Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study
Abstract
1. Introduction
2. Materials and Methods
3. Results
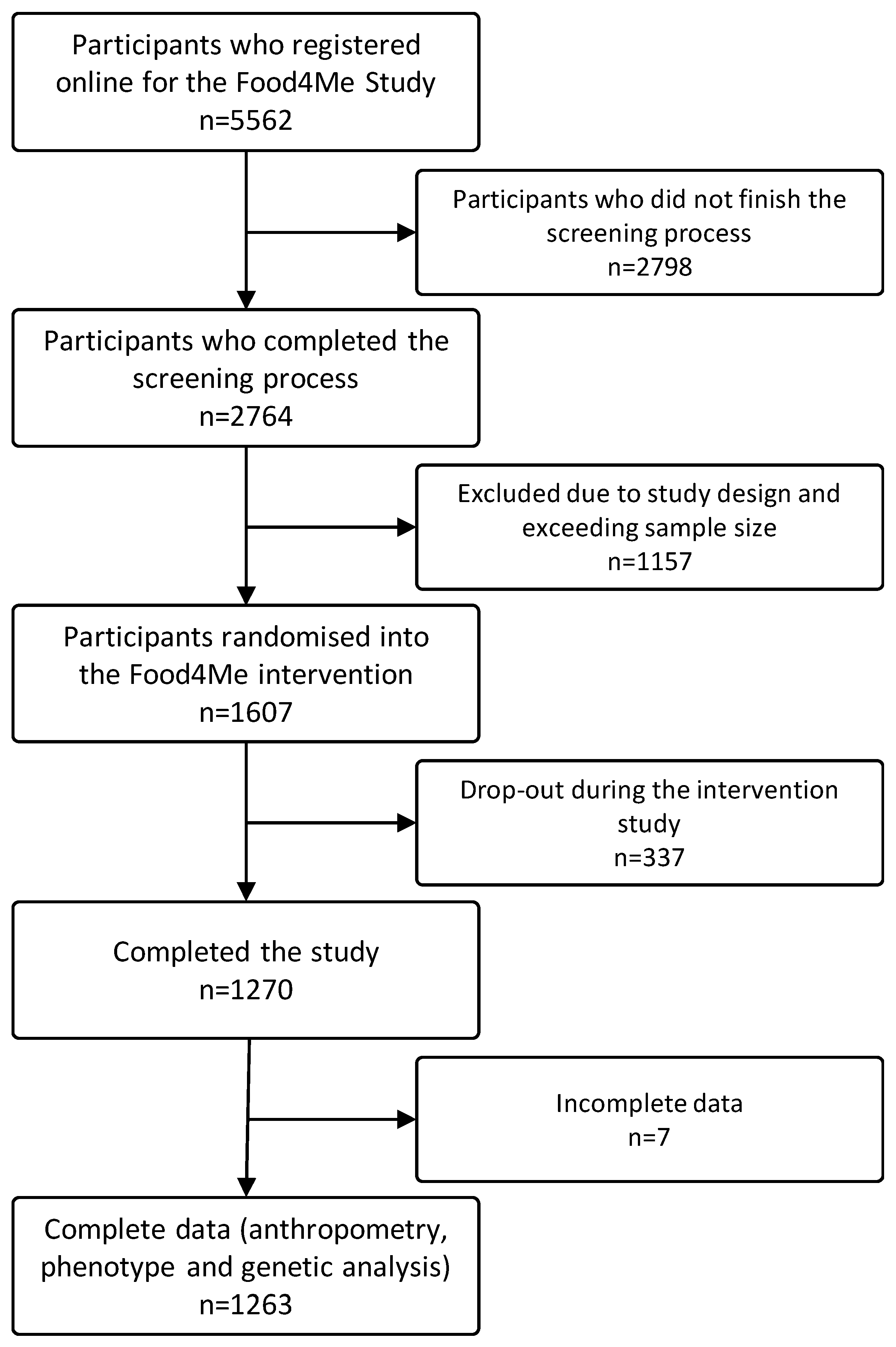
3.1. Baseline Characteristics of the Sample and Associations of GRS and MDS
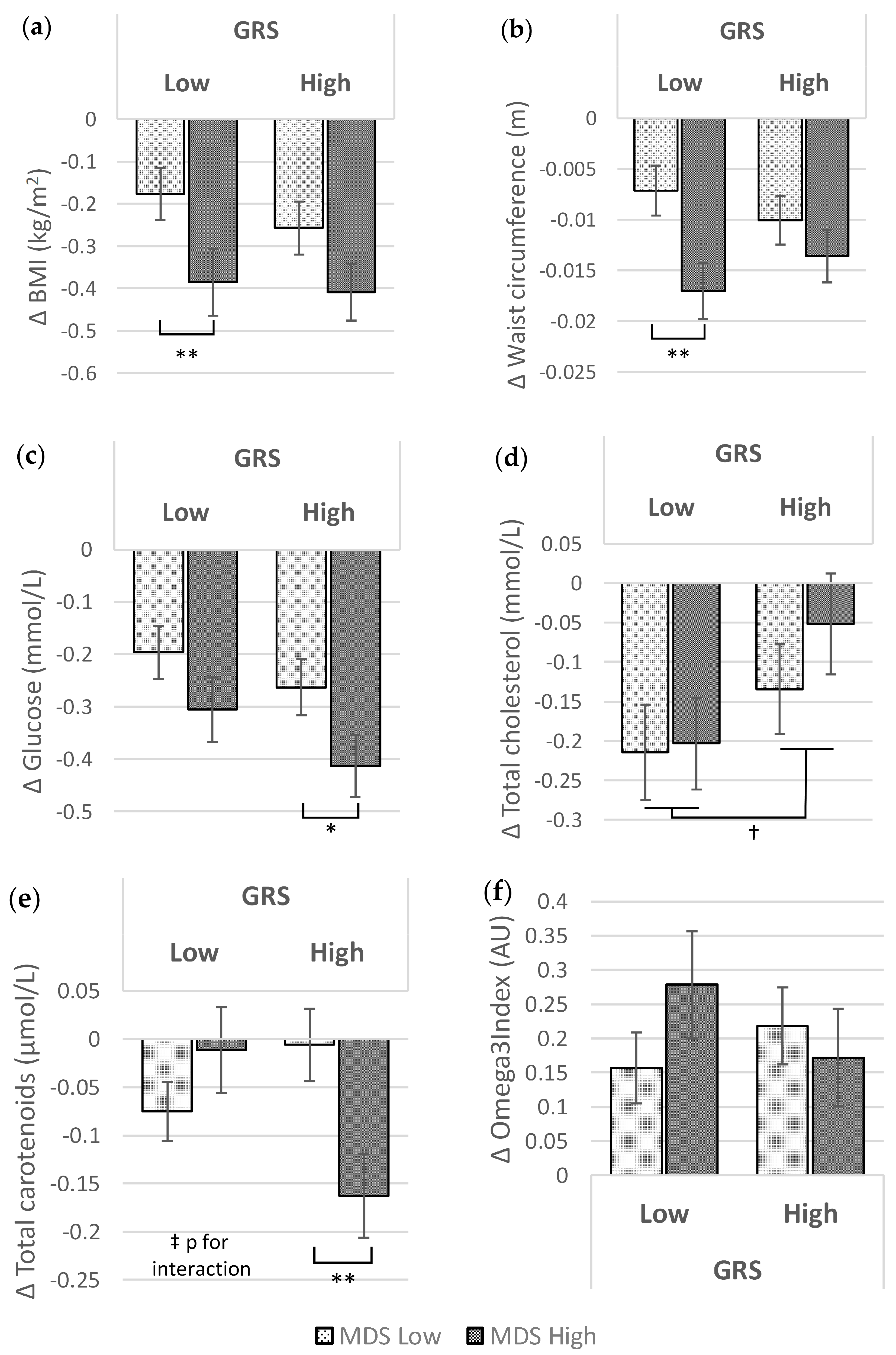
3.2. Associations of GRS and MDS at Baseline and after the Food4Me Intervention with Metabolic Traits
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- NCD Risk Factor Collaboration (NCD-RisC); Di Cesare, M.; Bentham, J.; Stevens, G.A.; Zhou, B.; Danaei, G.; Lu, Y.; Bixby, H.; Cowan, M.J.; Riley, L.M.; et al. Trends in adult body-mass index in 200 countries from 1975 to 2014: A pooled analysis of 1698 population-based measurement studies with 19·2 million participants. Lancet 2016, 387, 1377–1396. [Google Scholar] [CrossRef]
- GBD 2015 Obesity Collaborators; Afshin, A.; Forouzanfar, M.H.; Reitsma, M.B.; Sur, P.; Estep, K.; Lee, A.; Marczak, L.; Mokdad, A.H.; Moradi-Lakeh, M.; et al. Health Effects of Overweight and Obesity in 195 Countries over 25 Years. N. Engl. J. Med. 2017, 377, 13–27. [Google Scholar] [CrossRef]
- Vazquez, G.; Duval, S.; Jacobs, D.R.; Silventoinen, K. Comparison of Body Mass Index, Waist Circumference, and Waist/Hip Ratio in Predicting Incident Diabetes: A Meta-Analysis. Epidemiol. Rev. 2007, 29, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Danaei, G. Metabolic mediators of the effects of body-mass index, overweight, and obesity on coronary heart disease and stroke: A pooled analysis of 97 prospective cohorts with 1 8 million participants. Lancet 2014, 383, 970–983. [Google Scholar] [CrossRef]
- Livingston, G.; Sommerlad, A.; Orgeta, V.; Costafreda, S.G.; Huntley, J.; Ames, D.; Ballard, C.; Banerjee, S.; Burns, A.; Cohen-Mansfield, J.; et al. Dementia prevention, intervention, and care. Lancet 2017. [Google Scholar] [CrossRef]
- Popkin, B.M.; Gordon-Larsen, P. The nutrition transition: Worldwide obesity dynamics and their determinants. Int. J. Obes. Relat. Metab. Disord. 2004, 28 (Suppl. S3), S2–S9. [Google Scholar] [CrossRef] [PubMed]
- Jessri, M.; Wolfinger, R.D.; Lou, W.Y.; L’Abbé, M.R. Identification of dietary patterns associated with obesity in a nationally representative survey of Canadian adults: Application of a priori, hybrid, and simplified dietary pattern techniques. Am. J. Clin. Nutr. 2017, 105, 669–684. [Google Scholar] [CrossRef] [PubMed]
- Romieu, I.; Dossus, L.; Barquera, S.; Blottière, H.M.; Franks, P.W.; Gunter, M.; Hwalla, N.; Hursting, S.D.; Leitzmann, M.; Margetts, B.; et al. Energy balance and obesity: What are the main drivers? Cancer Causes Control 2017, 28, 247–258. [Google Scholar] [CrossRef] [PubMed]
- Moeller, S.M.; Reedy, J.; Millen, A.E.; Dixon, L.B.; Newby, P.K.; Tucker, K.L.; Krebs-Smith, S.M.; Guenther, P.M. Dietary Patterns: Challenges and Opportunities in Dietary Patterns Research. J. Am. Diet. Assoc. 2007, 107, 1233–1239. [Google Scholar] [CrossRef] [PubMed]
- Panagiotakos, D. α-priori versus α-posterior methods in dietary pattern analysis: A review in nutrition epidemiology. Nutr. Bull. 2008, 33, 311–315. [Google Scholar] [CrossRef]
- Widmer, R.J.; Flammer, A.J.; Lerman, L.O.; Lerman, A. The Mediterranean Diet, its Components, and Cardiovascular Disease. Am. J. Med. 2015, 128, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Estruch, R.; Martínez-González, M.A.; Corella, D.; Salas-Salvadó, J.; Fitó, M.; Chiva-Blanch, G.; Fiol, M.; Gómez-Gracia, E.; Arós, F.; Lapetra, J.; et al. Effect of a high-fat Mediterranean diet on bodyweight and waist circumference: A prespecified secondary outcomes analysis of the PREDIMED randomised controlled trial. Lancet Diabetes Endocrinol. 2016, 4, 666–676. [Google Scholar] [CrossRef]
- Martínez-González, M.A.; Salas-Salvadó, J.; Estruch, R.; Corella, D.; Fitó, M.; Ros, E. Benefits of the Mediterranean Diet: Insights From the PREDIMED Study. Prog. Cardiovasc. Dis. 2015, 58, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Grosso, G.; Marventano, S.; Yang, J.; Micek, A.; Pajak, A.; Scalfi, L.; Galvano, F.; Kales, S.N. A comprehensive meta-analysis on evidence of Mediterranean diet and cardiovascular disease: Are individual components equal? Crit. Rev. Food Sci. Nutr. 2017, 57, 3218–3232. [Google Scholar] [CrossRef] [PubMed]
