Expression of Granulisyn, Perforin and Granzymes in Human Milk over Lactation and in the Case of Maternal Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Human Milk Collection
2.2. Milk Cell Isolation
2.3. RNA Extraction
2.4. cDNA Generation
2.5. Quantitative Real Time Polymerase Chain Reaction (qRT-PCR)
2.6. Sequencing Library Research
2.7. Flow Cytometry
2.8. Statistical Analysis
3. Results
3.1. Analysis of mRNA Encoding Antimicrobial Proteins in Human Milk Cells
3.1.1. Higher Expression of Immune Cell Genes in the Mammary Gland and Milk Cells Taken during Pregnancy and Early Lactation
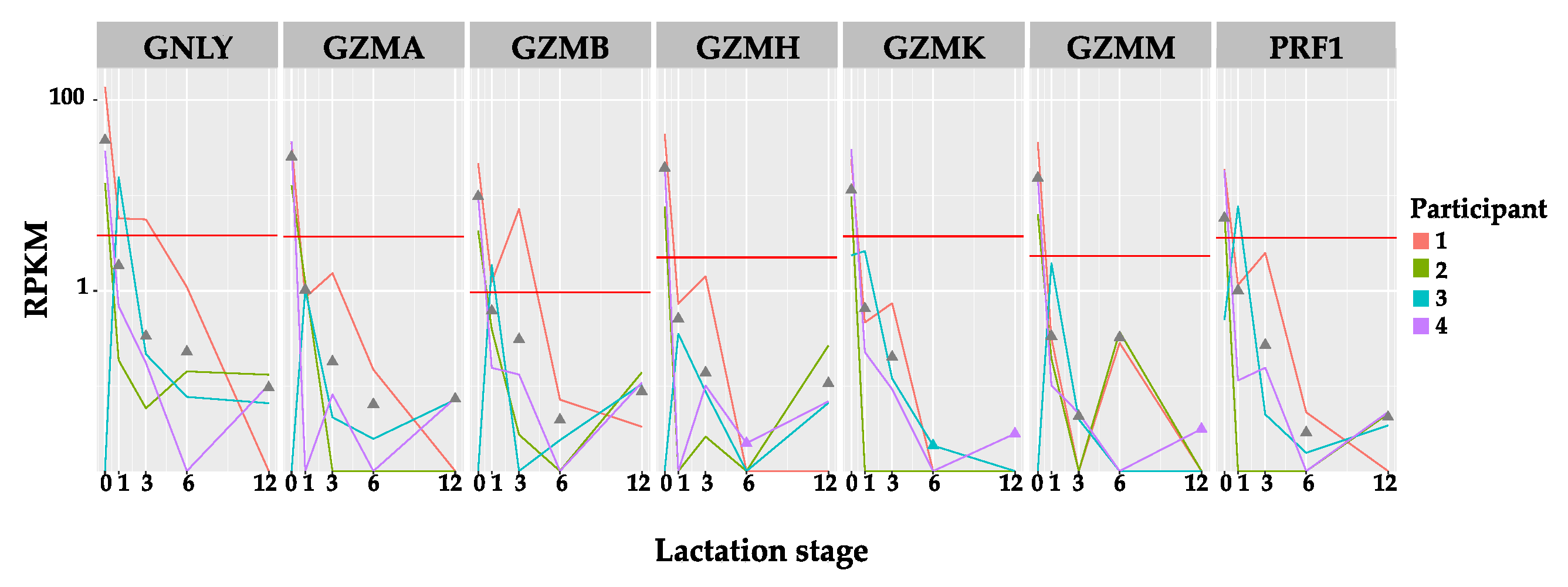
3.1.2. Expression Level of Immune Protein Encoding Genes Shows Little Association with Time Post-Partum
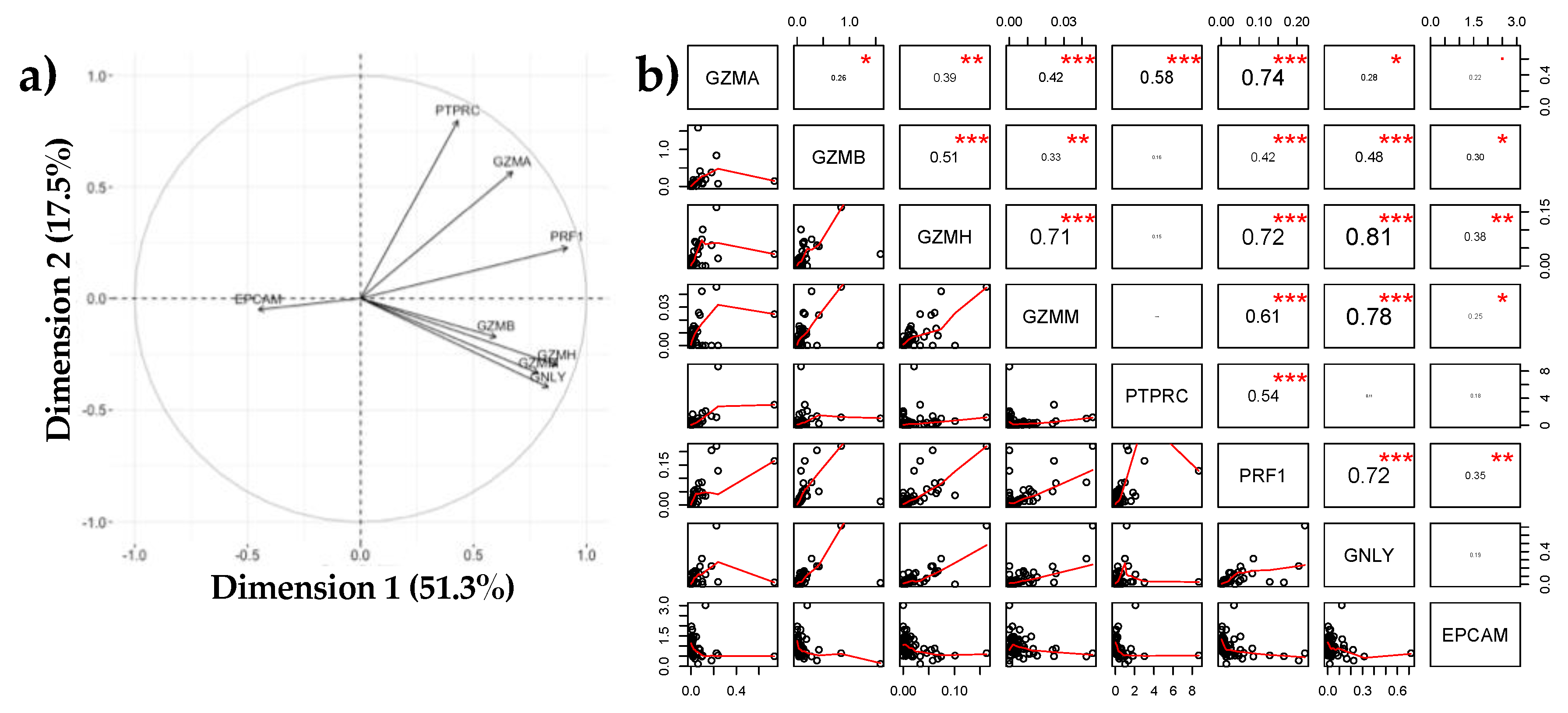
3.1.3. Negative Association between EPCAM and Immune Markers
3.1.4. Participants with Mastitis Show a Higher Expression of Immune Genes Compared to Healthy Participants
3.2. Analysis of Antimicrobial Proteins in Human Milk Cells
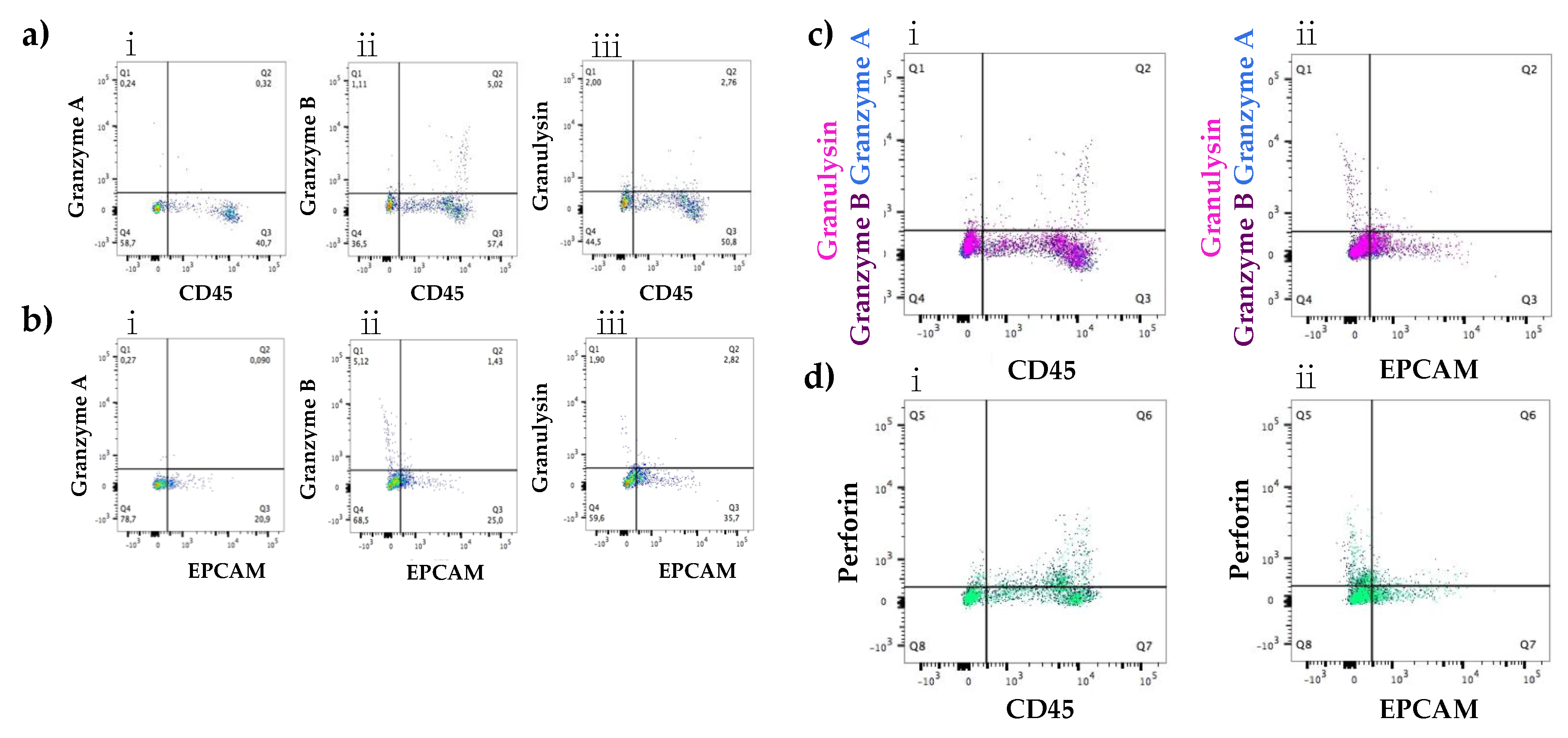
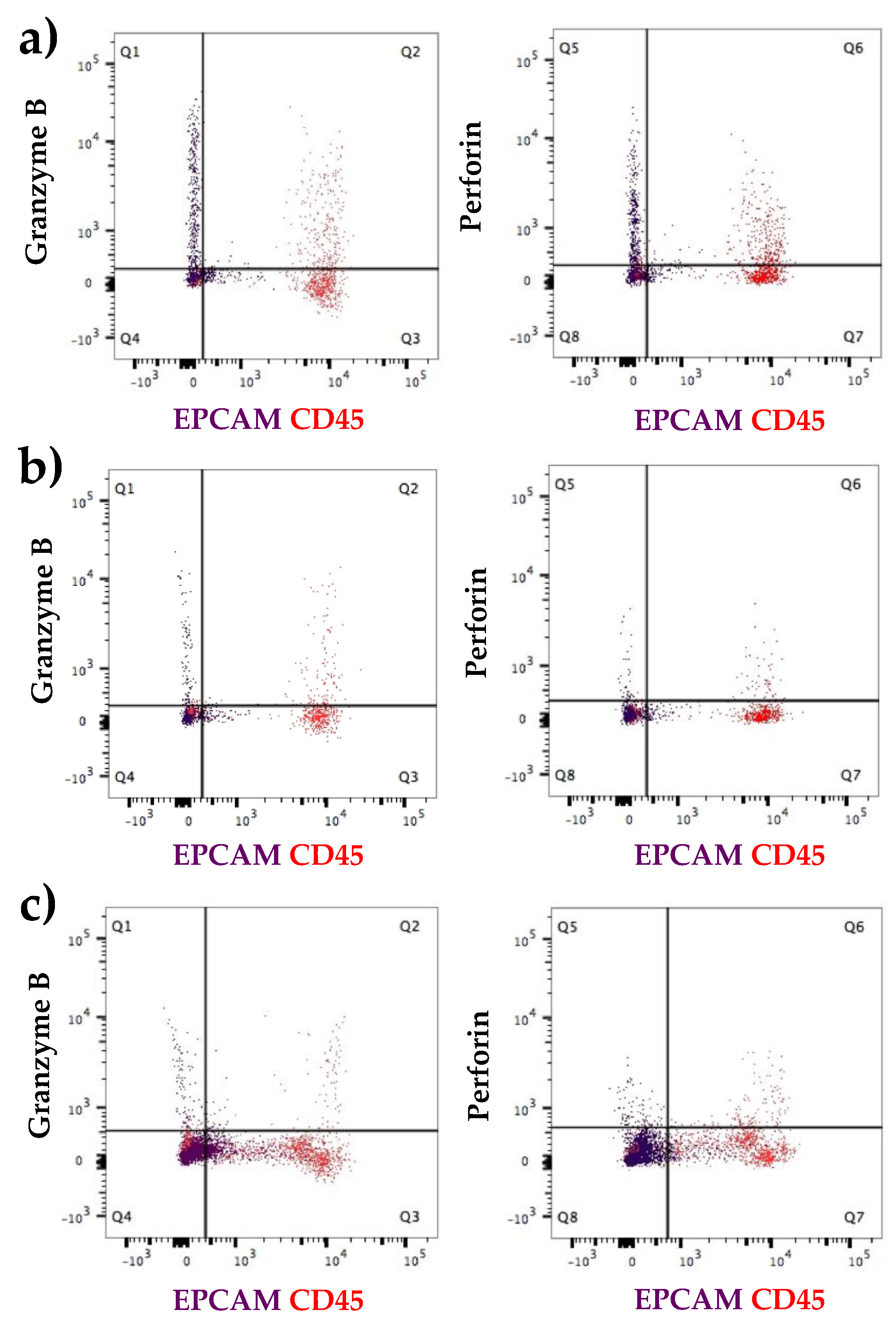
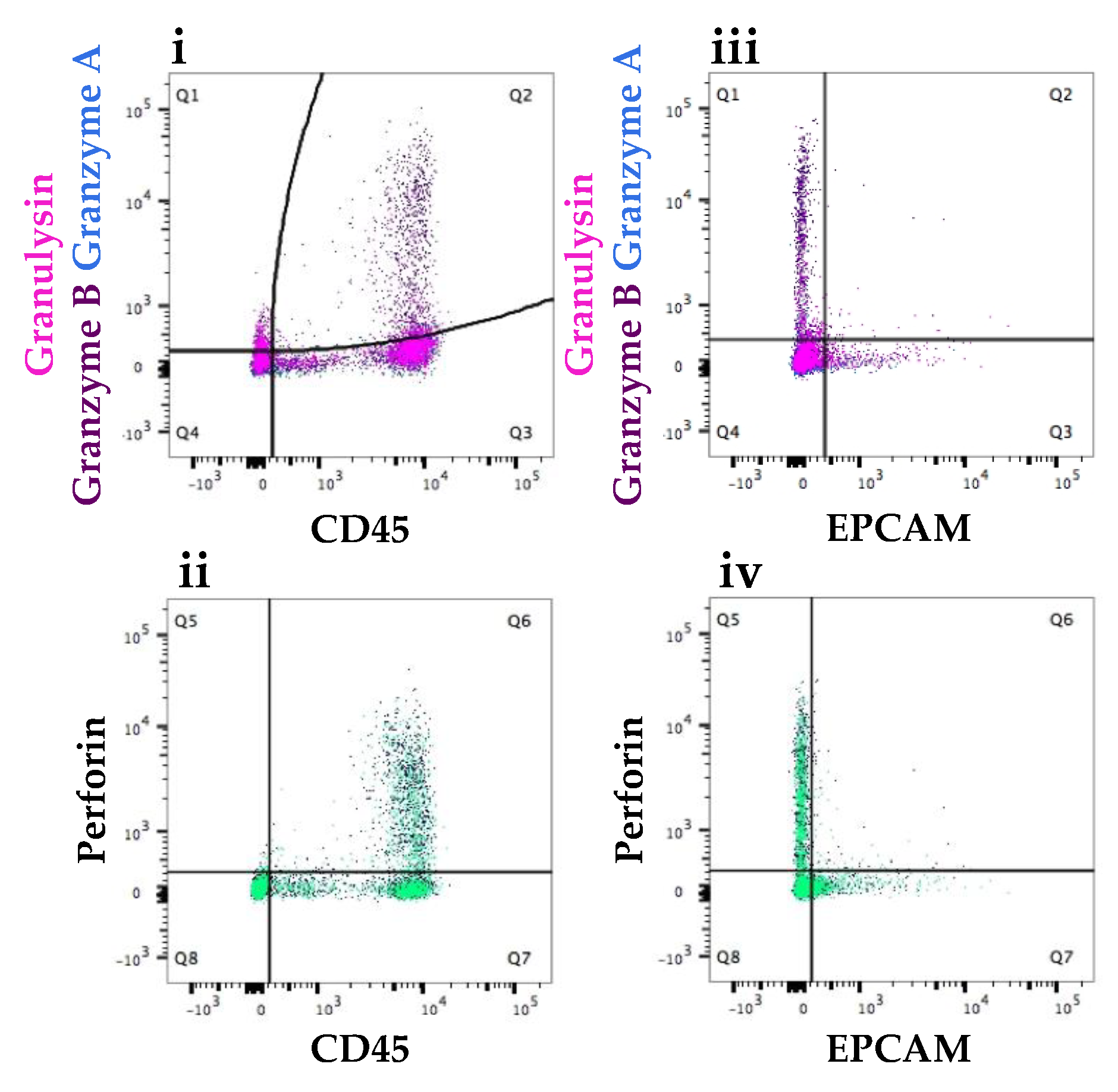
3.2.1. Expression at the Protein Level of All Immune Proteins in Healthy Participants
