Genetic Variations as Modifying Factors to Dietary Zinc Requirements—A Systematic Review
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Methods
- Zinc.ti,ab.
- Zn.ti,ab.
- Zinc/
- Zinc compounds/
- Finger*.ti,ab.
- 1 OR 2 OR 3 OR 4
- 6 NOT 5
- (Polymorph* OR Allel* OR Genet* OR Geno* OR Gene OR Genes).ti,ab.
- Exp Polymorphism, Genetic/
- 8 OR 9
- Exp Cation Transport Proteins/
- (Zinc adj2 deficienc*).ti,ab.
- (Zn adj2 deficienc*).ti,ab.
- (Zinc adj2 transporter*).ti,ab.
- (Zn adj2 transporter*).ti,ab.
- 11 OR 12 OR 13 OR 14 OR 15
- 7 AND 10 AND 16
2.2. Inclusion Criteria
2.3. Exclusion Criteria
2.4. Data Collection and Analysis
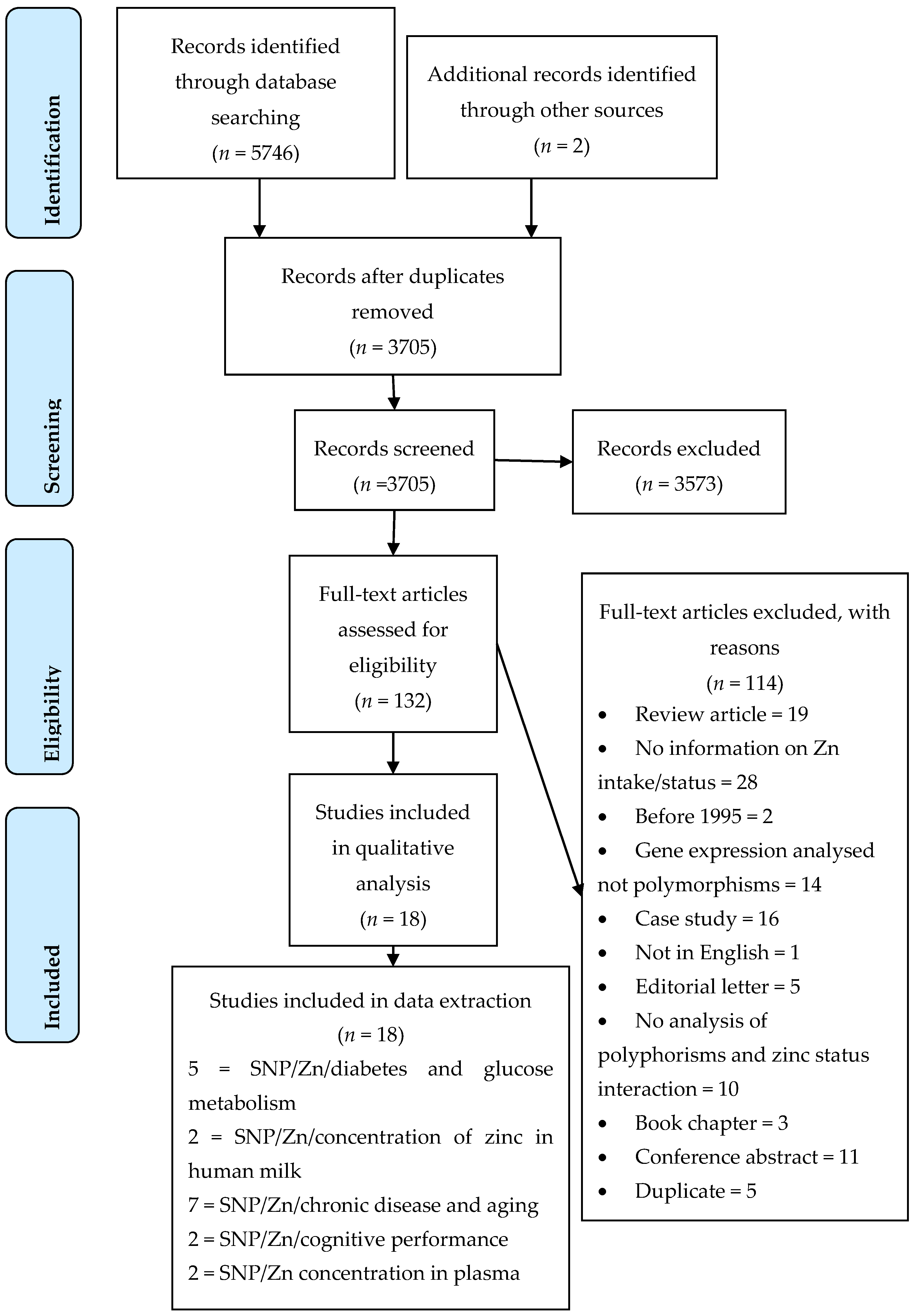
2.4.1. Selection of Studies
2.4.2. Data Extraction
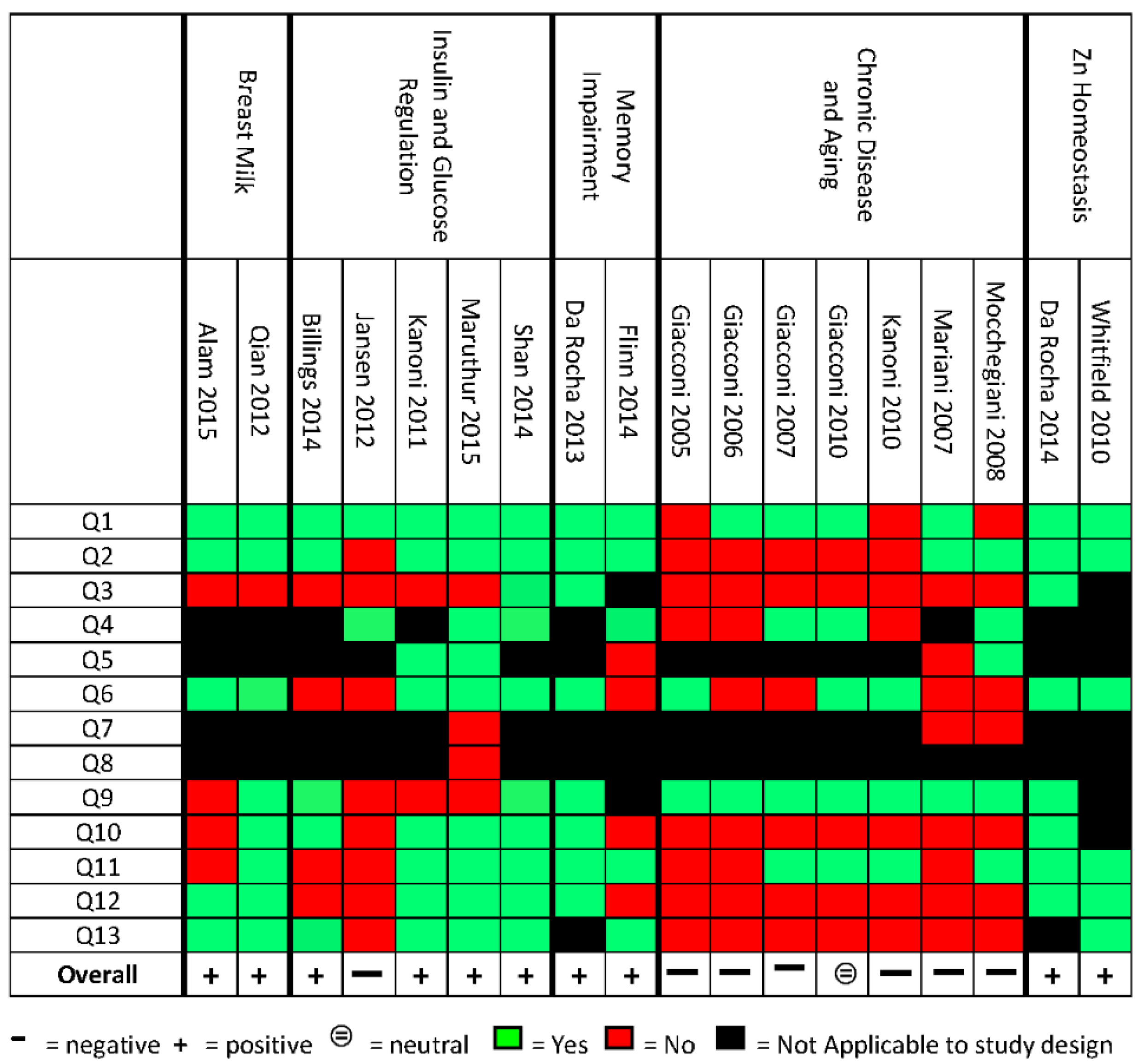
2.5. Quality Assessment
3. Results
3.1. Description of Included Studies
3.2. Does Dietary Zn Modulate the Effect of SLC30A2 Polymorphisms on Human Milk Zinc Content?
3.3. Does Dietary Zn Modulate the Association between Gene Variants and Glucose Metabolism Traits in Relation to Type-2 Diabetes?
3.4. Does Dietary Zn Modulate the Effect of Gene Variants on Cognitive Performance?
3.5. Does Dietary Zn Modulate the Association between Gene Variants and the Development of Chronic Diseases in the Aging Population?
