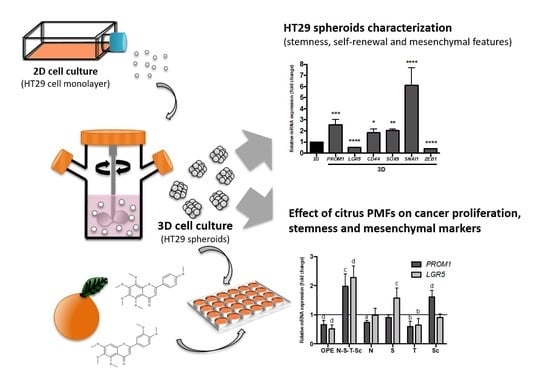
Polymethoxylated Flavones Target Cancer Stemness and Improve the Antiproliferative Effect of 5-Fluorouracil in a 3D Cell Model of Colorectal Cancer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Standard PMFs
2.2. Orange Peel Extract (OPE)
2.3. Cell Lines and Culture
2.4. 3D Cell Culture Using A Stirred-Tank Culture System
2.5. Antiproliferative Assay in HT29 Spheroids
2.6. Interaction Studies—Combination Assays of OPE/PMFs with 5-FU
2.7. Detection of ALDH1 Activity
2.8. Soft Agar Colony-Forming Unit Assay
2.9. Expression Analysis of Genes Involved in EMT, Cancer Stemness, and Wnt/β-Catenin Signaling
2.9.1. RNA Extraction and Reverse Transcription
2.9.2. Real-Time Polymerase Chain Reaction (qPCR)
2.10. Statistical Analysis
3. Results and Discussion
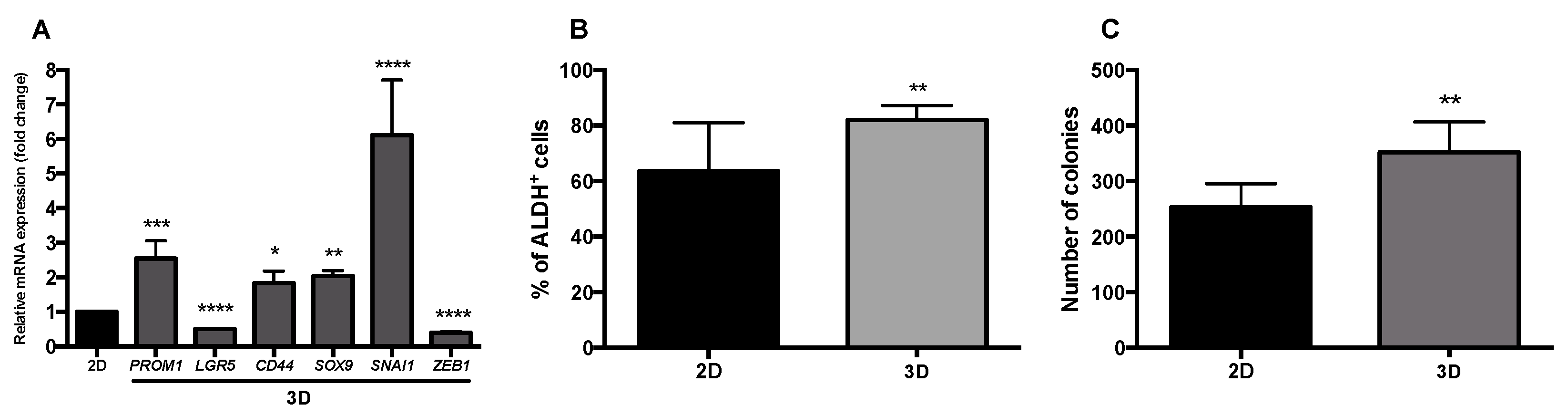
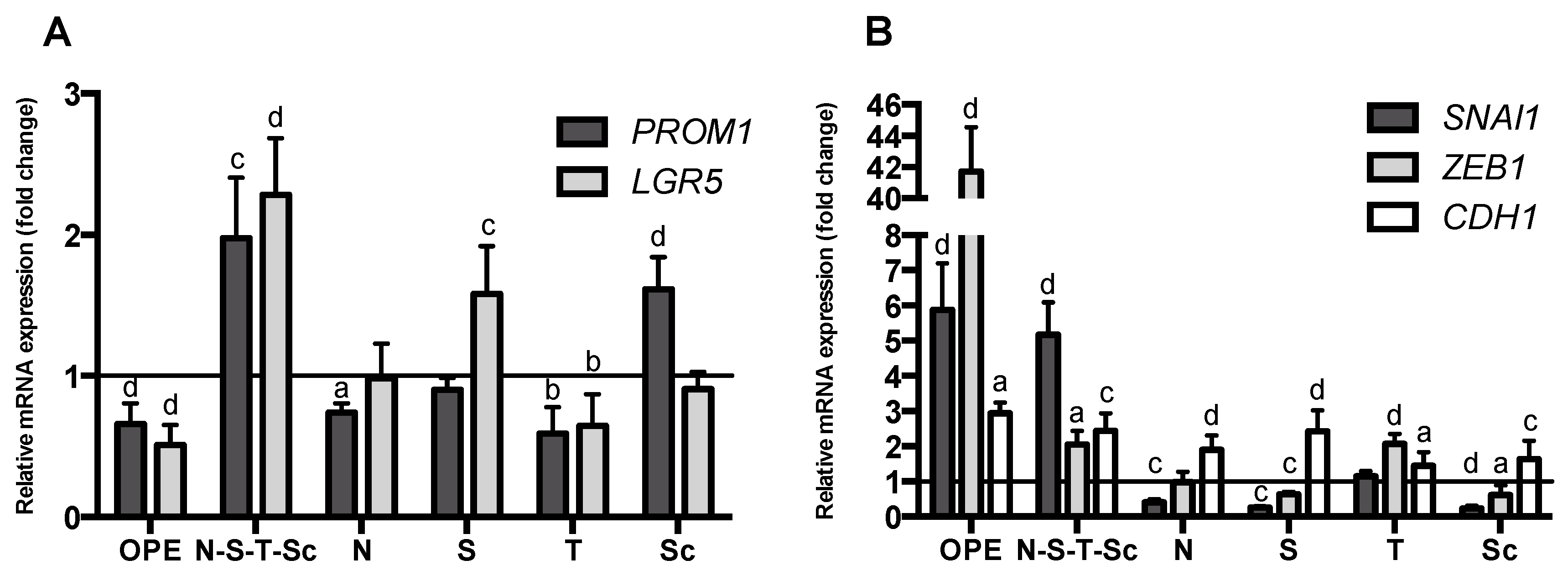
3.1. Characterization of Stemness, Self-Renewal, and Mesenchymal Features of HT29 Spheroids Cultured in Stirred Conditions
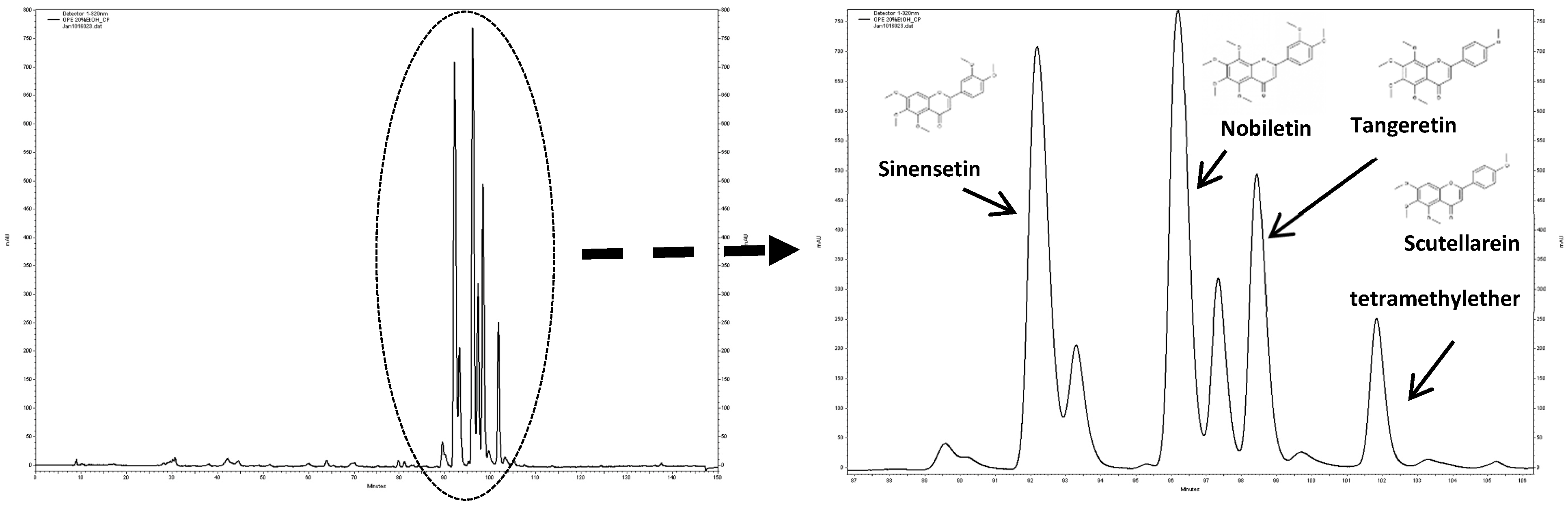
3.2. Characterization of the PMF Content of the OPE Extract
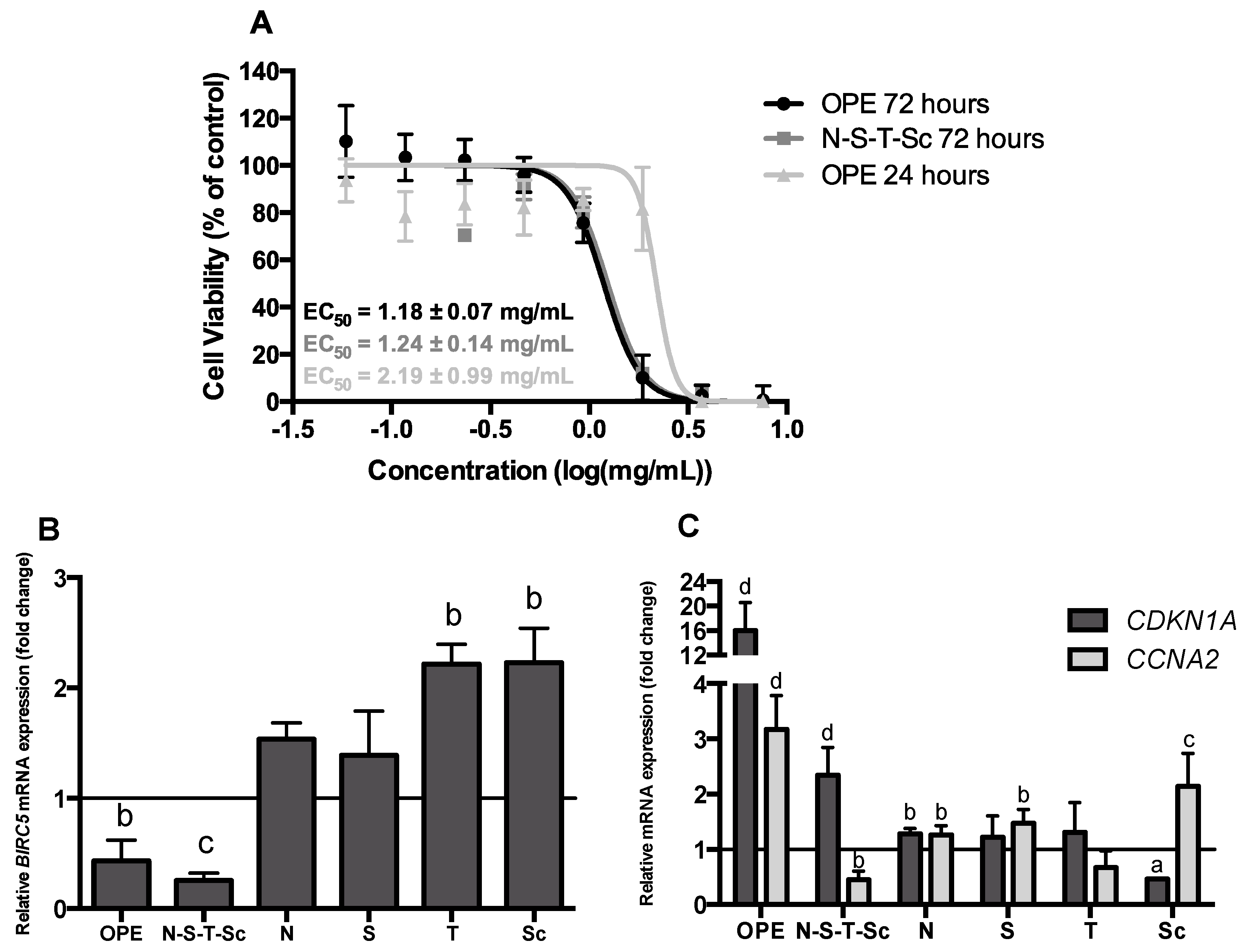
3.3. Effect of OPE and PMFs Modulating Cell Proliferation in a 3D CRC Cell Model
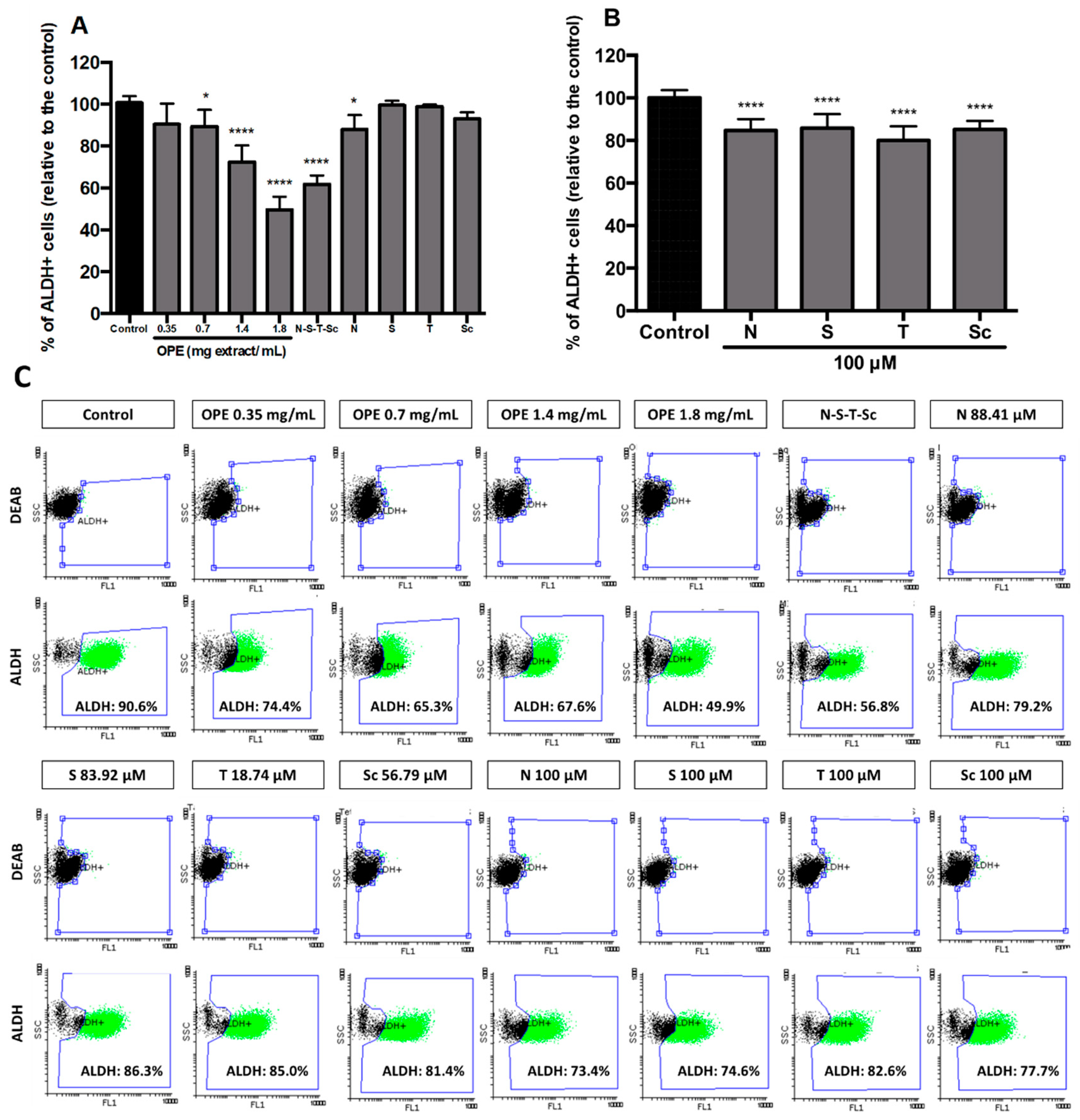
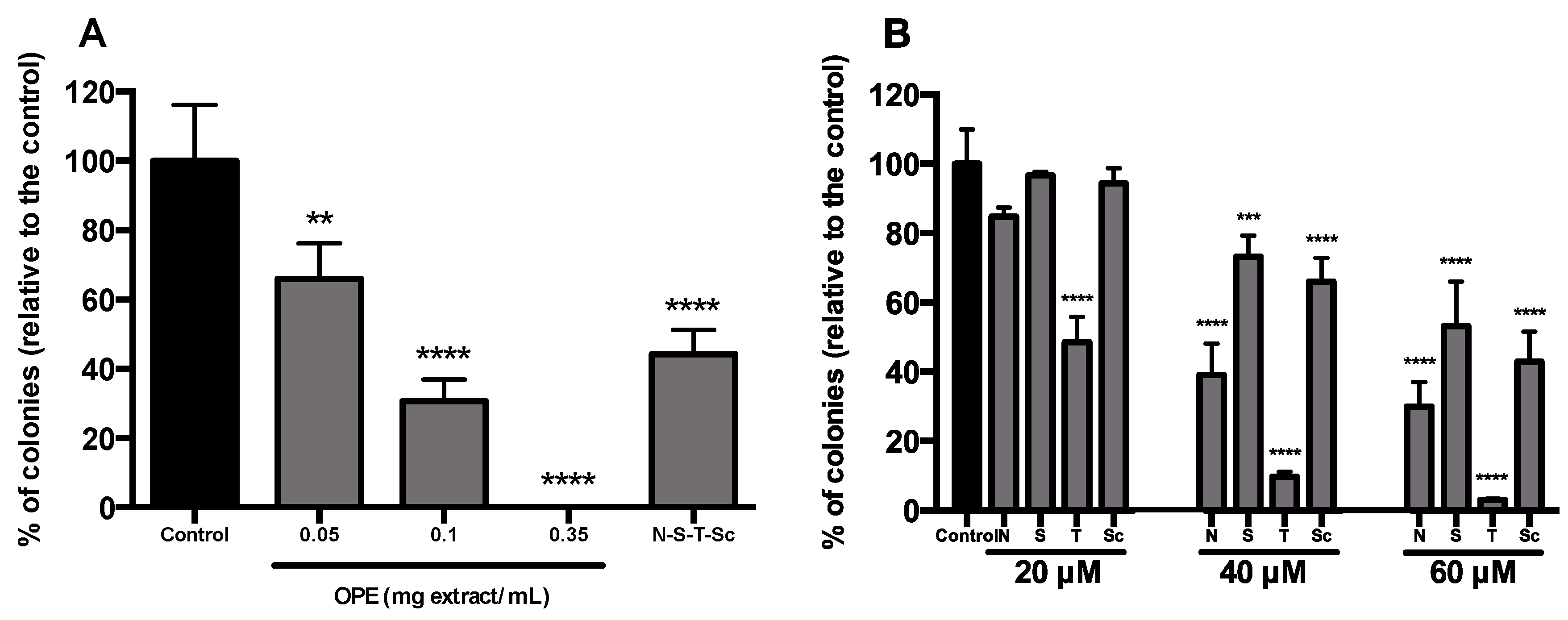
3.4. Effect of OPE and PMFs in Modulating Cancer Stemness and Self-Renewal in a 3D Cell Model of CRC
3.5. Effect of OPE and PMFs Combined with Chemotherapeutical Drug 5-FU
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arnold, M.; Sierra, M.S.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global patterns and trends in colorectal cancer incidence and mortality. Gut 2017, 66, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Reya, T.; Morrison, S.J.; Clarke, M.F.; Weissman, I.L. Stem cells, cancer, and cancer stem cells. Nature 2001, 414, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Visvader, J.E.; Lindeman, G.J. Cancer stem cells in solid tumours: Accumulating evidence and unresolved questions. Nat. Rev. Cancer 2008, 8, 755–768. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.A.; Kreso, A.; Jamieson, C.H.M. Cancer stem cells and self-renewal. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2010, 16, 3113–3120. [Google Scholar] [CrossRef] [PubMed]
- Mimeault, M.; Hauke, R.; Mehta, P.P.; Batra, S.K. Recent advances in cancer stem/progenitor cell research: Therapeutic implications for overcoming resistance to the most aggressive cancers. J. Cell. Mol. Med. 2007, 11, 981–1011. [Google Scholar] [CrossRef] [PubMed]
