Sex-Specific Changes in Gut Microbiome Composition following Blueberry Consumption in C57BL/6J Mice
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Design
2.2. Microbial Community Profiling Using 16S rRNA Amplicon Sequencing
2.3. Bioinformatics Analysis
2.4. Statistical Analysis
3. Results
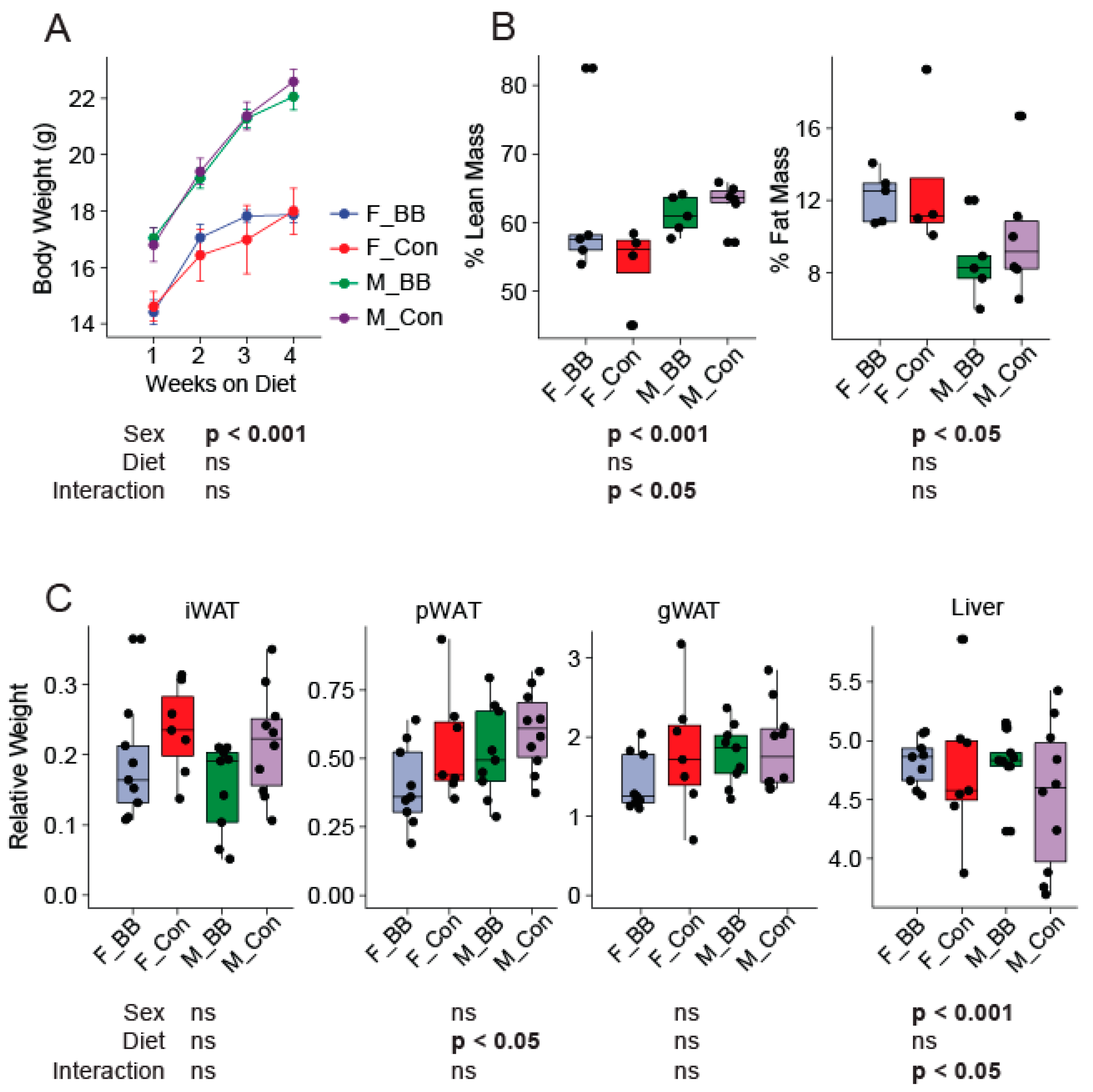
3.1. Body Weight, Body Composition, and Tissue Weights
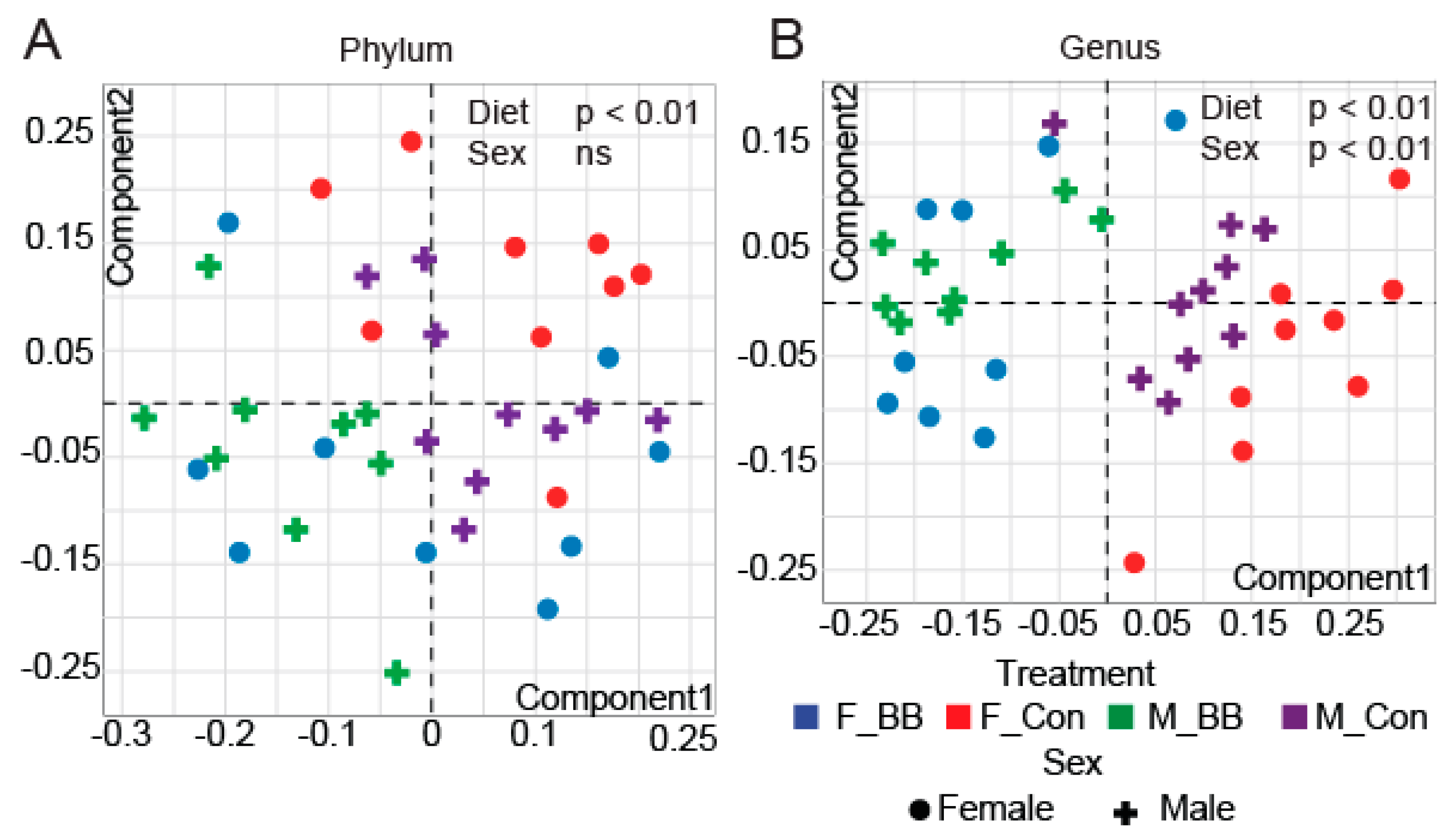
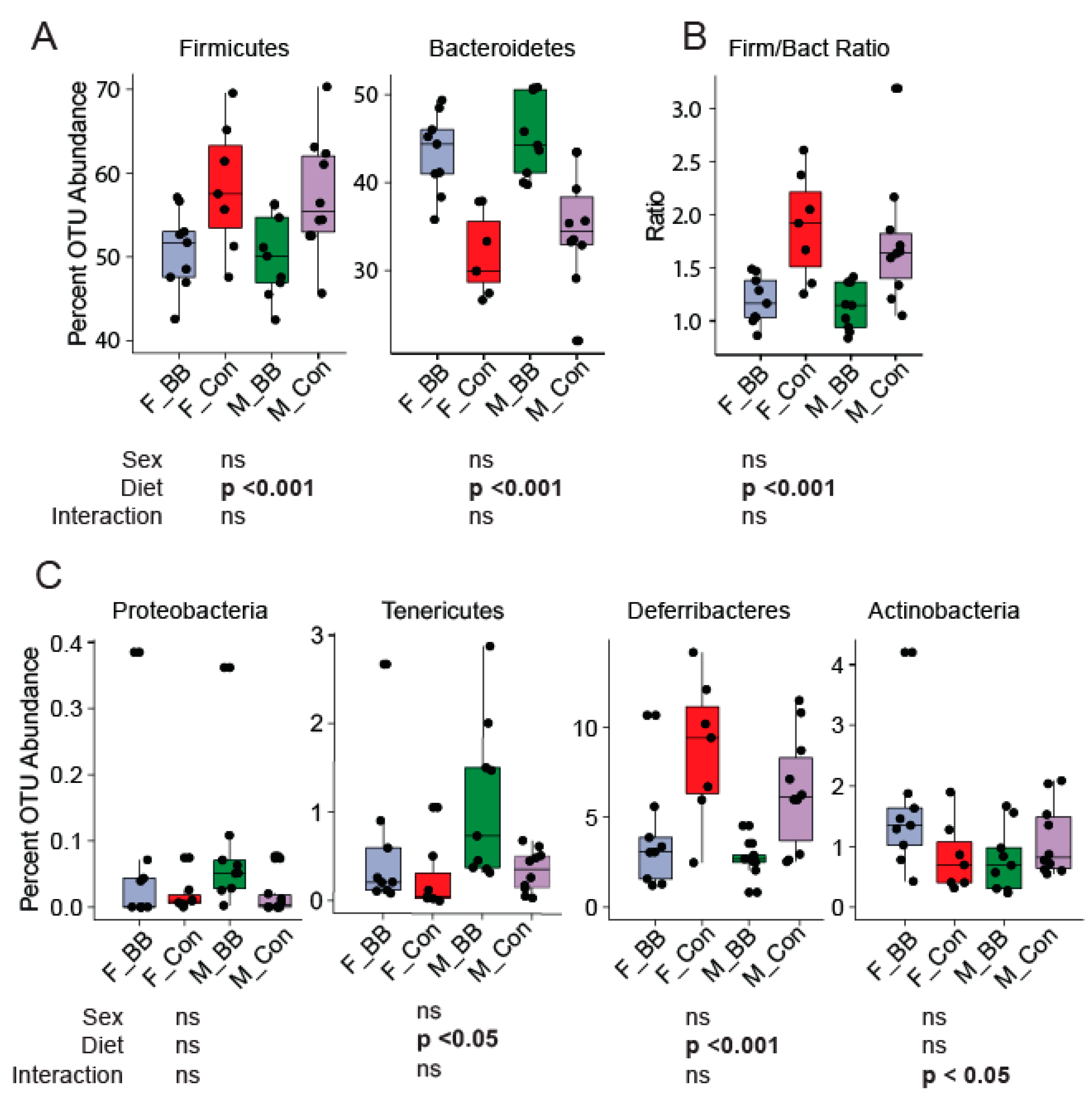
