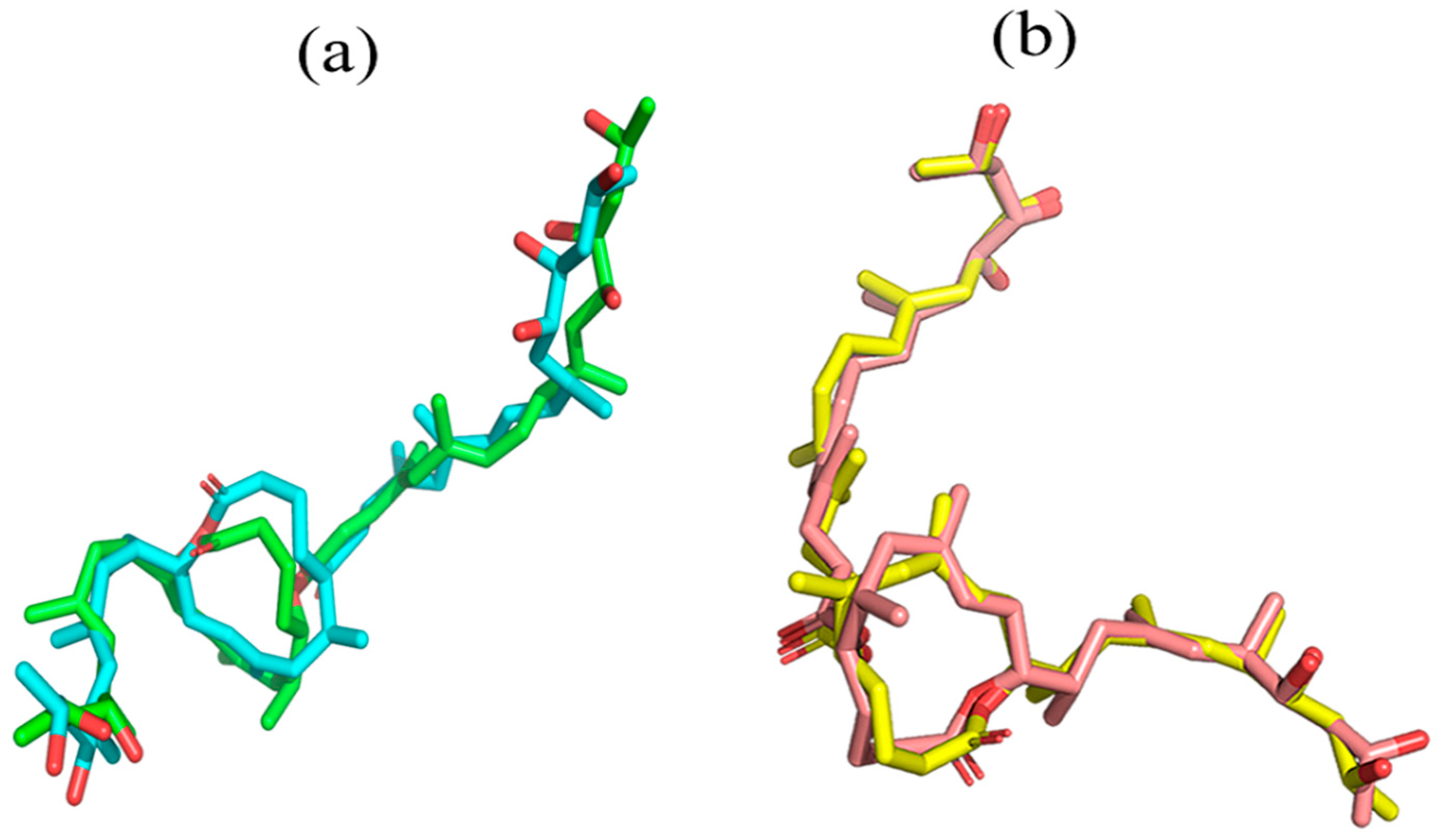
2.2.1. RMSD and RMSF Graphs show Instability in Munc18b’s Structure after the Binding of Mycolactone
2.2.2. G_mmpbsa Calculations show Strong Binding between Mycolactone and Munc18b
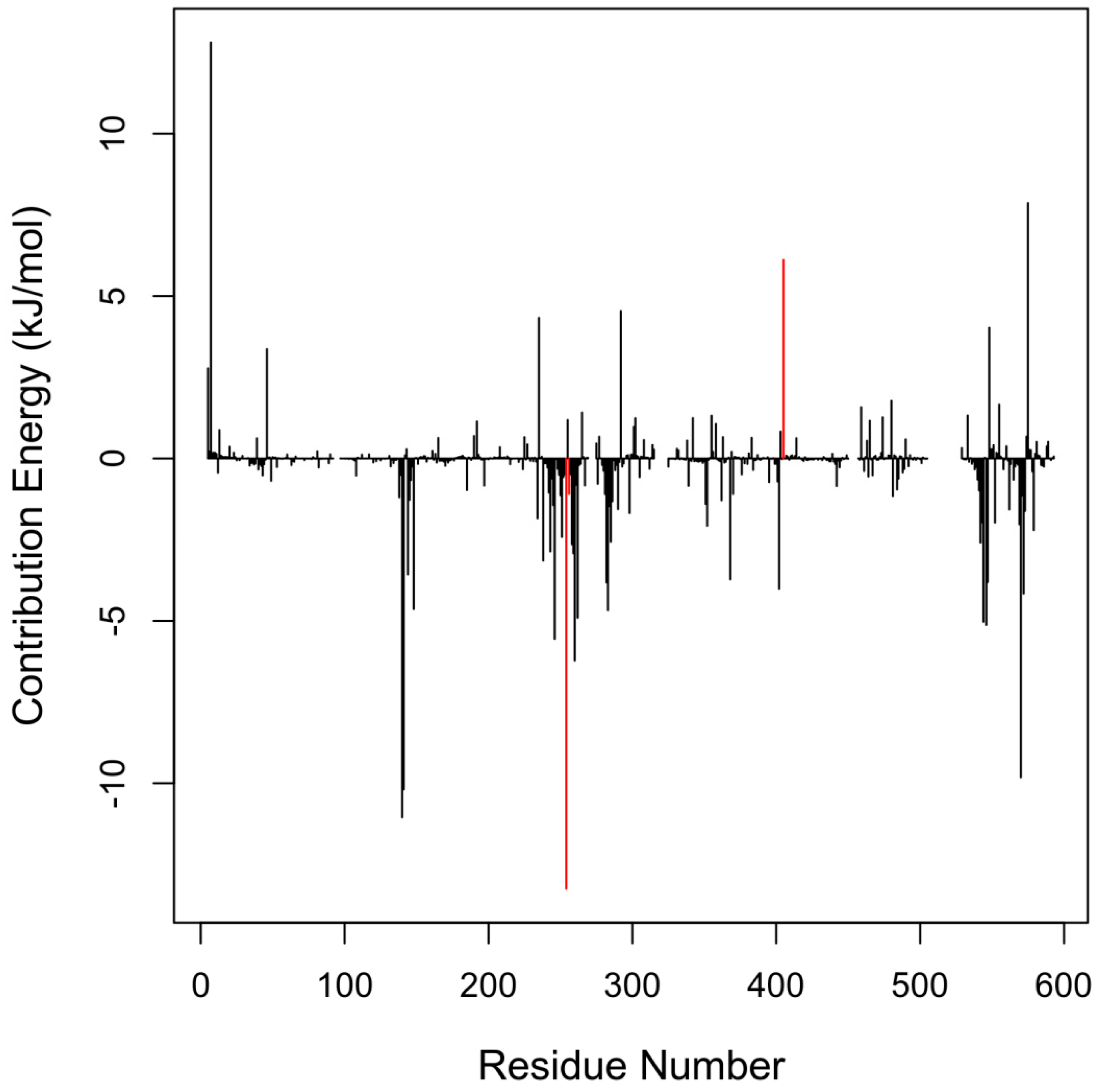
2.3. Critical Residues for Mycolactone and Munc18b Binding
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. The Crystal Structures
5.2. Molecular Docking
5.3. Molecular Dynamics Simulations
5.4. Binding Energy Calculations using the MM-PBSA Method
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Marion, E.; Song, O.R.; Christophe, T.; Babonneau, J.; Fenistein, D.; Eyer, J.; Letournel, F.; Henrion, D.; Clere, N.; Paille, V.; et al. Mycobacterial toxin induces analgesia in Buruli ulcer by targeting the angiotensin pathways. Cell 2014, 157, 1565–1576. [Google Scholar] [CrossRef] [PubMed]
- Chany, A.C.; Tresse, C.; Casarotto, V.; Blanchard, N. History, biology, and chemistry of Mycobacterium ulcerans infections (Buruli ulcer disease). Nat. Prod. Rep. 2013, 30, 1527–1567. [Google Scholar] [CrossRef]
- Ogbechi, J.; Ruf, M.T.; Hall, B.S.; Bodman-Smith, K.; Vogel, M.; Wu, H.L.; Stainer, A.; Esmon, C.T.; Ahnström, J.; Pluschke, G.; et al. Mycolactone-Dependent Depletion of Endothelial Cell Thrombomodulin Is Strongly Associated with Fibrin Deposition in Buruli Ulcer Lesions. PLoS Pathog. 2015, 11, e1005011. [Google Scholar] [CrossRef] [PubMed]
- Wansbrough-Jones, M.; Phillips, R. Buruli ulcer: Emerging from obscurity. Lancet 2006, 367, 1849–1858. [Google Scholar] [CrossRef]
- Wilson, M.D.; Boakye, D.A.; Mosi, L.; Asiedu, K.; Wilson, M.D. In the Case of Transmission of Mycobacterium ulcerans in Buruli Ulcer Disease Acanthamoeba Species Stand Accused. Ghana Med. J. 2011, 45, 31–34. [Google Scholar] [CrossRef] [PubMed]
- Hall, B.S.; Hill, K.; McKenna, M.; Ogbechi, J.; High, S.; Willis, A.E.; Simmonds, R.E. The Pathogenic Mechanism of the Mycobacterium ulcerans Virulence Factor, Mycolactone, Depends on Blockade of Protein Translocation into the ER. PLoS Pathog. 2014, 10, e1004061. [Google Scholar] [CrossRef] [PubMed]
- López, C.A.; Unkefer, C.J.; Swanson, B.I.; Swanson, J.M.J.; Gnanakaran, S. Membrane perturbing properties of toxin mycolactone from Mycobacterium ulcerans. PLoS Comput. Biol. 2018, 14, e1005972. [Google Scholar] [CrossRef] [PubMed]
