Human Milk-Derived Levels of let-7g-5p May Serve as a Diagnostic and Prognostic Marker of Low Milk Supply in Breastfeeding Women
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Participant Characteristics
2.3. Milk Collection
2.4. RNA Processing
2.5. Cell Culture and let-7g-5p Transfection
2.6. Immunoblotting
2.7. Statistical Methods
3. Results
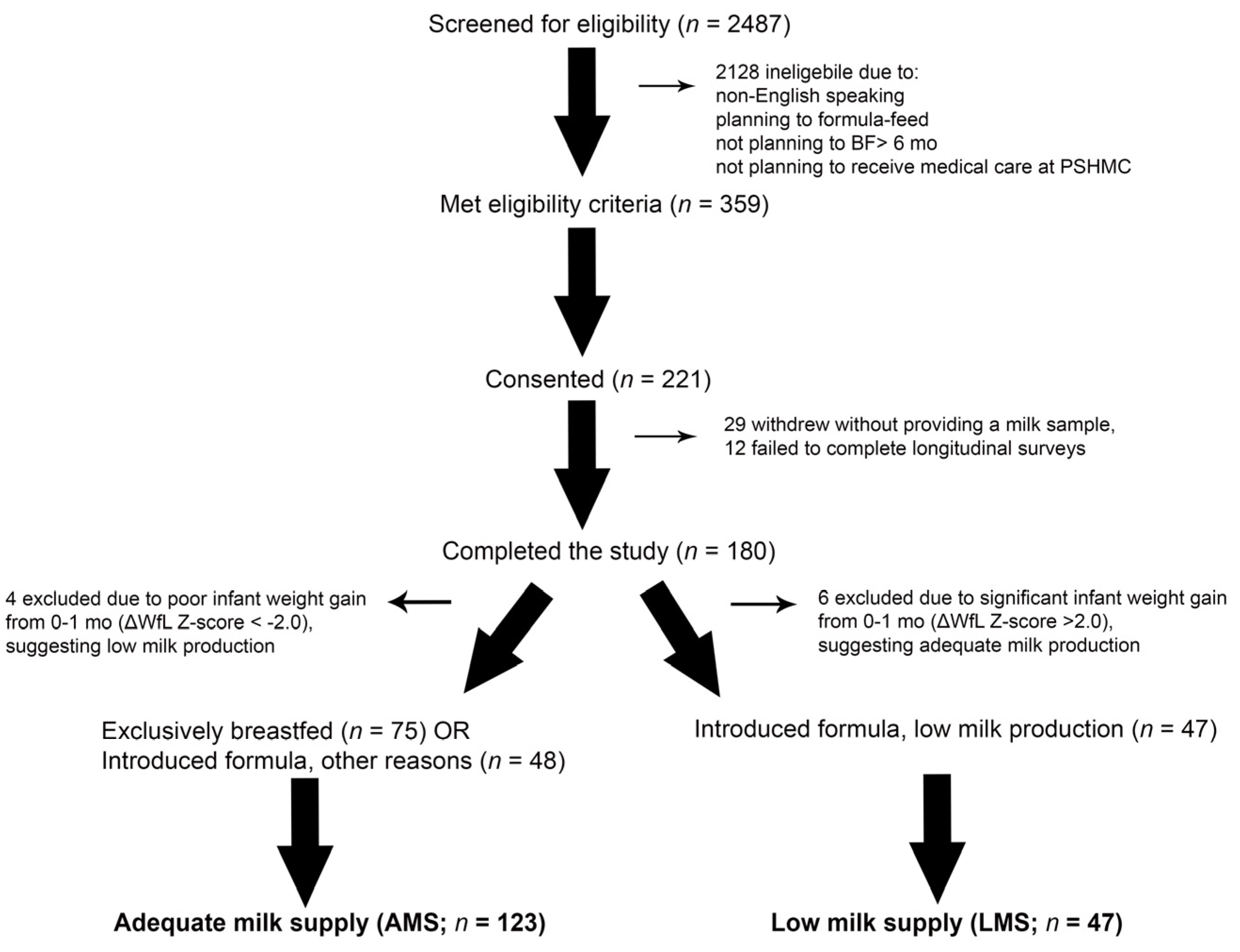
3.1. Participants
3.2. Milk Production and Infant Weight Trajectory
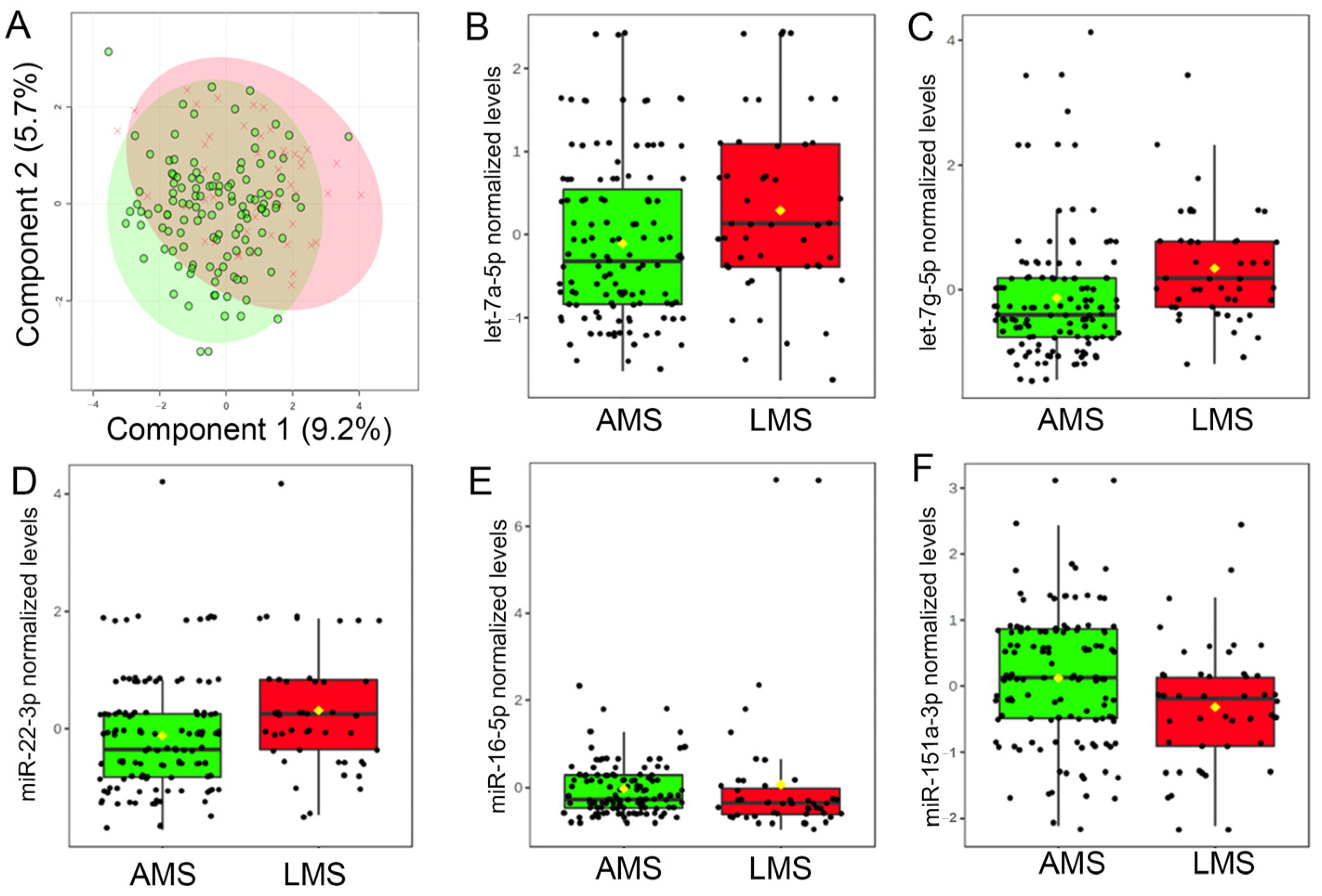
3.3. Milk miRNA Profiles
3.4. Longitudinal Changes in LMS-Related miRNAs
3.5. Effect of Modifiable Maternal Characteristics on LMS-Related miRNAs
3.6. Predicting Low Milk Supply Status
3.7. Pathway Analysis
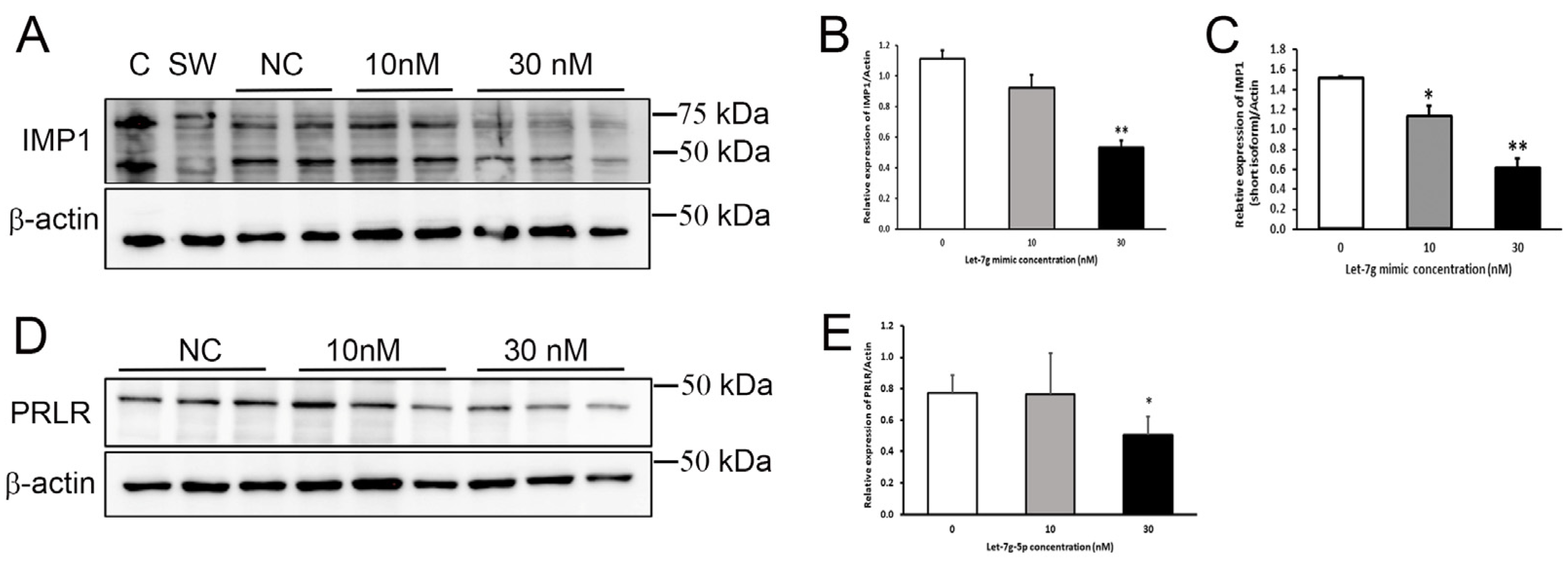
3.8. Confirmation of PRLR and IGF2BP1 Regulation in Mammary Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahluwalia, I.B.; Morrow, B.; Hsia, J. Why do women stop breastfeeding? Findings from the pregnancy risk assessment and monitoring system. Pediatrics 2005, 116, 1408–1412. [Google Scholar] [CrossRef] [PubMed]
- Neifert, M.; DeMarzo, S.; Seacat, J.; Young, D.; Leff, M.; Orleans, M. The influence of breast surgery, breast appearance, and pregnancy-induced breast changes on lactation sufficiency as measured by infant weight gain. Birth 1990, 17, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, B.; Dias, C.C.; Brandão, S.; Canário, C.; Nunes-Costa, R. Breastfeeding and postpartum depression: State of the art review. J. Pediatr. 2013, 89, 332–338. [Google Scholar] [CrossRef] [PubMed]
- Gilmour, C.; Hall, H.; McIntyre, M.; Gillies, L.; Harrison, B. Factors associated with early breastfeeding cessation in frankston, victoria: A descriptive study. Breastfeed. Rev. 2009, 17, 13–19. [Google Scholar]
- Meier, P.; Patel, A.L.; Wright, K.; Engstrom, J.L. Management of breastfeeding during and after the maternity hospitalization for late preterm infants. Clin. Perinatol. 2013, 40, 689–705. [Google Scholar] [CrossRef]
- Szabo, A.L. Review article: Intrapartum neuraxial analgesia and breastfeeding outcomes: Limitations of current knowledge. Anesth. Analg. 2013, 116, 399–405. [Google Scholar] [CrossRef]
- Ettyang, G.A.; van Marken Lichtenbelt, W.D.; Esamai, F.; Saris, W.H.; Westerterp, K.R. Assessment of body composition and breast milk volume in lactating mothers in pastoral communities in pokot, kenya, using deuterium oxide. Ann. Nutr. Metab. 2005, 49, 110–117. [Google Scholar] [CrossRef]
