Phylogeny and Mycotoxin Characterization of Alternaria Species Isolated from Wheat Grown in Tuscany, Italy
Abstract
:1. Introduction
2. Results
2.1. Detection of Fungal Contamination
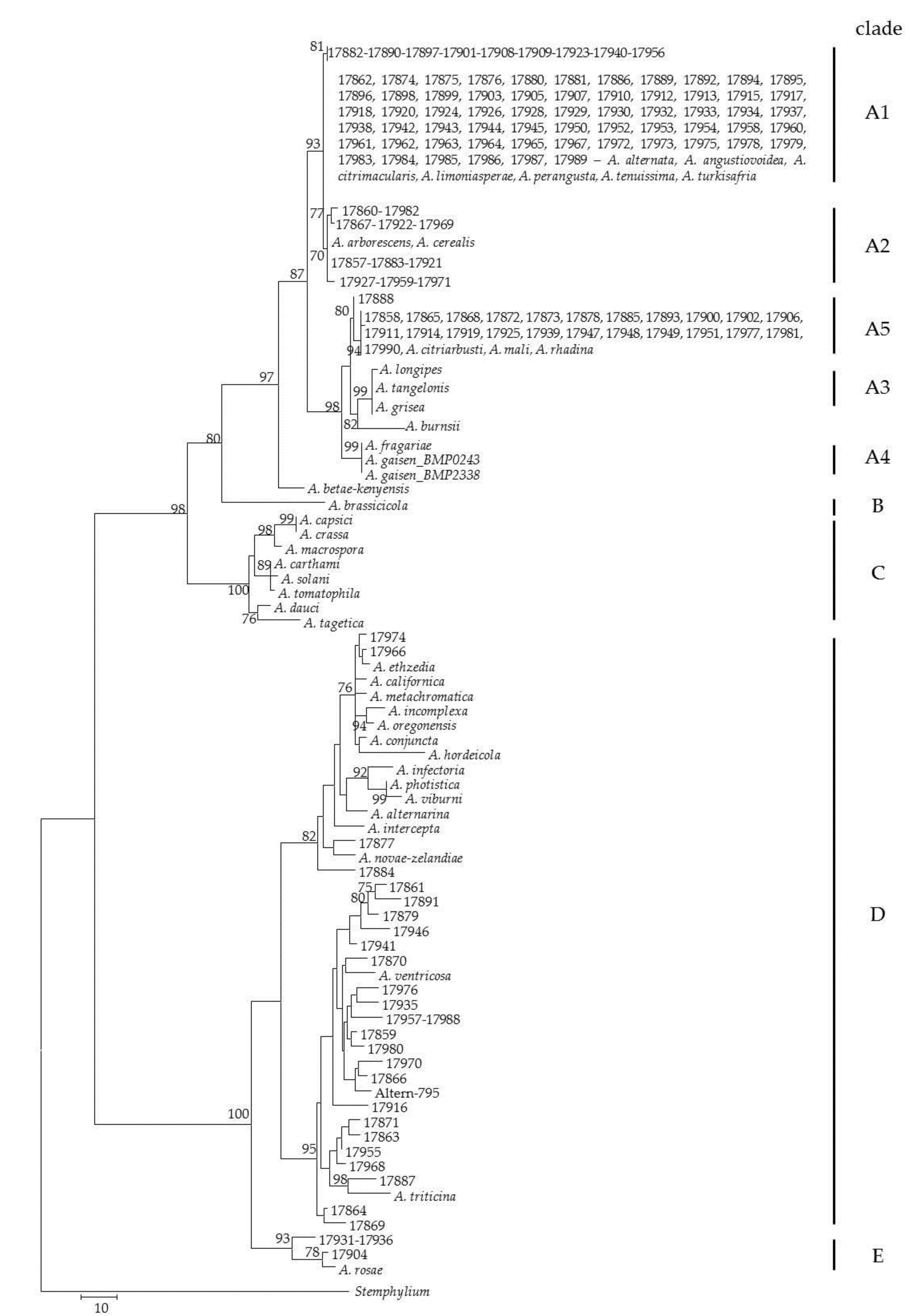
2.2. DNA-Based Identification
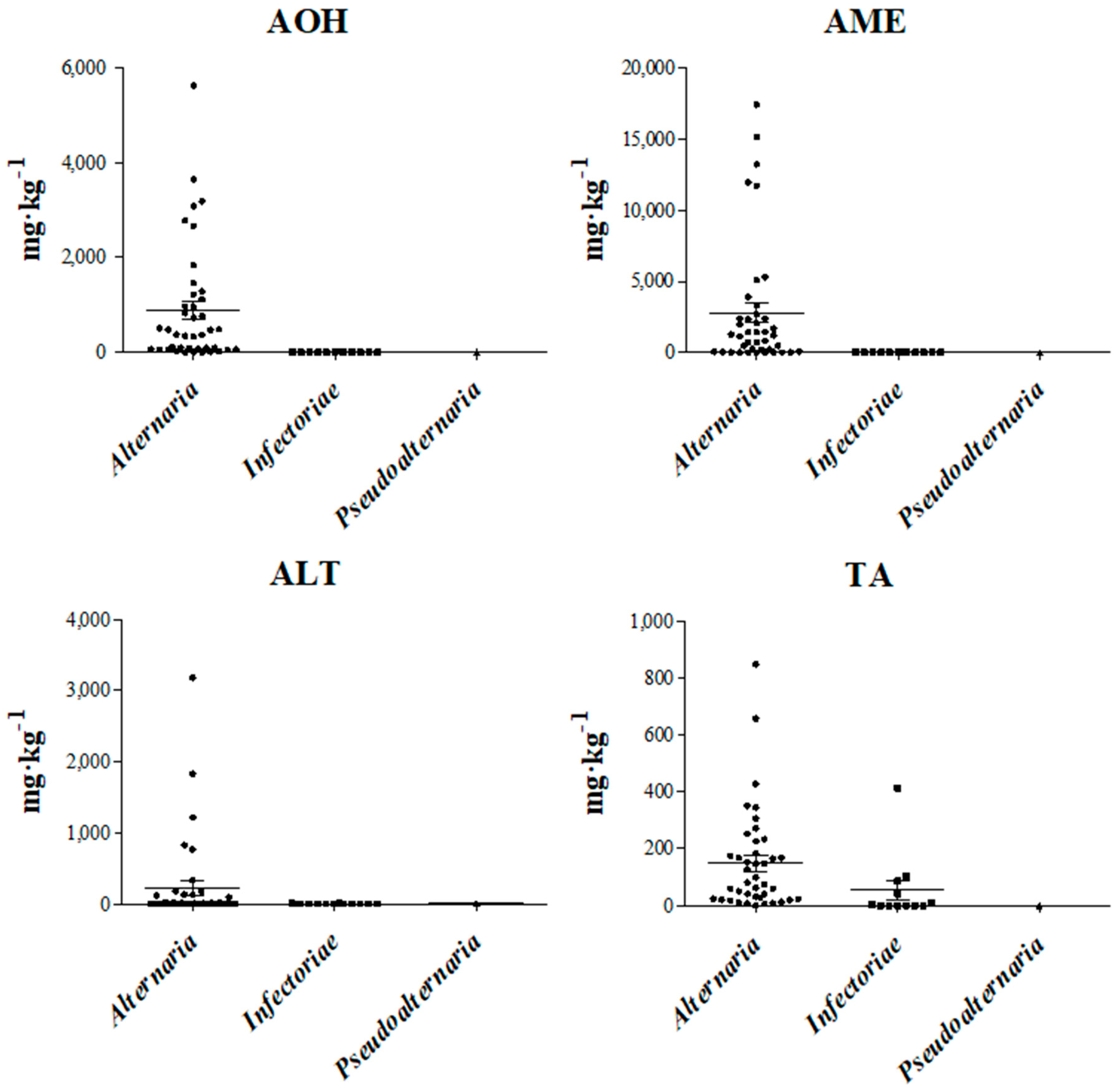
2.3. Mycotoxin Production Profile
3. Discussion
4. Conclusions
5. Material and Methods
5.1. Origin of Wheat Samples
5.2. Isolation and Morphological Characterization of Alternaria Strains
5.3. DNA Extraction
5.4. PCR Amplifications and DNA Sequencing
5.5. Phylogenetic Analyses
5.6. Mycotoxins Extraction
HPLC Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- ISTAT. Available online: www.agri.istat.it (accessed on 11 November 2018).