- Snyder, D.S.; Small, P.L.C. Uptake and cellular actions of mycolactone, a virulence determinant for Mycobacterium ulcerans. Microb. Pathog. 2003, 34, 91–101. [Google Scholar] [CrossRef]
- Phillips, R.; Sarfo, F.; Duah, M.S.; Wansbrough-Jones, M.; Frimpong, M. Buruli ulcer: Wound care and rehabilitation. Chronic Wound Care Manag. Res. 2016, 3, 73–84. [Google Scholar] [CrossRef]
- Velding, K.; Klis, S.A.; Abass, K.M.; Tuah, W.; Stienstra, Y.; van der Werf, T. Wound Care in Buruli ulcer Disease in Ghana and Benin. Am. J. Trop. Med. Hyg. 2014, 91, 313–318. [Google Scholar] [CrossRef] [PubMed]
- George, K.M.; Pascopella, L.; Welty, D.M.; Small, P.L.C. A Mycobacterium ulcerans Toxin, Mycolactone, Causes Apoptosis in Guinea Pig Ulcers and Tissue Culture Cells. Infect. Immun. 2000, 68, 877–883. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Young, A.; McNaught, C.-E. The Physiology of Wound Healing. Surgery 2017, 35, 473–477. [Google Scholar] [CrossRef]
- Stadelmann, W.K.; Digenis, A.G.; Tobin, G.R. Physiology and Healing Dynamics of ChronicCutaneous Wounds. Am. J. Surg. 1998, 176, 26S–38S. [Google Scholar] [CrossRef]
- Amin, K. The Role of Mast Cells In Allergic Inflammation. Respir. Med. 2012, 106, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Frenzel, L.; Hermine, O. Mast Cells and Inflammation. Joint Bone Spine 2013, 80, 141–145. [Google Scholar] [CrossRef]
- Lorentz, A.; Baumann, A.; Vitte, J.; Blank, U. The SNARE Machinery in Mast Cell Secretion. Front. Immunol. 2012, 3, 143. [Google Scholar] [CrossRef]
- Sharda, A.; Flaumenhaft, R. The Life Cycle of Platelet Granules. F1000Research 2018, 7, 236. [Google Scholar] [CrossRef]
- Sadiq, A.; Shah, A.; Jeschke, M.G.; Belo, C.; Hayat, M.Q.; Murad, S.; Amini-Nik, S. The Role of Serotonin During Skin Healing in Post-Thermal Injury. Int. J. Mol. Sci. 2018, 19, 1034. [Google Scholar] [CrossRef]
- Lansdown, A.B.G. Calcium: A Potential Central Regulator in Wound Healing in the Skin. Wound Repair Regen. 2002, 10, 271–285. [Google Scholar] [CrossRef]
- Orrenius, S.; Zhivotovsky, B.; Nicotera, P. Regulation of Cell Death: The Calcium—Apoptosis Link. Nat. Rev. Mol. Cell Biol. 2003, 4, 552–565. [Google Scholar] [CrossRef]
- Bechert, K.; Abraham, S.E. Pain Management and Wound Care. J. Am. Coll. Certif. Wound Spec. 2009, 1, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Hagenston, A.M.; Simonetti, M. Neuronal Calcium Signaling in Chronic Pain. Cell Tissue Res. 2014, 357, 407–426. [Google Scholar] [CrossRef] [PubMed]
