Citrus aurantium L. Dry Extracts Ameliorate Adipocyte Differentiation of 3T3-L1 Cells Exposed to TNFα by Down-Regulating miR-155 Expression
Abstract
1. Introduction
2. Materials and Methods
2.1. Citrus aurantium L. Dry Extract (CAde)
2.2. Cell Culture, Adipocyte Differentiation, and Treatments
2.3. miRNA Mimic and Inhibitor Transfection in Differentiating 3T3-L1
2.4. Image Acquisition, Oil-Red O Staining, and Triglyceride (TG) Quantification Assay
2.5. Total RNA and miRNA Purification, Reverse Transcription, and Quantitative Real-Time PCR
2.6. Western Blot (WB) Analysis
2.7. Statistical Procedures
3. Results
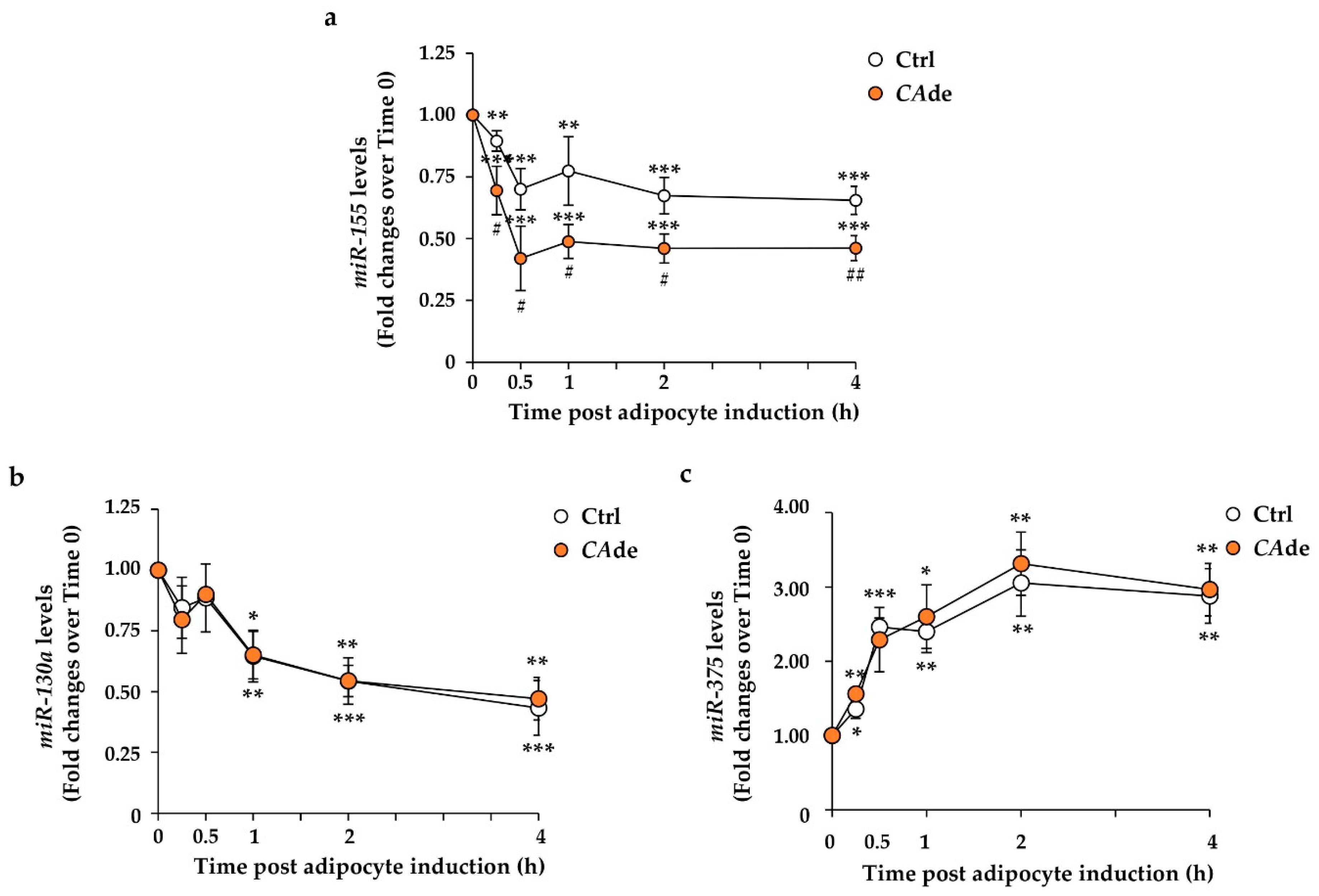
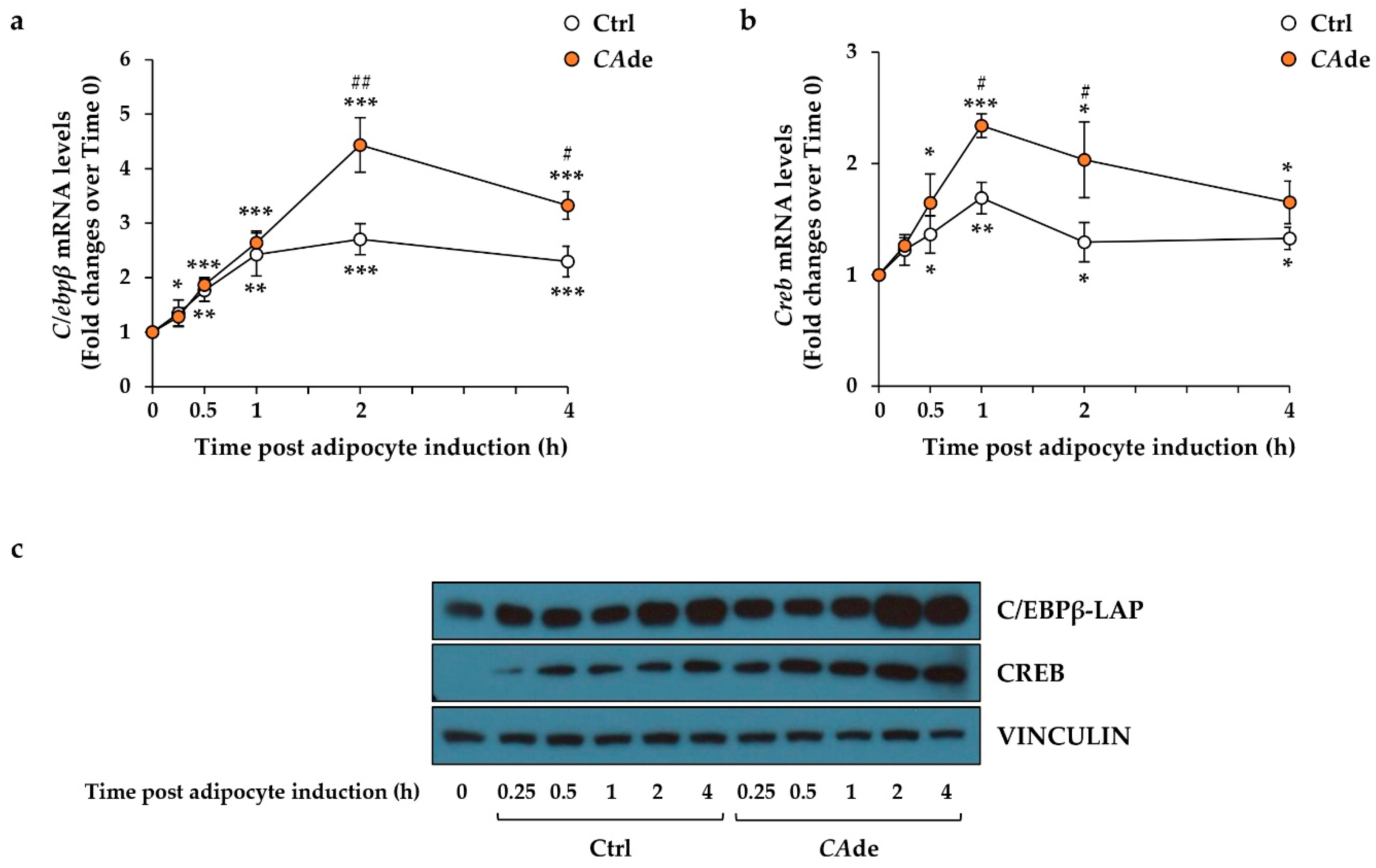
3.1. CAde Down-Regulated the Expression of miR-155 and Enhanced C/Ebpβ and Creb Levels during the Early Stage of Adipogenesis in 3T3-L1 Cells
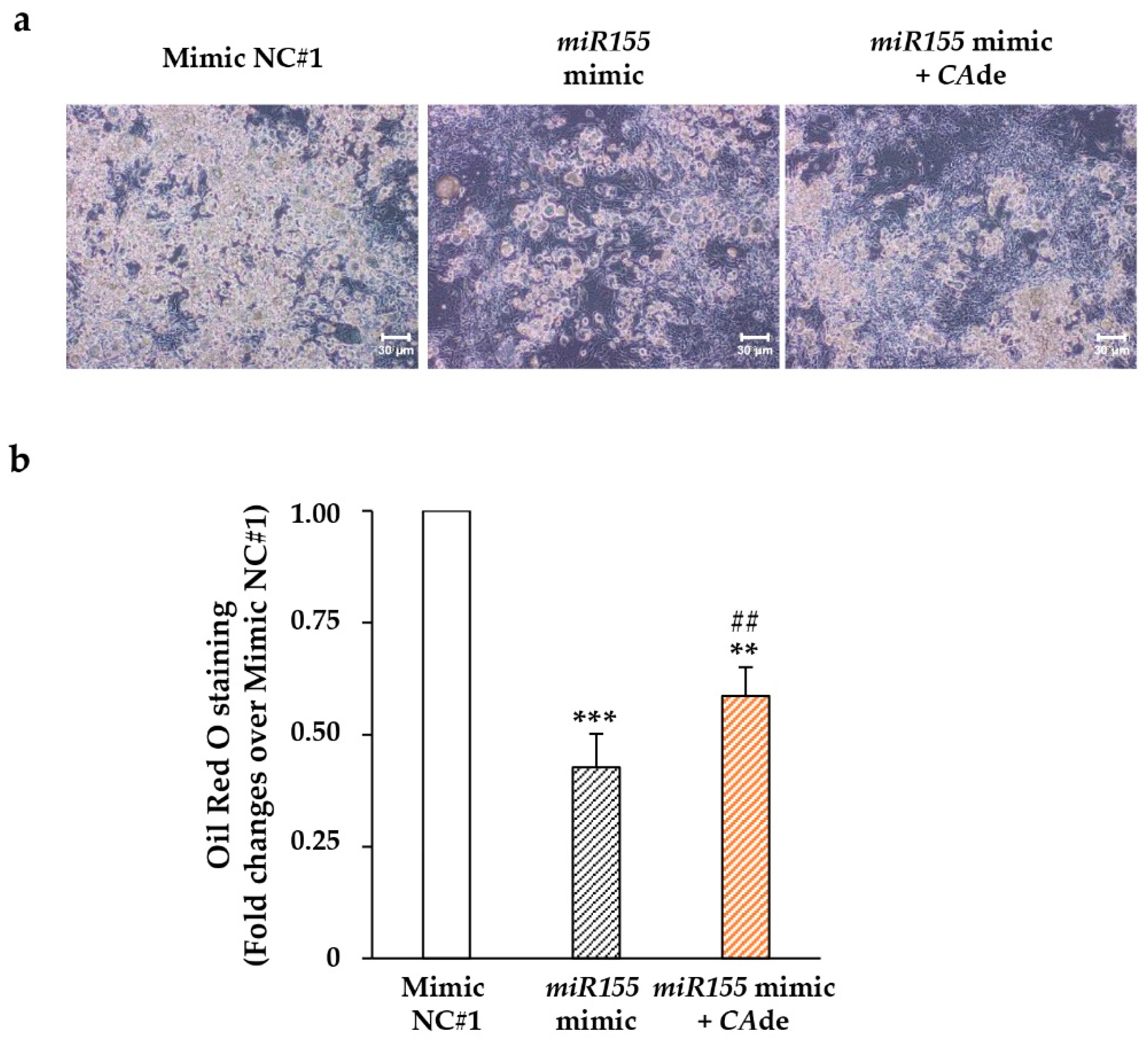
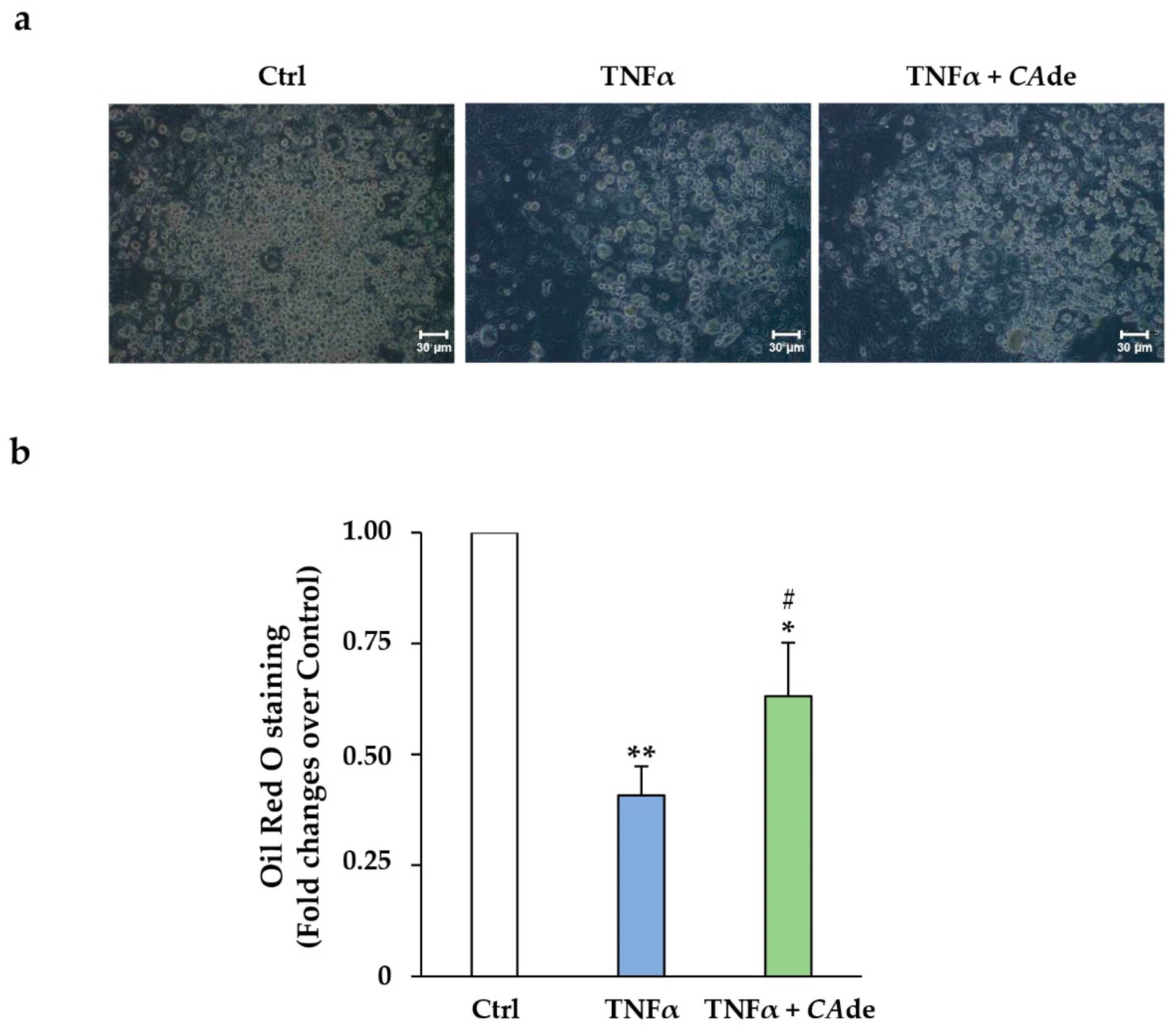
3.2. CAde Improved Terminal Adipocyte Differentiation of Both 3T3-L1 Cells Over-Expressing miR-155 by Mimic Transfection and 3T3-L1 Cells Exposed to TNFα
3.2.1. miR-155 Gain and Loss of Function Studies
3.2.2. Treatment with TNFα
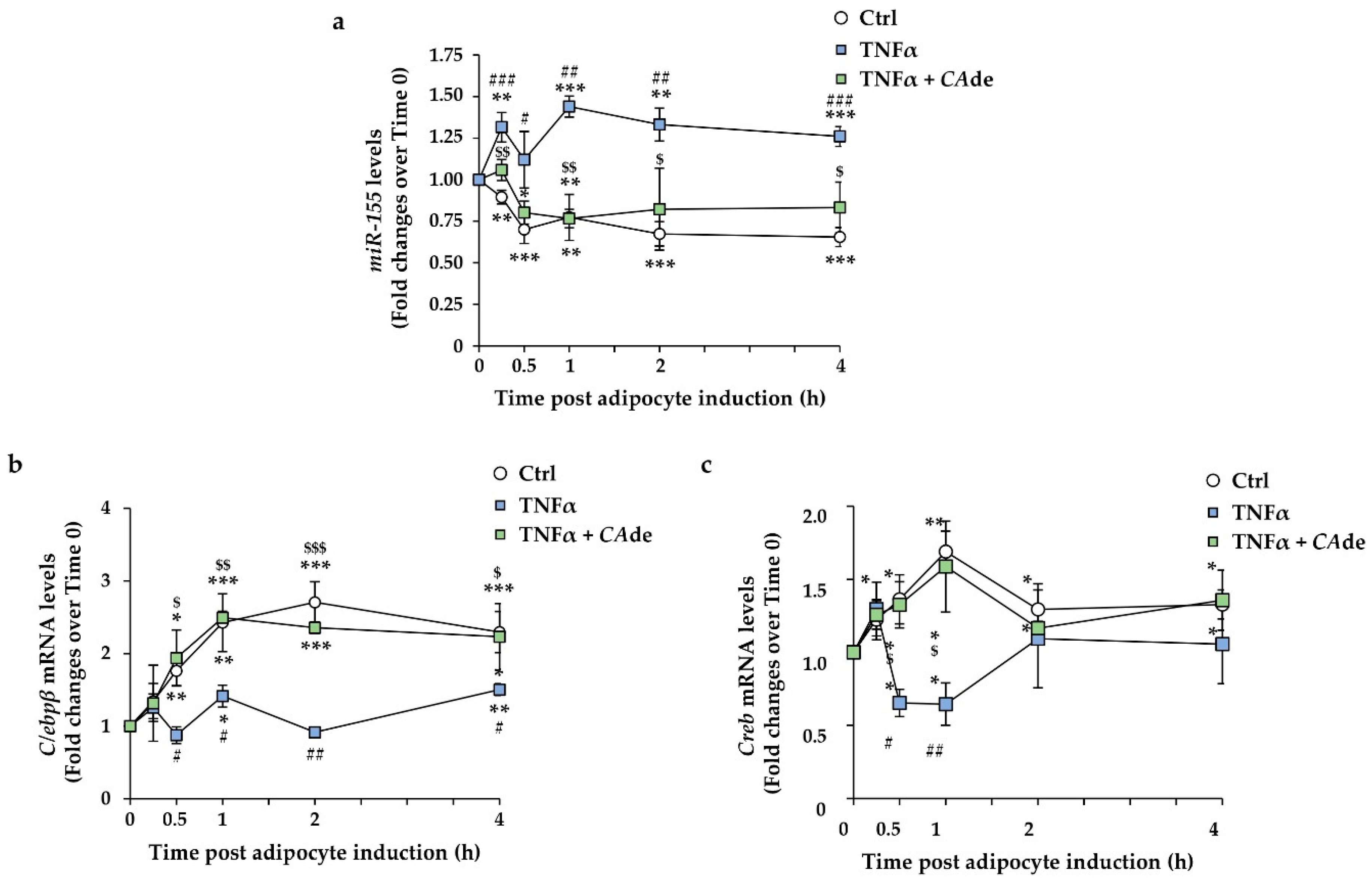
3.3. CAde Prevented TNFα-Induced Dysregulation of miR-155, C/Ebpβ, and Creb Expression during the Early Stage of Adipogenesis in 3T3-L1 Cells
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lowe, C.E.; O’Rahilly, S.; Rochford, J.J. Adipogenesis at a glance. J. Cell Sci. 2011, 124, 2681–2686. [Google Scholar] [CrossRef] [PubMed]
- Unger, R.H.; Clark, G.O.; Scherer, P.E.; Orci, L. Lipid homeostasis, lipotoxicity and the metabolic syndrome. Biochim. Biophys. Acta Mol. Cell Boil. Lipids 2010, 1801, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Gebert, L.; Macrae, I.J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell Boil. 2018, 20, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Oskowitz, A.Z.; Lu, J.; Penfornis, P.; Ylostalo, J.; McBride, J.; Flemington, E.K.; Prockop, D.J.; Pochampally, R. Human multipotent stromal cells from bone marrow and microRNA: Regulation of differentiation and leukemia inhibitory factor expression. Proc. Natl. Acad. Sci. USA 2008, 105, 18372–18377. [Google Scholar] [CrossRef]
- Brandão, B.B.; Guerra, B.A.; Mori, M.A. Shortcuts to a functional adipose tissue: The role of small non-coding RNAs. Redox Boil. 2017, 12, 82–102. [Google Scholar] [CrossRef]
- Chen, L.; Song, J.; Cui, J.; Hou, J.; Zheng, X.; Li, C.; Liu, L. MicroRNAs regulate adipocyte differentiation. Cell Boil. Int. 2013, 37, 533–546. [Google Scholar] [CrossRef]
- Lin, Q.; Gao, Z.; Alarcon, R.M.; Ye, J.; Yun, Z. A role of miR-27 in the regulation of adipogenesis. FEBS J. 2009, 276, 2348–2358. [Google Scholar] [CrossRef]
- Gerin, I.; Bommer, G.T.; McCoin, C.S.; Sousa, K.M.; Krishnan, V.; MacDougald, O.A. Roles for miRNA-378/378* in adipocyte gene expression and lipogenesis. Am. J. Physiol. Endocrinol. Metab. 2010, 299, 198–206. [Google Scholar] [CrossRef]
- Singh, S.K.; Bhadra, M.P.; Girschick, H.J.; Bhadra, U. MicroRNAs—Micro in size but macro in function. FEBS J. 2008, 275, 4929–4944. [Google Scholar] [CrossRef]
- Karbiener, M.; Fischer, C.; Nowitsch, S.; Opriessnig, P.; Papak, C.; Ailhaud, G.; Dani, C.; Amri, E.Z.; Scheideler, M. MicroRNA miR-27b impairs human adipocyte differentiation and targets PPARgamma. Biochem. Biophys. Res. Commun. 2009, 390, 247–251. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, A.Y.; Lee, H.W.; Son, Y.H.; Lee, G.Y.; Lee, J.-W.; Lee, Y.S.; Kim, J.B. MiR-27a is a negative regulator of adipocyte differentiation via suppressing PPARγ expression. Biochem. Biophys. Res. Commun. 2010, 392, 323–328. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.-F.; Zhang, Y.; Li, X.-Y.; Li, C.; Tian, W.; Liu, L. Expression of miR-31, miR-125b-5p, and miR-326 in the Adipogenic Differentiation Process of Adipose-Derived Stem Cells. OMICS A J. Integr. Boil. 2009, 13, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Wang, J.; Pan, Q.; Yu, Y.; Zhang, Y.; Wan, Y.; Wang, J.; Li, X.; Hong, A. Characterization of function and regulation of miR-24-1 and miR-31. Biochem. Biophys. Res. Commun. 2009, 380, 660–665. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.-Y.; Ou, H.-S.; Feng, S.-D.; Zhang, X.-Y.; Tuo, Q.-H.; Li, L.; Zhu, B.-Y.; Gao, Z.-P.; Tang, C.-K.; Yin, W.-D.; et al. Changes in microRNA (miR) Profile and Effectis of miR-320 in Insulin-Resistant 3T3-L1 Adipocytes. Clin. Exp. Pharmacol. Physiol. 2009, 36, 32–39. [Google Scholar] [CrossRef] [PubMed]
- He, A.; Zhu, L.; Gupta, N.; Chang, Y.; Fang, F.-D. Overexpression of Micro Ribonucleic Acid 29, Highly Up-Regulated in Diabetic Rats, Leads to Insulin Resistance in 3T3-L1 Adipocytes. Mol. Endocrinol. 2007, 21, 2785–2794. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Chen, Y.; Niu, Y.; Chen, W.; Wang, Q.; Xiao, S.; Li, A.; Xie, Y.; Li, J.; Zhao, X.; et al. A deep investigation into the adipogenesis mechanism: Profile of microRNAs regulating adipogenesis by modulating the canonical Wnt/β-catenin signaling pathway. BMC Genom. 2010, 11, 320. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.K.; Lee, M.J.; Abdelmohsen, K.; Kim, W.; Kim, M.M.; Srikantan, S.; Martindale, J.L.; Hutchison, E.R.; Kim, H.H.; Marasa, B.S.; et al. miR-130 suppresses adipogenesis by inhibiting peroxisome proliferator-activated receptor gamma expression. Mol. Cell. Biol. 2011, 31, 626–638. [Google Scholar] [CrossRef]