- Dalerba, P.; Dylla, S.J.; Park, I.K.; Liu, R.; Wang, X.; Cho, R.W.; Hoey, T.; Gurney, A.; Huang, E.H.; Simeone, D.M.; et al. Phenotypic characterization of human colorectal cancer stem cells. Proc. Natl. Acad. Sci. USA 2007, 104, 10158–10163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, P.; Clanton, D.J.; Snipas, T.S.; Lee, J.; Mitchell, E.; Nguyen, M.L.; Hare, E.; Peach, R.J. Characterization of a subpopulation of colon cancer cells with stem cell-like properties. Int. J. Cancer 2009, 124, 1312–1321. [Google Scholar] [CrossRef] [Green Version]
- Du, L.; Wang, H.; He, L.; Zhang, J.; Ni, B.; Wang, X.; Jin, H.; Cahuzac, N.; Mehrpour, M.; Lu, Y.; et al. CD44 is of Functional Importance for Colorectal Cancer Stem Cells. Clin. Cancer Res. 2008, 14, 6751–6760. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Xu, J.; Zhang, J.; Huang, J. Prognostic role of CD133 expression in colorectal cancer: A meta-analysis. BMC Cancer 2012, 12, 573. [Google Scholar] [CrossRef]
- Haraguchi, N.; Ohkuma, M.; Sakashita, H.; Matsuzaki, S.; Tanaka, F.; Mimori, K.; Kamohara, Y.; Inoue, H.; Mori, M. CD133+CD44+ Population Efficiently Enriches Colon Cancer Initiating Cells. Ann. Surg. Oncol. 2008, 15, 2927–2933. [Google Scholar] [CrossRef]
- Chen, K.; Pan, F.; Jiang, H.; Chen, J.F.; Pei, L.; Xie, F.W.; Liang, H.J. Highly enriched CD133+CD44+ stem-like cells with CD133+CD44high metastatic subset in HCT116 colon cancer cells. Clin. Exp. Metastasis 2011, 28, 751–763. [Google Scholar] [CrossRef] [PubMed]
- Baker, A.-M.; Graham, T.A.; Elia, G.; Wright, N.A.; Rodriguez-Justo, M. Characterization of LGR5 stem cells in colorectal adenomas and carcinomas. Sci. Rep. 2015, 5, 8654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, E.H.; Hynes, M.J.; Zhang, T.; Ginestier, C.; Dontu, G.; Appelman, H.; Fields, J.Z.; Wicha, M.S.; Boman, B.M. Aldehyde Dehydrogenase 1 Is a Marker for Normal and Malignant Human Colonic Stem Cells (SC) and Tracks SC Overpopulation during Colon Tumorigenesis. Cancer Res. 2009, 69, 3382–3389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Núñez-Sánchez, M.Á.; Karmokar, A.; González-Sarrías, A.; García-Villalba, R.; Tomás-Barberán, F.A.; García-Conesa, M.T.; Brown, K.; Espín, J.C. In vivo relevant mixed urolithins and ellagic acid inhibit phenotypic and molecular colon cancer stem cell features: A new potentiality for ellagitannin metabolites against cancer. Food Chem. Toxicol. 2016, 92, 8–16. [Google Scholar] [CrossRef]
- Anderson, E.C.; Hessman, C.; Levin, T.G.; Monroe, M.M.; Wong, M.H. The Role of Colorectal Cancer Stem Cells in Metastatic Disease and Therapeutic Response. Cancers 2011, 3, 319–339. [Google Scholar] [CrossRef] [Green Version]