- Alexander, J.; Benford, D.; Boobis, A.; Ceccatelli, S.; Cottrill, B.; Cravedi, J.; Di Domenico, A.; Doerge, D.; Dogliotti, E.; Edler, L.; et al. Scientific opinion on the risks for animal and public health related to the presence of Alternaria toxins in feed and food. EFSA J. 2011, 9, 2407. [Google Scholar]
- Prabhu, A.S.; Prasada, R. Pathological and epidemiological studies on leaf blight of wheat caused by Alternaria triticina. Indian Phytopathol. 1966, 19, 95–112. [Google Scholar]
- Kulshrestha, V.P.; Rao, M.V. Genetics of resistance to an isolate of Alternaria triticina causing leaf blight of wheat. Euphytica 1976, 25, 769–775. [Google Scholar] [CrossRef]
- Chaurasia, S.; Chand, R.; Joshi, A. Relative dominance of Alternaria triticina Pras. et Prab. and Bipolarissorokiniana (Sacc.) Shoemaker in different growth stages of wheat (T. aestivum L.). J. Plant Dis. Prot. 2000, 107, 176–181. [Google Scholar]
- Logrieco, A.; Moretti, A.; Solfrizzo, M. Alternaria toxins and plant diseases: And overview of origin, occurrence and risks. World Mycotoxin J. 2009, 2, 129–140. [Google Scholar] [CrossRef]
- Amatulli, M.T.; Fanelli, F.; Moretti, A.; Mulè, G.; Logrieco, A.F. Alternaria species and mycotoxins associated to black point of cereals. Mycotoxins 2013, 63, 39–46. [Google Scholar] [CrossRef]
- Somma, S.; Amatulli, M.T.; Masiello, M.; Moretti, A.; Logrieco, A.F. Alternaria species associated to wheat black point identified through a multilocus sequence approach. Int. J. Food Microbiol. 2018. under review. [Google Scholar]
- Kashem, M.A.; Sultana, N.; Samanta, S.C.; Kamal, A.M.A. Biochemical changes in wheat seed due to the effect of black-point at different levels of maturing. Pak. J. Sci. Ind. Res. 1999, 42, 89–92. [Google Scholar]
- Wang, W.; Jones, C.; Ciacci-Zanella, J.; Holt, T.; Gilchrist, D.G.; Dickman, M.B. Fumonisins and Alternaria alternata lycopersici toxins: Sphinganine analog mycotoxins induce apoptosis in monkey kidney cells. Proc. Natl. Acad. Sci. USA 1996, 93, 3461–3465. [Google Scholar] [CrossRef] [PubMed]
- Ostry, V. Alternaria mycotoxins: An overview of chemical characterization, producers, toxicity, analysis and occurrence in foodstuffs. World Mycotoxin J. 2008, 1, 175–188. [Google Scholar] [CrossRef]
- Patriarca, A.; Azcarate, M.P.; Terminiello, L.; Fernández Pinto, V. Mycotoxin production by Alternaria strains isolated from Argentinean wheat. Int. J. Food Microbiol. 2007, 119, 219–222. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.B.; Patriarca, A.; Magan, N. Alternaria in Food: Ecophysiology, Mycotoxin Production and Toxicology. Mycobiology 2015, 43, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Da Cruz Cabrala, L.; Terminiello, L.; Pinto, V.F.; Fog, K.; Patriarca, N.A. Natural occurrence of mycotoxins and toxigenic capacity of Alternaria strains from mouldy peppers. Int. J. Food Microbiol. 2016, 7, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.T.; Qian, Y.Z.; Zhang, P.; Dong, Z.M.; Shi, Z.Y.; Zhen, Y.Z.; Miao, J.; Xu, Y.M. Relationships between Alternaria alternata and oesophageal cancer. IARC Sci. Publ. 1991, 105, 258–262. [Google Scholar]