- Razquin, C.; Martinez, J.A.; Martinez-Gonzalez, M.A.; Mitjavila, M.T.; Estruch, R.; Marti, A. A 3 years follow-up of a Mediterranean diet rich in virgin olive oil is associated with high plasma antioxidant capacity and reduced body weight gain. Eur. J. Clin. Nutr. 2009, 63, 1387–1393. [Google Scholar] [CrossRef] [PubMed]
- Babio, N.; Bulló, M.; Salas-Salvadó, J. Mediterranean diet and metabolic syndrome: The evidence. Public Health Nutr. 2009, 12, 1607. [Google Scholar] [CrossRef] [PubMed]
- Schwingshackl, L.; Hoffmann, G. Mediterranean dietary pattern, inflammation and endothelial function: A systematic review and meta-analysis of intervention trials. Nutr. Metab. Cardiovasc. Dis. 2014, 24, 929–939. [Google Scholar] [CrossRef] [PubMed]
- Godos, J.; Zappalà, G.; Bernardini, S.; Giambini, I.; Bes-Rastrollo, M.; Martinez-Gonzalez, M. Adherence to the Mediterranean diet is inversely associated with metabolic syndrome occurrence: A meta-analysis of observational studies. Int. J. Food Sci. Nutr. 2017, 68, 138–148. [Google Scholar] [CrossRef] [PubMed]
- Salas-Salvadó, J.; Garcia-Arellano, A.; Estruch, R.; Marquez-Sandoval, F.; Corella, D.; Fiol, M.; Gómez-Gracia, E.; Viñoles, E.; Arós, F.; Herrera, C.; et al. Components of the mediterranean-type food pattern and serum inflammatory markers among patients at high risk for cardiovascular disease. Eur. J. Clin. Nutr. 2008, 62, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Schwingshackl, L.; Christoph, M.; Hoffmann, G. Effects of Olive Oil on Markers of Inflammation and Endothelial Function—A Systematic Review and Meta-Analysis. Nutrients 2015, 7, 7651–7675. [Google Scholar] [CrossRef] [PubMed]
- Bonaccio, M.; Di Castelnuovo, A.; Costanzo, S.; De Lucia, F.; Olivieri, M.; Donati, M.B.; de Gaetano, G.; Iacoviello, L.; Bonanni, A. Moli-sani Project Investigators Nutrition knowledge is associated with higher adherence to Mediterranean diet and lower prevalence of obesity. Results from the Moli-sani study. Appetite 2013, 68, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Georgoulis, M.; Kontogianni, M.; Yiannakouris, N. Mediterranean Diet and Diabetes: Prevention and Treatment. Nutrients 2014, 6, 1406–1423. [Google Scholar] [CrossRef] [PubMed]
- Nissensohn, M.; Román-Viñas, B.; Sánchez-Villegas, A.; Piscopo, S.; Serra-Majem, L. The Effect of the Mediterranean Diet on Hypertension: A Systematic Review and Meta-Analysis. J. Nutr. Educ. Behav. 2016, 48, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Venturini, D.; Simão, A.N.C.; Urbano, M.R.; Dichi, I. Effects of extra virgin olive oil and fish oil on lipid profile and oxidative stress in patients with metabolic syndrome. Nutrition 2015, 31, 834–840. [Google Scholar] [CrossRef] [PubMed]
- Sureda, A.; Bibiloni, M.D.; Martorell, M.; Buil-Cosiales, P.; Marti, A.; Pons, A.; Tur, J.A.; Martinez-Gonzalez, M.A.; PREDIMED Study Investigators. Mediterranean diets supplemented with virgin olive oil and nuts enhance plasmatic antioxidant capabilities and decrease xanthine oxidase activity in people with metabolic syndrome: The PREDIMED study. Mol. Nutr. Food Res. 2016, 60, 2654–2664. [Google Scholar] [CrossRef] [PubMed]
- Di Daniele, N.; Noce, A.; Vidiri, M.F.; Moriconi, E.; Marrone, G.; Annicchiarico-Petruzzelli, M.; D’Urso, G.; Tesauro, M.; Rovella, V.; De Lorenzo, A. Impact of Mediterranean diet on metabolic syndrome, cancer and longevity. Oncotarget 2017, 8, 8947–8979. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, D.; Stice, E.; Rodríguez-López, R.; Manco, L.; Nóbrega, C. Current review of genetics of human obesity: From molecular mechanisms to an evolutionary perspective. Mol. Genet. Genomics 2015, 290, 1191–1221. [Google Scholar] [CrossRef] [PubMed]
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J.; et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Papandonatos, G.D.; Erar, B.; Florez, J.C.; Jablonski, K.A.; Razquin, C.; Marti, A.; Heianza, Y.; Huang, T.; et al. FTO genotype and weight loss: Systematic review and meta-analysis of 9563 individual participant data from eight randomised controlled trials. BMJ 2016, 354, i4707. [Google Scholar] [CrossRef] [PubMed]