3.6. Do Polymorphisms in Genes Involved in Zn Transport and Metabolism Affect Plasma Zn Concentration?
4. Discussion
5. Perspectives
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ferguson, L.R.; De Caterina, R.; Görman, U.; Allayee, H.; Kohlmeier, M.; Prasad, C.; Kang, J.X. Guide and Position of the International Society of Nutrigenetics/Nutrigenomics on Personalised Nutrition: Part 1—Fields of Precision Nutrition. J. Nutrigenet. Nutrigenom. 2016, 9, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Lonnerdal, B. Dietary factors affecting trace element absorption in infants. Acta Paediatr. Scand. Suppl. 1989, 351, 109–113. [Google Scholar] [CrossRef] [PubMed]
- National Health and Medical Research Council. Nutrient Reference Values. 2006. Available online: http://www.nrv.gov.au (accessed on 24 May 2016). [Google Scholar]
- Maret, W.; Sandstead, H.H. Zinc requirements and the risks and benefits of zinc supplementation. J. Trace Elem. Med. Biol. 2006, 20, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Prasad, A.S.; Miale, A., Jr.; Farid, Z.; Sandstead, H.H.; Schulert, A.R. Zinc metabolism in patients with the syndrome of iron deficiency anemia, hepatosplenomegaly, dwarfism, and hypognadism. J. Lab. Clin. Med. 1963, 61, 537–549. [Google Scholar] [PubMed]
- Devirgiliis, C.; Zalewski, P.D.; Perozzi, G.; Murgia, C. Zinc fluxes and zinc transporter genes in chronic diseases. Mutat. Res. 2007, 622, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Hambidge, K.M.; Krebs, N.F. Zinc deficiency: A special challenge. J. Nutr. 2007, 137, 1101–1105. [Google Scholar] [PubMed]
- Ackland, M.L.; Michalczyk, A. Zinc deficiency and its inherited disorders—A review. Genes Nutr. 2006, 1, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Kambe, T. The Functions of Metallothionein and ZIP and ZnT Transporters: An Overview and Perspective. Int. J. Mol. Sci. 2016, 17, 336. [Google Scholar] [CrossRef] [PubMed]
- Kambe, T.; Tsuji, T.; Hashimoto, A.; Itsumura, N. The Physiological, Biochemical, and Molecular Roles of Zinc Transporters in Zinc Homeostasis and Metabolism. Physiol. Rev. 2015, 95, 749–784. [Google Scholar] [CrossRef] [PubMed]
- Huang, L. Zinc and its transporters, pancreatic beta-cells, and insulin metabolism. Vitam. Horm. 2014, 95, 365–390. [Google Scholar] [PubMed]
- Rutter, G.A.; Chimienti, F. SLC30A8 mutations in type 2 diabetes. Diabetologia 2015, 58, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Fan, M.; Li, W.; Wang, L.; Gu, S.; Dong, S.; Chen, M.; Jiang, X. Association of SLC30A8 gene polymorphism with type 2 diabetes, evidence from 46 studies: A meta-analysis. Endocrine 2016, 53, 381–394. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Hennigar, S.R.; Alam, S.; Nishida, K.; Kelleher, S.L. Essential Role for Zinc Transporter 2 (ZnT2)-mediated Zinc Transport in Mammary Gland Development and Function during Lactation. J. Biol. Chem. 2015, 290, 13064–13078. [Google Scholar] [CrossRef] [PubMed]
- Murgia, C.; Vespignani, I.; Rami, R.; Perozzi, G. The Znt4 mutation inlethal milk mice affects intestinal zinc homeostasis through the expression of other Zn transporters. Genes Nutr. 2006, 1, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Adlard, P.A.; Parncutt, J.; Lal, V.; James, S.; Hare, D.; Doble, P.; Bush, A.I. Metal chaperones prevent zinc-mediated cognitive decline. Neurobiol. Dis. 2015, 81, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Adlard, P.A.; Parncutt, J.M.; Finkelstein, D.I.; Bush, A.I. Cognitive loss in zinc transporter-3 knock-out mice: A phenocopy for the synaptic and memory deficits of Alzheimer’s disease? J. Neurosci. 2010, 30, 1631–1636. [Google Scholar] [CrossRef] [PubMed]
- Greenough, M.A.; Camakaris, J.; Bush, A.I. Metal dyshomeostasis and oxidative stress in Alzheimer’s disease. Neurochem. Int. 2013, 62, 540–555. [Google Scholar] [CrossRef] [PubMed]
- Hesketh, J. Personalised nutrition: How far has nutrigenomics progressed? Eur. J. Clin. Nutr. 2013, 67, 430–435. [Google Scholar] [CrossRef] [PubMed]
- Palmiter, R.D.; Findley, S.D. Cloning and functional characterization of a mammalian zinc transporter that confers resistance to zinc. EMBO J. 1995, 14, 639–649. [Google Scholar] [PubMed]
- Billings, L.K.; Jablonski, K.A.; Ackerman, R.J.; Taylor, A.; Fanelli, R.R.; McAteer, J.B.; Franks, P.W. The Influence of Rare Genetic Variation in SLC30A8 on Diabetes Incidence and beta-Cell Function. J. Clin. Endocrinol. Metab. 2014, 99, E926–E930. [Google Scholar] [CrossRef] [PubMed]
- Kanoni, S.; Nettleton, J.A.; Hivert, M.F.; Ye, Z.; Van Rooij, F.J.; Shungin, D.; Gustafsson, S. Total zinc intake may modify the glucose-raising effect of a zinc transporter (SLC30A8) variant: A 14-cohort meta-analysis. Diabetes 2011, 60, 2407–2416. [Google Scholar] [CrossRef] [PubMed]
