Probiotics, Prebiotics and Immunomodulation of Gut Mucosal Defences: Homeostasis and Immunopathology
Abstract
:1. Introduction
2. Commensalism
3. The Mucosal Epithelial Barrier of the GIT
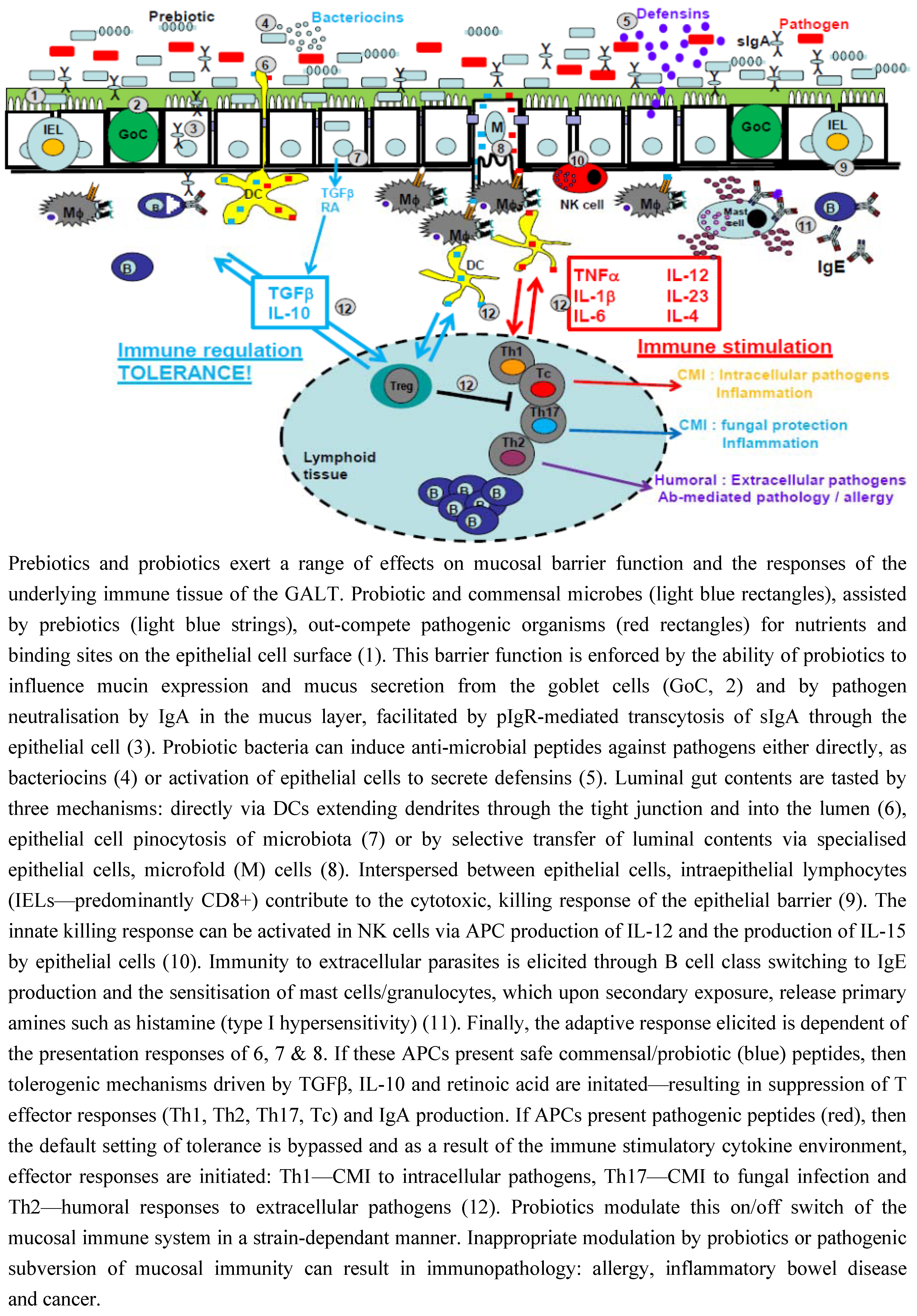
3.1. Intestinal Epithelial Cells
3.2. Mucus
3.3. IgA
3.4. Antimicrobial Peptides
3.5. Intraepithelial Lymphocytes
4. Immunomodulatory Role of Probiotic Bacteria
4.1. Tasting of Luminal Contents
4.2. Recognition of Pathogenic and Commensal Bacteria
4.3. Lumenal Contents Determine Immune Fate: Tolerance or Activation?
4.4. Cytokines Are Pivotal to This Immune Cell Fate
| Cytokines (Immune Response) | Cell system | Response | Probiotic strain | References |
|---|---|---|---|---|
| IFN-γ & IL-12 (Th1-associated, CMI and NK cell activity) | PBMCs | Increase | L. rhamnosus | [121] |
| L. plantarum L. lactis L. casei L. rhamnosus GG | [122] | |||
| L. lactis W58 | [123] | |||
| L. casei Shirota | [124] | |||
| L. casei Shirota | [125] | |||
| L. paracasei L. salivarius | [126] | |||
| B. longum W11 | [127] | |||
| L. rhamnosus L. gasseri B. bifidum E. coli (TG1) | [128] | |||
| L. casei Shirota | [129] | |||
| L. plantarum strains | [130] | |||
| PBMC-Mo | Increase | S. aureus L. johnsonii | [114] | |
| PBMC-DCs | Increase | L. salivarius L. rhamnosus Lcr35 | [131] [120] | |
| PBMC-NK cells | Increase | L. acidophilus L. reuteri | [132] | |
| Myeloid DCs | Increase | L. gasseri L. johnsonii L. reuteri | [133] | |
| PBMC-NK cells | Decrease | B. bifidum | [132] | |
| IL-23 & IL-17 (Th17-associated, pro-inflammatory) | Mo-DCs | Increase | L. rhamnosus Lcr35 | [120] |
| PBMCs | Decrease | B. breve LGG | [134] | |
| Caco-2 cell line | Decrease | L. plantarum | [135] | |
| IL-4 & IL-5 (Th2-associated, humoral) | PBMCs | Decrease | L. plantarum L. lactis L. casei L. rhamnosus GG | [122] |
| Bifidobacteria | [123] | |||
| L. rhamnosus L. gasseri B. bifidum | [128] | |||
| TGF-β (Treg-associated, anti-inflammatory) | PBMCs | Increase | B. longum | [136] |
| Epithelial cells | Increase | B. lactis L. johnsonii | [137] |
| Cytokines (Immune Response) | Cell system | Response | Probiotic strain | References |
|---|---|---|---|---|
| TNF-α and IL-1β (Pro-inflammatory) | PBMCs | Increase | L. rhamnosus L. bulgaricus S. pyogenes | [121] |
| Bifidobacteria | [142] | |||
| L. casei Shirota | [124] [129] | |||
| L. salivarius L. fermentum | [143] | |||
| L. plantarum strains | [130] | |||
| PBMC-DCs | Increase | L. rhamnosus Lcr35 | [120] | |
| Myeloid DCs | Increase | L. reuteri | [133] | |
| Epithelial cells | Increase | L. sakei | [137] | |
| Macrophage subset cell line | Increase and decrease (subset-specific) | L. casei Shirota | [139] [140] | |
| THP-1 cell line | Decrease | L. reuteri | [144] | |
| IL-6 (Pro-inflammatory) | PBMCs | Increase | L.rhamnosus L. bulgaricus S. pyogenes | [121] |
| Epithelial cells | Increase | B. lactis Bb12 L. casei CRL431 L. helveticus R389 | [145] [146] | |
| PBMCs | Decrease | L. casei Shirota | [129] | |
| IL-10 (Anti-inflammatory) | PBMCs | Increase | Bifidobacteria DNA | [147] [123] |
| Bifidobacteria | [142] | |||
| B. longum W11 | [127] | |||
| L. fermentum | [143] | |||
| L. acidophilus L. plantarum strains | [130] | |||
| L. acidophilus L. reuteri | [132] | |||
| PBMC-NK cells | Increase | B. bifidum VSL#3 L. reuteri | [147] [115] | |
| Blood-DCs | Increase | L. plantarum | ||
| Mo-DCs | Increase | L. casei L. rhamnosus Bifidobacteria | [148] [149] | |
| Mo-DCs | Increase | B. infantis | [150] | |
| Mo-DCs, mDCs, pDCs | Increase | |||
| PBMCs | Decrease | L. casei Shirota | [129] |
5. Immunopathology and Probiotic/Prebiotic Immunomodulation
5.1. Th1/Th17-Dominant Pathology, Crohn’s Disease
| Pathology | Response | Probiotic/Prebiotic | References |
|---|---|---|---|
| Crohn’s | ↓ IFN-γ, IL-12 | F. prausnitzii | [160] |
| ↑ IL-10 | Fructo-oligasaccharides | [167] | |
| Ulcerative colitis | ↓ IL-1β, TNF-α, IFN-γ, IL-12 | L. plantarum 299v | [168] |
| LGG | [169] | ||
| ↑ IL-10 | LGG | [169] | |
| ↓ β-defensins, TNF-α, IL-1, CRP | B. longum/Synergy 1 | [170] | |
| ↓ CRP | B. longum/psyllium | [171] | |
| ↓ adherence of B. vulgatas | LGG | [172] | |
| ↓ expression of tight junction proteins | VSL#3 | [172] | |
| ↓ tissue inflammation | VSL#3 | [173] | |
| ↑ no. γδ IEL ↓ no. γδ T-cells in lamina propria ↑ no. T-regs | L. acidophilus + B. longum | [87] | |
| Reshape microbiota composition | VSL#3 | [174] | |
| Colorectal Cancer | ↓ aberrant crypt formation ↓ cecal pH | Bifidobacteria+ Lactobacilli + Inulin+ Oligofructose | [175] |
| ↑ SCFA production ↓ IL-2 and iNOS | Bifidobacteria+ Lactobacilli + Inulin + Oligofructose | [176] | |
| Colorectal Cancer | ↓ H2O2 ↓ tissue inflammation ↑ catalase activity | L. lactis | [177] |
| ↓ IL-2 and iNOS | B. lactis | [176] | |
| L. rhamnosus | [178] | ||
| ↑ angiostatin ↑ no. CD4+ T-cells ↑ IL-17 and TNF-α | VSL#3 | [179] | |
| ↓ CXCR4 mRNA expression ↓ MHC-class 1 ↓ tumour growth ↑ CT-26 cancer cell apoptosis | L. acidophilus | [180] | |
| Allergy | ↓ IL-4 and IL-5 | L. plantarum | [122] |
| ↑ IL-12 and IFN-γ | L. lactis | [181] | |
| L. plantarum | [128] | ||
| ↓ IL-5, IL-4 and IL-13 | B. breve + oligosaccharide | [182] | |
| ↑ no. CD4+CD25+Foxp3+ T-regs | AhR ligand (TCDD) | [89] | |
| ↓ IL-12 ↔ no. T-regs | B. breve + oligosaccharide | [183] | |
| ↑ no. bifidobacteria colonies ↓ no. Clostridium colonies ↓ faecal pH ↔ IgE ↔ levels of acetic acid | B. breve + oligosaccharide | [184] | |
| Coeliac | ↑ no. CD4+ T-cells ↑ IFN-γ | L. casei | [185] |
| ↑ transepithelial resistance | B. lactis | [186] | |
| ↑ IL-12 and IFN-γ | Shigella CBD8 | [187] | |
| ↑ TNF-α | B. bifidum L. paracasei L. fermentum | [187] [188] |
5.2. Immunopathology: Th2-Dominant Pathology, Ulcerative Colitis
5.2.1. Anti-Tumoural Responses and NK Cell Activation
5.2.2. Colorectal Cancer
5.3. Immunopathology: Hypersensitivity
5.3.1. Allergy & Type I Hypersensitivity
5.3.2. Coeliac Disease
6. Future Perspectives
7. Conclusions
Conflict of Interest
References
- FAO/WHO. Guidelines for the Evaluation of Probiotics in Food. 2002. Available online: ftp://ftp.fao.org/es/esn/food/wgreport2.pdf (accessed on 1 May 2013).
