Effects of Commercial Apple Varieties on Human Gut Microbiota Composition and Metabolic Output Using an In Vitro Colonic Model
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fecal Donors
2.2. Materials
2.3. Apples and Controls
2.4. Preparation of Phospholipid Vesicles
2.5. In Vitro Gastric and Duodenal Digestion
2.6. Fecal Batch-Culture Fermentation and Samples Collection
2.7. DNA Extraction, Amplification and Sequencing
2.8. Sequence Data Analysis
2.9. Enumeration of Bacterial Groups with Fluorescence In Situ Hybridization (FISH)
2.10. Analysis of Short Chain Fatty Acids (SCFAs)
2.11. Analysis of Precursor Polyphenols and Polyphenol Microbial Metabolites
2.12. Statistical Analysis
3. Results
3.1. Composition of Apples
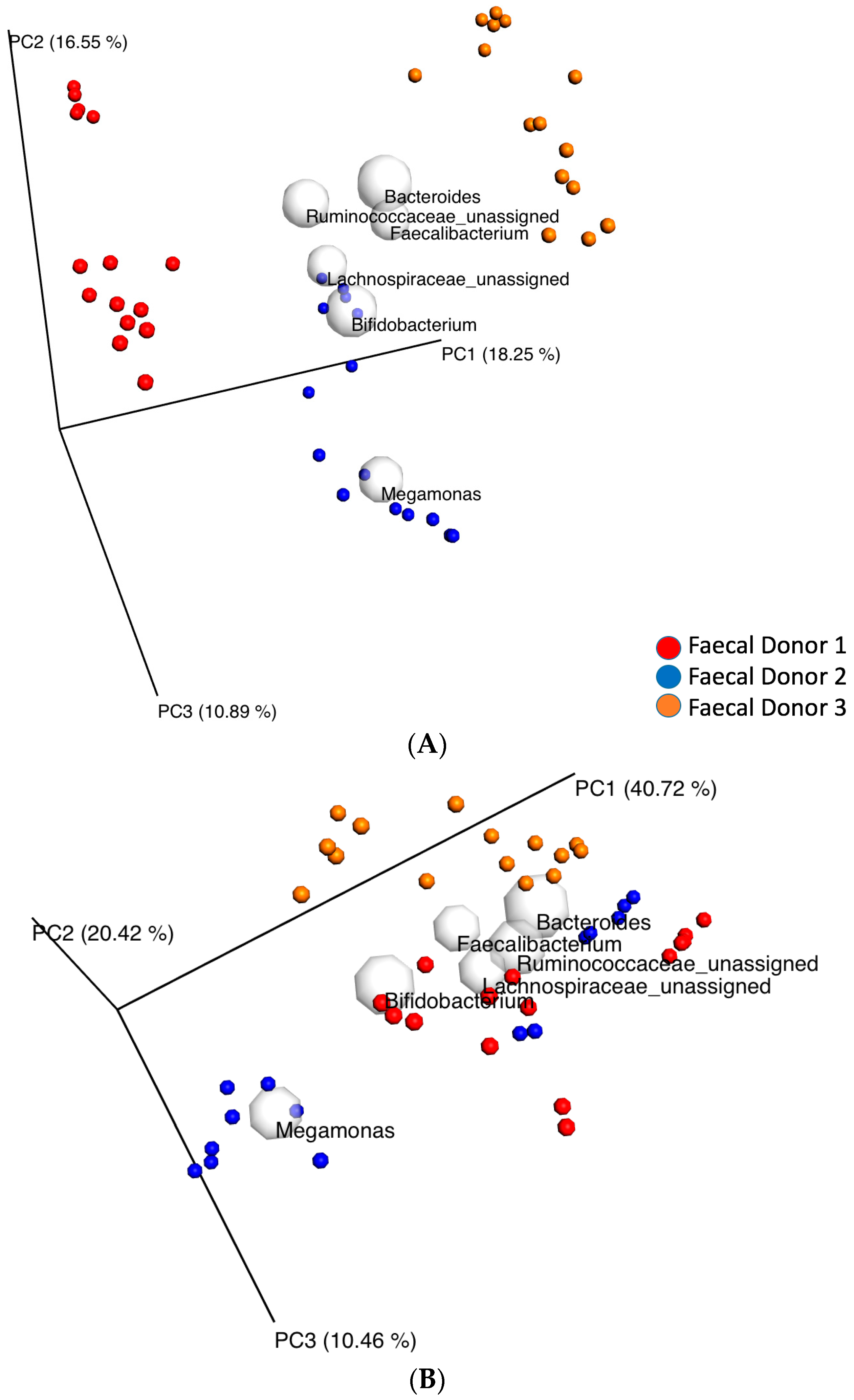
3.2. Changes in Fecal Bacterial Alpha and Beta Diversity
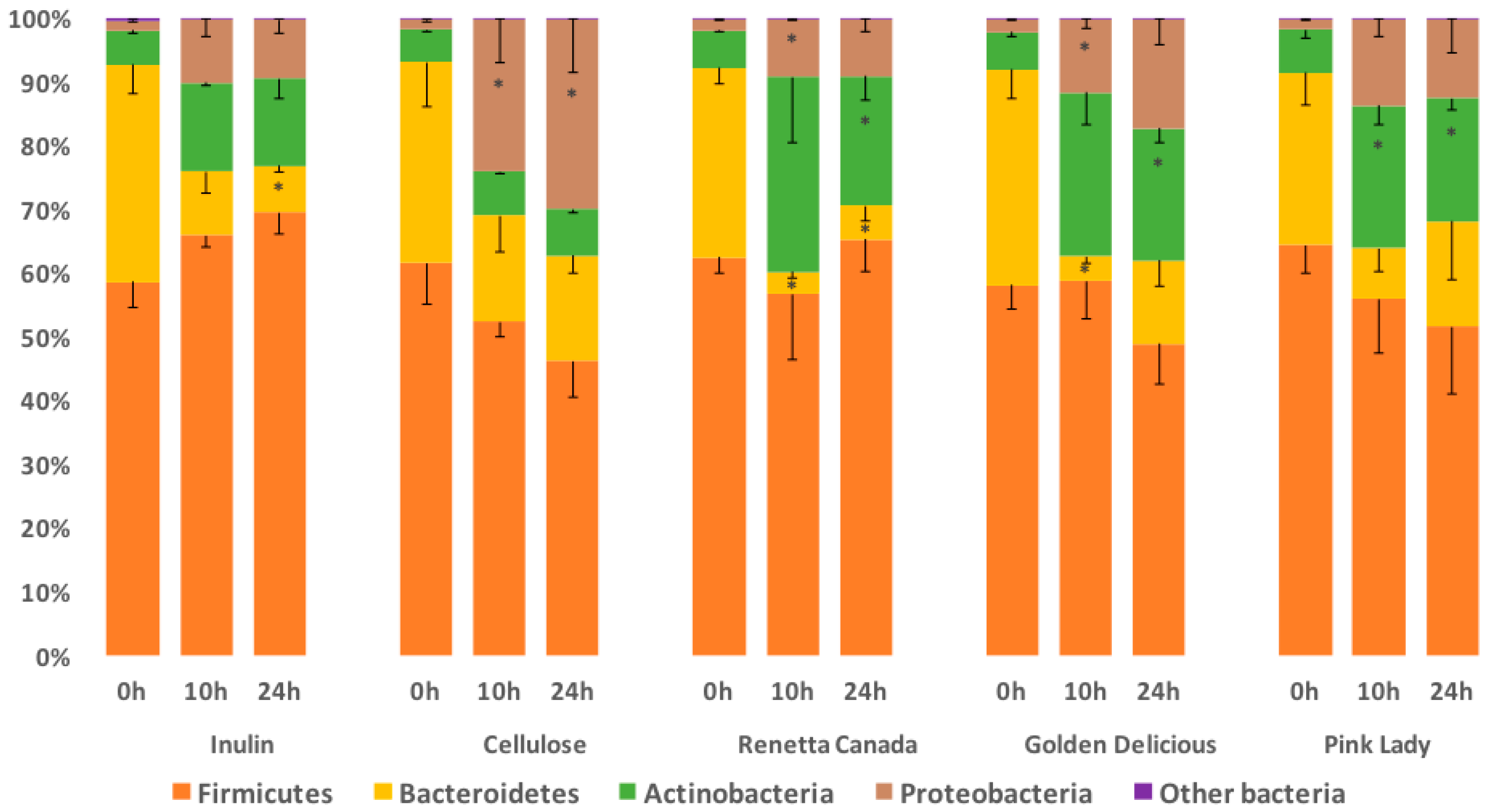
3.3. Fecal Bacterial Relative Abundance at the Phylum Level
3.4. Fecal Bacterial Relative Abundance at the Genus Level
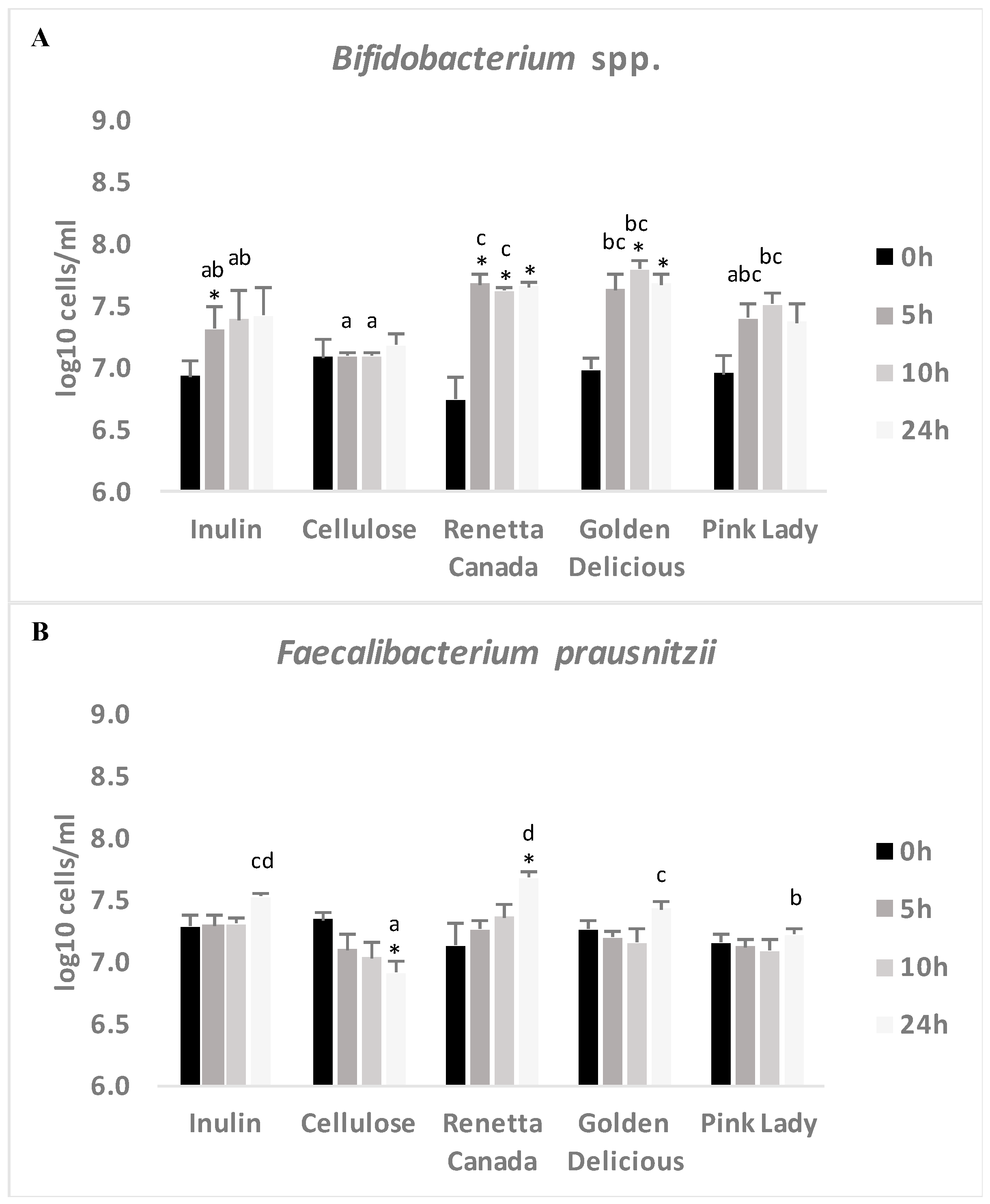
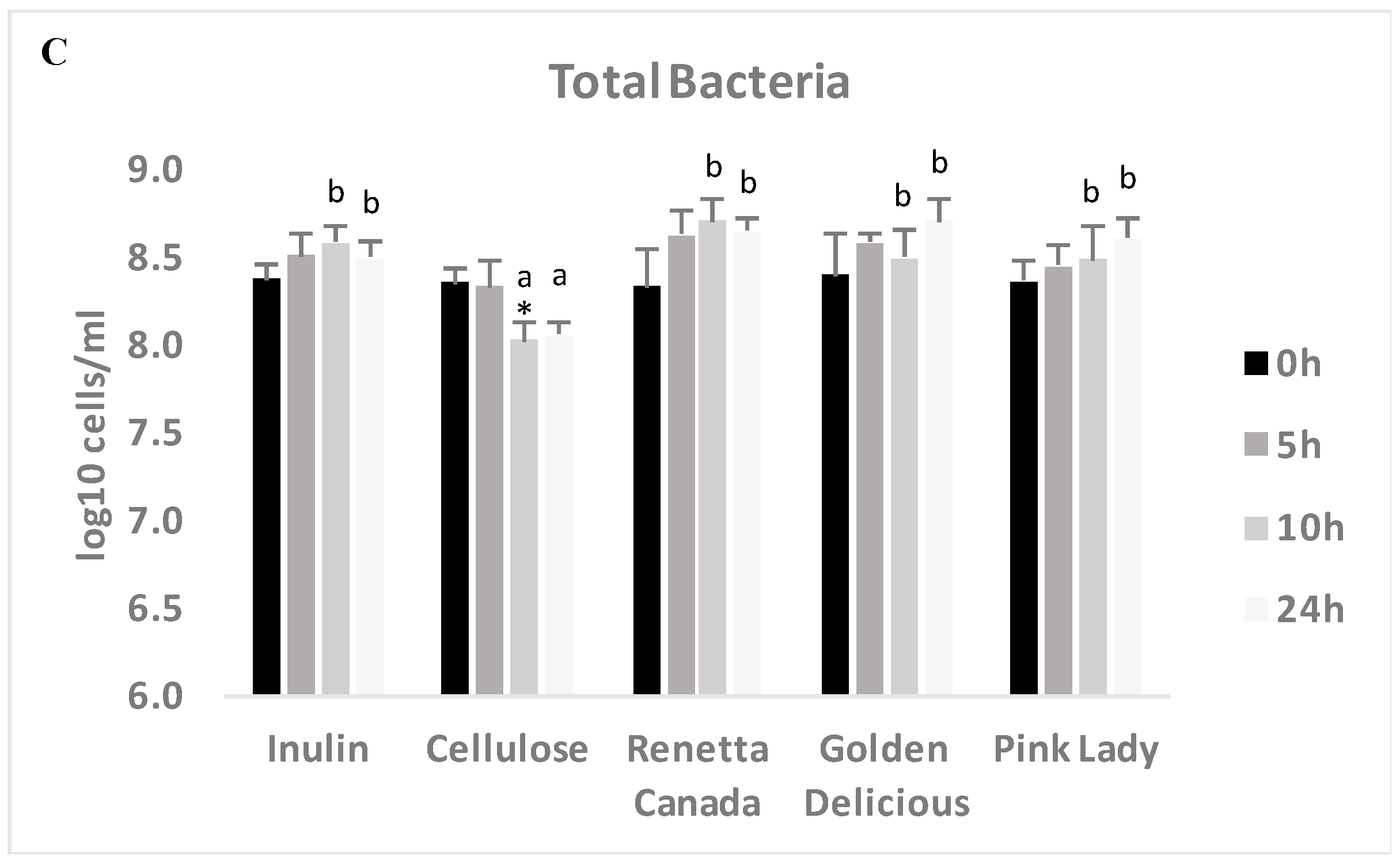
3.5. Changes in Selected Fecal Bacterial Populations Measured with FISH
3.6. SCFAs Production
3.7. Changes in Precursor Polyphenols
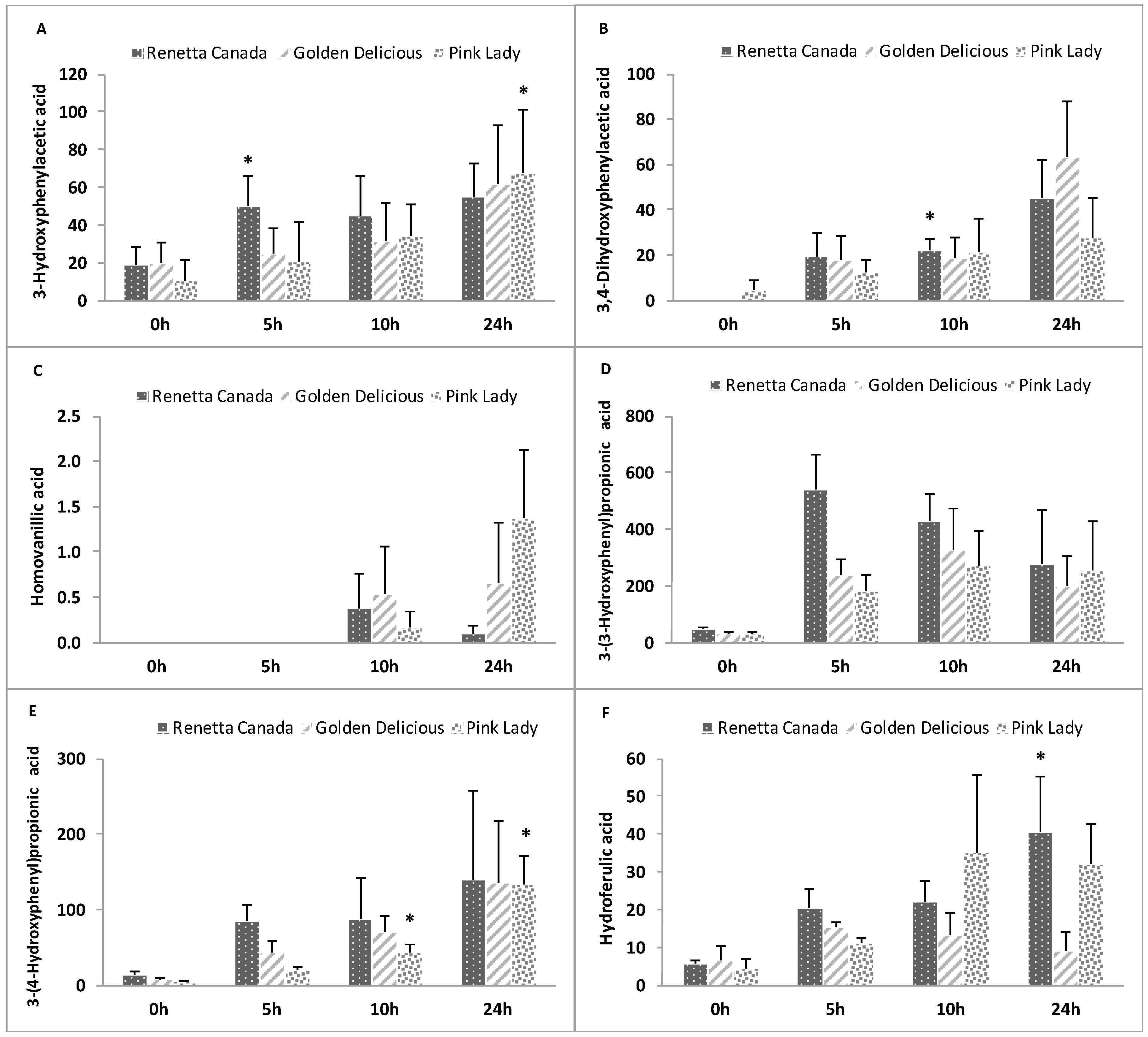
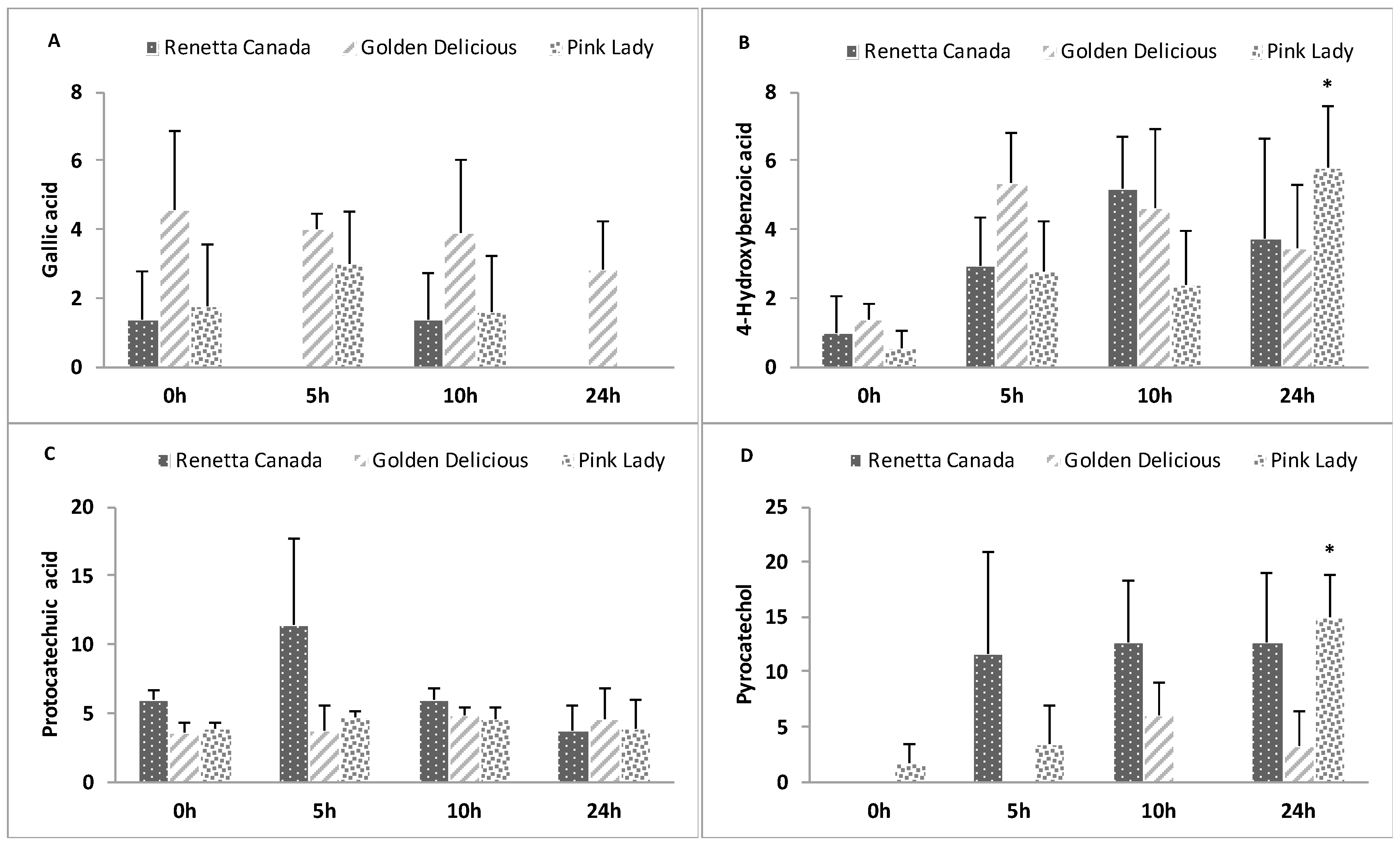
3.8. Formation of Polyphenol Microbial Metabolites
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, X.; Ouyang, Y.Y.; Liu, J.; Zhu, M.M.; Zhao, G.; Bao, W.; Hu, F.B. Fruit and vegetable consumption and mortality from all causes, cardiovascular disease, and cancer: Systematic review and dose-response meta-analysis of prospective cohort studies. BMJ 2014, 349, 14. [Google Scholar] [CrossRef] [PubMed]
- Koutsos, A.; Tuohy, K.M.; Lovegrove, J.A. Apples and cardiovascular health-is the gut microbiota a core consideration? Nutrients 2015, 7, 3959–3998. [Google Scholar] [CrossRef] [PubMed]
- Gerhauser, C. Cancer chemopreventive potential of apples, apple juice, and apple components. Planta Med. 2008, 74, 1608–1624. [Google Scholar] [CrossRef] [PubMed]
- Monagas, M.; Urpi-Sarda, M.; Sanchez-Patan, F.; Llorach, R.; Garrido, I.; Gomez-Cordoves, C.; Andres-Lacueva, C.; Bartolome, B. Insights into the metabolism and microbial biotransformation of dietary flavan-3-ols and the bioactivity of their metabolites. Food Funct. 2010, 1, 233–253. [Google Scholar] [CrossRef] [PubMed]
- Ozdal, T.; Sela, D.A.; Xiao, J.B.; Boyacioglu, D.; Chen, F.; Capanoglu, E. The reciprocal interactions between polyphenols and gut microbiota and effects on bioaccessibility. Nutrients 2016, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Olano-Martin, E.; Gibson, G.R.; Rastall, R.A. Comparison of the in vitro bifidogenic properties of pectins and pectic-oligosaccharides. J. Appl. Microbiol. 2002, 93, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Liang, R.-H.; Liu, W.; Li, T.; Liu, C.-M.; Wu, S.-S.; Wang, Z.-J. Pectic-oligosaccharides prepared by dynamic high-pressure nnicrofluidization and their in vitro fermentation properties. Carbohydr. Polym. 2013, 91, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Chung, W.S.F.; Walker, A.W.; Louis, P.; Parkhill, J.; Vermeiren, J.; Bosscher, D.; Duncan, S.H.; Flint, H.J. Modulation of the human gut microbiota by dietary fibres occurs at the species level. BMC Biol. 2016, 14, 13. [Google Scholar] [CrossRef] [PubMed]