- Steyn, P.S.; Rabie, C.J. Characterization of magnesium and calcium tenuazonate from Phomasorghina. Phytochemistry 1976, 15, 1977–1979. [Google Scholar] [CrossRef]
- Yekeler, H.; Bitmis, K.; Ozcelik, N.; Doymaz, M.Z.; Calta, M. Analysis of toxic effects of Alternaria toxins on oesophagus of mice by light and electron microscopy. Toxicol. Pathol. 2001, 29, 492–497. [Google Scholar] [CrossRef] [PubMed]
- Schrader, T.J.; Cherry, W.; Soper, K.; Langlois, I. Further examination of the effects of nitrosylation on Alternaria alternata mycotoxin mutagenicity in vitro. Mutat. Res. 2006, 606, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Huybrechts, I.; De Ruyck, K.; De Saeger, S.; De Boevre, M. Uniting large-scale databeses to unravel the impact of chronic multi-mycotoxins exposures on colorectal cancer incidence in Europe. In Proceedings of the 2nd MycoKey International Conference, Wuhan, China, 16–18 September 2018; China Agricultural Science and Technology Press: Beijing, China, 2018; pp. 181–183. [Google Scholar]
- Simmons, E.G. Alternaria: An Identification Manual, 6th ed.; CBS Fungal Biodiversity Centre: Utrecht, The Netherlands, 2007. [Google Scholar]
- Hong, S.G.; Cramer, R.A.; Lawrence, C.B.; Pryor, B.M. Alt a1 allergen homologs from Alternaria and related taxa: Analysis of phylogenetic content and secondary structure. Fungal Genet. Biol. 2005, 42, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.G.; Maccaroni, M.; Figuli, P.J.; Pryor, B.M.; Belisario, A. Polyphasic classification of Alternaria isolated from hazelnut and walnut fruit in Europe. Mycol. Res. 2006, 110, 1290–1300. [Google Scholar] [CrossRef] [PubMed]
- Andersen, B.; Sorensen, J.L.; Nielsen, K.F.; van den Ende, B.G.; de Hoog, S. A polyphasic approach to the taxonomy of the Alternaria infectoria species-group. Fungal Genet. Biol. 2009, 46, 642–656. [Google Scholar] [CrossRef] [PubMed]
- Somma, S.; Pose, G.; Pardo, A.; Mulè, G.; Pinto, V.F.; Moretti, A.; Logrieco, A.F. AFLP variability, toxin production, and pathogenicity of Alternaria species from Argentinean tomato fruits and puree. Int. J. Food Microbiol. 2011, 145, 414–419. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, D.P.; Gannibal, P.B.; Peever, T.L.; Pryor, B.M. The sections of Alternaria: Formalizing species-group concepts. Mycologia 2013, 105, 530–546. [Google Scholar] [CrossRef] [PubMed]
- Woudenberg, J.H.C.; Groenewald, J.Z.; Binder, M.; Crous, P.W. Alternaria redefined. Stud. Mycol. 2013, 75, 171–212. [Google Scholar] [CrossRef] [PubMed]
- Polizzotto, R.; Andersen, B.; Martini, M.; Grisan, S.; Assante, G.; Musetti, R. A polyphasic approach for the characterization of endophytic Alternaria strains isolated from grapevines. J. Microbiol. Methods 2012, 88, 162–171. [Google Scholar] [CrossRef] [PubMed]
- Bottalico, A.; Perrone, G. Toxigenic Fusarium species and mycotoxins associated with head blight in small-grain cereals in Europe. Eur. J. Plant Pathol. 2002, 108, 611–624. [Google Scholar] [CrossRef]