3.2. Gut Microbial Taxonomic Analysis
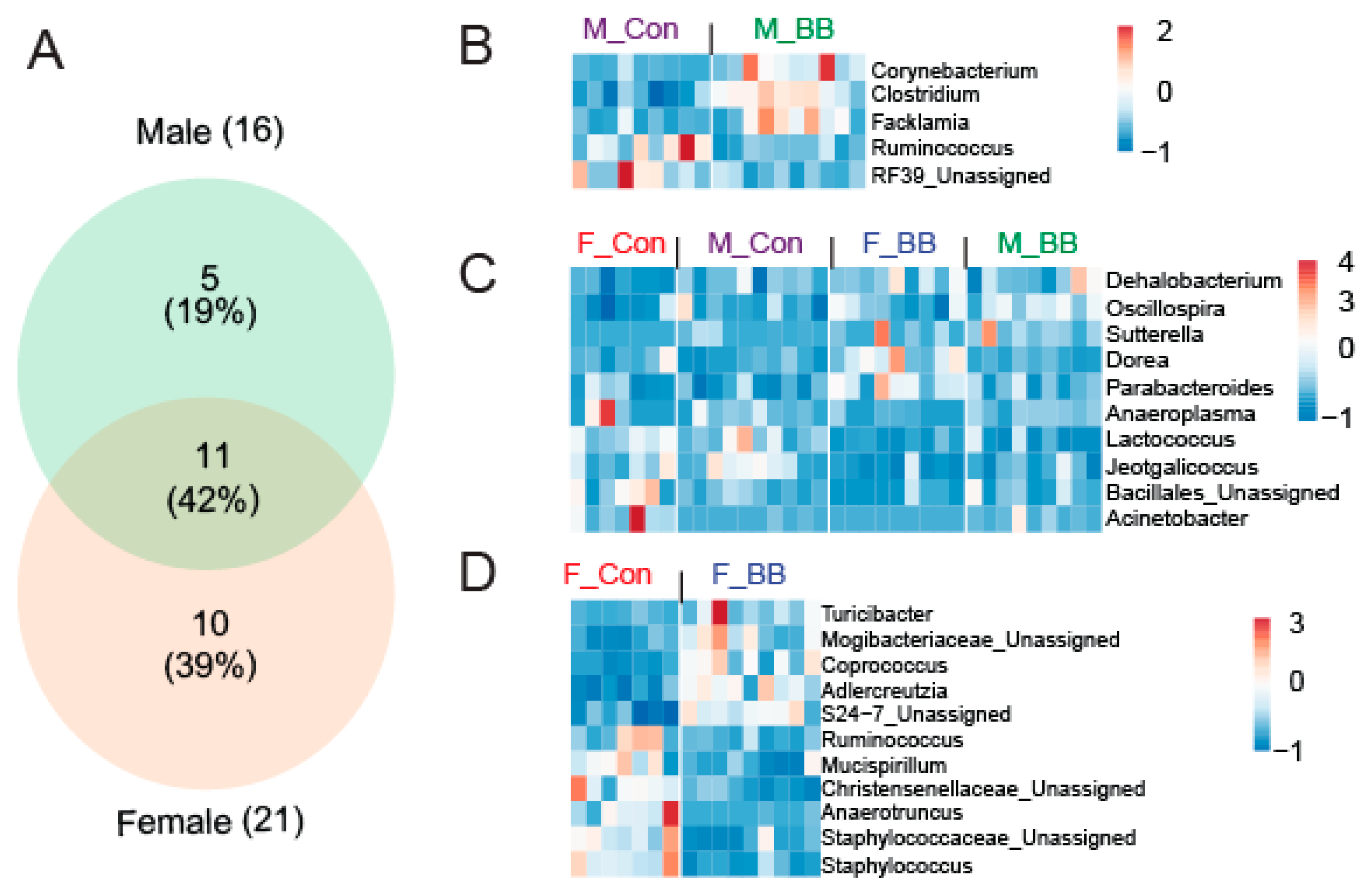
3.3. Blueberry Consumption-Associated Taxonomical Differences
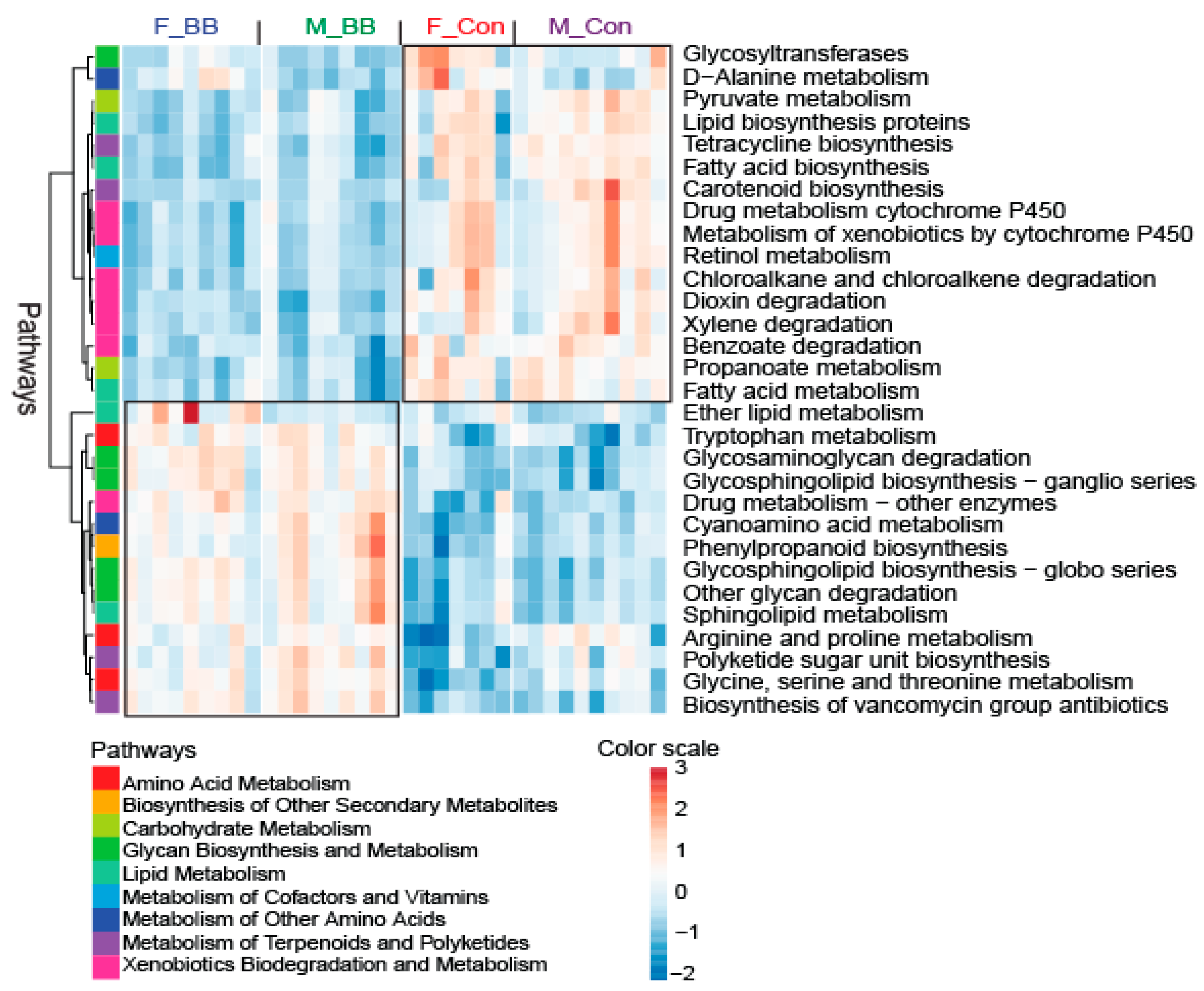
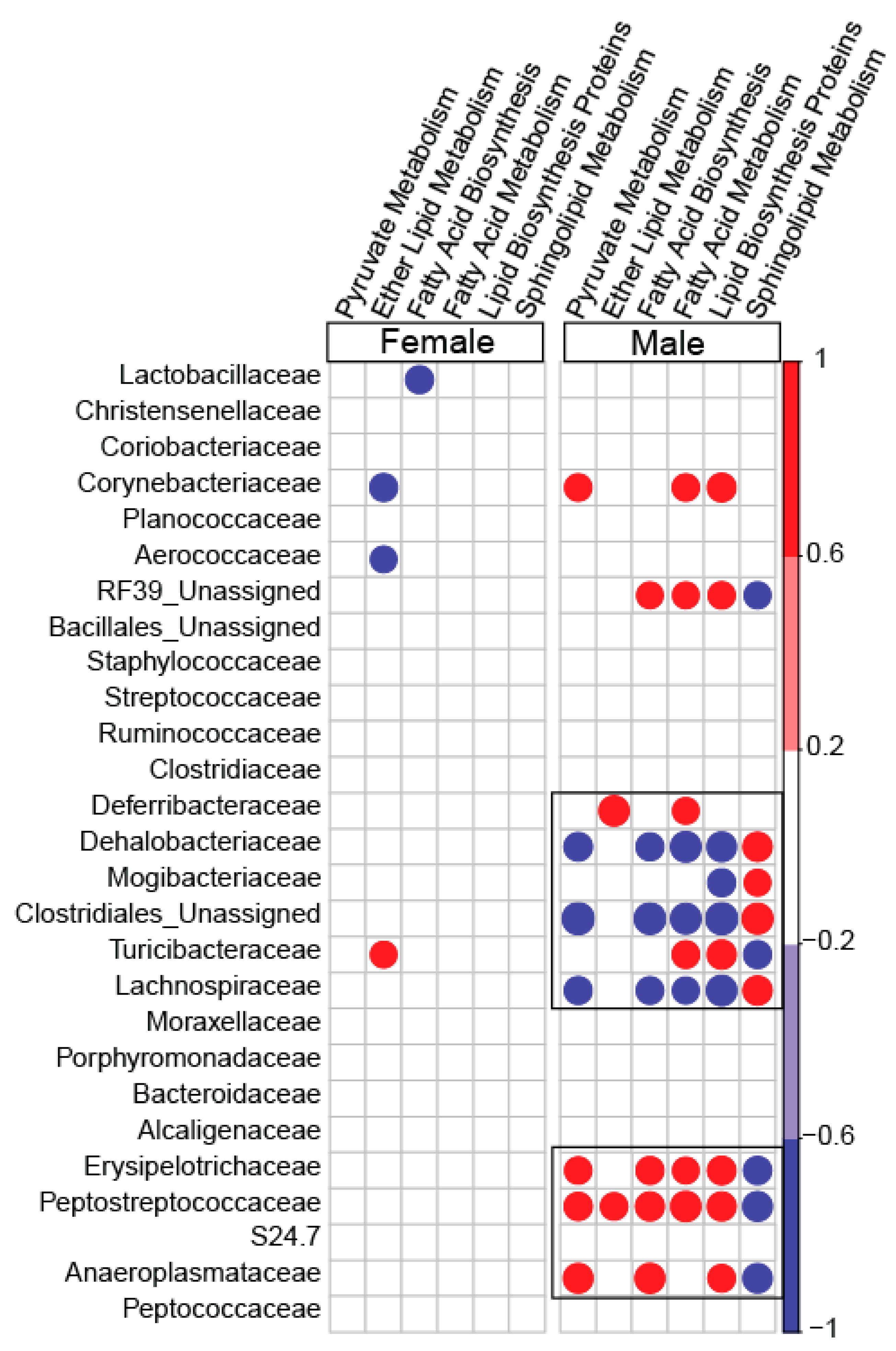
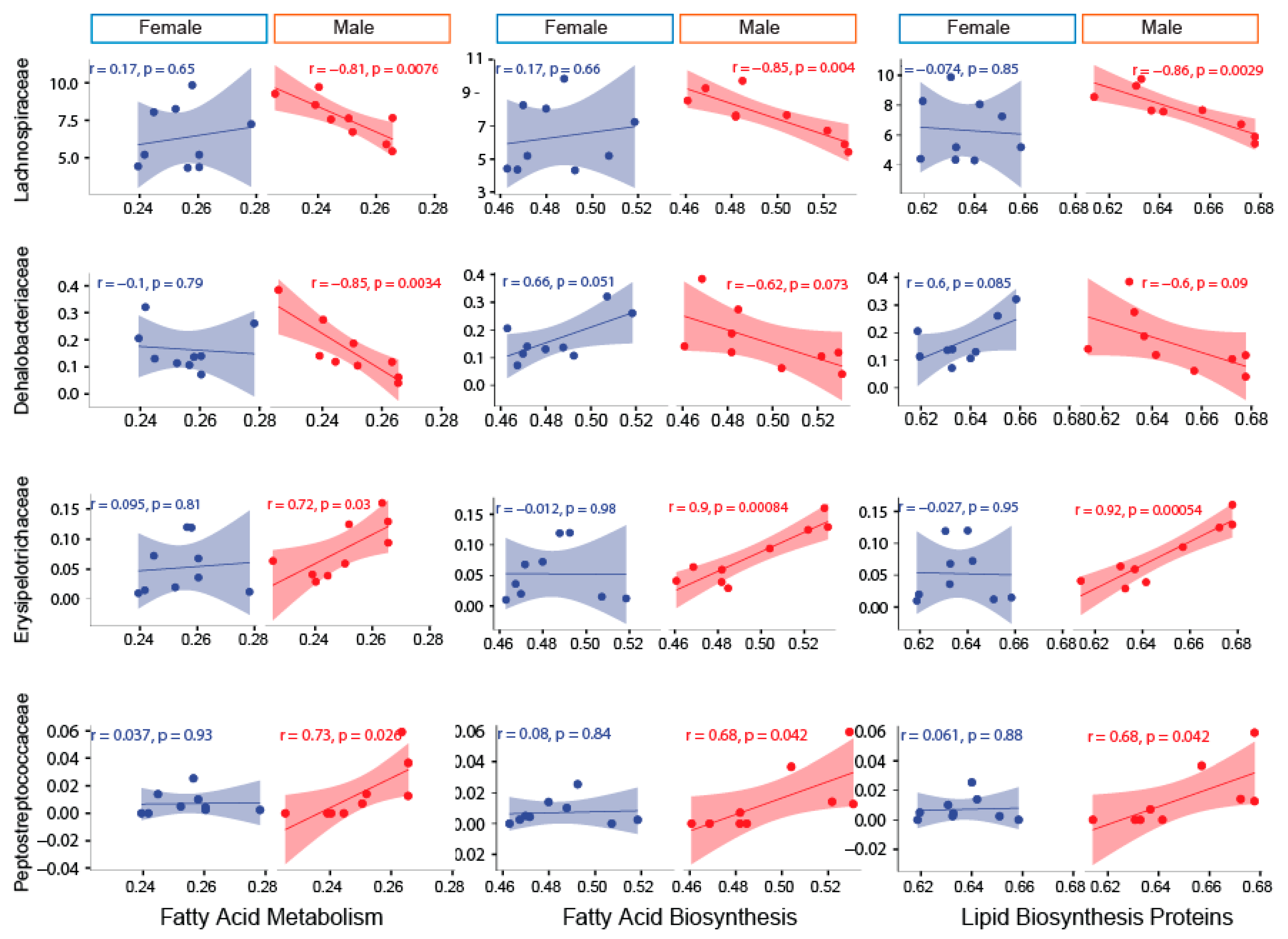
3.4. PICRUSt-Predicted Metabolic Pathways
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- National Nutrient Database for Standard Reference Release 28; ARS-2016; United States Department of Agriculture: Washington, DC, USA, 2016.
