Maternal Dietary Protein Intake Influences Milk and Offspring Gut Microbial Diversity in a Rat (Rattus norvegicus) Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal and Housing Conditions and Diet
2.2. Diets
2.3. Milk and Cecal Microbiota Experiment
2.4. Common Laboratory and Bioinformatics Methods
2.5. Statistical Analyses
3. Results
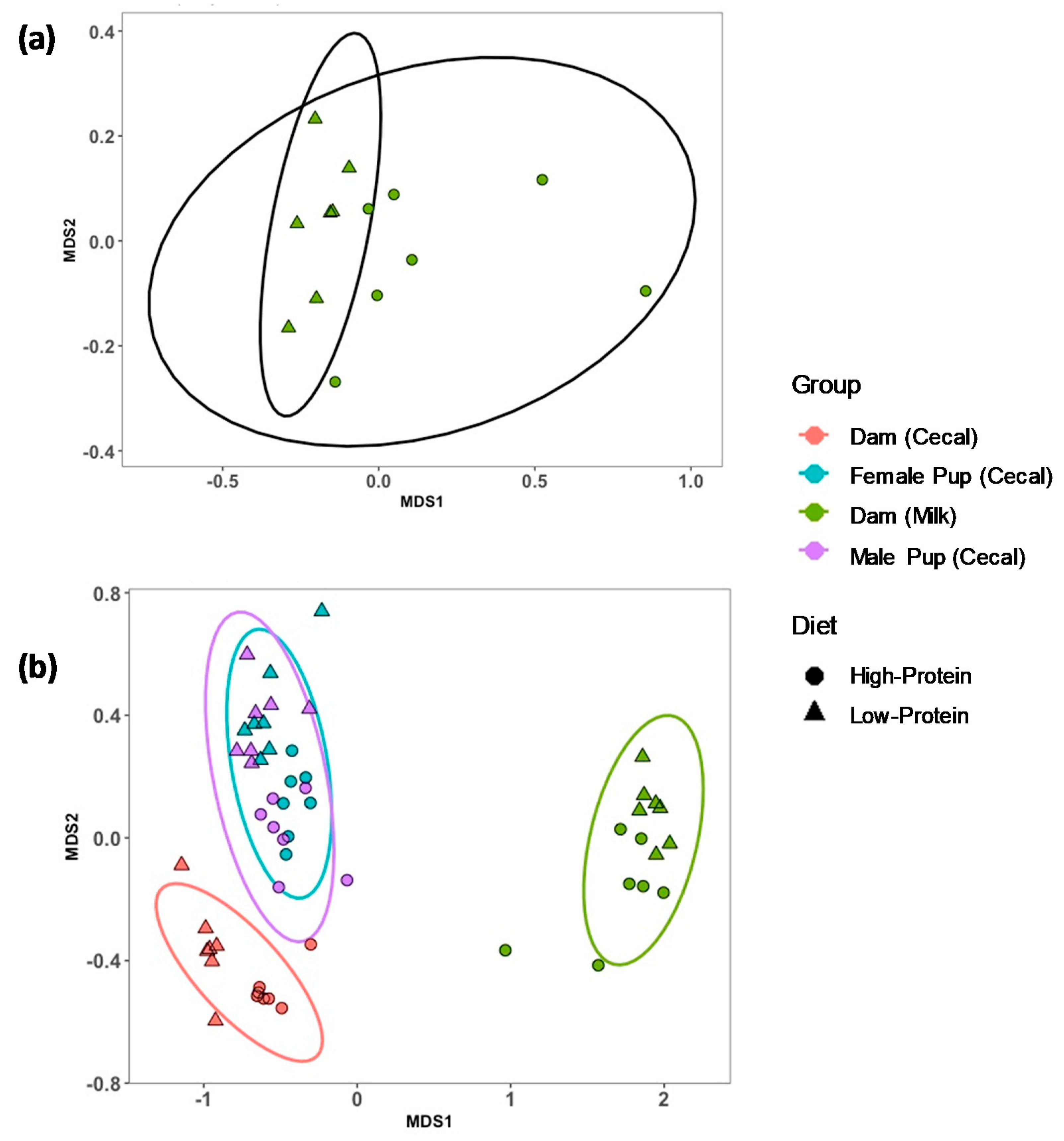
3.1. Alpha and Beta Diversity for Milk, and Dam and Offspring Cecal Contents
3.2. Impact of Maternal Diet on Specific Microorganisms between Maternal Ceca, Milk, and Offspring Ceca
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A. Fecal Microbiota Experiment
Appendix A.1. Methods
Appendix A.2. Results
References
- Sekirov, I.; Finlay, B.B. The role of the intestinal microbiota in enteric infection. J. Physiol. 2009, 587, 4159–4167. [Google Scholar] [CrossRef] [PubMed]
- Khodayar-Pardo, P.; Mira-Pascual, L.; Collado, M.C.; Martínez-Costa, C. Impact of lactation stage, gestational age and mode of delivery on breast milk microbiota. J. Perinatol. 2014, 34, 599. [Google Scholar] [CrossRef] [PubMed]
- Quinn, E.A.; Largado, F.; Power, M.; Kuzawa, C.W. Predictors of breast milk macronutrient composition in filipino mothers. J. Hum. Biol. 2012, 24, 533–540. [Google Scholar] [CrossRef]
- Fernández, L.; Langa, S.; Martín, V.; Maldonado, A.; Jiménez, E.; Martín, R.; Rodríguez, J.M. The human milk microbiota: Origin and potential roles in health and disease. Pharmacol. Res. 2013, 69, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, J.M. The Origin of Human Milk Bacteria: Is There a Bacterial Entero-Mammary Pathway during Late Pregnancy and Lactation? Adv. Nutr. 2014, 5, 779–784. [Google Scholar] [CrossRef] [PubMed]
- Perez, P.F.; Doré, J.; Leclerc, M.; Levenez, F.; Benyacoub, J.; Serrant, P.; Segura-Roggero, I.; Schiffrin, E.J.; Donnet-Hughes, A. Bacterial Imprinting of the Neonatal Immune System: Lessons From Maternal Cells? Pediatrics 2007, 119, e724–e732. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 2004, 303, 1662–1665. [Google Scholar] [CrossRef] [PubMed]
- Hood, W.R.; Voltura, M.B.; Oftedal, O.T. Methods of measuring milk composition and yield in small mammals. In Ecological and Behavioral Methods for the Study of Bats, 2nd ed.; Thomas, H.K., Parsons, S., Eds.; Johns Hopkins University Press: Baltimore, ML, USA, 2009; pp. 529–553. [Google Scholar]
- Butteiger, D.N.; Hibberd, A.A.; McGraw, N.J.; Napawan, N.; Hall-Porter, J.M.; Krul, E.S. Soy Protein Compared with Milk Protein in a Western Diet Increases Gut Microbial Diversity and Reduces Serum Lipids in Golden Syrian Hamsters. J. Nutr. 2016, 146, 697–705. [Google Scholar] [CrossRef] [PubMed]