- Demangel, C.; High, S. Sec61 Blockade by Mycolactone: A Central Mechanism in Buruli Ulcer Disease. Biol. Cell 2018, 110, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Guenin-Macé, L.; Veyron-Churlet, R.; Thoulouze, M.-I.; Romet-Lemonne, G.; Hong, H.; Leadlay, P.F.; Danckaert, A.; Ruf, M.-T.; Mostowy, S.; Zurzolo, C.; et al. Mycolactone Activation of Wiskott-Aldrich Syndrome Proteins Underpins Buruli Ulcer Formation. J. Clin. Investig. 2013, 123, 1501–1512. [Google Scholar] [CrossRef]
- Baron, L.; Paatero, A.O.; Morel, J.-D.; Impens, F.; Guenin-Mace, L.; Saint-Auret, S.; Blanchard, N.; Dillmann, R.; Niang, F.; Pellegrini, S.; et al. Mycolactone subverts immunity by selectively blocking the Sec61 translocon. J. Exp. Med. 2016, 213, 2885–2896. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the Speed and Accuracy of Docking with a New Scoring Function, Efficient Optimization, and Multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Dallakyan, S.; Olson, A.J. Small-Molecule Library Screening by Docking with PyRx. Methods Mol. Biol. 2015, 1263, 243–250. [Google Scholar] [PubMed]
- Shityakov, S.; Förster, C. In Silico Predictive Model to Determine Vector-Mediated Transport Properties for the Blood-Brain Barrier Choline Transporter. Adv. Appl. Bioinform. Chem. 2014, 7, 23–36. [Google Scholar] [CrossRef]
- Lang, S.; Benedix, J.; Fedeles, S.V.; Schorr, S.; Schirra, C.; Schauble, N.; Jalal, C.; Greiner, M.; Hassdenteufel, S.; Tatzelt, J.; et al. Different Effects of Sec61, Sec62 and Sec63 Depletion on Transport of Polypeptides into the Endoplasmic Reticulum of Mammalian Cells. J. Cell Sci. 2012, 125, 1958–1969. [Google Scholar] [CrossRef]
- Sarfo, F.S.; Phillips, R.; Wansbrough-Jones, M.; Simmonds, R.E. Recent Advances: Role of Mycolactone in the Pathogenesis and Monitoring of Mycobacterium Ulcerans Infection/Buruli Ulcer Disease. Cell. Microbiol. 2016, 18, 17–29. [Google Scholar] [CrossRef]
- Nouet, S.; Amzallag, N.; Li, J.M.; Louis, S.; Seitz, I.; Cui, T.X.; Alleaume, A.M.; di Benedetto, M.; Boden, C.; Masson, M.; et al. Trans-Inactivation of Receptor Tyrosine Kinases by Novel Angiotensin II AT2 Receptor-Interacting Protein, ATIP. J. Biol. Chem. 2004, 279, 28989–28997. [Google Scholar] [CrossRef]
