Salivary Microbiota Shifts under Sustained Consumption of Oolong Tea in Healthy Adults
Abstract
1. Introduction
2. Materials and Methods
2.1. Oolong Tea Infusion Preparation and Phenolic Profile Analysis
2.2. Subject Enrollment, Study Design, and Salivary Sample Collection
2.3. Salivary Bacterial DNA Extraction
2.4. Illumina Sequencing of Salivary Bacteria
2.5. Bioinformatic Analysis
3. Results
3.1. Phenolic Profile of Oolong Tea Infusion
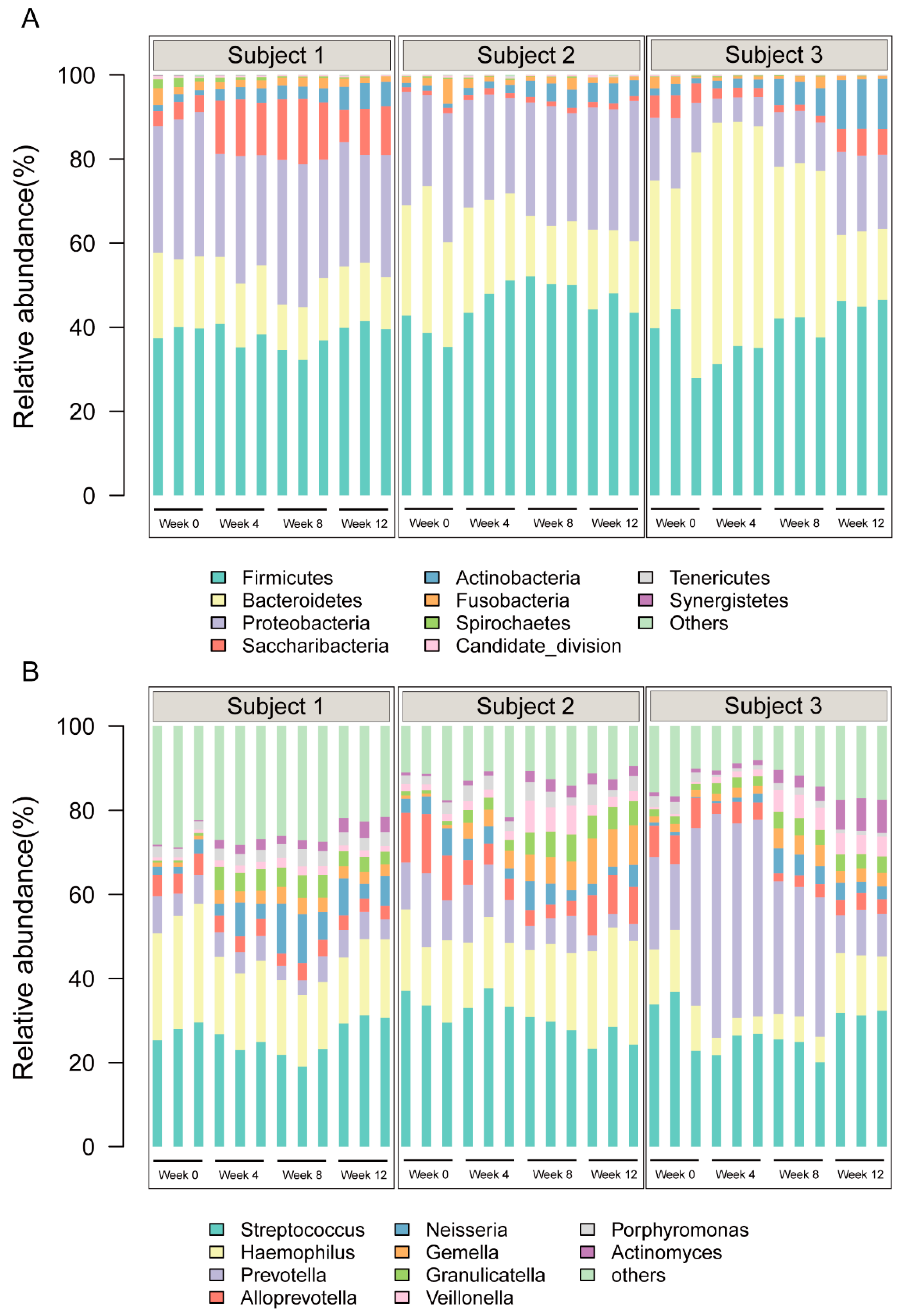
3.2. Overall Salivary Bacterial Structure
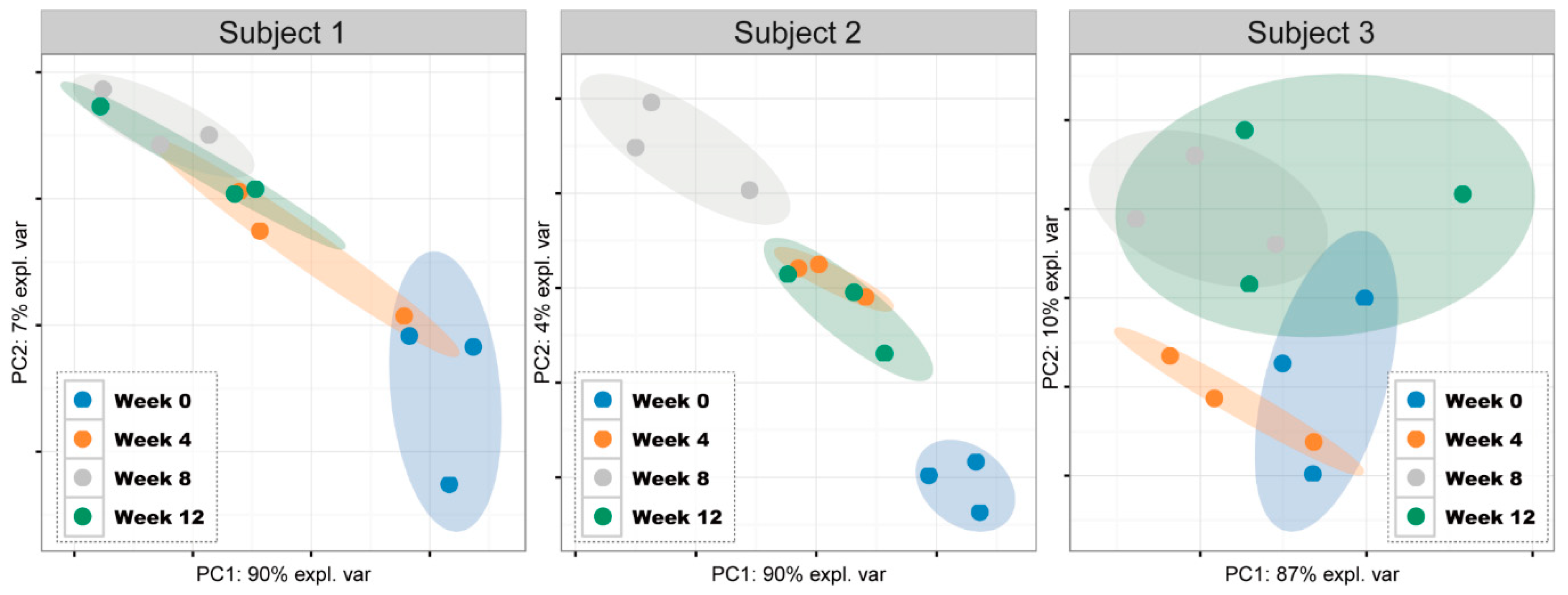
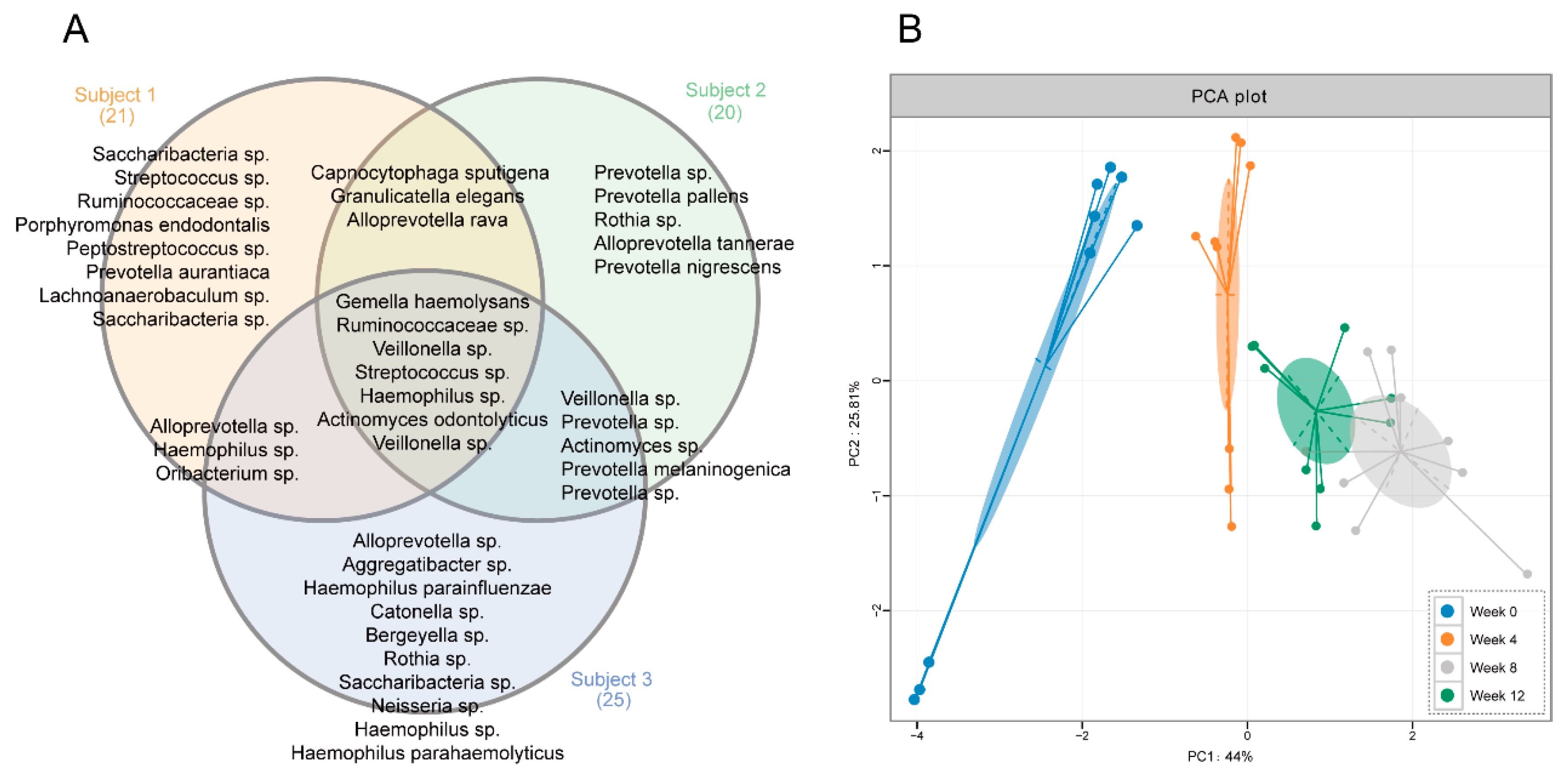
3.3. Comparisons of Salivary Bacterial Communities
3.4. Correlation Networks of Salivary Microbiota
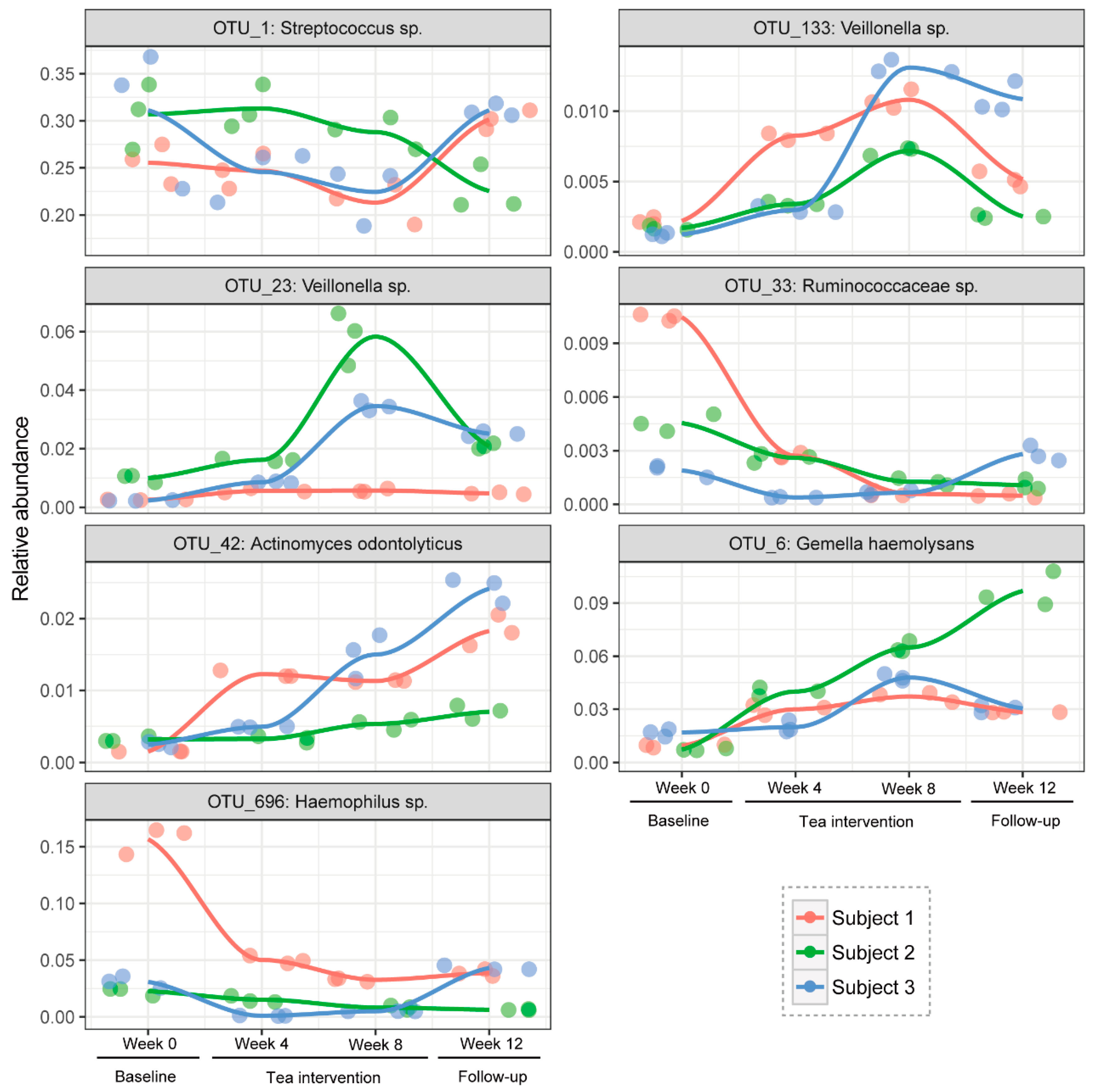
3.5. Hub Salivary Microbiota Identification
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jenkinson, H.F.; Lamont, R.J. Oral microbial communities in sickness and in health. Trends Microbiol. 2005, 13, 589–595. [Google Scholar] [CrossRef]
- Velden, U.V.D.; Winkelhoff, A.J.V.; Abbas, F.; Graaff, J.D. The habitat of periodontopathic micro-organisms. J. Clin. Periodontol. 1986, 13, 243–248. [Google Scholar] [CrossRef]
- Yang, F.; Zeng, X.; Ning, K.; Liu, K.-L.; Lo, C.-C.; Wang, W.; Chen, J.; Wang, D.; Huang, R.; Chang, X. Saliva microbiomes distinguish caries-active from healthy human populations. ISME J. 2012, 6, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Lazarevic, V.; Whiteson, K.; Hernandez, D.; François, P.; Schrenzel, J. Study of inter-and intra-individual variations in the salivary microbiota. BMC Genom. 2010, 11, 523. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, T.; Kageyama, S.; Furuta, M.; Tsuboi, H.; Takeuchi, K.; Shibata, Y.; Shimazaki, Y.; Akifusa, S.; Ninomiya, T.; Kiyohara, Y. Bacterial diversity in saliva and oral health-related conditions: The Hisayama Study. Sci. Rep. 2016, 6, 22164. [Google Scholar] [CrossRef] [PubMed]