- Maurice, C.F.; Haiser, H.J.; Turnbaugh, P.J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 2013, 152, 39–50. [Google Scholar] [CrossRef]
- Bentley, R.; Meganathan, R. Biosynthesis of vitamin K (menaquinone) in bacteria. Mircobiol. Rev. 1982, 46, 241–280. [Google Scholar]
- Nilsson, A.C.; Östman, E.M.; Holst, J.; Björk, I.M. Including indigestible carbohydrates in the evening meal of healthy subjects improves glucose tolerance, lowers inflammatory marks, and increases satiety after subsequent standardized breakfast. J. Nutr. 2008, 138, 732–739. [Google Scholar]
- Matsumoto, M.; Ishige, A.; Yazawa, Y.; Kondo, M.; Muramatsu, K.; Watanabe, K. Promotion of intestinal peristalsis by Bifidobacterium spp. Capable of hydrolysis sennosides in mice. PLoS One 2012, 7, e31700. [Google Scholar]
- Candela, M.; Perna, F.; Carnevali, P.; Vitali, B.; Ciati, R.; Gionchetti, P.; Rizzello, F.; Campieri, M.; Brigidi, P. Interaction of probiotic Lactobacillus and Bifidobacterium strains with human intestinal epithelial cells: Adhesion properties, competition against enteropathogens and modulation of IL-8 production. Int. J. Food Microbiol. 2008, 125, 286–292. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Lui, C.; Tzianaboz, A.O.; Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 2005, 122, 107–118. [Google Scholar] [CrossRef]
- Zschüttig, A.; Zimmermann, K.; Blom, J.; Goesmann, A.; Pöhlmann, C.; Gunzer, F. Identification and characterisation of Microcin S, a new antibacterial peptide produced by probiotic Escherichia coli G3/10. PLoS One 2012, 7, e33351. [Google Scholar]
- Gibson, G.R.; Roberfroid, M.B. Dietary modulation of the human colonic microbiota: Introducing the concept of prebiotics. J. Nutr. 1995, 125, 1401–1412. [Google Scholar]
- Van Zanten, G.C.; Knudsen, A.; Röytiö, H.; Forssten, S.; Lawther, M.; Blennow, A.; Lahtinen, S.J.; Jakobsen, M.; Svensson, B.; Jespersen, L. The effect of selected synbiotics on microbial composition and short-chain fatty acid production in model system of the human colon. PLoS One 2012, 7, e47212. [Google Scholar] [CrossRef]
- Knol, J.; Scholtens, P.; Kafka, C.; Steenbakkers, J.; Gro, S.; Helm, K.; Klarczyk, M.; Schöpfer, H.; Böckler, H.M.; Wells, J. Colon microflora in infants fed formula with galacto- and fructo- oligosaccharides: More like breast-fed infants. J. Pediatr. Gastroenterol. Nutr. 2005, 40, 36–42. [Google Scholar] [CrossRef]
- Holscher, H.D.; Faust, K.L.; Czerkies, L.A.; Litov, R.; Ziegler, E.E.; Lessin, H.; Hatch, T.; Sun, S.; Tappenden, K.A. Effects of prebiotic-containing infant formula on gastrointestinal tolerance and fecal microbiota in a randomized controlled trial. JPEN J. Parenter. Enteral Nutr. 2012, 36, 95S–105S. [Google Scholar] [CrossRef]
- Roberfroid, M.; Gibson, G.R.; Hoyles, L.; McCartney, N.L.; Rastall, R.; Rowland, I.; Wolvers, D.; Watzl, B.; Szajewska, H.; Stahl, B.; et al. Prebiotic effects metabolic and health benefits. Br. J. Nutr. 2010, 104, S1–S63. [Google Scholar]
- Yazawa, K.; Imai, K.; Tamura, Z. Oligosaccharides and polysaccharides specifically utilizable by bifidobacteria. Chem. Pharm. Bull. 1978, 26, 3306–3311. [Google Scholar] [CrossRef]
- Mitsuoak, T.; Hidaka, H.; Eida, T. Effect of fructo-oligosaccharides on intestinal microflora. Nahrung 1987, 31, 427–436. [Google Scholar] [CrossRef]
- Nepelska, M.; Cultrone, A.; Béguet-Crespel, F.; Le Roux, K.; Doré, J.; Arulampalam, V.; Blottière, H.M. Butyrate produced by commensal bacteria potentiates phorbol esters induced AP-1 response in human intestinal epithelial cells. PLoS One 2012, 7, e52869. [Google Scholar] [CrossRef]
- Ohigashi, S.; Sudo, K.; Kobayashi, D.; Takahashi, O.; Takahashi, T.; Asahara, T.; Nomoto, K.; Onodera, H. Changes of the intestinal microbiota, short chain fatty acids and fecal pH in patients with colorectal cancer. Digest. Dis. Sci. 2013, in press. [Google Scholar]
- Whitman, W.B.; Coleman, D.C.; Wiebe, W.J. Prokaryotes: The unseen majority. Proc. Natl. Acad. Sci. USA 1998, 95, 6578–6583. [Google Scholar] [CrossRef]
- Bäckhed, F.; Ley, R.E.; Sonnenburg, J.L.; Peterson, D.A.; Gordon, J.I. Host-bacterial mutualism in the human intestine. Science 2005, 307, 1915–1920. [Google Scholar] [CrossRef]
- Gill, S.R.; Pop, M.; DeBoy, R.T.; Eckburg, P.B.; Turnbaugh, P.J.; Samuel, B.S.; Gordon, J.I.; Relman, D.A.; Fraser-Liggett, C.M.; Nelson, K.E. Metagenomic analysis of the human distal gut microbiome. Science 2006, 312, 1355–1359. [Google Scholar] [CrossRef]
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef]
- Penders, J.; Thijs, C.; Vink, C.; Stelma, F.F.; Snijders, B.; Kummeling, I.; van den Brandt, P.A.; Stobberingh, E.E. Factors influencing the composition of the intestinal microbiota in early infancy. Paediatrics 2006, 118, 511–521. [Google Scholar] [CrossRef]
- Biasucci, G.; Benenati, B.; Morelli, L.; Bessi, E.; Boehm, G. Cesarean delivery may affect the early biodiversity of intestinal bacteria. J. Nutr. 2008, 138, 1796S–1800S. [Google Scholar]
- Huurre, A.; Kalliomäki, M.; Rautava, S.; Rinne, M.; Salminen, S.; Isolauri, E. Mode of delivery—Effects on gut microbiota and humoral immunity. Neonatology 2008, 93, 236–240. [Google Scholar] [CrossRef]
- Liao, Y.; Jiang, R.; Lönnerdal, B. Biochemical and molecular impacts of lactoferrin on small intestinal growth and development. Biochem. Cell Biol. 2012, 90, 476–484. [Google Scholar] [CrossRef]
- Martín, R.; Langa, S.; Reviriego, C.; Jimínez, E.; Marín, M.L.; Xaus, J.; Fernández, L.; Rodríguez, J.M. Human milk is a source of lactic acid bacteria for the infant gut. J. Pediatr. 2003, 143, 754–758. [Google Scholar] [CrossRef]
- Martín, R.; Jiménez, E.; Heilig, H.; Fernández, L.; Marín, M.L.; Zoetendal, E.G.; Rodríguez, J.M. Isolation of bifidobacteria from breast milk and assessment of the bifidobacterial population by PCR-denaturing gradient gel electrophoresis and quantitative real-time PCR. Appl. Environ. Microbiol. 2009, 74, 965–969. [Google Scholar]
- Albesharat, R.; Ehrmann, M.A.; Korakli, M.; Yazaji, S.; Vogel, R.F. Phenotypic and genotypic analyses of lactic acid bacteria in local fermented food, breast milk and faeces of mothers and their babies. Syst. Appl. Microbiol. 2011, 34, 148–155. [Google Scholar] [CrossRef]
- Martı́n, R.; Langa, S.; Reviriegoa, C.; Jiméneza, E.; Marı́na, M.L.; Olivaresb, M.; Bozab, J.; Jiménezb, J.; Fernándeza, L.; Xausb, J.; et al. The commensal microflora of human milk: New perspectives for food bacteriotherapy and probiotics. Trends Food Sci. Technol. 2004, 15, 121–127. [Google Scholar] [CrossRef]
- Garrido, D.; Nwosu, C.; Ruiz-Moyano, S.; Aldredge, D.; German, J.B.; Lebrilla, C.B.; Mills, D.A. Endo-β-n-acetylglucosaminidases from infant gut-associated bifidobacteria release complex n-glycans from human milk glycoproteins. Mol. Cell. Proteomics 2012, 11, 775–785. [Google Scholar] [CrossRef]
- Yolken, R.H.; Peterson, J.A.; Vonderfecht, S.L.; Fouts, E.T.; Midthun, K.; Newburg, D.S. Human milk mucin inhibits rotavirus replication and prevents experimental gastroenteritis. J. Clin. Invest. 1992, 90, 1984–1991. [Google Scholar] [CrossRef]
- Dobson, A.; Crispie, F.; Rea, M.C.; O’Sullivan, O.; Casey, P.G.; Lawlor, P.G.; Cotter, P.D.; Ross, P.; Gardiner, G.E.; Hill, C. Fate and efficacy of lacticin 3147-producing Lactococcus lactis in the mammalian gastrointestinal tract. FEMS Microbiol. Ecol. 2011, 76, 602–614. [Google Scholar] [CrossRef]
- Yu, Z.T.; Chen, C.; Kling, D.E.; Liu, B.; McCoy, J.M.; Merighi, M.; Heidtman, M.; Newburg, D.S. The principal fucosylated oligosaccharides of human milk exhibit prebiotic properties on cultured infant microbiota. Glycobiology 2013, 23, 169–177. [Google Scholar] [CrossRef]
- Eggesbø, M.; Moen, B.; Peddada, S.; Baird, D.; Rugtveit, J.; Midtvedt, T.; Bushel, P.R.; Sekelja, M.; Rudi, K. Development of gut microbiota in infants not exposed to medical interventions. APMIS 2011, 119, 17–35. [Google Scholar] [CrossRef]
- Savino, F.; Roana, J.; Mandras, N.; Tarasco, V.; Locatelli, E.; Tullio, V. Faecal microbiota in breast-fed infants after antibiotic therapy. Acta Paediatr. 2011, 100, 75–78. [Google Scholar] [CrossRef]
- Bartizal, K.F.; Wostmann, B.S.; Wagner, M. Distribution and effects of a defined six-member murine derived microflora in gnotobiotic gerbils. Appl. Environ. Microbiol. 1984, 47, 746–751. [Google Scholar]
- Pollard, M.; Sharon, N. Responses of the Peyer’s patches in germ-free mice to antigenic stimulation. Infect. Immun. 1970, 2, 96–100. [Google Scholar]
- Williams, A.M.; Probert, C.S.; Stepankova, R.; Tlaskalova-Hogenova, H.; Phillips, A.; Bland, P.W. Effects of microflora on the neonatal development of gut mucosal T cells and myeloid cells in the mouse. Immunology 2006, 119, 470–478. [Google Scholar] [CrossRef]
- Yamamoto, M.; Yamaguchi, R.; Munakata, K.; Takashima, K.; Nishiyama, M.; Hioki, K.; Ohnishi, Y.; Nagasaki, M.; Imoto, S.; Miyano, S.; et al. A microarray analysis of gnotobiotic mice indicating that microbial exposure during the neonatal period plays an essential role in immune system development. BMC Genomics 2012, 13, 335. [Google Scholar] [CrossRef]