- Parnell, L.D.; Blokker, B.A.; Dashti, H.S.; Nesbeth, P.-D.; Cooper, B.E.; Ma, Y.; Lee, Y.-C.; Hou, R.; Lai, C.-Q.; Richardson, K.; et al. CardioGxE, a catalog of gene-environment interactions for cardiometabolic traits. BioData Min. 2014, 7, 21. [Google Scholar] [CrossRef] [PubMed]
- Abraham, G.; Inouye, M. Genomic risk prediction of complex human disease and its clinical application. Curr. Opin. Genet. Dev. 2015, 33, 10–16. [Google Scholar] [CrossRef] [PubMed]
- McBride, C.M.; Koehly, L.M.; Sanderson, S.C.; Kaphingst, K.A. The Behavioral Response to Personalized Genetic Information: Will Genetic Risk Profiles Motivate Individuals and Families to Choose More Healthful Behaviors? Annu. Rev. Public Health 2010, 31, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Walter, S.; Mejía-Guevara, I.; Estrada, K.; Liu, S.Y.; Glymour, M.M. Association of a Genetic Risk Score With Body Mass Index Across Different Birth Cohorts. JAMA 2016, 316, 63. [Google Scholar] [CrossRef] [PubMed]
- Abraham, G.; Havulinna, A.S.; Bhalala, O.G.; Byars, S.G.; De Livera, A.M.; Yetukuri, L.; Tikkanen, E.; Perola, M.; Schunkert, H.; Sijbrands, E.J.; et al. Genomic prediction of coronary heart disease. Eur. Heart J. 2016, 37, 3267–3278. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Parast, L.; Cai, T.; Powers, C.; Gervino, E.V.; Hauser, T.H.; Hu, F.B.; Doria, A. Genetic susceptibility to coronary heart disease in type 2 diabetes: 3 independent studies. J. Am. Coll. Cardiol. 2011, 58, 2675–2682. [Google Scholar] [CrossRef] [PubMed]
- Meigs, J.B.; Shrader, P.; Sullivan, L.M.; McAteer, J.B.; Fox, C.S.; Dupuis, J.; Manning, A.K.; Florez, J.C.; Wilson, P.W.F.; D’Agostino, R.B.; et al. Genotype Score in Addition to Common Risk Factors for Prediction of Type 2 Diabetes. N. Engl. J. Med. 2008, 359, 2208–2219. [Google Scholar] [CrossRef] [PubMed]
- Kristiansson, K.; Perola, M.; Tikkanen, E.; Kettunen, J.; Surakka, I.; Havulinna, A.S.; Stancakova, A.; Barnes, C.; Widen, E.; Kajantie, E.; et al. Genome-Wide Screen for Metabolic Syndrome Susceptibility Loci Reveals Strong Lipid Gene Contribution But No Evidence for Common Genetic Basis for Clustering of Metabolic Syndrome Traits. Circ. Cardiovasc. Genet. 2012, 5, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Casas-Agustench, P.; Arnett, D.K.; Smith, C.E.; Lai, C.-Q.; Parnell, L.D.; Borecki, I.B.; Frazier-Wood, A.C.; Allison, M.; Chen, Y.-D.I.; Taylor, K.D.; et al. Saturated fat intake modulates the association between an obesity genetic risk score and body mass index in two US populations. J. Acad. Nutr. Diet. 2014, 114, 1954–1966. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Chu, A.Y.; Kang, J.H.; Huang, J.; Rose, L.M.; Jensen, M.K.; Liang, L.; Curhan, G.C.; Pasquale, L.R.; Wiggs, J.L.; et al. Fried food consumption, genetic risk, and body mass index: Gene-diet interaction analysis in three US cohort studies. BMJ 2014, 348, g1610. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Chu, A.Y.; Kang, J.H.; Jensen, M.K.; Curhan, G.C.; Pasquale, L.R.; Ridker, P.M.; Hunter, D.J.; Willett, W.C.; Rimm, E.B.; et al. Sugar-Sweetened Beverages and Genetic Risk of Obesity. N. Engl. J. Med. 2012, 367, 1387–1396. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.A.; Navas-Carretero, S.; Saris, W.H.M.; Astrup, A. Personalized weight loss strategies—The role of macronutrient distribution. Nat. Rev. Endocrinol. 2014, 10, 749–760. [Google Scholar] [CrossRef] [PubMed]
- Celis-Morales, C.; Livingstone, K.M.; Marsaux, C.F.M.; Forster, H.; O’Donovan, C.B.; Woolhead, C.; Macready, A.L.; Fallaize, R.; Navas-Carretero, S.; San-Cristobal, R.; et al. Design and baseline characteristics of the Food4Me study: A web-based randomised controlled trial of personalised nutrition in seven European countries. Genes Nutr. 2015, 10, 450. [Google Scholar] [CrossRef] [PubMed]