- Chinsembu, K.C. Plants and other natural products used in the management of oral infections and improvement of oral health. Acta Trop. 2016, 154, 6–18. [Google Scholar] [CrossRef] [PubMed]
- Musarra-Pizzo, M.; Ginestra, G.; Smeriglio, A.; Pennisi, R.; Sciortino, M.T.; Mandalari, G. The antimicrobial and antiviral activity of polyphenols from almond (Prunus dulcis L.) skin. Nutrients 2019, 11, 2355. [Google Scholar] [CrossRef] [PubMed]
- Tsou, S.-H.; Hu, S.-W.; Yang, J.-J.; Yan, M.; Lin, Y.-Y. Potential Oral Health Care Agent from Coffee against Virulence Factor of Periodontitis. Nutrients 2019, 11, 2235. [Google Scholar] [CrossRef]
- Isola, G.; Polizzi, A.; Muraglie, S.; Leonardi, R.; Lo Giudice, A. Assessment of Vitamin C and Antioxidant Profiles in Saliva and Serum in Patients with Periodontitis and Ischemic Heart Disease. Nutrients 2019, 11, 2956. [Google Scholar] [CrossRef]
- Isola, G. Current Evidence of Natural Agents in Oral and Periodontal Health. Nutrients 2020, 12, 585. [Google Scholar] [CrossRef]
- Chun, O.K.; Chung, S.J.; Song, W.O. Estimated dietary flavonoid intake and major food sources of US adults. J. Nutr. 2007, 137, 1244–1252. [Google Scholar] [CrossRef] [PubMed]
- Ferrazzano, G.F.; Roberto, L.; Amato, I.; Cantile, T.; Sangianantoni, G.; Ingenito, A. Antimicrobial properties of green tea extract against cariogenic microflora: An in vivo study. J. Med. Food 2011, 14, 907–911. [Google Scholar] [CrossRef] [PubMed]