- Wang, S.Y.; Lin, H.S. Antioxidant activity in fruits and leaves of blackberry, raspberry, and strawberry varies with cultivar and developmental stage. J. Agric. Food Chem. 2000, 48, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Beecher, G.R.; Holden, J.M.; Haytowitz, D.B.; Gebhardt, S.E.; Prior, R.L. Concentrations of anthocyanins in common foods in the United States and estimation of normal consumption. J. Agric. Food Chem. 2006, 54, 4069–4075. [Google Scholar] [CrossRef] [PubMed]
- Scalbert, A.; Manach, C.; Morand, C.; Remesy, C.; Jimenez, L. Dietary polyphenols and the prevention of diseases. Crit. Rev. Food Sci. Nutr. 2005, 45, 287–306. [Google Scholar] [CrossRef] [PubMed]
- Khurana, S.; Venkataraman, K.; Hollingsworth, A.; Piche, M.; Tai, T.C. Polyphenols: Benefits to the cardiovascular system in health and in aging. Nutrients 2013, 5, 3779–3827. [Google Scholar] [CrossRef] [PubMed]
- Kalt, W.; Forney, C.F.; Martin, A.; Prior, R.L. Antioxidant capacity, vitamin C, phenolics, and anthocyanins after fresh storage of small fruits. J. Agric. Food Chem. 1999, 47, 4638–4644. [Google Scholar] [CrossRef] [PubMed]
- Seeram, N.P.; Adams, L.S.; Zhang, Y.; Lee, R.; Sand, D.; Scheuller, H.S.; Heber, D. Blackberry, black raspberry, blueberry, cranberry, red raspberry, and strawberry extracts inhibit growth and stimulate apoptosis of human cancer cells in vitro. J. Agric. Food Chem. 2006, 54, 9329–9339. [Google Scholar] [CrossRef] [PubMed]
- Neto, C.C. Cranberry and blueberry: Evidence for protective effects against cancer and vascular diseases. Mol. Nutr. Food Res. 2007, 51, 652–664. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Li, X.Q.; Weber, C.; Lee, C.Y.; Brown, J.; Liu, R.H. Antioxidant and antiproliferative activities of raspberries. J. Agric. Food Chem. 2002, 50, 2926–2930. [Google Scholar] [CrossRef]
- Chen, J.R.; Lazarenko, O.P.; Wu, X.; Kang, J.; Blackburn, M.L.; Shankar, K.; Badger, T.M.; Ronis, M.J. Dietary-induced serum phenolic acids promote bone growth via p38 MAPK/beta-catenin canonical Wnt signaling. J. Bone Miner. Res. 2010, 25, 2399–2411. [Google Scholar] [CrossRef]
- Chen, J.R.; Lazarenko, O.P.; Zhang, J.; Blackburn, M.L.; Ronis, M.J.; Badger, T.M. Diet-derived phenolic acids regulate osteoblast and adipocyte lineage commitment and differentiation in young mice. J. Bone Miner. Res. 2014, 29, 1043–1053. [Google Scholar] [CrossRef]
- McGhie, T.K.; Walton, M.C. The bioavailability and absorption of anthocyanins: Towards a better understanding. Mol. Nutr. Food Res. 2007, 51, 702–713. [Google Scholar] [CrossRef] [PubMed]
- Bolca, S.; Van de Wiele, T.; Possemiers, S. Gut metabotypes govern health effects of dietary polyphenols. Curr. Opin. Biotechnol. 2013, 24, 220–225. [Google Scholar] [CrossRef] [PubMed]
- Rebello, C.J.; Burton, J.; Heiman, M.; Greenway, F.L. Gastrointestinal microbiome modulator improves glucose tolerance in overweight and obese subjects: A randomized controlled pilot trial. J. Diabetes Complicat. 2015, 29, 1272–1276. [Google Scholar] [CrossRef] [PubMed]
- Burton, J.H.; Johnson, M.; Johnson, J.; Hsia, D.S.; Greenway, F.L.; Heiman, M.L. Addition of a Gastrointestinal Microbiome Modulator to Metformin Improves Metformin Tolerance and Fasting Glucose Levels. J. Diabetes Sci. Technol. 2015, 9, 808–814. [Google Scholar] [CrossRef] [PubMed]
- Lacombe, A.; Li, R.W.; Klimis-Zacas, D.; Kristo, A.S.; Tadepalli, S.; Krauss, E.; Young, R.; Wu, V.C. Lowbush wild blueberries have the potential to modify gut microbiota and xenobiotic metabolism in the rat colon. PLoS ONE 2013, 8, e67497. [Google Scholar] [CrossRef] [PubMed]
- Muegge, B.D.; Kuczynski, J.; Knights, D.; Clemente, J.C.; Gonzalez, A.; Fontana, L.; Henrissat, B.; Knight, R.; Gordon, J.I. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 2011, 332, 970–974. [Google Scholar] [CrossRef] [PubMed]
- Markle, J.G.; Frank, D.N.; Mortin-Toth, S.; Robertson, C.E.; Feazel, L.M.; Rolle-Kampczyk, U.; von Bergen, M.; McCoy, K.D.; Macpherson, A.J.; Danska, J.S. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 2013, 339, 1084–1088. [Google Scholar] [CrossRef] [PubMed]
- Freire, A.C.; Basit, A.W.; Choudhary, R.; Piong, C.W.; Merchant, H.A. Does sex matter? The influence of gender on gastrointestinal physiology and drug delivery. Int. J. Pharm. 2011, 415, 15–28. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Baars, A.; Oosting, A.; Lohuis, M.; Koehorst, M.; El Aidy, S.; Hugenholtz, F.; Smidt, H.; Mischke, M.; Boekschoten, M.V.; Verkade, H.J.; et al. Sex differences in lipid metabolism are affected by presence of the gut microbiota. Sci. Rep. 2018, 8, 13426. [Google Scholar] [CrossRef] [PubMed]
- Bolnick, D.I.; Snowberg, L.K.; Hirsch, P.E.; Lauber, C.L.; Org, E.; Parks, B.; Lusis, A.J.; Knight, R.; Caporaso, J.G.; Svanback, R. Individual diet has sex-dependent effects on vertebrate gut microbiota. Nat. Commun. 2014, 5, 4500. [Google Scholar] [CrossRef] [PubMed]