- Pistollato, F.; Giampieri, F.; Battino, M. The use of plant-derived bioactive compounds to target cancer stem cells and modulate tumor microenvironment. Food Chem. Toxicol. 2015, 75, 58–70. [Google Scholar] [CrossRef]
- Burnett, J.P.; Lim, G.; Li, Y.; Lim, R.; Paholak, H.J.; McDermott, S.P.; Sun, L.; Tsume, Y.; Bai, S.; Wicha, M.S.; et al. Sulforaphane enhances the anticancer activity of taxanes against triple negative breast cancer by killing cancer stem cells. Cancer Lett. 2017, 394, 52–64. [Google Scholar] [CrossRef]
- Pandey, P.R.; Okuda, H.; Watabe, M.; Pai, S.K.; Liu, W.; Kobayashi, A.; Xing, F.; Fukuda, K.; Hirota, S.; Sugai, T.; et al. Resveratrol suppresses growth of cancer stem-like cells by inhibiting fatty acid synthase. Breast Cancer Res. Treat. 2011, 130, 387–398. [Google Scholar] [CrossRef]
- Chung, S.S.; Vadgama, J.V. Curcumin and Epigallocatechin Gallate Inhibit the Cancer Stem Cell Phenotype via Down-regulation of STAT3–NFκB signaling. Anticancer Res. 2015, 35, 39–46. [Google Scholar]
- Fabregat, I.; Malfettone, A.; Soukupova, J. New Insights into the Crossroads between EMT and Stemness in the Context of Cancer. J. Clin. Med. 2016, 5, 37. [Google Scholar] [CrossRef]
- Pradella, D.; Naro, C.; Sette, C.; Ghigna, C. EMT and stemness: Flexible processes tuned by alternative splicing in development and cancer progression. Mol. Cancer 2017, 16, 8. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Lo, C.-Y.; Ho, C.-T. Hydroxylated Polymethoxyflavones and Methylated Flavonoids in Sweet Orange (Citrus sinensis) Peel. J. Agric. Food Chem. 2006, 54, 4176–4185. [Google Scholar] [CrossRef] [PubMed]
- Benavente-García, O.; Castillo, J. Update on uses and properties of citrus flavonoids: New findings in anticancer, cardiovascular, and anti-inflammatory activity. J. Agric. Food Chem. 2008, 56, 6185–6205. [Google Scholar] [CrossRef] [PubMed]
- Rawson, N.E.; Ho, C.-T.; Li, S. Efficacious anti-cancer property of flavonoids from citrus peels. Food Sci. Hum. Wellness 2014, 3, 104–109. [Google Scholar] [CrossRef] [Green Version]
- Manthey, J.A.; Guthrie, N. Antiproliferative Activities of Citrus Flavonoids against Six Human Cancer Cell Lines. J. Agric. Food Chem. 2002, 50, 5837–5843. [Google Scholar] [CrossRef] [PubMed]
