A Theacrine-Based Supplement Increases Cellular NAD+ Levels and Affects Biomarkers Related to Sirtuin Activity in C2C12 Muscle Cells In Vitro
Abstract
:1. Introduction
2. Methods
2.1. Cell Culture
2.2. NAD3 Versus Control Treatments
2.3. Post-Treatment Processing of Cells and Real-Time PCR for mRNA Expression Analyses
2.4. Global SIRT Activity and Citrate Synthase Activity Assays
2.5. Follow-Up Culture Experiments for Western Blotting Targets and Determination of NAD+ Concentrations
2.6. Statistics
3. Results
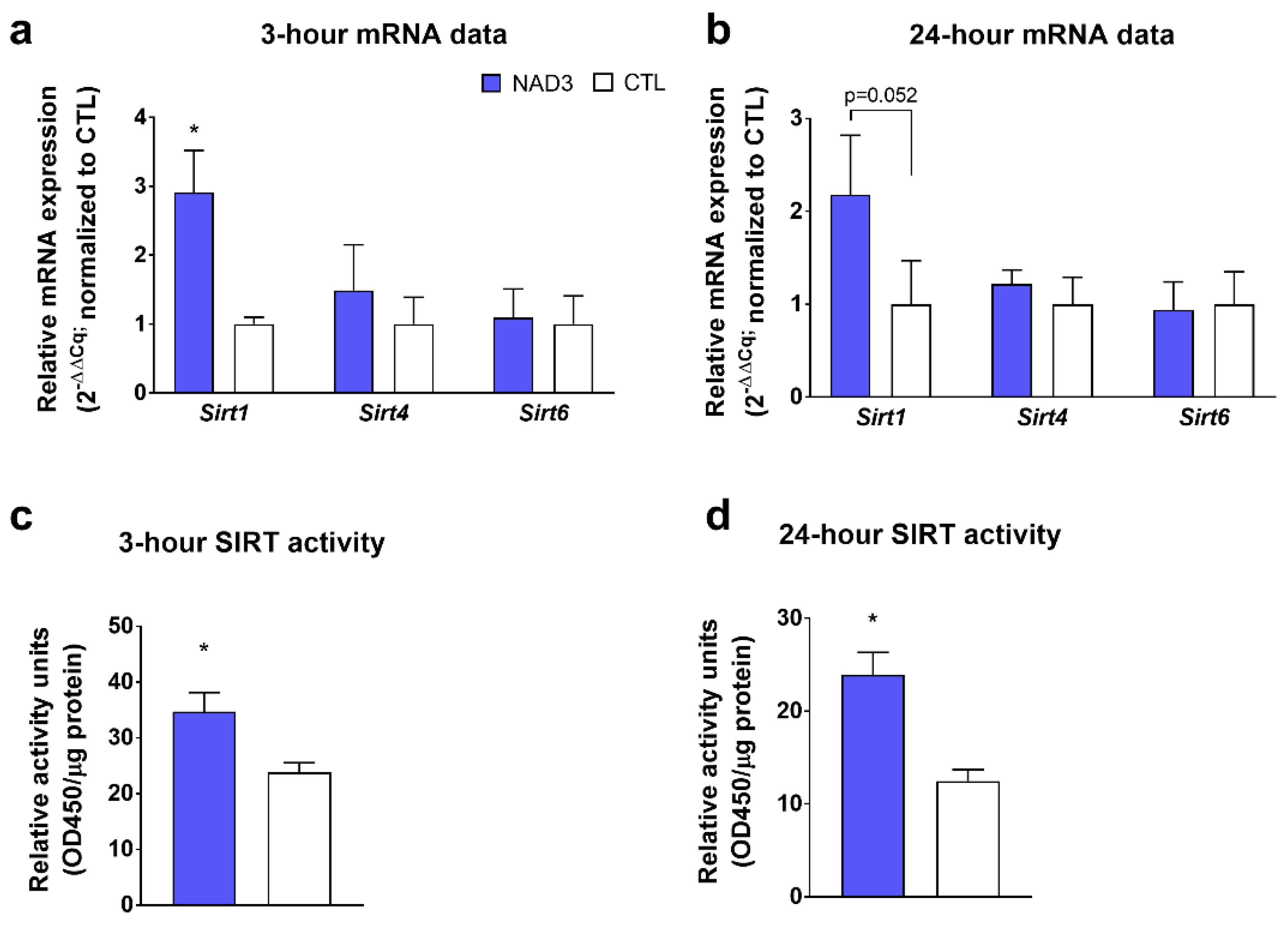
3.1. Effect of NAD3 Treatments on SIRT Markers
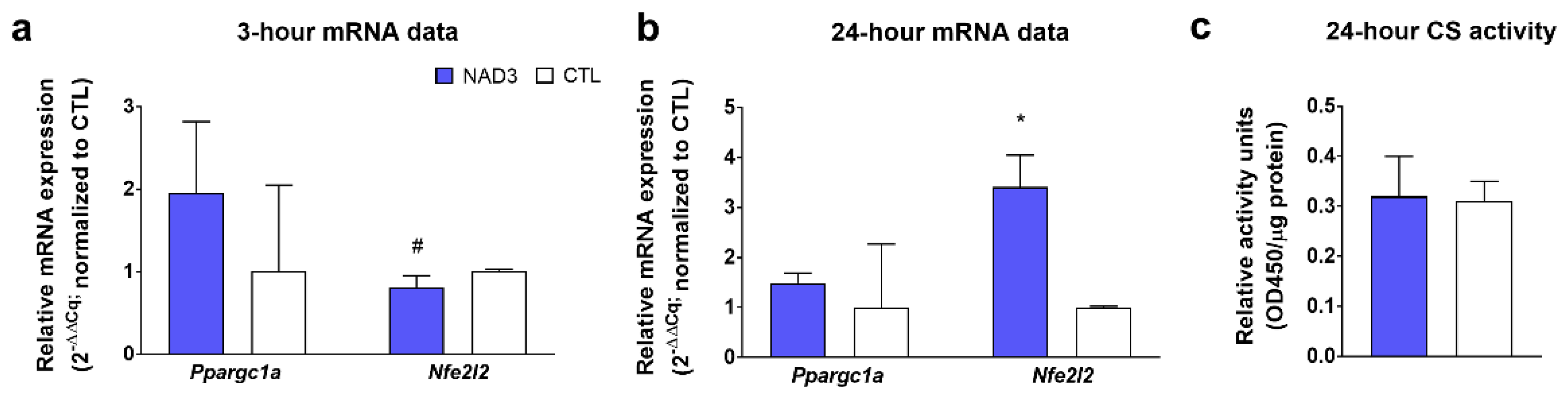
3.2. Effect of NAD3 Treatments on Mitochondrial Biogenesis Markers
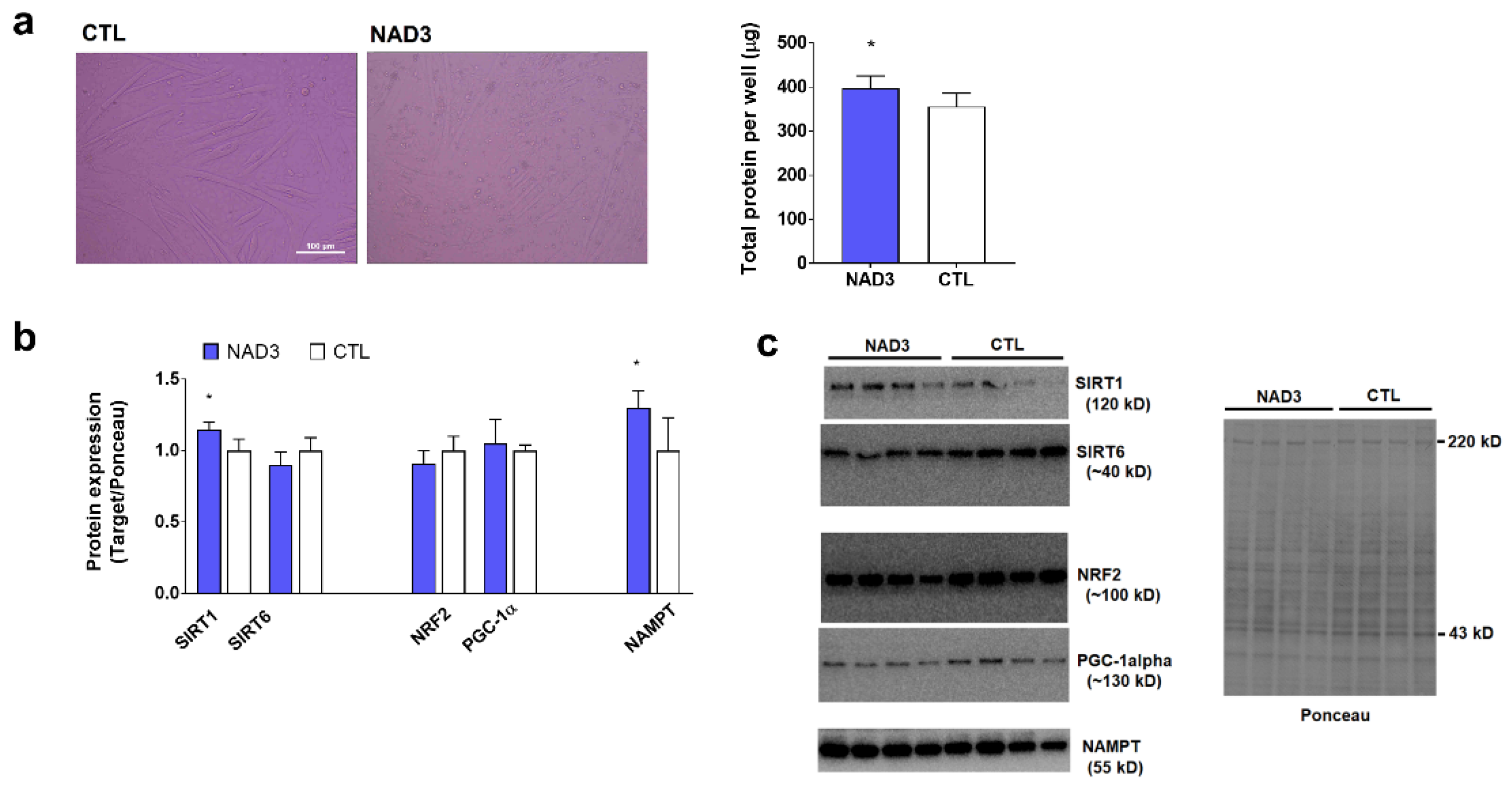
3.3. Effects of 24-Hour NAD3 Treatments on Protein Targets
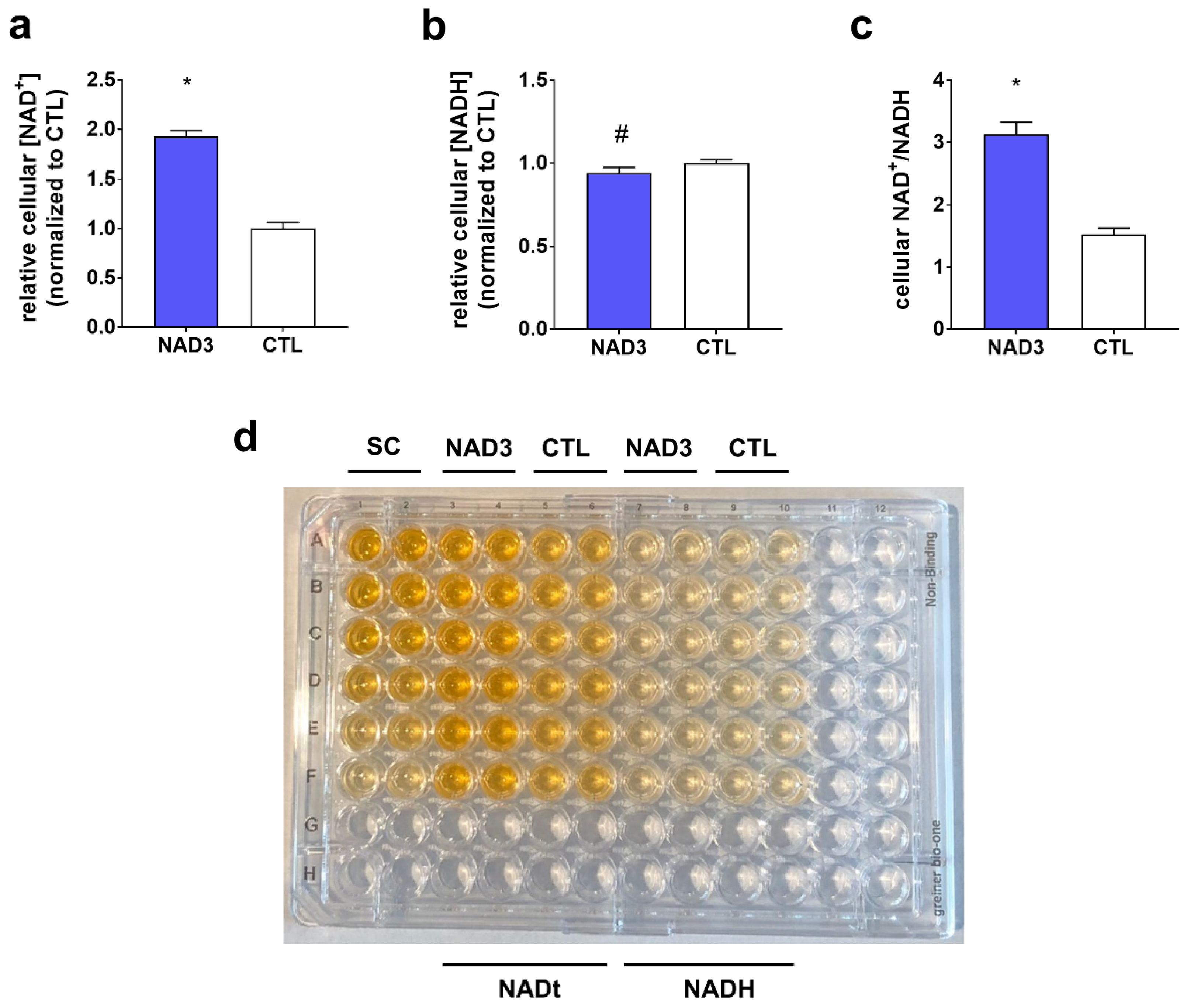
3.4. Effect of 24-hour NAD3 Treatments on Cellular NAD+, NADH, and NAD+/NADH
4. Discussion
Experimental Considerations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Johnson, M.L.; Robinson, M.M.; Nair, K.S. Skeletal muscle aging and the mitochondrion. Trends Endocrinol. Metab. 2013, 24, 247–256. [Google Scholar] [CrossRef] [Green Version]
- Gensous, N.; Bacalini, M.G.; Franceschi, C.; Meskers, C.G.M.; Maier, A.B.; Garagnani, P. Age-related DNA methylation changes: Potential impact on skeletal muscle aging in humans. Front. Physiol. 2019, 10, 996. [Google Scholar] [CrossRef] [PubMed]
