Abstract
The gut is a pivotal organ in health and disease. The events that take place in the gut during early life contribute to the programming, shaping and tuning of distant organs, having lifelong consequences. In this context, the maternal gut plays a quintessence in programming the mammary gland to face the nutritional, microbiological, immunological, and neuroendocrine requirements of the growing infant. Subsequently, human colostrum and milk provides the infant with an impressive array of nutrients and bioactive components, including microbes, immune cells, and stem cells. Therefore, the axis linking the maternal gut, the breast, and the infant gut seems crucial for a correct infant growth and development. The aim of this article is not to perform a systematic review of the human milk components but to provide an insight of their extremely complex interactions, which render human milk a unique functional food and explain why this biological fluid still truly remains as a scientific enigma.
1. Introduction
The intestine could be defined as the “ugly duckling” of our body; it has provided so many surprises to the scientific community in the last decades that it currently appears to us as a truly fascinating organ. First of all, large volumes of food, chemicals and microorganisms pass through this huge organ continuously, some of them potentially toxic or pathogenic. On the other hand, it is in charge of carrying out a remarkably efficient rescue operation: the rescue of the essential nutrients existing in an intermittent magma of food and water. To achieve this, it has to conduct this content at an adequate speed; provide various secretions; perform digestion tasks; absorb the digested products, electrolytes, and water; send this material into the circulatory stream; and, finally, expel waste products.
As if this were not enough, it is a keystone of the immune system, performing a conscientious task of sampling the enteric environment and having to maintain a difficult balance: firmly controlling pathogens, tolerating diners, and originating proportionate responses in front of the crowd of antigens with which it has contact. There is more still: the gut is the host of a certainly complex microbiota, which, directly or indirectly, contributes to most, if not all, host functions. Unveiling the roles of the gut microbiota in health and disease remains one of the most important challenges in biomedicine, and this may provide a key knowledge for medical personalization. Finally, the intestine is a complex neuroendocrine organ. It maintains a constant exchange of information with the brain and with other organs through a complex network of nervous, metabolic, and endocrine signals. The enteric nervous system is a vast chemical warehouse in which each of the neurotransmitters that operate in our brain is represented. Its mission goes far beyond the supervision of the already complex digestive processes and includes the production of psychoactive substances. The enteric nervous system is a place of neural integration and processing, which is why it has been considered our “second brain” [1]. It is illustrative that the intestine is precisely the organ in which the physical connections between the immune system and the nervous system were first revealed [2].
As a result, the gut is in the crossroads of health and disease, programming and modulating the function of distant organs. In fact, in the last decades, we have assisted in the blooming of an increasing number of axes names used to highlight the functional connections of the gut with almost any other human organ: gut‒brain axis [3], gut‒lung axis [4], gut‒liver axis [5], gut‒kidney axis [6], gut‒skin axis [7], gut‒joint axis [8], gut‒bone axis [9], gut‒pancreas axis [10], gut‒heart axis [11], gut‒spleen axis [12], gut‒vagina axis [13], and gut‒eye axis [14]. Among this myriad of interconnected axes, one of the most relevant ones for our biological history is usually forgotten: the gut‒breast axis.
2. The Maternal Gut‒Breast Axis
2.1. Gut and Breast Adaptations for Pregnancy and Lactation
The maternal gut plays a key role in programming and training the mammary gland to meet the nutritional, microbiological, immunological, and neuroendocrine requirements of the growing infant [15] and, therefore, in the short- and long-term benefits associated to breastfeeding [16,17]. Pregnancy and lactation require extensive changes in a woman’s body, involving most, if not all, organs and systems. Among them, the morphological and physiological adaptations of the maternal gastrointestinal tract are particularly relevant for the success of these two key life stages. These adaptations affect absorption ability, metabolism (including bacterial metabolism), and the composition of both the microbiota and the immune system [18,19,20,21,22]. Overall, the volume of the mucosal surface of the gut increases sharply to create an enormous absorptive area, especially in those parts (duodenum and ileum) that are highly involved in the breakdown of food components and nutrient absorption [19,23]. In addition, the conditions created during the last trimester of pregnancy favors a physiological translocation of immune and bacterial cells [24].
Parallel, there is a deep structural and functional remodeling of the mammary gland, including a huge process of angiogenesis, leading to the development of a highly specialized exocrine organ, ready for the transmission of a vast array of nutrients and bioactive compounds and cells to the infant gut. Gut and mammary adaptations are perfectly synchronized, since both organs establish complex and complementary interactions [25]. As stated by Weaver [18], “the relationship between lactating mammary function and neonatal gastrointestinal function is an example of the parallel evolution of two organs, the breast and gut, which after birth, together undertake functions performed by the placenta during intrauterine life.”
2.2. Maternal Gut and Breast: The Nutritional Connection
The main sources of milk components are maternal diet, maternal stores, and de novo biosynthesis by the lactocytes or by other cells (immune cells, microbes) inhabiting the mammary epithelium and/or carried by milk [26] (Figure 1). In this context, a correct functionality of the gastrointestinal tract (absorption, metabolism) is strictly necessary to assemble the components of human milk, either directly from the maternal diet or to provide the immediate principles (fatty acids, amino acids, glucose), water, and electrolytes required for the maternal cells to build them up (Figure 2).
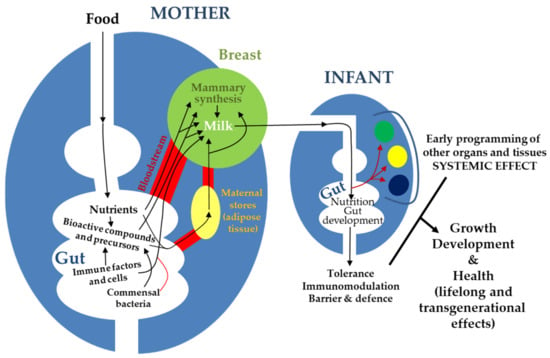
Figure 1.
Schematic representation of the complex interactions between maternal gut, milk, and infant gut and their relevance for infant growth, development, and health.
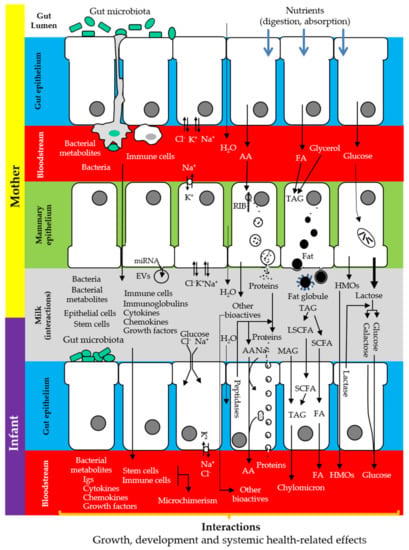
Figure 2.
Complementarity between the maternal gut, the breast, and the infant gut, illustrating the synthesis, transfer, use, and/or absorption of some of the components of human milk. AA: amino acids; EVs: extracellular vesicles; FA: fatty acids; HMOs: human milk oligosaccharides; LCFA: long chain fatty acids; MAG: monoacylglycerides; RIB: ribosomes; SCFA: short chain fatty acids; TAG: triacylglycerides. Adapted from [18].
The nutritional composition of human milk is quite conserved, but there are some nutrients (thiamine, riboflavin, vitamin B-6, vitamin B-12, choline, retinol, vitamin A, vitamin D, selenium, iodine, and fatty acids, including monounsaturated, n–6, and n–3 fatty acids) that are critical for infant development and health, and whose concentrations are highly affected by maternal diet or nutritional status [27,28,29,30,31,32]. In fact, the presence of these nutrients in milk is rapidly and/or substantially reduced by maternal deficit or depletion, and this situation has a negative impact on infant health and development. The concentrations of many of such nutrients are often below recommended levels in many populations. This situation is particularly worrying in low income countries, but it can be also observed in developed countries (e.g., omega-3 fatty acids, choline) [27,28,29,32,33,34]. This does not mean that breastfeeding is not the best feeding practice in these settings, but that the diet of the mother may require supplementation with some essential nutrients when deficiencies are observed. The concentrations of such nutrients can be restored following maternal supplementation, a practice that will benefit both maternal and infant health. This is in contrast with other nutrients, such as lactose, protein, calcium, or folate, which milk concentrations remain relatively largely unaffected by maternal diet [32]. However, in this case diet deficiencies will affect maternal health and, in the long term, they will indirectly affect infant’s health too. Unfortunately, research on how maternal diet affects the micro- and macronutrients’ content of human milk, both quantitatively and qualitatively, is surprisingly scarce. In this context, conflicting data have been obtained in relation to some minerals, such as iron or zinc, which are essential for a wide array of physiological functions [35,36], and which deficiencies in the mother‒infant dyad are particularly concerning [37,38]. Some studies have reported that their milk concentrations were associated with maternal dietary intake and/or supplementation, others have found no relationships with these factors and, finally, others have provided ambiguous results [39,40,41,42,43]. More recently, a positive correlation between maternal diet and the milk concentration of both minerals was detected only when the total intake (including diet and supplementation) was taken into account [44]. This and other studies have found that milk levels of these two minerals could be below the adequate intake for infants in a relatively high percentage of samples [34,35,36,37,38,39,40,41,42,43,44]. Positive associations between dietary iron and zinc intakes and their respective serum levels among premenopausal women have been described [45]. However, contrary to iron, zinc concentrations are higher in human milk than in the serum of a breastfeeding woman [46], and, therefore, it is possible that maternal factors may influence the concentration of these two minerals in a different way [44]. Iron and zinc are good examples to highlight that large scale studies are required to provide accurate information about levels, bioavailable forms, and critical factors that are associated with the concentrations of micronutrients in human milk.
Among human milk nutrients, fatty acids are extremely sensible to maternal nutrition [47]. This is a very relevant issue because of the critical roles of fatty acids for growth (energy source), cardiovascular health, and organ development, playing a keystone role in the composition of the cell membranes, neurodevelopment, immunomodulation, and as modulators of the milk microbiota [29,48,49]. There is evidence showing that, overall, the profile of fatty acids in human milk has changed with modern dietary practices and, at least in some cases, this may lead to profiles that are not adequate for an optimal neurological development of the infant [29]. In addition, intra-individual and inter-individual differences in the diet may account for differences in the composition and structure of milk fat and, as a result, in the concentration of the many fat-associated small molecules [50], and in the structure and composition of fat globules and their membranes, which represent complex colloidal assemblies with plenty of bioactive molecules that may be very relevant for infant outcomes [51]. The structure of the milk fat globule membrane is essential for correct delivery of nutritional and functional lipids and proteins to the infant gut [52] and, as a consequence, for many biological processes relevant for infant health [49,51,53]. This must be taken into account in Human Milk Banks, since processing of donor milk (e.g., pasteurization, freeze/unthaw) may disrupt the structure of the milk fat globules and their membranes and lead to relevant losses in the biological functions.
Oxidative stress is associated with respiratory and intestinal diseases in infants, especially among preterm ones [54]. Interestingly, human milk contains a wide spectrum of endogenous antioxidant compounds, including superoxide dismutase, glutathione peroxidase, catalase, glutathione, or melatonin [55]. The concentration of these antioxidants in human milk is adapted to gestational age, providing higher levels to infants with lower degree of maturation [56]. As a consequence, the level of oxidative stress in premature neonates fed with breast milk is lower than that observed among formula-fed ones [57,58]. Phytochemicals (polyphenols, carotenoids) in the maternal diet provides an exogenous source of antioxidants, since they are also transferred to milk [59]. Although many of the human milk phytochemicals have a wide range of biological activities [60,61,62], most of the research interest has been focused on polyphenols because of their potential as antioxidants. Through their antioxidant activities, diet-related human milk polyphenols might contribute to the benefits that human milk provides to this infant population [63]. Many different flavonoids (epicatechin, epicatechin gallate, epigallocatechin gallate, naringenin, kaempferol, hesperetin, quercetin) can be present, simultaneously, in the same human milk sample [64]. The flavonoid profile depends almost exclusively on maternal diet and rapid shifts can be observed depending on the intake of polyphenol-rich foods (berries, soya, dark chocolate, olive oil, wine) [59]. Polyphenol and other antioxidants are becoming increasingly popular as food supplements among the general population, but their actual impact in human milk composition and in infant outcomes is rather speculative and far from being fully elucidated because of the lack of well-designed epidemiological studies and human clinical trials.
The relevance of maternal diet for infant health has been highlighted by the use of murine models, since pups cross-fostered to dams that received a high-fat diet during lactation develop insulin resistance [65], obesity [66], and hypertension [67], which are symptoms typically related to metabolic syndrome. Metabolic diseases are associated with significant shifts in the gut microbiome [68]. In turn, the maternal gut microbiota can metabolize many food components. Some of the resulting metabolites, such as short chain fatty acids (SCFAs), will be absorbed and transferred to the milk [69]. It must be highlighted that there is an intense traffic of bacterial metabolites from the maternal gut to the breast or to maternal tissues (from which they can later reach the breast) through the bloodstream during late pregnancy and throughout the lactation period [70]. Many of these metabolites, including retinoid compounds, fatty acids, or aryl hydrocarbon receptor ligands, are transferred to the infant gut through breastfeeding and play relevant roles for immune development and infant health (including the metabolism of xenobiotic chemicals) [70,71]. At present, 8.3% of the human milk metabolites included in the Human Metabolome Database (http://www.hmdb.ca) are exclusively produced by certain microbes and, most probably, their presence is the result of the activity of the maternal gut microbiota.
2.3. Is There an Entero-Mammary Circulation of Immune and Microbial Cells?
Lactating mammary glands are a part of the maternal‒mucosal immune system [72] and, as such, their immune status has relevant implications for infant development and health [73]. In addition to a wide spectrum of soluble immune factors (which will be discussed later), human milk also contains a variety of leukocytes, including neutrophils, macrophages, and lymphocytes [74]. Although they may transmigrate to infants and seem to play a key role in infant defense and immune training during early life [75], their exact origin and functions remain largely unknown. Interestingly, other types of leukocytes (basophils, eosinophils, mast cells) and platelets, which are usually related to inflammation, are rarely found [74]. The concentration of total leukocytes is higher in colostrum than in mature milk, probably reflecting the fact that tight junctions of the mammary epithelium are looser during the first days after birth [76].