- Pan, M.-H.; Chen, W.-J.; Lin-Shiau, S.-Y.; Ho, C.-T.; Lin, J.-K. Tangeretin induces cell-cycle G1 arrest through inhibiting cyclin-dependent kinases 2 and 4 activities as well as elevating Cdk inhibitors p21 and p27 in human colorectal carcinoma cells. Carcinogenesis 2002, 23, 1677–1684. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kawabata, K.; Murakami, A.; Ohigashi, H. Nobiletin, a Citrus Flavonoid, Down-Regulates Matrix Metalloproteinase-7 (matrilysin) Expression in HT-29 Human Colorectal Cancer Cells. Biosci. Biotechnol. Biochem. 2005, 69, 307–314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Wang, J.; Fang, L.; Zheng, Z.; Zhi, D.; Wang, S.; Li, S.; Ho, C.T.; Zhao, H. Anticancer Activities of Citrus Peel Polymethoxyflavones Related to Angiogenesis and Others. BioMed Res. Int. 2014, 2014. [Google Scholar] [CrossRef]
- Silva, I.; Estrada, M.F.; Pereira, C.V.; da Silva, A.B.; Bronze, M.R.; Alves, P.M.; Duarte, C.M.M.; Brito, C.; Serra, A.T. Polymethoxylated flavones from orange peels inhibit cell proliferation in a 3D cell model of human colorectal cancer. Nutr. Cancer 2017, 70, 257–266. [Google Scholar] [CrossRef]
- Santo, V.E.; Estrada, M.F.; Rebelo, S.P.; Abreu, S.; Silva, I.; Pinto, C.; Veloso, S.C.; Serra, A.T.; Boghaert, E.; Alves, P.M.; et al. Adaptable stirred-tank culture strategies for large scale production of multicellular spheroid-based tumor cell models. J. Biotechnol. 2016, 221, 118–129. [Google Scholar] [CrossRef]
- Pereira, L.P.; Silva, P.; Duarte, M.; Rodrigues, L.; Duarte, C.M.M.; Albuquerque, C.; Serra, A.T. Targeting Colorectal Cancer Proliferation, Stemness and Metastatic Potential Using Brassicaceae Extracts Enriched in Isothiocyanates: A 3D Cell Model-Based Study. Nutrients 2017, 9, 4. [Google Scholar] [CrossRef] [PubMed]
- Silva, I. Evaluation of Chemotherapeutic Potential of Natural Extracts Using 3D Models of Colon Cancer. Master’s Thesis, Faculdade de Ciências e Tecnologia, Almada, Portugal, 2013. [Google Scholar]
- Chou, T.C.; Talalay, P. Quantitative analysis of dose-effect relationships: The combined effects of multiple drugs or enzyme inhibitors. Adv. Enzym. Regul. 1984, 22, 27–55. [Google Scholar] [CrossRef]
- Valdés, A.; García-Cañas, V.; Simó, C.; Ibáñez, C.; Micol, V.; Ferragut, J.A.; Cifuentes, A. Comprehensive foodomics study on the mechanisms operating at various molecular levels in cancer cells in response to individual rosemary polyphenols. Anal. Chem. 2014, 86, 9807–9815. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.-M.; Dong, T.T.; Wang, L.L.; Feng, B.; Zhao, H.C.; Fan, X.K.; Zheng, M.H. Suppression of colorectal cancer metastasis by nigericin through inhibition of epithelial-mesenchymal transition. World J. Gastroenterol. 2012, 18, 2640–2648. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, D.; Hu, Y.; Ried, T.; Moll, R.; Gaiser, T. Transcriptome profiling of LGR5 positive colorectal cancer cells. Genom. Data 2014, 2, 212–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hwang, W.; Yang, M.H.; Tsai, M.L.; Lan, H.Y.; Su, S.H.; Chang, S.C.; Teng, H.W.; Yang, S.H.; Lan, Y.T.; Chiou, S.H.; et al. SNAIL Regulates Interleukin-8 Expression, Stem Cell–Like Activity, and Tumorigenicity of Human Colorectal Carcinoma Cells. Gastroenterology 2011, 141, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Jägle, S.; Dertmann, A.; Schrempp, M.; Hecht, A. ZEB1 is neither sufficient nor required for epithelial-mesenchymal transition in LS174T colorectal cancer cells. Biochem. Biophys. Res. Commun. 2017, 482, 1226–1232. [Google Scholar] [CrossRef]
- Simpson, C.D.; Anyiwe, K.; Schimmer, A.D. Anoikis resistance and tumor metastasis. Cancer Lett. 2008, 272, 177–185. [Google Scholar] [CrossRef]
- Burrell, R.A.; McGranahan, N.; Bartek, J.; Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 2013, 501, 338–345. [Google Scholar] [CrossRef]
- Fan, X.; Ouyang, N.; Teng, H.; Yao, H. Isolation and characterization of spheroid cells from the HT29 colon cancer cell line. Int. J. Colorectal Dis. 2011, 26, 1279–1285. [Google Scholar] [CrossRef]
- Yeung, T.M.; Gandhi, S.C.; Wilding, J.L.; Muschel, R.; Bodmer, W.F. Cancer stem cells from colorectal cancer-derived cell lines. Proc. Natl. Acad. Sci. USA 2010, 107, 3722–3727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodriguez, F.J.; Lewis-Tuffin, L.J.; Anastasiadis, P.Z. E-cadherin’s dark side: Possible role in tumor progression. Biochim. Biophys. Acta 2012, 1826, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Jolly, M.K.; Boareto, M.; Huang, B.; Jia, D.; Lu, M.; Ben-Jacob, E.; Onuchic, J.N.; Levine, H. Implications of the Hybrid Epithelial/Mesenchymal Phenotype in Metastasis. Front. Oncol. 2015, 5, 155. [Google Scholar] [CrossRef] [PubMed]
- Rafiq, S.; Kaul, R.; Sofi, S.A.; Bashir, N.; Nazir, F.; Nayik, G.A. Citrus peel as a source of functional ingredient: A review. J. Saudi Soc. Agric. Sci. 2016, 17, 351–358. [Google Scholar] [CrossRef]
- Hernandez, J.M.; Farma, J.M.; Coppola, D.; Hakam, A.; Fulp, W.J.; Chen, D.T.; Siegel, E.M.; Yeatman, T.J.; Shibata, D. Expression of the Antiapoptotic Protein Survivin in Colon Cancer. Clin. Colorectal Cancer 2011, 10, 188–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H.; Zhang, X.; Wang, L.; Zheng, G.; Du, L.; Yang, Y.; Dong, Z.; Liu, Y.; Qu, A.; Wang, C. Investigation of cell free BIRC5 mRNA as a serum diagnostic and prognostic biomarker for colorectal cancer. J. Surg. Oncol. 2014, 109, 574–579. [Google Scholar] [CrossRef]