- Giacconi, R.; Cipriano, C.; Muti, E.; Costarelli, L.; Malavolta, M.; Caruso, C.; Mocchegiani, E. Involvement of-308 TNF-α and 1267 Hsp70–2 polymorphisms and zinc status in the susceptibility of coronary artery disease (CAD) in old patients. Biogerontology 2006, 7, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Mariani, E.; Neri, S.; Cattini, L.; Mocchegiani, E.; Malavolta, M.; Dedoussis, G.V.; Facchini, A. Effect of zinc supplementation on plasma IL-6 and MCP-1 production and NK cell function in healthy elderly: Interactive influence of +647 MT1a and -174 IL-6 polymorphic alleles. Exp. Gerontol. 2008, 43, 462–471. [Google Scholar] [CrossRef] [PubMed]
- Association, A.D. Evidence Analysis Manual: Steps in the ADA Evidence Analysis Process; American Dietetic Association: Chicago, IL, USA, 2008. [Google Scholar]
- Higgins, J.P.; Altman, D.G.; Gøtzsche, P.C.; Jüni, P.; Moher, D.; Oxman, A.D.; Sterne, J.A. The Cochrane Collaboration’s tool for assessing risk of bias in randomised trials. Br. Med. J. 2011, 343, d5928. [Google Scholar] [CrossRef] [PubMed]
- STROBE Statement. Checklist of items that should be included in reports of observational studies (STROBE initiative). Int. J. Public Health 2008, 53, 3–4. [Google Scholar]
- Salanti, G.; Amountza, G.; Ntzani, E.E.; Ioannidis, J.P. Hardy-Weinberg equilibrium in genetic association studies: An empirical evaluation of reporting, deviations, and power. Eur. J. Hum. Genet. 2005, 13, 840–848. [Google Scholar] [CrossRef] [PubMed]
- A Logo for Human Rights. ONE WORLD EQUAL RIGHTS. Available online: http://www.humanrightslogo.net/en/submission/one-world-%E2%80%93-equal-rights (accessed on 15 February 2017).
- Alam, S.; Hennigar, S.R.; Gallagher, C.; Soybel, D.I.; Kelleher, S.L. Exome Sequencing of SLC30A2 Identifies Novel Loss- and Gain-of-Function Variants Associated with Breast Cell Dysfunction. J. Mammary Gland Biol. Neoplasia 2015, 20, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.; Wang, B.; Tang, N.; Zhang, W.; Cai, W. Polymorphisms of SLC30A2 and selected perinatal factors associated with low milk zinc in Chinese breastfeeding women. Early Hum. Dev. 2012, 88, 663–668. [Google Scholar] [CrossRef] [PubMed]
- Da Rocha, T.J.; Blehm, C.J.; Bamberg, D.P.; Fonseca, T.L.R.; Tisser, L.A.; de Oliveira Junior, A.A.; Fiegenbaum, M. The effects of interactions between selenium and zinc serum concentration and SEP15 and SLC30A3 gene polymorphisms on memory scores in a population of mature and elderly adults. Genes Nutr. 2014, 9, 377. [Google Scholar] [CrossRef] [PubMed]
- Da Rocha, T.J.; Korb, C.; Schuch, J.B.; Bamberg, D.P.; de Andrade, F.M.; Fiegenbaum, M. SLC30A3 and SEP15 gene polymorphisms influence the serum concentrations of zinc and selenium in mature adults. Nutr. Res. 2014, 34, 742–748. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, J.B.; Dy, V.; McQuilty, R.; Zhu, G.; Heath, A.C.; Montgomery, G.W.; Martin, N.G. Genetic Effects on Toxic and Essential Elements in Humans: Arsenic, Cadmium, Copper, Lead, Mercury, Selenium, and Zinc in Erythrocytes. Environ. Health Perspect. 2010, 118, 776–782. [Google Scholar] [CrossRef] [PubMed]
- Kanoni, S.; Dedoussis, G.V.; Herbein, G.; Fulop, T.; Varin, A.; Jajte, J.; Giacconi, R. Assessment of gene-nutrient interactions on inflammatory status of the elderly with the use of a zinc diet score—ZINCAGE study. J. Nutr. Biochem. 2010, 21, 526–531. [Google Scholar] [CrossRef] [PubMed]
- Jansen, J.; Rosenkranz, E.; Overbeck, S.; Warmuth, S.; Mocchegiani, E.; Giacconi, R.; Rink, L. Disturbed zinc homeostasis in diabetic patients by in vitro and in vivo analysis of insulinomimetic activity of zinc. J. Nutr. Biochem. 2012, 23, 1458–1466. [Google Scholar] [CrossRef] [PubMed]
- Flinn, J.M.; Bozzelli, P.L.; Adlard, P.A.; Railey, A.M. Spatial memory deficits in a mouse model of late-onset Alzheimer’s disease are caused by zinc supplementation and correlate with amyloid-beta levels. Front. Aging Neurosci. 2014, 6, 174. [Google Scholar] [CrossRef] [PubMed]
- Giacconi, R.; Cipriano, C.; Muti, E.; Costarelli, L.; Maurizio, C.; Saba, V.; Mocchegiani, E. Novel -209A/G MT2A polymorphism in old patients with type 2 diabetes and atherosclerosis: Relationship with inflammation (IL-6) and zinc. Biogerontology 2005, 6, 407–413. [Google Scholar] [CrossRef] [PubMed]
- Giacconi, R.; Kanoni, S.; Mecocci, P.; Malavolta, M.; Richter, D.; Pierpaoli, S.; Piacenza, F. Association of MT1A haplotype with cardiovascular disease and antioxidant enzyme defense in elderly Greek population: Comparison with an Italian cohort. J. Nutr. Biochem. 2010, 21, 1008–1014. [Google Scholar] [CrossRef] [PubMed]