- Celis-Morales, C.; Livingstone, K.M.; Marsaux, C.F.M.; Macready, A.L.; Fallaize, R.; O’Donovan, C.B.; Woolhead, C.; Forster, H.; Walsh, M.C.; Navas-Carretero, S.; et al. Effect of personalized nutrition on health-related behaviour change: Evidence from the Food4me European randomized controlled trial. Int. J. Epidemiol. 2017, dyw186. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Macready, A.L.; Fallaize, R.; Forster, H.; Woolhead, C.; O’Donovan, C.B.; Marsaux, C.F.; Navas-Carretero, S.; San-Cristobal, R.; et al. Characteristics of European adults who dropped out from the Food4Me Internet-based personalised nutrition intervention. Public Health Nutr. 2017, 20, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Forster, H.; Fallaize, R.; Gallagher, C.; O’Donovan, C.B.; Woolhead, C.; Walsh, M.C.; Macready, A.L.; Lovegrove, J.A.; Mathers, J.C.; Gibney, M.J.; et al. Online Dietary Intake Estimation: The Food4Me Food Frequency Questionnaire. J. Med. Internet Res. 2014, 16, e150. [Google Scholar] [CrossRef] [PubMed]
- Fallaize, R.; Forster, H.; Macready, A.L.; Walsh, M.C.; Mathers, J.C.; Brennan, L.; Gibney, E.R.; Gibney, M.J.; Lovegrove, J.A. Online Dietary Intake Estimation: Reproducibility and Validity of the Food4Me Food Frequency Questionnaire Against a 4-Day Weighed Food Record. J. Med. Internet Res. 2014, 16, e190. [Google Scholar] [CrossRef] [PubMed]
- Marshall, S.J.; Livingstone, K.M.; Celis-Morales, C.; Forster, H.; Fallaize, R.; O’Donovan, C.B.; Woolhead, C.; Marsaux, C.F.; Macready, A.L.; Navas-Carretero, S.; et al. Reproducibility of the Online Food4Me Food-Frequency Questionnaire for Estimating Dietary Intakes across Europe. J. Nutr. 2016, 146, 1068–1075. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Navas-Carretero, S.; San-Cristobal, R.; Forster, H.; O’Donovan, C.B.; Woolhead, C.; Marsaux, C.F.M.; Macready, A.L.; Fallaize, R.; et al. Fat mass- and obesity-associated genotype, dietary intakes and anthropometric measures in European adults: The Food4Me study. Br. J. Nutr. 2016, 115, 440–448. [Google Scholar] [CrossRef] [PubMed]
- Celis-Morales, C.; Livingstone, K.M.; Woolhead, C.; Forster, H.; O’Donovan, C.B.; Macready, A.L.; Fallaize, R.; Marsaux, C.F.M.; Tsirigoti, L.; Efstathopoulou, E.; et al. How reliable is internet-based self-reported identity, socio-demographic and obesity measures in European adults? Genes Nutr. 2015, 10, 28. [Google Scholar] [CrossRef] [PubMed]
- Albani, V.; Celis-Morales, C.; Marsaux, C.F.M.; Forster, H.; O’Donovan, C.B.; Woolhead, C.; Macready, A.L.; Fallaize, R.; Navas-Carretero, S.; San-Cristobal, R.; et al. Exploring the association of dairy product intake with the fatty acids C15:0 and C17:0 measured from dried blood spots in a multipopulation cohort: Findings from the Food4Me study. Mol. Nutr. Food Res. 2016, 60, 834–845. [Google Scholar] [CrossRef] [PubMed]
- Markussen, M.S.; Veierød, M.B.; Sakhi, A.K.; Ellingjord-Dale, M.; Blomhoff, R.; Ursin, G.; Andersen, L.F. Evaluation of dietary patterns among Norwegian postmenopausal women using plasma carotenoids as biomarkers. Br. J. Nutr. 2015, 113, 672–682. [Google Scholar] [CrossRef] [PubMed]
- DeFina, L.F.; Vega, G.L.; Leonard, D.; Grundy, S.M. Fasting glucose, obesity, and metabolic syndrome as predictors of type 2 diabetes: The Cooper Center Longitudinal Study. J. Investig. Med. 2012, 60, 1164–1168. [Google Scholar] [CrossRef] [PubMed]
- Lamina, C.; Forer, L.; Schönherr, S.; Kollerits, B.; Ried, J.S.; Gieger, C.; Peters, A.; Wichmann, H.-E.; Kronenberg, F. Evaluation of gene-obesity interaction effects on cholesterol levels: A genetic predisposition score on HDL-cholesterol is modified by obesity. Atherosclerosis 2012, 225, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Gerber, M. Qualitative methods to evaluate Mediterranean diet in adults. Public Health Nutr. 2006, 9, 147–151. [Google Scholar] [CrossRef] [PubMed]
- Peterson, R.E.; Maes, H.H.; Holmans, P.; Sanders, A.R.; Levinson, D.F.; Shi, J.; Kendler, K.S.; Gejman, P.V.; Webb, B.T. Genetic risk sum score comprised of common polygenic variation is associated with body mass index. Hum. Genet. 2011, 129, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Cleves, M.A. Hardy-Weinberg Equilibrium Tests and Allele Frequency Estimation. Stata J. 1999, 48, 34–37. [Google Scholar]
- Machiela, M.J.; Chanock, S.J. LDlink: A web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 2015, 31, 3555–3557. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wu, J.; Yu, L. Association of Gln27Glu and Arg16Gly Polymorphisms in Beta2-Adrenergic Receptor Gene with Obesity Susceptibility: A Meta-Analysis. PLoS ONE 2014, 9, e100489. [Google Scholar] [CrossRef] [PubMed]
