Analysis of the Fungal Community in Ziziphi Spinosae Semen through High-Throughput Sequencing
Abstract
1. Introduction
2. Results
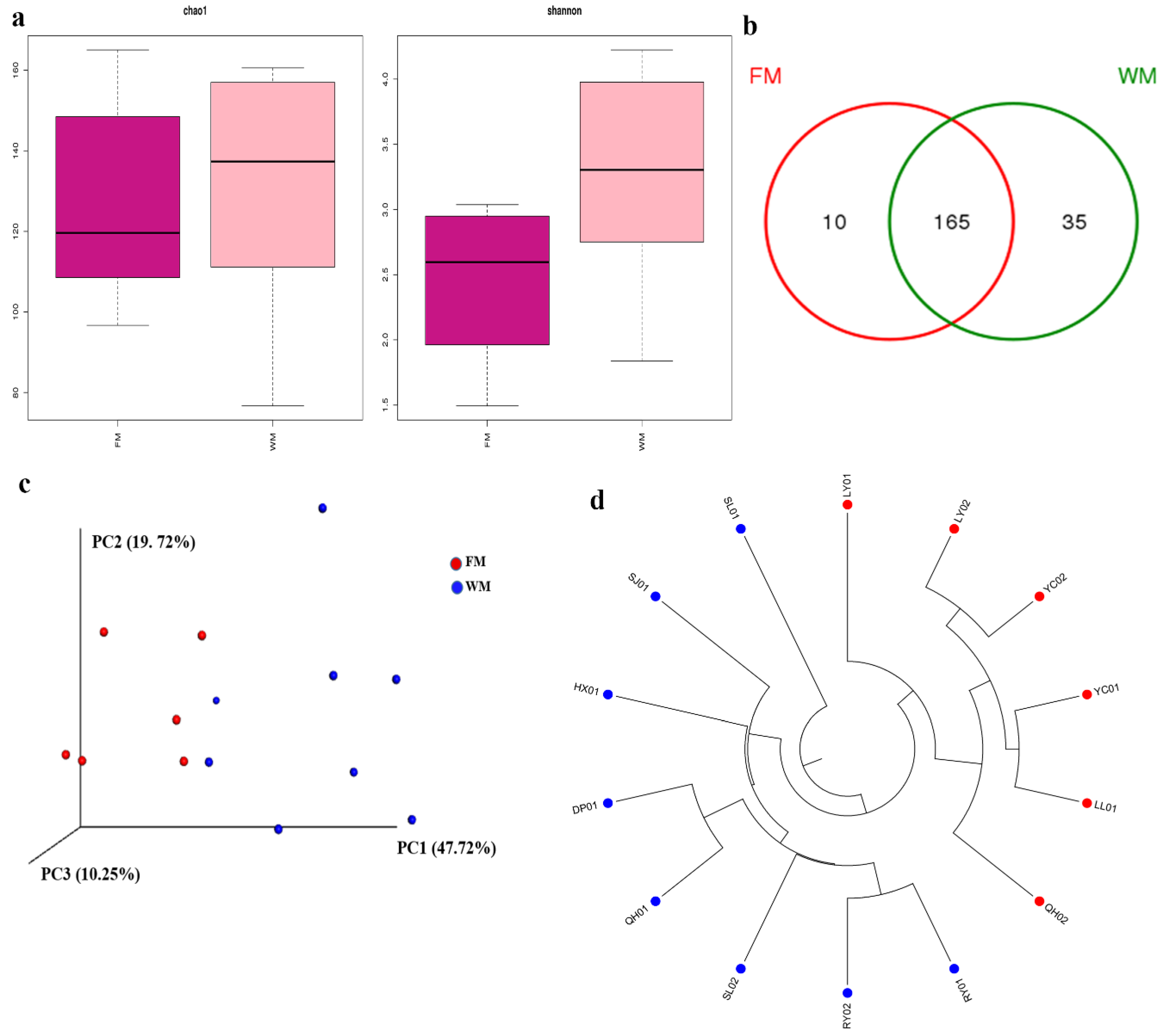
2.1. Analyses of the Diversity of Fungal Communities in the ZSS Samples
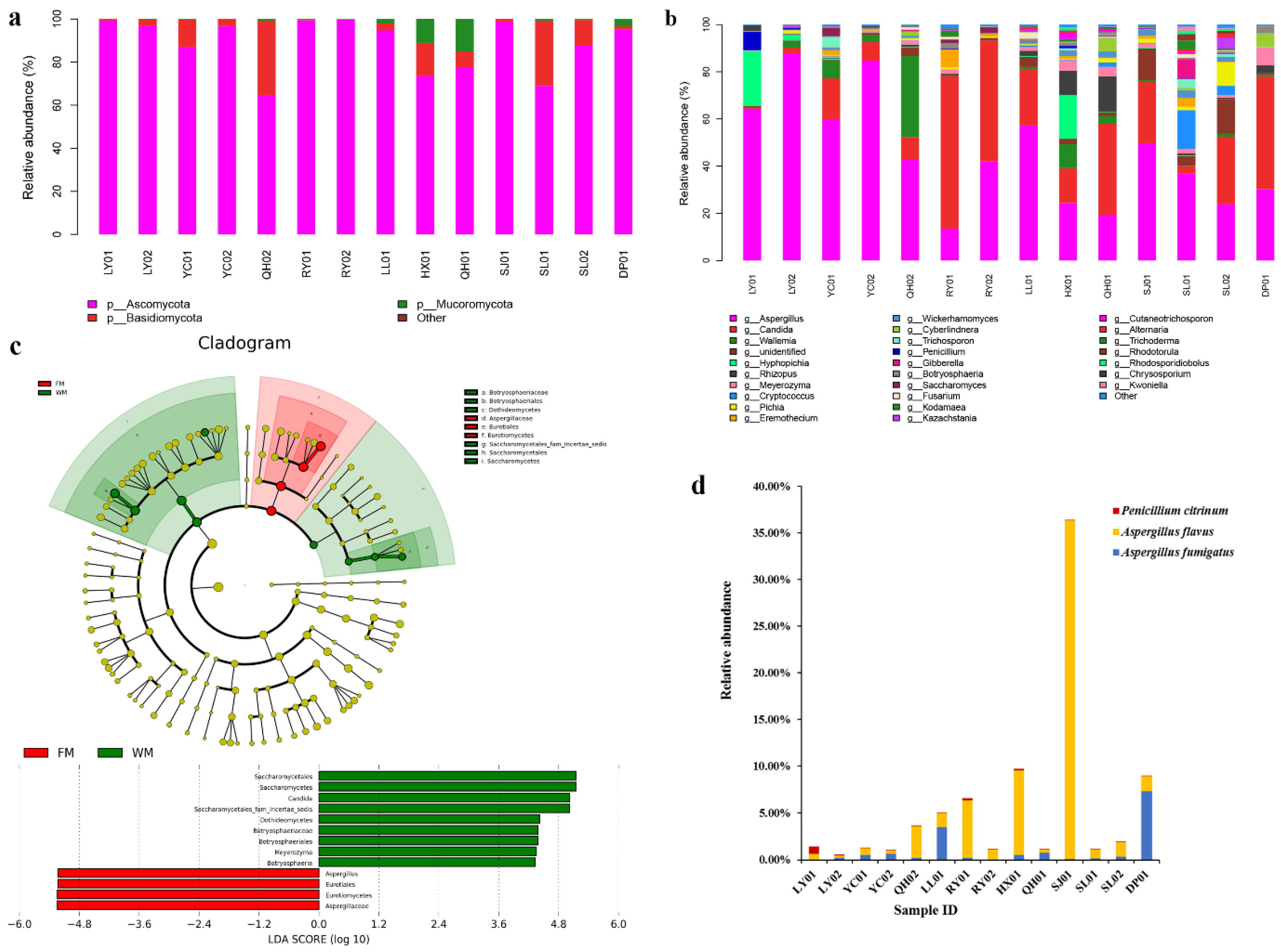
2.2. Fungal Community Composition in the ZSS Samples
3. Discussion
3.1. Prevalence of Fungal Contamination in the Commercial ZSS Samples
3.2. DNA Marker Selection to Analyze the Fungal Community in ZSS
3.3. Prospects of Applying Amplicon Sequencing for the Analysis of Fungal Diversity in Herbal Materials
4. Materials and Methods
4.1. Sampling
4.2. DNA Extraction
4.3. PCR Amplification and HTS
4.4. Sequence Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Moreira, D.D.L.; Teixeira, S.S.; Monteiro, M.H.D.; De-Oliveira, A.C.A.; Paumgartten, F.J. Traditional use and safety of herbal medicines. Rev. Bras. Farmacogn. 2014, 24, 248–257. [Google Scholar] [CrossRef]
- Shim, W.B.; Kim, K.; Ofori, J.A.; Chung, Y.C.; Chung, D.H. Occurrence of aflatoxins in herbal medicine distributed in South Korea. J. Food Prot. 2012, 75, 1991–1999. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, B.; Ashiq, S.; Hussain, A.; Bashir, S.; Hussain, M. Evaluation of mycotoxins, mycobiota, and toxigenic fungi in selected medicinal plants of Khyber Pakhtunkhwa, Pakistan. Fungal Biol. 2014, 118, 776–784. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Wei, R.; Logrieco, A.F.; Wei, J.; Wen, J.; Xiao, X.; Yang, M. Occurrence of toxigenic fungi and determination of mycotoxins by HPLC-FLD in functional foods and spices in China markets. Food Chem. 2014, 146, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Wei, R.; Qiu, F.; Kong, W.; Wei, J.; Yang, M.; Luo, Z.; Qin, J.; Ma, X. Co-occurrence of aflatoxin B1, B2, G1, G2 and ochrotoxin A in Glycyrrhiza uralensis analyzed by HPLC-MS/MS. Food Control 2013, 32, 216–221. [Google Scholar] [CrossRef]
- Shim, W.B.; Ha, K.S.; Kim, M.G.; Kim, J.S.; Chung, D.H. Evaluation of the transfer rate of ochratoxin A to decoctions of herbal medicines. Food Sci. Biotechnol. 2014, 23, 2103–2108. [Google Scholar] [CrossRef]
- Nian, Y.; Wang, H.; Ying, G.; Yang, M.; Wang, Z.; Kong, W.; Yang, S. Transfer rates of aflatoxins from herbal medicines to decoctions determined by an optimized high-performance liquid chromatography with fluorescence detection method. J. Pharm. Pharmacol. 2018, 70, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Roberts, S.M.; James, R.C.; Williams, P.L. Principles of Toxicology: Environmental and Industrial Applications; John Wiley & Sons: Hoboken, NJ, USA, 2015. [Google Scholar]