- Rhee, K.; Sethupathi, P.; Dirks, A.; Lanning, D.K.; Knight, K.L. Role of commensal bacteria in development of gut-associated lymphoid tissues and preimmune antibody repertoire. J. Immunol. 2004, 172, 1118–1124. [Google Scholar]
- Russell, S.L.; Gold, M.J.; Willing, B.P.; Thorson, L.; McNagny, K.M.; Finlay, B.B. Perinatal antibiotic treatment affects murine microbiota, immune responses and allergic asthma. Gut Microbes 2013, 4, 158–164. [Google Scholar] [CrossRef]
- King, C.; Sarvetnick, N. The incidence of type-1 diabetes in NOD mice is modulated by restricted flora not germ-free conditions. PLoS One 2011, 6, e17049. [Google Scholar] [CrossRef]
- Ochoa-Repáraz, J.; Mielcarz, D.W.; Wang, Y.; Begum-Haque, S.; Dasgupta, S.; Kasper, D.L.; Kasper, L.H. A polysaccharide from the human commensal Bacteroides fragilis protects against CNS demyelinating disease. Mucosal Immunol. 2010, 3, 487–495. [Google Scholar] [CrossRef]
- Strachan, D.P. Hay fever, hygiene, and household size. Br. Med. J. 1989, 299, 1259–1260. [Google Scholar] [CrossRef]
- Aumeunier, A.; Grela, F.; Ramadan, A.; Pham Van, L.; Bardel, E.; Gomez Alcala, A.; Jeannin, P.; Akira, S.; Bach, J.F.; Thieblemont, N. Systemic Toll-like receptor stimulation suppresses experimental allergic asthma and autoimmune diabetes in NOD mice. PLoS One 2010, 5, e11484. [Google Scholar] [CrossRef]
- Gross, F.; van der Meulen, J.; Snel, J.; van der Meer, R.; Kleerebezem, M.; Niewold, T.V.; Hulst, M.M.; Smiths, M.A. Mannose-specific interaction of Lactobacillus plantarum with porcine jejunal epithelium. FEMS Immunol. Med. Microbiol. 2008, 54, 215–223. [Google Scholar] [CrossRef]
- Wilson, K.H.; Perini, F. Role of competition for nutrients in suppression of Clostridium difficile by the colonic microflora. Infect. Immun. 1988, 56, 2610–2614. [Google Scholar]
- Maltby, R.; Leatham-Jensen, M.P.; Gibson, T.; Cohen, P.S.; Conway, T. Nutritional basis for colonization resistance by human commensal Escherichia coli strains HS and nissle 1917 against E. coli O157:H7 in the mouse intestine. PLoS One 2013, 8, e53957. [Google Scholar]
- Resta-Lenert, S.; Barrett, K.E. Live probiotics protect intestinal epithelial cells from the effects of infection with enteroinvasive Escherichia coli (EIEC). Gut 2003, 52, 988–997. [Google Scholar] [CrossRef]
- Cario, E.; Gerken, G.; Podolsky, D.K. Toll-like receptor 2 enhances ZO-1-associated intestinal epithelial barrier integrity via protein kinase C. Gastroenterology 2004, 127, 224–238. [Google Scholar] [CrossRef]
- Petrof, E.O.; Kojima, K.; Ropeleski, M.J.; Tao, Y.; de Simone, C.; Chang, E.B. Probiotics inhibit nuclear factor kappaB and induce heat shock proteins in colonic epithelial cells through proteasome inhibition. Gastroenterology 2004, 127, 1474–1487. [Google Scholar] [CrossRef]
- Tao, Y.; Drabik, K.A.; Waypa, T.S.; Musch, M.W.; Alverdy, J.C.; Schneewind, O.; Chang, E.B.; Petrof, E.O. Soluble factors from Lactobacillus GG activate MAPKs and induce cytoprotective heat shock proteins in intestinal epithelial cells. Am. J. Physiol. Cell Physiol. 2006, 290, C1018–C1030. [Google Scholar]
- Park, H.S.; Cho, S.G.; Kim, C.K.; Hwang, H.S.; Noh, K.T.; Kim, M.S.; Huh, S.H.; Kim, M.J.; Ryoo, K.; Kim, E.K.; et al. Heat shock protein hsp72 is a negative regulator of apoptosis signal-regulating kinase 1. Mol. Cell Biol. 2002, 22, 7721–7730. [Google Scholar] [CrossRef]
- Liu, T.S.; Musch, M.W.; Sugi, K.; Walsh-Reitz, M.M.; Ropeleski, M.J.; Hendrickson, B.A.; Pothoulakis, C.; Lamont, J.T.; Chang, E.B. Protective role of HSP72 against Clostridium difficile toxin A-induced intestinal epithelial cell dysfunction. Am. J. Physiol. Cell Physiol. 2003, 284, C1073–C1082. [Google Scholar] [CrossRef]
- Gendler, S.J.; Spicer, A.P. Epithelial mucin genes. Annu. Rev. Physiol. 1995, 57, 607–634. [Google Scholar] [CrossRef]
- Van Tassell, M.L.; Miller, M.J. Lactobacillus adhesion to mucus. Nutrients 2011, 3, 613–636. [Google Scholar] [CrossRef]
- Mack, D.R.; Michail, S.; Wei, S.; McDougall, L.; Hollingsworth, M.A. Probiotics inhibit enteropathogenic E. coli adherence in vitro by inducing intestinal mucin gene expression. Am. J. Physiol. 1999, 276, G941–G950. [Google Scholar]
- Caballero-Franco, C.; Keller, K.; de Simone, C.; Chadee, K. The VSL#3 probiotic formula induces mucin gene expression and secretion in colonic epithelial cells. Am. J. Physiol. Gastrointest. Liver Physoil. 2007, 292, 315–322. [Google Scholar] [CrossRef]
- Perez-Vilar, J.; Hill, R.L. The structure and assembly of secreted mucins. J. Biol. Chem. 1999, 274, 31751–31754. [Google Scholar] [CrossRef]
- Katayama, T.; Fujita, K.; Yamamoto, K. Novel bifidobacterial glycosides action on sugar chains of mucin glycoproteins. J. Biosci. Bioeng. 2005, 99, 457–465. [Google Scholar] [CrossRef]
- Kiyohara, M.; Nakatomi, T.; Kurihara, S.; Fushinobu, S.; Suzuki, H.; Tanaka, T.; Shoda, S.; Kitaoka, M.; Katayama, T.; Yamamoto, K.; et al. α-n-acetylgalactosaminidase from infant-associated bifidobacteria belonging to novel glycoside hydrolase family 129 is implicated in alternative mucin degradation pathway. J. Biol. Chem. 2012, 287, 693–700. [Google Scholar] [CrossRef]
- Biesbrock, A.R.; Reddy, M.S.; Levine, M.J. Interaction of a salivary mucin-secretory immunoglobulin A complex with mucosal pathogens. Infect. Immun. 1991, 59, 3492–3497. [Google Scholar]
- Rautava, S.; Arvilommi, H.; Isolaur, E. Specific probiotics in enhancing maturation of IgA responses in formula-fed infants. Pediatr. Res. 2006, 60, 221–224. [Google Scholar] [CrossRef]
- Rodrigues, A.C.; Cara, D.C.; Fretez, S.H.; Cunha, F.Q.; Vieira, E.C.; Nicoli, J.R.; Vieira, L.Q. Saccharomyces boulardii stimulates sIgA production and the phagocytic system of gnotobiotic mice. J. Appl. Microbiol. 2000, 89, 404–414. [Google Scholar] [CrossRef]
- He, B.; Xu, W.; Santini, P.A.; Polydorides, A.D.; Chiu, A.; Estrella, J.; Shan, M.; Chadburn, A.; Villanacci, V.; Plebani, A.; et al. Intestinal bacteria trigger T cell-independent immunoglobulin A2 class switching by inducing epithelial-cell secretion of the cytokine APRIL. Immunity 2007, 26, 812–826. [Google Scholar] [CrossRef]
- Shang, L.; Fukata, M.; Thirunarayanan, N.; Martin, A.P.; Arnaboldi, P.; Maussang, D.; Berin, C.; Unkeless, J.C.; Mayer, L.; Abreu, M.T.; et al. Toll-like receptor signaling in small intestinal epithelium promotes B-cell recruitment and IgA production in lamina propria. Gastroenterology 2008, 135, 529–538. [Google Scholar] [CrossRef]
- Reséndiz-Albor, A.A.; Reina-Garfias, H.; Rojas-Hernández, S.; Jarillo-Luna, A.; Rivera-Aguilar, V.; Miliar-García, A.; Campos-Rodríguez, R. Regionalization of pIgR expression in the mucosa of mouse small intestine. Immunol. Lett. 2010, 128, 59–67. [Google Scholar] [CrossRef]
- Gillor, O.; Giladi, I.; Riley, M.A. Persistance of colicinogenic Escherichia coli in the mouse gastrointestinal tract. BMC Microbiol. 2009, 9, 165. [Google Scholar] [CrossRef]
- Tong, Z.; Zhou, L.; Li, J.; Kuang, R.; Lin, Y.; Ni, L. An in vitro investigation of Lactococcus lactis antagonising cariogenic bacterium Streptococcus mutans. Arch. Oral Biol. 2012, 57, 376–382. [Google Scholar] [CrossRef]
- Corr, S.C.; Li, Y.; Riedel, C.U.; O’Toole, P.W.; Hill, C.; Gahan, C.G. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc. Natl. Acad. Sci. USA 2007, 104, 7617–7621. [Google Scholar] [CrossRef]
- Meijerink, M.; van Hemert, S.; Taverne, N.; Wels, M.; de Vos, P.; Bron, P.A.; Savelkoul, H.F.; van Bilsen, J.; Kleerebezem, M.; Wells, J.M. Identification of genetic loci in Lactobacillus plantarum that modulate the immune response of dendritic cells using comparative genome hybridization. PLoS One 2010, 5, e10632. [Google Scholar] [CrossRef]
- Casey, P.G.; Gardiner, G.E.; Casey, G.; Bradshaw, B.; Lawlor, P.G.; Lynch, P.B.; Leonard, F.C.; Stanton, C.; Ross, R.P.; Fitzgerald, G.F.; et al. A five-strain probiotic combination reduces pathogen shedding and alleviates disease signs in pigs challenged with Salmonella enterica Serovar Typhimurium. Appl. Envrion. Microb. 2007, 73, 1858–1863. [Google Scholar] [CrossRef]
- Dabour, N.; Zihler, A.; Kheadr, E.; Lacroix, C.; Fliss, I. In vivo study on the effectiveness of pediocin PA-1 and Pediococcus acidilactici UL5 at inhibiting Listeria monocytogenes. Int. J. Food Microbiol. 2009, 133, 225–233. [Google Scholar] [CrossRef]
- Rea, M.C.; Dobson, A.; O’Sullivan, O.; Crispie, F.; Fouhy, F.; Cotter, P.D.; Shanahan, F.; Kiely, B.; Hill, C.; Ross, R.P. Effect of broad- and narrow-spectrum antimicrobials on Clostridium difficile and microbial diversity in a model of the distal colon. Proc. Natl. Acad. Sci. USA 2011, 108, 4639–4644. [Google Scholar] [CrossRef]
- McAuliffe, O.; Ryan, M.P.; Ross, R.P.; Hill, C.; Breeuwer, P.; Abee, T. Lacticin 3147, a broad-spectrum bacteriocin which selectively dissipates the membrane potential. Appl. Environ. Microbiol. 1998, 64, 439–445. [Google Scholar]
- Ganz, T.; Selsted, M.E.; Szklarek, D.; Harwig, S.S.; Daher, K.; Bainton, D.F.; Lehrer, R.I. Defensins. Natural peptide antibiotics of human neutrophils. J. Clin. Invest. 1985, 76, 1427–1435. [Google Scholar] [CrossRef]
- Selsted, M.E.; Harwig, S.S.; Ganz, T.; Schilling, J.W.; Lehrer, R.I. Primary structures of three human neutrophil defensins. J. Clin. Invest. 1985, 76, 1436–1439. [Google Scholar] [CrossRef]