- Vrhovsek, U.; Rigo, A.; Tonon, D.; Mattivi, F. Quantitation of polyphenols in different apple varieties. J. Agric. Food Chem. 2004, 52, 6532–6538. [Google Scholar] [CrossRef] [PubMed]
- Bazzocco, S.; Mattila, I.; Guyot, S.; Renard, C.M.G.C.; Aura, A.-M. Factors affecting the conversion of apple polyphenols to phenolic acids and fruit matrix to short-chain fatty acids by human faecal microbiota in vitro. Eur. J. Nutr. 2008, 47, 442–452. [Google Scholar] [CrossRef] [PubMed]
- Sembries, S.; Dongowski, G.; Jacobasch, G.; Mehrlander, K.; Will, F.; Dietrich, H. Effects of dietary fibre-rich juice colloids from apple pomace extraction juices on intestinal fermentation products and microbiota in rats. Br. J. Nutr. 2003, 90, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Sembries, S.; Dongowski, G.; Mehrlaender, K.; Will, F.; Dietrich, H. Physiological effects of extraction juices from apple, grape, and red beet pomaces in rats. J. Agric. Food Chem. 2006, 54, 10269–10280. [Google Scholar] [CrossRef] [PubMed]
- Licht, T.R.; Hansen, M.; Bergstrom, A.; Poulsen, M.; Krath, B.N.; Markowski, J.; Dragsted, L.O.; Wilcks, A. Effects of apples and specific apple components on the cecal environment of conventional rats: Role of apple pectin. BMC Microbiol. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Gao, X.; Wu, C.; Tian, F.; Lei, Q.; Bi, J.; Xie, B.; Wang, H.Y.; Chen, S.; Wang, X. Apple-derived pectin modulates gut microbiota, improves gut barrier function, and attenuates metabolic endotoxemia in rats with diet-induced obesity. Nutrients 2016, 8, 126. [Google Scholar] [CrossRef] [PubMed]
- Masumoto, S.; Terao, A.; Yamamoto, Y.; Mukai, T.; Miura, T.; Shoji, T. Non-absorbable apple procyanidins prevent obesity associated with gut microbial and metabolomic changes. Sci. Rep. 2016, 6, 31208. [Google Scholar] [CrossRef] [PubMed]
- Aprikian, O.; Duclos, V.; Guyot, S.; Besson, C.; Manach, C.; Bernalier, A.; Morand, C.; Remesy, C.; Demigne, C. Apple pectin and a polyphenol-rich apple concentrate are more effective together than separately on cecal fermentations and plasma lipids in rats. J. Nutr. 2003, 133, 1860–1865. [Google Scholar] [PubMed]
- Shinohara, K.; Ohashi, Y.; Kawasumi, K.; Terada, A.; Fujisawa, T. Effect of apple intake on fecal microbiota and metabolites in humans. Anaerobe 2010, 16, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Ravn-Haren, G.; Dragsted, L.O.; Buch-Andersen, T.; Jensen, E.N.; Jensen, R.I.; Nemeth-Balogh, M.; Paulovicsova, B.; Bergstrom, A.; Wilcks, A.; Licht, T.R.; et al. Intake of whole apples or clear apple juice has contrasting effects on plasma lipids in healthy volunteers. Eur. J. Nutr. 2012. [Google Scholar] [CrossRef] [PubMed]
- Vrhovsek, U.; Masuero, D.; Gasperotti, M.; Franceschi, P.; Caputi, L.; Viola, R.; Mattivi, F. A versatile targeted metabolomics method for the rapid quantification of multiple classes of phenolics in fruits and beverages. J. Agric. Food Chem. 2012, 60, 8831–8840. [Google Scholar] [CrossRef] [PubMed]