- Marin, S.; Ramos, A.J.; Cano-Sancho, G.; Sanchis, V. Mycotoxins: Occurrence, toxicology, and exposure assessment. Food Chem. Toxicol. 2013, 60, 218–237. [Google Scholar] [CrossRef] [PubMed]
- Prouillac, C.; Koraichi, F.; Videmann, B.; Mazallon, M.; Rodriguez, F.; Baltas, M.; Lecoeur, S. In vitro toxicological effects of estrogenic mycotoxins on human placental cells: Structure activity relationships. Toxicol. Appl. Pharmacol. 2012, 259, 366–375. [Google Scholar] [CrossRef] [PubMed]
- Zinedine, A.; Mañes, J. Occurrence and legislation of mycotoxins in food and feed from Morocco. Food Control 2009, 20, 334–344. [Google Scholar] [CrossRef]
- Ashiq, S. Natural occurrence of mycotoxins in food and feed: Pakistan perspective. Compr. Rev. Food Sci. Food Saf. 2015, 14, 159–175. [Google Scholar] [CrossRef]
- El Darra, N.; Gambacorta, L.; Solfrizzo, M. Multimycotoxins occurrence in spices and herbs commercialized in Lebanon. Food Control 2019, 95, 63–70. [Google Scholar] [CrossRef]
- IARC. Some naturally occurring substances: Food items and constituents, heterocyclic aromatic amines and mycotoxins. In IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; International Agency for Research on Cancer: Lyon, France, 1993; Volume 56. [Google Scholar]
- Chawanthayatham, S.; Valentine, C.C.; Fedeles, B.I.; Fox, E.J.; Loeb, L.A.; Levine, S.S.; Slocum, S.L.; Wogan, G.N.; Croy, R.G.; Essigmann, J.M. Mutational spectra of aflatoxin B1 in vivo establish biomarkers of exposure for human hepatocellular carcinoma. Proc. Natl. Acad. Sci. USA 2017, 114, E3101–E3109. [Google Scholar] [CrossRef] [PubMed]
- Niessen, L. PCR-based diagnosis and quantification of mycotoxin producing fungi. Int. J. Food Microbiol. 2007, 119, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Srivastava, B.; Kumar, A.; Dubey, N.K. Fungal contamination of raw materials of some herbal drugs and recommendation of Cinnamomum camphora oil as herbal fungitoxicant. Microb. Ecol. 2008, 56, 555–560. [Google Scholar] [CrossRef] [PubMed]
- De Curtis, F.; De Felice, D.V.; Ianiri, G.; De Cicco, V.; Castoria, R. Environmental factors affect the activity of biocontrol agents against ochratoxigenic Aspergillus carbonarius on wine grape. Int. J. Food Microbiol. 2012, 159, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Jalili, M.; Jinap, S. Natural occurrence of aflatoxins and ochratoxin A in commercial dried chili. Food Control 2012, 24, 160–164. [Google Scholar] [CrossRef]