3.2.2. Increased Number of Immune Cells and Higher Expression of Immune Proteins in Mastitic Milk Samples in Comparison to Healthy Milk Samples
3.2.3. Participant Recovering from Non-Mammary Surgery Shows Higher Expression of Immune Proteins Compared to Healthy Participants
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hale, T.W.; Hartmann, P.E. Textbook of Human Lactation, 1st ed.; Hale Publishing L.P.: Amarillo, TX, USA, 2007; p. 662. [Google Scholar]
- Hanson, L.; Silfverdal, S.A.; Stromback, L.; Erling, V.; Zaman, S.; Olcen, P.; Telemo, E. The immunological role of breast feeding. Pediatr. Allergy Immunol. 2001, 12 (Suppl. S14), 15–19. [Google Scholar] [CrossRef]
- Hassiotou, F.; Geddes, D.T. Immune cell-mediated protection of the mammary gland and the infant during breastfeeding. Adv. Nutr. 2015, 6, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Liepke, C.; Zucht, H.D.; Forssmann, W.G.; Standker, L. Purification of novel peptide antibiotics from human milk. J. Chromatogr. B Biomed. Sci. Appl. 2001, 752, 369–377. [Google Scholar] [CrossRef]
- Zhang, F.; Cui, X.; Fu, Y.; Zhang, J.; Zhou, Y.; Sun, Y.; Wang, X.; Li, Y.; Liu, Q.; Chen, T. Antimicrobial activity and mechanism of the human milk-sourced peptide casein201. Biochem. Biophys. Res. Commun. 2017, 485, 698–704. [Google Scholar] [CrossRef] [PubMed]
