Antibiotic Treatment Does Not Ameliorate the Metabolic Changes in Rats Presenting Dysbiosis After Consuming a High Fructose Diet
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Antibiotic Treatment
2.3. Metabolic Parameters
2.4. Liver Studies
2.4.1. Liver Histological Studies
2.4.2. Real-Time Quantitative Reverse Transcription PCR of Liver Genes
2.5. SCFA Extraction and Analysis
2.6. Microbiome Analysis
2.6.1. DNA Extraction, PCR Amplification, and Sequencing
2.6.2. Microbiome Data Processing and Analysis
2.7. Statistical Analysis
3. Results
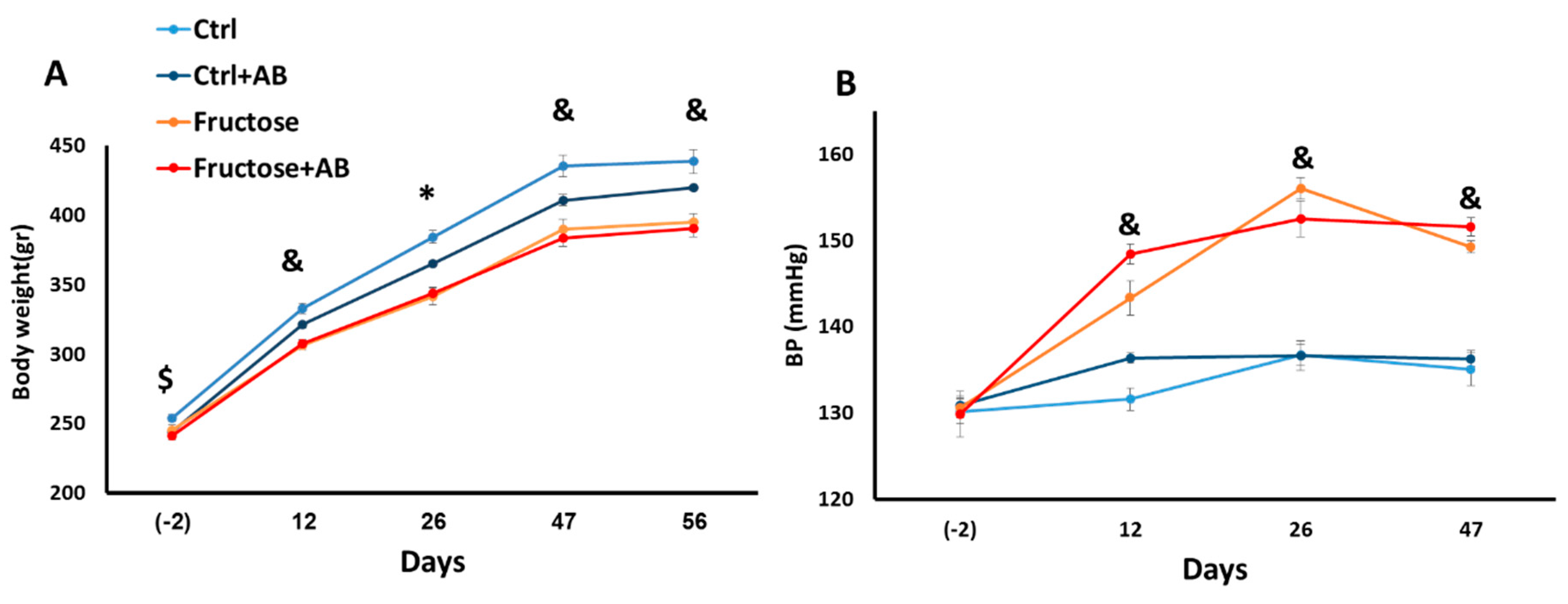
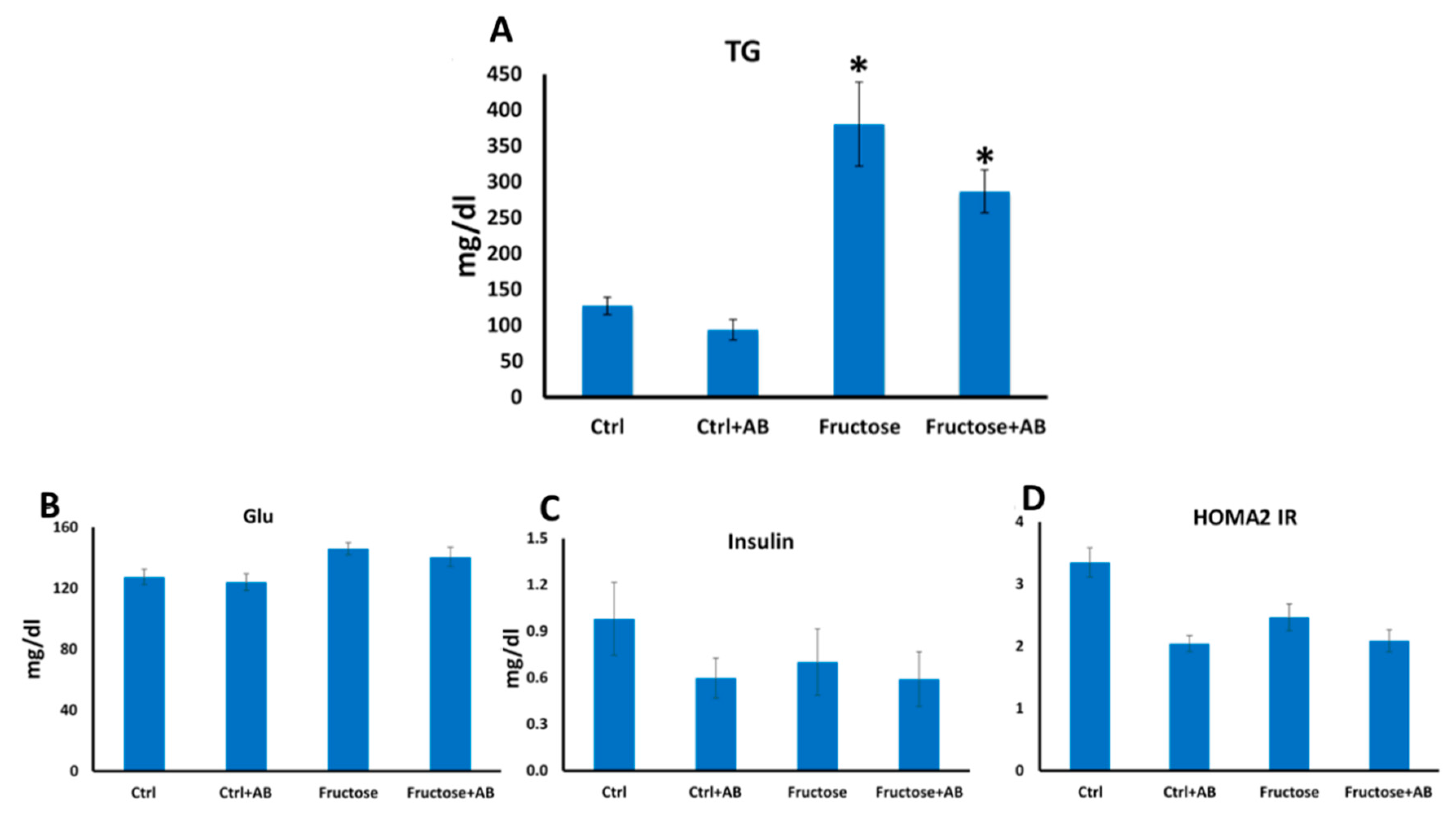
3.1. Metabolic and Hemodynamic Effects
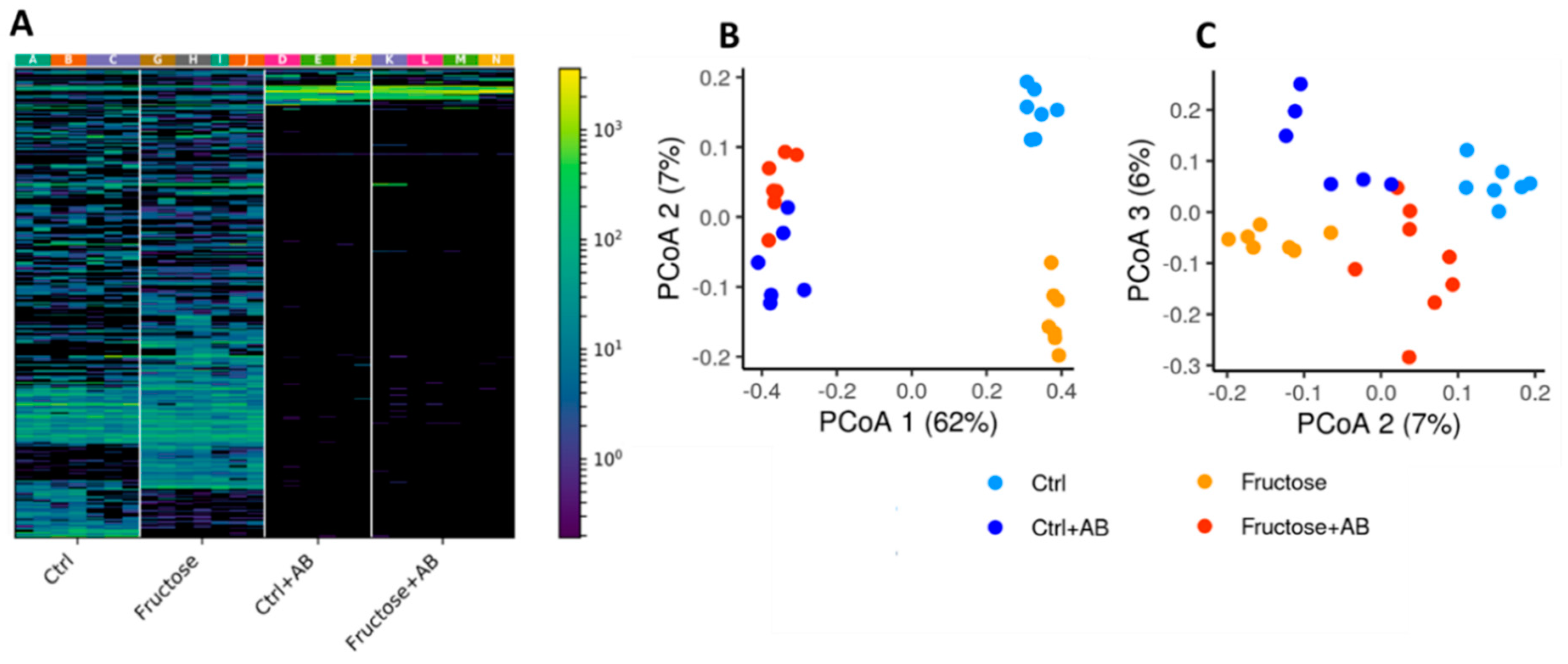
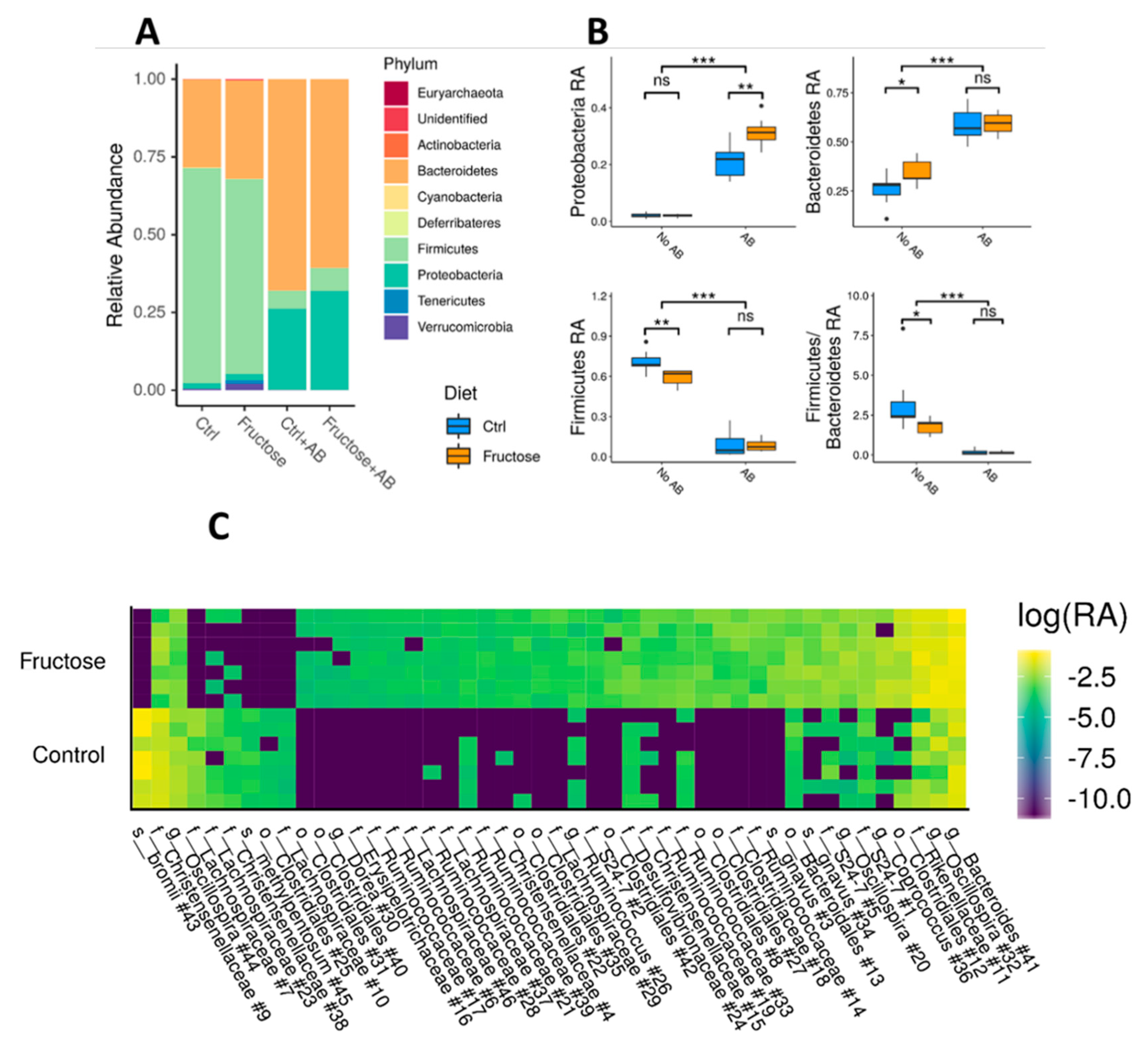
3.2. The Microbiome Profile
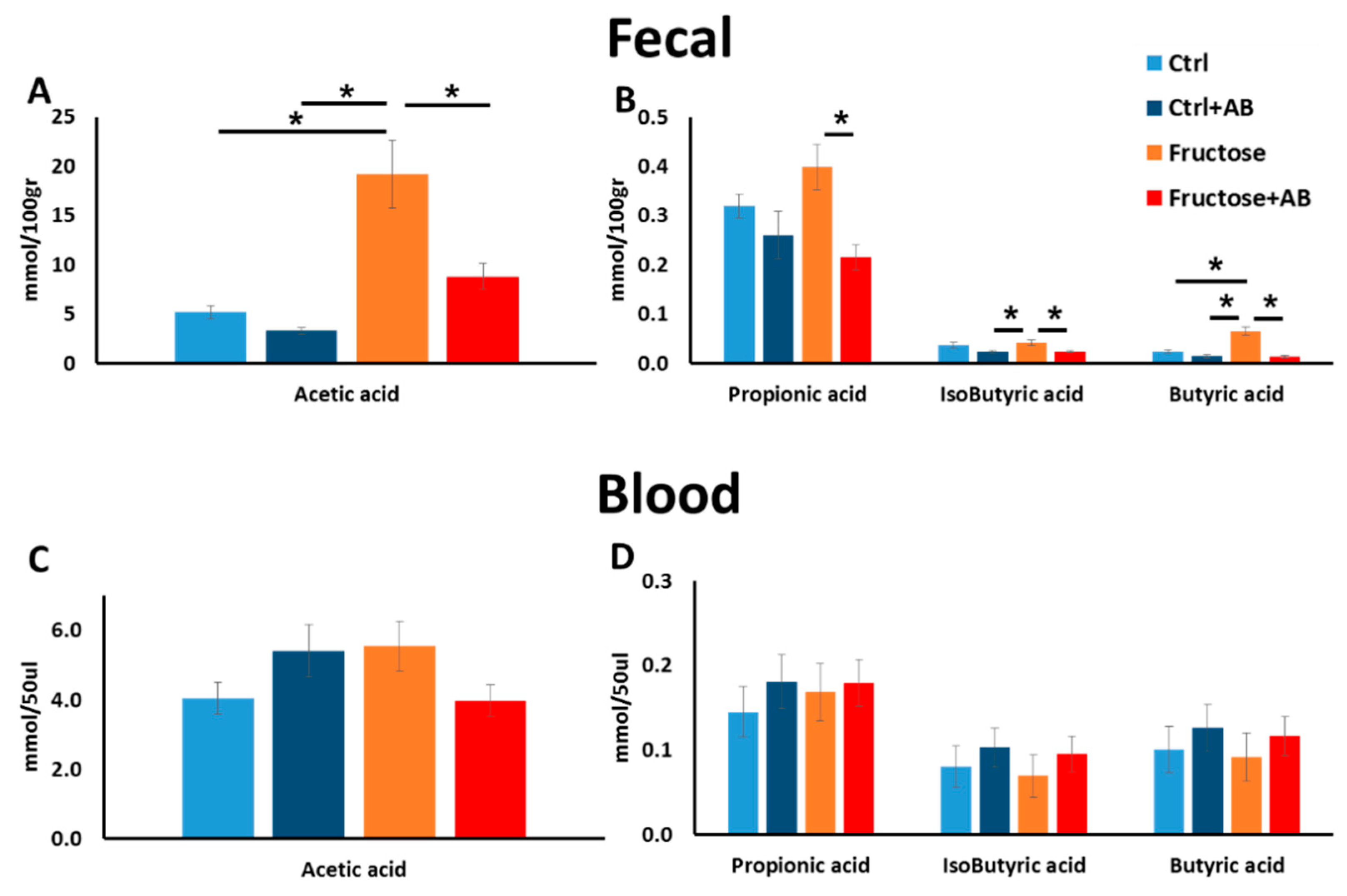
3.3. SCFA
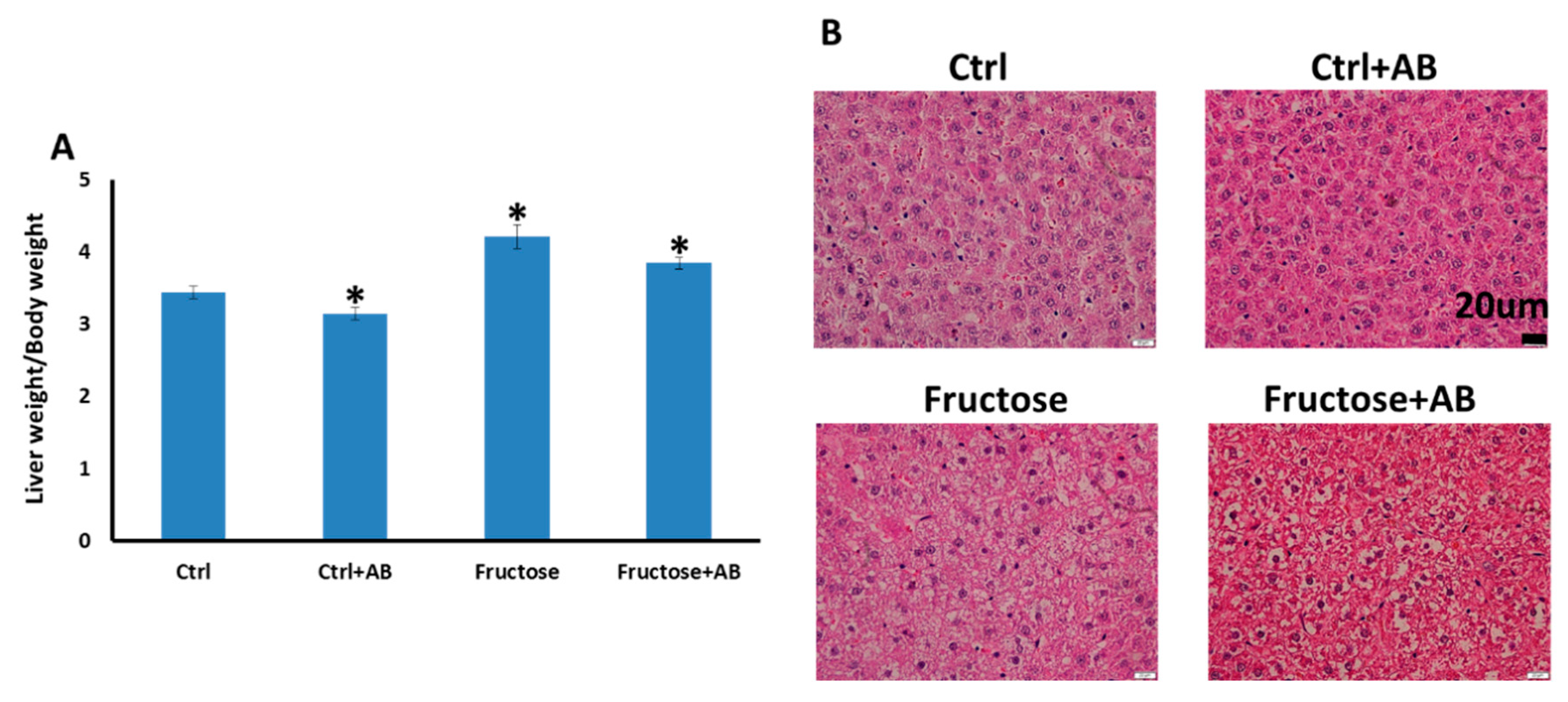
3.3.1. Fatty Liver Phenotype
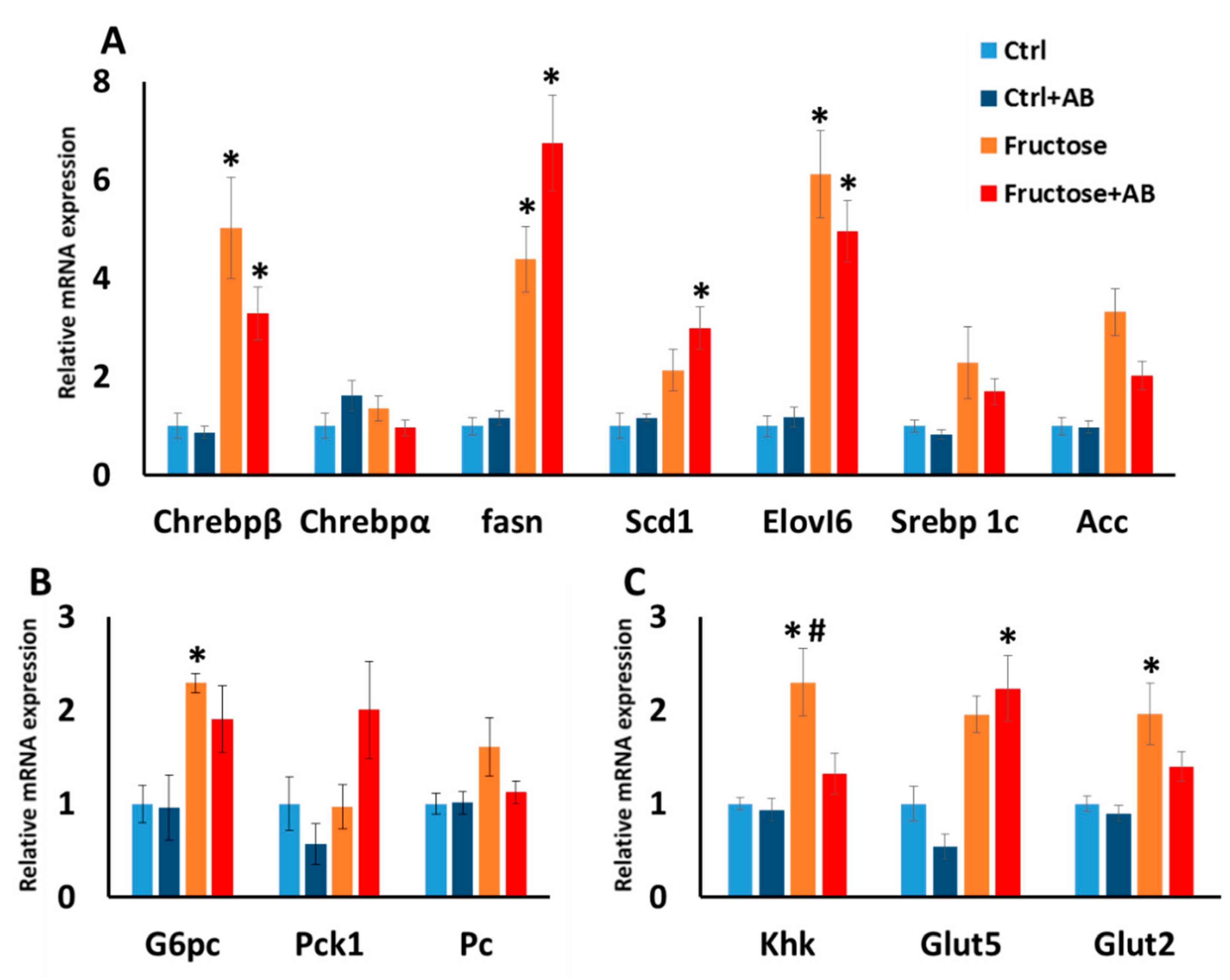
3.3.2. Hepatic Gene Expression
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yerlikaya, A.; Dagel, T.; King, C.; Kubawara, M.; Lanaspa, M.A.; Andres-Hernando, A.; Covic, A.; Manitius, J.; Sag, A.A.; Kanbay, M. Dietary and commercialized fructose: Sweet or sour? Int. Urol. Nephrol. 2017, 49, 1611–1620. [Google Scholar] [CrossRef]
- Heidenreich, P.A.; Trogdon, J.G.; Khavjou, O.A.; Butler, J.; Dracup, K.; Ezekowitz, M.D.; Finkelstein, E.A.; Hong, Y.; Johnston, S.C.; Khera, A.; et al. Forecasting the future of cardiovascular disease in the United States: A policy statement from the American Heart Association. Circulation 2011, 123, 933–944. [Google Scholar] [CrossRef]