- Ferrazzano, G.F.; Amato, I.; Ingenito, A.; De Natale, A.; Pollio, A. Anti-cariogenic effects of polyphenols from plant stimulant beverages (cocoa, coffee, tea). Fitoterapia 2009, 80, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Narotzki, B.; Reznick, A.Z.; Aizenbud, D.; Levy, Y. Green tea: A promising natural product in oral health. Arch. Oral Biol. 2012, 57, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Chen, Z.; Guo, H.; He, D.; Zhao, H.; Wang, Z.; Zhang, W.; Liao, L.; Zhang, C.; Ni, L. The modulatory effect of infusions of green tea, oolong tea, and black tea on gut microbiota in high-fat-induced obese mice. Food Funct. 2016, 7, 4869–4879. [Google Scholar] [CrossRef]
- Liu, Z.; Bruins, M.E.; Ni, L.; Vincken, J.-P. Green and black tea phenolics: Bioavailability, transformation by colonic microbiota, and modulation of colonic microbiota. J. Agric. Food Chem. 2018, 66, 8469–8477. [Google Scholar] [CrossRef]
- Hall, M.W.; Singh, N.; Ng, K.F.; Lam, D.K.; Goldberg, M.B.; Tenenbaum, H.C.; Neufeld, J.D.; Beiko, R.; Senadheera, D.B. Inter-personal diversity and temporal dynamics of dental, tongue, and salivary microbiota in the healthy oral cavity. NPJ Biofilm. Microbiomes 2017, 3, 2. [Google Scholar] [CrossRef]
- Obanda, M.; Owuor, P.O.; Taylor, S.J. Flavanol composition and caffeine content of green leaf as quality potential indicators of Kenyan black teas. J. Sci. Food Agric. 1997, 74, 209–215. [Google Scholar] [CrossRef]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, J.I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996. [Google Scholar] [CrossRef] [PubMed]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Yu, W.