- Heifets, A.; Lilien, R.H. LigAlign: Flexible ligand-based active site alignment and analysis. J. Mol. Graph. Model. 2010, 29, 93–101. [Google Scholar] [CrossRef]
- Liu, K.; Kokubo, H. Exploring the Stability of Ligand Binding Modes to Proteins by Molecular Dynamics Simulations: A Cross-docking Study. J. Chem. Inf. Model. 2017, 57, 2514–2522. [Google Scholar] [CrossRef]
- Kumari, R.; Kumar, R.; Lynn, A. G-mmpbsa-A GROMACS Tool for High-Throughput MM-PBSA Calculations. J. Chem. Inf. Model. 2014, 54, 1951–1962. [Google Scholar] [CrossRef]
- Baker, N.A.; Sept, D.; Joseph, S.; Holst, M.J.; McCammon, J.A. Electrostatics of Nanosystems: Application to Microtubules and the Ribosome. Proc. Natl. Acad. Sci. USA 2001, 98, 10037–10041. [Google Scholar] [CrossRef]
- Genheden, S.; Ryde, U. The MM/PBSA and MM/GBSA Methods to Estimate Ligand-Binding Affinities. Expert Opin. Drug Discov. 2015, 10, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Hackmann, Y.; Graham, S.C.; Ehl, S.; Honing, S.; Lehmberg, K.; Arico, M.; Owen, D.J.; Griffiths, G.M. Syntaxin Binding Mechanism and Disease-Causing Mutations in Munc18-2. Proc. Natl. Acad. Sci. USA 2013, 110, E4482–E4491. [Google Scholar] [CrossRef]
- Schrödinger. The PyMOL Molecular Graphics System. Schrödinger LLC wwwpymolorg Version 1. 2015. Available online: http://www.pymol.org (accessed on 10 January 2019).
- Stadt, U.Z.; Rohr, J.; Seifert, W.; Koch, F.; Grieve, S.; Pagel, J.; Strauß, J.; Kasper, B.; Nürnberg, G.; Becker, C.; et al. Familial Hemophagocytic Lymphohistiocytosis Type 5 (FHL-5) Is Caused by Mutations in Munc18-2 and Impaired Binding to Syntaxin 11. Am. J. Hum. Genet. 2009, 85, 482–492. [Google Scholar] [CrossRef]
- Dulubova, I. Munc18. Encycl. Neurosci. 2010, 24, 1131–1139. [Google Scholar]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Guex, N.; Peitsch, M.C.; Schwede, T. Automated Comparative Protein Structure Modeling with SWISS-MODEL and Swiss-PdbViewer: A Historical Perspective. Electrophoresis 2009, 30, S162–S173. [Google Scholar] [CrossRef]
- Freedman, S.J.; Song, H.K.; Xu, Y.; Sun, Z.-Y.J.; Eck, M.J. Homotetrameric Structure of the SNAP-23 N-terminal Coiled-Coil Domain. J. Biol. Chem. 2003, 278, 13462–13467. [Google Scholar] [CrossRef] [PubMed]