- Hanson, L.A.; Korotkova, M.; Lundin, S.; Haversen, L.; Silfverdal, S.A.; Mattsby-Baltzer, I.; Strandvik, B.; Telemo, E. The transfer of immunity from mother to child. Ann. N. Y. Acad. Sci. 2003, 987, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Geddes, D.T.; Hartmann, P.E. Cells in human milk: State of the science. J. Hum. Lact. 2013, 29, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Hepworth, A.R.; Metzger, P.; Tat Lai, C.; Trengove, N.; Hartmann, P.E.; Filgueira, L. Maternal and infant infections stimulate a rapid leukocyte response in breastmilk. Clin. Transl. Immunol. 2013, 2. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Abdubek, P.; Astakhova, T.; Axelrod, H.L.; Bakolitsa, C.; Cai, X.; Carlton, D.; Chen, C.; Chiu, H.J.; Clayton, T.; et al. Structure of a membrane-attack complex/perforin (macpf) family protein from the human gut symbiont bacteroides thetaiotaomicron. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2010, 66, 1297–1305. [Google Scholar] [CrossRef] [PubMed]
- Nobre, T.M.; Martynowycz, M.W.; Andreev, K.; Kuzmenko, I.; Nikaido, H.; Gidalevitz, D. Modification of salmonella lipopolysaccharides prevents the outer membrane penetration of novobiocin. Biophys. J. 2015, 109, 2537–2545. [Google Scholar] [CrossRef] [PubMed]