- Pagano, M.; Pepperkok, R.; Verde, F.; Ansorge, W.; Draetta, G. Cyclin A is required at two points in the human cell cycle. EMBO J. 1992, 11, 961–971. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Ambrosini, G.; Chu, E.Y.; Plescia, J.; Tognin, S.; Marchisio, P.C.; Altieri, D.C. Control of apoptosis and mitotic spindle checkpoint by survivin. Nature 1998, 396, 580–584. [Google Scholar] [CrossRef] [PubMed]
- Arakaki, N.; Toyofuku, A.; Emoto, Y.; Nagao, T.; Kuramoto, Y.; Shibata, H.; Higuti, T. Induction of G1 cell cycle arrest in human umbilical vein endothelial cells by flavone’s inhibition of the extracellular signal regulated kinase cascade. Biochem. Cell Biol. Biochim. Biol. Cell. 2004, 82, 583–588. [Google Scholar] [CrossRef]
- Haddad, A.Q.; Venkateswaran, V.; Viswanathan, L.; Teahan, S.J.; Fleshner, N.E.; Klotz, L.H. Novel antiproliferative flavonoids induce cell cycle arrest in human prostate cancer cell lines. Prostate Cancer Prostatic Dis. 2006, 9, 68–76. [Google Scholar] [CrossRef]
- Reiners, J.J.; Clift, R.; Mathieu, P. Suppression of cell cycle progression by flavonoids: Dependence on the aryl hydrocarbon receptor. Carcinogenesis 1999, 20, 1561–1566. [Google Scholar] [CrossRef] [PubMed]
- Nagappan, A.; Lee, H.J.; Saralamma, V.V.G.; Park, H.S.; Hong, G.E.; Yumnam, S.; Raha, S.; Charles, S.N.; Shin, S.C.; Kim, E.H.; et al. Flavonoids isolated from Citrus platymamma induced G2/M cell cycle arrest and apoptosis in A549 human lung cancer cells. Oncol. Lett. 2016, 12, 1394–1402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, S.Y.; Hyun, J.; Yu, J.-R.; Lim, Y.; Lee, Y.H. 5-Methoxyflavanone induces cell cycle arrest at the G2/M phase, apoptosis and autophagy in HCT116 human colon cancer cells. Toxicol. Appl. Pharmacol. 2011, 254, 288–298. [Google Scholar] [CrossRef] [PubMed]
- Koppaka, V.; Thonpson, D.C.; Chen, Y.; Ellerman, M.; Nicolaou, K.C.; Juvonen, R.O.; Petersen, D.; Deitrich, R.A.; Hurley, T.D.; Vasiliou, V. Aldehyde Dehydrogenase Inhibitors: A Comprehensive Review of the Pharmacology, Mechanism of Action, Substrate Specificity, and Clinical Application. Pharmacol. Rev. 2012, 64, 520–539. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, T.; Korkaya, H.; Liu, S.; Lee, H.F.; Newman, B.; Yu, Y.; Clouthier, S.G.; Schwartz, S.J.; Wicha, M.S.; et al. Sulforaphane, a Dietary Component of Broccoli/Broccoli Sprouts, Inhibits Breast Cancer Stem Cells. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2010, 16, 2580–2590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kakarala, M.; Brenner, D.E.; Korkaya, H.; Cheng, C.; Tazi, K.; Ginestier, C.; Liu, S.; Dontu, G.; Wicha, M.S. Targeting Breast Stem Cells with the Cancer Preventive Compounds Curcumin and Piperine. Breast Cancer Res. Treat. 2010, 122, 777–785. [Google Scholar] [CrossRef]