- Giacconi, R.; Muti, E.; Malavolta, M.; Cipriano, C.; Costarelli, L.; Bernardini, G.; Mocchegiani, E. The +838 C/G MT2A polymorphism, metals, and the inflammatory/immune response in carotid artery stenosis in elderly people. Mol. Med. 2007, 13, 388–395. [Google Scholar] [CrossRef]
- Shan, Z.L.; Bao, W.; Zhang, Y.; Rong, Y.; Wang, X.; Jin, Y.; Liu, L. Interactions Between Zinc Transporter-8 Gene ( SLC30A8) and Plasma Zinc Concentrations for Impaired Glucose Regulation and Type 2 Diabetes. Diabetes 2014, 63, 1796–1803. [Google Scholar] [CrossRef] [PubMed]
- Mocchegiani, E.; Giacconi, R.; Costarelli, L.; Muti, E.; Cipriano, C.; Tesei, S.; Gasparini, N. Zinc deficiency and IL-6−174G/C polymorphism in old people from different European countries: Effect of zinc supplementation. ZINCAGE study. Exp. Gerontol. 2008, 43, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Maruthur, N.M.; Clark, J.M.; Fu, M.; Kao, W.L.; Shuldiner, A.R. Effect of zinc supplementation on insulin secretion: Interaction between zinc and SLC30A8 genotype in Old Order Amish. Diabetologia 2015, 58, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Prisma Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Verghese, P.B.; Castellano, J.M.; Holtzman, D.M. Apolipoprotein E in Alzheimer’s disease and other neurological disorders. Lancet Neurol. 2011, 10, 241–252. [Google Scholar] [CrossRef]
- Mocchegiani, E.; Zincage, C. Zinc, metallothioneins, longevity: Effect of zinc supplementation on antioxidant response: A zincage study. Rejuvenation Res. 2008, 11, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Sladek, R.; Rocheleau, G.; Rung, J.; Dina, C.; Shen, L.; Serre, D.; Balkau, B. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007, 445, 881–885. [Google Scholar] [CrossRef] [PubMed]
- Diabetes Genetics Initiative of Broad Institute of Harvard and MIT, Lund University, and Novartis Institutes of BioMedical Research; Saxena, R.; Voight, B.F.; Lyssenko, V.; Burtt, N.P.; de Bakker, P.I.; Chen, H.; Roix, J.J.; Kathiresan, S.; Hirschhorn, J.N.; et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 2007, 316, 1331–1336. [Google Scholar] [PubMed]
- Zeggini, E.; Weedon, M.N.; Lindgren, C.M.; Frayling, T.M.; Elliott, K.S.; Lango, H.; Barrett, J.C. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 2007, 316, 1336–1341. [Google Scholar] [CrossRef] [PubMed]
- Scott, L.J.; Weedon, M.N.; Lindgren, C.M.; Frayling, T.M.; Elliott, K.S.; Lango, H.; Barrett, J.C. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 2007, 316, 1341–1345. [Google Scholar] [CrossRef] [PubMed]
- Mocchegiani, E.; Giacconi, R.; Malavolta, M. Zinc signalling and subcellular distribution: Emerging targets in type 2 diabetes. Trends Mol. Med. 2008, 14, 419–428. [Google Scholar] [CrossRef] [PubMed]
- Palmiter, R.D.; Cole, T.B.; Quaife, C.J.; Findley, S.D. ZnT-3, a putative transporter of zinc into synaptic vesicles. Proc. Natl. Acad. Sci. USA 1996, 93, 14934–14939. [Google Scholar] [CrossRef] [PubMed]
- Cole, T.B.; Quaife, C.J.; Findley, S.D. Elimination of zinc from synaptic vesicles in the intact mouse brain by disruption of the ZnT3 gene. Proc. Natl. Acad. Sci. USA 1999, 96, 1716–1721. [Google Scholar] [CrossRef] [PubMed]
- Tai, L.M.; Ghura, S.; Koster, K.P.; Liakaite, V.; Maienschein-Cline, M.; Kanabar, P.; Green, S.J. APOE-modulated Abeta-induced neuroinflammation in Alzheimer’s disease: Current landscape, novel data, and future perspective. J. Neurochem. 2015, 133, 465–488. [Google Scholar] [CrossRef] [PubMed]
- Lowe, N.M.; Fekete, K.; Decsi, T. Methods of assessment of zinc status in humans: A systematic review. Am. J. Clin. Nutr. 2009, 89, 2040S–2051S. [Google Scholar] [CrossRef] [PubMed]
| Study Reference | Study Design | Sample Description | SNPs Associated with a Phenotype (Gene, rs# and Nucleotide Change) | Description of the SNPs Association with Phenotypic Trait | SNPs Association with Plasma Zn Concentration ^ | SNPs Association with Zn Intake or Zn Supplementation | Outcomes |
|---|---|---|---|---|---|---|---|
| Polymorphisms in Zn transporters for Breast milk | |||||||
| Alam et al. 2015 [30] | Cross-sectional | 54 F, healthy American Inclusion: 18–40 years who were breastfeeding one healthy infant at ≈4 months post-partum. Exclusion: Pre-term births (<37 week gestation), multiple births, smokers | 12 novel SNPs identified as missense by sequencing SLC30A2 gene: A28D, K66N, Q71H, D103E, A105P Q117H, T288S, A310T, L311V, T312K, V313G, Q315R. | D103E found associated with low [Zn] breastmilk. | N/A | N/A | Variability in the concentration of Zn in human milk was associated with SLC30A2 SNPs. Possibly because of the small sample size, the other association did not reach significance. |