- Walch, M.; Dotiwala, F.; Mulik, S.; Thiery, J.; Kirchhausen, T.; Clayberger, C.; Krensky, A.M.; Martinvalet, D.; Lieberman, J. Cytotoxic cells kill intracellular bacteria through granulysin-mediated delivery of granzymes. Cell 2014, 157, 1309–1323. [Google Scholar] [CrossRef] [PubMed]
- Dotiwala, F.; Mulik, S.; Polidoro, R.B.; Ansara, J.A.; Burleigh, B.A.; Walch, M.; Gazzinelli, R.T.; Lieberman, J. Killer lymphocytes use granulysin, perforin and granzymes to kill intracellular parasites. Nat. Med. 2016, 22, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Lieberman, J. Granzyme a activates another way to die. Immunol. Rev. 2010, 235, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Stenger, S.; Hanson, D.A.; Teitelbaum, R.; Dewan, P.; Niazi, K.R.; Froelich, C.J.; Ganz, T.; Thoma-Uszynski, S.; Melian, A.; Bogdan, C.; et al. An antimicrobial activity of cytolytic t cells mediated by granulysin. Science 1998, 282, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Bryan, D.L.; Hart, P.H.; Forsyth, K.D.; Gibson, R.A. Immunomodulatory constituents of human milk change in response to infant bronchiolitis. Pediatr. Allergy Immunol. 2007, 18, 495–502. [Google Scholar] [CrossRef] [PubMed]