- Gasperotti, M.; Masuero, D.; Guella, G.; Mattivi, F.; Vrhovsek, U. Development of a targeted method for twenty-three metabolites related to polyphenol gut microbial metabolism in biological samples, using spe and uhplc-esi-ms/ms. Talanta 2014, 128, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Mandalari, G.; Faulks, R.M.; Rich, G.T.; Lo Turco, V.; Picout, D.R.; Lo Curto, R.B.; Bisignano, G.; Dugo, P.; Dugo, G.; Waldron, K.W.; et al. Release of protein, lipid, and vitamin e from almond seeds during digestion. J. Agric. Food Chem. 2008, 56, 3409–3416. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Patan, F.; Cueva, C.; Monagas, M.; Walton, G.E.; Gibson, G.R.; Quintanilla-Lopez, J.E.; Lebron-Aguilar, R.; Martin-Alvarez, P.J.; Moreno-Arribas, M.V.; Bartolome, B. In vitro fermentation of a red wine extract by human gut microbiota: Changes in microbial groups and formation of phenolic metabolites. J. Agric. Food Chem. 2012, 60, 2136–2147. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Aronesty, E. Comparison of sequencing utilitiy programs. Open Bioinform. J. 2013, 7, 1–8. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than blast. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Langendijk, P.S.; Schut, F.; Jansen, G.J.; Raangs, G.C.; Kamphuis, G.R.; Wilkinson, M.H.F.; Welling, G.W. Quantitative fluorescence in-situ hybridization of Bifidobacterium spp. with genus-specific 16s ribosomal-rna-targeted probes and its application in fecal samples. Appl. Environ. Microbiol. 1995, 61, 3069–3075. [Google Scholar] [PubMed]
- Hold, G.L.; Schwiertz, A.; Aminov, R.I.; Blaut, M.; Flint, H.J. Oligonucleotide probes that detect quantitatively significant groups of butyrate-producing bacteria in human feces. Appl. Environ. Microbiol. 2003, 69, 4320–4324. [Google Scholar] [CrossRef] [PubMed]
- Daims, H.; Stoecker, K.; Wagner, M. Fluorescence In Situ Hybridization for the detection of prokaryotes. In Molecular Microbial Ecology; Osborn, A.M., Smith, C.J., Eds.; Taylor & Francis: New York, NY, USA, 2005. [Google Scholar]
- Zhao, G.H.; Nyman, M.; Jonsson, J.A. Rapid determination of short-chain fatty acids in colonic contents and faeces of humans and rats by acidified water-extraction and direct-injection gas chromatography. Biomed. Chromatogr. 2006, 20, 674–682. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Martinez, I.; Walter, J.; Keshavarzian, A.; Rose, D.J. In vitro characterization of the impact of selected dietary fibers on fecal microbiota composition and short chain fatty acid production. Anaerobe 2013, 23, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Martinez, I.; Kim, J.; Duffy, P.R.; Schlegel, V.L.; Walter, J. Resistant starches types 2 and 4 have differential effects on the composition of the fecal microbiota in human subjects. PLoS ONE 2010, 5, 11. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.P.; Henning, S.M.; Lee, R.P.; Lu, Q.Y.; Summanen, P.H.; Thames, G.; Corbett, K.; Downes, J.; Tseng, C.H.; Finegold, S.M.; et al. Pomegranate extract induces ellagitannin metabolite formation and changes stool microbiota in healthy volunteers. Food Funct. 2015, 6, 2487–2495. [Google Scholar] [CrossRef] [PubMed]
- Lacombe, A.; Li, R.W.; Klimis-Zacas, D.; Kristo, A.S.; Tadepalli, S.; Krauss, E.; Young, R.; Wu, V.C.H. Lowbush wild blueberries have the potential to modify gut microbiota and xenobiotic metabolism in the rat colon. PLoS ONE 2013, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.; Corradini, C.; Amaretti, A.; Nicolini, M.; Pompei, A.; Zanoni, S.; Matteuzzi, D. Fermentation of fructooligosaccharides and inulin by bifidobacteria: A comparative study of pure and fecal cultures. Appl. Environ. Microbiol. 2005, 71, 6150–6158. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermudez-Humaran, L.G.; Gratadoux, J.-J.; Blugeon, S.; Bridonneau, C.; Furet, J.-P.; Corthier, G.; et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of crohn disease patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef] [PubMed]
