Buzz Kill: Function and Proteomic Composition of Venom from the Giant Assassin Fly Dolopus genitalis (Diptera: Asilidae)
Abstract
1. Introduction
2. Results
2.1. Bioactivity of Crude Venom
2.2. Proteomics Reveals Numerous Peptide and Protein Families in Asilid Venom
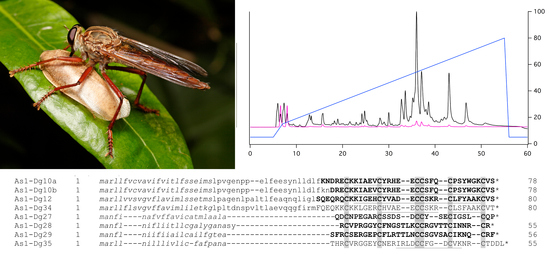
2.3. Disulfide-Rich Venom Peptides
2.4. Mature Primary Structures of Peptides in Venom
3. Discussion
4. Materials and Methods
4.1. Insects and Venom Collection
4.2. Toxicity Assays
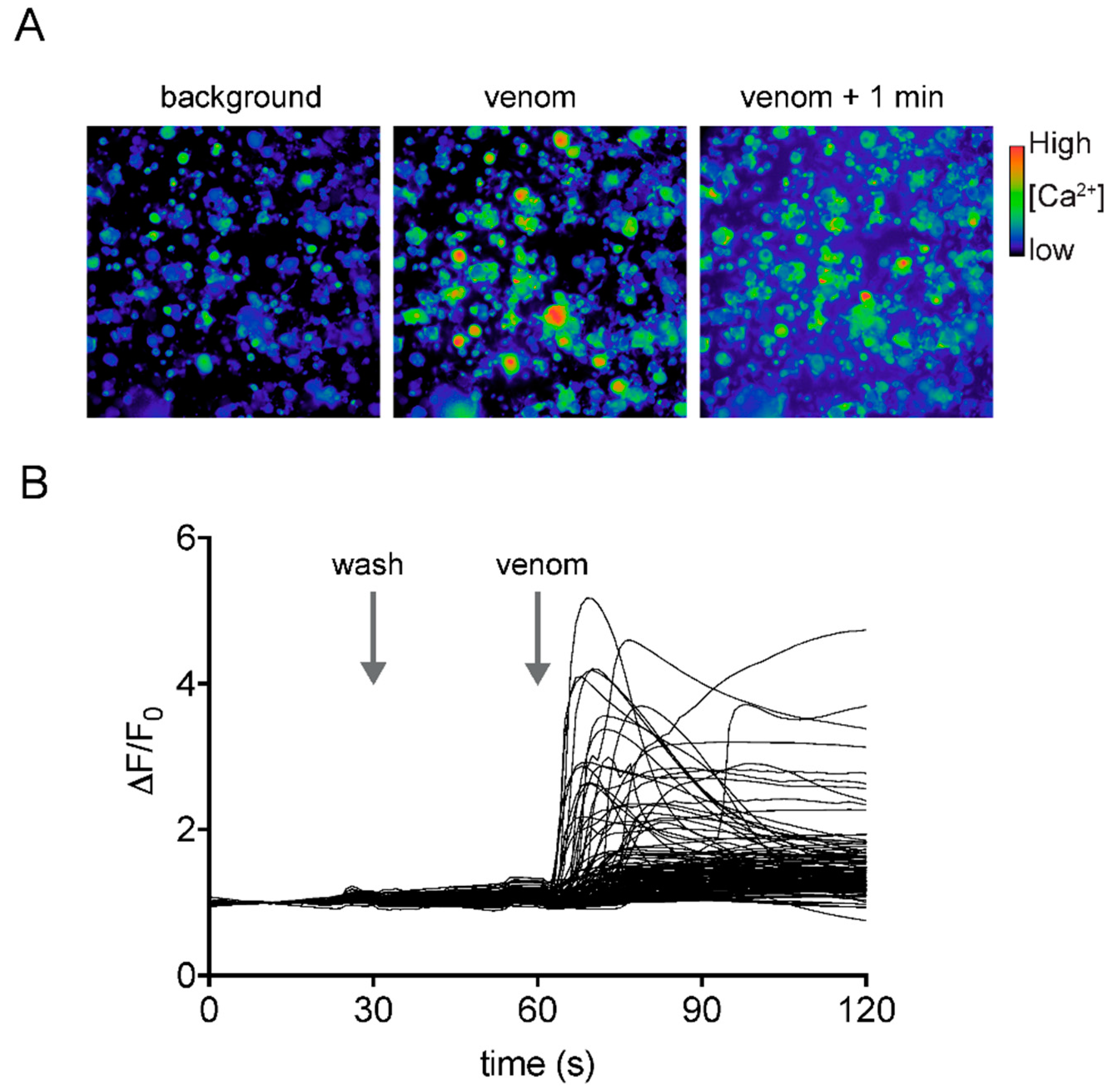
4.3. Calcium Imaging of Mouse Sensory Neurons
4.4. Calcium Imaging of Human Neuroblastoma SH-SY5Y Cells
4.5. Chromatography
4.6. Electrophoresis
4.7. Transcriptomics
4.8. Mass Spectrometry
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Walker, A.A.; Robinson, S.D.; Yeates, D.K.; Jin, J.; Baumann, K.; Dobson, J.; Fry, B.G.; King, G.F. Entomo-venomics: The evolution, biology, and biochemistry of insect venoms. Toxicon 2018, 154, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Suranse, V.; Srikanthan, A.; Sunagar, K. Animal venoms: Origin, diversity and evolution. In eLS; John Wiley & Sons, L., Ed.; American Cancer Society: Oklahoma City, OK, USA, 2018; pp. 1–20. [Google Scholar]
- Casewell, N.R.; Wüster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.S.W.; Morganstern, D.; Mofiz, E.; Gombert, S.; Morris, K.M.; Temple-Smith, P.; Renfree, M.B.; Whittington, C.M.; King, G.F.; Warren, W.C.; et al. Proteomics and deep sequencing comparison of seasonally active venom glands in the platypus reveals novel venom peptides and distinct expression profiles. Mol. Cell. Proteom. 2012, 11, 1354–1364. [Google Scholar] [CrossRef] [PubMed]
- Undheim, E.A.B.; Fry, B.G.; King, G.F. Centipede venom: Recent discoveries and current state of knowledge. Toxins 2015, 7, 679–704. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.A.; Madio, B.; Jin, J.; Undheim, E.A.; Fry, B.G.; King, G.F. Melt with this kiss: Paralyzing and liquefying venom of the assassin bug Pristhesancus plagipennis (Hemiptera: Reduviidae). Mol. Cell. Proteom. 2017, 16, 552–566. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.A.; Mayhew, M.L.; Jin, J.; Herzig, V.; Undheim, E.A.B.; Sombke, A.; Fry, B.G.; Meritt, D.J.; King, G.F. The assassin bug Pristhesancus plagipennis produces two distinct venoms in separate gland lumens. Nat. Commun. 2018, 9, 755. [Google Scholar] [CrossRef] [PubMed]