- Demontis, F.; Piccirillo, R.; Goldberg, A.L.; Perrimon, N. Mechanisms of skeletal muscle aging: Insights from Drosophila and mammalian models. Dis. Models Mech. 2013, 6, 1339–1352. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bielak-Zmijewska, A.; Grabowska, W.; Ciolko, A.; Bojko, A.; Mosieniak, G.; Bijoch, L.; Sikora, E. The role of curcumin in the modulation of ageing. Int. J. Mol. Sci. 2019, 20, 1239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harrison, F.E. A critical review of vitamin C for the prevention of age-related cognitive decline and Alzheimer’s disease. J. Alzheimers Dis. 2012, 29, 711–726. [Google Scholar] [CrossRef] [Green Version]
- Quinzii, C.M.; Hirano, M. Coenzyme Q and mitochondrial disease. Dev. Disabil. Res. Rev. 2010, 16, 183–188. [Google Scholar] [CrossRef]
- Anand David, A.V.; Arulmoli, R.; Parasuraman, S. Overviews of biological importance of quercetin: A bioactive flavonoid. Pharmacogn. Rev. 2016, 10, 84–89. [Google Scholar] [CrossRef] [Green Version]
- Dang, W. The controversial world of sirtuins. Drug Discov. Today Technol. 2014, 12, e9–e17. [Google Scholar] [CrossRef] [Green Version]
- Grabowska, W.; Sikora, E.; Bielak-Zmijewska, A. Sirtuins, a promising target in slowing down the ageing process. Biogerontology 2017, 18, 447–476. [Google Scholar] [CrossRef] [Green Version]