- Bruneau, M.L.; Johnson, B.T.; Huedo-Medina, T.B.; Larson, K.A.; Ash, G.I.; Pescatello, L.S. The blood pressure response to acute and chronic aerobic exercise: A meta-analysis of candidate gene association studies. J. Sci. Med. Sport 2016, 19, 424–431. [Google Scholar] [CrossRef] [PubMed]
- Kettunen, J.; Tukiainen, T.; Sarin, A.-P.; Ortega-Alonso, A.; Tikkanen, E.; Lyytikäinen, L.-P.; Kangas, A.J.; Soininen, P.; Würtz, P.; Silander, K.; et al. Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat. Genet. 2012, 44, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Frayling, T.M.; Timpson, N.J.; Weedon, M.N.; Zeggini, E.; Freathy, R.M.; Lindgren, C.M.; Perry, J.R.B.; Elliott, K.S.; Lango, H.; Rayner, N.W.; et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 2007, 316, 889–894. [Google Scholar] [CrossRef] [PubMed]
- Willer, C.J.; Speliotes, E.K.; Loos, R.J.F.; Li, S.; Lindgren, C.M.; Heid, I.M.; Berndt, S.I.; Elliott, A.L.; Jackson, A.U.; Lamina, C.; et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 2009, 41, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Perry, J.R.B.; Voight, B.F.; Yengo, L.; Amin, N.; Dupuis, J.; Ganser, M.; Grallert, H.; Navarro, P.; Li, M.; Qi, L.; et al. Stratifying Type 2 Diabetes Cases by BMI Identifies Genetic Risk Variants in LAMA1 and Enrichment for Risk Variants in Lean Compared to Obese Cases. PLoS Genet. 2012, 8, e1002741. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.J.; Zhang, F.; Richards, J.B.; Kestenbaum, B.; van Meurs, J.B.; Berry, D.; Kiel, D.P.; Streeten, E.A.; Ohlsson, C.; Koller, D.L.; et al. Common genetic determinants of vitamin D insufficiency: A genome-wide association study. Lancet 2010, 376, 180–188. [Google Scholar] [CrossRef]
- Ahn, J.; Yu, K.; Stolzenberg-Solomon, R.; Simon, K.C.; McCullough, M.L.; Gallicchio, L.; Jacobs, E.J.; Ascherio, A.; Helzlsouer, K.; Jacobs, K.B.; et al. Genome-wide association study of circulating vitamin D levels. Hum. Mol. Genet. 2010, 19, 2739–2745. [Google Scholar] [CrossRef] [PubMed]
- Teslovich, T.M.; Musunuru, K.; Smith, A.V.; Edmondson, A.C.; Stylianou, I.M.; Koseki, M.; Pirruccello, J.P.; Ripatti, S.; Chasman, D.I.; Willer, C.J.; et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 2010, 466, 707–713. [Google Scholar] [CrossRef] [PubMed]
- Cleves, M.A. Exploratory analysis of single nucleotide polymorphisms (SNPs) for quantitative traits. Stata J. 2005, 5, 141–153. [Google Scholar]
- Livingstone, K.M.; Celis-Morales, C.; Navas-Carretero, S.; San-Cristobal, R.; Macready, A.L.; Fallaize, R.; Forster, H.; Woolhead, C.; O’Donovan, C.B.; Marsaux, C.F.; et al. Effect of an Internet-based, personalized nutrition randomized trial on dietary changes associated with the Mediterranean diet: The Food4Me Study. Am. J. Clin. Nutr. 2016, 104, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Rokholm, B.; Silventoinen, K.; Tynelius, P.; Gamborg, M.; Sørensen, T.I.A.; Rasmussen, F. Increasing Genetic Variance of Body Mass Index during the Swedish Obesity Epidemic. PLoS ONE 2011, 6, e27135. [Google Scholar] [CrossRef] [PubMed]
- Roswall, N.; Angquist, L.; Ahluwalia, T.S.; Romaguera, D.; Larsen, S.C.; Ostergaard, J.N.; Halkjaer, J.; Vimaleswaran, K.S.; Wareham, N.J.; Bendinelli, B.; et al. Association between Mediterranean and Nordic diet scores and changes in weight and waist circumference: Influence of FTO and TCF7L2 loci. Am. J. Clin. Nutr. 2014, 100, 1188–1197. [Google Scholar] [CrossRef] [PubMed]
- Vaxillaire, M.; Yengo, L.; Lobbens, S.; Rocheleau, G.; Eury, E.; Lantieri, O.; Marre, M.; Balkau, B.; Bonnefond, A.; Froguel, P. Type 2 diabetes-related genetic risk scores associated with variations in fasting plasma glucose and development of impaired glucose homeostasis in the prospective DESIR study. Diabetologia 2014, 57, 1601–1610. [Google Scholar] [CrossRef] [PubMed]