- Diana, A.; Haszard, J.J.; Houghton, L.A.; Gibson, R.S. Breastmilk intake among exclusively breastfed indonesian infants is negatively associated with maternal fat mass. Eur. J. Clin. Nutr. 2019, 73, 1206–1208. [Google Scholar] [CrossRef]
- Suwaydi, M.A.; Wlodek, M.E.; Lai, C.T.; Prosser, S.A.; Geddes, D.T.; Perrella, S.L. Delayed secretory activation and low milk production in women with gestational diabetes: A case series. BMC Pregnancy Childbirth 2022, 22, 350. [Google Scholar] [CrossRef]
- Moriwaki, M.; Welt, C. Mon-434 familial alactogenesis associated with a prolactin mutation. J. Endocr. Soc. 2019, 3 (Suppl. 1), MON-434. [Google Scholar] [CrossRef]
- Kobayashi, T.; Usui, H.; Tanaka, H.; Shozu, M. Variant prolactin receptor in agalactia and hyperprolactinemia. N. Engl. J. Med. 2018, 379, 2230–2236. [Google Scholar] [CrossRef] [PubMed]
- Rivera, O.C.; Geddes, D.T.; Barber-Zucker, S.; Zarivach, R.; Gagnon, A.; Soybel, D.I.; Kelleher, S.L. A common genetic variant in zinc transporter znt2 (thr288ser) is present in women with low milk volume and alters lysosome function and cell energetics. Am. J. Physiol. Cell Physiol. 2020, 318, C1166–C1177. [Google Scholar] [CrossRef] [PubMed]
- Chandran, D.; Confair, A.; Warren, K.; Kawasawa, Y.I.; Hicks, S.D. Maternal variants in the mfge8 gene are associated with perceived breast milk supply. Breastfeed. Med. 2022, 17, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Li, J.; Cairns, M.J. Identifying mirnas, targets and functions. Brief Bioinform. 2014, 15, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Arantes, L.; De Carvalho, A.C.; Melendez, M.E.; Lopes Carvalho, A. Serum, plasma and saliva biomarkers for head and neck cancer. Expert Rev. Mol. Diagn. 2018, 18, 85–112. [Google Scholar] [CrossRef]
- Tölle, A.; Blobel, C.C.; Jung, K. Circulating mirnas in blood and urine as diagnostic and prognostic biomarkers for bladder cancer: An update in 2017. Biomark. Med. 2018, 12, 667–676. [Google Scholar] [CrossRef]
- Alsaweed, M.; Lai, C.T.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human milk cells contain numerous mirnas that may change with milk removal and regulate multiple physiological processes. Int. J. Mol. Sci. 2016, 17, 956. [Google Scholar] [CrossRef]
- Avril-Sassen, S.; Goldstein, L.D.; Stingl, J.; Blenkiron, C.; Le Quesne, J.; Spiteri, I.; Karagavriilidou, K.; Watson, C.J.; Tavaré, S.; Miska, E.A.; et al. Characterisation of microrna expression in post-natal mouse mammary gland development. BMC Genom. 2009, 10, 548. [Google Scholar] [CrossRef]