-H.; Izard, J.; Baranova, O.V.; Lakshmanan, A.; Dewhirst, F.E. The Human Oral Microbiome Database: A web accessible resource for investigating oral microbe taxonomic and genomic information. Database 2010, 2010. [Google Scholar] [CrossRef] [PubMed]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. ICWSM 2009, 8, 361–362. [Google Scholar]
- Faust, K.; Lima-Mendez, G.; Lerat, J.-S.; Sathirapongsasuti, J.F.; Knight, R.; Huttenhower, C.; Lenaerts, T.; Raes, J. Cross-biome comparison of microbial association networks. Front. Microbiol. 2015, 6, 1200. [Google Scholar] [CrossRef]
- Layeghifard, M.; Hwang, D.M.; Guttman, D.S. Disentangling interactions in the microbiome: A network perspective. Trends Microbiol. 2017, 25, 217–228. [Google Scholar] [CrossRef]
- Heberle, H.; Meirelles, G.V.; da Silva, F.R.; Telles, G.P.; Minghim, R. InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015, 16, 169. [Google Scholar] [CrossRef]
- Belstrøm, D.; Holmstrup, P.; Bardow, A.; Kokaras, A.; Fiehn, N.-E.; Paster, B.J. Temporal stability of the salivary microbiota in oral health. PLoS ONE 2016, 11, e0147472. [Google Scholar] [CrossRef]
- Romani Vestman, N.; Chen, T.; Lif Holgerson, P.; Öhman, C.; Johansson, I. Oral microbiota shift after 12-week supplementation with Lactobacillus reuteri DSM 17938 and PTA 5289; a randomized control trial. PLoS ONE 2015, 10, e0125812. [Google Scholar] [CrossRef]
- Wade, W.G. The oral microbiome in health and disease. Pharmacol. Res. 2013, 69, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Consortium, H.M.P. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar]
- Jiang, W.; Ling, Z.; Lin, X.; Chen, Y.; Zhang, J.; Yu, J.; Xiang, C.; Chen, H. Pyrosequencing analysis of oral microbiota shifting in various caries states in childhood. Microb. Ecol. 2014, 67, 962–969. [Google Scholar] [CrossRef] [PubMed]