- Andersson, E.A.; Allin, K.H.; Sandholt, C.H.; Borglykke, A.; Lau, C.J.; Ribel-Madsen, R.; Sparso, T.; Justesen, J.M.; Harder, M.N.; Jorgensen, M.E.; et al. Genetic Risk Score of 46 Type 2 Diabetes Risk Variants Associates With Changes in Plasma Glucose and Estimates of Pancreatic ?—Cell Function Over 5 Years of Follow-Up. Diabetes 2013, 62, 3610–3617. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ortega-Azorín, C.; Sorlí, J.V.; Asensio, E.M.; Coltell, O.; Martínez-González, M.; Salas-Salvadó, J.; Covas, M.-I.; Arós, F.; Lapetra, J.; Serra-Majem, L.; et al. Associations of the FTO rs9939609 and the MC4R rs17782313 polymorphisms with type 2 diabetes are modulated by diet, being higher when adherence to the Mediterranean diet pattern is low. Cardiovasc. Diabetol. 2012, 11, 137. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Huang, T.; Zheng, Y.; Rood, J.; Bray, G.A.; Sacks, F.M.; Qi, L. Genetic variation of fasting glucose and changes in glycemia in response to 2-year weight-loss diet intervention: The POUNDS LOST trial. Int. J. Obes. 2016, 40, 1164–1169. [Google Scholar] [CrossRef] [PubMed]
- Hivert, M.F.; Jablonski, K.A.; Perreault, L.; Saxena, R.; McAteer, J.B.; Franks, P.W.; Hamman, R.F.; Kahn, S.E.; Haffner, S.; Meigs, J.B.; et al. Updated Genetic Score Based on 34 Confirmed Type 2 Diabetes Loci Is Associated With Diabetes Incidence and Regression to Normoglycemia in the Diabetes Prevention Program. Diabetes 2011, 60, 1340–1348. [Google Scholar] [CrossRef] [PubMed]
- Kastorini, C.-M.; Milionis, H.J.; Esposito, K.; Giugliano, D.; Goudevenos, J.A.; Panagiotakos, D.B. The effect of Mediterranean diet on metabolic syndrome and its components: A meta-analysis of 50 studies and 534,906 individuals. J. Am. Coll. Cardiol. 2011, 57, 1299–1313. [Google Scholar] [CrossRef] [PubMed]
- Fitó, M.; Konstantinidou, V. Nutritional Genomics and the Mediterranean Diet’s Effects on Human Cardiovascular Health. Nutrients 2016, 8, 218. [Google Scholar] [CrossRef] [PubMed]
- De, R.; Verma, S.S.; Drenos, F.; Holzinger, E.R.; Holmes, M.V.; Hall, M.A.; Crosslin, D.R.; Carrell, D.S.; Hakonarson, H.; Jarvik, G.; et al. Identifying gene-gene interactions that are highly associated with Body Mass Index using Quantitative Multifactor Dimensionality Reduction (QMDR). BioData Min. 2015, 8, 41. [Google Scholar] [CrossRef] [PubMed]
- Nettleton, J.A.; Follis, J.L.; Ngwa, J.S.; Smith, C.E.; Ahmad, S.; Tanaka, T.; Wojczynski, M.K.; Voortman, T.; Lemaitre, R.N.; Kristiansson, K.; et al. Gene × dietary pattern interactions in obesity: Analysis of up to 68 317 adults of European ancestry. Hum. Mol. Genet. 2015, 24, 4728–4738. [Google Scholar] [CrossRef] [PubMed]
- Razquin, C.; Martinez, J.A.; Martinez-Gonzalez, M.A.; Bes-Rastrollo, M.; Fernández-Crehuet, J.; Marti, A. A 3-year intervention with a Mediterranean diet modified the association between the rs9939609 gene variant in FTO and body weight changes. Int. J. Obes. 2010, 34, 266–272. [Google Scholar] [CrossRef] [PubMed]
- Abete, I.; Parra, D.; Martinez, J.A. Legume-, Fish-, or High-Protein-Based Hypocaloric Diets: Effects on Weight Loss and Mitochondrial Oxidation in Obese Men. J. Med. Food 2009, 12, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Walker, C.G.; Holzapfel, C.; Loos, R.J.F.; Mander, A.P.; Klopp, N.; Illig, T.; Caterson, I.D.; Hauner, H.; Jebb, S.A. Genetic predisposition to an adverse lipid profile limits the improvement in total cholesterol in response to weight loss. Obesity 2013, 21, 2589–2595. [Google Scholar] [CrossRef] [PubMed]
- Walker, C.G.; Loos, R.J.F.; Olson, A.D.; Frost, G.S.; Griffin, B.A.; Lovegrove, J.A.; Sanders, T.A.B.; Jebb, S.A. Genetic predisposition influences plasma lipids of participants on habitual diet, but not the response to reductions in dietary intake of saturated fatty acids. Atherosclerosis 2011, 215, 421–427. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xu, M.; Ng, S.S.; Bray, G.A.; Ryan, D.H.; Sacks, F.M.; Ning, G.; Qi, L. Dietary Fat Intake Modifies the Effect of a Common Variant in the LIPC Gene on Changes in Serum Lipid Concentrations during a Long-Term Weight-Loss Intervention Trial. J. Nutr. 2015, 145, 1289–1294. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Carrasco, P.; Sorlí, J.V.; Estruch, R.; Rico-Sanz, J.; Martínez-González, M.Á.; Salas-Salvadó, J.; Covas, M.I.; Coltell, O.; Arós, F.; et al. Mediterranean diet reduces the adverse effect of the TCF7L2-rs7903146 polymorphism on cardiovascular risk factors and stroke incidence: A randomized controlled trial in a high-cardiovascular-risk population. Diabetes Care 2013, 36, 3803–3811. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, M.M.H.; Jones, P.J.H.; Eck, P.K. Nutrigenetics of cholesterol metabolism: Observational and dietary intervention studies in the postgenomic era. Nutr. Rev. 2015, 73, 523–543. [Google Scholar] [CrossRef] [PubMed]