- Benjamin, E.J.; Blaha, M.J.; Chiuve, S.E.; Cushman, M.; Das, S.R.; Deo, R.; de Ferranti, S.D.; Floyd, J.; Fornage, M.; Gillespie, C.; et al. Heart Disease and Stroke Statistics—2017 Update: A Report From the American Heart Association. Circulation 2017, 135, e146–e603. [Google Scholar] [CrossRef]
- Prabhakaran, D.; Roy, A.; Praveen, P.A.; Ramakrishnan, L.; Gupta, R.; Amarchand, R.; Kondal, D.; Singh, K.; Sharma, M.; Shukla, D.K.; et al. 20-Year Trend of CVD Risk Factors: Urban and Rural National Capital Region of India. Glob. Heart 2017, 12, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Dekker, M.J.; Su, Q.; Baker, C.; Rutledge, A.C.; Adeli, K. Fructose: A highly lipogenic nutrient implicated in insulin resistance, hepatic steatosis, and the metabolic syndrome. Am. J. Physiol. Endocrinol. Metab. 2010, 299, E685–E694. [Google Scholar] [CrossRef]
- Stanhope, K.L.; Schwarz, J.M.; Keim, N.L.; Griffen, S.C.; Bremer, A.A.; Graham, J.L.; Hatcher, B.; Cox, C.L.; Dyachenko, A.; Zhang, W.; et al. Consuming fructose-sweetened, not glucose-sweetened, beverages increase visceral adiposity and lipids and decrease insulin sensitivity in overweight/obese men. J. Clin. Investig. 2009, 1334, 1322–1334. [Google Scholar] [CrossRef]
- Stanhope, K.L.; Schwarz, J.M.; Havel, P.J. Adverse metabolic effects of dietary fructose: Results from the recent epidemiological, clinical, and mechanistic studies. Curr. Opin. Lipidol. 2013, 24, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Hannou, S.A.; Haslam, D.E.; McKeown, N.M.; Herman, M.A. Fructose metabolism and metabolic disease. J. Clin. Investig. 2018, 128, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Zmora, N.; Suez, J.; Elinav, E. You are what you eat: Diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 35–56. [Google Scholar] [CrossRef] [PubMed]