- Moretti, A.; Logrieco, A. Climate change effects on the biodiversity of mycotoxigenic fungi and their mycotoxins in preharvest conditions in Europe. In Climate Change and Mycotoxins; Botana, L.M., Sainz, J.M., Eds.; Walter de Gruyter GmbH & Co KG: Berlin, Germany, 2015; pp. 91–108. ISBN 978-3-11-033305-3. [Google Scholar]
- Arcella, D.; Eskola, M.; Gómez Ruiz, J.A. Scientific report on the dietary exposure assessment to Alternaria toxins in the European population. EFSA J. 2016, 14. [Google Scholar] [CrossRef]
- Solhaug, A.; Eriksen, G.S.; Holme, J.A. Mechanisms of action and toxicity of the mycotoxin alternariol: A review. Basic Clin. Pharmacol. Toxicol. 2016, 119, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Benassi, F.; Zid, M.; Rhouma, A.; Bacha, H.; Hajlaoui, M.R. First report of Alternaria species associated with black point of wheat in Tunisia. Ann. Microbiol. 2009, 59, 465–467. [Google Scholar] [CrossRef]
- Kahl, S.M.; Ulrich, A.; Kirichenko, A.A.; Muller, M.E.H. Phenotypic and phylogenetic segregation of Alternaria infectoria from small-spored Alternaria species isolated from wheat in Germany and Russia. J. Appl. Microbiol. 2015, 119, 1637–1650. [Google Scholar] [CrossRef] [PubMed]
- Perello, A.; Moreno, M.; Sisterna, M. Alternaria infectoria species-group associated with Black point of wheat in Argentina. Plant Pathol. 2008, 57, 379. [Google Scholar] [CrossRef]
- Kammoun, G.L.; Bensassi, F.; Hattab, M.M.; Rhouma, A.; Bacha, H.; Hajlaoui, M.R. Identification of Alternaria species recovered from stored durum wheat kernels in Tunisia. Tunis. J. Plant Prot. 2014, 9, 119–129. [Google Scholar]
- Woudenberg, J.H.C.; Seidl, M.F.; Groenewald, J.Z.; De Vries, M.; Stielow, J.B.; Thomma, B.P.H.J.; Crous, P.W. Alternaria section Alternaria: Species, formaespeciales or pathotypes? Stud. Mycol. 2015, 82, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, D.P.; Rotondo, F.; Gannibal, P.B. Biodiversity and taxonomy of the pleomorphic genus Alternaria. Mycol. Prog. 2016, 15, 3. [Google Scholar] [CrossRef]
- Lawrence, D.P.; Gannibal, P.B.; Peever, T.L.; Dugan, F.M.; Pryor, B. Characterization of Alternaria isolates from the infectoria species-group and a new taxon from Arrhenatherum, Pseudoalternaria arrhenatheria sp. nov. Mycol. Prog. 2014, 13, 257–276. [Google Scholar] [CrossRef]
- Simmons, E. Alternaria themes and variations (106–111). Mycotaxon 1994, 50, 409–427. [Google Scholar]
- Simmons, E.G. Alternaria themes and variations (22–26). Mycotaxon 1986, 25, 287–308. [Google Scholar]
- Simmons, E. Alternaria themes and variations (305–309) Lewia/Alternaria revisited. Mycotaxon 2002, 83, 127–145. [Google Scholar]
- Poursafar, A.; Ghosta, Y.; Orina, A.S.; Gannibal, P.B.; Nikkhah, M.J.; Lawrence, D.P. Taxonomic study on Alternaria sections Infectoriae and Pseudoalternaria associated with black (sooty) head mold of wheat and barley in Iran. Mycol. Prog. 2018, 17, 343–356. [Google Scholar] [CrossRef]