- De, R.; Verma, S.S.; Holzinger, E.; Hall, M.; Burt, A.; Carrell, D.S.; Crosslin, D.R.; Jarvik, G.P.; Kuivaniemi, H.; Kullo, I.J.; et al. Identifying gene–gene interactions that are highly associated with four quantitative lipid traits across multiple cohorts. Hum. Genet. 2017, 136, 165–178. [Google Scholar] [CrossRef] [PubMed]
- Greene, G.W.; Resnicow, K.; Thompson, F.E.; Peterson, K.E.; Hurley, T.G.; Hebert, J.R.; Toobert, D.J.; Williams, G.C.; Elliot, D.L.; Goldman Sher, T.; et al. Correspondence of the NCI Fruit and Vegetable Screener to repeat 24-H recalls and serum carotenoids in behavioral intervention trials. J. Nutr. 2008, 138, 200S–204S. [Google Scholar] [PubMed]
- Dahl, L.; Mæland, C.A.; Bjørkkjær, T. A short food frequency questionnaire to assess intake of seafood and n-3 supplements: Validation with biomarkers. Nutr. J. 2011, 10, 127. [Google Scholar] [CrossRef] [PubMed]
- Lietz, G.; Hesketh, J. A network approach to micronutrient genetics: Interactions with lipid metabolism. Curr. Opin. Lipidol. 2009, 20, 112–120. [Google Scholar] [CrossRef] [PubMed]
- Patel, C.J.; Cullen, M.R.; Ioannidis, J.P.A.; Butte, A.J. Systematic evaluation of environmental factors: Persistent pollutants and nutrients correlated with serum lipid levels. Int. J. Epidemiol. 2012, 41, 828–843. [Google Scholar] [CrossRef] [PubMed]
- Wallström, P.; Wirfält, E.; Lahmann, P.H.; Gullberg, B.; Janzon, L.; Berglund, G. Serum concentrations of beta-carotene and alpha-tocopherol are associated with diet, smoking, and general and central adiposity. Am. J. Clin. Nutr. 2001, 73, 777–785. [Google Scholar] [PubMed]
- Johnstone, A.M.; Lobley, G.E.; Horgan, G.W.; Bremner, D.M.; Fyfe, C.L.; Morrice, P.C.; Duthie, G.G. Effects of a high-protein, low-carbohydrate v. high-protein, moderate-carbohydrate weight-loss diet on antioxidant status, endothelial markers and plasma indices of the cardiometabolic profile. Br. J. Nutr. 2011, 106, 282–291. [Google Scholar] [CrossRef] [PubMed]
- Damms-Machado, A.; Weser, G.; Bischoff, S.C. Micronutrient deficiency in obese subjects undergoing low calorie diet. Nutr. J. 2012, 11, 34. [Google Scholar] [CrossRef] [PubMed]
- Lietz, G.; Oxley, A.; Leung, W.; Hesketh, J. Single Nucleotide Polymorphisms Upstream from the -Carotene 15,15′-Monoxygenase Gene Influence Provitamin A Conversion Efficiency in Female Volunteers. J. Nutr. 2012, 142, 161S–165S. [Google Scholar] [CrossRef] [PubMed]
- Borel, P.; Desmarchelier, C. Genetic Variations Associated with Vitamin A Status and Vitamin A Bioavailability. Nutrients 2017, 9, 246. [Google Scholar] [CrossRef] [PubMed]
- Lietz, G.; Oxley, A.; Boesch-Saadatmandi, C. Consequences of Common Genetic Variations on β-Carotene Cleavage for Vitamin A Supply. In Carotenoids and Vitamin A in Translational Medicine; Sommerburg, O., Siems, W., Kraemer, K., Eds.; CRC Press: Boca Raton, FL, USA, 2013; pp. 383–396. [Google Scholar]
| Overall | GRS | p † | MDS | p † | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Low | High | Low | High | |||||||||
| n (female) | 1263 | (722) | 640 | (354) | 623 | (368) | 0.177 | 747 | (419) | 516 | (303) | 0.353 |
| Age (years) | 40.8 | ±13.0 | 41.2 | ±12.8 | 40.4 | ±13.1 | 0.357 | 40.2 | ±12.9 | 41.7 | ±13.0 | 0.034 |
| Ethnicity n (% Caucasians) | 1224 | (96.9%) | 618 | (96.6%) | 606 | (97.3%) | 0.397 | 730 | (97.7%) | 494 | (95.7%) | 0.230 |
| Smoke habit n (%) | ||||||||||||
| Never smoker | 781 | (61.8%) | 392 | (61.3%) | 389 | (62.4%) | 0.902 | 473 | (63.3%) | 308 | (59.7%) | 0.294 |
| Former smoker | 333 | (26.4%) | 172 | (26.9%) | 161 | (25.8%) | 185 | (24.8%) | 148 | (28.7%) | ||
| Smoker | 149 | (11.8%) | 76 | (11.9%) | 73 | (11.7%) | 89 | (11.9%) | 60 | (11.6%) | ||
| MDS (over 14) | 5.1 | ±1.7 | 5.1 | ±1.6 | 5.2 | ±1.7 | 0.529 | 4.0 | ±1.0 | 6.8 | ±0.9 | <0.001 |
| GRS (over 28) | 10.5 | ±2.3 | 8.6 | ±1.3 | 12.4 | ±1.3 | <0.001 | 10.5 | ±2.3 | 10.5 | ±2.4 | 0.974 |
| BMI (kg/m2) | 25.4 | ±4.7 | 25.2 | ±4.5 | 25.6 | ±4.8 | 0.018 | 25.6 | ±4.7 | 25.1 | ±4.6 | 0.012 |