- Do, M.H.; Lee, E.; Oh, M.J.; Kim, Y.; Park, H.Y. High-Glucose or -Fructose Diet Cause Changes of the Gut Microbiota and Metabolic Disorders in Mice without Body Weight Change. Nutrients 2018, 10, 761. [Google Scholar] [CrossRef]
- Mastrocola, R.; Ferrocino, I.; Liberto, E.; Chiazza, F.; Cento, A.S.; Collotta, D.; Querio, G.; Nigro, D.; Bitonto, V.; Cutrin, J.C.; et al. Fructose liquid and solid formulations differently affect gut integrity, microbiota composition and related liver toxicity: A comparative in vivo study. J. Nutr. Biochem. 2018, 55, 185–199. [Google Scholar] [CrossRef] [PubMed]
- Di Luccia, B.; Crescenzo, R.; Mazzoli, A.; Cigliano, L.; Venditti, P.; Walser, J.C.; Widmer, A.; Baccigalupi, L.; Ricca, E.; Iossa, S. Rescue of fructose-induced metabolic syndrome by antibiotics or faecal transplantation in a rat model of obesity. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Sellmann, C.; Priebs, J.; Landmann, M.; Degen, C.; Engstler, A.J.; Jin, C.J.; Garttner, S.; Spruss, A.; Huber, O.; Bergheim, I. Diets rich in fructose, fat or fructose and fat alter intestinal barrier function and lead to the development of nonalcoholic fatty liver disease over time. J. Nutr. Biochem. 2015, 26, 1183–1192. [Google Scholar] [CrossRef] [PubMed]
- Volynets, V.; Louis, S.; Pretz, D.; Lang, L.; Ostaff, M.J.; Wehkamp, J.; Bischoff, S.C. Intestinal Barrier Function and the Gut Microbiome Are Differentially Affected in Mice Fed a Western-Style Diet or Drinking Water Supplemented with Fructose. J. Nutr. 2017, 147, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Bergheim, I.; Weber, S.; Vos, M.; Kramer, S.; Volynets, V.; Kaserouni, S.; McClain, C.J.; Bischoff, S.C. Antibiotics protect against fructose-induced hepatic lipid accumulation in mice: role of endotoxin. J. Hepatol. 2008, 48, 983–992. [Google Scholar] [CrossRef] [PubMed]
