Epidemiology of Shiga Toxin-Producing Escherichia coli O157 in the Province of Alberta, Canada, 2009–2016
Abstract
1. Introduction
2. Results
2.1. STEC O157 Isolate Characteristics
2.2. Outcomes of STEC O157 Infection and Demographic Characteristics of Cases
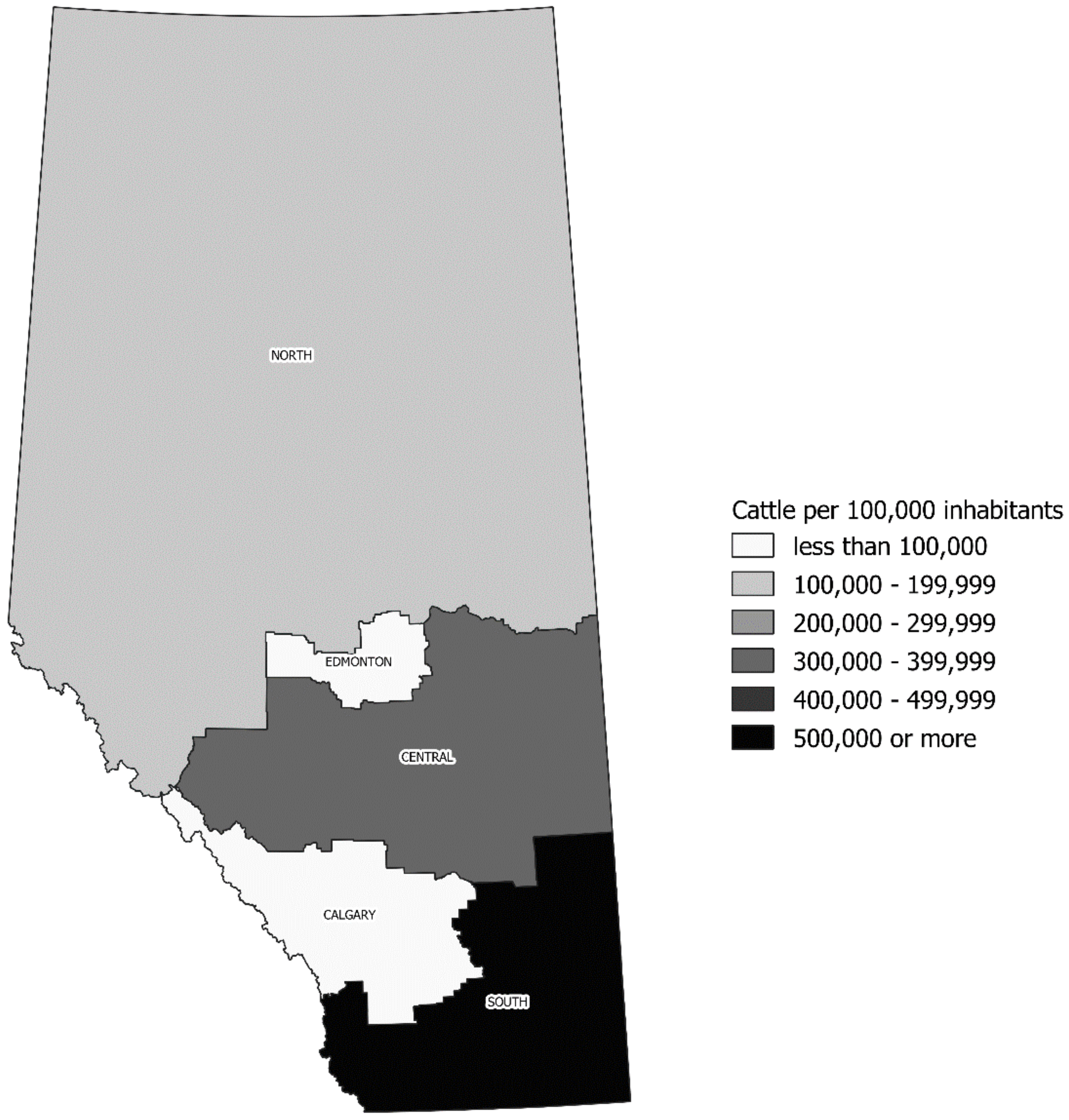
2.3. Seasonality of STEC O157 Infection and Its Geographic Distribution
2.4. Predictors of STEC O157 Morbidity
3. Discussion
4. Materials and Methods
4.1. Setting
4.2. Data Collection
4.3. Case Definitions and Inclusion Criteria
4.4. Virulence Gene Detection and Subtyping
4.5. Bacterial Typing and Clustering Analysis
4.6. Geographical Analysis
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Majowicz, S.E.; Scallan, E.; Jones-Bitton, A.; Sargeant, J.M.; Stapleton, J.; Angulo, F.J.; Yeung, D.H.; Kirk, M.D. Global incidence of human shiga toxin-producing Escherichia coli infections and deaths: A systematic review and knowledge synthesis. Foodborne Pathog. Dis. 2014, 11, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.K.; Murray, R.; Flockhart, L.; Pintar, K.; Fazil, A.; Nesbitt, A.; Marshall, B.; Tataryn, J.; Pollari, F. Estimates of Foodborne Illness-Related Hospitalizations and Deaths in Canada for 30 Specified Pathogens and Unspecified Agents. Foodborne Pathog. Dis. 2015, 12, 820–827. [Google Scholar] [CrossRef] [PubMed]
- Sockett, P.; Goebel, S.E.; Varela, N.P.; Guthrie, A.; Wilson, J.; Guilbault, L.A.; Clark, W.F. Verotoxigenic Escherichia coli: Costs of illness in Canada, including long-term health outcomes. J. Food Prot. 2014, 77, 216–226. [Google Scholar] [CrossRef]
- Statistics Canada Alberta Has the Most Beef Cattle in Canada and the Second Largest Total Farm Area. Available online: http://www.statcan.gc.ca/pub/95-640-x/2016001/article/14808-eng.htm (accessed on 20 January 2018).
- Waters, J.R.; Chaowagul, J.C.M.; Dev, V.J. Infection caused by Escherichia coli O157:H7 in Alberta, Canada, and in Scotland: A five-year review, 1987-1991. Clin. Infect. Dis. 1994, 19, 834–843. [Google Scholar] [CrossRef] [PubMed]
- Scheutz, F.; Teel, L.D.; Beutin, L.; Piérard, D.; Buvens, G.; Karch, H.; Mellmann, A.; Caprioli, A.; Tozzoli, R.; Morabito, S.; et al. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J. Clin. Microbiol. 2012, 50, 2951–2963. [Google Scholar] [CrossRef] [PubMed]
- Honish, L.; Punja, N.; Nunn, S.; Nelson, D.; Hislop, N.; Gosselin, G.; Stashko, N.; Dittrich, D. Escherichia coli O157:H7 infections associated with contaminated pork products—Alberta, Canada, July–October 2014. Morb. Mortal. Wkly. Rep. 2017, 65, 1477–1481. [Google Scholar] [CrossRef] [PubMed]
- Freedman, S.B.; Xie, J.; Neufeld, M.S.; Hamilton, W.L.; Hartling, L.; Tarr, P.I. Shiga toxin-producing Escherichia coli infection, antibiotics, and risk of developing hemolytic uremic syndrome: A meta-analysis. Clin. Infect. Dis. 2016, 62, 1251–1258. [Google Scholar] [CrossRef]