Some decades ago, flow cytometry studies showed that the proportions of neutrophils, macrophages, and lymphocytes in human milk are approximately 80%, 15%, and 4%, respectively [77,78]. However, leukocytes subsets are different in the mammary gland and in the milk, a fact that indicates the existence of selective mechanisms of cell transmigration to milk. A recent study compared leukocyte populations in the mammary gland and milk from lactating mice [79]. T cells (mainly CD4+ and CD8+ T cells with a memory phenotype) constituted the largest leukocyte fraction in milk but were minority in the mammary gland. Inversely, dendritic cells (DCs) represented the largest one in the mammary gland but were minority in milk. In addition, the populations of myeloid and B cells were significantly lower in milk than in mammary tissue. In contrast, another study working with lactating mice found that most leukocytes in the milk bolus were myeloid cells [75]. In relation to CD4+ and CD8+ T cells, their accumulation in human milk has been reported previously [77,78,80,81,82]. On the basis of animal model studies, it has been speculated that through a high expression of the genes (Cldn3, Cldn7, Tjp1) involved in the biosynthesis of the tight junction (TJ) protein ZO-1 and in the regulation of claudin polymerization [83], CD8+ T cells (and maybe CD4+ T, γδ T, and NK cells) may selectively translocate through the TJ region of the mammary epithelium while preserving its integrity [79]. As a result, they would reach milk and migrate into neonatal lymphoid tissues [75] to reinforce cell-mediated immunity in the neonatal period [84]. Interestingly, colostrum leukocytes are able to adhere to the gut mucosa of premature baboons, to persist in the wall of the large intestine for, at least, 60 h after a feed, and to reach organs that are particularly interesting in the frame of the development of the immune system, such as the spleen or the liver [85]. This actually represents a mode of maternal microchimerism, and this issue will be discussed below. In contrast, IgA-producing cells are retained inside the mammary gland and this may be regulated by the chemokine CCL28 [86,87].
It is generally agreed that, during pregnancy and lactation, leukocytes reach the mammary gland, but the homing to the mammary environment remains an open question, and few works have investigated the attractant and adherence molecules required for their presence there [88]. The T and B cells present in the lactating mammary ecosystem display phenotypes that are quite different to those found in blood and have a homing profile similar to the corresponding cells located in the gut-associated lymphoid tissue (GALT) [75,77,82,89]. The lactating mammary gland is an effector site of the mucosal-associated lymphoid tissue (MALT) system; this implies that following antigens’ exposure in a mucosal inductive site (gastrointestinal or respiratory tracts), lymphocytes may migrate to the unexposed mammary mucosal surface. A previous study reported that the lactating breast compartment is more closely related to the gut mucosa than to that of the upper respiratory tract [82].
Pioneer animal studies revealed that B cells from the maternal gut have the ability to migrate in a specific manner to the mammary gland (the so-called “entero-mammary” pathway or link) during late pregnancy and lactation [87,90,91,92,93,94,95,96]. The homing effect exerted by the mammary gland during such stages is not completely elucidated, but it seems to be under hormonal control [97]; more recent studies have shown the involvement of the interactions between the mucosal vascular addressin MadCAM-1, the gut homing receptor α4β7 integrin [98], and the mucosa-associated CCL28/CCR10 link [99]. Once in the mammary glands, the recruited B cells produce IgAs specifically against the pathogens prevalent in the mother’s environment. In 1975, and while studying the in vitro production of antibodies by human colostral cells after exposure to O antigens from Escherichia coli, Ahlstedt et al. [100] suggested that the IgA-forming cells may had been homed to the mammary gland from the gastrointestinal tract after exposure to enteric bacteria. Subsequently, the same research group showed that oral administration of a non-pathogenic E. coli strain to pregnant women led to an abundant presence of cells producing IgAs specific against the O antigen of E. coli in colostrum samples [101]. Some years later, it was found that the administration of respiratory syncytial virus (RSV) to pregnant rabbits only led to the presence of anti-RSV IgAs in colostrum and milk when viral inoculation was performed by oral or transtracheal routes but not when it was made by an intravenous one [102,103]. The gut-associated IgA system seems to be in a permanent state of adaptation to and protection from the gut microbiota [104]. In fact, the simultaneous vertical transfer of maternal IgA-producing cells and microbes during early life constitutes an elegant mode of inducing a tailored-adaptation of the infant gut to the own mother’s microbiota, fostering the development and maintenance of a healthy gut microbiota, enhancing the barrier function in the enteric environment, exerting immunomodulatory effects, and regulating gene expression of intestinal epithelial cells [105,106,107,108,109]. Studies with lactating mice and their offspring have revealed that the benefits of early exposure to the maternal assembly of IgAs persist into adulthood [89], and may condition allergic or autoimmune disease [110,111].
Recently, the existence of an immunologic entero-mammary system modulating the genetic tuning of gut immune responses and inflammatory disease susceptibility has been hypothesized on the basis of murine studies [112]. These authors suggested the existence of a narrow opportunity window (first days after birth) in which the level of bacterial IgA coating in milk inversely correlates with the number of colonic RORγ+ (transcription factors Helios or RORγ) regulatory T cells (Treg) that are induced. In turn, the number of these Treg cells is inversely correlated with the level of IgA in the adult intestine and, as a consequence, with the level of IgA in milk, which would be subsequently transmitted to the next generation. Therefore, this double-negative-feedback loop system would be vertically transmitted, but also serve as a mode of multi-generational matrilineal transmission of immunological information [113]. Intestinal Treg cells play crucial roles in the immune-regulation of host‒pathogen interactions [114] and suppress inadequate innate and adaptive immune responses, promoting tolerance to harmless antigens [113]. The rapid increase in the rates of autoimmune diseases and allergies over the past decades cannot be explained only on the basis of genetic changes [115], and it has been suggested that it may be related to an alteration of maternal immunologic transmission [112].
Similarly to IgA responses to gut microbes, contact with food antigens present in the gastrointestinal tract (such as cow’s milk proteins (CMP)) also direct the specificity of IgAs found in human colostrum and milk [116,117]. These authors showed that high levels of CMP-specific IgAs in human milk conferred protection against cow’s milk allergy, while cow’s milk exclusion from the maternal diet provoked a down-regulation of the levels of such specific IgAs in milk. They also reported that the IgA antibody repertoire towards CMP present in the lactating mammary environment is different to that in the systemic immune compartment, and hypothesized that it could be due to a distinct homing effect [117]. Overall, their results suggest, first, that there is an entero-mammary pathway for food-specific antibody-secreting cells, and second, that elimination diets during pregnancy and breastfeeding may result in decreased levels of protective milk-specific IgAs, and might be related with the development of food allergies in the infant population.
The finding that DCs are well represented among the leukocyte population of the lactating mammary gland [79] is also interesting in relation to the presence of bacterial cells in milk. DCs are among the professional antigen-presenting cells and play a pivotal role in the regulation and development of innate and adaptive immune responses. After sensing environmental signals (including microbes) from the mucosal surfaces (e.g., gut), they mature, adapt their cytokine microenvironment accordingly, and migrate from the peripheral tissues of the MALT system to secondary lymphoid organs in order to induce and coordinate effector T cell responses. Seminal studies showed DCs and CD18+ cells are able to open the tight junctions between gut epithelial cells, uptake non-invasive bacteria from the gut lumen, and transport them to other MALT sites [118,119,120]. The integrity of the epithelial barrier is preserved because of expression of tight junction proteins (ZO-1, occludin, claudin) by the DCs during the penetration of the dendrites [119]. Because of the fact that the lactating mammary gland is a member of the MALT system, Martín et al. [121] hypothesized the existence of an entero-mammary circulation of specific members of the gut microbiota upon complex interactions with gut immune and epithelial cells. Subsequently, it was shown that some lactic acid bacteria strains isolated from human milk had the ability to translocate across a Caco-2 cell monolayer through a DC-mediated mechanism [122,123]. Since then, several in vitro, animal, and human studies have confirmed the plausibility of a physiological bacterial translocation during late pregnancy and lactation [124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139].
In the last 20 years, several studies have shown that human milk harbors a low biomass microbiota under physiological conditions and that milk, maternal feces, and/or infant feces share some bacteria [24,140,141,142,143]. The ultimate origin (skin, oro-nasopharynx, gastrointestinal tract, vagina) of the microbes present in human milk is difficult to elucidate because of the complex and dynamic interactions between different body locations of the mother‒infant dyad. This is particularly true for the facultative anaerobic bacteria that are usually present in almost any mucosal/epithelial surface of the human body, including those of the digestive, upper respiratory, and genitourinary tracts and/or the skin. They include some of the most abundant bacteria in human milk, such as staphylococci, streptococci, corynebacteria, and cutibacteria. In addition, some gut-associated strict anaerobes have also been isolated or their DNA has been detected in human milk (Faecalibacterium, Roseburia, Bifidobacterium, Blautia, Bacteroides, Parabacteroides, etc.) [24,131,142,144,145]. Their presence is much more difficult to explain on the basis of mere contaminations arising from aerobic body locations, such as the skin or or nasopharynx. Interestingly, the vertical transmission of bifidobacterial communities through breastfeeding also leads to the inheritance of bifidobacterial phages [146]. In fact, bacteriophages represent the vast majority of the human milk viruses and have the ability to modulate the bacterial ecology in the mammary gland and in the infant gut [147]. In relation to the relevance of human milk as a source and/or promoter of Bifidobacterium populations in the infant gut, a recent study found that this genus was negatively correlated with the resistome (antibiotic resistance genes) and mobilome (mobile genetic elements) structure, which was shared between human milk and feces within each recruited mother‒infant pair [148]. These authors also found a link between enrichment of resistance genes and antibiotic consumption by mothers, a practice that may have enduring effects on the infant resistome. Similarly, early termination of breastfeeding (which also strongly affects the composition of the infant gut microbiota [149]) was also associated to a resistome enrichment. Therefore, the reduction of antibiotic resistance genes in the infant gut may constitute a new benefit of breastfeeding, especially in the context of the increasing public health threat posed by multi-resistant bacteria.
The potential translocation of intestinal bacteria has generally been related with disease states characterized by alterations in tight junction proteins that favor an increased gut leakiness and the transmigration of microbes and microbial products from the gut lumen into the bloodstream [150,151,152]. However, this process also happens, albeit at a low rate, in healthy hosts and involves the transmigration of mutualistic or beneficial bacteria, including Bacteroides, lactobacilli, and bifidobacteria [153,154,155,156,157]. It has been hypothesized that this phenomenon may be linked to physiological immunotraining or immunomodulation of the host [125]. Some studies have suggested that this may be a highly selective process, since some bacterial strains appear to mediate their own translocation without simultaneous translocation of other gut bacteria from the same host [158,159]. Some of the transient body adaptations that take place during pregnancy and lactation are compatible with an intensification of gut permeability and bacterial translocation during such stages [19,24,25]. Recently, Gosalbes et al. [22] investigated how late pregnancy conditions affect gene expression by the gut microbiota and found that during this life stage, some of the bacterial populations seemed to reach a stationary phase. This observation has relevant physiological implications, since growth phase and motility greatly increase the possibility of bacterial translocation [22].
Bacterial cells administrated per os may reach locations previously considered sterile environments, including tumors [158,159]. This fact has been explained on the basis of three circumstances that occur in solid tumors: increased nutrient availability (necrotic tumor core), increased vascularization, and suppressed immune surveillance [160,161,162,163,164,165]. It must be highlighted that the similar or identical circumstances also coincide in the mammary gland during late pregnancy: immune-privileged status to tolerate the fetus; intense angiogenesis and vasculogenesis; and a rich nutrient environment, since pre-colostrum is already produced during the last third of pregnancy. These circumstances may also contribute to explain the selective tropism of some maternal bacterial species for the mammary gland during late pregnancy and lactation. More studies are needed to elucidate the mechanisms by which non-pathogenic bacterial strains may translocate in certain life stages. This may provide new strategies for manipulating the infant gut microbiome when it may be required to improve infant health.
3. From the Breast to the Infant Gut
3.1. Milk: A Multi-Functional Food
Human milk is the best infant feeding option from the nutritional point of view. In addition, this biological fluid contains an impressive array of bioactive components (and it can be anticipated that many others will be discovered in the coming years), making it a very complex and non-imitable functional food. It has become evident that this wealth of nutrients and bioactive ingredients contribute, in a synergic manner, to growth (e.g., insulin-like growth factor (IGF) superfamily, somatostatin, calcitonin), organ development and tissue repair (e.g., stem cells, epidermal growth factor (EGF)), protection against inflammation and infections (e.g., acquired and innate immune factors and cells), enhancement of the mucosal barriers (e.g., SCFAs), immune maturation and tolerance (e.g., transforming growth factor β (TGFβ)), neurodevelopment (e.g., unsaturated fatty acids, neurotransmitters, neuronal growth factors (BDNF, GDNF)), regulation of metabolism and body composition (e.g., adiponectin, ghrelin, leptin), regulation of the vascular system (e.g., vascular endothelial growth factor (VEGF)), prevention of anemia (e.g., erythropoietin), acquisition and development of the microbiota (e.g., oligosaccharides, bacterial cells), and many others. In conclusion, there is no human physiological function that is not, directly or indirectly, influenced by human milk in early life. The complex composition of milk means that it connects with any of the body axes (gut‒brain and gut‒elsewhere axes, hypothalamic–pituitary–adrenal (HPA) axis), intervening in their programming and function. Epidemiological and clinical studies have provided overwhelming evidence that there is a consensus in recognizing that breastfeeding provides relevant short- and long-term benefits for health, both at the individual and at the community levels [16,17].
The contributions of human milk to infant health are the result of interactions among many milk components, and it must be highlighted that most of them (as those cited above) play multifunctional roles. The concentration of many of them is dynamic and, often, adapted to suit infant requirements (age, gestational age at birth, time of the day, environment). In addition, some of them show a high degree of inter-individual variability, and the factors determining their concentrations remain poorly known. Such a complexity makes very difficult, if not impossible, an accurate assessment of the individual and collective roles of human milk components for infant health; in fact, in most cases, this task has remained elusive to scientists, and we are just in the beginning of understanding the functions of individual molecules or cells.