- Ling, H.-Y.; Wen, G.-B.; Feng, S.-D.; Tuo, Q.-H.; Ou, H.-S.; Yao, C.H.; Zhu, B.-Y.; Gao, Z.-P.; Zhang, L.; Liao, D.-F. MicroRNA-375 promotes 3T3-L1 adipocyte differentiation through modulation of extracellular signal-regulated kinase signalling. Clin. Exp. Pharmacol. Physiol. 2011, 38, 239–246. [Google Scholar] [CrossRef]
- Tam, W.; Ben-Yehuda, D.; Hayward, W.S. Bic, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol. Cell. Boil. 1997, 17, 1490–1502. [Google Scholar] [CrossRef]
- Alivernini, S.; Gremese, E.; McSharry, C.; Tolusso, B.; Ferraccioli, G.; McInnes, I.B.; Kurowska-Stolarska, M. MicroRNA-155—At the Critical Interface of Innate and Adaptive Immunity in Arthritis. Front. Immunol. 2018, 8. [Google Scholar] [CrossRef]
- Faraoni, I.; Antonetti, F.R.; Cardone, J.; Bonmassar, E. MiR-155 gene: A typical multifunctional microRNA. Biochim. Biophys. Acta Mol. Basis Dis. 2009, 1792, 497–505. [Google Scholar] [CrossRef]
- Liu, S.; Yang, Y.; Wu, J. TNFα-induced up-regulation of miR-155 inhibits adipogenesis by down-regulating early adipogenic transcription factors. Biochem. Biophys. Res. Commun. 2011, 414, 618–624. [Google Scholar] [CrossRef]
- Karkeni, E.; Astier, J.; Tourniaire, F.; El-Abed, M.; Romier, B.; Gouranton, E.; Wan, L.; Borel, P.; Salles, J.; Walrand, S.; et al. Obesity-associated Inflammation Induces microRNA-155 Expression in Adipocytes and Adipose Tissue: Outcome on Adipocyte Function. J. Clin. Endocrinol. Metab. 2016, 101, 1615–1626. [Google Scholar] [CrossRef] [PubMed]
- McGregor, R.; Seo, D.Y. MiRNAs as Nutritional Targets in Aging. In Molecular Basis of Nutrition and Aging; Elsevier BV: Amsterdam, The Nederlands, 2016; pp. 277–291. [Google Scholar]
- García-Segura, L.; Pérez-Andrade, M.; Miranda-Ríos, J. The Emerging Role of MicroRNAs in the Regulation of Gene Expression by Nutrients. J. Nutr. Nutr. 2013, 6, 16–31. [Google Scholar] [CrossRef]