- Ayabe, T.; Satchell, D.P.; Pesendorfer, P.; Tanabe, H.; Wilson, C.L.; Hagen, S.J.; Ouellette, A.J. Activation of Paneth cell alpha-defensins in mouse small intestine. J. Biol. Chem. 2002, 277, 5219–5228. [Google Scholar]
- Möndel, M.; Schroeder, B.O.; Zimmermann, K.; Huber, H.; Nuding, S.; Beisner, J.; Fellermann, K.; Stange, E.F.; Wehkamp, J. Probiotic E. coli treatment mediates antimicrobial human beta-defensin synthesis and fecal excretion in humans. Mucosal Immunol. 2009, 2, 166–172. [Google Scholar] [CrossRef]
- Schlee, M.; Wehkamp, J.; Altenhoefer, A.; Oelschlaeger, T.A.; Stange, E.F.; Fellermann, K. Induction of human beta-defensin 2 by the probiotic Escherichia coli Nissle 1917 is mediated through flagellin. Infect. Immun. 2007, 75, 2399–2407. [Google Scholar] [CrossRef]
- Schlee, M.; Harder, J.; Köten, B.; Stange, E.F.; Wehkamp, J.; Fellermann, K. Probiotic lactobacilli and VSL#3 induce enterocyte beta-defensin 2. Clin. Exp. Immunol. 2008, 151, 528–535. [Google Scholar]
- Hugo, A.A.; de Antoni, G.L.; Pérez, P.F. Lactobacillus delbrueckii subsp lactis (strain CIDCA 133) resists the antimicrobial activity triggered by molecules derived from enterocyte-like Caco-2 cells. Lett. Appl. Microbiol. 2010, 50, 335–340. [Google Scholar] [CrossRef]
- Yang, D.; Chertov, O.; Bykovskaia, S.N.; Chen, Q.; Buffo, M.J.; Shogan, J.; Anderson, M.; Schröder, J.M.; Wang, J.M.; Howard, O.M.; Oppenheim, J.J. Beta-defensins: Linking innate and adaptive immunity through dendritic and T cell CCR6. Science 1999, 286, 525–528. [Google Scholar] [CrossRef]
- Ismail, A.S.; Severson, K.M.; Vaishnava, S.; Behrendt, C.L.; Yu, X.; Benjamin, J.L.; Ruhn, K.A.; Hou, B.; DeFranco, A.L.; Yarovinsky, F.; et al. γδ intraepithelial lymphocytes are essential mediators of host-microbial homeostasis at the intestinal mucosal surface. Proc. Natl. Acad. Sci. USA 2011, 108, 8743–8748. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, C.; Zhou, Z.; Zhang, J.; Zhang, J.; Tian, Z. Small intestinal CD8+TCRγδ+ intraepithelial lymphocytes (iIELs) are involved in bacterial clearance during Salmonella typhimurium infection. Infect. Immun. 2012, 80, 565–574. [Google Scholar] [CrossRef]
- Roselli, M.; Finamore, A.; Nuccitelli, S.; Carnevali, P.; Brigidi, P.; Vitali, B.; Nobili, F.; Rami, R.; Garaguso, I.; Mengheri, E. Prevention of TNBS-induced colitis by different Lactobacillus and Bifidobacterium strains is associated with as expansion of γδT and regulatory T cells of intestinal intraepithelial lymphocytes. Inflamm. Bowel Dis. 2009, 15, 1526–1536. [Google Scholar] [CrossRef]
- Li, Y.; Innocentin, S.; Withers, D.R.; Roberts, N.A.; Gallagher, A.R.; Grigorieva, E.F.; Wilhelm, C.; Veldhoen, M. Exogenous stimuli maintain intraepithelial lymphocytes via Aryl hydrocarbon receptor activation. Cell 2011, 147, 629–640. [Google Scholar] [CrossRef]
- Schulz, V.J.; Smit, J.J.; Willemsen, K.J.; Fiechter, D.; Hassing, I.; Bleumink, R.; Boon, L.; van den Berg, M.; van Duursen, M.B.M.; Pieters, R.H.H. Activation of the aryl hydrocarbon receptor suppresses sensitization in a mouse peanut allergy model. Toxicol. Sci. 2011, 123, 491–500. [Google Scholar] [CrossRef]
- Van Baarlen, P.; Troost, F.J.; van Hemert, S.; van der Meer, C.; de Vos, W.M.; de Groot, P.J.; Hooiveld, G.J.E.J.; Brummer, R.M.; Kleerebezem, M. Differential NF-κB pathways induction by Lactobacillus plantarum in the duodenum of healthy humans correlating with immune tolerance. Proc. Natl. Acad. Sci. USA 2009, 106, 2371–2376. [Google Scholar] [CrossRef]
- Hershberg, R.M.; Cho, D.H.; Youakim, A.; Bradley, M.B.; Lee, J.S.; Framson, P.E.; Nepom, G.T. Highly polarized HLA class II antigen processing and presentation by human intestinal epithelial cells. J. Clin. Invest. 1998, 102, 792–803. [Google Scholar] [CrossRef]
- Snoeck, V.; Goddeeris, B.; Cox, E. The role of enterocytes in the intestinal barrier function and antigen uptake. Microbes Infect. 2005, 7, 997–1004. [Google Scholar] [CrossRef]
- Bloom, S.; Simmons, D.; Jewell, D.P. Adhesion molecules intercellular adhesion mole-1 (ICAM-1), ICAM-3 and B7 are not expressed by epithelium in normal or inflamed colon. Clin. Exp. Immunol. 1995, 101, 157–163. [Google Scholar] [CrossRef]
- Rimoldi, M.; Chieppa, M.; Salucci, V.; Avogadri, F.; Sonzogni, A.; Sampietro, G.M.; Nespoli, A.; Viale, G.; Allavena, P.; Rescigno, M. Intestinal immune homeostasis is regulated by the crosstalk between epithelial cells and dendritic cells. Nat. Immunol. 2005, 6, 507–514. [Google Scholar]
- Taylor, B.C.; Zaph, C.; Troy, A.E.; Du, Y.; Guild, K.J.; Comeau, M.R.; Artis, D. TSLP regulates intestinal immunity and inflammation in mouse models of helminth infection and colits. J. Exp. Med. 2009, 206, 655–667. [Google Scholar] [CrossRef]
- Niess, J.H.; Brand, S.; Gu, X.; Landsman, L.; Jung, S.; McCormick, B.A.; Vyas, J.M.; Boes, M.; Ploegh, H.L.; Fox, J.G.; et al. CX3CR1-mediated dendritic cell access to the intestinal lumen and bacterial clearance. Science 2005, 307, 254–258. [Google Scholar] [CrossRef]
- Chieppa, M.; Rescigno, M.; Huang, A.Y.C.; Germain, R.N. Dynamic imaging of dendritic cell extension into the small bowel lumen in response to epithelial cell TLR engagement. J. Exp. Med. 2006, 203, 2841–2852. [Google Scholar] [CrossRef]
- Rescigno, M.; Urbano, M.; Valzasina, B.; Francolini, M.; Rotta, G.; Bonasio, R.; Granucci, F.; Krasehenbuhl, J.P.; Ricciardi-Castagnoli, P. Dendritic cells express tight junction proteins and penetrate gut epithelial monolayers to sample bacteria. Nat. Immunol. 2001, 2, 361–367. [Google Scholar] [CrossRef]
- Jang, M.H.; Kweon, M.; Iwatani, K.; Yamamoto, M.; Terahara, K.; Sasakawa, C.; Suzuki, T.; Nochi, T.; Yokota, Y.; Rennert, P.D.; et al. Intestinal villous M cells: An antigen entry site in the mucosal epithelium. Proc. Natl. Acad. Sci. USA 2004, 101, 6110–6115. [Google Scholar] [CrossRef]
- Neutra, M.R.; Mantis, N.J.; Frey, A.; Giannasca, P.J. The composition and function of M cell apical membranes: Implications for microbial pathogenesis. Semin. Immunol. 1999, 11, 171–181. [Google Scholar] [CrossRef]
- Abreu, M. Toll-like receptor signalling in the intestinal epithelium: How bacterial recognition shapes intestinal function. Nat. Rev. Immunol. 2010, 10, 131–144. [Google Scholar] [CrossRef]
- Gewirtz, A.T.; Navas, T.A.; Lyons, S.; Godowski, P.J.; Madara, J.L. Cutting edge: Bacterial flagellin activates basolaterally expressed TLR5 to induce epithelial proinflammatory gene expression. J. Immunol. 2001, 164, 1882–1885. [Google Scholar]
- Rhee, S.H.; Im, E.; Riegler, M.; Kokkotou, E.; O’Brien, M.; Pothoulakis, C. Pathophysiological role of Toll-like receptor 5 engagment by bacterial flagellin in colonic inflammation. Proc. Natl. Acad. Sci. USA 2005, 102, 13610–13615. [Google Scholar] [CrossRef]
- Eckmann, L.; Kagnoff, M.F.; Fierer, J. Epithelial cells secrete the chemokine interleukin-8 in response to bacterial entry. Infect. Immun. 1993, 61, 4569–4574. [Google Scholar]
- Otte, J.M.; Podolsky, D.K. Functional modulation of enterocytes by gram-positive and gram-negative microorganisms. Am. J. Physiol. Gastrointest. Liver Physiol. 2004, 286, G613–G626. [Google Scholar] [CrossRef]
- Barbier de La Serre, C.; Ellis, C.L.; Lee, J.; Hartman, A.L.; Rutledge, J.C.; Raybould, H.E. Propensity to high-fat diet-induced obesity in rats is associated with changes in the gut microbiota and gut inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 299, G440–G448. [Google Scholar] [CrossRef]
- Koyama, I.; Matsunaga, T.; Harada, T.; Hokari, S.; Komoda, T. Alkaline phosphatases reduce toxicity of lipopolysaccharides in vivo and in vitro through dephosphorylation. Clin. Biochem. 2002, 35, 455–461. [Google Scholar] [CrossRef]
- Bates, J.M.; Akerlund, J.; Mittge, E.; Guillemin, K. Intestional alkaline phosphatase detoxifies lipopolysaccharide and prevents inflammation in response to the gut microbiota. Cell Host Microbe 2007, 2, 371–382. [Google Scholar] [CrossRef]
- Lee, J.; Mo, J.H.; Katakura, K.; Alkalay, I.; Rucker, A.N.; Liu, Y.T.; Lee, H.K.; Shen, C.; Cojocaru, G.; Shenouda, S.; et al. Maintenance of colonic homeostasis by distinctive apical TLR9 signalling in intestinal epithelial cells. Nat. Cell Biol. 2006, 8, 1327–1336. [Google Scholar] [CrossRef]
- Shibolet, O.; Podolsky, D.K. TLRs in the Gut. IV. Negative regulation of Toll-like receptors and intestinal homeostasis: Addition by subtraction. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 292, G1469–G1473. [Google Scholar] [CrossRef]
- Rudensky, A.Y. Regulatory T-cells and Foxp3. Immunol. Rev. 2011, 241, 260–268. [Google Scholar] [CrossRef]
- Josefowicz, S.Z.; Lu, L.F.; Rudensky, A.Y. Regulatory T-cells: Mechanisms of differentiation and function. Annu. Rev. Immunol. 2012, 30, 531–364. [Google Scholar] [CrossRef]
- Lavasani, S.; Dzhambazov, B.; Nouri, M.; Fak, F.; Buske, S.; Molin, G.; Thorlacius, H.; Alenfall, J.; Jeppsson, B.; Westrom, B. A novel probiotic mixture exerts a therapeutic effect on experimental autoimmune encephalomyelitis mediated by IL-10 producing regulatory T-cells. PLoS One 2010, 5, e9009. [Google Scholar] [CrossRef]