- Bolnick, D.I.; Snowberg, L.K.; Hirsch, P.E.; Lauber, C.L.; Knight, R.; Caporaso, J.G.; Svanbäck, R. Individuals’ diet diversity influences gut microbial diversity in two freshwater fish (threespine stickleback and Eurasian perch). Ecol. Lett. 2014, 17, 979–987. [Google Scholar] [CrossRef]
- Scholtens, P.A.M.J.; Oozeer, R.; Martin, R.; Amor, K.B.; Knol, J. The Early Settlers: Intestinal Microbiology in Early Life. Annu. Rev. Food Sci. Technol. 2012, 3, 425–447. [Google Scholar] [CrossRef]
- Funkhouser, L.J.; Bordenstein, S.R. Mom knows best: The universality of maternal microbial transmission. PLoS Biol. 2013, 11, e1001631. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.A.; Nachman, M.W. Spatial Heterogeneity of Gut Microbial Composition along the Gastrointestinal Tract in Natural Populations of House Mice. PLoS ONE 2016, 11, e0163720. [Google Scholar] [CrossRef] [PubMed]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: a versatile open source tool for metagenomics. Peer J. 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Legendre, P.; Borcard, D.; Peres-Neto, P.R. Analyzing beta diversity: Partitioning the spatial variation of community composition data. Ecol. Monogr. 2005, 75, 435–450. [Google Scholar] [CrossRef]
- R. R Development Core Team. A language and environment for statistical computing. Computing 2006, 1. [Google Scholar] [CrossRef]
- Ward, T.L.; Hosid, S.; Ioshikhes, I.; Altosaar, I. Human milk metagenome: A functional capacity analysis. BMC Microbiol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Dill-McFarland, K.A.; Weimer, P.J.; Breaker, J.D.; Suen, G. Diet Influences Early Microbiota Development in Dairy Calves without Long-Term Impacts on Milk Production. Appl. Environ. Microbiol. 2019, 85. [Google Scholar] [CrossRef]
- Hermansson, H.; Kumar, H.; Collado, M.C.; Salminen, S.; Isolauri, E.; Rautava, S. Breast Milk Microbiota Is Shaped by Mode of Delivery and Intrapartum Antibiotic Exposure. Front. Nutr. 2019, 6, 4. [Google Scholar] [CrossRef]
- Read, T.; Fortun-Lamothe, L.; Pascal, G.; Le Boulch, M.; Cauquil, L.; Gabinaud, B.; Bannelier, C.; Balmisse, E.; Destombes, N.; Bouchez, O.; et al. Diversity and Co-occurrence Pattern Analysis of Cecal Microbiota Establishment at the Onset of Solid Feeding in Young Rabbits. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Mariat, D.; Firmesse, O.; Levenez, F.; Guimarăes, V.; Sokol, H.; Doré, J.; Corthier, G.; Furet, J. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 2009, 9, 123. [Google Scholar] [CrossRef] [PubMed]
- Agans, R.; Rigsbee, L.; Kenche, H.; Michail, S.; Khamis, H.J.; Paliy, O. Distal gut microbiota of adolescent children is different from that of adults. FEMS Microbiol. Ecol. 2011, 77, 404–412. [Google Scholar] [CrossRef] [PubMed]
