Evaluating Changes in Omega-3 Fatty Acid Intake after Receiving Personal FADS1 Genetic Information: A Randomized Nutrigenetic Intervention
Abstract
:1. Introduction
2. Materials and Methods
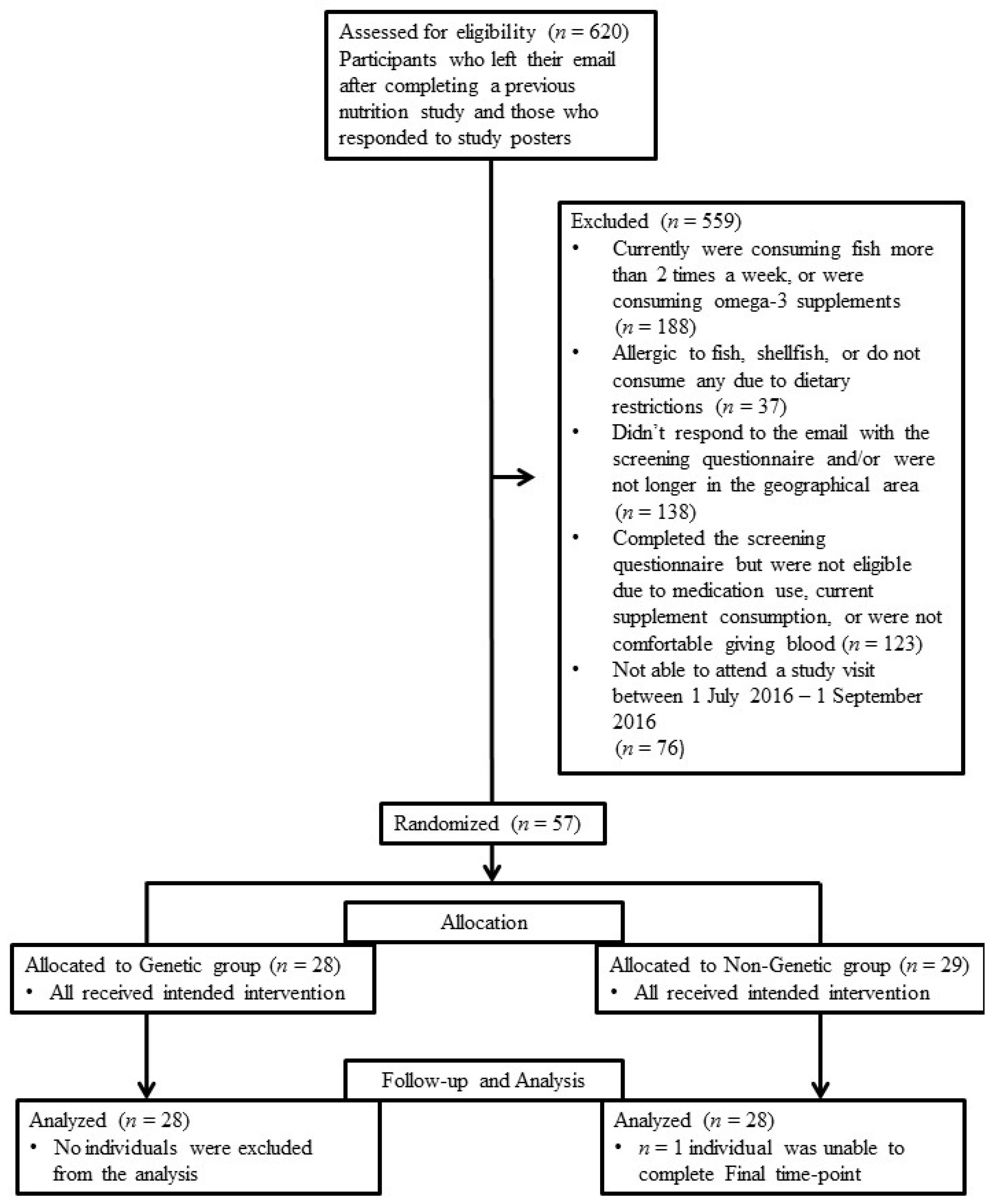
2.1. Participants and Ethics
2.2. Study Design
2.3. Online Surveys
2.3.1. Food Frequency Questionnaires
2.3.2. Diet and Genetics Questionnaire
2.4. Experimental Procedures
2.4.1. Genotyping
2.4.2. Gas Chromatography for FA Analysis
2.4.3. Clinical Measurements
2.5. Statistics
3. Results
3.1. Participant Characteristics
3.2. FFQ Analysis
3.3. FA Analysis
3.4. Clinical Blood Lipid Analysis
3.5. Diet and Genetics Questionnaires
3.5.1. Awareness of Omega-3 FA Terminology
3.5.2. Perceptions and Use of Generic Omega-3 Nutritional Information
3.5.3. Perceived Dietary Changes
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Mutch, D.M.; Wahli, W.; Williamson, G. Nutrigenomics and nutrigenetics: The emerging faces of nutrition. FASEB J. 2005, 19, 1602–1616. [Google Scholar] [CrossRef] [PubMed]
- Ordovas, J.M.; Mooser, V. Nutrigenomics and nutrigenetics. Curr. Opin. Lipidol. 2004, 15, 101–108. [Google Scholar] [CrossRef] [PubMed]
- McBride, C.M.; Koehly, L.M.; Sanderson, S.C.; Kaphingst, K.A. The behavioral response to personalized genetic information: Will genetic risk profiles motivate individuals and families to choose more healthful behaviors? Annu. Rev. Public Health 2010, 31, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Makeeva, O.A.; Markova, V.V.; Puzyrev, V.P. Public interest and expectations concerning commercial genotyping and genetic risk assessment. Pers. Med. 2009, 6, 329–341. [Google Scholar] [CrossRef]
- Stewart-Knox, B.J.; Bunting, B.P.; Gilpin, S.; Parr, H.J.; Pinhao, S.; Strain, J.; de Almeida, M.D.; Gibney, M. Attitudes toward genetic testing and personalised nutrition in a representative sample of European consumers. Br. J. Nutr. 2009, 101, 982–989. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, D.E.; Shih, S.; El-Sohemy, A. Perceptions of genetic testing for personalized nutrition: A randomized trial of dna-based dietary advice. J. Nutrigenet. Nutrigenom. 2014, 7, 94–104. [Google Scholar] [CrossRef] [PubMed]
- Cormier, H.; Tremblay, B.; Paradis, A.M.; Garneau, V.; Desroches, S.; Robitaille, J.; Vohl, M.C. Nutrigenomics—Perspectives from registered dietitians: A report from the quebec-wide e-consultation on nutrigenomics among registered dietitians. J. Hum. Nutr. Diet. 2014, 27, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Horne, J.; Madill, J.; O’Connor, C. Exploring knowledge and attitudes of personal nutrigenomics testing among dietetic students and its value as a component of dietetic education and practice. Can. J. Clin. Nutr. 2016, 4, 50–62. [Google Scholar] [CrossRef]