- Li, Z.; Liu, H.; Jin, X.; Lo, L.; Liu, J. Expression profiles of micrornas from lactating and non-lactating bovine mammary glands and identification of mirna related to lactation. BMC Genom. 2012, 13, 731. [Google Scholar] [CrossRef]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Time-dependent expression profiles of micrornas and mrnas in rat milk whey. PLoS ONE 2014, 9, e88843. [Google Scholar] [CrossRef]
- Alsaweed, M.; Lai, C.T.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human milk cells and lipids conserve numerous known and novel mirnas, some of which are differentially expressed during lactation. PLoS ONE 2016, 11, e0152610. [Google Scholar] [CrossRef]
- Hatmal, M.M.M.; Al-Hatamleh, M.A.; Olaimat, A.N.; Alshaer, W.; Hasan, H.; Albakri, K.A.; Alkhafaji, E.; Issa, N.N.; Al-Holy, M.A.; Abderrahman, S.M.; et al. Immunomodulatory properties of human breast milk: Microrna contents and potential epigenetic effects. Biomedicines 2022, 10, 1219. [Google Scholar] [CrossRef]
- Perri, M.; Lucente, M.; Cannataro, R.; De Luca, I.F.; Gallelli, L.; Moro, G.; De Sarro, G.; Caroleo, M.C.; Cione, E. Variation in immune-related micrornas profile in human milk amongst lactating women. MicroRNA 2018, 7, 107–114. [Google Scholar] [CrossRef]
- Tingö, L.; Ahlberg, E.; Johansson, L.; Pedersen, S.A.; Chawla, K.; Sætrom, P.; Cione, E.; Simpson, M.R. Non-coding rnas in human breast milk: A systematic review. Front. Immunol. 2021, 12, 725323. [Google Scholar] [CrossRef]
- Thompson, F.E.; Midthune, D.; Kahle, L.; Dodd, K.W. Development and evaluation of the national cancer institute’s dietary screener questionnaire scoring algorithms. J. Nutr. 2017, 147, 1226–1233. [Google Scholar] [CrossRef]
- Carney, M.C.; Tarasiuk, A.; DiAngelo, S.L.; Silveyra, P.; Podany, A.; Birch, L.L.; Paul, I.M.; Kelleher, S.; Hicks, S.D. Metabolism-related micrornas in maternal breast milk are influenced by premature delivery. Pediatr. Res. 2017, 82, 226–236. [Google Scholar] [CrossRef]
- Hicks, S.D.; Confair, A.; Warren, K.; Chandran, D. Levels of breast milk micrornas and other non-coding rnas are impacted by milk maturity and maternal diet. Front. Immunol. 2022, 12, 5754. [Google Scholar] [CrossRef]
- Vlachos, I.S.; Zagganas, K.; Paraskevopoulou, M.D.; Georgakilas, G.; Karagkouni, D.; Vergoulis, T.; Dalamagas, T.; Hatzigeorgiou, A.G. Diana-mirpath v3.0: Deciphering microrna function with experimental support. Nucleic Acids Res. 2015, 43, W460–W466. [Google Scholar] [CrossRef]