- Machiels, K.; Joossens, M.; Sabino, J.; De Preter, V.; Arijs, I.; Eeckhaut, V.; Ballet, V.; Claes, K.; Van Immerseel, F.; Verbeke, K.; et al. A decrease of the butyrate-producing species roseburia hominis and faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 2014, 63, 1275–1283. [Google Scholar] [CrossRef] [PubMed]
- Pryde, S.E.; Duncan, S.H.; Hold, G.L.; Stewart, C.S.; Flint, H.J. The microbiology of butyrate formation in the human colon. FEMS Microbiol. Lett. 2002, 217, 133–139. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Khan, T.M.; Duncan, S.H.; Harmsen, H.J.M.; Garcia-Gil, L.J.; Flint, H.J. Cultured representatives of two major phylogroups of human colonic faecalibacterium prausnitzii can utilize pectin, uronic acids, and host-derived substrates for growth. Appl. Environ. Microbiol. 2012, 78, 420–428. [Google Scholar] [CrossRef] [PubMed]
- Waldecker, M.; Kautenburger, T.; Daumann, H.; Busch, C.; Schrenk, D. Inhibition of histone-deacetylase activity by short-chain fatty acids and some polyphenol metabolites formed in the colon. J. Nutr. Biochem. 2008, 19, 587–593. [Google Scholar] [CrossRef] [PubMed]
- Dolara, P.; Luceri, C.; de Filippo, C.; Femia, A.P.; Giovannelli, L.; Caderni, G.; Cecchini, C.; Silvi, S.; Orpianesi, C.; Cresci, A. Red wine polyphenols influence carcinogenesis, intestinal microflora, oxidative damage and gene expression profiles of colonic mucosa in f344 rats. Mutat. Res.-Fundam. Mol. Mech. Mutag. 2005, 591, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Massot-Cladera, M.; Perez-Berezo, T.; Franch, A.; Castell, M.; Perez-Cano, F.J. Cocoa modulatory effect on rat faecal microbiota and colonic crosstalk. Arch. Biochem. Biophys. 2012, 527, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Kemperman, R.A.; Gross, G.; Mondot, S.; Possemiers, S.; Marzorati, M.; van de Wiele, T.; Dore, J.; Vaughan, E.E. Impact of polyphenols from black tea and red wine/grape juice on a gut model microbiome. Food Res. Int. 2013, 53, 659–669. [Google Scholar] [CrossRef]
- Eid, N.; Enani, S.; Walton, G.; Corona, G.; Costabile, A.; Gibson, G.; Rowland, I.; Spencer, J.P. The impact of date palm fruits and their component polyphenols, on gut microbial ecology, bacterial metabolites and colon cancer cell proliferation. J. Nutr. Sci. 2014, 3, e46. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.H.; Mackie, R.I. Effect of condensed tannins on bacterial diversity and metabolic activity in the rat gastrointestinal tract. Appl. Environ. Microbiol. 2004, 70, 1104–1115. [Google Scholar] [CrossRef] [PubMed]
- Isabel Queipo-Ortuno, M.; Boto-Ordonez, M.; Murri, M.; Miguel Gomez-Zumaquero, J.; Clemente-Postigo, M.; Estruch, R.; Cardona Diaz, F.; Andres-Lacueva, C.; Tinahones, F.J. Influence of red wine polyphenols and ethanol on the gut microbiota ecology and biochemical biomarkers. Am. J. Clin. Nutr. 2012, 95, 1323–1334. [Google Scholar] [CrossRef] [PubMed]