- Frese, S.A.; Parker, K.; Calvert, C.C.; Mills, D.A. Diet shapes the gut microbiome of pigs during nursing and weaning. Microbiome 2015, 3, 28. [Google Scholar] [CrossRef] [PubMed]
- Pannaraj, P.S.; Li, F.; Cerini, C.; Bender, J.M.; Yang, S.; Rollie, A.; Adisetiyo, H.; Zabih, S.; Lincez, P.J.; Bittinger, K.; et al. Association Between Breast Milk Bacterial Communities and Establishment and Development of the Infant Gut MicrobiomeAssociation Between Breast Milk Bacterial Communities and Infant Gut MicrobiomeAssociation Between Breast Milk Bacterial Communities and Infant Gut Microbiome. JAMA Pediatrics 2017, 171, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Ramanan, P.; Barreto, J.N.; Osmon, D.R.; Tosh, P.K. Rothia bacteremia: A 10-year experience at Mayo Clinic, Rochester, Minnesota. J. Clin. Microbiol. 2014, 52, 3184–3189. [Google Scholar] [CrossRef] [PubMed]
- Heikkilä, M.; Saris, P. Inhibition of Staphylococcus aureus by the commensal bacteria of human milk. J. Appl. Microbiol. 2003, 95, 471–478. [Google Scholar] [CrossRef]
- Könönen, E. Development of oral bacterial flora in young children. Ann. Med. 2000, 32, 107–112. [Google Scholar] [CrossRef]
- Hoekema, A.; Hirsch, P.; Hooykaas, P.; Schilperoort, R. A binary plant vector strategy based on separation of vir-and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 1983, 303, 179–180. [Google Scholar] [CrossRef]
- McInnis, E.A.; Kalanetra, K.M.; Mills, D.A.; Maga, E.A. Analysis of raw goat milk microbiota: Impact of stage of lactation and lysozyme on microbial diversity. Food Microbiol. 2015, 46, 121–131. [Google Scholar] [CrossRef]
- Ringø, E.; Olsen, R. The effect of diet on aerobic bacterial flora associated with intestine of Arctic charr (Salvelinus alpinus L.). J. Appl. Microbiol. 1999, 86, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Olivares, M.; Marín, M.; Xaus, J.; Fernández, L.; Rodríguez, J. Characterization of a reuterin-producing Lactobacillus coryniformis strain isolated from a goat’s milk cheese. Int. J. Food Microbiol. 2005, 104, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Nava, G.M.; Friedrichsen, H.J.; Stappenbeck, T.S. Spatial organization of intestinal microbiota in the mouse ascending colon. ISME J. 2010, 5, 627. [Google Scholar] [CrossRef] [PubMed]
- La Reau, A.J.; Suen, G. The Ruminococci: Key symbionts of the gut ecosystem. J. Microbiol. 2018, 56, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Cassir, N.; Benamar, S.; La Scola, B. Clostridium butyricum: From beneficial to a new emerging pathogen. Clin. Microbiol. Infect. 2016, 22, 37–45. [Google Scholar] [CrossRef]