- Twigger, A.J.; Kakulas, F. RNA-Sequencing of Milk Cells Extracted from Pre-Partum Secretions and Longitudinally from Mature Human Milk accros the First Year of Lactation. NCBI: Gene Expression Omnibus, 2016. Available online: https://www.ncbi.nlm.nih.gov/geo/ (accessed on 25 August 2016).
- Twigger, A.J.; Hepworth, A.R.; Lai, C.T.; Chetwynd, E.; Stuebe, A.M.; Blancafort, P.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Gene expression in breastmilk cells is associated with maternal and infant characteristics. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Hassiotou, F.; Beltran, A.; Chetwynd, E.; Stuebe, A.M.; Twigger, A.J.; Metzger, P.; Trengove, N.; Lai, C.T.; Filgueira, L.; Blancafort, P.; et al. Breastmilk is a novel source of stem cells with multilineage differentiation potential. Stem Cells 2012, 30, 2164–2174. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by rna-seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar]
- Sarkar, D. Lattice: Multivariate Data Visualization with R; Springer: New York, NY, USA, 2008. [Google Scholar]
- Pinheiro, J.; Bates, D.; DebRoy, S.; Sarkar, D.; Team, R.C. Nlme: Linear and Nonlinear Mixed Effects Models. R Package Version 3.1-131 2017. Available online: https://mran.microsoft.com/snapshot/2017-02-20/web/packages/nlme/index.html (accessed on 20 February 2017).