- Lyte, M.; Chapel, A.; Lyte, J.M.; Ai, Y.F.; Proctor, A.; Jane, J.L.; Phillips, G.J. Resistant starch alters the microbiota-gut brain axis: Implications for dietary modulation of behavior. PLoS ONE 2016, 11, 22. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.W.; Ince, J.; Duncan, S.H.; Webster, L.M.; Holtrop, G.; Ze, X.L.; Brown, D.; Stares, M.D.; Scott, P.; Bergerat, A.; et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J. 2011, 5, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Salonen, A.; Lahti, L.; Salojarvi, J.; Holtrop, G.; Korpela, K.; Duncan, S.H.; Date, P.; Farquharson, F.; Johnstone, A.M.; Lobley, G.E.; et al. Impact of diet and individual variation on intestinal microbiota composition and fermentation products in obese men. ISME J. 2014, 8, 2218–2230. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.W.; Duncan, S.H.; McWilliam Leitch, E.C.; Child, M.W.; Flint, H.J. Ph and peptide supply can radically alter bacterial populations and short-chain fatty acid ratios within microbial communities from the human colon. Appl. Environ. Microbiol. 2005, 71, 3692–3700. [Google Scholar] [CrossRef] [PubMed]
- Zambell, K.L.; Fitch, M.D.; Fleming, S.E. Acetate and butyrate are the major substrates for de novo lipogenesis in rat colonic epithelial cells. J. Nutr. 2003, 133, 3509–3515. [Google Scholar] [PubMed]
- Onumpai, C.; Kolida, S.; Bonnin, E.; Rastall, R.A. Microbial utilization and selectivity of pectin fractions with various structures. Appl. Environ. Microbiol. 2011, 77, 5747–5754. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Barcenilla, A.; Stewart, C.S.; Pryde, S.E.; Flint, H.J. Acetate utilization and butyryl coenzyme a (coa): Acetate-coa transferase in butyrate-producing bacteria from the human large intestine. Appl. Environ. Microbiol. 2002, 68, 5186–5190. [Google Scholar] [CrossRef] [PubMed]
- Rios-Covian, D.; Gueimonde, M.; Duncan, S.H.; Flint, H.J.; de los Reyes-Gavilan, C.G. Enhanced butyrate formation by cross-feeding between faecalibacterium prausnitzii and bifidobacterium adolescentis. FEMS Microbiol. Lett. 2015, 362. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.H.; Lai, M.H. Fermentation of resistant rice starch produces propionate reducing serum and hepatic cholesterol in rats. J. Nutr. 2000, 130, 1991–1995. [Google Scholar] [PubMed]
- Velderrain-Rodriguez, G.R.; Palafox-Carlos, H.; Wall-Medrano, A.; Ayala-Zavala, J.F.; Chen, C.Y.O.; Robles-Sanchez, M.; Astiazaran-Garcia, H.; Alvarez-Parrilla, E.; Gonzalez-Aguilar, G.A. Phenolic compounds: Their journey after intake. Food Funct. 2014, 5, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Larrosa, M.; Luceri, C.; Vivoli, E.; Pagliuca, C.; Lodovici, M.; Moneti, G.; Dolara, P. Polyphenol metabolites from colonic microbiota exert anti-inflammatory activity on different inflammation models. Mol. Nutr. Food Res. 2009, 53, 1044–1054. [Google Scholar] [CrossRef] [PubMed]
- Appeldoorn, M.M.; Vincken, J.P.; Aura, A.M.; Hollman, P.C.H.; Gruppen, H. Procyanidin dimers are metabolized by human microbiota with 2-(3,4-dihydroxyphenyl)acetic acid and 5-(3,4-dihydroxyphenyl)-gamma-valerolactone as the major metabolites. J. Agric. Food Chem. 2009, 57, 1084–1092. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Ruiz, J.A.; Leake, D.S.; Ames, J.M. In vitro antioxidant activity of coffee compounds and their metabolites. J. Agric. Food Chem. 2007, 55, 6962–6969. [Google Scholar] [CrossRef] [PubMed]