- Di Bella, S.; Ascenzi, P.; Siarakas, S.; Petrosillo, N.; di Masi, A. Clostridium difficile toxins A and B: Insights into pathogenic properties and extraintestinal effects. Toxins 2016, 8, 134. [Google Scholar] [CrossRef]
- Giménez, J.A.; Sugiyama, H. Comparison of toxins of Clostridium butyricum and Clostridium botulinum type E. Infect. Immun. 1988, 56, 926–929. [Google Scholar]
- Liu, X.; Blouin, J.-M.; Santacruz, A.; Lan, A.; Andriamihaja, M.; Wilkanowicz, S.; Benetti, P.-H.; Tomé, D.; Sanz, Y.; Blachier, F.; et al. High-protein diet modifies colonic microbiota and luminal environment but not colonocyte metabolism in the rat model: The increased luminal bulk connection. Am. J. Physiol. Gastroint. Liver Physiol. 2014, 307, G459–G470. [Google Scholar] [CrossRef]
- Zhou, L.; Fang, L.; Sun, Y.; Su, Y.; Zhu, W. Effects of the dietary protein level on the microbial composition and metabolomic profile in the hindgut of the pig. Anaerobe 2016, 38, 61–69. [Google Scholar] [CrossRef]
- Chen, W.; Mi, J.; Lv, N.; Gao, J.; Cheng, J.; Wu, R.; Ma, J.; Lan, T.; Liao, X. Lactation stage-dependency of the sow milk microbiota. Front. Microbiol. 2018, 9, 945. [Google Scholar] [CrossRef]
- Jost, T.; Lacroix, C.; Braegger, C.; Chassard, C. Impact of human milk bacteria and oligosaccharides on neonatal gut microbiota establishment and gut health. Nutr. Rev. 2015, 73, 426–437. [Google Scholar] [CrossRef] [PubMed]
- Liong, M.; Shah, N. Effects of a Lactobacillus casei synbiotic on serum lipoprotein, intestinal microflora, and organic acids in rats. J. Dairy Sci. 2006, 89, 1390–1399. [Google Scholar] [CrossRef]
- Charlier, C.; Cretenet, M.; Even, S.; Le Loir, Y. Interactions between Staphylococcus aureus and lactic acid bacteria: An old story with new perspectives. Int. J. Food Microbiol. 2009, 131, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Middelbos, I.S.; Boler, B.M.V.; Qu, A.; White, B.A.; Swanson, K.S.; Fahey, G.C., Jr. Phylogenetic characterization of fecal microbial communities of dogs fed diets with or without supplemental dietary fiber using 454 pyrosequencing. PLoS ONE 2010, 5, e9768. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Bäckhed, F.; Turnbaugh, P.; Lozupone, C.A.; Knight, R.D.; Gordon, J.I. Obesity alters gut microbial ecology. Proc. Natl. Acad. Sci. USA 2005, 102, 11070–11075. [Google Scholar] [CrossRef]
- Qu, A.; Brulc, J.M.; Wilson, M.K.; Law, B.F.; Theoret, J.R.; Joens, L.A.; Konkel, M.E.; Angly, F.; Dinsdale, E.A.; Edwards, R.A. Comparative metagenomics reveals host specific metavirulomes and horizontal gene transfer elements in the chicken cecum microbiome. PLoS ONE 2008, 3, e2945. [Google Scholar] [CrossRef] [PubMed]