- Wong, C.S.; Mooney, J.C.; Brandt, J.R.; Staples, A.O.; Jelacic, S.; Boster, D.R.; Watkins, S.L.; Tarr, P.I. Risk factors for the hemolytic uremic syndrome in children infected with Escherichia coli O157:H7: A multivariable analysis. Clin. Infect. Dis. 2012, 55, 33–41. [Google Scholar] [CrossRef]
- Freedman, S.B.; Eltorki, M.; Chui, L.; Xie, J.; Feng, S.; MacDonald, J.; Dixon, A.; Ali, S.; Louie, M.; Lee, B.E.; et al. Province-Wide Review of Pediatric Shiga Toxin-Producing Escherichia coli Case Management. J. Pediatr. 2017, 180, 184–190. [Google Scholar] [CrossRef]
- Ardissino, G.; Tel, F.; Possenti, I.; Testa, S.; Consonni, D.; Paglialonga, F.; Salardi, S.; Borsa-Ghiringhelli, N.; Salice, P.; Tedeschi, S.; et al. Early volume expansion and outcomes of hemolytic uremic syndrome. Pediatrics 2016, 137. [Google Scholar] [CrossRef]
- Berenger, B.; Chui, L.; Reimer, A.; Allen, V.; Alexander, D.; Domingo, M.-C.; Haldane, D.; Hoang, L.; Levett, P.; MacKeen, A.; et al. Canadian Public Health Laboratory Network position statement: Non-Culture based diagnostics for gastroenteritis and implications for public health investigations. Can. Commun. Dis. Rep. 2017, 43, 279–281. [Google Scholar] [CrossRef] [PubMed]
- Karmali, M.A. Host and pathogen determinants of verocytotoxin-producing Escherichia coli-associated hemolytic uremic syndrome. Kidney Int. 2009, 75, S4–S7. [Google Scholar] [CrossRef] [PubMed]
- Fuller, C.A.; Pellino, C.A.; Flagler, M.J.; Strasser, J.E.; Weiss, A.A. Shiga toxin subtypes display dramatic differences in potency. Infect. Immun. 2011, 79, 1329–1337. [Google Scholar] [CrossRef] [PubMed]
- Bunger, J.C.; Melton-Celsa, A.R.; Maynard, E.L.; O’Brien, A.D. Reduced toxicity of Shiga toxin (Stx) type 2c in mice compared to Stx2d is associated with instability of Stx2c holotoxin. Toxins 2015, 7, 2306–2320. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Zurek, L. Evaluation of the anti-terminator Q933 gene as a marker for Escherichia coli O157:H7 with high Shiga toxin production. Curr. Microbiol. 2006, 53, 324–328. [Google Scholar] [CrossRef]
- Taylor, E.V.; Shi, X.; Alam, M.J.; Peterson, G.; Narayanan, S.K.; Renter, D.G.; Nagaraja, T.G. Genetic Variations in Shiga Toxin-Producing Abilities of Bovine and Human Escherichia coli O157:H7. Zoonoses Public Health 2011, 58, 185–191. [Google Scholar] [CrossRef]
- Kawano, K.; Okada, M.; Haga, T.; Maeda, K.; Goto, Y. Relationship between pathogenicity for humans and stx genotype in Shiga toxin-producing Escherichia coli serotype O157. Eur. J. Clin. Microbiol. Infect. Dis. 2008, 27, 227–232. [Google Scholar] [CrossRef]
- Kawano, K.; Ono, H.; Iwashita, O.; Kurogi, M.; Haga, T.; Maeda, K.; Goto, Y. Relationship between stx genotype and stx2 expression level in Shiga toxin-producing Escherichia coli o157 strains. Jpn. J. Infect. Dis. 2012, 65, 322–325. [Google Scholar] [CrossRef][Green Version]