- Bonkowski, M.S.; Sinclair, D.A. Slowing ageing by design: The rise of NAD(+) and sirtuin-activating compounds. Nat. Rev. Mol. Cell Biol. 2016, 17, 679–690. [Google Scholar] [CrossRef]
- Hubbard, B.P.; Sinclair, D.A. Small molecule SIRT1 activators for the treatment of aging and age-related diseases. Trends Pharmacol. Sci. 2014, 35, 146–154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cesareo, K.R.; Mason, J.R.; Saracino, P.G.; Morrissey, M.C.; Ormsbee, M.J. The effects of a caffeine-like supplement, TeaCrine(R), on muscular strength, endurance and power performance in resistance-trained men. J. Int. Soc. Sports Nutr. 2019, 16, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuhman, D.J.; Joyner, K.J.; Bloomer, R.J. Cognitive Performance and mood following ingestion of a theacrine-containing dietary supplement, caffeine, or placebo by young men and women. Nutrients 2015, 7, 9618–9632. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bello, M.L.; Walker, A.J.; McFadden, B.A.; Sanders, D.J.; Arent, S.M. The effects of TeaCrine(R) and caffeine on endurance and cognitive performance during a simulated match in high-level soccer players. J. Int. Soc. Sports Nutr. 2019, 16, 20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, M.; Zheng, J.; Zheng, C.; Huang, Z.; Huang, Q. Theacrine alleviates chronic inflammation by enhancing TGF-beta-mediated shifts via TGF-beta/SMAD pathway in Freund’s incomplete adjuvant-induced rats. Biochem. Biophys. Res. Commun. 