- Beery, T.A.; Williams, J.K. Risk reduction and health promotion behaviors following genetic testing for adult-onset disorders. Genet. Test. 2007, 11, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, D.E.; El-Sohemy, A. Disclosure of genetic information and change in dietary intake: A randomized controlled trial. PLoS ONE 2014, 9, e112665. [Google Scholar] [CrossRef] [PubMed]
- Hietaranta-Luoma, H.-L.; Tahvonen, R.; Iso-Touru, T.; Puolijoki, H.; Hopia, A. An intervention study of individual, apoe genotype-based dietary and physical-activity advice: Impact on health behavior. J. Nutrigenet. Nutrigenom. 2014, 7, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Arkadianos, I.; Valdes, A.M.; Marinos, E.; Florou, A.; Gill, R.D.; Grimaldi, K.A. Improved weight management using genetic information to personalize a calorie controlled diet. Nutr. J. 2007, 6, 29. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Navas-Carretero, S.; San-Cristobal, R.; Macready, A.L.; Fallaize, R.; Forster, H.; Woolhead, C.; O’Donovan, C.B.; Marsaux, C.F. Effect of an internet-based, personalized nutrition randomized trial on dietary changes associated with the Mediterranean diet: The Food4Me study. Am. J. Clin. Nutr. 2016, 104, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.C.; Kris-Etherton, P.M. Achieving optimal essential fatty acid status in vegetarians: Current knowledge and practical implications. Am. J. Clin. Nutr. 2003, 78, S640–S646. [Google Scholar]
- Bell, G.A.; Kantor, E.D.; Lampe, J.W.; Kristal, A.R.; Heckbert, S.R.; White, E. Intake of long-chain ω-3 fatty acids from diet and supplements in relation to mortality. Am. J. Epidemiol. 2014, 179, 710–720. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-S.; Xun, P.; Iribarren, C.; Van Horn, L.; Steffen, L.; Daviglus, M.L.; Siscovick, D.; Liu, K.; He, K. Intake of fish and long-chain omega-3 polyunsaturated fatty acids and incidence of metabolic syndrome among American young adults: A 25-year follow-up study. Eur. J. Nutr. 2016, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Langlois, K.; Ratnayake, W.M. Omega-3 index of Canadian adults. Health Rep. 2015, 26, 3–11. [Google Scholar] [PubMed]
- Forsyth, S.; Gautier, S.; Salem, N., Jr. Global estimates of dietary intake of docosahexaenoic acid and arachidonic acid in developing and developed countries. Ann. Nutr. Metab. 2016, 68, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Merino, D.M.; Ma, D.W.; Mutch, D.M. Genetic variation in lipid desaturases and its impact on the development of human disease. Lipids Health Dis. 2010, 9, 63. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, M.T.; Nara, T.Y. Structure, function, and dietary regulation of δ6, δ5, and δ9 desaturases. Annu. Rev. Nutr. 2004, 24, 345–376. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Shen, J.; Abecasis, G.R.; Kisialiou, A.; Ordovas, J.M.; Guralnik, J.M.; Singleton, A.; Bandinelli, S.; Cherubini, A.; Arnett, D. Genome-wide association study of plasma polyunsaturated fatty acids in the inchianti study. PLoS Genet. 