- Romagnoli, B.; Menna, V.; Gruppion, N.; Bergamini, C. Afatoxins in spices, aromatic herbs, herb-teas and medicinal plants marketed in Italy. Food Control 2007, 18, 697–701. [Google Scholar] [CrossRef]
- Stevic, T.; Pavlovic, S.; Stankovic, S.; Savikin, K. Pathogenic microorganisms of medicinal herbal drugs. Arch. Biol. Sci. 2012, 64, 49–58. [Google Scholar] [CrossRef]
- Niessen, L.; Bechtner, J.; Fodil, S.; Taniwaki, M.H.; Vogel, R.F. LAMP-based group specific detection of aflatoxin producers within Aspergillus section Flavi in food raw materials, spices, and dried fruit using neutral red for visible-light signal detection. Int. J. Food Microbiol. 2018, 266, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Duarte, S.C.; Pena, A.; Lino, C.M. A review on ochratoxin A occurrence and effects of processing of cereal and cereal derived food products. Food Microbiol. 2010, 27, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Chinese Pharmacopeia Commission. Pharmacopoeia of People’s Republic of China (Part 1); Press of Chemical Industry: Beijing, China, 2015; pp. 366–367. [Google Scholar]
- Fang, X.S.; Hao, J.F.; Zhou, H.Y.; Zhu, L.X.; Wang, J.H.; Song, F.Q. Pharmacological studies on the sedative-hypnotic effect of Semen Ziziphi spinosae (Suanzaoren) and Radix et Rhizoma Salviae miltiorrhizae (Danshen) extracts and the synergistic effect of their combinations. Phytomedicine 2010, 17, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Guo, L.; Pang, G.; Wu, X.; Zhang, M. Modulation effect of Semen Ziziphi Spinosae extracts on IL-1β, IL-4, IL-6, IL-10, TNF-α and IFN-γ in mouse serum. Nat. Prod. Res. 2011, 25, 464–467. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Qiao, L.; Song, M.; Wang, L.; Xie, J.; Feng, H. Hplc-ESI-MS/MS analysis of the water-soluble extract from Ziziphi spinosae semen and its ameliorating effect of learning and memory performance in mice. Pharmacogn. Mag. 2014, 10, 509. [Google Scholar] [PubMed]
- Chen, A.J.; Huang, L.F.; Wang, L.Z.; Tang, D.; Cai, F.; Gao, W.W. Occurrence of toxigenic fungi in ochratoxin A contaminated liquorice root. Food Addit Contam A 2011, 28, 1091–1097. [Google Scholar] [CrossRef] [PubMed]
- Al-Hindi, R.R.; Aly, S.E.; Hathout, A.S.; Alharbi, M.G.; Al-Masaudi, S.; Al-Jaouni, S.K.; Harakeh, S.M. Isolation and molecular characterization of mycotoxigenic fungi in agarwood. Saudi J. Biol. Sci. 2017. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Põlme, S.; Kõljalg, U.; Yorou, N.S.; Wijesundera, R.; Ruiz, L.; Vasco-Palacios, A.; Thu, P.; Suija, A.; et al. Global diversity and geography of soil fungi. Science 2014, 346, 1256688. [Google Scholar] [CrossRef] [PubMed]