- Ross, S.A.; Davis, C.D. The Emerging Role of microRNAs and Nutrition in Modulating Health and Disease. Annu. Rev. Nutr. 2014, 34, 305–336. [Google Scholar] [CrossRef] [PubMed]
- Conroy, K.P.; Davidson, I.M.; Warnock, M. Pathogenic obesity and nutraceuticals. Proc. Nutr. Soc. 2011, 70, 426–438. [Google Scholar] [CrossRef] [PubMed]
- Raciti, G.; Fiory, F.; Campitelli, M.; Desiderio, A.; Spinelli, R.; Longo, M.; Nigro, C.; Pepe, G.; Sommella, E.; Campiglia, P.; et al. Citrus aurantium L. dry extracts promote C/Ebpβ expression and improve adipocyte differentiation in 3T3-L1 cells. PLoS ONE 2018, 13, e0193704. [Google Scholar] [CrossRef]
- Longo, M.; Spinelli, R.; D’Esposito, V.; Zatterale, F.; Fiory, F.; Nigro, C.; Raciti, G.; Miele, C.; Formisano, P.; Beguinot, F.; et al. Pathologic endoplasmic reticulum stress induced by glucotoxic insults inhibits adipocyte differentiation and induces an inflammatory phenotype. Biochim. Biophys. Acta Bioenerg. 2016, 1863, 1146–1156. [Google Scholar] [CrossRef]
- Raciti, G.; Spinelli, R.; Desiderio, A.; Longo, M.; Parrillo, L.; Nigro, C.; D’Esposito, V.; Mirra, P.; Fiory, F.; Pilone, V.; et al. Specific CpG hyper-methylation leads to Ankrd26 gene down-regulation in white adipose tissue of a mouse model of diet-induced obesity. Sci. Rep. 2017, 7, 43526. [Google Scholar] [CrossRef]
- Nigro, C.; Mirra, P.; Prevenzano, I.; Leone, A.; Fiory, F.; Longo, M.; Cabaro, S.; Oriente, F.; Beguinot, F.; Miele, C. miR-214-Dependent Increase of PHLPP2 Levels Mediates the Impairment of Insulin-Stimulated Akt Activation in Mouse Aortic Endothelial Cells Exposed to Methylglyoxal. Int. J. Mol. Sci. 2018, 19, 522. [Google Scholar] [CrossRef]
