1. Introduction
Forest ecosystems are fundamental to human society and the living world, which depends on them for survival and growth [
1,
2]. Forests serve as essential reservoirs of stable organic carbon, genetic diversity, and energy resources [
3,
4,
5], while playing a decisive role in maintaining ecological balance [
6,
7], supporting environmental conservation, and sustaining the fundamental conditions for human survival and development [
8,
9]. The natural environmental conditions and effects provided by forest ecosystems to sustain human life are called forest ecosystem services, which are formed by the structure and processes of forest ecosystems [
10,
11,
12]. Forest ecosystem services are divided into seven categories: water conservation, soil conservation, carbon sequestration and oxygen release, nutrient accumulation, environmental purification, biodiversity conservation and forest recreation [
11,
12,
13,
14,
15]. In recent years, the theory of “four reservoirs”, including water [
13], food [
14], money [
15] and carbon [
11], has further enriched our understanding of forest ecosystem services: forests support the water cycle (water reservoir) through water conservation, regulating the water cycle and supporting forest recreation. Forests regulate the water cycle through water harvesting (water bank), support agriculture and contribute to food security (food bank), provide economic values such as timber and ecotourism (money bank), and mitigate global warming by absorbing and storing carbon dioxide (carbon bank). For a long time, the view of forest ecosystem services as a source of public goods has led to a widespread undervaluation of their benefits in terms of forest resource utilization, as well as a lack of proper recognition of their importance [
16]. This has resulted in the predatory exploitation of forest resources, severely damaging forest ecosystems and, in turn, contributing to increasingly critical ecological issues such as climate warming, biodiversity loss, and natural disasters. These ecological issues pose a great threat to human life and production. History has proved that the rise and fall of forest ecosystems are closely related to the sustainable development of human production and life [
17,
18,
19], and the theory of “four reservoirs” emphasizes the central position of forest ecosystems in maintaining ecological security and socio-economic development. The protection and development of forest ecosystems is not only a necessity for environmental protection [
20], but also key to achieving long-term prosperity and stability in human society [
21,
22].
Xizang is known for its extreme natural environment. The thin air and low oxygen conditions brought by the high altitude, together with the significant temperature difference between day and night, pose a serious challenge to the survival of plants and animals [
23]. Mozhugongka County, where the study area is located, is part of the Lhasa Valley Plain and has a typical plateau temperate semi-arid monsoon climate. The high altitude results in a cold and dry climate with low oxygen content in the air. The annual precipitation is about 515.9 mm, mainly concentrated from June to September, while precipitation is scarce in winter. The annual sunshine hours reach up to 2813.5 h and the frost-free period is about 90 days. Strong windy weather is common in winter and spring, accelerating water evaporation and increasing the risk of soil erosion. Such extreme climatic conditions pose a serious challenge to the local ecosystem [
24,
25].
Seabuckthorn (
Hippophae rhamnoides L.), as a drought-resistant, cold-resistant and sand-resistant plant [
26], has significant ecological restoration functions. Seabuckthorn is botanically classified as a deciduous shrub or small tree, exhibiting remarkable morphological plasticity in response to environmental conditions. In our study area, it occurs both as low shrub thickets and as arborescent individuals exceeding 3 m in height with distinct trunk development. This growth-form continuum is central to our objective of extracting phenotypic traits from tree-like individuals to support germplasm selection and breeding programs. Seabuckthorn has a well-developed root system, which can effectively fix the soil [
16], reduce soil erosion, and prevent desertification and land degradation [
27]. Its drought-resistant and barrenness-resistant characteristics mean that it is able to grow under harsh environmental conditions and make it an important ecological restoration plant. Seabuckthorn also has strong nitrogen-fixing ability and can grow on wasteland, making it a pioneer plant for windy and sandy land [
28]. Therefore, seabuckthorn has a strong ability to improve the ecological environment and create a biological chain. The nitrogen-fixing ability of seabuckthorn effectively contributes to soil fertilization and yield improvement. Its root nodules are formed through the infection of the roots by the endophytic bacterium
Frankia [
29], and exhibit a higher nitrogen fixation capacity compared to leguminous plants [
30]. Through the nitrogen fixation of the seabuckthorn root system, soil organic matter and nitrogen content are notably increased [