- Lê, S.; Josse, J.; Husson, F. Factominer: An r package for multivariate analysis. J. Stat. Softw. 2008, 25, 1–18. [Google Scholar] [CrossRef]
- Chirico, G.; Marzollo, R.; Cortinovis, S.; Fonte, C.; Gasparoni, A. Antiinfective properties of human milk. J. Nutr. 2008, 138, 1801s–1806s. [Google Scholar] [CrossRef] [PubMed]
- Sabbaj, S.; Ghosh, M.K.; Edwards, B.H.; Leeth, R.; Decker, W.D.; Goepfert, P.A.; Aldrovandi, G.M. Breast milk-derived antigen-specific cd8+ t cells: An extralymphoid effector memory cell population in humans. J. Immunol. 2005, 174, 2951–2956. [Google Scholar] [CrossRef] [PubMed]
- Wirt, D.P.; Adkins, L.T.; Palkowetz, K.H.; Schmalstieg, F.C.; Goldman, A.S. Activated and memory t lymphocytes in human milk. Cytometry 1992, 13, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Peroni, D.G.; Chirumbolo, S.; Veneri, D.; Piacentini, G.L.; Tenero, L.; Vella, A.; Ortolani, R.; Raffaelli, R.; Boner, A.L. Colostrum-derived b and t cells as an extra-lymphoid compartment of effector cell populations in humans. J. Matern. Fetal Neonatal Med. 2013, 26, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Cullen, S.P.; Martin, S.J. Mechanisms of granule-dependent killing. Cell Death Differ. 2007, 15, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Costello, P.; Bresnihan, B.; O’Farrelly, C.; FitzGerald, O. Predominance of cd8+ t lymphocytes in psoriatic arthritis. J. Rheumatol. 1999, 26, 1117–1124. [Google Scholar] [PubMed]
- Veugelers, K.; Motyka, B.; Goping, I.S.; Shostak, I.; Sawchuk, T.; Bleackley, R.C. Granule-mediated killing by granzyme b and perforin requires a mannose 6-phosphate receptor and is augmented by cell surface heparan sulfate. Mol. Biol. Cell 2006, 17, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, A.E.; Chapman, K.E. The anti-inflammatory and immunosuppressive effects of glucocorticoids, recent developments and mechanistic insights. Mol. Cell. Endocrinol. 2011, 335, 2–13. [Google Scholar] [CrossRef] [PubMed]
- Field, C.J. The immunological components of human milk and their effect on immune development in infants. J. Nutr. 2005, 135, 1–4. [Google Scholar] [CrossRef] [PubMed]
 ). For each gene, the range of expression is shown by the whiskers of the plot and the interquartile ranges are displayed as the upper and lower sections of the boxes. The median expression in milk cells for each gene is represented by the horizontal line going through the box. In the case of milk cell outliers, these are represented by black circles. HeLa cells (◆) were used as a negative control whereas Lymphokine Activated Killer cells (LAK, ■) were a positive control for immune markers. All genes were normalized to resting mammary tissue (RMT, ●). The measured standard error of the mean for these reference samples are represented by error bars. All immune related genes were expressed in the HM cells, although in some cases having a lower expression compared to RMT and LAK.
). For each gene, the range of expression is shown by the whiskers of the plot and the interquartile ranges are displayed as the upper and lower sections of the boxes. The median expression in milk cells for each gene is represented by the horizontal line going through the box. In the case of milk cell outliers, these are represented by black circles. HeLa cells (◆) were used as a negative control whereas Lymphokine Activated Killer cells (LAK, ■) were a positive control for immune markers. All genes were normalized to resting mammary tissue (RMT, ●). The measured standard error of the mean for these reference samples are represented by error bars. All immune related genes were expressed in the HM cells, although in some cases having a lower expression compared to RMT and LAK.