- Liu, X.-F.; Bera, T.K.; Kahue, C.; Escobar, T.; Fei, Z.; Raciti, G.; Pastan, I. ANKRD26 and Its Interacting Partners TRIO, GPS2, HMMR and DIPA Regulate Adipogenesis in 3T3-L1 Cells. PLoS ONE 2012, 7, e38130. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Park, H.J.; Kang, S.N.; Jang, S.-H.; Lee, S.-J.; Ko, Y.-G.; Kim, G.-S.; Cho, J.-H. Blueberry Peel Extracts Inhibit Adipogenesis in 3T3-L1 Cells and Reduce High-Fat Diet-Induced Obesity. PLoS ONE 2013, 8, e69925. [Google Scholar] [CrossRef] [PubMed]
- Raciti, G.; Iadicicco, C.; Ulianich, L.; Vind, B.F.; Gaster, M.; Andreozzi, F.; Longo, M.; Teperino, R.; Ungaro, P.; Di Jeso, B.; et al. Glucosamine-induced endoplasmic reticulum stress affects GLUT4 expression via activating transcription factor 6 in rat and human skeletal muscle cells. Diabetologia 2010, 53, 955–965. [Google Scholar] [CrossRef] [PubMed]
- Iacomino, G.; Siani, A. Role of microRNAs in obesity and obesity-related diseases. Genes Nutr. 2017, 12, 23. [Google Scholar] [CrossRef] [PubMed]
- Arner, P.; Kulyté, A. MicroRNA regulatory networks in human adipose tissue and obesity. Nat. Rev. Endocrinol. 2015, 11, 276–288. [Google Scholar] [CrossRef]
- Klöting, N.; Berthold, S.; Kovács, P.; Schön, M.P.; Fasshauer, M.; Ruschke, K.; Stumvoll, M.; Blüher, M. MicroRNA Expression in Human Omental and Subcutaneous Adipose Tissue. PLoS ONE 2009, 4, e4699. [Google Scholar] [CrossRef]
- Hilton, C.; Neville, M.J.; Karpe, F. MicroRNAs in adipose tissue: Their role in adipogenesis and obesity. Int. J. Obes. 2012, 37, 325–332. [Google Scholar] [CrossRef]
- Neville, M.J.; Collins, J.M.; Gloyn, A.L.; McCarthy, M.I.; Karpe, F. Comprehensive human adipose tissue mRNA and microRNA endogenous control selection for quantitative real-time-PCR normalization. Obesity 2010, 19, 888–892. [Google Scholar] [CrossRef]
- Civelek, M.; Hagopian, R.; Pan, C.; Che, N.; Yang, W.-P.; Kayne, P.S.; Saleem, N.K.; Cederberg, H.; Kuusisto, J.; Gargalovic, P.S.; et al. Genetic regulation of human adipose microRNA expression and its consequences for metabolic traits. Hum. Mol. Genet. 2013, 22, 3023–3037. [Google Scholar] [CrossRef]