- Limsong, J.; Benjavongkulchai, E.; Kuvatanasuchati, J. Inhibitory effect of some herbal extracts on adherence of Streptococcus mutans. J. Ethnopharmacol. 2004, 92, 281–289. [Google Scholar] [CrossRef]
- Kawarai, T.; Narisawa, N.; Yoneda, S.; Tsutsumi, Y.; Ishikawa, J.; Hoshino, Y.; Senpuku, H. Inhibition of Streptococcus mutans biofilm formation using extracts from Assam tea compared to green tea. Arch. Oral Biol. 2016, 68, 73–82. [Google Scholar] [CrossRef]
- Lim, M.Y.; Yoon, H.S.; Rho, M.; Sung, J.; Song, Y.-M.; Lee, K.; Ko, G. Analysis of the association between host genetics, smoking, and sputum microbiota in healthy humans. Sci. Rep. 2016, 6, 23745. [Google Scholar] [CrossRef]
- Paropkari, A.D.; Leblebicioglu, B.; Christian, L.M.; Kumar, P.S. Smoking, pregnancy and the subgingival microbiome. Sci. Rep. 2016, 6, 30388. [Google Scholar] [CrossRef]
- Hara, K.; Ohara, M.; Hayashi, I.; Hino, T.; Nishimura, R.; Iwasaki, Y.; Ogawa, T.; Ohyama, Y.; Sugiyama, M.; Amano, H. The green tea polyphenol (−)-epigallocatechin gallate precipitates salivary proteins including alpha-amylase: Biochemical implications for oral health. Eur. J. Oral Sci. 2012, 120, 132–139. [Google Scholar] [CrossRef]
- Khan, N.; Mukhtar, H. Tea polyphenols in promotion of human health. Nutrients 2019, 11, 39. [Google Scholar] [CrossRef]
- Isola, G.; Alibrandi, A.; Currò, M.; Matarese, M.; Ricca, S.; Matarese, G.; Ientile, R.; Kocher, T. Evaluation of salivary and serum ADMA levels in patients with periodontal and cardiovascular disease as subclinical marker of cardiovascular risk. J. Periodontol. 2019. [Google Scholar] [CrossRef]