 ). For each gene, the range of expression is shown by the whiskers of the plot and the interquartile ranges are displayed as the upper and lower sections of the boxes. The median expression in milk cells for each gene is represented by the horizontal line going through the box. In the case of milk cell outliers, these are represented by black circles. HeLa cells (◆) were used as a negative control whereas Lymphokine Activated Killer cells (LAK, ■) were a positive control for immune markers. All genes were normalized to resting mammary tissue (RMT, ●). The measured standard error of the mean for these reference samples are represented by error bars. All immune related genes were expressed in the HM cells, although in some cases having a lower expression compared to RMT and LAK.
). For each gene, the range of expression is shown by the whiskers of the plot and the interquartile ranges are displayed as the upper and lower sections of the boxes. The median expression in milk cells for each gene is represented by the horizontal line going through the box. In the case of milk cell outliers, these are represented by black circles. HeLa cells (◆) were used as a negative control whereas Lymphokine Activated Killer cells (LAK, ■) were a positive control for immune markers. All genes were normalized to resting mammary tissue (RMT, ●). The measured standard error of the mean for these reference samples are represented by error bars. All immune related genes were expressed in the HM cells, although in some cases having a lower expression compared to RMT and LAK.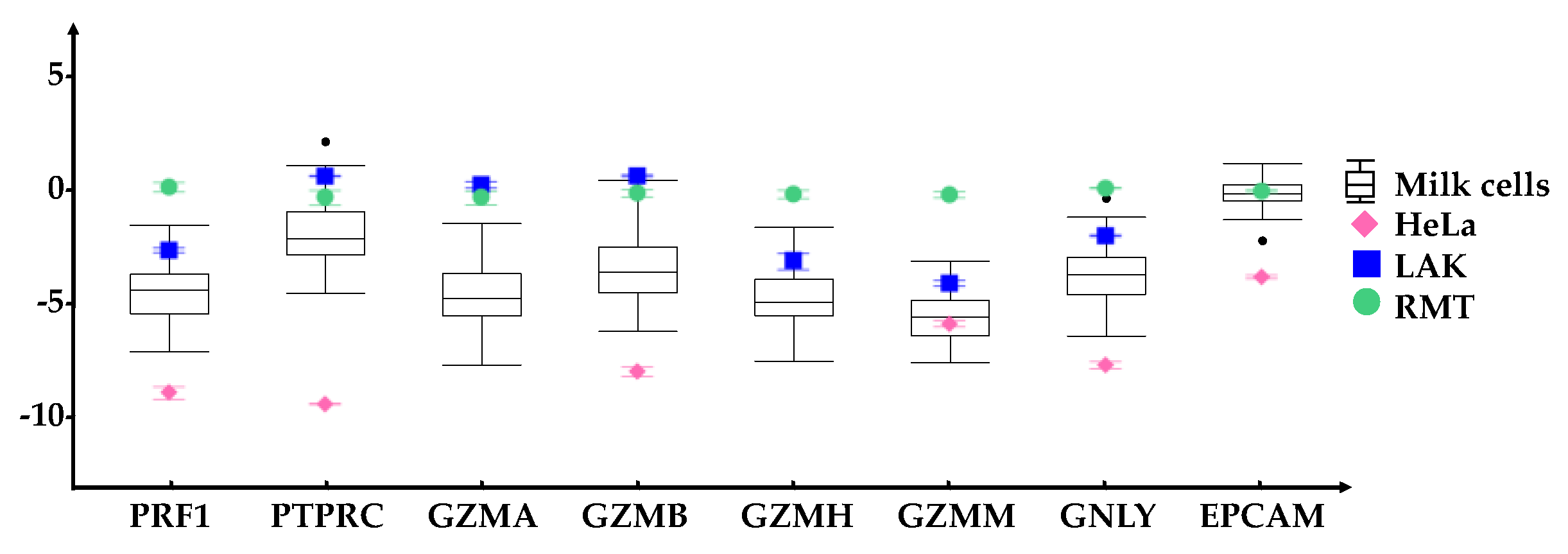

| Median (Range) | |||
|---|---|---|---|
| All Participants | PCR Participants | Flow Cytometry Participants | |
| Maternal characteristics | n = 36 | n = 24 | n = 13 |
| Age (years) | 34 (27–45) | 33 (27–45) | 35.5 (34–38) |
| Body Mass Index (BMI) | 24 (19.8–31.9) | 22.5 (19.8–27.9) | 25.6 (22–31.9) |
| Parity | 2 (1–4) | 2 (1–4) | 2 (1–3) |
| Infant characteristics | n = 36 | n = 24 | n = 13 |
| Gestational age (days) | 278 (249–301) | 278 (249–301) | 275 (252–281) |
| Infant age at collection (days) | 47 (4–142) | 45 (4–142) | 57 (37–84) |
| Milk characteristics | n = 85 | n = 70 | n = 15 |
| Volume of milk (mL) | 50 (0.61–490) | 50 (0.61–490) | 62 (35–195) |
| Total cell count (cells/mL) | 16.4 (1.9–214.5) | 16.55 (1.9–214.5) | 13.8 (4.9–63.8) |
| Viability (%) | 97.4 (53.2–100) | 98.1 (53.2–100) | 93.6 (84.7–96.5) |
| Healthy (n = 12) | Mastitis Sample (n = 1) | Non-Mammary Surgery Sample (n = 1) | ||
|---|---|---|---|---|
| Median (Range) | Mastitis Breast | Non-Mastitic Adjacent Breast | ||
| CD45+ (%) | ||||
| All single cells | 7.8 (2.1–13.4) | 12.2 | 8.4 | 26.7 |
| Population 1 | 51.5 (1.5–66.7) | 78.2 | 64.8 | 66.8 |
| Population 2 | 11.8 (1.9–51.5) | 69.6 | 73.5 | 71.8 |
| Population 3 | 2.7 (1.2–8.7) | 3.2 | 1.5 | 17.3 |
| Granzyme A+ (%) | ||||
| All single cells | 0.3 (0.0–10.6) | 0.8 | 0 | |
| Population 1 | 1.9 (0.4–82.0) | 7.1 | 4.3 | |
| Population 2 | 0.5 (0.0–19.2) | 1.6 | 0 | |
| Population 3 | 0.9 (0.00–1.3) | 0.4 | 0 | |
| Granzyme B+ (%) | ||||
| All single cells | 2.7 (0.0–4.1) | 0.4 | 1.2 | 2.6 |
| Population 1 | 11.9 (0.3–39.6) | 37.7 | 17.4 | 33.1 |
| Population 2 | 0.8 (0.0–9.5) | 1.1 | 1.7 | 7.8 |
| Population 3 | 0.4 (0.0–4.3) | 0.3 | 0.6 | 10.1 |
| Granulysin+ (%) | ||||
| All single cells | 1.6 (0.3–12.3) | 1.6 | 2.2 | |
| Population 1 | 11.7 (1.2–18.8) | 4.3 | 13 | |
| Population 2 | 2.0 (0.4–12.8) | 4.5 | 12.4 | |