- Fakhraldeen, S.; Clark, R.; Roopra, A.; Chin, E.; Huang, W.; Castorino, J.; Wisinski, T.; Spiegelman, V.; Alexander, C. Two isoforms of the rna binding protein, coding region determinant-binding protein (crd-bp/igf2bp1), are expressed in breast epithelium and support clonogenic growth of breast tumor cells. J. Biol. Chem. 2015, 290, 13386–13400. [Google Scholar] [CrossRef]
- Tian, L.; Li, Y.; Wang, C.; Li, Q. Let-7g-5p regulates mouse mammary cells differentiation and function by targeting prkca. J. Cell. Physiol. 2019, 234, 10101–10110. [Google Scholar] [CrossRef]
- Lee, H.; Han, S.; Kwon, C.S.; Lee, D. Biogenesis and regulation of the let-7 mirnas and their functional implications. Protein Cell 2016, 7, 100–113. [Google Scholar] [CrossRef] [PubMed]
- Boyerinas, B.; Park, S.M.; Hau, A.; Murmann, A.E.; Peter, M.E. The role of let-7 in cell differentiation and cancer. Endocr.-Relat. Cancer 2010, 17, F19–F36. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Shyh-Chang, N.; Segrè Ayellet, V.; Shinoda, G.; Shah Samar, P.; Einhorn William, S.; Takeuchi, A.; Engreitz Jesse, M.; Hagan John, P.; Kharas Michael, G.; et al. The Lin28/let-7 axis regulates glucose metabolism. Cell 2011, 147, 81–94. [Google Scholar] [CrossRef]
- Viswanathan, S.R.; Daley, G.Q.; Gregory, R.I. Selective blockade of microrna processing by lin28. Science 2008, 320, 97–100. [Google Scholar] [CrossRef]
- Mills WTt Nassar, N.N.; Ravindra, D.; Li, X.; Meffert, M.K. Multi-level regulatory interactions between nf-κb and the pluripotency factor lin28. Cells 2020, 9, 2710. [Google Scholar] [CrossRef]
- Ingman, W.V.; Glynn, D.J.; Hutchinson, M.R. Inflammatory mediators in mastitis and lactation insufficiency. J. Mammary Gland. Biol. Neoplasia 2014, 19, 161–167. [Google Scholar] [CrossRef]
- Nommsen-Rivers, L.A.; Chantry, C.J.; Peerson, J.M.; Cohen, R.J.; Dewey, K.G. Delayed onset of lactogenesis among first-time mothers is related to maternal obesity and factors associated with ineffective breastfeeding. Am. J. Clin. Nutr. 2010, 92, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Gilyazova, I.; Ivanova, E.; Gilyazova, G.; Sultanov, I.; Izmailov, A.; Safiullin, R.; Pavlov, V.; Khusnutdinova, E. Methylation and expression levels of microrna-23b/-24-1/-27b, microrna-30c-1/-30e, microrna-301a and let-7g are dysregulated in clear cell renal cell carcinoma. Mol. Biol. Rep. 2021, 48, 5561–5569. [Google Scholar] [CrossRef] [PubMed]