3.2. The Complex Case of Human Milk Oligosaccharides
Human milk oligosaccharides (HMOs) are a good example of the multi-functionality of many human milk compounds, but also of the difficulties in unveiling their actual impact on mammary or infant health. At present, approximately 200 different HMOs have been identified, which vary in size, structure, and complexity [166]. They can act as prebiotics, reduce binding of pathogenic microorganisms to the gut cells, and modulate host-epithelial immune responses among other functions, although the effects are often highly structure-specific or depend on the amount and proportion of certain HMOs [167]. Their effects may be not restricted to the gut, since some HMO structures may be absorbed in the gut and, in fact, some of them have already been found in the infant bloodstream [168,169]. The HMO profile of milk varies substantially among women, and the genetic and environmental factors responsible for the inter-individual variability are poorly understood. The peculiar structure of most HMOs makes them difficult to synthesize and/or commercialize from a technical or economical point of view. Therefore, most studies have been restricted to in vitro or animal model experiments, but there is a lack of well-designed and controlled infant studies. In addition, HMOs have mostly been investigated as individual items, when, in contrast, they are present as a diverse and interacting mix in milk and in the infant gut.
HMOs also interact with other milk compounds. It has been recently found that there is a direct interaction between HMOs and DCs, whereby tolerogenic DCs adopt a regulatory function that induces Treg expansion [170]. In addition, HMOs are able to inhibit LPS-induced pro-inflammatory responses through suppression of LPS-induced maturation of DCs [170]. HMOs selectively stimulate the growth of beneficial bacteria (e.g., bifidobacteria) in the infant gut. As a result of the fermentative activity of these and other bacteria on HMOS, there is an increase of SCFAs in the gut environment. In turn, both bacteria and SCFAs exert a strong influence on DCs phenotype and function [171,172,173,174]. SCFAs and other bacterial metabolites are essential for the cross-talk between the microbiota, the GALT, and the enteric nervous system, while direct interactions between gut DCs and the vagus nerve have also been demonstrated [2]. In relation to the nervous system, some of the bacteria whose growth is promoted by HMOs (including species that have been isolated from human milk) have the ability to participate in the biosynthesis of neurotransmitters (serotonin, dopamine) that have been detected in human milk [175,176,177], and whose presence deserves more research attention. This complex and personalized network of interactions, involving only a few of the human milk ingredients, serves to illustrate that novel combinations of methodologies are required to decipher the multifaceted interactions between them and their collective impact on infant health.
3.3. Role of Human Milk in Immunological Tolerance
While the importance of breast milk in infectious disease prevention is undisputable, there is uncertainty regarding its role in tolerance induction and allergy prevention [178,179,180,181]. However, recent evidence is accumulating and showing there is a potential of breastmilk for the latter as well that might be critically dependent on the maternal gut‒breast milk‒offspring axis. Two ways by which breast milk can promote tolerance induction in offspring can be distinguished: (a) by promoting immune regulation mechanisms in a non-antigen specific way, and (b) by inducing antigen specific immune tolerance.
In relation to the importance of breast milk for non-antigen specific immune regulation in offspring, and as stated above, breast milk has a major impact on child microbiota seeding and shaping [141,182]. This effect is most probably key in setting immune regulation in offspring, as largely illustrated in observational studies that associate microbiota composition and metabolites production in the first months of life with later allergy risk [183,184,185]. Currently, there are still major gaps of knowledge to be able to exploit the known impact of breast milk on tolerance induction: how to modulate microbiota shaping properties of mother’s milk and what is the ideal microbiota in early life for tolerance in the long term. In addition, and as already exposed in Section 2.2., an elegant study in mice uncovered the mechanisms of transgenerational effect of the maternal gut‒mammary axis on the levels of regulatory Tregs in offspring [112].
Regarding the importance of breast milk in the induction of antigen specific immune tolerance, the absorption of allergens in maternal gut and their shedding in maternal milk might be a key player in dictating allergy risk in offspring. The reports of allergic reactions in exclusively breastfed children demonstrate that the very low amounts of allergens in human milk are sensed by the developing immune system [186]. These observations do not allow a conclusion, however, on whether sensitization occurred through breast milk. Indeed, other sensitizing ways may have occurred, since the consumption of food in a household results in the presence of dietary allergen in dust, which may also lead to sensitization through the infant’s skin [187,188]. A tempting hypothesis to explore is that allergens in human milk might instruct the immune system and induce tolerance. This hypothesis has been widely demonstrated in rodent experimental models [189,190,191,192,193,194,195,196]. Importantly, mechanistic analyses have uncovered that the oral administration of allergen in early life is not sufficient to induce immune tolerance (in contrast to what is observed in adults) and that some maternal milk immune modulators are essential for oral tolerance induction in the developing immune system. Such key milk-derived immune modulators were identified as TGFβ [190], vitamin A [193], IGF-1 [196], and IgG [191,195]. They were found to act at the level of promoting gut barrier and controlling antigen transfer across the gut, antigen presentation by dendritic cells, and induction of regulatory T cells. Today, there is no existing study that formally demonstrates that maternal diet derived allergen influences allergy risk in offspring through its shedding in milk. Only one birth cohort study has analyzed the association between allergen content in human milk itself and allergy risk [197]. Although limited in size, the study showed that having the egg antigen ovalbumin (OVA) in human milk was associated with a four-fold reduction of egg allergy prevalence by the age of 2.5 [197]. Importantly, one cannot conclude on observational studies addressing the association between allergen consumption during lactation and allergy susceptibility in offspring, as multiple reports have shown the unpredictability of allergen content in breast milk. About half of lactating mothers do not secrete allergen in breast milk, even after ingestion of a well-controlled amount of egg, peanut, milk, or wheat [198,199,200]. What is even more surprising is that women consuming an egg exclusion diet were as likely to have detectable egg allergens (OVA and ovomucoid) in breast milk as women with an unmodified healthy diet [200,201,202].
In conclusion, while there is strong experimental support for a role of maternal diet-derived allergens in milk as key for oral tolerance induction in the developing system, much research is needed to elucidate fundamental questions. How is allergen transferred from the gut to the mammary gland? What controls the shedding of diet-derived allergen into maternal milk? Which compartment/configuration in milk make maternal diet-derived antigen the most suitable for tolerance induction? What is the role of micro-vesicles (see Section 3.4) and antigen-presenting cells in these processes? Which immune modulators in human milk are essential co-factors to induce oral tolerance to allergen that are transferred to the neonate through breast milk? How can we modulate those co-factors?
3.4. “Emerging” Components of Human Milk: Secreted Extracellular Vesicles and Mirnas
Human milk contains some biologically active entities that have been discovered in the last years. Secreted extracellular vesicles (EVs) are small particles (30–1000 nm) that are produced and exocytosed by many cell types throughout the body. They are packaged with a variety of regulatory molecules from the parent cell, including lipids, proteins, messenger RNA, and microRNAs (miRNAs) that participate in intercellular communication. EVs have attracted extensive research since their discovery in 1983 [203], because the presence of such cargo renders them as therapeutics and diagnosis targets [204]. EVs can be classified into exosomes or microvesicles on the basis of their size [205], although the methodologies currently available do not allow a clear discrimination between them [206].
EVs have been identified in milk from many mammalian species [207], including human milk [208,209]. Proteomic analysis of EVs from human milk has revealed the existence of a “novel” functional proteome within these particles, which is distinct to that typically found in milk [210]. In fact, this study identified 633 proteins in human milk-derived EVs that had not been identified in human milk previously, and they included proteins involved in the control of inflammatory signaling pathways and in the regulation of cell growth. The membrane of EVs protects enclosed molecules from digestion and, therefore, they can be delivered intact to the infant gut [211,212]. Gut epithelial cells are able to uptake milk EVs and deliver some of their molecules (e.g., miRNA) to the nucleus. In addition, it has also been suggested that EVs or their content may be transferred into the blood stream, thereby contributing to programming cellular events in distant tissues [209].
In vitro studies have demonstrated that human milk EVs promote the growth of intestinal epithelial cells [213] and protect them against oxidative stress [214], while it seems that they display a broad immunomodulatory function, including the activation of Treg cells [208]. The concentration of EVs may change through lactation, since milk collected within the first week after birth has higher concentration of exosomes than milk collected two months after birth [215]. In this context, it is interesting to note that EVs present in preterm and term milk exosomes show different peptide patterns, with an up-regulation of lactotransferrin and lactadherin in EVs from preterm milk, which are responsible for increased rates of cellular growth [216]. This fact has led to the hypothesis that human milk EVs or some of their components may have future applications for the prevention or treatment of necrotizing enterocolitis in preterm neonates [216,217].
Most of the research with human milk exosomes has targeted miRNAs, which are small non-coding RNA molecules implied in post-transcriptional regulation of target mRNA. Several abundant miRNAs (e.g., let-7a, let-7b, let-7f, miR-148a) are shared by the milk EVs of different mammalian species [207]. These evolutionarily conserved molecules are involved in regulation of cell growth and immune functions, which are absolutely crucial for the newborn. One study focused on other mammalian species showed that the miRNAs profile of colostrum is more associated with immune regulation, while that of mature milk was more directed to metabolism [218]. In contrast to EVs-associated peptides, the composition of those miRNAs that are highly abundant in human milk EVs is similar in milk from women delivering term and pre-term neonates [211,212]. An alteration of the miRNAs composition may have long-term consequences for health, since it has been shown that the miRNA profile of milk EVs from women with type-1 diabetes is different from that of healthy women, and some of the miRNAs that are differentially-expressed among diabetic women are able to induce the expression of pro-inflammatory cytokines by human monocytes [219].
Overall, studies performed so far indicate a relevant role of EVs and miRNAs in the development of the neonatal gastrointestinal tract and immune system and, probably, in other breastfeeding-associated health outcomes [220]. However, similarly to other human milk components, more studies are required to elucidate their roles and their interactions with other milk ingredients.
3.5. The Others That You Carry Inside: Immune Cells, Stem Cells, and Microchimerism
Different studies have provided evidence supporting the idea that some eukaryotic cells present in human milk may be transferred from the gut mucosa to the tissues of the infant [76,85,221,222,223]. Most of this postnatal maternal microchimerism probably occurs during early lactation when the permeability of both the mammary epithelium and the infant gut epithelium and the concentration of immune and epithelial cells are highest [224]. Little is known about the milk cells that may be involved in microchimerism but, in addition to mature immune cells, they include stem cells. Human milk harbors a whole hierarchy of stem and progenitor stem cells that account for up to 6% of the human cells in human milk and that englobe haemopoietic, mesenchymal, and neuroepithelial lineages [225,226,227]. Because of their properties, human milk stem cells are candidates for microchimerism in the infant tissue [225,228]. Maternal stem cells do not express MHC antigens and, therefore, they can be easily tolerated in infant tissues [224]. In fact, human milk stem cells seem to be able to go across the gut wall of nursed mice and rabbits and reach the bloodstream and, subsequently, different organs where they become functional [229,230,231,232]. More recently, it was found that breast milk stem cells were capable of reaching the brain, where they settled and differentiated into neuronal and glial cell types [233]. Previously, it was found that some stem cells of human milk are able to differentiate in vitro into a variety of cell types, including neurons [227,234].
Independently of the implied cells, breastfeeding-driven maternal microchimerism can complement pregnancy-related bidirectional microchimerism [235,236], contributing to the development of tolerance and to training and maturation of the infant immune system [223], and probably to many other relevant functions that are unknown at present. Given the diversity of cell populations in human milk (from memory lymphocytes and professional antigen-presenting cells to embryonic and mesenchymal stem cells), more research is required to elucidate how milk microchimeric cells may influence health and disease during life.
4. Conclusions
From the Industrial Revolution onwards, the rates of women that initiate breastfeeding and, especially, that of infants that are exclusively breastfed for their first six months of life, as recommended by the World Health Organization, started to fall, and the decrease was sharpest from the late 1800s, coinciding with the rapid urbanization of the societies in industrial countries. This trend is also observed in developing countries. However, here we have shown that human milk provides a complex nutritional, immunological, neuroendocrine, and microbiological integration between the mother and the infant, which is specifically tailored to the environment and requirements of the mother‒infant dyad. The axis established first between the maternal gut and breast and, then, between the breast and the infant gut has had, and will have, a paramount role in our existence as a species, serving as a pivotal node interacting with any other human axis and programming health for life. The complexity of the composition of human milk, together with the vast complexity of the interactions among all human milk components, renders this biological fluid a scientific enigma that we are unveiling at a very slow pace. As stated by Bode [166], such complexity “cannot be mimicked in artificial infant formula and provides yet another powerful reason to protect, promote, and support breastfeeding”.