- Berbee, M.L.; Pirseyedi, M.; Hubbard, S. Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 1999, 91, 964–977. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Rubert, J.; Dzuman, Z.; Vaclavikova, M.; Zachariasova, M.; Soler, C.; Hajslova, J. Analysis of mycotoxins in barley using ultra high liquid chromatography high resolution mass spectrometry: Comparison of efficiency and efficacy of different extraction procedures. Talanta 2012, 99, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Myresiotis, C.K.; Testempasis, S.; Vryzas, Z.; Karaoglanidis, G.S.; Mourkidou, P.E. Determination of mycotoxins in pomegranate fruits and juices using a QuEChERS-based method. Food Chem. 2015, 182, 81–88. [Google Scholar] [CrossRef] [PubMed]
| Host | Year of Sampling | Number of Fields | Fungal Contamination (%) | |||
|---|---|---|---|---|---|---|
| Alternaria spp. | Other Fungi | |||||
| Range | Mean Value | Range | Mean Value | |||
| Soft wheat | 2013 | 10 | 0–31 | 17 | 36–82 | 68 |
| 2014 | 16 | 0–17 | 4 | 19–95 | 73 | |
| 2015 | 11 | 4–73 | 26 | 15–81 | 54 | |
| 2016 | 11 | 11–50 | 33 | 36–83 | 57 | |
| Durum wheat | 2013 | 31 | 0–48 | 17 | 1–88 | 55 |
| 2014 | 15 | 0–11 | 7 | 57–93 | 76 | |
| 2015 | 4 | 8–43 | 26 | 36–84 | 64 | |
| 2016 | 2 | 22–35 | 29 | 60–77 | 68 | |
| Year of Sampling | Durum Wheat a | Soft Wheat a | Alternaria Contamination (%) b | Meteorological Data (from Seedling to Harvest) | Meteorological Data (from Heading to Harvest) | ||
|---|---|---|---|---|---|---|---|
| Rainfall (mm) | Mean T | Rainfall (mm) | Mean T | ||||
| 2013 | 31 | 10 | 17.1 | 820 | 9.1 | 181 | 14.8 |
| 2014 | 15 | 16 | 5.5 | 580 | 10.9 | 95 | 15 |
| 2015 | 4 | 11 | 26.2 | 525 | 11.1 | 115 | 16.6 |
| 2016 | 2 | 11 | 32.8 | 691 | 12.7 | 256 | 17.6 |
| Strain a | Mycotoxin (mg·kg−1) b | Strain a | Mycotoxin (mg·kg−1) b | ||||||
|---|---|---|---|---|---|---|---|---|---|
| AOH | AME | ALT | TA | AOH | AME | ALT | TA | ||
| Alternaria Section Sub-Clade A1 | Alternaria Section Sub-Clade A5 | ||||||||
| 17,862 | 60 | 710 | n.d. | 150 | 17,888 | 95 | 1450 | n.d. | 165 |
| 17,874 | 350 | 1440 | 140 | 305 | 17,893 | 335 | 3 | 10 | 17 |
| 17,875 | 475 | 2400 | 175 | 345 | 17,900 | 1225 | 2700 | 1 | 255 |
| 17,876 | 960 | 1700 | 120 | 350 | 17,902 | 835 | 15 | n.d. | 130 |
| 17,880 | 970 | 15,150 | n.d. | n.d. | 17,906 | 0.5 | 2 | 3 | 5.5 |
| 17,881 | 505 | 2385 | 15 | 60 | 17,865 | 385 | 1125 | 0.5 | 40 |
| 17,882 | 480 | 1420 | n.d. | 235 | 17,868 | 1290 | 3315 | n.d | 155 |
| 17,886 | 65 | 480 | 5 | 30 | 17,873 | 100 | 724 | 130 | 60 |
| 17,887 | 4.5 | 15 | 15 | 20 | 17,878 | 20 | 12.5 | 3 | 12.5 |
| 17,889 | 2780 | 13,230 | 180 | 230 | 17,885 | 3650 | 17,415 | 4 | 170 |
| 17,890 | 58 | 475 | 30 | 80 | Average | 794 | 2676 | 15 | 101 |
| 17,892 | 96 | 20 | 16 | 75 | Min Value | 1 | 2 | 0 | 6 |
| 17,894 | 375 | 1260 | 22 | 42 | Max value | 3650 | 17415 | 130 | 255 |
| 17,895 | 95 | 220 | n.d. | 272 | Infectoriae Section Clade D | ||||
| 17,896 | 1465 | 11,710 | 3.5 | 848 | 17,859 | 1.5 | 4 | n.d. | 105 |
| 17,897 | 3090 | 11,950 | 9 | 10 | 17,861 | 2.5 | 23 | 1.5 | 4 |