- Geoffroy, A.; Kerek, R.; Pourié, G.; Helle, D.; Guéant, J.L.; Daval, J.L.; Bossenmeyer-Pourié, C. Late maternal folate supplementation rescues from methyl donor deficiency-associated brain defects by restoring let-7 and mir-34 pathways. Mol. Neurobiol. 2017, 54, 5017–5033. [Google Scholar] [CrossRef]
- Qian, P.; Zuo, Z.; Wu, Z.; Meng, X.; Li, G.; Wu, Z.; Zhang, W.; Tan, S.; Pandey, V.; Yao, Y.; et al. Pivotal role of reduced let-7g expression in breast cancer invasion and metastasis. Cancer Res. 2011, 71, 6463–6474. [Google Scholar] [CrossRef]
- Bhat-Nakshatri, P.; Wang, G.; Collins, N.R.; Thomson, M.J.; Geistlinger, T.R.; Carroll, J.S.; Brown, M.; Hammond, S.; Srour, E.F.; Liu, Y.; et al. Estradiol-regulated micrornas control estradiol response in breast cancer cells. Nucleic Acids Res. 2009, 37, 4850–4861. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Zhang, A.; Zhang, J.; Wang, R.; Xia, H. Role of perinatal biological factors in delayed lactogenesis ii among women with pre-pregnancy overweight and obesity. Biol. Res. Nurs. 2022, 24, 459–471. [Google Scholar] [CrossRef] [PubMed]
- Bomfim, G.F.; Merighe, G.K.F.; de Oliveira, S.A.; Negrao, J.A. Acute and chronic effects of cortisol on milk yield, the expression of key receptors, and apoptosis of mammary epithelial cells in saanen goats. J. Dairy Sci. 2022, 105, 818–830. [Google Scholar] [CrossRef] [PubMed]
- Kelly, P.A.; Bachelot, A.; Kedzia, C.; Hennighausen, L.; Ormandy, C.J.; Kopchick, J.J.; Binart, N. The role of prolactin and growth hormone in mammary gland development. Mol. Cell. Endocrinol. 2002, 197, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Sureshbabu, A.; Tonner, E.; Flint, D.J. Insulin-like growth factor binding proteins and mammary gland development. Int. J. Dev. Biol. 2011, 55, 781–789. [Google Scholar] [CrossRef] [PubMed]
- Tsutsui, S.; Wakasa, H.; Tsugami, Y.; Suzuki, T.; Nishimura, T.; Kobayashi, K. Distinct expression patterns of fibrillar collagen types i, iii, and v in association with mammary gland remodeling during pregnancy, lactation and weaning. J. Mammary Gland. Biol. Neoplasia 2020, 25, 219–232. [Google Scholar] [CrossRef]
- McCormick, N.H.; Lee, S.; Hennigar, S.R.; Kelleher, S.L. Znt4 (slc30a4)-null (“lethal milk”) mice have defects in mammary gland secretion and hallmarks of precocious involution during lactation. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2016, 310, R33–R40. [Google Scholar] [CrossRef]