2009, 5, e1000338. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.E.; Follis, J.L.; Nettleton, J.A.; Foy, M.; Wu, J.H.; Ma, Y.; Tanaka, T.; Manichakul, A.W.; Wu, H.; Chu, A.Y. Dietary fatty acids modulate associations between genetic variants and circulating fatty acids in plasma and erythrocyte membranes: Meta-analysis of nine studies in the charge consortium. Mol. Nutr. Food Res. 2015, 59, 1373–1383. [Google Scholar] [CrossRef] [PubMed]
- Gillingham, L.G.; Harding, S.V.; Rideout, T.C.; Yurkova, N.; Cunnane, S.C.; Eck, P.K.; Jones, P.J. Dietary oils and FADS1-FADS2 genetic variants modulate [13C] α-linolenic acid metabolism and plasma fatty acid composition. Am. J. Clin. Nutr. 2013, 97, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Mumford, S.L.; Dasharathy, S.; Pollack, A.Z.; Schisterman, E.F. Variations in lipid levels according to menstrual cycle phase: Clinical implications. Clin. Lipidol. 2011, 6, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Dietitians of Canada. Eating Guidelines for Omega-3 Fats; Dietitians of Canada: Toronto, ON, Canada, 2013. [Google Scholar]
- Dietitians of Canada. Food Sources of Omega-3 Fats; Dietitians of Canada: Toronto, ON, Canada, 2013. [Google Scholar]
- Bokor, S.; Dumont, J.; Spinneker, A.; Gonzalez-Gross, M.; Nova, E.; Widhalm, K.; Moschonis, G.; Stehle, P.; Amouyel, P.; De Henauw, S. Single nucleotide polymorphisms in the FADS gene cluster are associated with delta-5 and delta-6 desaturase activities estimated by serum fatty acid ratios. J. Lipid Res. 2010, 51, 2325–2333. [Google Scholar] [CrossRef] [PubMed]
- Cormier, H.; Rudkowska, I.; Lemieux, S.; Couture, P.; Julien, P.; Vohl, M.-C. Effects of FADS and ELOVL polymorphisms on indexes of desaturase and elongase activities: Results from a pre-post fish oil supplementation. Genes Nutr. 2014, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Malerba, G.; Schaeffer, L.; Xumerle, L.; Klopp, N.; Trabetti, E.; Biscuola, M.; Cavallari, U.; Galavotti, R.; Martinelli, N.; Guarini, P. Snps of the FADS gene cluster are associated with polyunsaturated fatty acids in a cohort of patients with cardiovascular disease. Lipids 2008, 43, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Patterson, A.C.; Hogg, R.C.; Kishi, D.M.; Stark, K.D. Biomarker and dietary validation of a Canadian food frequency questionnaire to measure eicosapentaenoic and docosahexaenoic acid intakes from whole food, functional food, and nutraceutical sources. J. Acad. Nutr. Diet. 2012, 112, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Canadian Nutrient File. Health Canada: 2015. Available online: http://webprod3.hc-sc.gc.ca/cnf-fce/index-eng.jsp-fce/index-eng.jsp (accessed on 1 February 2016).