Funding
This work was supported by the University of Western Australia and the Larsson Rosenquist Foundation, and by project PID2019-105606RB-I00 from the Ministerio de Ciencia e Innovación (Spain).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Gershon, M.D. The enteric nervous system: A second brain. Hosp. Pract. (1995) 1999, 34, 31–32, 35–38, 41–42, passim. [Google Scholar] [CrossRef] [PubMed]
- Goehler, L.E.; Gaykema, R.P.; Nguyen, K.T.; Lee, J.E.; Tilders, F.J.; Maier, S.F.; Watkins, L.R. Interleukin-1beta in immune cells of the abdominal vagus nerve: A link between the immune and nervous systems? J. Neurosci. 1999, 19, 2799–2806. [Google Scholar] [CrossRef]
- Banks, W.A. Evidence for a cholecystokinin gut-brain axis with modulation by bombesin. Peptides 1980, 1, 347–351. [Google Scholar] [CrossRef]
- Cooke, K.R.; Hill, G.R.; Gerbitz, A.; Kobzik, L.; Martin, T.R.; Crawford, J.M.; Brewer, J.P.; Ferrara, J.L. Hyporesponsiveness of donor cells to lipopolysaccharide stimulation reduces the severity of experimental idiopathic pneumonia syndrome: Potential role for a gut-lung axis of inflammation. J. Immunol. 2000, 165, 6612–6619. [Google Scholar] [CrossRef] [PubMed]
- Towfigh, S.; Heisler, T.; Rigberg, D.A.; Hines, O.J.; Chu, J.; McFadden, D.W.; Chandler, C. Intestinal ischemia and the gut-liver axis: An in vitro model. J. Surg. Res. 2000, 88, 160–164. [Google Scholar] [CrossRef] [PubMed]
- Meijers, B.K.; Evenepoel, P. The gut-kidney axis: Indoxyl sulfate, p-cresyl sulfate and CKD progression. Nephrol. Dial. Transplant. 2011, 26, 759–761. [Google Scholar] [CrossRef] [PubMed]
- Saarialho-Kere, U. The gut-skin axis. J. Pediatr. Gastroenterol. Nutr. 2004, 39 (Suppl. 3), S734–S735. [Google Scholar] [CrossRef] [PubMed]
- Hvatum, M.; Kanerud, L.; Hällgren, R.; Brandtzaeg, P. The gut-joint axis: Cross reactive food antibodies in rheumatoid arthritis. Gut 2006, 55, 1240–1247. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Zhao, C.M. The possible existence of a gut-bone axis suggested by studies of genetically manipulated mouse models? Curr. Pharm. Des. 2011, 17, 1552–1555. [Google Scholar] [CrossRef] [PubMed]
- Polakof, S.; Míguez, J.M.; Soengas, J.L. Evidence for a gut-brain axis used by glucagon-like peptide-1 to elicit hyperglycaemia in fish. J. Neuroendocrinol. 2011, 23, 508–518. [Google Scholar] [CrossRef]
- Kim, M.; Platt, M.J.; Shibasaki, T.; Quaggin, S.E.; Backx, P.H.; Seino, S.; Simpson, J.A.; Drucker, D.J. GLP-1 receptor activation and Epac2 link atrial natriuretic peptide secretion to control of blood pressure. Nat. Med. 2013, 19, 567–575. [Google Scholar] [CrossRef]
- Carsetti, R.; Di Sabatino, A.; Rosado, M.M.; Cascioli, S.; Piano Mortari, E.; Milito, C.; Grimsholm, O.; Aranburu, A.; Giorda, E.; Paolo Tinozzi, F.; et al. Lack of gut secretory immunoglobulin A in memory B-cell dysfunction-associated disorders: A possible gut-spleen axis. Front. Immunol. 2020, 10, 2937. [Google Scholar] [CrossRef]
- Taghinezhad-S, S.; Keyvani, H.; Bermúdez-Humarán, L.G.; Donders, G.G.G.; Fu, X.; Mohseni, A.H. Twenty years of research on HPV vaccines based on genetically modified lactic acid bacteria: An overview on the gut-vagina axis. Cell. Mol. Life Sci. 2020, 26, 1–16. [Google Scholar] [CrossRef]
- Trujillo-Vargas, C.M.; Schaefer, L.; Alam, J.; Pflugfelder, S.C.; Britton, R.A.; de Paiva, C.S. The gut-eye-lacrimal gland-microbiome axis in Sjögren Syndrome. Ocul. Surf. 2020, 18, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.M.; Meyer, K.M.; Prince, A.L.; Aagaard, K.M. Impact of maternal nutrition in pregnancy and lactation on offspring gut microbial composition and function. Gut Microbes 2016, 7, 459–470. [Google Scholar] [CrossRef] [PubMed]
- U.S. Department of Health and Human Services. The Surgeon General’s Call to Action to Support Breastfeeding; U.S. Department of Health and Human Services, Office of the Surgeon General: Washington, DC, USA, 2011.
- Renfrew, M.; Pokhrel, S.; Quigley, M.; McCormick, F.; Fox-Rushby, J.; Dodds, R.; Duffy, S.; Trueman, P.; Williams., A. Preventing Disease and Saving Resources: The Potential Contribution of Increasing Breastfeeding Rates in the UK; UNICEF: London, UK, 2012. [Google Scholar]
- Weaver, L.T. Breast and gut: The relationship between lactating mammary function and neonatal gastrointestinal function. Proc. Nutr. Soc. 1992, 51, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Hammond, K.A. Adaptation of the maternal intestine during lactation. J. Mammary Gland. Biol. Neoplasia 1997, 2, 243–252. [Google Scholar] [CrossRef]
- Koren, O.; Goodrich, J.K.; Cullender, T.C.; Spor, A.; Laitinen, K.; Bäckhed, H.K.; Gonzalez, A.; Werner, J.J.; Angenent, L.T.; Knight, R.; et al. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 2012, 150, 470–480. [Google Scholar] [CrossRef] [PubMed]
- DiGiulio, D.B.; Callahan, B.J.; McMurdie, P.J.; Costello, E.K.; Lyell, D.J.; Robaczewska, A.; Sun, C.L.; Goltsman, D.S.; Wong, R.J.; Shaw, G.; et al. Temporal and spatial variation of the human microbiota during pregnancy. Proc. Natl. Acad. Sci. USA 2015, 112, 11060–11065. [Google Scholar] [CrossRef] [PubMed]
- Gosalbes, M.J.; Compte, J.; Moriano-Gutierrez, S.; Vallès, Y.; Jiménez-Hernández, N.; Pons, X.; Artacho, A.; Francino, M.P. Metabolic adaptation in the human gut microbiota during pregnancy and the first year of life. EBioMedicine 2019, 39, 497–509. [Google Scholar] [CrossRef]
- Prieto, R.M.; Ferrer, M.; Fe, J.M.; Rayó, J.M.; Tur, J.A. Morphological adaptive changes of small intestinal tract regions due to pregnancy and lactation in rats. Ann. Nutr. Metab. 1994, 38, 295–300. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.M. The origin of human milk bacteria: Is there a bacterial entero-mammary pathway during late pregnancy and lactation? Adv. Nutr. 2014, 5, 779–784. [Google Scholar] [CrossRef]
- Mira, A.; Rodríguez, J.M. The origin of human milk bacteria. In Prebiotics and Probiotics in Human Milk; McGuire, M., McGuire, M., Bode, L., Eds.; Academic Press: London, UK, 2017; pp. 349–364. [Google Scholar]
- Ballard, O.; Morrow, A.L. Human milk composition: Nutrients and bioactive factors. Pediatr. Clin. N. Am. 2013, 60, 49–74. [Google Scholar] [CrossRef] [PubMed]
- Valentine, C.J.; Wagner, C.L. Nutritional management of the breastfeeding dyad. Pediatr. Clin. N. Am. 2013, 60, 261–274. [Google Scholar] [CrossRef] [PubMed]
- Allen, L.H. B vitamins in breast milk: Relative importance of maternal status and intake, and effects on infant status and function. Adv. Nutr. 2012, 3, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Innis, S.M. Impact of maternal diet on human milk composition and neurological development of infants. Am. J. Clin. Nutr. 2014, 99, 734S–741S. [Google Scholar] [CrossRef]
- Leermakers, E.T.M.; Moreira, E.M.; Kiefte-de Jong, J.C.; Darweesh, S.K.L.; Visser, T.; Voortman, T.; Bautista, P.K.; Chowdhury, R.; Gorman, D.; Bramer, W.M.; et al. Effects of choline on health across the life course: A systematic review. Nutr. Rev. 2015, 73, 500–522. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, C.S. Maternal mineral and bone metabolism during pregnancy, lactation, and post-weaning recovery. Physiol. Rev. 2016, 96, 449–547. [Google Scholar] [CrossRef]
- Dror, D.K.; Allen, L.H. Overview of nutrients in human milk. Adv. Nutr. 2018, 9 (Suppl. 1), 278S–294S. [Google Scholar] [CrossRef]
- Wallace, T.C.; Blusztajn, J.K.; Caudill, M.A.; Klatt, K.C.; Natker, E.; Zeisel, S.H.; Zelman, K.M. Choline: The underconsumed and underappreciated essential nutrient. Nutr. Today 2018, 53, 240–253. [Google Scholar] [CrossRef] [PubMed]
- Daniels, L.; Gibson, R.S.; Diana, A.; Haszard, J.J.; Rahmannia, S.; Luftimas, D.E.; Hampel, D.; Shahab-Ferdows, S.; Reid, M.; Melo, L.; et al. Micronutrient intakes of lactating mothers and their association with breast milk concentrations and micronutrient adequacy of exclusively breastfed Indonesian infants. Am. J. Clin. Nutr. 2019, 110, 391–400. [Google Scholar] [CrossRef]
- Eussen, S.; Alles, M.; Uijterschout, L.; Brus, F.; van der Horst-Graat, J. Iron intake and status of children aged 6–36 months in Europe: A systematic review. Ann. Nutr. Metab. 2015, 66, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Ackland, L.; Michalczyk, A. Zinc and infant nutrition. Arch. Biochem. Biophys. 2016, 611, 51–57. [Google Scholar] [CrossRef]
- Petry, N.; Olofin, I.; Boy, E.; Donahue Angel, M.; Rohner, F. The effect of low dose iron and zinc intake on child micronutrient status and development during the first 1000 days of life: A systematic review and meta-Analysis. Nutrients 2016, 30, 773. [Google Scholar] [CrossRef]
- Bzikowska-Jura, A.; Czerwonogrodzka-Senczyna, A.; Olędzka, G.; Szostak-Węgierek, D.; Weker, H.; Wesołowska, A. Maternal nutrition and body composition during breastfeeding: Association with human milk composition. Nutrients 2018, 10, 1379. [Google Scholar] [CrossRef]
- Choi, Y.K.; Kim, J.M.; Lee, J.E.; Cho, M.S.; Kang, B.S.; Choi, H.; Kim, Y. Association of maternal diet with zinc, copper, and iron concentrations in transitional human milk produced by Korean mothers. Clin. Nutr. Res. 2016, 5, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Hannan, M.A.; Faraji, B.; Tanguma, J.; Longoria, N.; Rodriguez, R.C. Maternal milk concentration of zinc, iron, selenium, and iodine and its relationship to dietary intakes. Biol. Trace Elem. Res. 2009, 127, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Yalçin, S.S.; Baykan, A.; Yurdakök, K.; Yalçin, S.; Gücüş, A.I. The factors that affect milk-to-serum ratio for iron during early lactation. J. Pediatr. Hematol. Oncol. 2009, 31, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Mahdavi, R.; Nikniaz, L.; Gayemmagami, S.J. Association between zinc, copper, and iron concentrations in breast milk and growth of healthy infants in Tabriz, Iran. Biol. Trace Elem. Res. 2010, 135, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Silvestre, M.D.; Lagarda, M.J.; Farre, R.; Martinez-Costa, C.; Brines, J.; Molina, A.; Clemente, G. A study of factors that may influence the determination of copper, iron, and zinc in human milk during sampling and in sample individuals. Biol. Trace Elem. Res. 2000, 76, 217–227. [Google Scholar] [CrossRef]
- Bzikowska-Jura, A.; Sobieraj, P.; Michalska-Kacymirow, M.; Wesołowska, A. Investigation of iron and zinc concentrations in human milk in correlation to maternal factors: An observational pilot study in Poland. Nutrients 2021, 13, 303. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.; Booth, A.; Szymlek-Gay, E.A.; Gibson, R.S.; Bailey, K.B.; Irving, D.; Nowson, C.; Riddell, L. Associations between dietary iron and zinc intakes, and between biochemical iron and zinc status in women. Nutrients 2015, 7, 2983–2999. [Google Scholar] [CrossRef] [PubMed]
- Nakamori, M.; Ninh, N.X.; Isomura, H.; Yoshiike, N.; Hien, V.T.T.; Nhug, B.T.; Van Nhien, N.; Nakano, T.; Khan, N.C.; Yamamoto, S. Nutritional status of lactating mothers and their breast milk concentration of iron, zinc and copper in rural Vietnam. J. Nutr. Sci. Vitaminol. 2009, 55, 338–345. [Google Scholar] [CrossRef]
- Innis, S.M. Maternal nutrition, genetics and human milk lipids. Curr. Nutr. Rep. 2013, 2, 151–158. [Google Scholar] [CrossRef]
- Azagra-Boronat, I.; Tres, A.; Massot-Cladera, M.; Franch, A.; Castell, M.; Guardiola, F.; Pérez-Cano, F.J.; Rodríguez-Lagunas, M.J. Associations of breast milk microbiota, immune factors, and fatty acids in the rat mother-offspring pair. Nutrients 2020, 12, 319. [Google Scholar] [CrossRef] [PubMed]
- Ramiro-Cortijo, D.; Singh, P.; Liu, Y.; Medina-Morales, E.; Yakah, W.; Freedman, S.D.; Martin, C.R. Breast milk lipids and fatty acids in regulating neonatal intestinal development and protecting against intestinal injury. Nutrients 2020, 12, 534. [Google Scholar] [CrossRef] [PubMed]