- Haller, D.; Serrant, P.; Granato, D.; Schiffrin, E.J.; Blum, S. Activation of human NK cells by Staphylocci and Lactobacilli requires cell contact-dependent costimulation by autologous monocytes. Clin. Diagn. Lab. Immunol. 2002, 9, 649–657. [Google Scholar]
- Smits, H.H.; Engering, A.; ven der Kleij, D.; de Jong, E.C.; Schipper, K.; van Capel, T.M.; Zaat, B.A.; Yazdanbakhsh, M.; Wierenga, E.A.; van Kooyk, Y.; et al. Selective probiotic bacteria induce IL-10-producing regulatory T-cells in vitro by modulating dendritic cell function through dendritic cell-specific intercellular adhesion molecule 3-grabbing nonintegrin. J. Allergy Clin. Immunol. 2005, 115, 1260–1267. [Google Scholar] [CrossRef]
- Jeon, S.G.; Kayama, H.; Ueda, Y.; Takahashi, T.; Asahara, T.; Tsuji, H.; Tsuji, N.M.; Kiyono, H.; Ma, J.S.; Kusu, T.; et al. Probiotic Bifidobacterium breve induces IL-10 producing Tr1 cells in the colon. PLoS One 2012, 8, e1002714. [Google Scholar]
- Peterson, E.R.; Claesson, H.; Schmidt, E.G.; Jensen, S.S.; Ravn, P.; Olsen, J.; Ouwehand, A.; Kristensen, N. Consumption of probiotics increases the effect of regulatory T-cells in transfer colitis. Inflamm. Bowel Dis. 2012, 18, 1–12. [Google Scholar] [CrossRef]
- Re, F.; Strominger, J.L. IL-10 released by concomitant TLR-2 stimulation blocks the induction of a subset of Th1 cytokines that are specifically induced by TLR-4 or TLR-3 in human dendritic cells. J. Immunol. 2004, 173, 7548–7555. [Google Scholar]
- Coombes, J.L.; Siddiqui, K.R.; Arancibia-Carcamo, C.V.; Hall, J.; Sun, C.M.; Belkaid, Y.; Powrie, F. A functionally specialised population of mucosal CD103+ DCs induces Foxp3+ regulatory T-cells via a TGF-β and retinoic acid-dependent manner. J. Exp. Med. 2007, 204, 1757–1764. [Google Scholar] [CrossRef]
- Evrard, B.; Coudeyras, S.; Dosgilbert, A.; Charbonnel, N.; Alame, J.; Tridon, A.; Forestier, C. Dose-dependent immunomodulation of human dendritic cells by the probiotic Lactobacillus rhamnosus Lcr35. PLoS One 2011, 6, e18735. [Google Scholar]
- Miettinen, M.; Matikainen, S.; Vuopio-Varkila, J.; Pirhonen, J.; Varkila, K.; Kurimoto, M.; Julkunen, I. Lactobacilli and streptococci induce interleukin-12 (IL-12), IL-18 and gamma interferon production in human peripheral blood mononuclear cells. Infect. Immun. 1998, 66, 6058–6062. [Google Scholar]
- Pochard, P.; Gossett, P.; Grangette, C.; Andre, C.; Tonnel, A.; Pestel, J.; Mercenier, A. Lactic acid bacteria inhibit Th2 cytokine production by mononuclear cells from allergic patients. Basic Clin. Immunol. 2002, 110, 617–623. [Google Scholar]
- Niers, L.E.; Timmerman, H.M.; Rijkers, G.T.; van Bleek, G.M.; van Uden, N.O.; Knol, E.F.; Kapsenberg, M.L.; Kimpen, J.L.; Hoekstra, M.O. Identification of strong interleukin-10 inducing lactic acid bacteria which down-regulate T helper type 2 cytokines. Clin. Exp. Allergy 2005, 35, 1481–1489. [Google Scholar] [CrossRef]
- Shida, K.; Suzuki, T.; Kiyoshima-Shibata, J.; Shimada, S.I.; Nanno, M. Essential roles of monocytes in stimulating human peripheral blood mononuclear cells with Lactobacillus casei to produce cytokines and augment natural killer cell activity. Clin. Vaccine Immunol. 2006, 13, 997–1003. [Google Scholar] [CrossRef]
- Takeda, K.; Suzuki, T.; Shimada, S.I.; Shida, K.; Nanno, M.; Okumura, K. Interleukin-12 is involved in the enhancement of human natural killer cell activity by Lactobacillus casei Shirota. Clin. Exp. Immunol. 2006, 146, 109–115. [Google Scholar] [CrossRef]
- Castellazi, A.M.; Valsecchi, C.; Montagna, L.; Malfa, P.; Ciprandi, G.; Avanzini, M.A.; Marseglia, G.L. In vitro activation of mononuclear cells by two probiotics: Lactobacillus paracasei I 1688, Lactobacillus salivarus I 1794, and their mixture (PSMIX). Immunol. Invest. 2007, 36, 413–421. [Google Scholar] [CrossRef]
- Medina, M.; Izquierdo, E.; Ennahar, S.; Sanz, Y. Differential immunomodulatory properties of Bifidobacterium longum strains: Relevance to probiotics selection and clinical applications. Clin. Exp. Immunol. 2007, 150, 531–538. [Google Scholar] [CrossRef]
- Ghadimi, D.; Folster-Holst, R.; de Vrese, M.; Winkler, P.; Heller, K.J.; Schrezenmeir, J. Effects of probiotic bacteria and their genomic DNA on TH1/TH2-cytokine production by peripheral blood mononuclear cells (PBMCs) of healthy and allergic subjects. Immunobiology 2008, 213, 677–692. [Google Scholar] [CrossRef]
- Dong, H.; Rowland, I.; Tuohy, K.M.; Thomas, L.V.; Yaqoob, P. Selective effects of Lactobacillus casei Shirota on T-cell activation, natural killer cell activity and cytokine production. Clin. Exp. Immunol. 2010, 161, 378–388. [Google Scholar]
- Vissers, Y.M.; Snel, J.; Zuurendonk, P.F.; Smit, B.A.; Wichers, H.J.; Savelkoul, H.F. Differential effects of Lactobacillus acidophilus and Lactobacillus plantarum strains on cytokine induction in human peripheral blood mononuclear cells. FEMS Immunol. Med. Microbiol. 2010, 59, 60–70. [Google Scholar] [CrossRef]
- O’Mahony, L.; O’Callaghan, L.; McCarthy, J.; Shilling, D.; Scully, P.; Sibartie, S.; Kavanagh, E.; Kirwan, W.O.; Redmond, H.P.; Collins, J.K.; et al. Differential cytokine response from dendritic cells to commensal and pathogenic bacteria in different lymphoid compartments in humans. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 290, 839–845. [Google Scholar] [CrossRef]
- Fink, L.N.; Zeuthen, L.H.; Christensen, H.R.; Morandi, B.; Frokiaer, H.; Ferlazzo, G. Distinct gut-derived lactic acid bacteria elicit divergent dendritic cell-mediated NK cell responses. Int. Immunol. 2007, 19, 1319–1327. [Google Scholar] [CrossRef]
- Mohamadzadeh, M.; Olson, S.; Kalina, W.V.; Ruthel, G.; Demmin, G.L.; Warfield, K.L.; Bavari, S.; Klaenhammer, T.R. Lactobacilli activate human dendritic cells that skew T cells toward T helper 1 polarisation. Proc. Natl. Acad. Sci. USA 2005, 102, 288–2885. [Google Scholar]
- Ghadimi, D.; Helwig, U.; Schrezenmeir, J.; Heller, K.J.; de Vrese, M. Epigenetic imprinting by commensal probiotics inhibits the IL-23/IL-17 axis in an in vitro model of the intestinal mucosal immune system. J. Leukoc. Biol. 2012, 92, 895–911. [Google Scholar] [CrossRef]
- Paolillo, R.; Carratelli, C.R.; Sorrentino, S.; Mazzola, N.; Rizzo, A. Immunomodulatory effects of Lactobacillus plantarum on human colon cancer cells. Int. Immunopharmacol. 2009, 9, 1265–1271. [Google Scholar] [CrossRef]
- Donkor, O.N.; Ravikumar, M.; Proudfoot, O.; Day, S.L.; Apostolopoulos, V.; Paukovics, G.; Vasiljevic, T.; Nutt, S.L.; Gill, H. Cytokine profile and induction of T helper type 17 and regulatory T cells by human peripheral mononuclear cells after microbial exposure. Clin. Exp. Immunol. 2012, 167, 282–295. [Google Scholar] [CrossRef]
- Haller, D.; Bode, C.; Hammes, W.P.; Pfeifer, A.M.; Schiffrin, E.J.; Blum, S. Non-pathogenic bacteria elicit a differential cytokine response by intestinal epithelial cell/leucocyte co-cultures. Gut 2000, 47, 79–87. [Google Scholar] [CrossRef]
- Foey, A.D. Mucosal Macrophages: Phenotypr and Functionality in Homeostasis and Pathology. In Handbook of Macrophages: Life Cycle, Functions and Diseases; Takahashi, R., Kai, H., Eds.; Nova Science Publishers Inc.: New York, NY, USA, 2012; Chapter 4; pp. 121–146. [Google Scholar]
- Habil, N.; Al-Murrani, W.; Beal, J.; Foey, A.D. Probiotic bacterial strains differentially modulate macrophage cytokine production in a strain-dependent and cell subset-specific manner. Benef. Microbes 2011, 2, 283–293. [Google Scholar] [CrossRef]
- Habil, N.; Beal, J.; Foey, A. Lactobacillus casei strain Shirota selectively modulates macrophage subset cytokine production. Int. J. Probiotics Prebiotics 2012, 7, 1–12. [Google Scholar]
- Foey, A.D. Butyrate regulation of distinct macrophage subsets: Opposing effects on M1 and M2 macrophages. Int. J. Probiotics Prebiotics 2011, 6, 147–158. [Google Scholar]
- Helwig, U.; Lammers, K.M.; Rizzello, F.; Brigidi, P.; Rohleder, V.; Caramelli, E.; Gionchetti, P.; Schrezenmeir, J.; Foelsch, U.R.; Schreiber, S.; et al. Lactobacilli, bifidobacteria and E. coli Nissle induce pro- and anti-inflammatory cytokines in peripheral blood mononuclear cells. World J. Gastroenterol. 2006, 12, 5978–5986. [Google Scholar]
- Perez-Cano, F.J.; Dong, H.; Yaqoob, P. In vitro immunomodulatory activity of Lactobacillus fermentum CECT5716 and Lactobacillus salivarius CECT5713: Two probiotic strains isolated from human breast milk. Immunobiology 2010, 215, 996–1004. [Google Scholar] [CrossRef]
- Thomas, C.M.; Hong, T.; van Pijkeren, J.P.; Hemarajata, P.; Trinh, D.V.; Hu, W.; Britton, R.A.; Kalkum, M.; Versalovic, J. Histamine derived from probiotic Lactobacillus reuteri suppresses TNF via modulation of PKA and ERK signalling. PLoS One 2012, 7, e31951. [Google Scholar] [CrossRef]
- Ruiz, P.A.; Hoffmann, M.; Szcesny, S.; Blaut, M.; Haller, D. Innate mechanisms for Bifidobacterium lactis to activate transient pro-inflammatory host responses in intestinal epithelial cells after the colonisation of germ-free rats. Immunology 2005, 115, 441–450. [Google Scholar] [CrossRef]
- Vinderola, G.; Matar, C.; Perdigon, G. Role of intestinal epitelial cells in immune effects mediated by gram-positive probiotic bactéria: Involvement of toll-like receptors. Clin. Diagn. Lab. Immunol. 2005, 12, 1075–1084. [Google Scholar]