- Isola, G.; Giudice, A.L.; Polizzi, A.; Alibrandi, A.; Patini, R.; Ferlito, S. Periodontitis and Tooth Loss Have Negative Systemic Impact on Circulating Progenitor Cell Levels: A Clinical Study. Genes 2019, 10, 1022. [Google Scholar] [CrossRef] [PubMed]
- Isola, G.; Polizzi, A.; Alibrandi, A.; Indelicato, F.; Ferlito, S. Analysis of Endothelin-1 concentrations in individuals with periodontitis. Sci. Rep. 2020, 10, 1652. [Google Scholar] [CrossRef] [PubMed]

| Peak No. a | tR (Min) | Tentative Identification | Chemical Formula | [M-H]− (m/z) | ||
|---|---|---|---|---|---|---|
| Measured Mass (Da) | Theoretical Exact Mass (Da) | Mass Accuracy (ppm) | ||||
| 1 | 1.05 | Caffeoyl-hexoside | C15H18O9 | 341.0875 | 341.0873 | 0.58 |
| 2 | 1.40 | L-Theanine | C7H14N2O3 | 173.0931 | 173.0927 | 2.53 |
| 3 | 1.97 | Epigallocatechin-glucuronide | C21H22O13 | 481.0991 | 481.0983 | 1.74 |
| 4 | 2.49 | Theasinensin C | C30H26O14 | 609.1235 | 609.1245 | −1.60 |
| 5 | 2.74 | Gallic acid | C7H6O5 | 169.0140 | 169.0137 | 1.52 |
| 6 | 2.92 | Theogallin | C14H16O10 | 343.0665 | 343.0666 | −0.19 |
| 7 | 3.80 | Theobromine b | C7H8N4O2 | 181.0736 | 181.0725 | 6.04 |
| 8 | 3.84 | Gallocatechin | C15H14O7 | 305.0662 | 305.0662 | 0.09 |
| 9 | 4.37 | Theasinensin B | C37H30O18 | 761.1348 | 761.1354 | −0.83 |
| 10 | 4.41 | Digalloyl-hexoside | C20H20O14 | 483.0758 | 483.0775 | −3.57 |
| 11 | 4.54 | O-Methylgallic acid | C8H8O5 | 183.0295 | 183.0294 | 0.58 |
| 12 | 4.81 | Theacitrin A | C37H28O18 | 759.1196 | 759.1198 | −0.24 |
| 13 | 4.91 | Epigallocatechin | C15H14O7 | 305.0689 | 305.0662 | 8.94 |
| 14 | 5.16 | p-Coumaroylquinic acid | C16H18O8 | 337.0923 | 337.0924 | −0.26 |