- Bittinger, K.; Charlson, E.S.; Loy, E.; Shirley, D.J.; Haas, A.R.; Laughlin, A.; Yi, Y.; Wu, G.; Lewis, J.; Frank, I.; et al. Improved characterization of medically relevant fungi in the human respiratory tract using next-generation sequencing. Genome Biol. 2014, 15, 487. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; An, C.; Xu, S.; Yamamoto, N. High-throughput sequencing reveals unprecedented diversities of Aspergillus species in outdoor air. Lett. Appl. Microbiol. 2016, 63, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Aiko, V.; Mehta, A. Prevalence of toxigenic fungi in common medicinal herbs and spices in India. 3 Biotech. 2016, 6, 159. [Google Scholar] [CrossRef] [PubMed]
- Zheng, R.S.; Wang, W.L.; Tan, J.; Xu, H.; Zhan, R.T.; Chen, W.W. An investigation of fungal contamination on the surface of medicinal herbs in China. Chin. Med. 2017, 12, 2. [Google Scholar] [CrossRef] [PubMed]
- Su, C.; Hu, Y.; Gao, D.; Luo, Y.I.; Chen, A.J.; Jiao, X.; Gao, W. Occurrence of Toxigenic Fungi and Mycotoxins on Root Herbs from Chinese Markets. J. Food Prot. 2018, 81, 754–761. [Google Scholar] [CrossRef] [PubMed]
- Mandeel, Q.A. Fungal contamination of some imported spices. Mycopathologia 2005, 159, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.J.; Jiao, X.; Hu, Y.; Lu, X.; Gao, W. Mycobiota and mycotoxins in traditional medicinal seeds from China. Toxins 2015, 7, 3858–3875. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Fungal Barcoding Consortium. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Bellemain, E.; Carlsen, T.; Brochmann, C.; Coissac, E.; Taberlet, P.; Kauserud, H. ITS as an environmental DNA barcode for fungi: An in silico approach reveals potential PCR biases. BMC Microbiol. 2010, 10, 189. [Google Scholar] [CrossRef] [PubMed]
- De Filippis, F.; Laiola, M.; Blaiotta, G.; Ercolini, D. Different amplicon targets for sequencing-based studies of fungal diversity. Appl. Environ. Microbiol. 2017, 83, pii: e00905-17. [Google Scholar] [CrossRef]
- Tedersoo, L.; Anslan, S.; Bahram, M.; Põlme, S.; Riit, T.; Liiv, I.; Urmas, K.; Veljo, K.; Henrik, N.; Falk, H.; et al. Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. MycoKeys 2015, 10, 1–43. [Google Scholar] [CrossRef]
- Samson, R.A.; Visagie, C.M.; Houbraken, J.; Hong, S.B.; Hubka, V.; Klaassen, C.H.; Perrone, G.; Seifert, K.A.; Susca, A.; Tanney, J.B.; et al. Phylogeny, identification and nomenclature of the genus Aspergillus. Stud. Mycol. 2014, 78, 141–173. [Google Scholar] [CrossRef] [PubMed]