- Crescenzo, R.; Mazzoli, A.; Di Luccia, B.; Bianco, F.; Cancelliere, R.; Cigliano, L.; Liverini, G.; Baccigalupi, L.; Iossa, S. Dietary fructose causes defective insulin signalling and ceramide accumulation in the liver that can be reversed by gut microbiota modulation. Food Nutr. Res. 2017, 61, 1331657. [Google Scholar] [CrossRef] [PubMed]
- Li, J.M.; Yu, R.; Zhang, L.P.; Wen, S.Y.; Wang, S.J.; Zhang, X.Y.; Xu, Q.; Kong, L.D. Dietary fructose-induced gut dysbiosis promotes mouse hippocampal neuroinflammation: A benefit of short-chain fatty acids. Microbiome 2019, 7, 98. [Google Scholar] [CrossRef]
- Sanchez-Lozada, L.G.; Tapia, E.; Jimenez, A.; Bautista, P.; Cristobal, M.; Nepomuceno, T.; Soto, V.; Avila-Casado, C.; Nakagawa, T.; Johnson, R.J.; et al. Fructose-induced metabolic syndrome is associated with glomerular hypertension and renal microvascular damage in rats. Am. J. Physiol. Renal. Physiol. 2007, 292, F423–F429. [Google Scholar] [CrossRef]
- Wong, S.K.; Chin, K.Y.; Suhaimi, F.H.; Fairus, A.; Ima-Nirwana, S. Animal models of metabolic syndrome: A review. Nutr. Metab. 2016, 13, 65. [Google Scholar] [CrossRef]
- Kucera, O.; Cervinkova, Z. Experimental models of non-alcoholic fatty liver disease in rats. World J. Gastroenterol. 2014, 20, 8364–8376. [Google Scholar] [CrossRef]
- Kawasaki, T.; Igarashi, K.; Koeda, T.; Sugimoto, K.; Nakagawa, K.; Hayashi, S.; Yamaji, R.; Inui, H.; Fukusato, T.; Yamanouchi, T. Rats fed fructose-enriched diets have characteristics of nonalcoholic hepatic steatosis. J. Nutr. 2009, 139, 2067–2071. [Google Scholar] [CrossRef] [PubMed]