- Diao, J.; Liu, R.; Rong, Y.; Zhao, M.; Zhang, J.; Lai, Y.; Zhou, Q.; Wilz, L.M.; Li, J.; Vivona, S.; et al. ATG14 Promotes Membrane Tethering and Fusion of Autophagosomes to Endolysosomes. Nature 2015, 520, 563–566. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Machius, M.; Dulubova, I.; Dai, H.; Sudhof, T.C.; Tomchick, D.R.; Rizo, J. Structural Basis for a Munc13-1 Dimeric to Munc13-1/Rim Heterodimer Switch. PLoS Biol. 2006, 4, E192. [Google Scholar] [CrossRef]
- Colbert, K.N.; Hattendorf, D.A.; Weiss, T.M.; Burkhardt, P.; Fasshauer, D.; Weis, W.I. Syntaxin1a Variants Lacking an N-peptide or Bearing the LE Mutation Bind to Munc18a in a Closed Conformation. Proc. Natl. Acad. Sci. USA 2013, 110, 12637–12642. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology Modelling of Protein Structures and Complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Kim, A.S.; Kakalis, L.T.; Abdul-Manan, N.; Liu, G.A.; Rosen, M.K. Autoinhibition and Activation Mechanisms of the Wiskott-Aldrich Syndrome Protein. Nature 2000, 404, 151–158. [Google Scholar] [CrossRef]
- Voorhees, R.M.; Hegde, R.S. Structure of the Sec61 Channel Opened by a Signal Sequence. Science 2016, 351, 88–91. [Google Scholar] [CrossRef]
- Zhang, H.; Han, G.W.; Batyuk, A.; Ishchenko, A.; White, K.L.; Patel, N.; Sadybekov, A.; Zamlynny, B.; Rudd, M.T.; Hollenstein, K.; et al. Structural Basis for Selectivity and Diversity in Angiotensin II Receptors. Nature 2017, 544, 327–332. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple Ligand–Protein Interaction Diagrams for Drug Discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High Performance Molecular Simulations Through Multi-Level Parallelism from Laptops to Supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Turner, P.J. XMGRACE, Version 5.1.25; Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology: Beaverton, OR, USA, 2015; p. 2005. [Google Scholar]
- R. C. Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
| SNARE Protein Target | H-Bond Residues | Hydrophobic Bond Residues |
|---|---|---|
| Munc18b | His245, Asp255, Leu256, Asp262, Arg402, Arg405, Thr574, Arg575 | Lys7, Ser43, Leu138, Tyr140, Ser146, Tyr254, Gln261, Val361, Ile570, Thr572, Asp578. |
| SNAP23 | Ser39 | Gln40, Gly43, Thr46, Ile47, Leu50, Asp51, Lys54 |
| VAMP8 | Thr48, Ser55 | Leu51, Glu52, Thr54, Glu56, Phe58, Lys59 |
| Syntaxin 11 | None | Gln140, His144, Asn147, Met151, Arg154, Glu218, Ile221, Arg222, Phe228, Leu229, Ala232, His255 |