- Field, C.J. The immunological components of human milk and their effect on immune development in infants. J. Nutr. 2005, 135, 1–4. [Google Scholar] [CrossRef]
- Paape, M.J.; Shafer-Weaver, K.; Capuco, A.V.; Van Oostveldt, K.; Burvenich, C. Immune surveillance of mammary tissue by phagocytic cells. In Biology of the Mammary Gland; Springer: Berlin/Heidelberg, Germany, 2002; pp. 259–277. [Google Scholar]
- Merad, M.; Sathe, P.; Helft, J.; Miller, J.; Mortha, A. The dendritic cell lineage: Ontogeny and function of dendritic cells and their subsets in the steady state and the inflamed setting. Annu. Rev. Immunol. 2013, 31, 563–604. [Google Scholar] [CrossRef]
- Worbs, T.; Hammerschmidt, S.I.; Förster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 2017, 17, 30. [Google Scholar] [CrossRef]
- Mastromarino, P.; Capobianco, D.; Miccheli, A.; Praticò, G.; Laforgia, N.; Capursi, T.; Baldassarre, M.E. Administration of a multistrain probiotic product (VSL#3) to women in the perinatal period differentially affects breast milk beneficial microbiota in relation to mode of delivery. Pharmacol. Res. 2015, 95, 63–70. [Google Scholar]
- Moossavi, M.; Sepehri, S.; Robertson, B.; Bode, L.; Goruk, S.; Field, C.J.; Lix, L.M.; de Souza, R.J.; Becker, A.B.; Mandhane, P.J.; et al. Composition and Variation of the Human Milk Microbiota are Influenced by Maternal and Early-Life Factors. Cell Host Microbe 2019, 25, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Derrickson, E.M.; Lowas, S.R. The Effects of Dietary Protein Levels on Milk Protein Levels and Postnatal Growth in Laboratory Mice (Mus musculus). J. Mammal. 2007, 88, 1475–1481. [Google Scholar] [CrossRef]


| Components # | Low-Protein | Moderate-Protein | High-Protein |
|---|---|---|---|
| Dry matter (DM, % as is) | 90.5 | 90.6 | 90.5 |
| Crude protein (% DM) | 10.3 | 15.1 | 20.9 |
| Fat (acid hydrolysis, % DM) | 5.7 | 5.8 | 5.5 |
| Ash (% DM) | 3.9 | 4.7 | 6.0 |
| Crude fiber (% DM) | 4.4 | 4.5 | 4.8 |
| Neutral detergent fiber (% fiber) | 16.7 | 16.0 | 16.3 |
| Acid detergent fiber (% fiber) | 5.3 | 5.5 | 5.9 |
| Carbohydrates (% DM) | 66.2 | 60.5 | 53.3 |
| Starch | 43.1 | 36.7 | 33.9 |
| Glucose | 0.4 | 0.3 | 0.2 |
| Fructose | 0.4 | 0.3 | 0.2 |
| Sucrose | 2.8 | 3.0 | 3.2 |
| Lactose | 1.3 | 1.3 | 1.3 |
| Metabolizable energy (kcal/g) | 3.5 | 3.5 | 3.4 |
| Phyla/Family | Genera | p-Value | ||
|---|---|---|---|---|
| Diet | Sample Type | Interaction | ||
| Actinobacteria | 0.290 | <0.001 | 0.41 | |
| Micrococcaceae | Rothia | 0.380 | <0.001 | 0.39 |
| Bacteroidetes | 0.500 | <0.001 | 0.88 | |
| Bacteroidaceae | Bacteroides | 0.480 | <0.001 | 0.31 |