- Strauch, E.; Schaudinn, C.; Beutin, L. First-time isolation and characterization of a bacteriophage encoding the Shiga toxin 2c variant, which is globally spread in strains of Escherichia coli O157. Infect. Immun. 2004, 72, 7030–7039. [Google Scholar] [CrossRef]
- Strauch, E.; Hammerl, J.A.; Konietzny, A.; Schneiker-Bekel, S.; Arnold, W.; Goesmann, A.; Pühler, A.; Beutin, L. Bacteriophage 2851 is a prototype phage for dissemination of the Shiga toxin variant gene 2c in Escherichia coli O157:H7. Infect. Immun. 2008, 76, 5466–5477. [Google Scholar] [CrossRef]
- Tarr, G.A.M.; Shringi, S.; Oltean, H.N.; Mayer, J.; Rabinowitz, P.; Wakefield, J.; Tarr, P.I.; Besser, T.E.; Phipps, A.I. Importance of case age in the purported association between phylogenetics and hemolytic uremic syndrome in Escherichia coli O157:H7 infections. Epidemiol. Infect. 2018, 146, 1550–1555. [Google Scholar] [CrossRef] [PubMed]
- Ziebell, K.; Steele, M.; Zhang, Y.; Benson, A.; Taboada, E.N.; Laing, C.; McEwen, S.; Ciebin, B.; Johnson, R.; Gannon, V. Genotypic characterization and prevalence of virulence factors among Canadian Escherichia coli O157:H7 strains. Appl. Environ. Microbiol. 2008, 74, 4314–4323. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Laing, C.R.; Buchanan, C.; Taboada, E.N.; Zhang, Y.; Karmali, M.A.; Thomas, J.E.; Gannon, V.P. In silico genomic analyses reveal three distinct lineages of Escherichia coli O157:H7, one of which is associated with hyper-virulence. BMC Genom. 2009, 10, 287. [Google Scholar] [CrossRef] [PubMed]
- Manning, S.D.; Motiwala, A.S.; Springman, A.C.; Qi, W.; Lacher, D.W.; Ouellette, L.M.; Mladonicky, J.M.; Somsel, P.; Rudrik, J.T.; Dietrich, S.E.; et al. Variation in virulence among clades of Escherichia coli O157:H7 associated with disease outbreaks. Proc. Natl. Acad. Sci. USA 2008, 105, 4868–4873. [Google Scholar] [CrossRef]
- Iyoda, S.; Manning, S.D.; Seto, K.; Kimata, K.; Isobe, J.; Etoh, Y.; Ichihara, S.; Migita, Y.; Ogata, K.; Honda, M.; et al. Phylogenetic clades 6 and 8 of enterohemorrhagic Escherichia coli O157: H7 With particular stx subtypes are more frequently found in isolates from hemolytic uremic syndrome patients than from asymptomatic carriers. Open Forum Infect. Dis. 2014, 1, ofu061. [Google Scholar] [CrossRef]
- Byrne, L.; Dallman, T.J.; Adams, N.; Mikhail, A.F.W.; McCarthy, N.; Jenkins, C. Highly pathogenic clone of shiga toxin-producing escherichia coli O157:H7, England and Wales. Emerg. Infect. Dis. 2018, 24, 2303. [Google Scholar] [CrossRef]
- Brandal, L.T.; Wester, A.L.; Lange, H.; Løbersli, I.; Lindstedt, B.A.; Vold, L.; Kapperud, G. Shiga toxin-producing escherichia coli infections in Norway, 1992-2012: Characterization of isolates and identification of risk factors for haemolytic uremic syndrome. BMC Infect. Dis. 2015, 15, 324. [Google Scholar] [CrossRef]