- Lafave, L.M.Z.; Lafave, M.R.; Nordstrom, P. Development of a Canadian Behaviour, Attitude and Nutrition Knowledge Survey (BANKS); The Canadian Council on Learning (CCL): Calgary, AB, Canada, 2009. [Google Scholar]
- Nielsen, D.E.; El-Sohemy, A. A randomized trial of genetic information for personalized nutrition. Genes Nutr. 2012, 7, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Roke, K.; Mutch, D.M. The role of FADS1/2 polymorphisms on cardiometabolic markers and fatty acid profiles in young adults consuming fish oil supplements. Nutrients 2014, 6, 2290–2304. [Google Scholar] [CrossRef] [PubMed]
- Roke, K.; Jannas-Vela, S.; Spriet, L.L.; Mutch, D.M. FADS2 genotype influences whole-body resting fat oxidation in young adult men. Appl. Physiol. Nutr. Metab. 2016, 41, 791–794. [Google Scholar] [CrossRef] [PubMed]
- Davidson, E.A.; Pickens, C.A.; Fenton, J. Supplementation with dietary EPA/DHA influences red blood cell fatty acid desaturase estimates and reflects tissue changes in fatty acids in systemic organs. FASEB J. 2016, 30, 267.1. [Google Scholar]
- Martinelli, N.; Girelli, D.; Malerba, G.; Guarini, P.; Illig, T.; Trabetti, E.; Sandri, M.; Friso, S.; Pizzolo, F.; Schaeffer, L. FADS genotypes and desaturase activity estimated by the ratio of arachidonic acid to linoleic acid are associated with inflammation and coronary artery disease. Am. J. Clin. Nutr. 2008, 88, 941–949. [Google Scholar] [PubMed]
- Vessby, B.; Gustafsson, I.-B.; Tengblad, S.; Berglund, L. Indices of fatty acid desaturase activity in healthy human subjects: Effects of different types of dietary fat. Br. J. Nutr. 2013, 110, 871–879. [Google Scholar] [CrossRef] [PubMed]
- Roke, K.; Mutch, David M. Unpublished work. 2017.
- Folch, J.; Lees, M.; Sloane-Stanley, G. A simple method for the isolation and purification of total lipids from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar] [PubMed]
- Merino, D.M.; Johnston, H.; Clarke, S.; Roke, K.; Nielsen, D.; Badawi, A.; El-Sohemy, A.; Ma, D.W.; Mutch, D.M. Polymorphisms in FADS1 and FADS2 alter desaturase activity in young caucasian and asian adults. Mol. Genet. Metab. 2011, 103, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Harris, W.S. The omega-3 index as a risk factor for coronary heart disease. Am. J. Clin. Nutr. 2008, 87, S1997–S2002. [Google Scholar]
- Zulyniak, M.A.; Perreault, M.; Gerling, C.; Spriet, L.L.; Mutch, D.M. Fish oil supplementation alters circulating eicosanoid concentrations in young healthy men. Metabolism 2013, 62, 1107–1113. [Google Scholar] [CrossRef] [PubMed]
- Al-Hilal, M.; AlSaleh, A.; Maniou, Z.; Lewis, F.J.; Hall, W.L.; Sanders, T.A.; O’Dell, S.D. Genetic variation at the FADS1-FADS2 gene locus influences delta-5 desaturase activity and lc-pufa proportions after fish oil supplement. J. Lipid Res. 2013, 54, 542–551. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 289–300. [Google Scholar]
- Benjamini, Y.; Drai, D.; Elmer, G.; Kafkafi, N.; Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 2001, 125, 279–284. [Google Scholar] [CrossRef]