31], while the presence of and improvement in organic matter can reduce soil bulk weight and increase porosity. This is more conducive to improving the water storage capacity of the soil, and thus improves the soil’s geotechnical capacity [
32]. Therefore, planting seabuckthorn is considered an effective means to solve the problems of soil erosion and sanding [
33]. In China, the government attaches great importance to ecological protection and restoration. In the ecological management of the western and highland areas, seabuckthorn has become an important plant in the implementation of ecological restoration projects [
34]. Policies such as “returning farmland to forests and grasslands”, natural forest protection projects and desertification prevention and control projects implemented by the state have mentioned seabuckthorn as a key plant for greening and ecological restoration. In recent years, seabuckthorn has been widely planted in ecological restoration projects in Xizang and other places to restore grassland, woodland and desertified areas. Through the support of these policies, the scale of seabuckthorn cultivation has gradually expanded and begun to drive the dual development of the local ecology and economy. The reason for choosing the river valley plain of Mozhugongka County as the study area is that the diversity of seabuckthorn forms in this area increases the generality and representativeness of the results [
35]. The ecological characteristics and expression of seabuckthorn in this area can provide valuable reference cases for other, similar alpine, arid and semi-arid regions in China and worldwide [
36].
Unmanned aerial vehicle (UAV) remote sensing technology, compared with traditional satellite remote sensing, is more flexible and efficient, especially in ecological environment monitoring in areas with poor natural environments and sparse human trails [
37]. Compared with satellite-borne remote sensing data, UAV remote sensing data are more refined [
38], and can be better applied to individual tree-level research, especially on seabuckthorn (with different morphologies, such as shrubs, small trees, tall trees, etc.). UAV remote sensing shows the potential ability to extract the position, structure, and physiological and biochemical traits of seabuckthorn with high precision [
39] compared with ground-based remote sensing technology (e.g., ground-based LIDAR). UAV remote sensing can be used to more widely and quickly gather heterogeneous data from multiple sources [
40,
41,
42]. With their ease of deployment and operation, drones can quickly cover a designated area and acquire high-resolution images instantly, regardless of ground conditions. Equipped with advanced multispectral (or hyperspectral) and LiDAR sensors, drones are able to provide detailed surface information, such as land cover types, vegetation growth and health, and soil erosion. UAV remote sensing is particularly advantageous in arid and semi-arid regions [
20,
43,
44]. It can monitor the progress of desertification and land degradation with higher efficiency and frequency, as well as the process of plant recovery in real time, and has potential in monitoring natural seabuckthorn forests. The high-precision data collected by drones can make it possible to more precisely assess seabuckthorn vegetation cover, its health status and its relationship with soil and climate factors. In addition, it can quantify the impact of seabuckthorn planting on soil and water conservation, ecological restoration and land productivity enhancement, which provides strong data support for the effectiveness of ecological protection measures. In conclusion, the convenient deployment of UAV remote sensing and the high accuracy of the multi-source heterogeneous data it provides not only improve the efficiency of small- and medium-scale monitoring, but also provide technical support for optimizing ecological restoration strategies. The introduction of UAV remote sensing technology marks a new era of more efficient and accurate ecological environment monitoring.
Taking an individual tree as the basic unit of a forest and extracting its structural traits is the focus of forest resources investigation, and the most important traits of the individual tree structure are the tree height and crown [
45]. Tree height is the main index for forest resource investigation and monitoring, and accurately obtaining the tree height of forest trees is of great significance to the management of forest resources [
46]. Tree height can show the biological characteristics and growth capacity of trees, and is a critical indicator of tree growth [
47]. Tree height is also an indicator for determining the stand quality of a community, which determines the biomass of the community. Tree crowns are important for the survival and growth of trees, and they are indispensable for physiological activities, such as respiration, photosynthesis, and transpiration and can directly reflect the growth and dynamic changes in individual trees. Therefore, obtaining accurate structural information of individual trees can facilitate understanding of the competitive relationship among trees and allow for the monitoring and prediction of tree growth, helping to determine the health status and biomass (e.g., carbon storage) of the trees [
48].