| Porphyromonadaceae | Parabacteroides | 0.630 | <0.001 | 0.91 |
| Rikenellaceae | Rikenellaceae | 0.970 | <0.001 | 0.2 |
| S24-7 | S24-7 | 0.980 | <0.001 | 0.24 |
| Firmicutes | 0.005 | <0.001 | 0.42 | |
| Planococcaceae | Planococcaceae * | 0.160 | <0.001 | 0.06 |
| Staphylococcaceae | Staphylococcus | 0.370 | 0.002 | 0.32 |
| Lactobacillaceae | Lactobacillus | 0.014 | <0.001 | 0.59 |
| Streptococcaceae | Streptococcus | 0.052 | <0.001 | 0.16 |
| Clostridiales | Clostridiales & | 0.300 | <0.001 | 0.013 |
| Lachnospiraceae | Lachnospiraceae * | 0.440 | <0.001 | 0.08 |
| Ruminococcaceae | Ruminococcaceae * | 0.001 | <0.001 | 0.07 |
| Ruminococcaceae | Oscillospira | 0.570 | <0.001 | 0.2 |
| Ruminococcaceae | Ruminococcus | 0.003 | <0.001 | 0.026 |
| Proteobacteria | 0.910 | <0.001 | 0.97 | |
| Rhizobiaceae | Agrobacterium | 1 | <0.001 | 1 |
| Comamonadaceae | Delftia | 0.340 | <0.001 | 0.26 |
| Enterobacteriaceae | Enterobacteriaceae | 0.600 | 0.001 | 0.85 |
| Pasteurellaceae | Other | 0.840 | <0.001 | 0.93 |
| Pasteurellaceae | Haemophilus | 0.044 | <0.001 | 0.062 |
| Pasteurellaceae | Mannheimia | 0.005 | <0.001 | <0.001 |
| Moraxellaceae | Acinetobacter | 0.740 | 0.068 | 0.43 |
| Xanthomonadaceae | Xanthomonadaceae * | 0.480 | <0.001 | 0.48 |
| Verrucomicrobia | 0.002 | <0.001 | <0.001 | |
| Verrucomicrobiaceae | Akkermansia | 0.002 | <0.001 | <0.001 |
| Phyla/Family | Genera | Maternal Cecal | Milk | Offspring Cecal | |||
|---|---|---|---|---|---|---|---|
| LP | HP | LP | HP | LP | HP | ||
| Actinobacteria | 1.4 ± 1.1% | 0.1 ± 0.0% | 26.0 ± 0.0% | 19.2 ± 0.1% | 1.3 ± 0.2% | 1.0 ± 0.2% | |
| Micrococcaceae | Rothia | neg | neg | 24.8 ± 4.3% | 18.2 ± 7.0% | 1.1 ± 0.2% | 0.9 ± 0.2% |
| Bacteroidetes | 24.8 ± 3.5% | 21.5 ± 3.6% | neg | neg | 30.8 ± 3.0% | 29.1 ± 2.7% | |
| Bacteroidaceae | Bacteroides | 4.2 ± 0.9% | 7.2 ± 1.4% | neg | neg | 16.1 ± 3.6% | 11.7 ± 1.7% |
| Porphyromonadaceae | Parabacteroides | neg | neg | absent | absent | 10.6 ± 2.0% | 11.9 ± 1.8% |
| Rikenellaceae | Rikenellaceae | 5.8 ± 1.1% | 4.2 ± 1.2% | neg | neg | 0.3 ± 0.1% | 1.1 ± 0.7% |
| S24-7 | S24-7 | 11.9 ± 2.2% | 9.4 ± 1.7% | absent | absent | 3.2 ± 0.5% | 4.4 ± 1.0% |
| Firmicutes | 66.1 ± 2.8% | 71.9 ± 1.5% | 33.9 ± 0.1% | 39.7 ± 0.1% | 39.2 ± 4.2% | 54.9 ± 2.1% | |
| Planococcaceae | Planococcaceae * | neg | neg | 6.7 ± 3.4% | 1.9 ± 0.6% | neg | neg |
| Staphylococcaceae | Staphylococcus | absent | absent | 8.8 ± 5.5% | 3.8 ± 1.7% | neg | neg |
| Lactobacillaceae | Lactobacillus | 0.3 ± 0.0% | 4.7 ± 3.0% | 0.8 ± 0.1% | 12.5 ± 5.1% | 19.6 ± 2.6% | 25.6 ± 3.8% |
| Streptococcaceae | Streptococcus | neg | neg | 15.3 ± 4.8% | 8.4 ± 2.3% | 2.8 ± 0.4% | 1.0 ± 0.1% |