| Qian et al. 2012 [31] | Cross-sectional | 750 F, healthy Chinese Inclusion: 18–36 years breastfeeding day 42 postpartum. Exclusion: multiple gestations, multiparity, pre-existing maternal diseases, foetal malformation and lactation failure, complicated pregnancy | 1 SNP in the promoter region and 1 in the coding region of the coding sequence of SLC30A2 gene (rs117153535 (G/T); No rs-SLC30A2/1031A > G) | Association between genetic polymorphisms and milk Zn concentration: rs117153535 T allele associated with lower [Zn]; SLC30A2/1031A > G allele associated with lower [Zn]. | N/A | N/A | Variability in the concentration of Zn in human milk was associated with SLC30A2 SNPs. |
| Polymorphisms in genes relating to Insulin and Glucose Regulation | |||||||
| Billings et al. 2014 [21] | Cross-sectional from RCT | 2997 male and female participants in the Diabetes Prevention Program (US) Inclusion: high risk of developing T2D (overweight with elevated fasting glucose and IGT) randomised to placebo, metformin 850 mg twice daily, or lifestyle intervention and consented to genetic testing | 44 novel SNPs including: SLC30A8: rs2464591 (C/T), rs2466296 (C/T), rs2466297 (A/G), rs2466299 (C/T) | Associated with improvement in β-cell function. | No association between Zn intake and SNPs on diabetes incidence | N/A | SLC30A8 variants influence T2D risk, insulin secretion traits and β-cell function. Zn intake did not modify the genetic risk. None of the SNPs had a large effect. |
| SLC30A8: rs2466293 (C/T) | Decrease in β-cell function. | ||||||
| Shan et al. 2014 [41] | Cross-sectional Case-control study | 1796 male and female Chinese Han ethnicity Inclusion: newly diagnosed IGR and T2D, ≥30 years, BMI ≥ 40 kg/m2, no history of diabetes diagnosis or pharmacological treatment for hyperlipidaemia. Exclusion: clinically significant neurological, endocrine, or other systemic diseases, as well as acute illness, chronic inflammatory, or infectious diseases. | SLC308A, rs13266634 (C/T) | C allele was associated with increased odds of T2D. | Decreased risk of T2D in carriers of risk (C) allele with high plasma Zn concentration. | N/A | SLC30A8 rs13266634 is associated with T2D. CC genotype and low plasma Zn increase T2D risk and Impaired Glucose Regulation. High plasma Zn concentration (third highest tertile ≥197.58 µg/dL) decreased risk of T2D in carriers of risk C allele. |
| Kanoni et al. 2011 [22] | Cross-sectional Meta-analysis | Meta-analysis from 14 cohort studies, 46,021 participants. Inclusion: no diabetes (fasting glucose ≥ 7 mmol/L or use of antidiabetic medications). Sample size for interaction analysis between dietary Zn intake and SNPs ranged from 27,010 to 45,821. | 20 SNPs analysed including: SLC30A8, rs11558471 (A/G) | N/A | Zn intake of 14 mg/day associated with 0.024 mmol/L decrease in fasting glucose concentration in A carriers in comparison to GG (0.048 mmol/L reduction for AA homozygotes) | Interaction only significant with zinc diet and supplementation | Increased Zn intake improves diabetes risk dependent for A allele carriers of SNP rs11558471.S |
| Jansen et al. 2012 [36] | Case-control study | 150 male and female (22 T1D; 53 T2D; 7 matched controls). Inclusion: ≥18 years, patients with type 1 or 2 diabetes; Exclusion: acute infection, recent, cancer, liver or renal disease. | SLC30A8 rs13266634 (C/T) MT1A rs11640851 (A/C), rs8052394 (A/G) | No significance in SNP association T1D or T2D | N/A | N/A | Serum Zn decreased in patients with diabetes, no association with any SNPs and disease biomarkers. Small sample size. |
| Maruthur et al. 2015 [43] | Non-randomised supplement clinical trial | 55 male and female Old Order Amish from Lancaster, PA, USA. Inclusion: 21–70 years no diagnosis of diabetes and random glucose less than 11.10 mmol/L [200 mg/dL]. | SLC30A8, rs13266634 (C/T) (R325W). The (C) allele encodes the arginine (R), and the (T) allele encodes the tryptophan (W) | Carriers of allele T showed increased insulin response after supplementation | Serum Zn concentration increased by 23% for CC genotypes and 33% for CT/TT genotypes after supplementation | Oral Zn acetate 50 mg, twice daily × 14 days | Carriers of risk T allele showed increased insulin response after supplementation with Zn; these participants may benefit most from Zn supplementation |
| Polymorphisms in genes relating to Memory Impairment | |||||||