- Von Reumont, B.M.; Campbell, L.I.; Richter, S.; Hering, L.; Sykes, D.; Hetmank, J.; Jenner, R.A.; Bleidorn, C. A polychaete’s powerful punch: Venom gland transcriptomics of Glycera reveals a complex cocktail of toxin homologs. Genome Biol. Evol. 2014, 6, 2406–2423. [Google Scholar] [CrossRef] [PubMed]
- Dikow, T.; Grimaldi, D.A. Robber flies in Cretaceous ambers (Insecta: Diptera: Asilidae). Am. Mus Novit. 2014, 3799, 1–19. [Google Scholar] [CrossRef]
- Dikow, T. A phylogenetic hypothesis for Asilidae based on a total evidence analysis of morphological and DNA sequence data (Insecta: Diptera: Brachycera: Asiloidea). Org. Divers. Evol. 2009, 9, 165–188. [Google Scholar] [CrossRef]
- Wardill, T.J.; Fabian, S.T.; Pettigrew, A.C.; Stavenga, D.G.; Nordström, K.; Gonzalez-Bellido, P.T. A novel interception strategy in a miniature robber fly with extreme visual acuity. Curr. Biol. 2017, 27, 854–859. [Google Scholar] [CrossRef] [PubMed]
- Ghahari, H.; Lehr, P.A.; Lavigne, R.J.; Hayat, R.; Ostovan, H. New records of robber flies (Diptera, Asilidae) for the Iranian fauna with their prey records. Far East. Entomol. 2007, 179, 1–9. [Google Scholar]
- Whitfield, F.G.S. The relation between the feeding-habits and the structure of the mouth-parts in the Asilidæ (Diptera). Proc. Zool. Soc. Lond. 1925, 95, 599–638. [Google Scholar] [CrossRef]
- Dennis, D.S.; Lavigne, R.J.; Dennis, J.G. Spiders (Araneae) as prey of robber flies (Diptera: Asilidae). J. Entomol. Res. Soc. 2012, 14, 65–76. [Google Scholar]
- Owsley, W.M.B. The comparative morphology of internal structures of the Asilidae (Diptera). Ann. Entomol. Soc. Am. 1946, 39, 33–68. [Google Scholar] [CrossRef]
- Drukewitz, S.; Fuhrmann, N.; Undheim, E.; Blanke, A.; Giribaldi, J.; Mary, R.; Laconde, G.; Dutertre, S.; von Reumont, B. A dipteran’s novel sucker punch: Evolution of arthropod atypical venom with a neurotoxic component in robber flies (Asilidae, Diptera). Toxins 2018, 10, 29. [Google Scholar] [CrossRef] [PubMed]
- Kahan, D. The toxic effect of the bite and the proteolytic activity of the saliva and stomach contents of the robber flies (Diptera: Asilidae). Isr. J. Zool. 1964, 13, 47–57. [Google Scholar]
- Musso, J.J.; Garnier, R.; Legier, F. Comparison of toxicity of venom of some asilids (Diptera: Brachycera) on locusts. Ann. Entomol. Fr. 1978, 14, 177–184. [Google Scholar]
- BromLey, S.W. Ohio robber flies IV (Diptera: Asilidae). Ohio J. Sci. 1947, 47, 67–68. [Google Scholar]