- Stark, K.D.; Van Elswyk, M.E.; Higgins, M.R.; Weatherford, C.A.; Salem, N. Global survey of the omega-3 fatty acids, docosahexaenoic acid and eicosapentaenoic acid in the blood stream of healthy adults. Prog. Lipid Res. 2016, 63, 132–152. [Google Scholar] [CrossRef] [PubMed]
- Global Organization for EPA and DHA Omega-3. Global Recommendations for EPA and DHA Intake; Global Organization for EPA and DHA Omega-3: Salt Lake City, UT, USA, 2014. [Google Scholar]
| Genetic (n = 28) | Non-Genetic (n = 29) | χ2 p-Value | |
|---|---|---|---|
| FADS1 (rs174537) Genoptype Frequency | |||
| Major (GG) | 13/28 (46.4%) | 13/29 (44.8%) | 0.79 |
| Minor (GT + TT) | 15/28 (53.6%) | 16/29 (55.2%) | |
| Field of study/area of work | |||
| Life Science | 19/28 (67.9%) | 21/29 (72.4%) | 0.46 |
| Social Science | 7/28 (25.0%) | 4/29 (13.8%) | |
| Other | 2/28 (7.1%) | 4/29 (13.8%) | |
| Current Position | |||
| Undergraduate student | 20/28 (71.4%) | 19/29 (65.5%) | 0.36 |
| Graduate student | 7/28 (25.0%) | 8/29 (27.6%) | |
| Working full-time | 1/28 (3.6%) | 2/29 (6.9%) | |
| Self-Reported Ethnicity | |||
| White/Caucasian | 18/28 (64.3%) | 22/29 (75.9%) | 0.47 |
| Asian | 4/28 (14.3%) | 3/29 (10.3%) | |
| European | 4/28 (14.3%) | 1/29 (3.5%) | |
| Other | 2/28 (7.1%) | 3/29 (10.3%) | |
| Contraceptive use | |||
| Not taking contraceptives | 8/28 (28.6%) | 9/29 (31.0%) | 0.89 |
| Oral contraceptive | 17/28 (60.7%) | 18/29 (62.1%) | |
| IUD contraceptive | 3/28 (10.7%) | 2/29 (6.9%) | |
| Genetic (n = 28) | Non-Genetic (n = 28) | Group | Time | Group × Time Interaction | |||
|---|---|---|---|---|---|---|---|
| Baseline | Final | Baseline | Final | p-Value | p-Value | p-Value | |
| Omega-3 Dietary Intake (FFQ) | |||||||
| EPA and DHA (mg/day) | 211.50 ± 43.16 | 323. 23 ± 65.27 | 190.16 ± 39.21 | 395.82 ± 70.60 | 0.70 | 1.0 × 10−4 | 0.27 |
| RBC FA levels | |||||||
| ALA (%) | 0.43 ± 0.02 | 0.44 ± 0.02 | 0.41 ± 0.02 | 0.42 ± 0.02 | 0.66 | 0.14 | 0.79 |
| EPA (%) | 0.45 ± 0.02 | 0.51 ± 0.03 | 0.55 ± 0.02 | 0.61 ± 0.04 | 3.9 × 10−3 * | 0.02 | 0.89 |
| DHA (%) | 3.40 ± 0.11 | 3.42 ± 0.09 | 3.42 ± 0.11 | 3.54 ± 0.11 | 0.64 | 0.20 | 0.36 |
| Omega-3 Index | 3.86 ± 0.11 | 3.97 ± 0.11 | 3.97 ± 0.12 | 4.15 ± 0.13 | 0.34 | 0.04 | 0.66 |
| Clinical Data | |||||||
| BMI (kg/m2) | 23.01 ± 0.60 | 22.87 ± 0.61 | 23.41 ± 0.50 | 23.55 ± 0.53 | 0.50 | 0.99 | 0.11 |
| TAG (mmol/L) | 0.90 ± 0.07 | 1.02 ± 0.08 | 1.02 ± 0.07 | 1.02 ± 0.06 | 0.24 | 0.06 | 0.06 |
| Cholesterol (mmol/L) | 4.40 ± 0.19 | 4.67 ± 0.20 | 4.43 ± 0.14 | 4.72 ± 0.14 | 0.87 | 2.0 × 10−4 * | 0.85 |
| HDL (mmol/L) | 1.76 ± 0.06 | 1.75 ± 0.07 | 1.75 ± 0.09 | 1.78 ± 0.09 | 0.94 | 0.66 | 0.48 |
| Chol/HDL ratio | 2.58 ± 0.13 | 2.70 ± 0.11 | 2.67 ± 0.12 | 2.77 ± 0.12 | 0.60 | 0.04 | 0.81 |
| LDL (mmol/L) | 2.18 ± 0.15 | 2.31 ± 0.14 | 2.22 ± 0.13 | 2.47 ± 0.12 | 0.58 | 2.6 × 10−3 * | 0.32 |
| Non-HDL Chol | 2.65 ± 0.18 | 2.88 ± 0.18 | 2.68 ± 0.13 | 2.94 ± 0.12 | 0.83 | 2.0 × 10−4 * | 0.92 |
| Questionnaire Data (Omega-3 terminology) | |||||||
| Alpha-linolenic acid | 16/28 (57.1%) | 23/28 (82.1%) | 18/29 (62.1%) | 20/28 (71.%) | 0.73 | 2.0 × 10−3 * | 0.33 |
| Eicosapentaenoic acid | 12/28 (42.9%) | 22/28 (78.6%) | 12/29 (41.4%) | 13/28 (46.4%) | 0.08 | 1.0 × 10−3 * | 0.07 |