- Lee, S.; Rivera, O.C.; Kelleher, S.L. Zinc transporter 2 interacts with vacuolar atpase and is required for polarization, vesicle acidification, and secretion in mammary epithelial cells. J. Biol. Chem. 2017, 292, 21598–21613. [Google Scholar] [CrossRef]
- Reinhardt, T.A.; Lippolis, J.D. Mammary gland involution is associated with rapid down regulation of major mammary ca2+-atpases. Biochem. Biophys. Res. Commun. 2009, 378, 99–102. [Google Scholar] [CrossRef]
- Hanna, S.; Khalil, B.; Nasrallah, A.; Saykali, B.A.; Sobh, R.; Nasser, S.; El-Sibai, M. Stard13 is a tumor suppressor in breast cancer that regulates cell motility and invasion. Int. J. Oncol. 2014, 44, 1499–1511. [Google Scholar] [CrossRef]
- Degrauwe, N.; Suvà, M.L.; Janiszewska, M.; Riggi, N.; Stamenkovic, I. Imps: An rna-binding protein family that provides a link between stem cell maintenance in normal development and cancer. Genes Dev. 2016, 30, 2459–2474. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.; Zhao, C.; Zhu, Q.; Katakura, Y.; Tanaka, H.; Ohnuki, K.; Shimizu, K. Ursolic acid isolated from the leaves of loquat (Eriobotrya japonica) inhibited osteoclast differentiation through targeting exportin 5. J. Agric. Food Chem. 2019, 67, 3333–3340. [Google Scholar] [CrossRef] [PubMed]
- Seo, D.Y.; Lee, S.R.; Heo, J.W.; No, M.H.; Rhee, B.D.; Ko, K.S.; Kwak, H.B.; Han, J. Ursolic acid in health and disease. Korean J. Physiol. Pharmacol. 2018, 22, 235–248. [Google Scholar] [CrossRef] [PubMed]
- Lam, T.K.; Shao, S.; Zhao, Y.; Marincola, F.; Pesatori, A.; Bertazzi, P.A.; Caporaso, N.E.; Wang, E.; Landi, M.T. Influence of quercetin-rich food intake on microrna expression in lung cancer tissues. Cancer Epidemiol. Biomark. Prev. 2012, 21, 2176–2184. [Google Scholar] [CrossRef]
- Kupsco, A.; Prada, D.; Valvi, D.; Hu, L.; Petersen, M.S.; Coull, B.; Grandjean, P.; Weihe, P.; Baccarelli, A.A. Human milk extracellular vesicle mirna expression and associations with maternal characteristics in a population-based cohort from the faroe islands. Sci. Rep. 2021, 11, 5840. [Google Scholar] [CrossRef] [PubMed]
- Alsaweed, M.; Hepworth, A.R.; Lefèvre, C.; Hartmann, P.E.; Geddes, D.T.; Hassiotou, F. Human milk microrna and total rna differ depending on milk fractionation. J. Cell. Biochem. 2015, 116, 2397–2407. [Google Scholar] [CrossRef] [PubMed]
- Munch, E.M.; Harris, R.A.; Mohammad, M.; Benham, A.L.; Pejerrey, S.M.; Showalter, L.; Hu, M.; Shope, C.D.; Maningat, P.D.; Gunaratne, P.H.; et al. Transcriptome profiling of microrna by next-gen deep sequencing reveals known and novel mirna species in the lipid fraction of human breast milk. PLoS ONE 2013, 8, e50564. [Google Scholar] [CrossRef]