| 17,898 | 110 | 230 | 1.0 | 170 | 17,863 | 2.5 | 1 | 8.5 | n.d. |
| 17,899 | 1120 | 2050 | 2.5 | 660 | 17,864 | 3 | 3 | n.d. | 10 |
| 17,901 | 60 | 75 | n.d. | 145 | 17,866 | 3.5 | 10 | n.d. | n.d. |
| 17,903 | 730 | 3910 | n.d. | 430 | 17,869 | 0.5 | 6 | n.d. | n.d. |
| 17,905 | 70 | 80 | n.d. | 23 | 17,870 | 0.5 | 2 | n.d. | n.d. |
| 17,907 | 485 | 2335 | n.d. | 175 | 17,871 | 10 | 6 | n.d. | n.d. |
| 17,908 | 75 | 215 | n.d. | 20 | 17,877 | 2.5 | 2.5 | n.d. | n.d. |
| 17,909 | 14 | 7 | 7.5 | 25 | 17,879 | 0.5 | 0.5 | n.d. | 415 |
| 17,910 | 2670 | 5096 | n.d. | 185 | 17,884 | 20 | 15 | 10 | 40 |
| 17,912 | 25 | 15 | 7 | 50 | 17,891 | 2 | 1 | 0.2 | 90 |
| 17,858 | 5620 | 5290 | 1.5 | 65 | Average | 4 | 6 | 2 | 55 |
| 17,911 | 770 | 1970 | 0.5 | 30 | Min Value | 0.3 | 0.5 | 0 | 0 |
| Average | 842 | 3066 | 27 | 180 | Max value | 20 | 23 | 10 | 415 |
| Min Value | 5 | 7 | 0 | 0 | - | Pseudoalternaria Section Clade E | |||
| Max value | 5620 | 15150 | 180 | 848 | 17,904 | 1.5 | 5.5 | 10 | n.d. |
| Alternaria Section Sub-Clade A2 | |||||||||
| 17,857 | 3180 | 1215 | 0.5 | 7 | |||||
| 17,867 | 1835 | 810 | n.d. | 7.5 | |||||
| 17,883 | 2.5 | 10 | n.d. | 100 | |||||
| Average | 1673 | 678 | 0.2 | 38 | |||||
| Min Value | 2.5 | 10 | 0.0 | 7 | |||||
| Max value | 3180 | 1215 | 0.5 | 100 | |||||
| Alternaria Species | Strain Number | GenBank Accession Numbers | |||
|---|---|---|---|---|---|
| Alt-a-1 | Gpd | Tef | Database * | ||
| A. alternarina | CBS 119396 | JQ905113 | JQ905170 | JQ905142 | NCBI |
| A. alternata | ATCC11680 | ATNCTG00005 | ATNCTG00056 | ATNCTG00621 | AGD |
| A. alternata | ATCC66891 | AATCTG00058 | AATCTG00420 | AATCTG00079 | AGD |
| A. alternata | BMP 0270 | AA2CTG00036 | AA2CTG00134 | AA2CTG00228 | AGD |
| A. angustiovoidea | EGS 36-172 | JQ646398 | JQ646315 | JQ672465 | NCBI |
| A. arborescens | BMP 0308 | AABCTG00367 | AABCTG03225 | AABCTG04967 | AGD |
| A. betae-kenyensis | CBS 118810 | JQ905104 | JQ905161 | KP125197 | NCBI |
| A. brassicicola | EEB 2232 | AY563311 | AY278813 | JQ672450 | NCBI |
| A. burnsii | CBS 107.38 | JQ646388 | JQ646305 | KP125198 | NCBI |
| A. californica | EGS 52-082 | JQ646373 | JQ646285 | JQ672433 | NCBI |
| A. capsici | BMP 0180 | AY563298 | AY562408 | ACSCTG01642 | NCBI/AGD |
| A. cerealis | EGS 43-072 | JQ646405 | JQ646321 | JQ672467 | NCBI |
| A. citriarbusti | BMP 2343 | ACSCTG04746 | ACSCTG00332 | ACSCTG01642 | AGD |
| A. citrimacularis | BC2-RLR-17s | JQ646407 | JQ646323 | JQ672466 | NCBI |
| A. conjuncta | EGS 37-139 | AY563281 | AY562401 | JQ672415 | NCBI |
| A. crassa | BMP 0172 | AY563293 | AY278804 | JQ672489 | NCBI |
| A. dauci | BMP 0167 | AY563292 | AY278803 | ADCCTG02999 | NCBI/AGD |
| A. ethzedia | EGS 37-143 | AY563284 | AY278795 | JQ672427 | NCBI |
| A. fragariae | BMP 3062 | ACTCTG00345 | ACTCTG00074 | ACTCTG00439 | AGD |
| A. gaisen | BMP 2338 | ACRCTG04151 | ACRCTG04221 | ACRCTG02961 | AGD |
| A. gaisen | BMP 0243 | JQ646400 | JQ646317 | JQ672463 | NCBI |