- Demmelmair, H.; Koletzko, B. Variation of metabolite and hormone contents in human milk. Clin. Perinatol. 2017, 44, 151–164. [Google Scholar] [CrossRef]
- Lee, H.; Padhi, E.; Hasegawa, Y.; Larke, J.; Parenti, M.; Wang, A.; Hernell, O.; Lönnerdal, B.; Slupsky, C. Compositional dynamics of the milk fat globule and its role in infant development. Front. Pediatr. 2018, 6, 313. [Google Scholar] [CrossRef]
- Contarini, G.; Povolo, M. Phospholipids in milk fat: Composition, biological and technological significance, and analytical strategies. Int. J. Mol. Sci. 2013, 14, 2808–2831. [Google Scholar] [CrossRef] [PubMed]
- Petersen, E.N.; Chung, H.W.; Nayebosadri, A.; Hansen, S.B. Kinetic disruption of lipid rafts is a mechanosensor for phospholipase D. Nat. Commun. 2016, 7, 13873. [Google Scholar] [CrossRef]
- Thibeault, D.W. The precarious antioxidant defenses of the preterm infant. Am. J. Perinatol. 2000, 17, 167–182. [Google Scholar] [CrossRef] [PubMed]
- Gila-Diaz, A.; Arribas, S.M.; Algara, A.; Martín-Cabrejas, M.A.; López de Pablo, Á.L.; Sáenz de Pipaón, M.; Ramiro-Cortijo, D. A review of bioactive factors in human breastmilk: A focus on prematurity. Nutrients 2019, 11, 1307. [Google Scholar] [CrossRef]
- Gila-Diaz, A.; Herranz Carrillo, G.; Cañas, S.; Saenz de Pipaón, M.; Martínez-Orgado, J.A.; Rodríguez-Rodríguez, P.; López de Pablo, Á.L.; Martin-Cabrejas, M.A.; Ramiro-Cortijo, D.; Arribas, S.M. Influence of maternal age and gestational age on breast milk antioxidants during the first month of lactation. Nutrients 2020, 12, 2569. [Google Scholar] [CrossRef]
- Friel, J.K.; Martin, S.M.; Langdon, M.; Herzberg, G.R.; Buettner, G.R. Milk from mothers of both premature and full-term infants provides better antioxidant protection than does infant formula. Pediatr. Res. 2002, 51, 612–618. [Google Scholar] [CrossRef]
- Ledo, A.; Arduini, A.; Asensi, M.A.; Sastre, J.; Escrig, R.; Brugada, M.; Aguar, M.; Saenz, P.; Vento, M. Human milk enhances antioxidant defenses against hydroxyl radical aggression in preterm infants. Am. J. Clin. Nutr. 2009, 89, 210–215. [Google Scholar] [CrossRef]
- Tsopmo, A. Phytochemicals in human milk and their potential antioxidative protection. Antioxidants 2018, 7, 32. [Google Scholar] [CrossRef] [PubMed]
- Landete, J.M. Updated knowledge about polyphenols: Functions, bioavailability, metabolism, and health. Crit. Rev. Food Sci. Nutr. 2012, 52, 936–948. [Google Scholar] [CrossRef]
- Pérez-Cano, F.J.; Massot-Cladera, M.; Rodríguez-Lagunas, M.J.; Castell, M. Flavonoids affect host-microbiota crosstalk through TLR modulation. Antioxidants 2014, 3, 649–670. [Google Scholar] [CrossRef]
- Koudoufio, M.; Desjardins, Y.; Feldman, F.; Spahis, S.; Delvin, E.; Levy, E. Insight into polyphenol and gut microbiota crosstalk: Are their metabolites the key to understand protective effects against metabolic disorders? Antioxidants 2020, 9, 982. [Google Scholar] [CrossRef]
- Tsopmo, A.; Friel, J.K. Human milk has anti-oxidant properties to protect premature infants. Curr. Pediatr. Rev. 2007, 3, 45–51. [Google Scholar] [CrossRef]
- Song, B.J.; Jouni, Z.E.; Ferruzzi, M.G. Assessment of phytochemical content in human milk during different stages of lactation. Nutrition 2013, 29, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Du, Q.; Hosoda, H.; Umekawa, T.; Kinouchi, T.; Ito, N.; Miyazato, M.; Kangawa, K.; Ikeda, T. Postnatal weight gain induced by overfeeding pups and maternal high-fat diet during the lactation period modulates glucose metabolism and the production of pancreatic and gastrointestinal peptides. Peptides 2015, 70, 23–31. [Google Scholar] [CrossRef]
- Tsuduki, T.; Kitano, Y.; Honma, T.; Kijima, R.; Ikeda, I. High dietary fat intake during lactation promotes development of diet-induced obesity in male offspring of mice. J. Nutr. Sci. Vitaminol. (Tokyo) 2013, 59, 384–392. [Google Scholar] [CrossRef]
- Khan, I.Y. A high-fat diet during rat pregnancy or suckling induces cardiovascular dysfunction in adult offspring. AJP Regul. Integr. Comp. Physiol. 2004, 288, R127–R133. [Google Scholar] [CrossRef] [PubMed]
- Tilg, H.; Kaser, A. Gut microbiome, obesity, and metabolic dysfunction. J. Clin. Investig. 2011, 121, 2126–2132. [Google Scholar] [CrossRef]
- Priyadarshini, M.; Thomas, A.; Reisetter, A.C.; Scholtens, D.M.; Wolever, T.M.; Josefson, J.L.; Layden, B.T. Maternal short-chain fatty acids are associated with metabolic parameters in mothers and newborns. Transl. Res. 2014, 164, 153–157. [Google Scholar] [CrossRef]
- Gay, M.; Koleva, P.T.; Slupsky, C.M.; Toit, E.D.; Eggesbo, M.; Johnson, C.C.; Wegienka, G.; Shimojo, N.; Campbell, D.E.; Prescott, S.L.; et al. Worldwide variation in human milk metabolome: Indicators of breast physiology and maternal lifestyle? Nutrients 2018, 10, 1151. [Google Scholar] [CrossRef]
- Prentice, P.; Ong, K.K.; Schoemaker, M.H.; van Tol, E.A.; Vervoort, J.; Hughes, I.A.; Acerini, C.L.; Dunger, D.B. Breast milk nutrient content and infancy growth. Acta Paediatr. 2016, 105, 641–647. [Google Scholar] [CrossRef]
- Brandtzaeg, P. The mucosal immune system and its integration with the mammary glands. J Pediatr. 2010, 156 (Suppl. 2), S8–S15. [Google Scholar] [CrossRef] [PubMed]
- Hanson, L.A. Feeding and infant development breast-feeding and immune function. Proc. Nutr. Soc. 2007, 66, 384–396. [Google Scholar] [CrossRef]
- Goldman, A.S.; Goldblum, R.M. Transfer of maternal leukocytes to the infant by human milk. Curr. Top. Microbiol. Immunol. 1997, 222, 205–213. [Google Scholar] [CrossRef]
- Cabinian, A.; Sinsimer, D.; Tang, M.; Zumba, O.; Mehta, H.; Toma, A.; Sant’Angelo, D.; Laouar, Y.; Laouar, A. Transfer of maternal immune cells by breastfeeding: Maternal cytotoxic T lymphocytes present in breast milk localize in the Peyer’s patches of the nursed infant. PLoS ONE 2016, 11, e0156762. [Google Scholar] [CrossRef] [PubMed]
- Trend, S.; de Jong, E.; Lloyd, M.L.; Kok, C.H.; Richmond, P.; Doherty, D.A.; Simmer, K.; Kakulas, F.; Strunk, T.; Currie, A. Leukocyte populations in human preterm and term breast milk identified by multicolour flow cytometry. PLoS ONE 2015, 10, e0135580. [Google Scholar] [CrossRef]
- Bertotto, A.; Castellucci, G.; Fabietti, G.; Scalise, F.; Vaccaro, R. Lymphocytes bearing the T cell receptor gamma delta in human breast milk. Arch. Dis. Child. 1990, 65, 1274–1275. [Google Scholar] [CrossRef]
- Wirt, D.; Adkins, L.T.; Palkowetz, K.H.; Schmalstieg, F.C.; Goldman, A.S. Activated-memory T cells in human milk. Cytometry 1992, 13, 282–290. [Google Scholar] [CrossRef]
- Ikebuchi, R.; Fujimoto, M.; Moriya, T.; Kusumoto, Y.; Kobayashi, K.; Tomura, M. T cells are the main population in mouse breast milk and express similar profiles of tight junction proteins as those in mammary alveolar epithelial cells. J. Reprod. Immunol. 2020, 140, 103137. [Google Scholar] [CrossRef] [PubMed]
- Sabbaj, S.; Ghosh, M.K.; Edwards, B.H.; Leeth, R.; Decker, W.D.; Goepfert, P.A.; Aldrovandi, G.M. Breast milk-derived antigen-specific CD8+ T cells: An extralymphoid effector memory cell population in humans. J. Immunol. 2005, 174, 2951–2956. [Google Scholar] [CrossRef]
- Kourtis, A.P.; Ibegbu, C.C.; Theiler, R.; Xu, Y.X.; Bansil, P.; Jamieson, D.J.; Lindsay, M.; Butera, S.; Duerr, A. Breast milk CD4+ T cells express high levels of C chemokine receptor 5 and CXC chemokine receptor 4 and are preserved in HIV-infected mothers receiving highly active antiretroviral therapy. J. Infect. Dis. 2007, 195, 965–972. [Google Scholar] [CrossRef]
- Tuaillon, E.; Valea, D.; Becquart, P.; Al Tabaa, Y.; Meda, N.; Bollore, K.; Van de Perre, P.; Vendrell, J.P. Human milk-derived B cells: A highly activated switched memory cell population primed to secrete antibodies. J. Immunol. 2009, 182, 7155–7162. [Google Scholar] [CrossRef] [PubMed]
- Baumgartner, H.K.; Rudolph, M.C.; Ramanathan, P.; Burns, V.; Webb, P.; Bitler, B.G.; Stein, T.; Kobayashi, K.; Neville, M.C. Developmental expression of claudins in the mammary gland. J. Mammary Gland. Biol. Neoplasia 2017, 22, 141–157. [Google Scholar] [CrossRef]
- Bandrick, M.; Ariza-Nieto, C.; Baidoo, S.K.; Molitor, T.W. Colostral antibody-mediated and cell-mediated immunity contributes to innate and antigen-specific immunity in piglets. Dev. Comp. Immunol. 2014, 43, 114–120. [Google Scholar] [CrossRef]
- Jain, L.; Vidyasagar, D.; Xanthou, M.; Ghai, V.; Shimada, S.; Blend, M. In vivo distribution of human milk leucocytes after ingestion by newborn baboons. Arch. Dis. Child. 1989, 64, 930–933. [Google Scholar] [CrossRef] [PubMed]
- Lazarus, N.H.; Kunkel, E.J.; Johnston, B.; Wilson, E.; Youngman, K.R.; Butcher, E.C. A common mucosal chemokine (mucosae-associated epithelial chemokine/CCL28) selectively attracts IgA plasmablasts. J. Immunol. 2003, 170, 3799–3805. [Google Scholar] [CrossRef]
- Wilson, E.; Butcher, E.C. CCL28 controls immunoglobulin (Ig)A plasma cell accumulation in the lactating mammary gland and IgA antibody transfer to the neonate. J. Exp. Med. 2004, 200, 805–809. [Google Scholar] [CrossRef]
- Goldman, A.S. Future research in the immune system of human milk. J. Pediatr. 2019, 206, 274–279. [Google Scholar] [CrossRef] [PubMed]
- Bertotto, A.; Gerli, R.; Castellucci, G.; Scalise, F.; Vaccaro, R. Human milk lymphocytes bearing the gamma/delta T-cell receptor are mostly delta TCS1-positive cells. Immunology 1991, 74, 360–361. [Google Scholar] [PubMed]
- Roux, M.E.; McWilliams, M.; Phillips-Quagliata, J.M.; Weisz-Carrington, P.; Lamm, M.E. Origin of IgA-secreting plasma cells in the mammary gland. J. Exp. Med. 1977, 146, 1311–1322. [Google Scholar] [CrossRef]
- Hanson, L.A.; Ahlstedt, S.; Carlsson, B.; Fallstrom, S.P.; Kaijser, B.; Lindblad, B.S.; Akerlund, A.S.; Edén, C.S. New knowledge in human milk immunoglobulin. Acta Paediatr. Scand. 1978, 67, 577–582. [Google Scholar] [CrossRef] [PubMed]
- Jelliffe, D.B.; Jelliffe, E.F. Breast milk and infection. Lancet 1981, 2, 419. [Google Scholar] [CrossRef]
- Roux, M.E.; McWilliams, M.; Phillips-Quagliata, J.M.; Lamm, M.E. Differentiation pathway of Peyer’s patch precursors of IgA plasma cells in the secretory immune system. Cell. Immunol. 1981, 61, 141–153. [Google Scholar] [CrossRef]
- Hanson, L.A.; Ahlstedt, S.; Andersson, B.; Cruz, J.R.; Dahlgren, U.; Fällström, S.P.; Porras, O.; Svanborg Edén, C.; Söderström, T.; Wettergren, B. The immune response of the mammary gland and its significance for the neonate. Ann. Allergy 1984, 53 (6 Pt 2), 576–582. [Google Scholar] [PubMed]
- Dahlgren, U.I.; Ahlstedt, S.; Hanson, L.A. The localization of the antibody response in milk or bile depends on the nature of the antigen. J. Immunol. 1987, 138, 1397–1402. [Google Scholar] [PubMed]
- Nathavitharana, K.A.; Catty, D.; McNeish, A.S. IgA antibodies in human milk: Epidemiological markers of previous infections? Arch. Dis. Child. Fetal Neonatal Ed. 1994, 71, F192–F197. [Google Scholar] [CrossRef] [PubMed]
- Weisz-Carrington, P.; Roux, M.E.; McWilliams, M.; Phillips-Quagliata, J.M.; Lamm, M.E. Organ and isotype distribution of plasma cells producing specific antibody after oral immunization: Evidence for a generalized secretory immune system. J. Immunol. 1979, 123, 1705–1708. [Google Scholar] [PubMed]
- Tanneau, G.M.; Hibrand-Saint Oyant, L.; Chevaleyre, C.C.; Salmon, H.P. Differential recruitment of T- and IgA B-lymphocytes in the developing mammary gland in relation to homing receptors and vascular addressins. J. Histochem. Cytochem. 1999, 47, 1581–1592. [Google Scholar] [CrossRef]
- Morteau, O.; Gerard, C.; Lu, B.; Ghiran, S.; Rits, M.; Fujiwara, Y.; Law, Y.; Distelhorst, K.; Nielsen, E.M.; Hill, E.D.; et al. An indispensable role for the chemokine receptor CCR10 in IgA antibody-secreting cell accumulation. J. Immunol. 2008, 181, 6309–6315. [Google Scholar] [CrossRef]