| Waist circumference (m) | 0.859 | ±0.136 | 0.857 | ±0.133 | 0.861 | ±0.140 | 0.052 | 0.866 | ±0.138 | 0.848 | ±0.134 | 0.001 |
| Physical activity factor (AU) | 1.521 | ±0.104 | 1.525 | ±0.106 | 1.517 | ±0.101 | 0.094 | 1.516 | ±0.104 | 1.527 | ±0.103 | 0.021 |
| Energy intake reported (kcal/day) | 2552 | ±1066 | 2609 | ±1086 | 2493 | ±1042 | 0.079 | 2512 | ±1060 | 2609 | ±1072 | 0.069 |
| Glucose (mmol/L) | 3.73 | ±0.80 | 3.69 | ±0.80 | 3.77 | ±0.79 | 0.067 | 3.71 | ±0.75 | 3.76 | ±0.86 | 0.499 |
| Total cholesterol (mmol/L) | 4.61 | ±0.95 | 4.61 | ±0.93 | 4.60 | ±0.97 | 0.601 | 4.64 | ±0.96 | 4.55 | ±0.93 | 0.008 |
| Total carotenoids (μmol/L) | 1.52 | ±0.67 | 1.50 | ±0.64 | 1.55 | ±0.71 | 0.285 | 1.45 | ±0.60 | 1.64 | ±0.76 | <0.001 |
| Omega3 index (AU) | 5.71 | ±1.22 | 5.70 | ±1.20 | 5.73 | ±1.24 | 0.377 | 5.53 | ±1.08 | 5.97 | ±1.35 | <0.001 |
| Baseline GRS Category | p ‡ for Differences | ||||||
| Low | p † | High | p † | ||||
| BMI (kg/m2) | −0.281 | ±0.047 | <0.001 | −0.333 | ±0.044 | <0.001 | 0.417 |
| Waist circumference (m) | −0.012 | ±0.002 | <0.001 | −0.012 | ±0.002 | <0.001 | 0.920 |
| Glucose (mmol/L) | −0.251 | ±0.039 | <0.001 | −0.338 | ±0.039 | <0.001 | 0.114 |
| Total cholesterol (mmol/L) | −0.209 | ±0.040 | <0.001 | −0.093 | ±0.041 | 0.024 | 0.043 |
| Total carotenoids (μmol/L) | −0.043 | ±0.026 | 0.102 | −0.085 | ±0.028 | 0.003 | 0.282 |
| Omega3 index (AU) | 0.217 | ±0.045 | <0.001 | 0.195 | ±0.044 | <0.001 | 0.718 |
| Baseline MDS Category | p § for Differences | ||||||
| Low | p † | High | p † | ||||
| BMI (kg/m2) | −0.217 | ±0.044 | <0.001 | −0.397 | ±0.052 | <0.001 | 0.011 |
| Waist circumference (m) | −0.009 | ±0.002 | <0.001 | −0.015 | ±0.002 | <0.001 | 0.010 |
| Glucose (mmol/L) | −0.230 | ±0.036 | <0.001 | −0.360 | ±0.043 | <0.001 | 0.022 |
| Total cholesterol (mmol/L) | −0.174 | ±0.042 | <0.001 | −0.127 | ±0.044 | 0.003 | 0.453 |
| Total carotenoids (μmol/L) | −0.041 | ±0.024 | 0.097 | −0.087 | ±0.031 | 0.005 | 0.244 |
| Omega3 index (AU) | 0.187 | ±0.038 | <0.001 | 0.225 | ±0.053 | <0.001 | 0.573 |
| p † | GRS | p ‡ | ||||||
|---|---|---|---|---|---|---|---|---|
| Overall | Low | High | ||||||
| BMI (kg/m2) | −0.065 | ±0.022 | 0.003 | −0.060 | ±0.024 | −0.069 | ±0.021 | 0.457 |
| Waist circumference (m) | −0.002 | ±0.001 | 0.003 | −0.002 | ±0.001 | −0.002 | ±0.001 | 0.709 |
| Glucose (mmol/L) | −0.050 | ±0.017 | 0.003 | −0.043 | ±0.018 | −0.057 | ±0.017 | 0.182 |
| Total cholesterol (mmol/L) | 0.006 | ±0.018 | 0.753 | −0.004 | ±0.018 | 0.015 | ±0.018 | 0.058 |
| Total carotenoids (μmol/L) | −0.018 | ±0.012 | 0.118 | −0.014 | ±0.012 | −0.023 | ±0.012 | 0.206 |
| Omega3 index (AU) | −0.010 | ±0.019 | 0.611 | −0.007 | ±0.021 | −0.012 | ±0.020 | 0.677 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
San-Cristobal, R.; Navas-Carretero, S.; Livingstone, K.M.; Celis-Morales, C.; Macready, A.L.; Fallaize, R.; O’Donovan, C.B.; Lambrinou, C.P.; Moschonis, G.; Marsaux, C.F.M.; et al. Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study. Nutrients 2017, 9, 1107. https://doi.org/10.3390/nu9101107
San-Cristobal R, Navas-Carretero S, Livingstone KM, Celis-Morales C, Macready AL, Fallaize R, O’Donovan CB, Lambrinou CP, Moschonis G, Marsaux CFM, et al. Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study. Nutrients. 2017; 9(10):1107. https://doi.org/10.3390/nu9101107
Chicago/Turabian StyleSan-Cristobal, Rodrigo, Santiago Navas-Carretero, Katherine M. Livingstone, Carlos Celis-Morales, Anna L. Macready, Rosalind Fallaize, Clare B. O’Donovan, Christina P. Lambrinou, George Moschonis, Cyril F. M. Marsaux, and et al. 2017. "Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study" Nutrients 9, no. 10: 1107. https://doi.org/10.3390/nu9101107
APA StyleSan-Cristobal, R., Navas-Carretero, S., Livingstone, K. M., Celis-Morales, C., Macready, A. L., Fallaize, R., O’Donovan, C. B., Lambrinou, C. P., Moschonis, G., Marsaux, C. F. M., Manios, Y., Jarosz, M., Daniel, H., Gibney, E. R., Brennan, L., Drevon, C. A., Gundersen, T. E., Gibney, M., Saris, W. H. M., ... Martinez, J. A. (2017). Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study. Nutrients, 9(10), 1107. https://doi.org/10.3390/nu9101107