- Bakker, M.G. A fungal mock community control for amplicon sequencing experiments. Mol. Ecol. Resour. 2018, 18, 541–556. [Google Scholar] [CrossRef] [PubMed]
- Visagie, C.M.; Houbraken, J.; Frisvad, J.C.; Hong, S.B.; Klaassen, C.H.W.; Perrone, G.; Seifert, K.A.; Varga, J.; Yaguchi, T.; Samson, R.A. Identification and nomenclature of the genus Penicillium. Stud. Mycol. 2014, 78, 343–371. [Google Scholar] [CrossRef] [PubMed]
- De Llanos Frutos, R.; Fernández-Espinar, M.T.; Querol, A. Identification of species of the genus Candida by analysis of the 5.8 S rRNA gene and the two ribosomal internal transcribed spacers. Anton. Leeuw. 2004, 85, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Daniel, R. The soil metagenome–a rich resource for the discovery of novel natural products. Curr. Opin. Biotechnol. 2004, 15, 199–204. [Google Scholar] [CrossRef] [PubMed]
- Xia, F.; Chen, X.; Guo, M.Y.; Bai, X.H.; Liu, Y.; Shen, G.R.; Li, Y.; Lin, J.; Zhou, X.W. High-throughput sequencing-based analysis of endogenetic fungal communities inhabiting the Chinese Cordyceps reveals unexpectedly high fungal diversity. Sci. Rep. 2016, 6, 33437. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Leff, J.W.; Adams, B.J.; Nielsen, U.N.; Bates, S.T.; Lauber, C.L.; Owens, S.; Gilbert, J.; Wall, D.; Caporaso, J.G. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc. Natl. Acad. Sci. USA. 2012, 109, 21390–21395. [Google Scholar] [CrossRef] [PubMed]
- Žifčáková, L.; Větrovský, T.; Howe, A.; Baldrian, P. Microbial activity in forest soil reflects the changes in ecosystem properties between summer and winter. Environ. Microbiol. 2016, 18, 288–301. [Google Scholar] [CrossRef] [PubMed]
- Ercolini, D. High-throughput sequencing and metagenomics: Moving forward in the culture-independent analysis of food microbial ecology. Appl. Environ. Microbiol. 2013, 79, 3148–3155. [Google Scholar] [CrossRef] [PubMed]
- Marsh, A.J.; O’Sullivan, O.; Hill, C.; Ross, R.P.; Cotter, P.D. Sequence-based analysis of the bacterial and fungal compositions of multiple kombucha (tea fungus) samples. Food Microbiol. 2014, 38, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Wang, B.; Li, F.; Qiu, L.; Li, F.; Wang, S.; Cui, J. Analysis of bacterial communities on aging flue-cured tobacco leaves by 16S rDNA PCR–DGGE technology. Appl. Microbiol. Biotechnol. 2007, 73, 1435–1440. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols a Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.; Goodrich, J.; Gordon, J.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Med. 2010, 7, 335–336. [Google Scholar]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Kõljalg, U.; Nilsson, R.H.; Abarenkov, K.; Tedersoo, L.; Taylor, A.F.; Bahram, M.; Bates, S.; Bruns, T.; Bengtsson-Palme, J.; Callaghan, T.; et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 2013, 22, 5271–5277. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