- Ahlstedt, S.; Carlsson, B.; Hanson, L.A.; Goldblum, R.M. Antibody production by human colostral cells. I. Immunoglobulin class, specificity, and quantity. Scand. J. Immunol 1975, 4, 535–539. [Google Scholar] [CrossRef] [PubMed]
- Goldblum, R.M.; Ahlstedt, S.; Carlsson, B.; Hanson, L.A.; Jodal, U.; Lidin-Janson, G.; Sohl-Akerlund, A. Antibody-forming cells in human colostrum after oral immunisation. Nature 1975, 257, 797–798. [Google Scholar] [CrossRef]
- Peri, B.A.; Theodore, C.M.; Losonsky, G.A.; Fishaut, J.M.; Rothberg, R.M.; Ogra, P.L. Antibody content of rabbit milk and serum following inhalation or ingestion of respiratory syncytial virus and bovine serum albumin. Clin. Exp. Immunol. 1982, 48, 91–101. [Google Scholar] [PubMed]
- Ogra, P.L.; Losonsky, G.A.; Fishaut, M. Colostrum-derived immunity and maternal-neonatal interaction. Ann. N. Y. Acad. Sci. 1983, 409, 82–95. [Google Scholar] [CrossRef]
- Hapfelmeier, S.; Lawson, M.A.; Slack, E.; Kirundi, J.K.; Stoel, M.; Heikenwalder, M.; Cahenzli, J.; Velykoredko, Y.; Balmer, M.L.; Endt, K.; et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 2010, 328, 1705–1709. [Google Scholar] [CrossRef] [PubMed]
- Rogier, E.W.; Frantz, A.L.; Bruno, M.E.; Wedlund, L.; Cohen, D.A.; Stromberg, A.J.; Kaetzel, C.S. Lessons from mother: Long-term impact of antibodies in breast milk on the gut microbiota and intestinal immune system of breastfed offspring. Gut Microbes 2014, 5, 663–668. [Google Scholar] [CrossRef]
- Harris, N.L.; Spoerri, I.; Schopfer, J.F.; Nembrini, C.; Merky, P.; Massacand, J.; Urban, J.F., Jr.; Lamarre, A.; Burki, K.; Odermatt, B.; et al. Mechanisms of neonatal mucosal antibody protection. J. Immunol. 2006, 177, 6256–6262. [Google Scholar] [CrossRef]
- Koch, M.A.; Reiner, G.L.; Lugo, K.A.; Kreuk, L.S.; Stanbery, A.G.; Ansaldo, E.; Seher, T.D.; Ludington, W.B.; Barton, G.M. Maternal IgG and IgA antibodies dampen mucosal T helper cell responses in early life. Cell 2016, 165, 827–841. [Google Scholar] [CrossRef] [PubMed]
- Bunker, J.J.; Bendelac, A. IgA responses to microbiota. Immunity 2018, 49, 211–224. [Google Scholar] [CrossRef]
- Gopalakrishna, K.P.; Macadangdang, B.R.; Rogers, M.B.; Tometich, J.T.; Firek, B.A.; Baker, R.; Ji, J.; Burr, A.; Ma, C.; Good, M.; et al. Maternal IgA protects against the development of necrotizing enterocolitis in preterm infants. Nat. Med. 2019, 25, 1110–1115. [Google Scholar] [CrossRef]
- Rogier, E.W.; Frantz, A.L.; Bruno, M.E.; Wedlund, L.; Cohen, D.A.; Stromberg, A.J.; Kaetzel, C.S. Secretory antibodies in breast milk promote long-term intestinal homeostasis by regulating the gut microbiota and host gene expression. Proc. Natl. Acad. Sci. USA 2014, 111, 3074–3079. [Google Scholar] [CrossRef] [PubMed]
- Zinkernagel, R.M. Maternal antibodies, childhood infections, and autoimmune diseases. N. Engl. J. Med. 2001, 345, 1331–1335. [Google Scholar] [CrossRef]
- Ramanan, D.; Sefik, E.; Galván-Peña, S.; Wu, M.; Yang, L.; Yang, Z.; Kostic, A.; Golovkina, T.V.; Kasper, D.L.; Mathis, D.; et al. An immunologic mode of multigenerational transmission governs a gut Treg setpoint. Cell 2020, 181, 1276–1290.e13. [Google Scholar] [CrossRef]
- Minton, K. Mother knows best (about gut Treg cells). Nat. Rev. Immunol. 2020, 20, 405. [Google Scholar] [CrossRef]
- Schiering, C.; Krausgruber, T.; Chomka, A.; Fröhlich, A.; Adelmann, K.; Wohlfert, E.A.; Pott, J.; Griseri, T.; Bollrath, J.; Hegazy, A.N.; et al. The alarmin IL-33 promotes regulatory T-cell function in the intestine. Nature 2014, 513, 564–568. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.H.; Gregersen, P.K. Genomics and the multifactorial nature of human autoimmune disease. N. Engl. J. Med. 2011, 365, 1612–1623. [Google Scholar] [CrossRef]
- Järvinen, K.M.; Westfall, J.E.; Seppo, M.S.; James, A.K.; Tsuang, A.J.; Feustel, P.J.; Sampson, H.A.; Berin, C. Role of maternal elimination diets and human milk IgA in the development of cow’s milk allergy in the infants. Clin. Exp. Allergy 2014, 44, 69–78. [Google Scholar] [CrossRef]
- Seppo, A.E.; Savilahti, E.M.; Berin, M.C.; Sampson, H.A.; Järvinen, K.M. Breast milk IgA to foods has different epitope specificity than serum IgA-Evidence for entero-mammary link for food-specific IgA? Clin. Exp. Allergy 2017, 47, 1275–1284. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Torres, A.; Jones-Carson, J.; Baumler, A.J.; Falkow, S.; Valdivia, R.; Brown, W.; Le, M.; Berggren, R.; Parks, W.T.; Fang, F.C. Extraintestinal dissemination of salmonella by CD18-expressing phagocytes. Nature 1999, 401, 804–808. [Google Scholar] [CrossRef]
- Rescigno, M.; Urbano, M.; Valzasina, B.; Francolini, M.; Rotta, G.; Bonasio, R.; Granucci, F.; Kraehenbuhl, J.P.; Ricciardi-Castagnoli, P. Dendritic cells express tight junction proteins and penetrate gut epithelial monolayers to sample bacteria. Nat. Immunol. 2001, 2, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 2004, 303, 1662–1665. [Google Scholar] [CrossRef]
- Martín, R.; Langa, S.; Reviriego, C.; Jiménez, E.; Marín, M.L.; Olivares, M.; Boza, J.; Jiménez, J.; Fernández, L.; Xaus, J.; et al. The commensal microflora of human milk: New perspectives for food bacteriotherapy and probiotics. Trends Food Sci. Technol. 2004, 15, 121–127. [Google Scholar] [CrossRef]
- Langa, S. Interactions between Lactic acid Bacteria, Intestinal Epithelial Cells and Immune Cells. Development of In Vitro Models. Ph.D. Thesis, Complutense University of Madrid, Madrid, Spain, 2006. [Google Scholar]
- Langa, S.; Maldonado-Barragán, A.; Delgado, S.; Martín, R.; Martín, V.; Jiménez, E.; Ruíz-Barba, J.L.; Mayo, B.; Connor, R.I.; Suárez, J.E.; et al. Characterization of Lactobacillus salivarius CECT 5713, a strain isolated from human milk: From genotype to phenotype. Appl. Microbiol. Biotechnol. 2012, 94, 1279–1287. [Google Scholar] [CrossRef]
- Martin, R.; Jimenez, E.; Olivares, M.; Marin, M.L.; Fernandez, L.; Xaus, J.; Rodriguez, J.M. Lactobacillus salivarius CECT 5713, a potential probiotic strain isolated from infant feces and breast milk of a mother-child pair. Int. J. Food Microbiol. 2006, 112, 35–43. [Google Scholar] [CrossRef]
- Perez, P.F.; Dore, J.; Leclerc, M.; Levenez, F.; Benyacoub, J.; Serrant, P.; Segura-Roggero, I.; Schiffrin, E.J.; Donnet-Hughes, A. Bacterial imprinting of the neonatal immune system: Lessons from maternal cells? Pediatrics 2007, 119, e724–e732. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, E.; Fernandez, L.; Maldonado, A.; Martin, R.; Olivares, M.; Xaus, J.; Rodriguez, J.M. Oral administration of Lactobacillus strains isolated from breast milk as an alternative for the treatment of infectious mastitis during lactation. Appl. Environ. Microbiol. 2008, 74, 4650–4655. [Google Scholar] [CrossRef] [PubMed]
- Abrahamsson, T.R.; Sinkiewicz, G.; Jakobsson, T.; Fredrikson, M.; Bjorksten, B. Probiotic lactobacilli in breast milk and infant stool in relation to oral intake during the first year of life. J. Pediatr. Gastroenterol. Nutr. 2009, 49, 349–354. [Google Scholar] [CrossRef]
- Arroyo, R.; Martin, V.; Maldonado, A.; Jimenez, E.; Fernandez, L.; Rodriguez, J.M. Treatment of infectious mastitis during lactation: Antibiotics versus oral administration of lactobacilli isolated from breast milk. Clin. Infect. Dis. 2010, 50, 1551–1558. [Google Scholar] [CrossRef] [PubMed]
- Makino, H.; Kushiro, A.; Ishikawa, E.; Muylaert, D.; Kubota, H.; Sakai, T.; Oishi, K.; Martin, R.; Ben Amor, K.; Oozeer, R.; et al. Transmission of intestinal Bifidobacterium longum subsp. longum strains from mother to infant, determined by multilocus sequencing typing and amplified fragment length polymorphism. Appl. Environ. Microbiol. 2011, 77, 6788–6793. [Google Scholar] [CrossRef] [PubMed]
- Martin, V.; Maldonado-Barragan, A.; Moles, L.; Rodriguez-Banos, M.; Campo, R.D.; Fernandez, L.; Rodriguez, J.M.; Jimenez, E. Sharing of bacterial strains between breast milk and infant feces. J. Hum. Lact. 2012, 28, 36–44. [Google Scholar] [CrossRef] [PubMed]
- Jost, T.; Lacroix, C.; Braegger, C.P.; Rochat, F.; Chassard, C. Vertical mother-neonate transfer of maternal gut bacteria via breastfeeding. Environ. Microbiol. 2014, 16, 2891–2904. [Google Scholar] [CrossRef]
- Schanche, M.; Avershina, E.; Dotterud, C.; Øien, T.; Storrø, O.; Johnsen, R.; Rudi, K. High-resolution analyses of overlap in the microbiota between mothers and their children. Curr. Microbiol. 2015, 71, 283–290. [Google Scholar] [CrossRef]
- Milani, C.; Mancabelli, L.; Lugli, G.A.; Duranti, S.; Turroni, F.; Ferrario, C.; Mangifesta, M.; Viappiani, A.; Ferretti, P.; Gorfer, V.; et al. Exploring vertical transmission of bifidobacteria from mother to child. Appl. Environ. Microbiol. 2015, 81, 7078–7087. [Google Scholar] [CrossRef]
- Treven, P.; Mrak, V.; Bogovič Matijašić, B.; Horvat, S.; Rogelj, I. Administration of probiotics Lactobacillus rhamnosus GG and Lactobacillus gasseri K7 during pregnancy and lactation changes mouse mesenteric lymph nodes and mammary gland microbiota. J. Dairy Sci. 2015, 98, 2114–2128. [Google Scholar] [CrossRef]
- Fernandez, L.; Cardenas, N.; Arroyo, R.; Manzano, S.; Jimenez, E.; Martin, V.; Rodriguez, J.M. Prevention of infectious mastitis by oral administration of Lactobacillus salivarius PS2 during late pregnancy. Clin. Infect. Dis. 2016, 62, 568–573. [Google Scholar] [CrossRef]
- De Andrés, J.; Jiménez, E.; Chico-Calero, I.; Fresno, M.; Fernández, L.; Rodríguez, J.M. Physiological translocation of lactic acid bacteria during pregnancy contributes to the composition of the milk microbiota in mice. Nutrients 2018, 10, 14. [Google Scholar] [CrossRef] [PubMed]
- Asnicar, F.; Manara, S.; Zolfo, M.; Truong, D.T.; Scholz, M.; Armanini, F.; Ferretti, P.; Gorfer, V.; Pedrotti, A.; Tett, A.; et al. Studying vertical microbiome transmission from mothers to infants by strain-level metagenomic profiling. mSystems 2017, 2, e00164-16. [Google Scholar] [CrossRef]
- Murphy, K.; Curley, D.; O’Callaghan, T.F.; O’Shea, C.A.; Dempsey, E.M.; O’Toole, P.W.; Ross, R.P.; Ryan, C.A.; Stanton, C. The composition of human milk and infant fecal microbiota over the first three months of life: A pilot study. Sci. Rep. 2017, 7, 40597. [Google Scholar] [CrossRef]
- Kordy, K.; Gaufin, T.; Mwangi, M.; Li, F.; Cerini, C.; Lee, D.J.; Adisetiyo, H.; Woodward, C.; Pannaraj, P.S.; Tobin, N.H.; et al. Contributions to human breast milk microbiome and enteromammary transfer of Bifidobacterium breve. PLoS ONE 2020, 15, e0219633. [Google Scholar] [CrossRef]
- Fernandez, L.; Langa, S.; Martin, V.; Maldonado, A.; Jimenez, E.; Martin, R.; Rodriguez, J.M. The human milk microbiota: Origin and potential roles in health and disease. Pharmacol. Res. 2013, 69, 1–10. [Google Scholar] [CrossRef]
- Pannaraj, P.S.; Li, F.; Cerini, C.; Bender, J.M.; Yang, S.; Rollie, A.; Adisetiyo, H.; Zabih, S.; Lincez, P.J.; Bittinger, K.; et al. Association between breast milk bacterial communities and establishment and development of the infant gut microbiome. JAMA Pediatr. 2017, 171, 647–654. [Google Scholar] [CrossRef]
- Fernández, L.; Rodríguez, J.M. Human milk microbiota: Origin and potential uses. Nestle Nutr. Inst. Workshop Ser. 2020, 94, 75–85. [Google Scholar] [CrossRef]
- Fernández, L.; Pannaraj, P.S.; Rautava, S.; Rodríguez, J.M. The microbiota of the human mammary ecosystem. Front. Cell. Infect. Microbiol. 2020, 10, 586667. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Jiménez, E.; Heilig, H.; Fernandez, L.; Marin, M.L.; Zoetendal, E.G.; Rodríguez, J.M. Isolation of bifidobacteria from breast milk and assessment of the bifidobacterial population by PCR-DGGE and qRTi-PCR. Appl. Environ. Microbiol. 2009, 75, 965–969. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, E.; de Andres, J.; Manrique, M.; Pareja-Tobes, P.; Tobes, R.; Martinez-Blanch, J.F.; Codoner, F.M.; Ramon, D.; Fernandez, L.; Rodriguez, J.M. Metagenomic analysis of milk of healthy and mastitis-suffering women. J. Hum. Lact. 2015, 31, 406–415. [Google Scholar] [CrossRef]