| Clostridiales | Clostridiales & | 26.6 ± 3.3% | 18.6 ± 3.2% | 0.2 ± 0.0% | 3.0 ± 1.9% | 2.8 ± 0.7% | 3.4 ± 0.9% |
| Lachnospiraceae | Lachnospiraceae * | 9.3 ± 2.1% | 6.8 ± 1.8% | 0.1 ± 0.0% | 1.7 ± 0.9% | 1.5 ± 0.2% | 3.2 ± 0.6% |
| Ruminococcaceae | Ruminococcaceae * | 6.1 ± 1.2% | 14.4 ± 2.6% | 0.1 ± 0.0% | 1.6 ± 0.8% | 3.4 ± 0.6% | 8.9 ± 1.3% |
| Ruminococcaceae | Oscillospira | 10.5 ± 1.1% | 7.4 ± 1.5% | neg | neg | neg | neg |
| Ruminococcaceae | Ruminococcus | 3.5 ± 0.8% | 6.2 ± 1.3% | neg | neg | neg | neg |
| Proteobacteria | 3.3 ± 0.6% | 5.3 ± 3.2% | 39.5 ± 0.1% | 38.9 ± 0.1% | 13.1 ± 2.3% | 13.1 ± 3.5% | |
| Rhizobiaceae | Agrobacterium | absent | absent | 18.8 ± 6.4% | 18.9 ± 6.1% | absent | absent |
| Comamonadaceae | Delftia | absent | absent | 1.0 ± 0.4% | 1.6 ± 0.5% | absent | absent |
| Enterobacteriaceae | Enterobacteriaceae | neg | neg | neg | neg | 6.5 ± 2.3% | 8.6 ± 2.8% |
| Pasteurellaceae | Other | absent | absent | 4.4 ± 1.1% | 5.3 ± 3.9% | neg | neg |
| Pasteurellaceae | Haemophilus | neg | neg | 0.8 ± 0.3% | 2.2 ± 0.9% | neg | neg |
| Pasteurellaceae | Mannheimia | absent | neg | 5.3 ± 1.3% | 2.3 ± 0.6% | neg | neg |
| Moraxellaceae | Acinetobacter | 0.3 ± 0.1% | 3.6 ± 3.4% | neg | neg | 4.4 ± 1.2% | 3.7 ± 1.5% |
| Xanthomonadaceae | Xanthomonadaceae * | absent | absent | 4.68 ± 1.32% | 3.54 ± 0.98% | absent | absent |
| Verrucomicrobia | 1.4 ± 0.8% | 0.01 ± 0.0% | absent | absent | 15.2 ± 3.1% | 0.4 ± 0.2% | |
| Verrucomicrobiaceae | Akkermansia | 1.4 ± 0.8% | 0.01 ± 0.0% | absent | absent | 15.2 ± 3.1% | 0.4 ± 0.2% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Warren, M.F.; Hallowell, H.A.; Higgins, K.V.; Liles, M.R.; Hood, W.R. Maternal Dietary Protein Intake Influences Milk and Offspring Gut Microbial Diversity in a Rat (Rattus norvegicus) Model. Nutrients 2019, 11, 2257. https://doi.org/10.3390/nu11092257
Warren MF, Hallowell HA, Higgins KV, Liles MR, Hood WR. Maternal Dietary Protein Intake Influences Milk and Offspring Gut Microbial Diversity in a Rat (Rattus norvegicus) Model. Nutrients. 2019; 11(9):2257. https://doi.org/10.3390/nu11092257
Chicago/Turabian StyleWarren, Matthew F., Haley A. Hallowell, Keah V. Higgins, Mark R. Liles, and Wendy R. Hood. 2019. "Maternal Dietary Protein Intake Influences Milk and Offspring Gut Microbial Diversity in a Rat (Rattus norvegicus) Model" Nutrients 11, no. 9: 2257. https://doi.org/10.3390/nu11092257
APA StyleWarren, M. F., Hallowell, H. A., Higgins, K. V., Liles, M. R., & Hood, W. R. (2019). Maternal Dietary Protein Intake Influences Milk and Offspring Gut Microbial Diversity in a Rat (Rattus norvegicus) Model. Nutrients, 11(9), 2257. https://doi.org/10.3390/nu11092257