| All (n = 170) | AMS (n = 123) | LMS (n = 47) | |
|---|---|---|---|
| Age in years, mean (SD) | 30 (4.6) | 29.5 (4.5) | 31.3 (4.7) * |
| Asian, n (%) | 8 (4.7) | 6 (4.9) | 2 (4.3) |
| Bi-racial, n (%) | 6 (3.5) | 4 (3.3) | 2 (4.3) |
| African American, n (%) | 12 (7.1) | 8 (6.5) | 4 (8.5) |
| Other—not specified, n (%) | 3 (1.8) | 3 (2.4) | 0 (0.0) |
| Other—specified, n (%) | 6 (3.5) | 4 (3.3) | 2 (4.3) |
| White, n (%) | 135 (79.4) | 98 (79.7) | 37 (78.7) |
| Parity, # (SD) | 2 (1) | 2 (1) | 2 (1) |
| Pre-pregnancy BMI, kg/m2, mean (SD) | 27.8 (6.7) | 27.2 (6.1) | 29.2 (8.0) |
| Gestational diabetes, n (%) | 20 (11.8) | 13 (10.6) | 7 (14.9) |
| Prior tobacco use, n (%) | 26 (15.3) | 17 (13.8) | 9 (19.1) |
| Some HS, n (%) | 2 (1.2) | 2 (1.6) | 0 (0.0) |
| Completed HS, n (%) | 22 (12.9) | 19 (15.4) | 3 (6.4) * |
| Some college, n (%) | 25 (14.7) | 16 (13.0) | 9 (19.1) |
| Completed college, n (%) | 69 (40.6) | 56 (45.5) | 13 (27.7) |
| Post-graduate degree, n (%) | 52 (30.6) | 30 (24.4) | 22 (46.8) * |
| Single, n (%) | 11 (6.5) | 9 (7.3) | 2 (4.3) |
| Divorced, n (%) | 2 (1.2) | 1 (0.8) | 1 (2.1) |
| Co-habitating, n (%) | 23 (13.5) | 16 (13.0) | 7 (14.9) |
| Married, n (%) | 134 (78.8) | 97 (78.9) | 37 (78.7) |
| Never, n (%) | 67 (39.4) | 42 (34.1) | 25 (53.2) * |
| <1 month, n (%) | 4 (2.4) | 1 (0.8) | 3 (6.4) |
| 1–2 months, n (%) | 4 (2.4) | 4 (3.3) | 0 (0.0) |
| 2–4 months, n (%) | 5 (2.9) | 4 (3.3) | 1 (2.1) |
| >4 months, n (%) | 90 (52.9) | 72 (58.5) | 18 (38.3) * |
| Gestational age, weeks (SD) | 38.9 (1.0) | 38.9 (1.0) | 39.0 (1.0) |
| Infant Sex, female, n (%) | 99 (58.2) | 74 (60.1) | 25 (53.1) |
| Birth weight, grams (SD) | 3364 (440) | 3377 (440) | 3329 (443) |
| All (n = 170) | AMS (n = 123) | LMS (n = 47) | |
|---|---|---|---|
| Formula intro by 4 weeks, n (%) | 55 (32.3) | 27 (21.9) | 28 (59.5) * |
| Breastfeeding at 6 months, n (%) | 134 (78.8) | 105 (85.4) | 29 (61.7) * |
| Milk production at 4 weeks, oz/day (SD) | 26.0 (15.0) | 28.1 (15.6) | 20.6 (11.8) * |
| Milk production at 16 weeks, oz/day (SD) | 29.6 (22.5) | 33.3 (24.0) | 20.1 (14.1) * |
| Infant WfL Z-score at birth | −0.72 (1.1) | −0.71 (1.1) | −0.76 (1.0) |
| Infant WfL Z-score at 4 weeks | 0.38 (1.2) | 0.50 (1.1) | 0.05 (1.2) * |
| ∆ WfL Z-score from birth to 4 weeks | 1.1 (1.4) | 1.2 (1.4) | 0.8 (1.3) * |
| KEGG Pathway | p-Value | Genes (#) | miRNAs (#) |
|---|---|---|---|
| Prion diseases | 6.88 × 10−17 | 13 | 2 |