- Gould, L.H.; Demma, L.; Jones, T.F.; Hurd, S.; Vugia, D.J.; Smith, K.; Shiferaw, B.; Segler, S.; Palmer, A.; Zansky, S.; et al. Hemolytic Uremic Syndrome and Death in Persons with Escherichia coli O157:H7 Infection, Foodborne Diseases Active Surveillance Network Sites, 2000–2006. Clin. Infect. Dis. 2009, 49, 1480–1485. [Google Scholar] [CrossRef]
- Sharma, R.; Stanford, K.; Louie, M.; Munns, K.; John, S.J.; Zhang, Y.; Gannon, V.; Chui, L.; Read, R.; Topp, E.; et al. Escherichia coli O157:H7 lineages in healthy beef and dairy cattle and clinical human cases in Alberta, Canada. J. Food Prot. 2009, 72, 601–607. [Google Scholar] [CrossRef]
- Tostes, R.; Goji, N.; Amoako, K.; Chui, L.; Kastelic, J.; Devinney, R.; Stanford, K.; Reuter, T. Subtyping Escherichia coli Virulence Genes Isolated from Feces of Beef Cattle and Clinical Cases in Alberta. Foodborne Pathog. Dis. 2017, 14, 35–42. [Google Scholar] [CrossRef]
- Aslam, M.; Stanford, K.; McAllister, T.A. Characterization of antimicrobial resistance and seasonal prevalence of Escherichia coli O157:H7 recovered from commercial feedlots in Alberta, Canada. Lett. Appl. Microbiol. 2010, 50, 320–326. [Google Scholar] [CrossRef]
- Stanford, K.; Reuter, T.; Bach, S.J.; Chui, L.; Ma, A.; Conrad, C.C.; Tostes, R.; McAllister, T.A. Effect of severe weather events on the shedding of Shiga toxigenic Escherichia coli in slaughter cattle and phenotype of serogroup O157 isolates. FEMS Microbiol. Ecol. 2017, 93. [Google Scholar] [CrossRef]
- Rivero, M.A.; Passucci, J.A.; Rodríguez, E.M.; Parma, A.E. Seasonal variation of HUS occurrence and VTEC infection in children with acute diarrhoea from Argentina. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 1131–1135. [Google Scholar] [CrossRef] [PubMed]
- Vally, H.; Hall, G.; Dyda, A.; Raupach, J.; Knope, K.; Combs, B.; Desmarchelier, P. Epidemiology of Shiga toxin producing Escherichia coli in Australia, 2000-2010. BMC Public Health 2012, 12, 63. [Google Scholar] [CrossRef] [PubMed]
- Bifolchi, N.; Michel, P.; Talbot, J.; Svenson, L.; Simmonds, K.; Checkley, S.; Chui, L.; Dick, P.; Wilson, J.B. Weather and livestock risk factors for Escherichia coli O157 human infection in Alberta, Canada. Epidemiol. Infect. 2014, 142, 2302–2313. [Google Scholar] [CrossRef] [PubMed]
- Pearl, D.L.; Louie, M.; Chui, L.; Doré, K.; Grimsrud, K.M.; Martin, S.W.; Michel, P.; Svenson, L.W.; McEwen, S.A. A multi-level approach for investigating socio-economic and agricultural risk factors associated with rates of reported cases of escherichia coli O157 in humans in Alberta, Canada. Zoonoses Public Health 2009, 56, 455–464. [Google Scholar] [CrossRef]