- Duranti, S.; Lugli, G.A.; Mancabelli, L.; Armanini, F.; Turroni, F.; James, K.; Ferretti, P.; Gorfer, V.; Ferrario, C.; Milani, C.; et al. Maternal inheritance of bifidobacterial communities and bifidophages in infants through vertical transmission. Microbiome 2017, 5, 66. [Google Scholar] [CrossRef]
- Pannaraj, P.S.; Ly, M.; Cerini, C.; Saavedra, M.; Aldrovandi, G.M.; Saboory, A.A.; Johnson, K.M.; Pride, D.T. Shared and distinct features of human milk and infant stool viromes. Front. Microbiol. 2018, 9, 1162. [Google Scholar] [CrossRef] [PubMed]
- Pärnänen, K.; Karkman, A.; Hultman, J.; Lyra, C.; Bengtsson-Palme, J.; Larsson, D.; Rautava, S.; Isolauri, E.; Salminen, S.; Kumar, H.; et al. Maternal gut and breast milk microbiota affect infant gut antibiotic resistome and mobile genetic elements. Nat. Commun. 2018, 9, 3891. [Google Scholar] [CrossRef]
- Bäckhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 2015, 17, 852. [Google Scholar] [CrossRef]
- Berg, R.D. Bacterial translocation from the gastrointestinal tract. Adv. Exp. Med. Biol. 1999, 473, 11–30. [Google Scholar] [CrossRef] [PubMed]
- Balzan, S.; de Almeida Quadros, C.; de Cleva, R.; Zilberstein, B.; Cecconello, I. Bacterial translocation: Overview of mechanisms and clinical impact. J. Gastroenterol. Hepatol. 2007, 22, 464–471. [Google Scholar] [CrossRef]
- Ravisankar, S.; Tatum, R.; Garg, P.M.; Herco, M.; Shekhawat, P.S.; Chen, Y.H. Necrotizing enterocolitis leads to disruption of tight junctions and increase in gut permeability in a mouse model. BMC Pediatr. 2018, 18, 372. [Google Scholar] [CrossRef]
- Yamazaki, S.; Machii, K.; Tsuyuki, S.; Momose, H.; Kawashima, T.; Ueda, K. Immunological responses to monoassociated Bifidobacterium longum and their relation to prevention of bacterial invasion. Immunology 1985, 56, 43–50. [Google Scholar] [PubMed]
- Sedman, P.C.; Macfie, J.; Sagar, P.; Mitchell, C.J.; May, J.; Mancey-Jones, B.; Johnstone, D. The prevalence of gut translocation in humans. Gastroenterology 1994, 107, 643–649. [Google Scholar] [CrossRef]
- Berg, R.D. Bacterial translocation from the gastrointestinal tract. Trends Microbiol. 1995, 3, 149–154. [Google Scholar] [CrossRef]
- Rodriguez, A.V.; Baigori, M.D.; Alvarez, S.; Castro, G.R.; Oliver, G. Phosphatidylinositol-specific phospholipase C activity in Lactobacillus rhamnosus with capacity to translocate. FEMS Microbiol. Lett. 2001, 204, 33–38. [Google Scholar] [CrossRef]
- Dasanayake, A.P.; Li, Y.; Wiener, H.; Ruby, J.D.; Lee, M.J. Salivary Actinomyces naeslundii genospecies 2 and Lactobacillus casei levels predict pregnancy outcomes. J. Periodontol. 2005, 76, 171–177. [Google Scholar] [CrossRef]
- Cronin, M.; Morrissey, D.; Rajendran, S.; El Mashad, S.M.; van Sinderen, D.; O’Sullivan, G.C.; Tangney, M. Orally administered bifidobacteria as vehicles for delivery of agents to systemic tumors. Mol. Ther. 2010, 18, 1397–1407. [Google Scholar] [CrossRef]
- Danino, T.; Prindle, A.; Kwong, G.A.; Skalak, M.; Li, H.; Allen, K.; Hasty, J.; Bhatia, S.N. Programmable probiotics for detection of cancer in urine. Sci. Transl. Med. 2015, 7, 289ra84. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Fu, G.F.; Fan, Y.R.; Liu, W.H.; Liu, X.J.; Wang, J.J.; Xu, G.X. Bifidobacterium adolescentis as a delivery system of endostatin for cancer gene therapy: Selective inhibitor of angiogenesis and hypoxic tumor growth. Cancer Gene Ther. 2003, 10, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.A.; Shabahang, S.; Timiryasova, T.M.; Zhang, Q.; Beltz, R.; Gentschev, I.; Goebel, W.; Szalay, A.A. Visualization of tumors and metastases in live animals with bacteria and vaccinia virus encoding light-emitting proteins. Nat. Biotechnol. 2004, 22, 313–320. [Google Scholar] [CrossRef]
- Zhao, M.; Yang, M.; Li, X.M.; Jiang, P.; Baranov, E.; Li, S.; Xu, M.; Penman, S.; Hoffman, R.M. Tumor-targeting bacterial therapy with amino acid auxotrophs of GFP-expressing Salmonella typhimurium. Proc. Natl. Acad. Sci. USA 2005, 102, 755–760. [Google Scholar] [CrossRef]
- Zhao, M.; Yang, M.; Ma, H.; Li, X.; Tan, X.; Li, S.; Yang, Z.; Hoffman, R.M. Targeted therapy with a Salmonella Typhimurium leucine-arginine auxotroph cures orthotopic human breast tumors in nude mice. Cancer Res. 2006, 66, 7647–7652. [Google Scholar] [CrossRef]
- Nagakura, C.; Hayashi, K.; Zhao, M.; Yamauchi, K.; Yamamoto, N.; Tsuchiya, H.; Tomita, K.; Bouvet, M.; Hoffman, R.M. Efficacy of a genetically-modified Salmonella typhimurium in an orthotopic human pancreatic cancer in nude mice. Anticancer. Res. 2009, 29, 1873–1878. [Google Scholar] [PubMed]
- Morrissey, D.; O’Sullivan, G.C.; Tangney, M. Tumour targeting with systemically administered bacteria. Curr. Gene Ther. 2010, 10, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Bode, L. Human milk oligosaccharides: Next-generation functions and questions. Nestle Nutr. Inst. Workshop Ser. 2019, 90, 191–201. [Google Scholar] [CrossRef]
- Bode, L. Human milk oligosaccharides: Structure and functions. Nestle Nutr. Inst. Workshop Ser. 2020, 94, 115–123. [Google Scholar] [CrossRef]
- Ruhaak, L.R.; Stroble, C.; Underwood, M.A.; Lebrilla, C.B. Detection of milk oligosaccharides in plasma of infants. Anal. Bioanal. Chem. 2014, 406, 5775–5784. [Google Scholar] [CrossRef]
- Goehring, K.C.; Kennedy, A.D.; Prieto, P.A.; Buck, R.H. Direct evidence for the presence of human milk oligosaccharides in the circulation of breastfed infants. PLoS ONE 2014, 9, e101692. [Google Scholar] [CrossRef]
- Xiao, L.; van De Worp, W.R.; Stassen, R.; van Maastrigt, C.; Kettelarij, N.; Stahl, B.; Blijenberg, B.; Overbeek, S.A.; Folkerts, G.; Garssen, J.; et al. Human milk oligosaccharides promote immune tolerance via direct interactions with human dendritic cells. Eur. J. Immunol. 2019, 49, 1001–1014. [Google Scholar] [CrossRef]
- 171. Hart, A.L.; Lammers, K.; Bigidi, P.; Vitali, B.; Rizzello, F.; Gionchetti, P.; Campieri, M.; Kamm, M.A.; Knight, S.C.; Stagg, A.J. Modulation of human dendritic cell phenotype and function by probiotic bacteria. Gut 2004, 53, 1602–1609. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.-K.; Lee, C.-G.; So, J.-S.; Chae, C.-S.; Hwang, J.-S.; Sahoo, A.; Nam, J.H.; Rhee, J.H.; Hwang, K.C.; Im, S.H. Generation of regulatory dendritic cells and CD4+Foxp3+ T cells by probiotics administration suppresses immune disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 2159–2164. [Google Scholar] [CrossRef] [PubMed]
- Nastasi, C.; Candela, M.; Bonefeld, C.M.; Geisler, C.; Hansen, M.; Krejsgaard, T.; Biagi, E.; Andersen, M.H.; Brigidi, P.; Ødum, N.; et al. The effect of short-chain fatty acids on human monocyte-derived dendritic cells. Sci. Rep. 2015, 5, 16148. [Google Scholar] [CrossRef] [PubMed]
- Kaisar, M.M.M.; Pelgrom, L.R.; van der Ham, A.J.; Yazdanbakhsh, M.; Everts, B.; Pelgrom, L. Butyrate conditions human dendritic cells to prime type 1 regulatory T cells via both histone deacetylase inhibition and G protein-coupled receptor 109A signaling. Front. Immunol. 2017, 8, 1429. [Google Scholar] [CrossRef]
- Ertl, T.; Sulyok, E.; Hartmann, G. Dopamine and prolactin content of the human milk. Pediatr. Res. 1992, 32, 617. [Google Scholar] [CrossRef][Green Version]
- Chiba, T.; Maeda, T.; Tairabune, T.; Tomita, T.; Sanbe, A.; Takeda, R.; Kikuchi, A.; Kudo, K. Analysis of serotonin concentrations in human milk by high-performance liquid chromatography with fluorescence detection. Biochem. Biophys. Res. Commun. 2017, 485, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Maeda, T.; Shioyama, A.; Tairabune, T.; Takeda, R.; Sanbe, A.; Kikuchi, A.; Kudo, K.; Chiba, T. Physiologic changes in serotonin concentrations in breast milk during lactation. Nutrition 2020, 79–80, 110969. [Google Scholar] [CrossRef]
- Victora, C.G.; Bahl, R.; Barros, A.J.; França, G.V.; Horton, S.; Krasevec, J.; Murch, S.; Sankar, M.J.; Walker, N.; Rollins, N.C. Breastfeeding in the 21st century: Epidemiology, mechanisms, and lifelong effect. Lancet 2016, 387, 475–490. [Google Scholar] [CrossRef]
- Güngör, D.; Nadaud, P.; LaPergola, C.C.; Dreibelbis, C.; Wong, Y.P.; Terry, N.; Abrams, S.A.; Beker, L.; Jacobovits, T.; Järvinen, K.M.; et al. Infant milk-feeding practices and food allergies, allergic rhinitis, atopic dermatitis, and asthma throughout the life span: A systematic review. Am. J. Clin. Nutr. 2019, 109 (Suppl. 7), 772S–799S. [Google Scholar] [CrossRef]
- Greer, F.R.; Sicherer, S.H.; Burks, A.W.; Committee on Nutrition; Section on Allergy and Immunology. The effects of early nutritional interventions on the development of atopic disease in infants and children: The role of maternal dietary restriction, breastfeeding, hydrolyzed formulas, and timing of introduction of allergenic complementary foods. Pediatrics 2019, 143, e20190281. [Google Scholar] [CrossRef] [PubMed]
- de Silva, D.; Halken, S.; Singh, C.; Muraro, A.; Angier, E.; Arasi, S.; Arshad, H.; Beyer, K.; Boyle, R.; du Toit, G.; et al. Preventing food allergy in infancy and childhood: Systematic review of randomised controlled trials. Pediatr. Allergy Immunol. 2020, 31, 813–826. [Google Scholar] [CrossRef] [PubMed]
- Fehr, K.; Moossavi, S.; Sbihi, H.; Boutin, R.; Bode, L.; Robertson, B.; Yonemitsu, C.; Field, C.J.; Becker, A.B.; Mandhane, P.J.; et al. Breastmilk feeding practices are associated with the co-occurrence of bacteria in mothers’ milk and the infant gut: The CHILD Cohort Study. Cell Host Microbe 2020, 28, 285–297.e4. [Google Scholar] [CrossRef]
- van den Elsen, L.W.J.; Garssen, J.; Burcelin, R.; Verhasselt, V. Shaping the gut microbiota by breastfeeding: The gateway to allergy prevention? Front. Pediatr. 2019, 7, 47. [Google Scholar] [CrossRef]
- Feehley, T.; Plunkett, C.H.; Bao, R.; Choi Hong, S.M.; Culleen, E.; Belda-Ferre, P.; Campbell, E.; Aitoro, R.; Nocerino, R.; Andrade, J.; et al. Healthy infants harbor intestinal bacteria that protect against food allergy. Nat Med. 2019, 25, 448–453. [Google Scholar] [CrossRef]
- Depner, M.; Taft, D.H.; Kirjavainen, P.V.; Kalanetra, K.M.; Karvonen, A.M.; Peschel, S.; Schmausser-Hechfellner, E.; Roduit, C.; Frei, R.; Lauener, R.; et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat Med. 2020, 26, 1766–1775. [Google Scholar] [CrossRef] [PubMed]
- Host, A.; Husby, S.; Osterballe, O. A prospective study of cow’s milk allergy in exclusively breast-fed infants. Incidence, pathogenetic role of early inadvertent exposure to cow’s milk formula, and characterization of bovine milk protein in human milk. Acta Paediatr. Scand. 1988, 77, 663–670. [Google Scholar] [CrossRef] [PubMed]
- Brough, H.A.; Simpson, A.; Makinson, K.; Hankinson, J.; Brown, S.; Douiri, A.; Belgrave, D.C.; Penagos, M.; Stephens, A.C.; McLean, W.H.; et al. Peanut allergy: Effect of environmental peanut exposure in children with filaggrin loss-of-function mutations. J. Allergy Clin. Immunol. 2014, 134, 867–875.e1. [Google Scholar] [CrossRef] [PubMed]
- Brough, H.A.; Liu, A.H.; Sicherer, S.; Makinson, K.; Douiri, A.; Brown, S.J.; Stephens, A.C.; Irwin McLean, W.H.; Turcanu, V.; Wood, R.A.; et al. Atopic dermatitis increases the effect of exposure to peanut antigen in dust on peanut sensitization and likely peanut allergy. J. Allergy Clin. Immunol. 2015, 135, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Korotkova, M.; Telemo, E.; Yamashiro, Y.; Hanson, L.A.; Strandvik, B. The ratio of n-6 to n-3 fatty acids in maternal diet influences the induction of neonatal immunological tolerance to ovalbumin. Clin. Exp. Immunol. 2004, 137, 237–244. [Google Scholar] [CrossRef]