- Walker, A.A.; Rosenthal, M.; Undheim, E.A.B.; King, G.F. Harvesting venom toxins from assassin bugs and other heteropteran insects. JoVE 2018, e57729. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.D.; Mueller, A.; Clayton, D.; Starobova, H.; Hamilton, B.R.; Payne, R.J.; Vetter, I.; King, G.F.; Undheim, E.A.B. A comprehensive portrait of the venom of the giant red bull ant, Myrmecia gulosa, reveals a hyperdiverse hymenopteran toxin gene family. Sci. Adv. 2018, 4, eaau4640. [Google Scholar] [CrossRef] [PubMed]
- Vetter, I. Development and optimization of FLIPR high throughput calcium assays for ion channels and GPCRs. In Calcium Signaling; Islam, M.S., Ed.; Springer: Dordrecht, The Netherlands, 2012; Volume 740, pp. 45–82. [Google Scholar]
- Vetter, I.; Dekan, Z.; Knapp, O.; Adams, D.J.; Alewood, P.F.; Lewis, R.J. Isolation, characterization and total regioselective synthesis of the novel muO-conotoxin MfVIA from Conus magnificus that targets voltage-gated sodium channels. Biochem. Pharmacol. 2012, 84, 540–548. [Google Scholar] [CrossRef] [PubMed]
- King, G.F.; Gentz, M.C.; Escoubas, P.; Nicholson, G.M. A rational nomenclature for naming peptide toxins from spiders and other venomous animals. Toxicon 2008, 52, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Garnier, J.; Gibrat, J.-F.; Robson, B. GOR secondary structure prediction method version IV. Methods Enzymol. 1996, 266, 540–553. [Google Scholar] [PubMed]
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.; King, G.F.; Nevalainen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The toxicogenomic multiverse: Convergent recruitment of proteins into animal venoms. Ann. Rev. Genom. Hum. Gen. 2009, 10, 483–511. [Google Scholar] [CrossRef] [PubMed]
- Norton, R.S.; Pallaghy, P.K. The cystine knot structure of ion channel toxins and related polypeptides. Toxicon 1998, 36, 1573–1583. [Google Scholar] [CrossRef]
- Undheim, E.A.; Mobli, M.; King, G.F. Toxin structures as evolutionary tools: Using conserved 3D folds to study the evolution of rapidly evolving peptides. Bioessays 2015, 38, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Dutertre, S.; Jin, A.-H.; Vetter, I.; Hamilton, B.; Sunagar, K.; Lavergne, V.; Dutertre, V.; Fry, B.G.; Antunes, A.; Venter, D.J.; et al. Evolution of separate predation- and defence-evoked venoms in carnivorous cone snails. Nat. Commun. 2014, 5, 3521. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Deuis, J.; Dashevsky, D.; Dobson, J.; Jackson, T.; Brust, A.; Xie, B.; Koludarov, I.; Debono, J.; Hendrikx, I.; et al. The snake with the scorpion’s sting: Novel three-finger toxin sodium channel activators from the venom of the long-glanded blue coral snake (Calliophis bivirgatus). Toxins 2016, 8, 303. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.A.; Hernández-Vargas, M.J.; Corzo, G.; Fry, B.G.; King, G.F. Giant fish-killing water bug reveals ancient and dynamic venom evolution in Heteroptera. Cell. Mol. Life Sci. 2018, 75, 3215–3229. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.C. Solid-to-liquid feeding: The inside(s) out story of extra-oral digestion in predaceous arthropods. Am. Entomol. 1998, 44, 103–116. [Google Scholar] [CrossRef]
- Escoubas, P.; Diochot, S.; Corzo, G. Structure and pharmacology of spider venom neurotoxins. Biochimie 2000, 82, 893–907. [Google Scholar] [CrossRef]
- Vassilevski, A.A.; Kozlov, S.A.; Grishin, E.V. Molecular diversity of spider venom. Biochemistry 2010, 74, 1505–1534. [Google Scholar] [CrossRef]
- Rungsa, P.; Incamnoi, P.; Sukprasert, S.; Uawonggul, N.; Klaynongsruang, S.; Daduang, J.; Patramanon, R.; Roytrakul, S.; Daduang, S. Comparative proteomic analysis of two wasps venom, Vespa tropica and Vespa affinis. Toxicon 2016, 119, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Pineda, S.S.; Sollod, B.L.; Wilson, D.; Darling, A.; Sunagar, K.; Undheim, E.A.; Kely, L.; Antunes, A.; Fry, B.G.; King, G.F. Diversification of a single ancestral gene into a successful toxin superfamily in highly venomous Australian funnel-web spiders. BMC Genom. 2014, 15, 177. [Google Scholar] [CrossRef] [PubMed]
- Sunagar, K.; Undheim, E.; Chan, A.; Koludarov, I.; Muñoz-Gómez, S.; Antunes, A.; Fry, B. Evolution stings: The origin and diversification of scorpion toxin peptide scaffolds. Toxins 2013, 5, 2456. [Google Scholar] [CrossRef] [PubMed]
- Nagao, J.; Miyashita, M.; Nakagawa, Y.; Miyagawa, H. Chemical synthesis of La1 isolated from the venom of the scorpion Liocheles australasiae and determination of its disulfide bonding pattern. J. Pept. Sci. 2015, 21, 636–643. [Google Scholar] [CrossRef] [PubMed]