| Munc13-4 | Arg86, Glu95, Asp124, Ala125 | Val78, Trp79, Ile80, Thr84, Ile85, Leu97, Thr98, Leu99, Asp100, Phe118 |
| Protein | Binding Energy /kcal/mol | Location in Human Cells | Functional Roles |
|---|---|---|---|
| Munc18b | −8.5 | Localized on the plasma membrane in platelet cells and vesicle membranes of mast cells | Interacts with STX 3 in mast cells and STX11 in platelets to regulate granule exocytosis [16,17]. |
| Sec61 | −8.9 | Endoplasmic reticulum(ER) | Responsible for protein translocation into the endoplasmic reticulum [29] |
| Wiskott-Aldrich protein (WASP/NWASP) | −7.1 | Cytoskeleton | Dynamic extensive alteration of actin filament through its interaction with Arp2/3 complex [30] |
| Type 2 angiotensin II receptor (AT2R) | −9.0 | Plasma membrane Sensory neurons | Involved in cell proliferation and functional inhibition of ERK2 receptor [31]; involved in nociception and neuronal regeneration [1]. |
| Vesicle associated membrane protein 8 (VAMP8) | −5.7 | Vesicular, Secretory granules | Forms an extended parallel four alpha-helical trans-SNARE complex with STX11 and SNAP23 upon stimulation, causing membrane fusion and driving platelet exocytosis [17]. |
| Syntaxin 11 (STX11) | −6.0 | Plasma membrane | Interacts and binds selectively with SNAP23 and VAMP8 to form a complex during membrane fusion and they facilitate granule exocytosis [17]. |
| N-ethylmaleimide-sensitive factor attachment protein 23 (SNAP23) | −4.4 | Plasma membrane | Highly involved in membrane fusion regulation during granule exocytosis in platelet and mast cells. Usually binds to VAMP8 and STX4 in mast cells and VAMP8 and STX 11 in platelets during exocytosis [16,17]. |
| Munc13-4 (Isoform Munc13-1 was used for docking) | −6.2 | Highly localized on the plasma membrane, endosome, lysosome, and the cytoplasm | Involved in granule maturation, docking, and vesicle fusion: playing a major role in vesicle priming. They bind to STX4 in mast cells during degranulation process [16]. |
| Energy Terms | Munc18b-Bound Energy Values (kJ/mol) |
|---|---|
| van der Waal energy | −313.404 ± 35.505 |
| Electrostatic energy | −49.944 ± 28.281 |
| Polar solvation energy | 144.781 ± 43.370 |
| Nonpolar solvation energy | −29.005 ± 3.066 |
| Binding energy | −247.571 ± 37.471 |
| Molecular Interactions before MD Simulations | ||