- Hart, A.L.; Lammers, K.; Brigidi, P.; Vitali, B.; Rizzello, F.; Gionchetti, P.; Campieri, M.; Kamm, M.A.; Knight, S.C.; Stagg, A.J. Modulation of human dendritic cell phenotype and function by probiotic bacteria. Gut 2004, 53, 1602–1609. [Google Scholar] [CrossRef]
- Braat, H.; van den Brande, J.; van Tol, E.; Hommes, D.; Peppelenbosch, M.; van Deventer, S. Lactobacillus rhamnosus induces peripheral hyporesponsiveness in stimulated CD4+ Tcells via modulation of dendritic cell function. Am. J. Nutr. 2004, 80, 1618–1625. [Google Scholar]
- Latvala, S.; Pietila, T.E.; Veckman, V.; Kekkonen, R.A.; Tynkknen, S.; Korpela, R.; Julkunen, I. Potentially probiotic bacteria induce efficient maturation but differential cytokine production in human monocyte-derived dendritic cells. World J. Gastroenterol. 2008, 14, 5570–5583. [Google Scholar] [CrossRef]
- Koniecza, P.; Groeger, D.; Ziegler, M.; Frei, R.; Ferstl, R.; Shanahan, F.; Quigley, E.M.; Kiely, B.; Akdis, C.A.; O’Mahony, L. Bifidobacterium infantis 35624 administration induces Foxp3 T regulatory cells in human peripheral blood: Potential role for myeloid and plasmacytoid dendritic cells. Gut 2012, 61, 354–366. [Google Scholar] [CrossRef]
- Strober, W.; Zhang, F.; Kitani, A.; Fuss, I.; Fichtner-Feigl, S. Proinflammatory cytokines underlying the inflammation of Crohn’s disease. Curr. Opin. Gastroenterol. 2010, 26, 310–307. [Google Scholar] [CrossRef]
- Chen, Z.; O’Shea, J.J. Regulation of IL-17 production in human lymphocytes. Cytokine 2008, 41, 71–78. [Google Scholar] [CrossRef]
- Cooke, A. Th17 cells in inflammatory conditions. Rev. Diabet. Stud. 2006, 3, 72–75. [Google Scholar] [CrossRef]
- Dicksved, J.; Halfvarson, J.; Rosenquist, M.; Jarnernot, G.; Tysk, C.; Apajalahti, J.; Engstrand, L.; Jansson, J.K. Molecular analysis of the gut microbiota of identical twins with Crohn’s disease. ISME 2008, 2, 716–727. [Google Scholar] [CrossRef]
- Erickson, A.R.; Cantarel, B.L.; Lamendella, R.; Darzi, Y.; Mongodin, E.F.; Pan, C.; Shah, M.; Halfvarson, J.; Tysk, C.; Henrissat, B.; et al. Integrated metagenomics/metaproteomics reveals human host-micrbiota signatures of Crohns’ disease. PLoS One 2012, 7, e49138. [Google Scholar] [CrossRef]
- Maeda, S.; Hsu, L.C.; Liu, H.; Bankston, L.A.; Limura, M.; Kagnoff, M.F.; Eckmann, L.; Karin, M. Nod2 mutation in Crohn’s disease potentiates NF-κB activity and IL-1β processing. Science 2006, 307, 734–738. [Google Scholar]
- Hong, J.; Leung, E.; Fraser, A.G.; Merriman, T.R.; Vishnu, P.; Krissansen, G.W. TLR2, TLR4 and TLR9 polymorphisms and Crohn’s disease in a New Zealand Caucasian cohort. J. Gastroenterol. Hepatol. 2006, 22, 1760–1766. [Google Scholar]
- Hugot, J. CARD15/NOD2 mutations in Crohn’s disease. Ann. N. Y. Acad. Sci. 2006, 1072, 9–18. [Google Scholar]
- Netea, M.G.; Kullberg, B.J.; de Jong, D.J.; Franke, B.; Sprong, T.; Naber, T.H.J.; Drenth, J.P.H.; van der Meer, J.W. NOD2 mediates anti-inflammatory signals induced by TLR2 ligands: Implications for Crohn’s disease. Eur. J. Immunol. 2004, 34, 2052–2059. [Google Scholar] [CrossRef]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermudez-Humaran, L.G.; Gratadoux, J.J.; Blugeon, S.; Bridonneau, C.; Furet, J.P.; Corthier, G.; et al. Feacalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef]
- Jia, W.; Whitehead, R.N.; Griffiths, L.; Dawson, C.; Waring, R.H.; Ramsden, D.B.; Hunter, J.O.; Cole, J.A. Is the abundance of Faecalibacterium prausnitzii relevant to Crohn’s disease? FEMS Microbiol. Lett. 2010, 310, 138–144. [Google Scholar] [CrossRef]
- Van Immerseel, F.; Ducatelle, R.; de Vos, M.; Boon, N.; van de Wiele, T.; Verbeke, K.; Rutgeerts, P.; Sas, B.; Louis, P.; Flint, H.J. Butyric acid-producing anaerobic bacteria as a novel probiotic treatment approach for inflammatory bowel disease. J. Med. Microbiol. 2010, 59, 141–143. [Google Scholar] [CrossRef]
- Duncan, S.H.; Hold, G.L.; Harmsen, H.J.M.; Stewart, C.S.; Flint, H.J. Growth requirements and fermentation products of Fusobacterium prausnitzii, and a proposal to reclassify it as Faecalibacterium prausnitzii gen. nov., comb. nov. Int. J. Syst. Evol. Microbiol. 2002, 52, 2141–2146. [Google Scholar] [CrossRef]
- Pryde, S.E.; Duncan, S.H.; Hold, G.L.; Stewart, C.S.; Flint, H.J. The microbiology of butyrate formation in the human colon. FEMS Micrbiol. Lett. 2002, 217, 133–139. [Google Scholar] [CrossRef]
- Tuohy, K.M.; Rouzaud, G.C.M.; Bruck, W.M.; Gibson, G.R. Modulation of the human gut microflora towards improved health using prebiotics—assessment of efficacy. Curr. Pharm. Des. 2005, 11, 75–90. [Google Scholar] [CrossRef]
- Ramirez-Farias, C.; Slezak, K.; Fuller, Z.; Duncan, A.; Holtrop, G.; Louis, P. Effect of inulin on the human gut microbiota: Stimulation of Bifidobacterium adolescentis and Faecalibacterium prausnitzii. Br. J. Nutr. 2009, 101, 541–550. [Google Scholar]
- Benjamin, J.L.; Hedin, C.R.H.; Koutsoumpas, A.; Ng, S.C.; McCarthy, N.E.; Hart, A.L.; Kamm, M.A.; Sanderson, J.D.; Knight, S.C.; Forbes, A.; et al. Randomised, double-blind, placebo-controlled trail of fructo-oligosaccharides in active Crohn’s disease. Gut 2011, 60, 923–929. [Google Scholar] [CrossRef]
- Schultz, M.; Veltkamp, C.; Dieleman, L.A.; Grenther, W.B.; Wyrick, P.B.; Tonkonogy, S.L.; Sartor, R.B. Lactobacillus plantarum 299v in the treatment and prevention of spontaneous colitis in interleukin-10-deficient mice. Inflamm. Bowel Dis. 2002, 8, 71–80. [Google Scholar] [CrossRef]
- Dieleman, L.A.; Goerres, M.S.; Arends, A.; Sprengers, D.; Torrice, C.; Hoentjen, F.; Grenther, W.B.; Sartor, R.B. Lactobacillus GG prevents recurrence of colitis in HLA-B27 transgenic rats after antibiotic treatment. Gut 2003, 52, 370–376. [Google Scholar] [CrossRef]
- Furrie, E.; Macfarlane, S.; Kennedy, A.; Cummings, J.H.; Walsh, S.V.; O’Neil, D.A.; Macfarlane, G.T. Synbiotic therapy (Bifidobacterium longum/Synergy1) initiates resolution of inflammation in patients with active ulcerative colitis: A randomised controlled pilot trial. Gut 2005, 54, 242–249. [Google Scholar] [CrossRef]
- Fujimori, S.; Gudis, K.; Mitsui, K.; Seo, T.; Yonezawa, M.; Tanaka, S.; Tatsuguchi, A.; Sakamoto, C. A randomized controlled trial on the efficacy of synbiotic versus probiotic and prebiotic treatment to improve the quality of life in patients with ulcerative colitis. Nutrition 2009, 25, 520–525. [Google Scholar] [CrossRef]
- Mennigen, R.; Nolte, K.; Rijcken, E.; Utech, M.; Loeffler, B.; Senninger, N.; Bruewer, M. Probiotic mixture VSL#3 protects the epithelial barrier by maintaining tight junction protein expression and preventing apoptosis in a murine model of colitis. Am. J. Physiol. Gastrointest. Liver Physiol. 2009, 296, 1140–1149. [Google Scholar] [CrossRef]
- Miele, E.; Pascarella, F.; Giannetti, E.; Quaglietta, L.; Baldassano, R.N.; Staiano, A. Effect of a probiotic preparation (VSL#3) on induction and maintenance of remission in children with ulcerative colitis. Am. J. Gastroenterol. 2009, 104, 437–443. [Google Scholar] [CrossRef]
- Uronis, J.M.; Arthur, J.C.; Keku, T.; Fodor, A.; Carroll, I.M.; Cruz, M.L.; Appleyard, C.B.; Jobin, C. Gut microbial diversity is reduced by the probiotic VSL#3 and correlates with decreased TNBS-induced colitis. Inflamm. Bowel Dis. 2011, 17, 289–297. [Google Scholar] [CrossRef]
- Rowland, I.R.; Rumney, C.J.; Coutts, J.T.; Lievense, L.C. Effect of Bifidobacterium longum and inulin on gut bacterial metabolism and carcinogen-induced aberrant crypt foci in rats. Carcinogenesis 1998, 19, 281–285. [Google Scholar] [CrossRef]
- Femia, A.P.; Luceri, C.; Dolara, P.; Giannini, A.; Biggeri, A.; Salvadori, M.; Clune, Y.; Collins, K.J.; Paglierani, M.; Caderni, G. Antitumorigenic activity of the prebiotic inulin enriched with oligofructose in combination with the prebiotics Lactobacillus rhamnosus and Bifidobacterium lactis in azoxymethane-induced colon carcinogenesis in rats. Carcinogenesis 2002, 23, 1953–1960. [Google Scholar] [CrossRef]
- Moreno de LeBlanc, A.; Matar, C.; Perdigon, G. The application of probiotics in cancer. Br. J. Nutr. 2007, 98, 105–110. [Google Scholar]
- Rafter, J.; Bennett, M.; Caderni, G.; Clune, Y.; Hughes, R.; Karlsson, P.C.; Klinder, A.; O’Riordan, M.; O’Sullivan, G.C.; Pool-Zobel, B.; et al. Dietary synbiotics reduce cancer risk factors in polypectomised and colon cancer patients. Am. J. Clin. Nutr. 2007, 85, 488–496. [Google Scholar]
- Bassaganya-Riera, J.; Viladomiu, M.; Pedragosa, M.; de Simone, C.; Carbo, A.; Shaykhutdinov, R.; Jobin, C.; Arthur, J.C.; Corl, B.A.; Vogel, H.; et al. Probiotic bacteria produce conjugated linoleic acid locally in the gut that targets macrophage PPAR-gamma to suppress colitis. PLoS One 2012, 7, e31238. [Google Scholar]
- Chen, C.C.; Lin, W.C.; Kong, M.S.; Shi, H.N.; Walker, W.A.; Lin, C.Y.; Huang, C.T.; Lin, Y.C.; Jung, S.M.; Lin, T.Y. Oral inoculation of probiotics Lactobacillus acidophilus NCFM suppresses tumour growth both in segmental orthotopic colon cancer and extra-intestinal tissue. Br. J. Nutr. 2012, 107, 1623–1634. [Google Scholar] [CrossRef]