- Min, S.J.; Lim, J.Y.; Kim, H.R.; Kim, S.-J.; Kim, Y. Sasa quelpaertensis Leaf Extract Inhibits Colon Cancer by Regulating Cancer Cell Stemness in Vitro and in Vivo. Int. J. Mol. Sci. 2015, 16, 9976–9997. [Google Scholar] [CrossRef] [Green Version]
- Charoensinphon, N. Inhibition of Lung Carcinogenesis by Polymethoxyflavones. Ph.D. Dissertations, University of Massachusetts Amherst, Amherst, MA, USA, 2013. [Google Scholar]
- Moselhy, J.; Srinivasan, S.; Ankem, M.K.; Damodaran, C. Natural Products That Target Cancer Stem Cells. Anticancer Res. 2015, 35, 5773–5788. [Google Scholar]
- Sak, K. Chemotherapy and dietary phytochemical agents. Chemother. Res. Pract. 2012, 2012, 282570. [Google Scholar] [CrossRef]
- Wang, T.; Li, Q.; Bi, K. Bioactive flavonoids in medicinal plants: Structure, activity and biological fate. Asian J. Pharm. Sci. 2018, 13, 12–23. [Google Scholar] [CrossRef]
- Fantini, M.; Benvenuto, M.; Masuelli, L.; Frajese, G.V.; Tresoldi, I.; Modesti, A.; Bei, R. In Vitro and in Vivo Antitumoral Effects of Combinations of Polyphenols, or Polyphenols and Anticancer Drugs: Perspectives on Cancer Treatment. Int. J. Mol. Sci. 2015, 16, 9236–9282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nautiyal, J.; Banerjee, S.; Kanwar, S.S.; Yu, Y.; Patel, B.B.; Sarkar, F.H.; Majumdar, A.P. Curcumin Enhances Dasatinib Induced Inhibition of Growth and Transformation of Colon Cancer Cells. Int. J. Cancer J. Int. Cancer 2011, 128, 951–961. [Google Scholar] [CrossRef] [PubMed]
- Shakibaei, M.; Mobasheri, A.; Lueders, C.; Busch, F.; Shayan, P.; Goel, A. Curcumin Enhances the Effect of Chemotherapy against Colorectal Cancer Cells by Inhibition of NF-κB and Src Protein Kinase Signaling Pathways. PLoS ONE 2013, 8, e57218. [Google Scholar] [CrossRef] [PubMed]
- Aires, V.; Limagne, E.; Cotte, A.K.; Latruffe, N.; Ghiringhelli, F.; Delmas, D. Resveratrol metabolites inhibit human metastatic colon cancer cells progression and synergize with chemotherapeutic drugs to induce cell death. Mol. Nutr. Food Res. 2013, 57, 1170–1181. [Google Scholar] [CrossRef] [PubMed]
- Suganuma, M.; Saha, A.; Fujiki, H. New cancer treatment strategy using combination of green tea catechins and anticancer drugs. Cancer Sci. 2011, 102, 317–323. [Google Scholar] [CrossRef] [PubMed]