- Ferrier, L.; Berard, F.; Debrauwer, L.; Chabo, C.; Langella, P.; Bueno, L.; Fioramonti, J. Impairment of the intestinal barrier by ethanol involves enteric microflora and mast cell activation in rodents. Am. J. Pathol. 2006, 168, 1148–1154. [Google Scholar] [CrossRef] [PubMed]
- Kamari, Y.; Shaish, A.; Vax, E.; Shemesh, S.; Kandel-Kfir, M.; Arbel, Y.; Olteanu, S.; Barshack, I.; Dotan, S.; Voronov, E.; et al. Lack of interleukin-1alpha or interleukin-1beta inhibits transformation of steatosis to steatohepatitis and liver fibrosis in hypercholesterolemic mice. J. Hepatol. 2011, 55, 1086–1094. [Google Scholar] [CrossRef] [PubMed]
- Braun, T.; Di Segni, A.; BenShoshan, M.; Asaf, R.; Squires, J.E.; Farage Barhom, S.; Glick Saar, E.; Cesarkas, K.; Smollan, G.; Weiss, B.; et al. Fecal microbial characterization of hospitalized patients with suspected infectious diarrhea shows significant dysbiosis. Sci. Rep. 2017, 7, 1088. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef]
- Flores, G.E.; Henley, J.B.; Fierer, N. A direct PCR approach to accelerate analyses of human-associated microbial communities. PLoS ONE 2012, 7, e44563. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Amir, A.; McDonald, D.; Navas-Molina, J.A.; Kopylova, E.; Morton, J.T.; Zech Xu, Z.; Kightley, E.P.; Thompson, L.R.; Hyde, E.R.; Gonzalez, A.; et al. Deblur Rapidly Resolves Single-Nucleotide Community Sequence Patterns. mSystems 2017, 2. [Google Scholar] [CrossRef]
- Lozupone, C.; Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef]