- Repa, A.; Grangette, C.; Daniel, C.; Hochreiter, R.; Hoffmann-Sommergruber, K.; Thlhamer, J.; Kraft, D.; Breiteneder, H.; Mercenier, A.; Wiedermann, U. Mucosal co-application of lactic acid bacteria and allergen induces counter-regulatory immune responses in a murine model of birch pollen allergy. Vaccine 2003, 22, 87–95. [Google Scholar] [CrossRef]
- Van de Pol, M.A.; Lutter, R.; Smids, B.S.; Weersink, E.J.M.; van der Zee, J.S. Synbiotics reduce allergen-induced T-helper 2 response and improve peak expiratory flow in allergic asthmatics. Allergy 2010, 66, 39–47. [Google Scholar]
- Van der Aa, L.B.; Lutter, R.; Heymans, H.S.; Smids, B.S.; Dekker, T.; van Aalderen, W.M.; Sillevis Smitt, J.H.; Knippels, L.M.J.; Garssen, J.; Nauta, A.J.; et al. No detectable beneficial systemic immunomodulatory effects of a specific synbiotic mixture in infants with atopic dermatitis. Clin. Exp. Allergy 2012, 42, 531–539. [Google Scholar] [CrossRef]
- Van der Aa, L.B.; Heymans, H.S.; van Aalderen, W.M.; Sillevis Smitt, J.H.; Knol, J.; Ben Amor, K.; Goossens, D.A.; Sprikkelman, A.B.; Synbad Study Group. Effect of a new synbiotic mixture on atopic dermatitis in infants: A randomised-controlled trial. Clin. Exp. Allergy 2010, 40, 795–804. [Google Scholar]
- D’Arienzo, R.; Maurano, F.; Luongo, D.; Mazzarella, G.; Stefanile, R.; Troncone, R.; Auricchio, S.; Ricca, E.; David, C.; Rossi, M. Adjuvant effect of Lactobacillus casei in a mouse model of gluten sensitivity. Immunol. Lett. 2008, 119, 78–83. [Google Scholar] [CrossRef]
- Lindfors, K.; Blomqvist, T.; Juuti-Uusitalo, K.; Stenman, S.; Venalainen, J.; Maki, M.; Kaukinen, K. Live probiotic Bifidobacterium lactis bacteria inhibit the toxic effects induced by wheat gliadin in epithelial cell culture. Clin. Exp. Immunol. 2008, 152, 552–558. [Google Scholar] [CrossRef]
- De Palma, G.; Cinova, J.; Stepankova, R.; Tuckova, L.; Sanz, Y. Pivotal advance: Bifidobacteria and Gram-negative bacteria differentially influence immune responses in the proinflammatory milieu of celiac disease. J. Leukoc. Biol. 2010, 87, 765–778. [Google Scholar] [CrossRef]
- D’Arienzo, R.; Maurano, F.; Lavermicocca, P.; Ricca, E.; Rossi, M. Modulation of the immune response by probiotic strains in a mouse model of gluten sensitivity. Cytokine 2009, 48, 254–259. [Google Scholar] [CrossRef]
- Kullberg, M.C.; Jankovic, D.; Feng, C.G.; Hue, S.; Gorelick, P.L.; McKenzie, B.S.; Cua, D.J.; Powrie, F.; Cheever, A.W. IL-23 plays a key role in Helicobacter hepaticus-induced T cell-dependent colitis. J. Exp. Med. 2006, 203, 2485–2494. [Google Scholar] [CrossRef]
- Brand, S. Crohn’s disease: Th1, Th17 or both? The change of a paradigm: New immunological and genetic insights implicate Th17 cells in the pathogenesis of Crohn’s disease. Gut 2009, 58, 1152–1167. [Google Scholar]
- Fujimori, S.; Tatsuguchi, A.; Gudis, K.; Kishida, T.; Mitsui, K.; Ehara, A.; Kobayashi, T.; Sekita, Y.; Seo, T.; Sakamoto, C. High dose probiotic and prebiotic cotherapy for the remission induction of active Crohn’s disease. J. Gastroenterol. Hepatol. 2006, 22, 1199–1204. [Google Scholar]
- Popivanova, B.K.; Kitamura, K.; Wu, Y.; Kondo, T.; Kagaya, T.; Kaneko, S.; Oshima, M.; Fuji, C.; Mukaida, N. Blocking TNF-α in mice reduces colorectal carcinogenesis associated with chronic colitis. J. Clin. Invest. 2008, 118, 560–570. [Google Scholar]
- Gophna, U.; Sommerfeld, K.; Gophna, S.; Doolittle, W.F.; Veldhuyzen van Zanten, S.J. Differences between tissue-associated intestinal microfloras of patients with Crohn’s disease and Ulcerative colitis. J. Clin. Microbiol. 2006, 44, 4136–4141. [Google Scholar] [CrossRef]
- Sood, A.; Midha, V.; Makharia, G.K.; Ahuja, V.; Singal, D.; Goswami, P.; Tandon, R.K. The probiotic preparation, VSL#3, induces remission in patients with mild-to-moderately active ulcerative colitis. Clin. Gastroenterol. Heptol. 2009, 7, 1202–1209. [Google Scholar] [CrossRef]
- Henriksen, M.; Jahnsen, J.; Lygren, I.; Stray, N.; Sauar, J.; Vatn, M.H.; Moum, B.; IBSEN Study Group. C-reactive protein: A predictive factor and marker of inflammation in inflammatory bowel disease. Results from a prospective population-based study. Gut 2008, 57, 1518–1523. [Google Scholar] [CrossRef]
- Rizello, V.; Bonaccorsi, I.; Dongarra, M.L.; Fink, L.N.; Ferlazzo, G. Role of natural killer and dendritic cell crosstalk in immunomodulation by commensal bacteria probiotics. J. Biomed. Biotechnol. 2011, 2011, 1–10. [Google Scholar]
- Szaradkiewicz, A.; Marciniak, R.; Chudzicka-Strugala, I.; Wasilewska, A.; Drews, M.; Majewski, P.; Karpinski, T.; Zwozdziak, B. Proinflammatory cytokines and IL-10 in inflammatory bowel disease and colorectal cancer patients. Arch. Immunol. Ther. Exp. 2009, 57, 291–294. [Google Scholar] [CrossRef]
- Kane, K.F.; Langman, M.J.; Williams, G.R. Antiproliferative responses of two human colon cancer cell lines to Vitamin D3 are differentially modified by 9-cis-retionic acid. Cancer Res. 1996, 56, 623–632. [Google Scholar]
- Muscat, J.; Wynder, E.L. The consumption of well-done red meat and the risk of colorectal cancer. Am. J. Public Health 1994, 84, 856–858. [Google Scholar] [CrossRef]
- Le Marchand, L.; Hankin, J.H.; Wilkens, L.R.; Pierce, L.M.; Franke, A.; Kolonel, L.N.; Seifried, A.; Custer, L.J.; Chang, W.; Lum-Jones, A.; et al. Combined effects of well-done red meat, smoking and rapid n-acetyltransferase 2 and CYP1A2 phenotypes in increasing colorectal cancer risk. Cancer Epidemiol. Biomarkers Prev. 2001, 10, 1259–1266. [Google Scholar]
- Herbst, T.; Sichelstiel, A.; Schär, C.; Yadava, K.; Bürki, K.; Cahenzli, J.; McCoy, K.; Marsland, B.J.; Harris, N.L. Dysregulation of allergic airway inflammation in the absence of microbial colonization. Am. J. Respir. Crit. Care Med. 201 2011, 184, 198–205. [Google Scholar]
- Abrahamsson, T.R.; Jakobsson, H.E.; Andersson, A.F.; Björkstén, B.; Engstrand, L.; Jenmalm, M.C. Low diversity of the gut microbiota in infants with atopic eczema. J. Allery Clin. Immun. 2012, 129, 434–440. [Google Scholar] [CrossRef]
- Nylund, L.; Satokari, R.; Nikkilä, J.; Rajilić-Stojanović, M.; Kalliomäki, M.; Isolauri, E.; Salminen, S.; de Vos, W.M. Microarray analysis reveals marked intestinal microbiota aberrancy in infants having eczema compared to healthy children in at-risk for atopic disease. BMC Microbiol. 2013, 13, 12. [Google Scholar] [CrossRef]
- Lee, H.; Kim, H.; Lee, E.; Jang, M.; Kim, S.; Park, J.; Seoh, J.; Jung, Y. Characterisation of CCR9 expression and thymus-expressed chemokine responsiveness of the murine thymus, spleen and mesenteric lymph nodes. Immunobiology 2012, 217, 402–411. [Google Scholar] [CrossRef]
- Kalliomäki, M.; Salminen, S.; Arvilommi, H.; Kero, P.; Koskinen, P.; Isolauri, E. Probiotics in primary prevention of atopic disease: A randomised placebo-controlled trial. Lancet 2001, 357, 1076–1079. [Google Scholar] [CrossRef]
- Böttcher, M.F.; Nordin, E.K.; Sandin, A.; Midtvedt, T.; Björkstén, B. Microflora-associated characteristics in faeces from allergic and nonallergic infants. Clin. Exp. Allergy 2000, 30, 1590–1596. [Google Scholar]
- Wakabayashi, H.; Nariai, C.; Takemura, G.; Nakao, W.; Fujiwara, D. Dietary supplementation with lactic acid bacteria attenuates the development of atopic-dermatitis-like skin lesions in NC/Nga mice in a strain-dependent manner. Int. Arch. Allergy Immunol. 2008, 145, 141–151. [Google Scholar] [CrossRef]
- Tanaka, A.; Jung, K.; Benyacoub, J.; Prioult, G.; Okamoto, N.; Ohmori, K.; Blum, S.; Mercenier, A.; Mastuda, H. Oral supplementation with Lactobacillus rhamnosus CGMCC 1.3724 prevents development of atopic dermatitis in NC/NgaTnd mice possibly by modulating local production of IFN-gamma. Exp. Dermatol. 2009, 18, 1022–1027. [Google Scholar] [CrossRef]
- Brouwer, M.L.; Wolt-Plompen, S.A.; Dubois, A.E.; van der Heide, S.; Jansen, D.F.; Hoijer, M.A.; Kauffman, H.F.; Duiverman, E.J. No effects of probiotics on atopic dermatitis in infancy: A randomized placebo-controlled trial. Clin. Exp. Allergy 2006, 36, 899–906. [Google Scholar] [CrossRef]
- Tanaka, A.; Fukushima, Y.; Bevacoub, J.; Blum, S.; Matsuda, H. Prophylactic effect of oral administration of Lactobacillus johnsonii NCC533 (La1) during the weaning period on atopic dermatitis in NC/NgaTnd mice. Eur. J. Dermatol. 2008, 18, 136–140. [Google Scholar]
- Inoue, R.; Nishio, A.; Fukushima, Y.; Ushida, K. Oral treatment with probiotic Lactobacillus johnsonii NCC533 (La1) for a specific part of the weaning period prevents the development of atopic dermatitis induced after maturation in model mice, NC/Nga. Br. J. Dermatol. 2007, 156, 499–509. [Google Scholar] [CrossRef]
- Wickens, K.; Black, P.; Stanley, T.V.; Mitchell, E.; Fitzharris, P.; Tannock, G.W.; Purdie, G.; Crane, J.; Probiotic Study Group. A differential effect of 2 probiotics in the prevention of eczema and atopy: A double-blind, randomized, placebo controlled trial. J. Allergy Clin. Immunol. 2008, 122, 788–794. [Google Scholar] [CrossRef]
- Wickens, K.; Black, P.; Stanley, T.V.; Mitchell, E.; Barthow, C.; Fitzharris, P.; Purdie, G.; Crane, J. A protective effect of Lactobacillus rhamnosus HN001 against eczema in the first 2 years of life persists to age 4 years. Clin. Exp. Allergy 2012, 42, 1071–1079. [Google Scholar] [CrossRef]