| Name | Voucher No. | Sources | Group | Mildewy | Group | GenBank Accession No. |
|---|---|---|---|---|---|---|
| Ziziphi Spinosae Semen | RY01 | Ruyang, Henan | HN | No | WM | SAMN10275060 |
| Ziziphi Spinosae Semen | RY02 | Ruyang, Henan | HN | No | WM | SAMN10275061 |
| Ziziphi Spinosae Semen | HX01 | Hui, Henan | HN | No | WM | SAMN10275062 |
| Ziziphi Spinosae Semen | SJ01 | Shijiazhuang, Hebei | HB | No | WM | SAMN10275069 |
| Ziziphi Spinosae Semen | QH01 | Qinhuangdao, Hebei | HB | No | WM | SAMN10275067 |
| Ziziphi Spinosae Semen | SL01 | Shengli, Liaoning | LN | No | WM | SAMN10275065 |
| Ziziphi Spinosae Semen | SL02 | Shengli, Liaoning | LN | No | WM | SAMN10275066 |
| Ziziphi Spinosae Semen | DP01 | Dongping, Shandong | SD | No | WM | SAMN10275070 |
| Ziziphi Spinosae Semen | LL01 | Lanling, Shandong | SD | Yes | FM | SAMN10275071 |
| Ziziphi Spinosae Semen | QH02 | Qinhuangdao, Hebei | HB | Yes | FM | SAMN10275068 |
| Ziziphi Spinosae Semen | YC01 | Yuncheng, Shanxi | SX | Yes | FM | SAMN10275058 |
| Ziziphi Spinosae Semen | YC02 | Yuncheng, Shanxi | SX | Yes | FM | SAMN10275059 |
| Ziziphi Spinosae Semen | LY01 | Lingyuan, Liaoning | LN | Yes | FM | SAMN10275063 |
| Ziziphi Spinosae Semen | LY02 | Lingyuan, Liaoning | LN | Yes | FM | SAMN10275064 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, M.; Jiang, W.; Luo, J.; Yang, M.; Pang, X. Analysis of the Fungal Community in Ziziphi Spinosae Semen through High-Throughput Sequencing. Toxins 2018, 10, 494. https://doi.org/10.3390/toxins10120494
Guo M, Jiang W, Luo J, Yang M, Pang X. Analysis of the Fungal Community in Ziziphi Spinosae Semen through High-Throughput Sequencing. Toxins. 2018; 10(12):494. https://doi.org/10.3390/toxins10120494
Chicago/Turabian StyleGuo, Mengyue, Wenjun Jiang, Jiaoyang Luo, Meihua Yang, and Xiaohui Pang. 2018. "Analysis of the Fungal Community in Ziziphi Spinosae Semen through High-Throughput Sequencing" Toxins 10, no. 12: 494. https://doi.org/10.3390/toxins10120494
APA StyleGuo, M., Jiang, W., Luo, J., Yang, M., & Pang, X. (2018). Analysis of the Fungal Community in Ziziphi Spinosae Semen through High-Throughput Sequencing. Toxins, 10(12), 494. https://doi.org/10.3390/toxins10120494