The existing research on the use of remote sensing to monitor seabuckthorn is still in the preliminary stage, with very limited relevant research, a fragmented research scope, and a lack of systematicity and continuity [
24,
49,
50]. Several studies focus on classifying seabuckthorn as a shrubland component at the patch or community scale [
51], while others extract parameters from tree-form individuals in simplified, managed settings like plantations or shelterbelts [
52]. This highlights a critical gap, where no complete technical system exists that is capable of operationally distinguishing, segmenting, and phenotyping the intermixed growth forms within natural, structurally complex seabuckthorn forests. A complete technical system, from data acquisition and processing methods to information extraction, has not yet been formed, especially for the automatic extraction of individual wood-scale traits, which is almost a blank state. Therefore, this study aims to develop a suite of individual tree remote sensing monitoring solutions that are applicable to seabuckthorn. Systematic experiments and a full-process validation are conducted, covering multi-source remote sensing data acquisition, preprocessing, and feature extraction, as well as individual tree crown segmentation and trait prediction. This work aims to address the research gap in this field and to provide a repeatable, transferable technical reference for the standardized and operational application of remote sensing in seabuckthorn monitoring.
This study develops an integrated technical solution for the remote sensing-based monitoring of seabuckthorn at the individual tree level, utilizing UAV-based multispectral and LiDAR data. The specific research objectives include (1) proposing a method for precise tree-shrub classification from LiDAR point clouds through the integration of multi-scale spatial analysis and hierarchical clustering, supported by geometric constraints, density features, and dynamic threshold optimization; (2) extracting tree height and crown width from the individual tree segmentation results, and using hand-held LiDAR to measure Diameter at Breast Height (DBH) for constructing DBH estimation models; (3) predicting leaf nitrogen content at the individual tree level by combining multispectral data with spectra simulations from the N-PROSAIL model.
2. Materials and Methods
2.1. Study Area and Technology Workflow
The study area was located in Mozhugongka County, Lhasa City, within the Xizang Autonomous Region. This area is located in central Xizang, on the west side of the Mira Mountains, and encompasses part of the middle and upper reaches of the Lhasa River Valley, forming an important section of the mid-Yarlung Zangbo River valley system. The region belongs to the Lhasa Valley Plain and falls within the plateau temperate semi-arid monsoon climate zone.
Due to its high elevation, the climate of Mozhugongka County is characterized by cold, arid conditions with thin air. Winters and springs are windy, and the annual temperature variation is small, but the diurnal temperature range is large. The area had an annual frost-free period of approximately 90 days, with sunshine hours exceeding 2813 per year. Annual precipitation was about 515 mm, predominantly occurring between June and September. The geographical coordinates of the study area ranged from 91.74°E to 91.77°E and 29.82°N to 29.84°N. The study area comprises natural, monodominant stands of seabuckthorn. The vegetation structure is characterized by a continuous morphological gradient within the species, ranging from low, multi-stemmed shrubs to distinct, single-stemmed arborescent individuals.
A comprehensive field survey was conducted on 120 sample trees. Tree height was measured using a Vertex IV ultrasonic hypsometer (Haglöf, Switzerland), diameter at breast height (DBH) was obtained with a diameter tape, and precise individual tree locations were recorded using a Qianxun SR6 RTK system (Qianxun SI, Chengdu, China). Additionally, three-dimensional point cloud data of the sample plot were collected using a LiGrip H120 handheld laser scanner (Green Valley Technology Co., Ltd., Beijing, China). Boundary markers and closed scanning paths were employed to ensure data completeness and spatial alignment accuracy. Leaf samples were collected from 37 representative trees. Leaves were then dried at 80 °C for 48 h, after which the dried seabuckthorn leaves were ground, sieved (0.25 mm), and analyzed with the micro-Kjeldahl method [
53] to determine leaf nitrogen content (LNC) (mg/g). This integrated dataset provides a multi-source foundation for subsequent analyses, encompassing tree structural parameters, spatial distribution, three-dimensional morphology, and biochemical traits.