| Population 3 | 15.5 (0.6–30.3) | 1.1 | 15.7 | |
| Perforin+ (%) | ||||
| All single cells | 1.4 (0.5–2.5) | 2.3 | 1.4 | 2.4 |
| Population 1 | 8.4 (0.0–31.7) | 37.4 | 8.6 | 27 |
| Population 2 | 2.2 (0.5–13.9) | 4.5 | 9.7 | 10.3 |
| Population 3 | 1.3 (0.8–9.5) | 0.3 | 0.6 | 1.1 |
| EPCAM+ (%) | ||||
| All single cells | 4.5 (3.8–12.8) | 6.6 | 5.1 | 3.4 |
| Population 1 | 18.2 (3.5–36.8) | 18.2 | 11.4 | 13.4 |
| Population 2 | 20.2 (5.8–26.0) | 23.4 | 12.9 | 9.5 |
| Population 3 | 19.9 (9.4–30.3) | 19.3 | 16.5 | 7.1 |
| Healthy (n = 12) | Mastitis Sample (n = 1) | Non-Mammary Surgery Sample (n = 1) | ||
|---|---|---|---|---|
| Median (Range) | Mastitis Breast | Adjacent Breast | ||
| CD45–Granzyme A (%) | ||||
| All single cells | 0.2 (0.0–14.9) | 0.2 | 0.4 | |
| Population 1 | 1.0 (0.3–32.6) | 10.9 | 5.4 | |
| Population 2 | 0.2 (0.0–50.0) | 3.5 | 1.3 | |
| Population 3 | 0.2 (0.0–0.4) | 0.2 | 0 | |
| CD45–Granzyme B (%) | ||||
| All single cells | 0.7 (0.0–5.5) | 0.5 | 0.4 | 4.4 |
| Population 1 | 5.2 (0.0–26.8) | 27 | 17.2 | 33.4 |
| Population 2 | 0.3 (0.0–13.9) | 0.3 | 4.2 | 20.3 |
| Population 3 | 0.0 (0.0–0.3) | 0 | 0.3 | 1.4 |
| CD45–Granulysin (%) | ||||
| All single cells | 0.7 (0.0–10.4) | 0.3 | 4.1 | |
| Population 1 | 3.2 (0.8–21.1) | 4.4 | 12.9 | |
| Population 2 | 3.6 (0.0–13.4) | 5.8 | 34 | |
| Population 3 | 13.6 (0.2–27.1) | 0.1 | 0.9 | |
| CD45–Perforin (%) | ||||
| All single cells | 0.9 (0.6–8.3) | 0.8 | 0.9 | 23.7 |
| Population 1 | 11.0 (0.1–35.2) | 34.1 | 8.2 | 27.7 |
| Population 2 | 2.1 (0.1–26.8) | 3.9 | 10.6 | 48.2 |
| Population 3 | 0.7 (0.1–1.8) | 0.7 | 0.1 | 4.8 |
| EPCAM–Granzyme A (%) | ||||
| All single cells | 0.0 (0.0–0.2) | 0.1 | 0 | |
| Population 1 | 0.1 (0.0–35.2) | 0.1 | 0.1 | |
| Population 2 | 0.4 (0.0–1.5) | 0.6 | 0 | |
| Population 3 | 1.0 (0.2–1.8) | 0.2 | 0 | |
| PCAM–Granzyme B (%) | ||||
| All single cells | 0.2 (0.1–1.0) | 1.3 | 0 | 1.5 |
| Population 1 | 0.5 (0.3–3.5) | 2.2 | 0.6 | 0.9 |
| Population 2 | 1.4 (0.0–3.0) | 0.3 | 0.5 | 6.1 |
| Population 3 | 1.3 (0.2–2.3) | 0 | 0.2 | 1.1 |
| EPCAM–Granulysin (%) | ||||
| All single cells | 0.2 (0.0–2.1) | 0.1 | 2.4 | |
| Population 1 | 1.5 (0.2–3.1) | 1.8 | 1 | |
| Population 2 | 0.6 (0.1–1.4) | 1.4 | 7.9 | |
| Population 3 | 2.6 (0.2–5.0) | 0.1 | 1.9 | |
| EPCAM–Perforin (%) | ||||
| All single cells | 0.2 (0.0–5.5) | 0.2 | 0.1 | 5.2 |
| Population 1 | 3.3 (1.1–9.5) | 2.1 | 0.5 | 2.2 |
| Population 2 | 1.7 (0.3–8.4) | 0.7 | 1.1 | 7.4 |
| Population 3 | 0.4 (0.3–0.4) | 0.3 | 0 | 1.8 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Twigger, A.-J.; Küffer, G.K.; Geddes, D.T.; Filgueria, L. Expression of Granulisyn, Perforin and Granzymes in Human Milk over Lactation and in the Case of Maternal Infection. Nutrients 2018, 10, 1230. https://doi.org/10.3390/nu10091230
Twigger A-J, Küffer GK, Geddes DT, Filgueria L. Expression of Granulisyn, Perforin and Granzymes in Human Milk over Lactation and in the Case of Maternal Infection. Nutrients. 2018; 10(9):1230. https://doi.org/10.3390/nu10091230
Chicago/Turabian StyleTwigger, Alecia-Jane, Gwendoline K. Küffer, Donna T. Geddes, and Luis Filgueria. 2018. "Expression of Granulisyn, Perforin and Granzymes in Human Milk over Lactation and in the Case of Maternal Infection" Nutrients 10, no. 9: 1230. https://doi.org/10.3390/nu10091230
APA StyleTwigger, A.-J., Küffer, G. K., Geddes, D. T., & Filgueria, L. (2018). Expression of Granulisyn, Perforin and Granzymes in Human Milk over Lactation and in the Case of Maternal Infection. Nutrients, 10(9), 1230. https://doi.org/10.3390/nu10091230