- Cueva, C.; Sanchez-Patan, F.; Monagas, M.; Walton, G.E.; Gibson, G.R.; Martin-Alvarez, P.J.; Bartolome, B.; Moreno-Arribas, M.V. In vitro fermentation of grape seed flavan-3-ol fractions by human faecal microbiota: Changes in microbial groups and phenolic metabolites. FEMS Microbiol. Ecol. 2013, 83, 792–805. [Google Scholar] [CrossRef] [PubMed]
- Selma, M.V.; Espin, J.C.; Tomas-Barberan, F.A. Interaction between phenolics and gut microbiota: Role in human health. J. Agric. Food Chem. 2009, 57, 6485–6501. [Google Scholar] [CrossRef] [PubMed]
- Dall’Asta, M.; Calani, L.; Tedeschi, M.; Jechiu, L.; Brighenti, F.; Del Rio, D. Identification of microbial metabolites derived from in vitro fecal fermentation of different polyphenolic food sources. Nutrition 2012, 28, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Deprez, S.; Brezillon, C.; Rabot, S.; Philippe, C.; Mila, I.; Lapierre, C.; Scalbert, A. Polymeric proanthocyanidins are catabolized by human colonic microflora into low-molecular-weight phenolic acids. J. Nutr. 2000, 130, 2733–2738. [Google Scholar] [PubMed]
- Rios, L.Y.; Gonthier, M.P.; Remesy, C.; Mila, L.; Lapierre, C.; Lazarus, S.A.; Williamson, G.; Scalbert, A. Chocolate intake increases urinary excretion of polyphenol-derived phenolic acids in healthy human subjects. Am. J. Clin. Nutr. 2003, 77, 912–918. [Google Scholar] [PubMed]
- Williams, C.F.; Walton, G.E.; Jiang, L.; Plummer, S.; Garaiova, I.; Gibson, G.R. Comparative analysis of intestinal tract models. In Annual Review of Food Science and Technology; Doyle, M.P., Klaenhammer, T.R., Eds.; Annual Reviews: Palo Alto, CA, USA, 2015; Volume 6, pp. 329–350. [Google Scholar]
- Johnson, L.P.; Walton, G.E.; Psichas, A.; Frost, G.S.; Gibson, G.R.; Barraclough, T.G. Prebiotics modulate the effects of antibiotics on gut microbial diversity and functioning in vitro. Nutrients 2015, 7, 4480–4497. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, D.; Walton, G.; Sousa, S.; Rocha-Santos, T.A.P.; Duarte, A.C.; Freitas, A.C.; Gomes, A.M.P. In vitro fermentation and prebiotic potential of selected extracts from seaweeds and mushrooms. LWT -Food Sci. Technol. 2016, 73, 131–139. [Google Scholar] [CrossRef]
- Blatchford, P.; Stoklosinski, H.; Walton, G.; Swann, J.; Gibson, G.; Gearry, R.; Ansell, J. Kiwifruit fermentation drives positive gut microbial and metabolic changes irrespective of initial microbiota composition. Bioact. Carbohydr. Diet. Fibre 2015, 6, 37–45. [Google Scholar] [CrossRef]
- Walton, G.E.; van den Heuvel, E.; Kosters, M.H.W.; Rastall, R.A.; Tuohy, K.M.; Gibson, G.R. A randomised crossover study investigating the effects of galacto-oligosaccharides on the faecal microbiota in men and women over 50 years of age. Br. J. Nutr. 2012, 107, 1466–1475. [Google Scholar] [CrossRef] [PubMed]
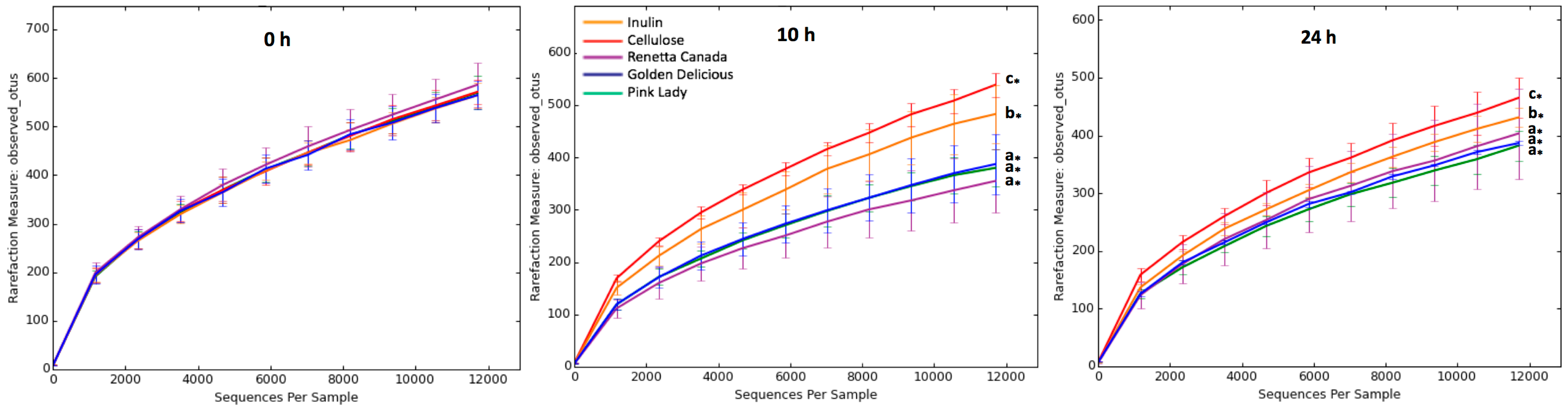
| Treatment | Chao1 (0 h) | Shannon (0 h) | Chao1 (10 h) | Shannon (10 h) | Chao1 (24 h) | Shannon (24 h) | ||
|---|---|---|---|---|---|---|---|---|
| Inulin | 1076 ± 43 | 6.36 ± 0.36 | 931 ± 91 b,* | 5.32 ± 0.53 b,* | 811 ± 37 b,* | 4.88 ± 0.32 a,* | ||
| Cellulose | 1091 ± 26 | 6.39 ± 0.43 | 1030 ± 66 c,* | 5.99 ± 0.27 c,* | 849 ± 89 c,* | 5.72 ± 0.29 c,* | ||
| Renetta Canada | 1124 ± 84 | 6.49 ± 0.28 | 673 ± 113 a,* | 4.28 ± 0.71 a,* | 774 ± 135 a,* | 4.78 ± 0.63 a,* | ||
| Golden Delicious | 1064 ± 46 | 6.35 ± 0.41 | 724 ± 125 a,* | 4.57 ± 0.64 a,* | 737 ± 31 a,* | 5.20 ± 0.07 b,* | ||
| Pink Lady | 1115 ± 67 | 6.47 ± 0.27 | 748 ± 69 a,* | 4.63 ± 0.66 a,* | 769 ± 48 a,* | 5.14 ± 0.47 b,* |