- Mirarab, S.; Nguyen, N.; Warnow, T. SEPP: SATe-enabled phylogenetic placement. Pac. Symp. Biocomput. 2012, 247–258. [Google Scholar] [CrossRef]
- Xu, Z.Z.; Amir, A.; Sanders, J.; Zhu, Q.; Morton, J.T.; Bletz, M.C.; Tripathi, A.; Huang, S.; McDonald, D.; Jiang, L.; et al. Calour: An Interactive, Microbe-Centric Analysis Tool. mSystems 2019, 4. [Google Scholar] [CrossRef]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; LeLeiko, N.; Snapper, S.B.; et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 2012, 13, R79. [Google Scholar] [CrossRef]
- Cho, Y.E.; Kim, D.K.; Seo, W.; Gao, B.; Yoo, S.H.; Song, B.J. Fructose Promotes Leaky Gut, Endotoxemia, and Liver Fibrosis Through Ethanol-Inducible Cytochrome P450-2E1-Mediated Oxidative and Nitrative Stress. Hepatology 2019. [Google Scholar] [CrossRef]
- Kim, M.S.; Krawczyk, S.A.; Doridot, L.; Fowler, A.J.; Wang, J.X.; Trauger, S.A.; Noh, H.L.; Kang, H.J.; Meissen, J.K.; Blatnik, M.; et al. ChREBP regulates fructose-induced glucose production independently of insulin signaling. J. Clin. Investig. 2016, 126, 4372–4386. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Astapova, I.I.; Flier, S.N.; Hannou, S.A.; Doridot, L.; Sargsyan, A.; Kou, H.H.; Fowler, A.J.; Liang, G.; Herman, M.A. Intestinal, but not hepatic, ChREBP is required for fructose tolerance. JCI Insight 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Burgeiro, A.; Cerqueira, M.G.; Varela-Rodriguez, B.M.; Nunes, S.; Neto, P.; Pereira, F.C.; Reis, F.; Carvalho, E. Glucose and Lipid Dysmetabolism in a Rat Model of Prediabetes Induced by a High-Sucrose Diet. Nutrients 2017, 9, 638. [Google Scholar] [CrossRef] [PubMed]