| Concentration | 5-FU (mg/mL) | 0.15 | 0.30 | 0.60 | 1.20 |
| OPE (mg/mL) | 0.30 | 0.60 | 1.20 | 2.40 | |
| Nobiletin (μM) | 14.70 | 29.39 | 58.78 | 117.56 | |
| Sinensetin (μM) | 13.95 | 27.90 | 55.79 | 111.58 | |
| Tangeretin (μM) | 3.12 | 6.23 | 12.46 | 24.92 | |
| Scutellarein tetramethylether (μM) | 9.44 | 18.88 | 37.76 | 75.52 | |
| 5-FU OPE | CI | 0.31 | 0.52 | 0.52 | 0.46 |
| DRI for 5-FU | 7.54 | 13.44 | 46.63 | 1080.81 | |
| 5-FU N–S–T–Sc | CI | 0.41 | 0.29 | 0.67 | 0.56 |
| DRI for 5-FU | 6.61 | 21.06 | 7.97 | 25.60 | |
| 5-FU Nobiletin | CI | 0.26 | 0.27 | 0.64 | 0.75 |
| DRI for 5-FU | 4.56 | 4.01 | 1.72 | 1.41 | |
| 5-FU Sinensetin | CI | 0.08 | 0.13 | 0.38 | 0.46 |
| DRI for 5-FU | 12.95 | 7.98 | 2.88 | 2.29 | |
| 5-FU Tangeretin | CI | 0.17 | 0.22 | 0.43 | 2.72 |
| DRI for 5-FU | 10.04 | 9.44 | 5.07 | 0.48 | |
| 5-FU Scutellarein tetramethylether | CI | 0.06 | 0.03 | 0.04 | 0.15 |
| DRI for 5-FU | 17.40 | 35.35 | 22.04 | 6.70 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pereira, C.V.; Duarte, M.; Silva, P.; Bento da Silva, A.; Duarte, C.M.M.; Cifuentes, A.; García-Cañas, V.; Bronze, M.R.; Albuquerque, C.; Serra, A.T. Polymethoxylated Flavones Target Cancer Stemness and Improve the Antiproliferative Effect of 5-Fluorouracil in a 3D Cell Model of Colorectal Cancer. Nutrients 2019, 11, 326. https://doi.org/10.3390/nu11020326
Pereira CV, Duarte M, Silva P, Bento da Silva A, Duarte CMM, Cifuentes A, García-Cañas V, Bronze MR, Albuquerque C, Serra AT. Polymethoxylated Flavones Target Cancer Stemness and Improve the Antiproliferative Effect of 5-Fluorouracil in a 3D Cell Model of Colorectal Cancer. Nutrients. 2019; 11(2):326. https://doi.org/10.3390/nu11020326
Chicago/Turabian StylePereira, Carolina V., Marlene Duarte, Patrícia Silva, Andreia Bento da Silva, Catarina M. M. Duarte, Alejandro Cifuentes, Virginia García-Cañas, Maria R. Bronze, Cristina Albuquerque, and Ana Teresa Serra. 2019. "Polymethoxylated Flavones Target Cancer Stemness and Improve the Antiproliferative Effect of 5-Fluorouracil in a 3D Cell Model of Colorectal Cancer" Nutrients 11, no. 2: 326. https://doi.org/10.3390/nu11020326
APA StylePereira, C. V., Duarte, M., Silva, P., Bento da Silva, A., Duarte, C. M. M., Cifuentes, A., García-Cañas, V., Bronze, M. R., Albuquerque, C., & Serra, A. T. (2019). Polymethoxylated Flavones Target Cancer Stemness and Improve the Antiproliferative Effect of 5-Fluorouracil in a 3D Cell Model of Colorectal Cancer. Nutrients, 11(2), 326. https://doi.org/10.3390/nu11020326