- Son, Y.H.; Ka, S.; Kim, A.Y.; Kim, J.B. Regulation of Adipocyte Differentiation via MicroRNAs. Endocrinol. Metab. 2014, 29, 122–135. [Google Scholar] [CrossRef]
- Kim, G.S.; Park, H.J.; Woo, J.-H.; Kim, M.-K.; Koh, P.-O.; Min, W.; Ko, Y.-G.; Kim, C.-H.; Won, C.; Cho, J.-H. Citrus aurantium flavonoids inhibit adipogenesis through the Akt signaling pathway in 3T3-L1 cells. BMC Complement. Altern. Med. 2012, 12, 31. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Kim, H.-L.; Jung, J.; Ahn, K.S.; Kwak, H.J.; Um, Y.J.; Park, J. Bitter Orange (Citrus aurantium Linné) Improves Obesity by Regulating Adipogenesis and Thermogenesis through AMPK Activation. Nutriens 2019, 11, 1988. [Google Scholar] [CrossRef] [PubMed]
- Lechner, S.; Mitterberger, M.C.; Mattesich, M.; Zwerschke, W. Role of C/Ebpβ-LAP and C/Ebpβ-LIP in early adipogenic differentiation of human white adipose-derived progenitors and at later stages in immature adipocytes. Differentioation 2013, 85, 20–31. [Google Scholar] [CrossRef]
- Eseberri, I.; Lasa, A.; Miranda, J.; Gràcia, A.; Portillo, M.P. Potential miRNA involvement in the anti-adipogenic effect of resveratrol and its metabolites. PLoS ONE 2017, 12, e0184875. [Google Scholar] [CrossRef]
- Saadatmandi, C.B.; Łoboda, A.; Wagner, A.E.; Stachurska, A.; Jozkowicz, A.; Dulak, J.; Döring, F.; Wolffram, S.; Rimbach, G. Effect of quercetin and its metabolites isorhamnetin and quercetin-3-glucuronide on inflammatory gene expression: Role of miR-155. J. Nutr. Biochem. 2011, 22, 293–299. [Google Scholar] [CrossRef]