- Davis, D.J.; Hecht, P.M.; Jasarevic, E.; Beversdorf, D.Q.; Will, M.J.; Fritsche, K.; Gillespie, C.H. Sex-specific effects of docosahexaenoic acid (DHA) on the microbiome and behavior of socially-isolated mice. Brain Behav. Immun. 2017, 59, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, J.I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57–59. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Wankhade, U.D.; Zhong, Y.; Kang, P.; Alfaro, M.; Chintapalli, S.V.; Thakali, K.M.; Shankar, K. Enhanced offspring predisposition to steatohepatitis with maternal high-fat diet is associated with epigenetic and microbiome alterations. PLoS ONE 2017, 12, e0175675. [Google Scholar] [CrossRef]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Rideout, J.R.; He, Y.; Navas-Molina, J.A.; Walters, W.A.; Ursell, L.K.; Gibbons, S.M.; Chase, J.; McDonald, D.; Gonzalez, A.; Robbins-Pianka, A.; et al. Subsampled open-reference clustering creates consistent, comprehensive OTU definitions and scales to billions of sequences. PeerJ 2014, 2, e545. [Google Scholar] [CrossRef] [PubMed]
- Knights, D.; Kuczynski, J.; Charlson, E.S.; Zaneveld, J.; Mozer, M.C.; Collman, R.G.; Bushman, F.D.; Knight, R.; Kelley, S.T. Bayesian community-wide culture-independent microbial source tracking. Nat. Methods 2011, 8, 761–763. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Piccolo, B.D.; Wankhade, U.D.; Chintapalli, S.V.; Bhattacharyya, S.; Luo, C.; Shankar, K. Dynamic Assessment of Microbial Ecology (DAME): A web app for interactive analysis and visualization of microbial sequencing data. Bioinformatics 2017, 34, 1050–1052. [Google Scholar] [CrossRef]
- Van Nas, A.; Guhathakurta, D.; Wang, S.S.; Yehya, N.; Horvath, S.; Zhang, B.; Ingram-Drake, L.; Chaudhuri, G.; Schadt, E.E.; Drake, T.A.; et al. Elucidating the role of gonadal hormones in sexually dimorphic gene coexpression networks. Endocrinology 2009, 150, 1235–1249. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.; Saunier, K.; Hanisch, C.; Norin, E.; Alm, L.; Midtvedt, T.; Cresci, A.; Silvi, S.; Orpianesi, C.; Verdenelli, M.C.; et al. Differences in fecal microbiota in different European study populations in relation to age, gender, and country: A cross-sectional study. Appl. Environ. Microbiol. 2006, 72, 1027–1033. [Google Scholar] [CrossRef] [PubMed]
- Shastri, P.; McCarville, J.; Kalmokoff, M.; Brooks, S.P.; Green-Johnson, J.M. Sex differences in gut fermentation and immune parameters in rats fed an oligofructose-supplemented diet. Biol. Sex Differ. 2015, 6, 13. [Google Scholar] [CrossRef] [PubMed]
- Gomez, A.; Luckey, D.; Taneja, V. The gut microbiome in autoimmunity: Sex matters. Clin. Immunol. 2015, 159, 154–162. [Google Scholar] [CrossRef]
- Yurkovetskiy, L.; Burrows, M.; Khan, A.A.; Graham, L.; Volchkov, P.; Becker, L.; Antonopoulos, D.; Umesaki, Y.; Chervonsky, A.V. Gender bias in autoimmunity is influenced by microbiota. Immunity 2013, 39, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Org, E.; Mehrabian, M.; Parks, B.W.; Shipkova, P.; Liu, X.; Drake, T.A.; Lusis, A.J. Sex differences and hormonal effects on gut microbiota composition in mice. Gut Microbes 2016, 7, 313–322. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; McClusky, R.; Chen, J.; Beaven, S.W.; Tontonoz, P.; Arnold, A.P.; Reue, K. The number of x chromosomes causes sex differences in adiposity in mice. PLoS Genet. 2012, 8, e1002709. [Google Scholar] [CrossRef] [PubMed]
- Laaksonen, D.E.; Niskanen, L.; Punnonen, K.; Nyyssonen, K.; Tuomainen, T.P.; Salonen, R.; Rauramaa, R.; Salonen, J.T. Sex hormones, inflammation and the metabolic syndrome: A population-based study. Eur. J. Endocrinol. 2003, 149, 601–608. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.C. Sexual dimorphism of body composition. Best Pract. Res. Clin. Endocrinol. Metab. 2007, 21, 415–430. [Google Scholar] [CrossRef]
- Palmer, B.F.; Clegg, D.J. The sexual dimorphism of obesity. Mol. Cell. Endocrinol. 2015, 402, 113–119. [Google Scholar] [CrossRef]
- Ter Horst, K.W.; Gilijamse, P.W.; de Weijer, B.A.; Kilicarslan, M.; Ackermans, M.T.; Nederveen, A.J.; Nieuwdorp, M.; Romijn, J.A.; Serlie, M.J. Sexual Dimorphism in Hepatic, Adipose Tissue, and Peripheral Tissue Insulin Sensitivity in Obese Humans. Front. Endocrinol. 2015, 6, 182. [Google Scholar] [CrossRef]
- Dulebohn, R.V.; Yi, W.; Srivastava, A.; Akoh, C.C.; Krewer, G.; Fischer, J.G. Effects of blueberry (Vaccinium ashei) on DNA damage, lipid peroxidation, and phase II enzyme activities in rats. J. Agric. Food Chem. 2008, 56, 11700–11706. [Google Scholar] [CrossRef]
- Riso, P.; Klimis-Zacas, D.; Del Bo, C.; Martini, D.; Campolo, J.; Vendrame, S.; Moller, P.; Loft, S.; De Maria, R.; Porrini, M. Effect of a wild blueberry (Vaccinium angustifolium) drink intervention on markers of oxidative stress, inflammation and endothelial function in humans with cardiovascular risk factors. Eur. J. Nutr. 2013, 52, 949–961. [Google Scholar] [CrossRef]
- DeFuria, J.; Bennett, G.; Strissel, K.J.; Perfield, J.W., II; Milbury, P.E.; Greenberg, A.S.; Obin, M.S. Dietary blueberry attenuates whole-body insulin resistance in high fat-fed mice by reducing adipocyte death and its inflammatory sequelae. J. Nutr. 2009, 139, 1510–1516. [Google Scholar] [CrossRef]
- Petersen, C.; Bharat, D.; Cutler, B.R.; Gholami, S.; Denetso, C.; Mueller, J.E.; Cho, J.M.; Kim, J.S.; Symons, J.D.; Anandh Babu, P.V. Circulating metabolites of strawberry mediate reductions in vascular inflammation and endothelial dysfunction in db/db mice. Int. J. Cardiol. 2018, 263, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Bharat, D.; Cavalcanti, R.R.M.; Petersen, C.; Begaye, N.; Cutler, B.R.; Costa, M.M.A.; Ramos, R.; Ferreira, M.R.; Li, Y.; Bharath, L.P.; et al. Blueberry Metabolites Attenuate Lipotoxicity-Induced Endothelial Dysfunction. Mol. Nutr. Food Res. 2018, 62, 1700601. [Google Scholar] [CrossRef] [PubMed]