| Da Rocha et al. 2014a [32] | Cross-sectional study | 240 male and female mature, elderly adults Inclusion: ≥50 years, absence of dementia, owning intellect enough to continue production, present complaint of progressive memory loss, show objective evidence of memory deficits. Exclusion: symptoms of depression, anxiety, or stress, IQ < 70, use of vitamin supplements containing micronutrients of interest | SLC30A3, rs73924411 (C/T) rs11126936 (G/T) | Increased frequency of rs11126936 TT carriers in people with memory deficits. | When serum Zn was below recommended concentration: rs11126936 T carriers had better memory scores, whereas CC carriers’ performance decreased. | N/A | Recommended Zn concentration may be neurotoxic for T carriers suffering from memory deficit. CC genotype may benefit from increased Zn intake when Zn serum is low |
| Flinn et al. 2014 [37] | Case-control on mouse-model | 22 Mouse strain CRND8 with human ApoE ε4; 23 Mouse strain CRND8; 24 WT controls | ApoE ε4 (Apoliprotein E isoform SNP in NCBI rs7412) | Carriers of human ApoE ε4 showed impaired spatial memory | N/A | 10 ppm of ZnCO3 in water | Increased dietary Zn significantly impaired spatial memory of mice carrying ApoE ε4 human compared with WT and CRND8. The amount of Zn consumed daily was not reported |
| Polymorphisms in genes relating to chronic disease in the aging population | |||||||
| Giacconi et al. 2007 [40] | Case-control study | 506 male and female elderly adults (288 with coronary artery stenosis; 218 healthy, age and sex matched controls living at home) Inclusion: older individuals born and living in Central Italy admitted to INRCA Geriatric Hospital, Ancona, Italy, for endarterectomy | MT2A rs10636 (C/G) | C allele more frequent amongst patients with carotid stenosis | Low plasma [Zn] associated with C allele | N/A | MT2A rs10636 C allele is associated with low plasma Zn and independently with carotid stenosis. No conclusive evidence that modifying Zn intake is beneficial for C allele carriers |
| Kanoni et al. 2010 [35] | Cross-sectional study | 819 male and female elderly adults (272 from Italy, 163 from Greece, 137 from Germany, 128 from France, and 119 from Poland); Inclusion: (ZINCAGE Project: www.zincage.org). ≥ 60 years old, non-institutionalised, free of medication and supplements. Exclusion: autoimmune, neurodegenerative, cardiovascular, kidney, or liver disease, diabetes, infections, cancer, sickle cell, skin ulcerations, and endocrine disorders. | −174 IL-6 G/C (inNational Center for Biotechnology Information (NCBI)rs1800795 C/G) | GG genotype had greater increase in IL-6 levels with increased ‘Zn diet score’ than CG and CC genotypes | Significant interaction of ‘Zn diet score’ and GG genotype of rs1800795 | N/A | Method developed to calculate Zn dietary intake and correlate it with plasma Zn “Zn score”. Association observed between Zn score, G allele and IL-6 levels |
| Giacconi et al. 2005 [38] | Case-control study | 279 male and female born and living in Central Italy, (91 T2D; 188 age and sex-matched controls living at home). Inclusion: (ZINCAGE Project: www.zincage.org) older adults, diagnosis of T2D with carotid stenosis, control individuals living at home, no hypertension, diabetes or carotid stenosis, no history of Coronary Heart Disease (CHD,) normal electrocardiography and no sign of myocardial ischemia | MT2A rs1610216 A/G (in NCBI database C/T) | AA genotype associated with carotid stenosis and T2D, inflammation markers. A allele more frequent in patients than controls | Plasma Zn concentration decreased in AA genotype compared with AG genotypes | N/A | MT2A rs1610216, AA genotype associated with disease biomarkers and low serum Zn concentration. No evidence supporting that increased serum Zn would be beneficial |
| Giacconi et al. 2010 [39] | Case-control study | 459 male and female elderly Italian adults (215 Cardio-Vascular Diseases (CVD,) 244 age and sex-matched healthy controls); 374 male and female elderly Greek adults (154 CVD 220 age and sex-matched healthy controls). Inclusion: (ZINCAGE Project: www.zincage.org) Italian: diagnosis of ischemic heart disease and/or carotid heart disease; Greek: diagnoses history of angina, heart failure, coronary heart disease, stroke, myocardial infarction, or heart surgery. Exclusion: Italian controls: diabetes diagnosis | MT1A rs8052394 (A/C), rs11640851 (A/G) | Increased frequency of rs11640851 G allele carriers in Greek patients with ischemic heart and/or carotid heart disease in comparison to Greek controls. No difference between Italian patients and controls. | Intracellular zinc of peripheral blood cells decreased in CVD patients MT1A haplotype CG+ compared with MT1A CG−/CG− haplotype | N/A | No evidence of association between genotype, Zn, and biomarkers of disease, including inflammation and circulating lipids |