| Docosahexaenoic acid | 14/28 (50.0%) | 22/28 (78.6%) | 12/29 (41.4%) | 14/28 (50.0%) | 0.04 | 5.0 × 10−3 * | 0.21 |
| ALA | 12/28 (42.9%) | 23/28 (82.1%) | 19/29 (65.5%) | 20/28 (71.4%) | 0.71 | 0.01 * | 0.04 |
| EPA | 15/28 (53.6%) | 23/28 (82.1%) | 20/29 (69.0%) | 20/28 (71.4%) | 0.86 | 0.01 * | 0.05 |
| DHA | 19/28 (67.9%) | 25/28 (89.3%) | 19/29 (65.5%) | 22/28 (78.6%) | 0.46 | 0.01 * | 0.49 |
| Genetic (n = 28) | Non-Genetic (n = 28) | p-Value | |
|---|---|---|---|
| Average | Average | ||
| I understood the nutrition information about omega-3 fats provided at the start of the study | 6.21 ± 0.26 | 6.21 ± 0.23 | 1.0 |
| The recommendations about omega-3 fats that were provided in the document at the start of the study were new to me | 4.64 ± 1.75 | 3.71 ± 0.31 | 0.05 |
| I enjoyed learning about the dietary recommendations related to omega-3 fats | 6.14 ± 1.18 | 5.89 ± 1.07 | 0.41 |
| The dietary recommendations were useful when I considered my diet throughout the study | 5.68 ± 1.21 | 4.93 ± 1.25 | 0.03 |
| When I am in the grocery store or supplement store, I can confidently determine foods that have been fortified, or have added EPA and DHA omega-3 fats | 5.43 ± 1.48 | 5.32 ± 1.19 | 0.77 |
| I would like to know more about the dietary recommendations related to omega-3 fats | 5.64 ± 1.47 | 5.39 ± 1.47 | 0.53 |
| I am interested in the relationship between diet and genetics | 6.12 ± 1.45 | 6.61 ± 0.79 | 0.11 |
| Genetic (n = 28) | Non-Genetic (n = 28) | χ2 p-Value | |
|---|---|---|---|
| Omega-3 foods are expensive | 9/28 (32.1%) | 17/28 (60.7%) | 0.17 |
| When I get busy I don’t make time to eat healthy foods | 8/28 (28.6%) | 4/28 (14.3%) | |
| I didn’t experience any barriers to change throughout this study | 3/28 (10.7%) | 1/28 (3.6%) | |
| Other obstacles/barriers # | 8/28 (28.6%) | 6/28 (21.4%) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Roke, K.; Walton, K.; Klingel, S.L.; Harnett, A.; Subedi, S.; Haines, J.; Mutch, D.M. Evaluating Changes in Omega-3 Fatty Acid Intake after Receiving Personal FADS1 Genetic Information: A Randomized Nutrigenetic Intervention. Nutrients 2017, 9, 240. https://doi.org/10.3390/nu9030240
Roke K, Walton K, Klingel SL, Harnett A, Subedi S, Haines J, Mutch DM. Evaluating Changes in Omega-3 Fatty Acid Intake after Receiving Personal FADS1 Genetic Information: A Randomized Nutrigenetic Intervention. Nutrients. 2017; 9(3):240. https://doi.org/10.3390/nu9030240
Chicago/Turabian StyleRoke, Kaitlin, Kathryn Walton, Shannon L. Klingel, Amber Harnett, Sanjeena Subedi, Jess Haines, and David M. Mutch. 2017. "Evaluating Changes in Omega-3 Fatty Acid Intake after Receiving Personal FADS1 Genetic Information: A Randomized Nutrigenetic Intervention" Nutrients 9, no. 3: 240. https://doi.org/10.3390/nu9030240
APA StyleRoke, K., Walton, K., Klingel, S. L., Harnett, A., Subedi, S., Haines, J., & Mutch, D. M. (2017). Evaluating Changes in Omega-3 Fatty Acid Intake after Receiving Personal FADS1 Genetic Information: A Randomized Nutrigenetic Intervention. Nutrients, 9(3), 240. https://doi.org/10.3390/nu9030240