- Verhasselt, V.; Milcent, V.; Cazareth, J.; Kanda, A.; Fleury, S.; Dombrowicz, D.; Glaichenhaus, N.; Julia, V. Breast milk-mediated transfer of an antigen induces tolerance and protection from allergic asthma. Nat. Med. 2008, 14, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Mosconi, E.; Rekima, A.; Seitz-Polski, B.; Kanda, A.; Fleury, S.; Tissandie, E.; Monteiro, R.; Dombrowicz, D.D.; Julia, V.; Glaichenhaus, N.; et al. Breast milk immune complexes are potent inducers of oral tolerance in neonates and prevent asthma development. Mucosal Immunol. 2010, 3, 461–474. [Google Scholar] [CrossRef]
- Yamamoto, T.; Tsubota, Y.; Kodama, T.; Kageyama-Yahara, N.; Kadowaki, M. Oral tolerance induced by transfer of food antigens via breast milk of allergic mothers prevents offspring from developing allergic symptoms in a mouse food allergy model. Clin. Dev. Immunol. 2012, 2012, 721085. [Google Scholar] [CrossRef]
- Turfkruyer, M.; Rekima, A.; Macchiaverni, P.; Le Bourhis, L.; Muncan, V.; van den Brink, G.R.; Tulic, M.K.; Verhasselt, V. Oral tolerance is inefficient in neonatal mice due to a physiological vitamin A deficiency. Mucosal Immunol. 2016, 9, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Rekima, A.; Macchiaverni, P.; Turfkruyer, M.; Holvoet, S.; Dupuis, L.; Baiz, N.; Annesi-Maesano, I.; Mercenier, A.; Nutten, S.; Verhasselt, V. Long-term reduction in food allergy susceptibility in mice by combining breastfeeding-induced tolerance and TGF-β-enriched formula after weaning. Clin. Exp. Allergy 2017, 47, 565–576. [Google Scholar] [CrossRef]
- Ohsaki, A.; Venturelli, N.; Buccigrosso, T.M.; Osganian, S.K.; Lee, J.; Blumberg, R.S.; Oyoshi, M.K. Maternal IgG immune complexes induce food allergen-specific tolerance in offspring. J. Exp. Med. 2018, 215, 91–113. [Google Scholar] [CrossRef] [PubMed]
- Dawod, B.; Haidl, I.D.; Azad, M.B.; Marshall, J.S. Toll-like receptor 2 impacts the development of oral tolerance in mouse pups via a milk-dependent mechanism. J. Allergy Clin. Immunol. 2020, 146, 631–641.e8. [Google Scholar] [CrossRef] [PubMed]
- Verhasselt, V.; Genuneit, J.; Metcalfe, J.R.; Tulic, M.K.; Rekima, A.; Palmer, D.J.; Prescott, S.L. Ovalbumin in breastmilk is associated with a decreased risk of IgE-mediated egg allergy in children. Allergy 2020, 75, 1463–1466. [Google Scholar] [CrossRef]
- Palmer, D.J.; Gold, M.S.; Makrides, M. Effect of cooked and raw egg consumption on ovalbumin content of human milk: A randomized, double-blind, cross-over trial. Clin. Exp. Allergy 2005, 35, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Palmer, D.J.; Makrides, M. Diet of lactating women and allergic reactions in their infants. Curr. Opin. Clin. Nutr. Metab. Care 2006, 9, 284–288. [Google Scholar] [CrossRef]
- Metcalfe, J.R.; Marsh, J.A.; D’Vaz, N.; Geddes, D.T.; Lai, C.T.; Prescott, S.L.; Palmer, D.J. Effects of maternal dietary egg intake during early lactation on human milk ovalbumin concentration: A randomized controlled trial. Clin. Exp. Allergy 2016, 46, 1605–1613. [Google Scholar] [CrossRef]
- Vance, G.H.; Lewis, S.A.; Grimshaw, K.E.; Wood, P.J.; Briggs, R.A.; Thornton, C.A.; Warner, J.O. Exposure of the fetus and infant to hens’ egg ovalbumin via the placenta and breast milk in relation to maternal intake of dietary egg. Clin. Exp. Allergy 2005, 35, 1318–1326. [Google Scholar] [CrossRef]
- Hirose, J.; Ito, S.; Hirata, N.; Kido, S.; Kitabatake, N.; Narita, H. Occurrence of the major food allergen, ovomucoid, in human breast milk as an immune complex. Biosci. Biotechnol. Biochem. 2001, 65, 1438–1440. [Google Scholar] [CrossRef]
- Pan, B.T.; Johnstone, R.M. Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: Selective externalization of the receptor. Cell 1983, 33, 967–978. [Google Scholar] [CrossRef]
- Galley, J.D.; Besner, G.E. The therapeutic potential of breast milk-derived extracellular vesicles. Nutrients 2020, 12, 745. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Raposo, G.; Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 2014, 30, 255–289. [Google Scholar] [CrossRef]
- van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell. Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef] [PubMed]
- van Herwijnen, M.J.C.; Driedonks, T.A.P.; Snoek, B.L.; Kroon, A.M.T.; Kleinjan, M.; Jorritsma, R.; Pieterse, C.M.J.; Hoen, E.; Wauben, M.H.M. Abundantly present miRNAs in milk-derived extracellular vesicles are conserved between mammals. Front. Nutr. 2018, 5, 81. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Johansson, S.M.; Qazi, K.R.; Filén, J.J.; Lahesmaa, R.; Norman, M.; Neve, E.P.; Scheynius, A.; Gabrielsson, S. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 2007, 179, 1969–1978. [Google Scholar] [CrossRef]
- Lönnerdal, B. Human milk microRNAs/exosomes: Composition and biological effects. Nestle Nutr. Inst. Workshop Ser. 2019, 90, 83–92. [Google Scholar] [CrossRef]
- van Herwijnen, M.J.; Zonneveld, M.I.; Goerdayal, S.; Nolte-’t Hoen, E.N.; Garssen, J.; Stahl, B.; Maarten Altelaar, A.F.; Redegeld, F.A.; Wauben, M.H. Comprehensive proteomic analysis of human milk-derived extracellular vesicles unveils a novel functional proteome distinct from other milk components. Mol. Cell. Proteomics. 2016, 15, 3412–3423. [Google Scholar] [CrossRef]
- Liao, Y.; Du, X.; Li, J.; Lönnerdal, B. Human milk exosomes and their microRNAs survive digestion in vitro and are taken up by human intestinal cells. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Kahn, S.; Liao, Y.; Du, X.; Xu, W.; Li, J.; Lönnerdal, B. Exosomal microRNAs in milk from mothers delivering preterm infants survive in vitro digestion and are taken up by human intestinal cells. Mol. Nutr. Food Res. 2018, 62, e1701050. [Google Scholar] [CrossRef]
- Hock, A.; Miyake, H.; Li, B.; Lee, C.; Ermini, L.; Koike, Y.; Chen, Y.; Maattanen, P.; Zani, A.; Pierro, A. Breast milk-derived exosomes promote intestinal epithelial cell growth. J. Pediatr. Surg. 2017, 52, 755–759. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.; Patel, M.; Williams, S.; Arora, H.; Brawner, K.; Sims, B. Human breast milk-derived exosomes attenuate cell death in intestinal epithelial cells. Innate Immun. 2018, 24, 278–284. [Google Scholar] [CrossRef]
- Torregrosa Paredes, P.; Gutzeit, C.; Johansson, S.; Admyre, C.; Stenius, F.; Alm, J.; Scheynius, A.; Gabrielsson, S. Differences in exosome populations in human breast milk in relation to allergic sensitization and lifestyle. Allergy 2014, 69, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yan, X.; Zhang, L.; Cai, J.; Zhou, Y.; Liu, H.; Hu, Y.; Chen, W.; Xu, S.; Liu, P.; et al. Identification and peptidomic profiling of exosomes in preterm human milk: Insights into necrotizing enterocolitis prevention. Mol. Nutr. Food Res. 2019, 63, e1801247. [Google Scholar] [CrossRef]
- Pisano, C.; Galley, J.; Elbahrawy, M.; Wang, Y.; Farrell, A.; Brigstock, D.; Besner, G.E. Human breast milk-derived extracellular vesicles in the protection against experimental necrotizing enterocolitis. J. Pediatr. Surg. 2020, 55, 54–58. [Google Scholar] [CrossRef]
- Ma, J.; Wang, C.; Long, K.; Zhang, H.; Zhang, J.; Jin, L.; Tang, Q.; Jiang, A.; Wang, X.; Tian, S.; et al. Exosomal microRNAs in giant panda (Ailuropoda melanoleuca) breast milk: Potential maternal regulators for the development of newborn cubs. Sci. Rep. 2017, 7, 3507. [Google Scholar] [CrossRef]
- Mirza, A.H.; Kaur, S.; Nielsen, L.B.; Storling, J.; Yarani, R.; Roursgaard, M.; Mathiesen, E.R.; Damm, P.; Svare, J.; Mortensen, H.B.; et al. Breast milk-derived extracellular vesicles enriched in exosomes from mothers with type 1 diabetes contain aberrant levels of microRNAs. Front. Immunol. 2019, 10, 2543. [Google Scholar] [CrossRef] [PubMed]
- Alsaweed, M.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. MicroRNAs in breastmilk and the lactating breast: Potential immunoprotectors and developmental regulators for the infant and the mother. Int. J. Environ. Res. Public Health 2015, 12, 13981–14020. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Yoshimura, Y.; Huang, Y.; Suzuki, R.; Yokoyama, M.; Okabe, M.; Shimamura, M. Two independent pathways of maternal cell transmission to offspring: Through placenta during pregnancy and by breast-feeding after birth. Immunology 2000, 101, 570–580. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.J.; Walter, B.; Deguzman, A.; Muller, H.K.; Walker, A.M. Trans-epithelial immune cell transfer during suckling modulates delayed-type hypersensitivity in recipients as a function of gender. PLoS ONE 2008, 3, e3562. [Google Scholar] [CrossRef] [PubMed]
- Molès, J.P.; Tuaillon, E.; Kankasa, C.; Bedin, A.S.; Nagot, N.; Marchant, A.; McDermid, J.M.; Van de Perre, P. Breastmilk cell trafficking induces microchimerism-mediated immune system maturation in the infant. Pediatr. Allergy Immunol. 2018, 29, 133–143. [Google Scholar] [CrossRef]
- Molès, J.P.; Tuaillon, E.; Kankasa, C.; Bedin, A.S.; Nagot, N.; Marchant, A.; McDermid, J.M.; Van de Perre, P. Breastfeeding-related maternal microchimerism. Nat. Rev. Immunol. 2017, 17, 729–731. [Google Scholar] [CrossRef] [PubMed]
- Cregan, M.D.; Fan, Y.; Appelbee, A.; Brown, M.L.; Klopcic, B.; Koppen, J.; Mitoulas, L.R.; Piper, K.M.; Choolani, M.A.; Chong, Y.S.; et al. Identification of nestin-positive putative mammary stem cells in human breastmilk. Cell Tissue Res. 2007, 329, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Chong, Y.S.; Choolani, M.A.; Cregan, M.D.; Chan, J.K. Unravelling the mystery of stem/progenitor cells in human breast milk. PLoS ONE 2010, 5, e14421. [Google Scholar] [CrossRef]
- Hassiotou, F.; Beltran, A.; Chetwynd, E.; Stuebe, A.M.; Twigger, A.J.; Metzger, P.; Trengove, N.; Lai, C.T.; Filgueira, L.; Blancafort, P.; et al. Breastmilk is a novel source of stem cells with multilineage differentiation potential. Stem Cells 2012, 30, 2164–2174. [Google Scholar] [CrossRef] [PubMed]
- Ninkina, N.; Kukharsky, M.S.; Hewitt, M.V.; Lysikova, E.A.; Skuratovska, L.N.; Deykin, A.V.; Buchman, V.L. Stem cells in human breast milk. Hum. Cell. 2019, 32, 223–230. [Google Scholar] [CrossRef]
- Twigger, A.J.; Hodgetts, S.; Filgueira, L.; Hartmann, P.E.; Hassiotou, F. From breast milk to brains: The potential of stem cells in human milk. J. Hum. Lact. 2013, 29, 136–139. [Google Scholar] [CrossRef]
- Hassiotou, F.; Mobley, A.; Geddes, D.; Hartmann, P.; Wilkie, T. Breastmilk imparts the mother’s stem cells to the infant. FASEB J. 2015, 29, 876–878. [Google Scholar] [CrossRef]
- Abd Allah, S.H.; Shalaby, S.M.; El-Shal, A.S.; El Nabtety, S.M.; Khamis, T.; Abd El Rhman, S.A.; Ghareb, M.A.; Kelani, H.M. Breast milk MSCs: An explanation of tissue growth and maturation of offspring. IUBMB Life 2016, 68, 935–942. [Google Scholar] [CrossRef]
- Ghosh, A. Breast milk stem cell survival in neonate’s gut, entery into neonate circulation and adaption by the body. Curr. Stem Cell Res. Ther. 2020, 15, 98–101. [Google Scholar] [CrossRef]
- Aydın, M.Ş.; Yiğit, E.N.; Vatandaşlar, E.; Erdoğan, E.; Öztürk, G. Transfer and integration of breast milk stem cells to the brain of suckling pups. Sci. Rep. 2018, 8, 14289. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, S.M.; Talaei-Khozani, T.; Sani, M.; Owrangi, B. Differentiation of human breast-milk stem cells to neural stem cells and neurons. Neurol. Res. Int. 2014, 2014, 807896. [Google Scholar] [CrossRef] [PubMed]
- Kinder, J.M.; Jiang, T.T.; Ertelt, J.M.; Xin, L.; Strong, B.S.; Shaaban, A.F.; Way, S.S. Cross-generational reproductive fitness enforced by microchimeric maternal cells. Cell 2015, 162, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Kinder, J.M.; Stelzer, I.A.; Arck, P.C.; Way, S.S. Immunological implications of pregnancy-induced microchimerism. Nat. Rev. Immunol. 2017, 17, 483–494. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).