2020, 522, 743–748. [Google Scholar] [CrossRef] [PubMed]
- Li, W.X.; Li, Y.F.; Zhai, Y.J.; Chen, W.M.; Kurihara, H.; He, R.R. Theacrine, a purine alkaloid obtained from Camellia assamica var. kucha, attenuates restraint stress-provoked liver damage in mice. J. Agric. Food Chem. 2013, 61, 6328–6335. [Google Scholar] [CrossRef]
- Wang, G.E.; Li, Y.F.; Zhai, Y.J.; Gong, L.; Tian, J.Y.; Hong, M.; Yao, N.; Wu, Y.P.; Kurihara, H.; He, R.R. Theacrine protects against nonalcoholic fatty liver disease by regulating acylcarnitine metabolism. Metabolism 2018, 85, 227–239. [Google Scholar] [CrossRef]
- Mumford, P.W.; Romero, M.A.; Mao, X.; Mobley, C.B.; Kephart, W.C.; Haun, C.T.; Roberson, P.A.; Young, K.C.; Martin, J.S.; Yarrow, J.F.; et al. Cross talk between androgen and Wnt signaling potentially contributes to age-related skeletal muscle atrophy in rats. J. Appl. Physiol. (1985) 2018, 125, 486–494. [Google Scholar] [CrossRef]
- Romero, M.A.; Mumford, P.W.; Roberson, P.A.; Osburn, S.C.; Parry, H.A.; Kavazis, A.N.; Gladden, L.B.; Schwartz, T.S.; Baker, B.A.; Toedebusch, R.G.; et al. Five months of voluntary wheel running downregulates skeletal muscle LINE-1 gene expression in rats. Am. J. Physiol. Cell Physiol. 2019, 317, C1313–C1323. [Google Scholar] [CrossRef]