- Lozano, I.; Van der Werf, R.; Bietiger, W.; Seyfritz, E.; Peronet, C.; Pinget, M.; Jeandidier, N.; Maillard, E.; Marchioni, E.; Sigrist, S.; et al. High-fructose and high-fat diet-induced disorders in rats: Impact on diabetes risk, hepatic and vascular complications. Nutr. Metab. 2016, 13, 15. [Google Scholar] [CrossRef]
- D’Angelo, G.; Elmarakby, A.A.; Pollock, D.M.; Stepp, D.W. Fructose feeding increases insulin resistance but not blood pressure in Sprague-Dawley rats. Hypertens (Dallas, Tex 1979) 2005, 46, 806–811. [Google Scholar] [CrossRef]
- Jang, C.; Hui, S.; Lu, W.; Cowan, A.J.; Morscher, R.J.; Lee, G.; Liu, W.; Tesz, G.J.; Birnbaum, M.J.; Rabinowitz, J.D. The Small Intestine Converts Dietary Fructose into Glucose and Organic Acids. Cell Metab. 2018, 27, 351–361.e3. [Google Scholar] [CrossRef]
| Control (TD.170308) (g/Kg) | HFrD (TD.89247) (g/Kg) | |||
| Casein | 207 | 207 | ||
| DL-Methionine | 3 | 3 | ||
| Corn starch | 400.15 | --- | ||
| Maltodextrin | 200 | --- | ||
| Fructose | --- | 600 | ||
| Lard | 50 | 50 | ||
| Cellulose | 79.81 | 79.81 | ||
| Mineral mix rogers-harper (170760) | 50 | 50 | ||
| Zinc carbonate | 0.04 | 0.04 | ||
| Vitamin mix Teklad (40,060) | 10 | 10 | ||
| % by weight | % kcal from | % by weight | % kcal from | |
| Protein | 18.3 | 21.4 | 18.3 | 20.2 |
| Carbohydrate | 55.4 | 64.9 | 60.4 | 66.8 |
| fat | 5.2 | 13.7 | 5.2 | 13 |
| Kcal/g | 3.4 | 3.6 | ||
| Gene Name | Forward | Reverse |
|---|---|---|
| Chrebpβ | TCTGCAGATCGCGCGGAG | CTTGTCCCGGCATAGCAAC |
| Chrebpα | TGCATCGATCACAGGTCATT | AGGCTCAAGCATTCGAAGAG |
| Srebp 1c | AGTTCCAGCATGGCTACCAC | GGGGTCTCTCAGTTTCCTGC |
| Scd1 | TGCTCTGGGGGATATTTTACTACC | GAGAAGAAAAAGCCACGGCG |
| Elovl6 | GAGGCGCAGAGAACACGTAG | CGCTTGTTCATCAGATGCCG |
| Fasn | AGCCTGAGCTTGTCCCTAGA | CACTGGTACACTTTCCCGCT |