| Giacconi et al. 2006 [23] | Case-control study | 406 male and female older adults born and living in central Italy (105 with carotid stenosis and CVD (C); 111 with carotid stenosis, no symptoms or cardioischaemia (D); 190 age and sex-matched controls). Inclusion: ≥70 years, older individuals born and living in Central Italy admitted to INRCA Geriatric Hospital, Ancona, Italy, for endarterectomy | 1267 Hsp70-2 (A/G); (in NCBI database rs780016316 C/T) -308 TNFα G/A (in NCBI rs1800629 A/G) | 1267 Hsp70-2 G allele more frequent in (C) group than (D) | Plasma Zn similar across genotypic groups | N/A | No evidence of association between genotype, Zn, and biomarkers of disease, including hypertension and circulating lipids. |
| Mocchegiani et al. 2008 [42] | Non-randomised supplement clinical trial | 110 male and female healthy, non-institutionalised older adults from Italy, France, Germany, Poland, and Greece. Inclusion: (ZINCAGE Project: www.zincage.org). 60–84 years old, free of medication, plasma zinc ≤ 10.5 μM. Exclusion: autoimmune, neurodegenerative, cardiovascular, kidney or liver diseases, diabetes, infections, cancer, chronic inflammatory bowel disease or acrodermatitis enteropathica, sickle cell anaemia, chronic skin ulcerations, and endocrine disorders | −174 IL-6 G/C (in NCBI rs1800795 C/G) | No significant difference between carriers of alleles for immune parameters | GG genotype had significantly lower plasma Zn than C carriers. GG genotypes with normal plasma Zn still presented with impaired Zn status | 10 mg/day Zn-aspartate for 48 ± 2 days | Low plasma Zn associated with impaired immune response and psychological function independent of genotype GG genotype carriers are more predisposed to Zn deficiency and suggested as better candidates for supplementation |
| Mariani et al. 2008 [24] | Non-randomised supplement clinical trial | 39 male and female healthy older adults Inclusion: (ZINCAGE Project: www.zincage.org). 60–83 years old, healthy old people, still living independently, plasma Zn < 11μmol/L. Exclusion: taking medication, nutritional integrators, or vitamin complexes. | +647 A/C MT1A −174 IL-6 (in NCBI rs1800795 (C/G) | +647 MT1A genotype associated with increased inflammatory biomarkers | +647 MT1A C− allele associated with lower plasma Zn than C+ at basal and after supplementation. | 10 mg/day Zn-aspartate for 48 ± 2 days. Plasma Zn concentration significantly increased after supplementation in C+ carriers of −174 IL6. | Carriers of C- genotype of MT1A had lowest concentration of plasma zinc. Increment after supplementation was more pronounced in subjects carrying C- allele of MT1A /C+ −174 IL-6. Carries of C− MT1A/C −174 IL-6 did not respond to zinc supplementation. |
| Polymorphisms relating to Zn homeostasis | |||||||
| Whitfield et al. 2010 [34] | Cross-sectional study | 2926 male and female adult twins living in Australia Inclusion: Born between 1903 and 1964 (30–92 years old), enrolled in the Australian Twin Registry | N/A | N/A | 20% of the variation in plasma Zn concentration is due to genetic factors. | N/A | Genetic variability is contributing factor to Zn plasma variability. Specific genotypes not reported |
| Da Rocha et al. 2014b [33] | Cross-sectional study | 110 male and female older adults Inclusion: ≥50 years old adults Exclusion: use of vitamin supplements containing micronutrients | SLC30A3 rs11126936 (A/C) | N/A | CC genotypes had lower plasma Zn concentration than A carriers. CC genotype more frequent in participants with low plasma Zn | N/A | SLC30A3 polymorphism rs11126936 was associated with differences in plasma Zn concentration |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Day, K.J.; Adamski, M.M.; Dordevic, A.L.; Murgia, C. Genetic Variations as Modifying Factors to Dietary Zinc Requirements—A Systematic Review. Nutrients 2017, 9, 148. https://doi.org/10.3390/nu9020148
Day KJ, Adamski MM, Dordevic AL, Murgia C. Genetic Variations as Modifying Factors to Dietary Zinc Requirements—A Systematic Review. Nutrients. 2017; 9(2):148. https://doi.org/10.3390/nu9020148
Chicago/Turabian StyleDay, Kaitlin J., Melissa M. Adamski, Aimee L. Dordevic, and Chiara Murgia. 2017. "Genetic Variations as Modifying Factors to Dietary Zinc Requirements—A Systematic Review" Nutrients 9, no. 2: 148. https://doi.org/10.3390/nu9020148
APA StyleDay, K. J., Adamski, M. M., Dordevic, A. L., & Murgia, C. (2017). Genetic Variations as Modifying Factors to Dietary Zinc Requirements—A Systematic Review. Nutrients, 9(2), 148. https://doi.org/10.3390/nu9020148