- Jokinen, C.; Edge, T.A.; Ho, S.; Koning, W.; Laing, C.; Mauro, W.; Medeiros, D.; Miller, J.; Robertson, W.; Taboada, E.; et al. Molecular subtypes of Campylobacter spp., Salmonella enterica, and Escherichia coli O157:H7 isolated from faecal and surface water samples in the Oldman River watershed, Alberta, Canada. Water Res. 2011, 45, 1247–1257. [Google Scholar] [CrossRef]
- Gannon, V.P.J.; Graham, T.A.; Read, S.; Ziebell, K.; Muckle, A.; Mori, J.; Thomas, J.; Selinger, B.; Townshend, I.; Byrne, J. Bacterial pathogens in rural water supplies in Southern Alberta, Canada. J. Toxicol. Environ. Health Part A 2004, 67, 1643–1653. [Google Scholar] [CrossRef]
- Gould, L.H.; Bopp, C.; Strockbine, N.; Atkinson, R.; Baselski, V.; Body, B.; Carey, R.; Crandall, C.; Hurd, S.; Kaplan, R.; et al. Recommendations for diagnosis of shiga toxin-producing Escherichia coli infections by clinical laboratories. MMWR Recomm. Rep. 2009, 58, 1–14. [Google Scholar] [CrossRef]
- Government of Alberta Notifiable Disease Report (NDR) Manual. Available online: https://open.alberta.ca/publications/ndr-manual-9th-edition/ (accessed on 10 October 2018).
- Holland, J.L.; Louie, L.; Simor, A.E.; Louie, M. PCR detection of Escherichia coli O157:H7 directly from stools: Evaluation of commercial extraction methods for purifying fecal DNA. J. Clin. Microbiol. 2000, 38, 4108–4113. [Google Scholar]
- Nielsen, E.M.; Andersen, M.T. Detection and characterization of verocytotoxin-producing Escherichia coli by automated 5′ nuclease PCR assay. J. Clin. Microbiol. 2003, 41, 2884–2893. [Google Scholar] [CrossRef] [PubMed]
- Chui, L.; Lee, M.C.; Allen, R.; Bryks, A.; Haines, L.; Boras, V. Comparison between ImmunoCard STAT!® and real-time PCR as screening tools for both O157: H7 and non-O157 Shiga toxin-producing Escherichia coli in Southern Alberta, Canada. Diagn. Microbiol. Infect. Dis. 2013, 77, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Chui, L.; Li, V.; Fach, P.; Delannoy, S.; Malejczyk, K.; Patterson-Fortin, L.; Poon, A.; King, R.; Simmonds, K.; Scott, A.N.; et al. Molecular profiling of Escherichia coli O157:H7 and non-O157 strains isolated from humans and cattle in Alberta, Canada. J. Clin. Microbiol. 2015, 53, 986–990. [Google Scholar] [CrossRef] [PubMed]
- Zhi, S.; Szelewicki, J.; Ziebell, K.; Parsons, B.; Chui, L. General detection of Shiga toxin 2 and subtyping of Shiga toxin 1 and 2 in Escherichia coli using qPCR. J. Microbiol. Methods 2019, 159, 51–55. [Google Scholar] [CrossRef]
- PulseNet International. PFGE Protocols. Available online: http://www.pulsenetinternational.org/protocols/ (accessed on 10 October 2018).
- PulseNet International. MLVA Protocols. Available online: http://www.pulsenetinternational.org/protocols/mlva/ (accessed on 10 October 2018).
- QGIS Development Team (2017). QGIS Geographic Information System; Open Source Geospacial Foundation Project. Available online: http://qgis.osgeo.org. (accessed on 10 October 2018).