- Lamb, D.A.; Moore, J.H.; Mesquita, P.H.C.; Smith, M.A.; Vann, C.G.; Osburn, S.C.; Fox, C.D.; Lopez, H.L.; Ziegenfuss, T.N.; Huggins, K.W.; et al. Resistance training increases muscle NAD(+) and NADH concentrations as well as NAMPT protein levels and global sirtuin activity in middle-aged, overweight, untrained individuals. Aging (Albany NY) 2020, 12, 9447–9460. [Google Scholar] [CrossRef]
- Larsen, S.; Nielsen, J.; Hansen, C.N.; Nielsen, L.B.; Wibrand, F.; Stride, N.; Schroder, H.D.; Boushel, R.; Helge, J.W.; Dela, F.; et al. Biomarkers of mitochondrial content in skeletal muscle of healthy young human subjects. J. Physiol. 2012, 590, 3349–3360. [Google Scholar] [CrossRef] [PubMed]
- Mesquita, P.H.C.; Lamb, D.A.; Parry, H.A.; Moore, J.H.; Smith, M.A.; Vann, C.G.; Osburn, S.C.; Fox, C.D.; Ruple, B.A.; Huggins, K.W.; et al. Acute and chronic effects of resistance training on skeletal muscle markers of mitochondrial remodeling in older adults. Physiol. Rep. 2020, 8, e14526. [Google Scholar] [CrossRef] [PubMed]
- Song, L.L.; Zhang, Y.; Zhang, X.R.; Song, Y.N.; Dai, H.Z. Theacrine attenuates myocardial fibrosis after myocardial infarction via the SIRT3/beta-catenin/PPARgamma pathway in estrogen-deficient mice. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 5477–5486. [Google Scholar] [CrossRef] [PubMed]