| Acc | CTTGGGGTGATGCTCCCATT | GCTGGGCTTAAACCCCTCAT |
| G6pc | CGTCACCTGTGAGACTGGAC | ACGACATTCAAGCACCGGAA |
| pck1 | GGATGTGGCCAGGATCGAAA | ATACATGGTGCGGCCTTTCA |
| Pc | CCAAGCAGGTTGGCTATGAGAA | GATGTTTTCCTGCCGCAGCC |
| Khk | ATGGCCATGTTGCCGACTT | TCTGGCAGGTTCGTGTCGTA |
| Glut5 | CATGGTCACGGTTTTTGTGG | AGACGATGCTGACATAGGGC |
| Glut2 | CGCACGCAACATGTCAGAAG | TTATTACCTCTTGAGGTGCATTGA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bier, A.; Khasbab, R.; Haberman, Y.; Braun, T.; Hadar, R.; Sosnovski, K.; Amir, A.; Leibowitz, A.; Grossman, E. Antibiotic Treatment Does Not Ameliorate the Metabolic Changes in Rats Presenting Dysbiosis After Consuming a High Fructose Diet. Nutrients 2020, 12, 203. https://doi.org/10.3390/nu12010203
Bier A, Khasbab R, Haberman Y, Braun T, Hadar R, Sosnovski K, Amir A, Leibowitz A, Grossman E. Antibiotic Treatment Does Not Ameliorate the Metabolic Changes in Rats Presenting Dysbiosis After Consuming a High Fructose Diet. Nutrients. 2020; 12(1):203. https://doi.org/10.3390/nu12010203
Chicago/Turabian StyleBier, Ariel, Rawan Khasbab, Yael Haberman, Tzipi Braun, Rotem Hadar, Katya Sosnovski, Amnon Amir, Avshalom Leibowitz, and Ehud Grossman. 2020. "Antibiotic Treatment Does Not Ameliorate the Metabolic Changes in Rats Presenting Dysbiosis After Consuming a High Fructose Diet" Nutrients 12, no. 1: 203. https://doi.org/10.3390/nu12010203
APA StyleBier, A., Khasbab, R., Haberman, Y., Braun, T., Hadar, R., Sosnovski, K., Amir, A., Leibowitz, A., & Grossman, E. (2020). Antibiotic Treatment Does Not Ameliorate the Metabolic Changes in Rats Presenting Dysbiosis After Consuming a High Fructose Diet. Nutrients, 12(1), 203. https://doi.org/10.3390/nu12010203