| Components | Renetta Canada | Golden Delicious | Pink Lady |
|---|---|---|---|
| Total dietary fiber (AOAC) (g/100 g) | 2.6 | 2.4 | 2.4 |
| Soluble fiber (AOAC) (g/100 g) | 1.6 | 1.3 | 0.9 |
| Insoluble fiber (AOAC) (g/100 g) | 1.0 | 1.1 | 1.5 |
| Polyphenols (mg/100 g) | |||
| Flavanols | |||
| (+)—Catechin | 1.07 | 0.16 | 0.17 |
| (−)—Epicatechin | 10.9 | 2.8 | 2.8 |
| Procyanidin B1 | 6.6 | 0.95 | 0.78 |
| Procyanidin B2 + B4 (as B2) | 18.3 | 6.1 | 4.8 |
| Proanthocyanidin (as cyanidin) | 169.2 | 91.5 | 62.1 |
| Hydroxycinnamates | |||
| Chlorogenic acid | 61 | 18.7 | 17.5 |
| Neochlorogenic acid | 0.04 | 0.04 | 0.01 |
| Cryptochlorogenic acid | 0.98 | 0.67 | 0.15 |
| Flavonols | |||
| Quercetin-3-glucoside | 0.99 | 6.9 | 3.1 |
| Quercetin-3-rhamnoside | 0.65 | 2.7 | 1.7 |
| Rutin | 0.09 | 0.44 | 0.19 |
| Kaempferol-3-rutinoside | 0.002 | 0.011 | 0.002 |
| Isorhamnetin-3-glucoside | 0.001 | 0.002 | 0.001 |
| Dihydrochalcones | |||
| Phlorizin | 5.9 | 1.5 | 0.8 |
| Anthocyanins | |||
| Cyanidin 3-galactoside | 0.006 | 0.017 | 0.027 |
| Benzoic Acid Derivatives | |||
| Vanillin | 0.008 | 0.006 | 0.003 |
| Vanillic acid | 0.001 | 0.003 | 0.002 |
| Phylum | Class | Order | Family | Genus | Time Point | Inulin (%) | Cellulose (%) | Renetta Canada (%) | Golden Delicious (%) | Pink Lady (%) | p * | p # (FDR-Corrected) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Firmicutes | Clostridia | Clostridiales | Lachnospiraceae | unassigned | 10 h | 4.3 ± 1.3 a | 9 ± 0.9 b | 2.6 ± 0.7 a | 5 ± 2 a,b | 2.9 ± 1.2 a | 0.040 | 0.336 |
| Firmicutes | Clostridia | Clostridiales | [Mogibacteriaceae] | unassigned | 10 h | 0.15 ± 0.01 b | 0.14 ± 0.01 b | 0.08 ± 0.02 a | 0.05 ± 0.01 a | 0.06 ± 0.01 a | 0.001 | 0.040 |
| Firmicutes | Clostridia | Clostridiales | Ruminococcaceae | Oscillospira | 10 h | 0.66 ± 0.1 b | 1 ± 0.1 c | 0.32 ± 0.03 a | 0.3 ± 0.1 a | 0.23 ± 0.07 a | 0.001 | 0.052 |
| Firmicutes | Clostridia | Clostridiales | Ruminococcaceae | Ruminococcus | 10 h | 1.4 ± 0.3 a,b | 1.8 ± 0.5 b | 0.63 ± 0.43 a | 0.41 ± 0.18 a | 0.45 ± 0.17 a | 0.050 | 0.369 |
| Firmicutes | Clostridia | Clostridiales | unassigned | unassigned | 10 h | 1.8 ± 0.1 b | 2.8 ± 0.3 c | 0.82 ± 0.24 a | 0.76 ± 0.16 a | 0.81 ± 0.12 a | 0.000 | 0.009 |
| Firmicutes | Clostridia | Clostridiales | unclassified | unclassified | 10 h | 0.68 ± 0.12 b | 1.4 ± 0.23 c | 0.47 ± 0.1 a,b | 0.33 ± 0.03 a,b | 0.59 ± 0.27 a,b | 0.013 | 0.215 |
| Bacteroidetes | Bacteroidia | Bacteroidales | Porphyromonadaceae | Parabacteroides | 10 h | 0.72 ± 0.2 b | 1.9 ± 0.6 c | 0.4 ± 0.1 a,b | 0.37 ± 0.04 a,b | 0.56 ± 0.1 a,b | 0.022 | 0.264 |
| Actinobacteria | Actinobacteria | Bifidobacteriales | Bifidobacteriaceae | Bifidobacterium | 10 h | 4.7 ± 2.4 a | 2.8 ± 0.2 a | 24.3 ± 8.7 b | 19.4 ± 4.2 b | 16.2 ± 2.1 a,b | 0.028 | 0.273 |
| Proteobacteria | Deltaproteobacteria | Desulfovibrionales | Desulfovibrionaceae | Bilophila | 10 h | 0.33 ± 0.03 a | 1.7 ± 0.6 b | 0.17 ± 0.01 a | 0.24 ± 0.08 a | 0.26 ± 0.05 a | 0.024 | 0.264 |
| Firmicutes | Clostridia | Clostridiales | Ruminococcaceae | Faecalibacterium | 24 h | 17 ± 3.7 c | 4.6 ± 2.2 a | 16 ± 5.5 b,c | 5.9 ± 1.9 a,b | 5.4 ± 1.4 a | 0.049 | 0.950 |
| Firmicutes | Clostridia | Clostridiales | unassigned | unassigned | 24 h | 1.2 ± 0.3 a | 2.5 ± 0.6 b | 0.36 ± 0.05 a | 0.75 ± 0.19 a | 0.86 ± 0.19 a | 0.007 | 0.482 |