- Arango, D.; Diosa-Toro, M.; Rojas-Hernandez, L.S.; Cooperstone, J.L.; Schwartz, S.J.; Mo, X.; Jiang, J.; Schmittgen, T.D.; Doseff, A.I. Dietary apigenin reduces LPS-induced expression of miR-155 restoring immune balance during inflammation. Mol. Nutr. Food Res. 2015, 59, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Kalliolias, G.; Ivashkiv, L.B. TNF biology, pathogenic mechanisms and emerging therapeutic strategies. Nat. Rev. Rheumatol. 2015, 12, 49–62. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Campitelli, M.; Desiderio, A.; Cacace, G.; Nigro, C.; Prevenzano, I.; Leone, A.; de Simone, S.; Campiglia, P.; Formisano, P.; Raciti, G.A.; et al. Citrus aurantium L. Dry Extracts Ameliorate Adipocyte Differentiation of 3T3-L1 Cells Exposed to TNFα by Down-Regulating miR-155 Expression. Nutrients 2020, 12, 1587. https://doi.org/10.3390/nu12061587
Campitelli M, Desiderio A, Cacace G, Nigro C, Prevenzano I, Leone A, de Simone S, Campiglia P, Formisano P, Raciti GA, et al. Citrus aurantium L. Dry Extracts Ameliorate Adipocyte Differentiation of 3T3-L1 Cells Exposed to TNFα by Down-Regulating miR-155 Expression. Nutrients. 2020; 12(6):1587. https://doi.org/10.3390/nu12061587
Chicago/Turabian StyleCampitelli, Michele, Antonella Desiderio, Giuseppe Cacace, Cecilia Nigro, Immacolata Prevenzano, Alessia Leone, Sonia de Simone, Pietro Campiglia, Pietro Formisano, Gregory A. Raciti, and et al. 2020. "Citrus aurantium L. Dry Extracts Ameliorate Adipocyte Differentiation of 3T3-L1 Cells Exposed to TNFα by Down-Regulating miR-155 Expression" Nutrients 12, no. 6: 1587. https://doi.org/10.3390/nu12061587
APA StyleCampitelli, M., Desiderio, A., Cacace, G., Nigro, C., Prevenzano, I., Leone, A., de Simone, S., Campiglia, P., Formisano, P., Raciti, G. A., Beguinot, F., & Miele, C. (2020). Citrus aurantium L. Dry Extracts Ameliorate Adipocyte Differentiation of 3T3-L1 Cells Exposed to TNFα by Down-Regulating miR-155 Expression. Nutrients, 12(6), 1587. https://doi.org/10.3390/nu12061587