- Swanson, K.V.; Deng, M.; Ting, J.P. The NLRP3 inflammasome: Molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 2019, 19, 477–489. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mumford, P.W.; Osburn, S.C.; Fox, C.D.; Godwin, J.S.; Roberts, M.D. A Theacrine-Based Supplement Increases Cellular NAD+ Levels and Affects Biomarkers Related to Sirtuin Activity in C2C12 Muscle Cells In Vitro. Nutrients 2020, 12, 3727. https://doi.org/10.3390/nu12123727
Mumford PW, Osburn SC, Fox CD, Godwin JS, Roberts MD. A Theacrine-Based Supplement Increases Cellular NAD+ Levels and Affects Biomarkers Related to Sirtuin Activity in C2C12 Muscle Cells In Vitro. Nutrients. 2020; 12(12):3727. https://doi.org/10.3390/nu12123727
Chicago/Turabian StyleMumford, Petey W., Shelby C. Osburn, Carlton D. Fox, Joshua S. Godwin, and Michael D. Roberts. 2020. "A Theacrine-Based Supplement Increases Cellular NAD+ Levels and Affects Biomarkers Related to Sirtuin Activity in C2C12 Muscle Cells In Vitro" Nutrients 12, no. 12: 3727. https://doi.org/10.3390/nu12123727
APA StyleMumford, P. W., Osburn, S. C., Fox, C. D., Godwin, J. S., & Roberts, M. D. (2020). A Theacrine-Based Supplement Increases Cellular NAD+ Levels and Affects Biomarkers Related to Sirtuin Activity in C2C12 Muscle Cells In Vitro. Nutrients, 12(12), 3727. https://doi.org/10.3390/nu12123727