- Fairlie-Jones, L.; Davison, K.; Fromentin, E.; Hill, A.M. The Effect of Anthocyanin-Rich Foods or Extracts on Vascular Function in Adults: A Systematic Review and Meta-Analysis of Randomised Controlled Trials. Nutrients 2017, 9, 908. [Google Scholar] [CrossRef] [PubMed]
- Faria, A.; Fernandes, I.; Norberto, S.; Mateus, N.; Calhau, C. Interplay between anthocyanins and gut microbiota. J. Agric. Food Chem. 2014, 62, 6898–6902. [Google Scholar] [CrossRef] [PubMed]
- Roopchand, D.E.; Carmody, R.N.; Kuhn, P.; Moskal, K.; Rojas-Silva, P.; Turnbaugh, P.J.; Raskin, I. Dietary Polyphenols Promote Growth of the Gut Bacterium Akkermansia muciniphila and Attenuate High-Fat Diet-Induced Metabolic Syndrome. Diabetes 2015, 64, 2847–2858. [Google Scholar] [CrossRef] [PubMed]
- Henning, S.M.; Yang, J.; Shao, P.; Lee, R.P.; Huang, J.; Ly, A.; Hsu, M.; Lu, Q.Y.; Thames, G.; Heber, D.; et al. Health benefit of vegetable/fruit juice-based diet: Role of microbiome. Sci. Rep. 2017, 7, 2167. [Google Scholar] [CrossRef]
- Lee, S.; Keirsey, K.I.; Kirkland, R.; Grunewald, Z.I.; Fischer, J.G.; de La Serre, C.B. Blueberry Supplementation Influences the Gut Microbiota, Inflammation, and Insulin Resistance in High-Fat-Diet-Fed Rats. J. Nutr. 2018, 148, 209–219. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Liu, W.; Zhao, S.; Wang, J.; Shi, J.; Sun, Y.; Wang, W.; Ning, G.; Hong, J.; Liu, R. Grape seed proanthocyanidin extract ameliorates inflammation and adiposity by modulating gut microbiota in high-fat diet mice. Mol. Nutr. Food Res. 2017, 61, 1601082. [Google Scholar] [CrossRef]
- Wu, T.; Qi, X.; Liu, Y.; Guo, J.; Zhu, R.; Chen, W.; Zheng, X.; Yu, T. Dietary supplementation with purified mulberry (Morus australis Poir) anthocyanins suppresses body weight gain in high-fat diet fed C57BL/6 mice. Food Chem. 2013, 141, 482–487. [Google Scholar] [CrossRef]
- Anhe, F.F.; Nachbar, R.T.; Varin, T.V.; Vilela, V.; Dudonne, S.; Pilon, G.; Fournier, M.; Lecours, M.A.; Desjardins, Y.; Roy, D.; et al. A polyphenol-rich cranberry extract reverses insulin resistance and hepatic steatosis independently of body weight loss. Mol. Metab. 2017, 6, 1563–1573. [Google Scholar] [CrossRef]
- Anhe, F.F.; Roy, D.; Pilon, G.; Dudonne, S.; Matamoros, S.; Varin, T.V.; Garofalo, C.; Moine, Q.; Desjardins, Y.; Levy, E.; et al. A polyphenol-rich cranberry extract protects from diet-induced obesity, insulin resistance and intestinal inflammation in association with increased Akkermansia spp. population in the gut microbiota of mice. Gut 2015, 64, 872–883. [Google Scholar] [CrossRef] [PubMed]
- Takikawa, M.; Inoue, S.; Horio, F.; Tsuda, T. Dietary anthocyanin-rich bilberry extract ameliorates hyperglycemia and insulin sensitivity via activation of AMP-activated protein kinase in diabetic mice. J. Nutr. 2010, 140, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Laparra, J.M.; Sanz, Y. Interactions of gut microbiota with functional food components and nutraceuticals. Pharmacol. Res. 2010, 61, 219–225. [Google Scholar] [CrossRef]
- Milbury, P.E.; Kalt, W. Xenobiotic metabolism and berry flavonoid transport across the blood-brain barrier. J. Agric. Food Chem. 2010, 58, 3950–3956. [Google Scholar] [CrossRef] [PubMed]
| Ingredients | 7% Corn Oil Diet | 5% Blueberry Diet |
|---|---|---|
| TD.95092 | TD.10679 | |
| g/kg | g/kg | |
| Casein | 200.0 | 198.2 |
| L-Cysteine | 3 | 3 |
| Corn Starch | 397.5 | 351.9 |
| Maltodextrin | 132 | 132 |
| Sucrose | 100 | 100 |
| Corn Oil | 70.0 | 69.5 |
| Cellulose | 50.0 | 47.9 |
| Min Mix (AIN-93G-MX(94046)) | 35 | 35 |
| Vitamin Mix (AIN-93-VX(94047)) | 10 | 10 |
| Choline Bitartrate | 2.5 | 2.5 |
| TBHQ, Anti-oxident | 0.014 | 0.014 |
| Blueberry Powder | - | 50 |
| Both Diets | ||
| Macronutrient Info | % by weight | % kcal from |
| Protein | 17.7 | 18.8 |
| Carbohydrate | 60.1 | 63.9 |
| Fat | 7.2 | 17.2 |
| Kcal/g | 3.8 | 3.8 |
| Phylum | |||||||
| Index | F_BB | F_Con | M_BB | M_Con | Sex | Diet | Diet:Sex |
| Chao1 | 5.444 ± 0.176 | 5.714 ± 0.184 | 6 ± 0 | 5.5 ± 0.167 | 0.225 | 0.381 | 0.017 |
| Fisher | 0.48 ± 0.017 | 0.5 ± 0.018 | 0.534 ± 0.001 | 0.485 ± 0.016 | 0.163 | 0.299 | 0.025 |
| InvSimpson | 2.22 ± 0.041 | 2.214 ± 0.099 | 2.176 ± 0.03 | 2.181 ± 0.067 | 0.526 | 0.955 | 0.924 |
| Observed | 5.444 ± 0.176 | 5.714 ± 0.184 | 6 ± 0 | 5.5 ± 0.167 | 0.225 | 0.381 | 0.017 |
| Shannon | 0.908 ± 0.025 | 0.929 ± 0.034 | 0.887 ± 0.022 | 0.915 ± 0.027 | 0.512 | 0.389 | 0.880 |
| Simpson | 0.548 ± 0.008 | 0.543 ± 0.022 | 0.54 ± 0.006 | 0.538 ± 0.015 | 0.599 | 0.737 | 0.890 |
| Genus | |||||||
| Index | F_BB | F_Con | M_BB | M_Con | Sex | Diet | Diet:Sex |
| Chao1 | 31.833 ± 0.677 | 34.286 ± 0.778 | 33.667 ± 1.003 | 34.833 ± 0.564 | 0.125 | 0.022 | 0.412 |
| Fisher | 3.368 ± 0.071 | 3.614 ± 0.091 | 3.568 ± 0.101 | 3.719 ± 0.065 | 0.071 | 0.017 | 0.570 |
| InvSimpson | 5.186 ± 0.233 | 5.49 ± 0.32 | 4.883 ± 0.21 | 6.277 ± 0.258 | 0.398 | 0.001 | 0.041 |
| Observed | 31.667 ± 0.601 | 34.143 ± 0.738 | 33.333 ± 0.882 | 34.6 ± 0.521 | 0.128 | 0.009 | 0.392 |
| Shannon | 2.091 ± 0.039 | 2.067 ± 0.061 | 2.124 ± 0.039 | 2.227 ± 0.028 | 0.029 | 0.191 | 0.128 |
| Simpson | 0.804 ± 0.009 | 0.814 ± 0.011 | 0.792 ± 0.009 | 0.838 ± 0.006 | 0.534 | 0.001 | 0.046 |