- Shah, M.M.; Saio, M.; Yamashita, H.; Tanaka, H.; Takami, T.; Ezaki, T.; Inagaki, N. Lactobacillus acidophilus strain L-92 induces CD4+ CD25+ Foxp3+ regulatory T cells and suppresses allergic contact dermatitis. Biol. Pharm. Bull. 2012, 35, 612–616. [Google Scholar] [CrossRef]
- Karimi, K.; Inman, M.D.; Bienenstock, J.; Forsythe, P. Lactobacillus reuteri-induced regulatory T cells protect against an allergic airway response in mice. Am. J. Respir. Crit. Care Med. 2009, 197, 186–193. [Google Scholar]
- Kwon, H.; Lee, C.; So, J.; Chae, C.; Hwang, J.; Sahoo, A.; Nam, J.; Rhee, J.; Hwang, K.; Im, S. Generation of regulatory dendritic cells and CD4+ Foxp3+ T-cells by probiotics administration suppresses immune disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 2159–2164. [Google Scholar]
- Shida, K.; Takahashi, E.; Iwadate, E.; Takamizawa, K.; Yasui, H.; Sato, T.; Habu, S.; Hachimura, S.; Kaminogawa, S. Lactobacillus casei strain Shirota suppresses serum immunoglobulin E and immunoglobulin G1 responses and systemic anaphylaxis in a food allergy model. Clin. Exp. Allergy 2002, 32, 563–570. [Google Scholar] [CrossRef]
- Takahashi, N.; Kitazawa, H.; Iwabuchi, N.; Xiao, J.Z.; Miyaji, K.; Iwatsuki, K.; Saito, T. Immunostimulatory oligodeoxynucleotide from Bifidobacterium longum suppresses Th2 immune responses in a murine model. Clin. Exp. Immunol. 2006, 145, 130–138. [Google Scholar] [CrossRef]
- Schiffer, C.; Lalanne, A.I.; Cassard, L.; Mancardi, D.A.; Malbec, O.; Bruhns, P.; Dif, F.; Daeron, M. A strain of Lactobacillus casei inhibits the effector phase of immune inflammation. J. Immunol. 2011, 187, 1–10. [Google Scholar] [CrossRef]
- Dev, S.; Mizuguchi, H.; Das, A.K.; Matsushita, C.; Maeyama, K.; Umehara, H.; Ohtoshi, T.; Kojima, J.; Nishida, K.; Takahashi, K.; et al. Suppression of histamine signalling by probiotic Lac-B: A possible mechanism of its anti-allergic effect. J. Pharmacol. Sci. 2008, 107, 159–166. [Google Scholar] [CrossRef]
- Van Heel, D.A.; West, J. Recent advances in coeliac disease. Gut 2006, 55, 1037–1046. [Google Scholar] [CrossRef]
- Jabri, B.; Patey-Mariaud De Serre, N.; Cellier, C.; Evans, K.; Gache, C.; Carvalho, C.; Mougenot, J.F.; Allez, M.; Jian, R.; Desreumaux, P.; et al. Selectivr expanision of intraepithelial lymphocytes expressing the HLA-E-specific natural killer receptor CD94 in celiac disease. Gastroenterology 2000, 118, 867–879. [Google Scholar] [CrossRef]
- Maiuri, L.; Ciacci, C.; Vacca, L.; Ricciardelli, I.; Auricchio, S.; Quaratino, S.; Londei, M. IL-15 drives the specific migration of CD94+ and TCR-γδ+ intraepithelial lymphocytes in organ cultures of treated celiac patients IL-15 and Intraepithelial Migration of CD3 γδand CD94. Am. J. Gastroenterol. 2001, 96, 150–156. [Google Scholar]
- Meresse, B.; Chen, Z.; Ciszewski, C.; Tretiakova, M.; Bhagat, G.; Krausz, T.N.; Raulet, D.H.; Lanier, L.L.; Groh, V.; Spies, T.; et al. Coordinated induction of IL15 of a TCR-independent NKG2D signaling pathway converts CTL into lymphokine-activated killer cells in celiac disease. Immunity 2004, 21, 357–366. [Google Scholar] [CrossRef]
- Groh, V.; Steinle, A.; Bauer, S.; Spies, T. Recognition of stress-induced MHC molecules by intestinal epithelial γδ T cells. Science 1998, 279, 1737–1740. [Google Scholar] [CrossRef]
- Bauer, S.; Groh, V.; Wu, J.; Steinle, A.; Phillips, J.H.; Lanier, L.L.; Spies, T. Activation of NK cells and T cells by NKG2D, a receptor for stress-inducible MICA. Science 1999, 285, 727–729. [Google Scholar] [CrossRef]
- Hüe, S.; Mention, J.J.; Monteiro, R.C.; Zhang, S.; Cellier, C.; Schmitz, J.; Verkarre, V.; Fodil, N.; Bahram, S.; Cerf-Bensussan, N.; et al. A direct role for NKG2D/MICA interaction in villous atrophy during celiac disease. Immunity 2004, 21, 367–377. [Google Scholar] [CrossRef]
- Salvati, V.M.; MacDonald, T.T.; Bajaj-Elliott, M.; Borrelli, M.; Staiano, A.; Auricchio, S.; Troncone, R.; Monteleone, G. Interleukin 18 and associated markers of T helper cell type 1 activity in coeliac disease. Gut 2002, 50, 186–190. [Google Scholar] [CrossRef]
- Matysiak-B, T.; Moura, I.C.; Arcos-Fajardo, M.; Lebreton, C.; Menard, S.; Candalh, C.; Ben-Khalifa, K.; Dugave, C.; Tamouza, H.; van Niel, G.; et al. Sectretory IgA mediates retrotranscytosis of intact gliadin peptides via the transferrin receptor in celiac disease. J. Exp. Med. 2007, 205, 143–154. [Google Scholar]
- Dieterich, W.; Ehnis, T.; Bauer, M.; Donner, P.; Volta, U.; Riecken, E.O.; Schuppan, D. Identification of tissue transglutaminase as the autoantigen of celiac disease. Nat. Med. 1997, 3, 797–801. [Google Scholar] [CrossRef]
- Dahldom, I.; Korponay-Szabo, I.R.; Kovacs, J.B.; Szalai, Z.; Maki, M.; Hansson, T. Prediction of clinical and mucosal severity of coeliac disease and dermatitis herpetiformis by quantification of IgA/IgG serum antibodies to tissue transglutaminase. J. Pediatr. Gastroenterol. Nutr. 2010, 50, 140–146. [Google Scholar] [CrossRef]
- Sanz, Y.; Sanchez, E.; Marzotto, M.; Calabuig, M.; Torriani, S.; Dellaglio, F. Differences in faecal bacterial communities in coeliac and healthy children as detected by PCR and denaturing gradient gel electrophoresis. FEMS Immunol. Med. Microbiol. 2007, 51, 562–568. [Google Scholar] [CrossRef]
- Collado, M.; Donat, E.; Ribes-Koninckx, C.; Calabuig, M.; Sanz, Y. Imbalances in faecal and duodenal Bifidobacterium species composition in active and non-active coeliac disease. BMC Microbiol. 2008, 8, 1–9. [Google Scholar] [CrossRef]
- Silano, M.; Agostoni, C.; Guandalini, S. Effect of the timing of gluten introduction on the development of celiac disease. World J. Gastroenterol. 2010, 16, 1939–1942. [Google Scholar] [CrossRef]
- D’Arienzo, R.; Stefanile, R.; Maurano, F.; Mazzarella, G.; Ricca, E.; Troncone, R.; Auricchio, S.; Rossi, M. Immunomodulatory effects of Lactobacillus casei administration in a mouse model of gliadin-sensitive enteropathy. Basic Immunol. 2011, 74, 335–341. [Google Scholar]
- Krupa-Kozak, U.; Altamirano-Fortoul, R.; Wronkowska, M.; Rosell, C.M. Breadmaking performance and technological characteristic of gluten-free bread with inulin supplemented with calcium salts. Eur. Food Res. Technol. 2012, 235, 545–554. [Google Scholar] [CrossRef]
- Czesnikiewicz-Guzik, M.; Lee, W.-W.; Cui, D.; Hiruma, Y.; Lamar, D.L.; Yang, Z.-Z.; Ouslander, J.G.; Weyand, C.M.; Goronzy, J.J. T cell subset-specific susceptibility to aging. Clin. Immunol. 2008, 127, 107–118. [Google Scholar] [CrossRef]
- Zhang, Y.; Wallace, D.L.; de Lara, C.M.; Ghattas, H.; Asquith, B.; Worth, A.; Griffin, G.E.; Taylor, G.P.; Tough, D.F.; Beverley, P.C.; et al. In vivo kinetics of human natural killer cells: The effects of ageing and acute and chronic viral infection. Immunology 2007, 121, 258–265. [Google Scholar] [CrossRef]
- Dong, H.; Rowland, I.; Thomas, L.V.; Yaqoob, P. Immunomodulatory effects of a probiotic drink containing Lactobacillus casei Shirota in healthy older volunteers. Eur. J. Nutr. 2013. [Google Scholar] [CrossRef]
- Kim, H.G.; Lee, S.Y.; Kim, N.R.; Ko, M.Y.; Lee, J.M.; Yi, T.H.; Chung, S.K.; Chung, D.K. Inhibitory effects of Lactobacillus plantarum Lipoteichoic acid (LTA) on Staphylococcus aureus LTA-induced tumour necrosis factor-alpha production. J. Microbiol. Biotechnol. 2008, 18, 1191–1196. [Google Scholar]
- Kim, H.G.; Lee, S.Y.; Kim, N.R.; Lee, H.Y.; Ko, M.Y.; Jung, B.J.; Kim, C.M.; Lee, J.M.; Park, J.H.; Han, S.H.; Chung, D.K. Lactobacillus plantarum lipoteichoic acid down regulated Shigella flexneri peptidoglycan-induced inflammation. Mol. Immunol. 2011, 48, 382–291. [Google Scholar] [CrossRef]
- Kleerebezem, M.; Hols, P.; Bernard, E.; Rolain, T.; Zhou, M.; Siezen, R.J.; Bron, P.A. The extracellular biology of the lactobacilli. FEMS Microbiol. Rev. 2010, 34, 199–230. [Google Scholar] [CrossRef]
- Preidis, G.A.; Versalovic, J. Targeting the human microbiome with antibiotics, probiotics and prebiotics: Gastroenterology enters the metagenomics era. Gastroenterology 2009, 136, 2015–2031. [Google Scholar] [CrossRef]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 9, 222–227. [Google Scholar]
- Anderson, J.L.; Edney, R.J.; Whelan, K. Systematic review: Faecal microbiota transplantation in the management of inflammatory bowel disease. Aliment. Pharmacol. Ther. 2012, 36, 503–516. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hardy, H.; Harris, J.; Lyon, E.; Beal, J.; Foey, A.D. Probiotics, Prebiotics and Immunomodulation of Gut Mucosal Defences: Homeostasis and Immunopathology. Nutrients 2013, 5, 1869-1912. https://doi.org/10.3390/nu5061869
Hardy H, Harris J, Lyon E, Beal J, Foey AD. Probiotics, Prebiotics and Immunomodulation of Gut Mucosal Defences: Homeostasis and Immunopathology. Nutrients. 2013; 5(6):1869-1912. https://doi.org/10.3390/nu5061869
Chicago/Turabian StyleHardy, Holly, Jennifer Harris, Eleanor Lyon, Jane Beal, and Andrew D. Foey. 2013. "Probiotics, Prebiotics and Immunomodulation of Gut Mucosal Defences: Homeostasis and Immunopathology" Nutrients 5, no. 6: 1869-1912. https://doi.org/10.3390/nu5061869
APA StyleHardy, H., Harris, J., Lyon, E., Beal, J., & Foey, A. D. (2013). Probiotics, Prebiotics and Immunomodulation of Gut Mucosal Defences: Homeostasis and Immunopathology. Nutrients, 5(6), 1869-1912. https://doi.org/10.3390/nu5061869