- Government of Alberta Geographic and Geospatial Statistics. Available online: https://www.alberta.ca/geographic-geospatial-statistics.aspx (accessed on 10 October 2018).
- Government of Alberta Open Government Program. Available online: https://open.alberta.ca/opendata (accessed on 10 October 2018).
- Government of Alberta Interactive Health Data Application. Available online: http://www.ahw.gov.ab.ca/IDHA_Retrieval/ (accessed on 10 October 2018).
| Variable | With HUS | Without HUS | Univariate OR (CI 95%) | p-Value | Multivariate OR (CI 95%) | p-Value |
|---|---|---|---|---|---|---|
| Sex | 0.793 | |||||
| Female | 21 (5.2) | 383 (94.8) | 0.917 (0.480–1.752) | |||
| Male | 18 (5.6) | 301 (94.4) | 1.091 (0.571–2.084) | |||
| Age group | ||||||
| <5 | 21 (14.6) | 123 (85.4) | 5.319 (2.75–10.309) | <0.001 | 9.311 (3.588–24.161) | <0.001 |
| 5–9 | 10 (10.3) | 87 (89.7) | 2.364 (1.11–5.025) | 0.021 | 5.208 (1.791–15.143) | 0.002 |
| 10–19 | 2 (2.1) | 95 (97.9) | 0.335 (0.079–1.414) | 0.148 | ||
| 20–29 | 2 (1.3) | 155 (98.7) | 0.184 (0.044–0.774) | 0.008 | 0.776 (0.153–3.942) | 0.76 |
| 30–39 | 0 | 50 (100.0) | 0.102 | |||
| 40–49 | 0 | 40 (100.0) | 0.159 | |||
| 50–59 | 2 (3.5) | 55 (96.5) | 0.618 (0.145–2.632) | 0.511 | ||
| ≥60 | 2 (2.5) | 79 (97.5) | 0.414 (0.098–1.751) | 0.299 | ||
| Season | ||||||
| Jan–Mar | 5 (5.9) | 80 (94.1) | 1.110 (0.422–2.924) | 0.798 | ||
| Apr–Jun | 12 (10.2) | 106 (89.8) | 2.421 (1.190–4.926) | 0.012 | 2.314 (1.066–5.024) | 0.034 |
| Jul–Sep | 21 (5.2) | 385 (94.8) | 0.906 (0.474–1.730) | 0.765 | ||
| Oct–Dec | 1 (0.9) | 113 (99.1) | 0.133 (0.018–0.978) | 0.02 | 0.191 (0.025–1.454) | 0.11 |
| Shiga toxin genotype | ||||||
| stx1a stx2a | 20 (3.8) | 504 (96.2) | 0.376 (0.196–0.720) | 0.002 | 0.466 (0.198–1.097) | 0.08 |
| stx1a stx2a stx2c | 0 | 12 (100) | 1 | |||
| stx1a stx2c | 4 (25.0) | 12 (75.0) | 6.410 (1.965–20.833) | <0.001 | 2.679 (0.636–11.277) | 0.179 |
| stx2a | 7 (7.4) | 88 (92.6) | 1.481 (0.635–3.460) | 0.361 | ||
| stx2a stx2c | 6 (12.5) | 42 (87.5) | 2.778 (1.103–6.993) | 0.024 | 1.657 (0.502–5.465) | 0.407 |
| stx2c | 2 (7.1) | 26 (92.9) | 1.368 (0.313–5.988) | 0.659 | ||
| Isolate relatedness | 0.933 | |||||
| Outbreak-related | 21 (5.3) | 373 (94.7) | 0.973 (0.509–1.858) | |||
| Sporadic | 18 (5.5) | 311 (94.5) | 1.028 (0.538–1.965) |
| Variable | Hospitalized | Not Hospitalized | Univariate OR (CI 95%) | p-Value | Multivariate OR (CI 95%) | p-Value |
|---|---|---|---|---|---|---|
| Sex | 0.217 | |||||
| Female | 114 (28.2) | 290 (71.8) | 1.235 (0.883–1.728) | |||
| Male | 77 (24.1) | 242 (75.9) | 0.809 (0.579–1.132) | |||
| Age group | ||||||
| <5 | 39 (27.1) | 105 (72.9) | 1.044 (0.692–1.575) | 0.84 | ||
| 5–9 | 30 (30.9) | 67 (69.1) | 1.294 (0.811–2.062) | 0.279 | ||