| Cell cycle | 1.11 × 10−16 | 63 | 3 |
| Hepatitis B | 3.77 × 10−15 | 72 | 3 |
| Proteoglycans in cancer | 4.77 × 10−14 | 85 | 2 |
| Fatty acid biosynthesis | 1.63 × 10−13 | 4 | 2 |
| Adherens junction | 1.29 × 10−11 | 46 | 2 |
| Hippo signaling pathway | 1.42 × 10−10 | 57 | 3 |
| TGF-beta signaling pathway | 1.32 × 10−09 | 31 | 1 |
| Lysine degradation | 1.77 × 10−09 | 15 | 1 |
| Oocyte meiosis | 6.19 × 10−09 | 35 | 1 |
| Viral carcinogenesis | 9.29 × 10−9 | 102 | 4 |
| ECM-receptor interaction | 8.02 × 10−08 | 16 | 2 |
| Transcriptional misregulation | 2.10 × 10−05 | 38 | 1 |
| in cancer |
| KEGG Pathway | Genes (#) |
|---|---|
| Metabolic pathways | 72 |
| Pathways in cancer | 39 |
| PI3K-Akt signaling pathway | 33 |
| Human Papillomavirus infection | 30 |
| Herpes simplex virus infection | 29 |
| Pathways of neurodegeneration | 28 |
| Proteoglycans in cancer | 23 |
| MicroRNAs in cancer | 23 |
| Focal adhesion | 21 |
| Axon guidance | 21 |
| Human T-cell leukemia virus 1 infection | 20 |
| Cytokine-cytokine receptor interaction | 20 |
| Transcriptional misregulation in cancer | 19 |
| FOXO signaling pathway | 19 |
| Ras signaling pathway | 19 |
| Calcium signaling pathway | 18 |
| mTOR signaling pathway | 18 |
| JAK-STAT signaling pathway | 18 |
| Lipid and atherosclerosis | 18 |
| AGE-RAGE signaling pathway | 18 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hicks, S.D.; Chandran, D.; Confair, A.; Ward, A.; Kelleher, S.L. Human Milk-Derived Levels of let-7g-5p May Serve as a Diagnostic and Prognostic Marker of Low Milk Supply in Breastfeeding Women. Nutrients 2023, 15, 567. https://doi.org/10.3390/nu15030567
Hicks SD, Chandran D, Confair A, Ward A, Kelleher SL. Human Milk-Derived Levels of let-7g-5p May Serve as a Diagnostic and Prognostic Marker of Low Milk Supply in Breastfeeding Women. Nutrients. 2023; 15(3):567. https://doi.org/10.3390/nu15030567
Chicago/Turabian StyleHicks, Steven D., Desirae Chandran, Alexandra Confair, Anna Ward, and Shannon L. Kelleher. 2023. "Human Milk-Derived Levels of let-7g-5p May Serve as a Diagnostic and Prognostic Marker of Low Milk Supply in Breastfeeding Women" Nutrients 15, no. 3: 567. https://doi.org/10.3390/nu15030567
APA StyleHicks, S. D., Chandran, D., Confair, A., Ward, A., & Kelleher, S. L. (2023). Human Milk-Derived Levels of let-7g-5p May Serve as a Diagnostic and Prognostic Marker of Low Milk Supply in Breastfeeding Women. Nutrients, 15(3), 567. https://doi.org/10.3390/nu15030567