| Bacteroidetes | Bacteroidia | Bacteroidales | [Odoribacteraceae] | Butyricimonas | 24 h | 0.04 ± 0.01 a,b | 0.09 ± 0.03 b | 0.03 ± 0.01 a | 0.02 ± 0.01 a | 0.02 ± 0.01 a | 0.044 | 0.950 |
| Actinobacteria | Actinobacteria | Bifidobacteriales | Bifidobacteriaceae | Bifidobacterium | 24 h | 6.3 ± 4.6 a,b | 2.6 ± 0.2 a | 14.9 ± 2.5 b | 15.2 ± 2.6 b | 10.4 ± 3 a,b | 0.050 | 0.950 |
| Substrate | Time (h) | Acetic Acid (mmol/L) | p a | Propionic Acid (mmol/L) | p a | Butyric Acid (mmol/L) | p a | Total SCFAs (mmol/L) | p a |
|---|---|---|---|---|---|---|---|---|---|
| Inulin | 0 | 2.2 ± 0.1 | 0.050 | 0.8 ± 0.1 | 0.185 | 1 ± 0.3 | 0.013 | 4.1 ± 0.6 | 0.049 |
| 5 | 9.5 ± 5.4 | 2.8 ± 1.4 | 4.4 ± 2.8 | 16.8 ± 9.6 | |||||
| 10 | 11 ± 2.1 | 2.9 ± 0.8 | 5.5 ± 1 * | 19.3 ± 3.3 * | |||||
| 24 | 17.3 ± 2.1 * | 6.9 ± 2.6 | 11 ± 0.5 * | 35.4 ± 4.6 * | |||||
| Cellulose | 0 | 4.8 ± 2.9 | 0.062 | 1.4 ± 0.8 | 0.062 | 1.2 ± 0.6 | 0.041 | 7.5 ± 4.2 | 0.022 |
| 5 | 5 ± 1.3 | 1.9 ± 0.5 | 2.3 ± 0.2 * | 9.2 ± 2.3 | |||||
| 10 | 13.5 ± 5.2 | 3.3 ± 1 | 2.8 ± 0.6 | 19.5 ± 5.6 * | |||||
| 24 | 13.1 ± 1.1 | 3.4 ± 0.1 | 3.5 ± 1 | 20 ± 1.5 | |||||
| Renetta Canada | 0 | 1 ± 0.1 | 0.034 | 0.5 ± 0.1 | 0.020 | 0.5 ± 0.1 | 0.025 | 3 ± 0.3 | 0.044 |
| 5 | 11.5 ± 2.7 | 4.6 ± 2.6 | 2 ± 0.5 | 18.1 ± 3.9 | |||||
| 10 | 19 ± 2.2 * | 5.6 ± 2.1 | 3.7 ± 0.6 *,# | 28.4 ± 1.8 * | |||||
| 24 | 28.1 ± 6 *,# | 8.8 ± 2.6 #,^ | 16.9 ± 6.1 | 53.7 ± 11.4 * | |||||
| Golden Delicious | 0 | 2 ± 0.4 | 0.034 | 0.6 ± 0.1 | 0.016 | 0.5 ± 0.2 | 0.057 | 3.1 ± 0.7 | 0.008 |
| 5 | 13.9 ± 5.8 | 4.2 ± 1.6 | 1.9 ± 0.7 | 20 ± 6.7 | |||||
| 10 | 15.3 ± 3 * | 4.7 ± 1.9 | 4.9 ± 1.6 | 25 ± 3.1 * | |||||
| 24 | 23.7 ± 2.8 * | 7.8 ± 1.9 #,^ | 13.3 ± 5.1 | 44.8 ± 7.2 *,# | |||||
| Pink Lady | 0 | 1.8 ± 0.3 | 0.049 | 0.5 ± 0.1 | 0.020 | 0.4 ± 0.1 | 0.044 | 2.7 ± 0.5 | 0.040 |
| 5 | 8.3 ± 2.3 | 3.2 ± 1.5 | 1.1 ± 0.2 | 12.6 ± 2.9 | |||||
| 10 | 22.8 ± 8 | 6 ± 2.2 | 4 ± 1.2 | 32.8 ± 9.7 | |||||
| 24 | 26.2 ± 3.8 *,# | 8.8 ± 2 #,^ | 13 ± 2.8 | 48 ± 7.9 *,#,^ |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koutsos, A.; Lima, M.; Conterno, L.; Gasperotti, M.; Bianchi, M.; Fava, F.; Vrhovsek, U.; Lovegrove, J.A.; Tuohy, K.M. Effects of Commercial Apple Varieties on Human Gut Microbiota Composition and Metabolic Output Using an In Vitro Colonic Model. Nutrients 2017, 9, 533. https://doi.org/10.3390/nu9060533
Koutsos A, Lima M, Conterno L, Gasperotti M, Bianchi M, Fava F, Vrhovsek U, Lovegrove JA, Tuohy KM. Effects of Commercial Apple Varieties on Human Gut Microbiota Composition and Metabolic Output Using an In Vitro Colonic Model. Nutrients. 2017; 9(6):533. https://doi.org/10.3390/nu9060533
Chicago/Turabian StyleKoutsos, Athanasios, Maria Lima, Lorenza Conterno, Mattia Gasperotti, Martina Bianchi, Francesca Fava, Urska Vrhovsek, Julie A. Lovegrove, and Kieran M. Tuohy. 2017. "Effects of Commercial Apple Varieties on Human Gut Microbiota Composition and Metabolic Output Using an In Vitro Colonic Model" Nutrients 9, no. 6: 533. https://doi.org/10.3390/nu9060533
APA StyleKoutsos, A., Lima, M., Conterno, L., Gasperotti, M., Bianchi, M., Fava, F., Vrhovsek, U., Lovegrove, J. A., & Tuohy, K. M. (2017). Effects of Commercial Apple Varieties on Human Gut Microbiota Composition and Metabolic Output Using an In Vitro Colonic Model. Nutrients, 9(6), 533. https://doi.org/10.3390/nu9060533