| Protein Complex | H-Bond Residues | Hydrophobic Bonded Residues |
| Munc18b-Mycolactone complex | His245, Asp255, Leu256, Asp262, Arg402, Arg405, Thr574, and Arg575. | Lys7, Ser43, Leu138, Tyr140, Ser146, Tyr254, Gln261, Val361, Ile570, Thr572, and Asp578. |
| Molecular Interactions after MD Simulations | ||
| Protein Complex | H-Bond Residues | Hydrophobic Bonded Residues |
| Munc18b-Mycolactone complex | Val542, Leu571 | Tyr140, Val144, Pro242, Leu243,Leu244,His245,Ala251, Tyr254,Asp255,Leu256, Tyr401,Arg405, Gly541, Ser567, Ile570, Thr572, Pro573 |
| Residues | Energies <−5.0 kJ/mol | Residues | Energies >5.0 kJ/mol |
|---|---|---|---|
| Met544 | −5.0279 | Arg405 | 6.1118 |
| Glu546 | −5.1263 | Arg575 | 7.8640 |
| Glu246 | −5.5520 | Lys7 | 12.8054 |
| Glu260 | −6.2233 | ||
| Ile570 | −9.8204 | ||
| Glu141 | −10.1902 | ||
| Tyr140 | −11.0537 | ||
| Tyr254 | −13.2513 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kwofie, S.K.; Dankwa, B.; Enninful, K.S.; Adobor, C.; Broni, E.; Ntiamoah, A.; Wilson, M.D. Molecular Docking and Dynamics Simulation Studies Predict Munc18b as a Target of Mycolactone: A Plausible Mechanism for Granule Exocytosis Impairment in Buruli Ulcer Pathogenesis. Toxins 2019, 11, 181. https://doi.org/10.3390/toxins11030181
Kwofie SK, Dankwa B, Enninful KS, Adobor C, Broni E, Ntiamoah A, Wilson MD. Molecular Docking and Dynamics Simulation Studies Predict Munc18b as a Target of Mycolactone: A Plausible Mechanism for Granule Exocytosis Impairment in Buruli Ulcer Pathogenesis. Toxins. 2019; 11(3):181. https://doi.org/10.3390/toxins11030181
Chicago/Turabian StyleKwofie, Samuel K., Bismark Dankwa, Kweku S. Enninful, Courage Adobor, Emmanuel Broni, Alfred Ntiamoah, and Michael D. Wilson. 2019. "Molecular Docking and Dynamics Simulation Studies Predict Munc18b as a Target of Mycolactone: A Plausible Mechanism for Granule Exocytosis Impairment in Buruli Ulcer Pathogenesis" Toxins 11, no. 3: 181. https://doi.org/10.3390/toxins11030181
APA StyleKwofie, S. K., Dankwa, B., Enninful, K. S., Adobor, C., Broni, E., Ntiamoah, A., & Wilson, M. D. (2019). Molecular Docking and Dynamics Simulation Studies Predict Munc18b as a Target of Mycolactone: A Plausible Mechanism for Granule Exocytosis Impairment in Buruli Ulcer Pathogenesis. Toxins, 11(3), 181. https://doi.org/10.3390/toxins11030181