- Fernández, J.; Gutiérrez, J.M.; Calvete, J.J.; Sanz, L.; Lomonte, B. Characterization of a novel snake venom component: Kazal-type inhibitor-like protein from the arboreal pitviper Bothriechis schlegelii. Biochimie 2016, 125, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Ye, X.-H.; Liu, Y.; Yan, Z.-C.; Stanley, D.; Ye, G.-Y.; Fang, Q. A venom gland extracellular chitin-binding-like protein from pupal endoparasitoid wasps, Pteromalus puparum, selectively binds chitin. Toxins 2015, 7, 4867. [Google Scholar] [CrossRef] [PubMed]
- Touchard, A.; Aili, S.R.; Fox, E.G.P.; Escoubas, P.; Orivel, J.; Nicholson, G.M.; Dejean, A. The biochemical toxin arsenal from ant venoms. Toxins 2016, 8, 30. [Google Scholar] [CrossRef] [PubMed]
- King, G.F.; Escoubas, P.; Nicholson, G.M. Peptide toxins that selectively target insect NaV and CaV channels. Channels 2008, 2, 1–17. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L. Antimicrobial and cytolytic peptides of venomous arthropods. Cell. Mol. Life Sci. 2003, 60, 2651–2668. [Google Scholar] [CrossRef] [PubMed]
- Dutertre, S.; Lewis, R.J. Use of venom peptides to probe ion channel structure and function. J. Biol. Chem. 2010, 285, 13315–13320. [Google Scholar] [CrossRef] [PubMed]
- King, G.F.; Hardy, M.G. Spider-venom peptides: Structure, pharmacology, and potential for control of insect pests. Ann. Rev. Entomol. 2013, 58, 475–496. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.D.; Undheim, E.A.B.; Ueberheide, B.; King, G.F. Venom peptides as therapeutics: Advances, challenges and the future of venom-peptide discovery. Exp. Rev. Proteom. 2017, 14, 931–939. [Google Scholar] [CrossRef] [PubMed]





| Peptide | Amino Acid Sequence | Amino Acids | Putative Fold | LC-MS/MS Monoisotopic Mass, Untreated (Da) | LC-MS/MS Monoisotopic Mass, after RAa (Da) | LC-MS/MS Abundance Rank b | MALDI Mono-Isotopic Mass b | MALDI Peak Signal Rank | Transcript Expression (FPKM) | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Predicted | Measured | Predicted | Measured | ||||||||
| U-Asilidin12-Dg3a | ITCDLIGNERLCVLHCLAKGFRGGWCDGRKVCNCRR | 36 | CS-α/β defensin | 4057.97 | 4057.87 | 4328.15 | 4328.04 | 1 | 4057.85 | 4 | 5439 |
| U-Asilidin12-Dg3b | ITCDLIGNERLCVVHCLAKGFRGGWCDSRKVCNCRR | 36 | CS-α/β defensin | 4073.97 | 4073.85 | 4344.15 | 4344.03 | 2 | 4073.85 | 7 | 2558 |
| U-Asilidin14-Dg4 | YDDHDPCFLKCKFWRSISRICVKTEDGTEKTLINESVLLCAKGCNRNWTLLHHGACPSDPGG | 62 | Kazal? | 6970.31 | 6970.17 | 7240.49 | 7241.37 | 25 | nd | nd | 26,171 |
| U-Asilidin1-Dg10a | KNDRECKKIAEVCYRHEECCSFQCPSYWGKCVS | 33 | ICK | 3921.70 | 3921.57 | 4191.88 | 4191.77 | 78 | 3921.5 | 9 | 2721 |