| 15 | 5.36 | Catechin | C15H14O6 | 289.0718 | 289.0713 | 1.87 |
| 16 | 5.60 | Caffeine b | C8H10N4O2 | 195.0888 | 195.0882 | 3.30 |
| 17 | 5.68 | Procyanidin | C30H26O12 | 577.1356 | 577.1346 | 1.65 |
| 18 | 5.79 | Epicatechin-epicatechin | C30H26O12 | 577.1356 | 577.1346 | 1.65 |
| 19 | 6.14 | p-Coumaroylquinic acid | C16H18O8 | 337.0923 | 337.0924 | −0.26 |
| 20 | 6.23 | Epicatechin | C15H14O6 | 289.0734 | 289.0713 | 7.41 |
| 21 | 6.34 | Epigallocatechin gallate | C22H18O11 | 457.0777 | 457.0771 | 1.24 |
| 22 | 6.41 | p-Coumaroylquinic acid | C16H18O8 | 337.0918 | 337.0924 | −1.74 |
| 23 | 6.68 | Gallocatechin gallate | C22H18O11 | 457.0773 | 457.0771 | 0.37 |
| 24 | 6.92 | Theaflavin | C29H24O12 | 563.1199 | 563.1190 | 1.60 |
| 25 | 7.01 | Myricetin-hexoside | C21H20O13 | 479.0827 | 479.0826 | 0.19 |
| 26 | 7.11 | Myricetin-hexoside | C21H20O13 | 479.0825 | 479.0826 | −0.23 |
| 27 | 7.21 | Quercetin-hexosyl-hexosyl-deoxyhexoside | C33H40O21 | 771.1986 | 771.1984 | 0.22 |
| 28 | 7.36 | Quercetin-hexosyl-hexosyl-deoxyhexoside | C33H40O21 | 771.1982 | 771.1984 | −0.30 |
| 29 | 7.62 | Kaempferol-deoxyhexosyl-deoxyhexoside | C27H30O14 | 577.1555 | 577.1558 | −0.48 |
| 30 | 7.72 | Kaempferol-hexosyl-hexosyl-deoxyhexoside | C33H40O20 | 755.2029 | 755.2035 | −0.81 |
| 31 | 8.00 | Kaempferol-hexosyl-hexosyl-deoxyhexoside | C33H40O20 | 755.2048 | 755.2035 | 1.70 |
| 32 | 8.43 | Kaempferol-hexosyl-hexoside | C27H30O15 | 593.1508 | 593.1507 | 0.18 |
| 33 | 8.78 | Kaempferol-hexoside | C21H20O11 | 447.0927 | 447.0928 | −0.18 |
| Baseline | Tea Intervention | Follow-Up | ||
|---|---|---|---|---|
| Week 0 | Week 4 | Week 8 | Week 12 | |
| Subject 1 | ||||