| 10–19 | 19 (19.6) | 78 (80.4) | 0.643 (0.378–1.094) | 0.101 | ||
| 20–29 | 32 (20.4) | 125 (79.6) | 0.655 (0.427–1.007) | 0.053 | ||
| 30–39 | 7 (14.0) | 43 (86.0) | 0.433 (0.191–0.979) | 0.039 | 0.445 (0.194–1.021) | 0.056 |
| 40–49 | 4 (10.0) | 36 (90.0) | 0.295 (0.103–0.840) | 0.015 | 0.328 (0.114–0.943) | 0.039 |
| 50–59 | 19 (33.3) | 38 (66.7) | 1.437 (0.806–2.558) | 0.217 | ||
| ≥60 | 41 (50.6) | 40 (49.4) | 3.367 (2.096–5.405) | <0.001 | 3.054 (1.879–4.961) | <0.001 |
| Season | ||||||
| Jan–Mar | 20 (23.5) | 65 (76.5) | 0.840 (0.494–1.429) | 0.52 | ||
| Apr–Jun | 41 (34.7) | 77 (65.3) | 1.616 (1.059–2.463) | 0.025 | 1.795 (1.157–2.786) | 0.009 |
| Jul–Sep | 101 (24.9) | 305 (75.1) | 0.835 (0.599–1.164) | 0.288 | ||
| Oct–Dec | 29 (25.4) | 85 (74.6) | 0.942 (0.595–1.488) | 0.796 | ||
| Shiga toxin genotype | ||||||
| stx1a stx2a | 140 (26.7) | 384 (73.3) | 1.058 (0.729–1.536) | 0.767 | ||
| stx1a stx2a stx2c | 1 (8.3) | 11 (91.7) | 0.249 (0.032–1.946) | 0.199 | ||
| stx1a stx2c | 5 (31.3) | 11 (68.8) | 1.274 (0.437–3.717) | 0.774 | ||
| stx2a | 21 (22.1) | 74 (77.9) | 0.765 (0.457–1.280) | 0.306 | ||
| stx2a stx2c | 19 (39.6) | 29 (60.4) | 1.916 (1.047–3.509) | 0.032 | 1.878 (1.006–3.505) | 0.048 |
| stx2c | 5 (17.9) | 23 (82.1) | 0.595 (0.223–1.587) | 0.295 | ||
| Isolate relatedness | 0.027 | 0.158 | ||||
| Outbreak-related | 91 (23.1) | 303 (76.9) | 0.688 (0.494–0.958) | 0.780 (0.552–1.101) | ||
| Sporadic | 100 (30.4) | 229 (69.6) | 1.454 (1.043–2.026) | 1.282 (0.908–1.811) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lisboa, L.F.; Szelewicki, J.; Lin, A.; Latonas, S.; Li, V.; Zhi, S.; Parsons, B.D.; Berenger, B.; Fathima, S.; Chui, L. Epidemiology of Shiga Toxin-Producing Escherichia coli O157 in the Province of Alberta, Canada, 2009–2016. Toxins 2019, 11, 613. https://doi.org/10.3390/toxins11100613
Lisboa LF, Szelewicki J, Lin A, Latonas S, Li V, Zhi S, Parsons BD, Berenger B, Fathima S, Chui L. Epidemiology of Shiga Toxin-Producing Escherichia coli O157 in the Province of Alberta, Canada, 2009–2016. Toxins. 2019; 11(10):613. https://doi.org/10.3390/toxins11100613
Chicago/Turabian StyleLisboa, Luiz F., Jonas Szelewicki, Alex Lin, Sarah Latonas, Vincent Li, Shuai Zhi, Brendon D. Parsons, Byron Berenger, Sumana Fathima, and Linda Chui. 2019. "Epidemiology of Shiga Toxin-Producing Escherichia coli O157 in the Province of Alberta, Canada, 2009–2016" Toxins 11, no. 10: 613. https://doi.org/10.3390/toxins11100613
APA StyleLisboa, L. F., Szelewicki, J., Lin, A., Latonas, S., Li, V., Zhi, S., Parsons, B. D., Berenger, B., Fathima, S., & Chui, L. (2019). Epidemiology of Shiga Toxin-Producing Escherichia coli O157 in the Province of Alberta, Canada, 2009–2016. Toxins, 11(10), 613. https://doi.org/10.3390/toxins11100613