| U-Asilidin1-Dg10b | DRECKKIAEVCYRHEECCSFQCPSYWGKCVS | 31 | ICK | 3679.56 | 3679.45 | 3949.74 | 3949.64 | 11 | 3679.5 | 2 | 2721 |
| U-Asilidin1-Dg12 | SQEQRQCKKIGEHCYVADECCSKRCLFYAAKCVS | 34 | ICK | 3906.76 | 3906.64 | 4176.94 | 4176.83 | 28 | 3906.6 | 3 | 2507 |
| U-Asilidin15-Dg14a | RECPTVENEKDIAVHLPHKDCSKYYACVKGKKIERKCPRGLLFNKTLQVCDFPERVKC | 58 | Peritrophin domain | 6755.44 | 6755.28 | 7025.62 | 7026.48 | 26 | nd | nd | 1303 |
| Kazal protein Dg21 domain 2 | SDFCPEVCPLLYKPVCGSYGDIKKIFPNECELKRANCKFGEAWEKINMDICRNIS | 55 | Kazal | 6307.00 | 6306.9 | 6577.18 | 6577.09 | 24 | nd | nd | 2172 |
| U-Asilidin1-Dg27 | pyroQ-DCNPEGARCSSDSDCCYSECIGSLCQP | 27 | ICK | 2946.03 | 2945.94 | 3216.21 | nd | 116 | 2945.9 | 11 | 4377 |
| U-Asilidin1-Dg28 | RCVPRGGYCFNGSTLKCCRGVTTCINNRCR | 30 | ICK | 3330.54 | 3330.42 | 3600.72 | 3601.63 | 464 | nd | nd | 93 |
| U-Asilidin1-Dg29 | SFRCSERGEPCFLRTTLNCCSGVSACIKNQCRF | 33 | ICK | 3708.67 | 3709.56 | 3978.85 | 3978.74 | 51 | nd | nd | 1827 |
| Kazal protein Dg51 domain 4 | DFQKKCKLICPALYAPVCGFNGETYKWFQNKCIMEMDNCLFNHNWVADKMENCKA | 55 | Kazal | 6472.94 | 6473.8 | 6743.12 | 6742.94 | 31 | nd | nd | 2339 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Walker, A.A.; Dobson, J.; Jin, J.; Robinson, S.D.; Herzig, V.; Vetter, I.; King, G.F.; Fry, B.G. Buzz Kill: Function and Proteomic Composition of Venom from the Giant Assassin Fly Dolopus genitalis (Diptera: Asilidae). Toxins 2018, 10, 456. https://doi.org/10.3390/toxins10110456
Walker AA, Dobson J, Jin J, Robinson SD, Herzig V, Vetter I, King GF, Fry BG. Buzz Kill: Function and Proteomic Composition of Venom from the Giant Assassin Fly Dolopus genitalis (Diptera: Asilidae). Toxins. 2018; 10(11):456. https://doi.org/10.3390/toxins10110456
Chicago/Turabian StyleWalker, Andrew A., James Dobson, Jiayi Jin, Samuel D. Robinson, Volker Herzig, Irina Vetter, Glenn F. King, and Bryan G. Fry. 2018. "Buzz Kill: Function and Proteomic Composition of Venom from the Giant Assassin Fly Dolopus genitalis (Diptera: Asilidae)" Toxins 10, no. 11: 456. https://doi.org/10.3390/toxins10110456
APA StyleWalker, A. A., Dobson, J., Jin, J., Robinson, S. D., Herzig, V., Vetter, I., King, G. F., & Fry, B. G. (2018). Buzz Kill: Function and Proteomic Composition of Venom from the Giant Assassin Fly Dolopus genitalis (Diptera: Asilidae). Toxins, 10(11), 456. https://doi.org/10.3390/toxins10110456