| A. grisea | CBS 107.36 | JQ646393 | JQ646310 | JQ672471 | NCBI |
| A. hordeicola | EGS 50-184 | JQ646372 | JQ646284 | JQ672425 | NCBI |
| A. incomplexa | EGS 17-103 | JQ646374 | JQ646287 | JQ672422 | NCBI |
| A. infectoria | EGS 27.193 | FJ266502 | AY278793 | JQ672436. | NCBI |
| A. intercepta | EGS 49-137 | JQ646380 | JQ646297 | JQ672431 | NCBI |
| A. limoniasperae | BMP 2335 | AFGCTG00301 | AFGCTG00774 | AFGCTG00104 | AGD |
| A. longipes | BMP 0313 | ADTCTG24504 | ADTCTG20582 | ADTCTG20250 | AGD |
| A. mali | BMP 3064 | AGSCTG02862 | AGSCTG00707 | AGSCTG00243 | AGD |
| A. macrospora | BMP 1949 | AMRCTG01538 | AMRCTG02006 | AMRCTG00463 | AGD |
| A. metachromatica | EGS 38-132 | AY563285 | AY762956 | JQ672437 | NCBI |
| A. novae-zelandiae | EGS 48-092 | JQ646379 | JQ646296 | JQ672418 | NCBI |
| A. oregonensis | EGS 29-194 | AY563283 | AY762957 | JQ672428 | NCBI |
| A. perangusta | BMP 2336 | JQ646403 | JQ646319 | JQ672477 | NCBI |
| A. photistica | EGS 35-172 | AY563282 | AY562402 | JQ672417 | NCBI |
| A. rhadina | CBS 595.93 | JQ646399 | JQ646316 | JQ672470 | NCBI |
| A. rosae | EGS 41-130 | JQ646370 | JQ646279 | JQ672414 | NCBI |
| A.solani | CBS 116651 | AY563299 | KC584139 | KC584688 | NCBI |
| A. tangelonis | BMP 2327 | ADCCTG06617 | ADCCTG03746 | ADCCTG02999 | AGD |
| A. tagetica | BMP 0179 | AY563297 | AY562407 | JQ672490 | NCBI |
| A. tenuissima | BMP 0304 | ALGCTG02124 | ALGCTG02071 | ALGCTG00260 | AGD |
| A. tomatophila | BMP 2032 | GQ180101 | GQ180085 | ATMCTG00738 | NCBI/AGD |
| A. triticina | EGS 17-061 | JQ646371 | JQ646281 | JQ672426 | NCBI |
| A. turkisafria | BMP 3436 | ATKCTG00833 | ATKCTG00298 | ATKCTG00533 | AGD |
| A. ventricosa | EGS 52-075 | JQ646377 | JQ646290 | JQ672426 | NCBI |
| A. viburni | EGS 49-147 | JQ646375 | JQ646288 | JQ672420 | NCBI |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramires, F.A.; Masiello, M.; Somma, S.; Villani, A.; Susca, A.; Logrieco, A.F.; Luz, C.; Meca, G.; Moretti, A. Phylogeny and Mycotoxin Characterization of Alternaria Species Isolated from Wheat Grown in Tuscany, Italy. Toxins 2018, 10, 472. https://doi.org/10.3390/toxins10110472
Ramires FA, Masiello M, Somma S, Villani A, Susca A, Logrieco AF, Luz C, Meca G, Moretti A. Phylogeny and Mycotoxin Characterization of Alternaria Species Isolated from Wheat Grown in Tuscany, Italy. Toxins. 2018; 10(11):472. https://doi.org/10.3390/toxins10110472
Chicago/Turabian StyleRamires, Francesca A., Mario Masiello, Stefania Somma, Alessandra Villani, Antonia Susca, Antonio F. Logrieco, Carlos Luz, Giuseppe Meca, and Antonio Moretti. 2018. "Phylogeny and Mycotoxin Characterization of Alternaria Species Isolated from Wheat Grown in Tuscany, Italy" Toxins 10, no. 11: 472. https://doi.org/10.3390/toxins10110472
APA StyleRamires, F. A., Masiello, M., Somma, S., Villani, A., Susca, A., Logrieco, A. F., Luz, C., Meca, G., & Moretti, A. (2018). Phylogeny and Mycotoxin Characterization of Alternaria Species Isolated from Wheat Grown in Tuscany, Italy. Toxins, 10(11), 472. https://doi.org/10.3390/toxins10110472