| Family_Genus | F_Con | F_BB | M_Con | M_BB | Sex | Diet | Diet × Sex |
|---|---|---|---|---|---|---|---|
| Clostridiales_Unassigned | 12.93 ± 3.25 | 15.8 ± 1.04 | 12.42 ± 1.51 | 16.68 ± 1.85 | 0.954 | 0.065 | 0.720 |
| Staphylococcaceae_Unassigned | 0.04 ± 0.01 | 0.01 ± 0 | 0.02 ± 0.01 | 0 ± 0 | 0.064 | 0.000 | 0.434 |
| Alcaligenaceae_Sutterella | 0 ± 0 | 0.06 ± 0.04 | 0.02 ± 0.01 | 0.08 ± 0.04 | 0.652 | 0.047 | 0.985 |
| Coriobacteriaceae_Adlercreutzia | 0.29 ± 0.07 | 1.19 ± 0.15 | 0.38 ± 0.09 | 0.72 ± 0.15 | 0.050 | 0.000 | 0.033 |
| Erysipelotrichaceae_Unassigned | 0.07 ± 0.02 | 0.05 ± 0.01 | 0.05 ± 0.02 | 0.08 ± 0.02 | 0.740 | 0.615 | 0.131 |
| Clostridiaceae_Unassigned | 5.52 ± 1.59 | 8.35 ± 1.6 | 12.04 ± 1.3 | 6.06 ± 0.65 | 0.123 | 0.144 | 0.002 |
| Staphylococcaceae_Staphylococcus | 0.19 ± 0.04 | 0.04 ± 0.01 | 0.14 ± 0.04 | 0.02 ± 0.01 | 0.479 | 0.000 | 0.613 |
| Lactobacillaceae_Lactobacillus | 23.83 ± 3.73 | 9.93 ± 1.96 | 9.67 ± 2.6 | 4.47 ± 1.38 | 0.001 | 0.001 | 0.086 |
| Ruminococcaceae_Oscillospira | 1.8 ± 0.32 | 2.72 ± 0.39 | 2.39 ± 0.39 | 3.42 ± 0.29 | 0.132 | 0.011 | 0.885 |
| Staphylococcaceae_Jeotgalicoccus | 0.05 ± 0.01 | 0.01 ± 0.01 | 0.05 ± 0.01 | 0.02 ± 0.01 | 0.204 | 0.002 | 0.920 |
| Dehalobacteriaceae_Dehalobacterium | 0.07 ± 0.01 | 0.17 ± 0.03 | 0.12 ± 0.02 | 0.16 ± 0.04 | 0.589 | 0.027 | 0.311 |
| X.Mogibacteriaceae._Unassigned | 0.03 ± 0.01 | 0.11 ± 0.02 | 0.04 ± 0.01 | 0.1 ± 0.01 | 0.654 | 0.000 | 0.566 |
| Ruminococcaceae_Ruminococcus | 3.8 ± 1.24 | 1.12 ± 0.25 | 6.64 ± 1.26 | 2.46 ± 0.62 | 0.018 | 0.001 | 0.432 |
| Lachnospiraceae_Dorea | 0.05 ± 0.02 | 0.11 ± 0.02 | 0.02 ± 0.01 | 0.03 ± 0 | 0.000 | 0.050 | 0.064 |
| Streptococcaceae_Lactococcus | 0.95 ± 0.14 | 0.28 ± 0.04 | 0.86 ± 0.19 | 0.38 ± 0.14 | 0.695 | 0.000 | 0.503 |
| Ruminococcaceae_Unassigned | 2.84 ± 0.73 | 1.83 ± 0.21 | 3.28 ± 0.31 | 2.37 ± 0.36 | 0.162 | 0.024 | 0.901 |
| Porphyromonadaceae_Parabacteroides | 3.44 ± 0.72 | 6.44 ± 0.91 | 2.75 ± 0.45 | 3.74 ± 0.52 | 0.007 | 0.008 | 0.143 |
| S24.7_Unassigned | 27.48 ± 1.08 | 36.57 ± 1.32 | 30.15 ± 1.68 | 39.53 ± 1.23 | 0.167 | 0.000 | 0.920 |
| Planococcaceae_Sporosarcina | 0.01 ± 0 | 0 ± 0 | 0.01 ± 0 | 0.01 ± 0.01 | 0.036 | 0.647 | 0.458 |
| Lachnospiraceae_Coprococcus | 0.21 ± 0.03 | 0.92 ± 0.16 | 0.37 ± 0.08 | 1.44 ± 0.23 | 0.089 | 0.000 | 0.243 |
| RF39_Unassigned | 0.23 ± 0.15 | 0.57 ± 0.28 | 0.33 ± 0.08 | 1.11 ± 0.3 | 0.220 | 0.014 | 0.330 |
| Christensenellaceae_Unassigned | 0.46 ± 0.09 | 0.11 ± 0.03 | 0.28 ± 0.05 | 0.08 ± 0.01 | 0.114 | 0.000 | 0.123 |
| Anaeroplasmataceae_Anaeroplasma | 0.03 ± 0.02 | 0 ± 0 | 0.02 ± 0 | 0.01 ± 0 | 0.941 | 0.143 | 0.166 |
| Lachnospiraceae_.Ruminococcus. | 0.59 ± 0.14 | 0.35 ± 0.08 | 0.26 ± 0.04 | 0.72 ± 0.15 | 0.837 | 0.186 | 0.002 |
| Lachnospiraceae_Unassigned | 3.73 ± 1.24 | 4.94 ± 0.59 | 5.84 ± 0.74 | 5.42 ± 0.25 | 0.102 | 0.668 | 0.274 |
| Turicibacteraceae_Turicibacter | 0.18 ± 0.1 | 2.22 ± 0.91 | 1.51 ± 0.67 | 4.54 ± 0.76 | 0.034 | 0.001 | 0.508 |
| Ruminococcaceae_Anaerotruncus | 0.02 ± 0.01 | 0 ± 0 | 0.02 ± 0.01 | 0 ± 0 | 0.793 | 0.000 | 0.799 |
| Deferribacteraceae_Mucispirillum | 8.72 ± 1.5 | 3.75 ± 0.98 | 6.45 ± 1.02 | 2.71 ± 0.34 | 0.219 | 0.000 | 0.542 |
| Corynebacteriaceae_Corynebacterium | 0.55 ± 0.2 | 0.37 ± 0.24 | 0.73 ± 0.2 | 0.07 ± 0.04 | 0.879 | 0.026 | 0.211 |
| Peptococcaceae_Unassigned | 0.03 ± 0.01 | 0.06 ± 0.02 | 0.08 ± 0.02 | 0.08 ± 0.01 | 0.169 | 0.474 | 0.393 |
| Peptococcaceae_rc4.4 | 0.76 ± 0.26 | 1.46 ± 0.3 | 0.83 ± 0.27 | 1.46 ± 0.39 | 0.941 | 0.042 | 0.908 |
| Bacillales_Unassigned | 0.03 ± 0.01 | 0 ± 0 | 0.01 ± 0 | 0.01 ± 0 | 0.351 | 0.035 | 0.004 |
| Aerococcaceae_Facklamia | 0.05 ± 0.01 | 0.04 ± 0.02 | 0.06 ± 0.01 | 0.01 ± 0.01 | 0.832 | 0.056 | 0.208 |
| Moraxellaceae_Acinetobacter | 0.02 ± 0.01 | 0 ± 0 | 0 ± 0 | 0.01 ± 0 | 0.258 | 0.200 | 0.012 |
| Bacteroidaceae_Bacteroides | 0.96 ± 0.36 | 0.33 ± 0.1 | 1.92 ± 0.52 | 1.95 ± 0.54 | 0.005 | 0.539 | 0.455 |
| Peptostreptococcaceae_Unassigned | 0.01 ± 0 | 0.01 ± 0 | 0.02 ± 0.01 | 0.01 ± 0.01 | 0.059 | 0.394 | 0.433 |
| Clostridiaceae_Clostridium | 0.07 ± 0.02 | 0.08 ± 0.02 | 0.21 ± 0.02 | 0.07 ± 0.01 | 0.002 | 0.000 | 0.000 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wankhade, U.D.; Zhong, Y.; Lazarenko, O.P.; Chintapalli, S.V.; Piccolo, B.D.; Chen, J.-R.; Shankar, K. Sex-Specific Changes in Gut Microbiome Composition following Blueberry Consumption in C57BL/6J Mice. Nutrients 2019, 11, 313. https://doi.org/10.3390/nu11020313
Wankhade UD, Zhong Y, Lazarenko OP, Chintapalli SV, Piccolo BD, Chen J-R, Shankar K. Sex-Specific Changes in Gut Microbiome Composition following Blueberry Consumption in C57BL/6J Mice. Nutrients. 2019; 11(2):313. https://doi.org/10.3390/nu11020313
Chicago/Turabian StyleWankhade, Umesh D., Ying Zhong, Oxana P. Lazarenko, Sree V. Chintapalli, Brian D. Piccolo, Jin-Ran Chen, and Kartik Shankar. 2019. "Sex-Specific Changes in Gut Microbiome Composition following Blueberry Consumption in C57BL/6J Mice" Nutrients 11, no. 2: 313. https://doi.org/10.3390/nu11020313
APA StyleWankhade, U. D., Zhong, Y., Lazarenko, O. P., Chintapalli, S. V., Piccolo, B. D., Chen, J.-R., & Shankar, K. (2019). Sex-Specific Changes in Gut Microbiome Composition following Blueberry Consumption in C57BL/6J Mice. Nutrients, 11(2), 313. https://doi.org/10.3390/nu11020313