| Shannon | 5.28 ± 0.41 a | 4.68 ± 0.27 ab | 4.00 ± 0.39 b | 4.17 ± 0.40 b |
| Simpson | 0.94 ± 0.02 a | 0.89 ± 0.03 ab | 0.81 ± 0.04 b | 0.84 ± 0.06 b |
| Subject 2 | ||||
| Shannon | 4.79 ± 0.58 a | 4.63 ± 0.22 ab | 4.02 ± 0.27 b | 4.37 ± 0.13 ab |
| Simpson | 0.91 ± 0.06 a | 0.88 ± 0.06 a | 0.83 ± 0.05 b | 0.90 ± 0.02 a |
| Subject 3 | ||||
| Shannon | 3.99 ± 0.57 a | 3.93 ± 0.27 a | 3.90 ± 0.17 a | 4.07 ± 0.65 a |
| Simpson | 0.85 ± 0.10 a | 0.83 ± 0.03 a | 0.80 ± 0.03 b | 0.83 ± 0.09 a |
| Compared Data Sets | MRPP | Anosim | ||
|---|---|---|---|---|
| Delta | p-Value | R | p-Value | |
| Subject 1 vs. Subject 2 | 0.1969 | 0.001 | 0.7105 | 0.001 |
| Subject 1 vs. Subject 3 | 0.1479 | 0.001 | 0.4886 | 0.001 |
| Subject 2 vs. Subject 3 | 0.1919 | 0.001 | 0.7562 | 0.001 |
| Subject 1 vs. Subject 2 vs. Subject 3 | 0.2227 | 0.001 | 0.6482 | 0.001 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Z.; Guo, H.; Zhang, W.; Ni, L. Salivary Microbiota Shifts under Sustained Consumption of Oolong Tea in Healthy Adults. Nutrients 2020, 12, 966. https://doi.org/10.3390/nu12040966
Liu Z, Guo H, Zhang W, Ni L. Salivary Microbiota Shifts under Sustained Consumption of Oolong Tea in Healthy Adults. Nutrients. 2020; 12(4):966. https://doi.org/10.3390/nu12040966
Chicago/Turabian StyleLiu, Zhibin, Hongwen Guo, Wen Zhang, and Li Ni. 2020. "Salivary Microbiota Shifts under Sustained Consumption of Oolong Tea in Healthy Adults" Nutrients 12, no. 4: 966. https://doi.org/10.3390/nu12040966
APA StyleLiu, Z., Guo, H., Zhang, W., & Ni, L. (2020). Salivary Microbiota Shifts under Sustained Consumption of Oolong Tea in Healthy Adults. Nutrients, 12(4), 966. https://doi.org/10.3390/nu12040966