The technical workflow was structured into four key steps, as outlined below. Firstly, data were collected sequentially using three systems: UAV LiDAR data acquired with a DJI M350 RTK UAV (DJI, Shenzhen, China) equipped with a Huace AA-10 LiDAR sensor (Huace, Beijing, China); handheld LiDAR data obtained using a Ligrip H120 scanner; and multispectral imagery captured by a DJI Mavic 3M UAV (DJI, Shenzhen, China). The UAV and handheld LiDAR data were preprocessed through stitching, cropping, and denoising, while the multispectral imagery underwent stitching, cropping, radiometric calibration, and atmospheric correction. Next, tree and shrub point clouds were separated. Based on the preprocessed UAV LiDAR data, a Cloth Simulation Filter (CSF) [
54] was applied to isolate ground points and extract above-ground vegetation. The normalized point cloud was then processed at multiple scales to identify candidate tree points, which were subsequently clustered using the Ordering Points to identify the clustering structure algorithm (OPTICS) [
55] and finalize the separation. Subsequently, individual tree segmentation was performed. From the separated tree point cloud, a Digital Elevation Model (DEM) and a Digital Surface Model (DSM) were generated by interpolation, and a Canopy Height Model (CHM) was derived. A marker-controlled watershed algorithm [
17] was applied to the CHM for crown segmentation, while a hierarchical clustering algorithm was used on the normalized point cloud for direct point-based segmentation. Finally, individual tree traits were extracted. Tree height and crown width were derived from the UAV LiDAR data, and DBH was obtained from the handheld LiDAR data. Predictive models were developed to estimate these structural parameters directly from the UAV LiDAR data. For biochemical traits, LNC was retrieved by integrating the segmented crown boundaries with the preprocessed multispectral data, supported by both field measurements and N-PROSAIL model simulations.
2.2. UAV Data Acquisition and Pre-Processing
This study was carried out within the natural seabuckthorn forests in Mozhugongka County, Lhasa City, Xizang Autonomous Region. The LiDAR point cloud data acquisition was carried out using a DJI M350 RTK UAV equipped with a Huace AA-10 LiDAR sensor. A total of nine flights were carried out, with the flight strip width set to 180 m, a flight speed of 8 m/s, and a relative altitude of 190 m (the ground elevation of the survey area ranges from 3800 m to 3950 m). The overlapping degree of heading sideways was controlled at 30–40%, the laser emission frequency was 100 kHz, the scanning speed was 90 RPM, and the starting and stopping scanning angles were set at 135–225°, so that the density of the point cloud acquired finally reached 80–120 points/m
2. The LiDAR acquisition equipment is shown in
Figure 1a.
The multispectral data were collected by a DJI Mavic 3M multispectral UAV. The central wavelength and bandwidth information of each band are shown in
Figure 1, and the sensor spectral response curve is shown in
Figure 1. The preset spatial resolution of the mission was 0.12 m. The shutter-priority mode was used, with a range of shutter speeds from 1/1200 to 1/2000, and the ISO setting was set to automatic. The overlap rates of heading and sidetracking were set to 80% and 75%, respectively, and the flight speed was 14 m/s.
The point cloud data were subsequently subjected to pre-processing steps, such as splicing, cropping and denoising. The multispectral data were processed with radiometric calibration, smoothing and denoising, and precise geo-alignment. The multispectral UAV acquisition equipment is shown in
Figure 1a.
2.3. Individual Tree Segmentation Methods
Addressing the significant interference caused by the mixed growth of trees and shrubs and their distinct morphological disparities in individual tree segmentation within natural seabuckthorn forests, this study adopts a two-stage strategy [
56]. First, a high-precision separation of trees and shrubs is performed on the point cloud data by integrating multi-scale dynamic thresholds, using hierarchical clustering techniques to accurately extract tree points. Subsequently, individual tree segmentation is implemented using both point cloud-based clustering and marker-controlled watershed algorithms on the purified tree point cloud to enhance the overall segmentation accuracy.
This study employed a vegetation point cloud classification method that integrates multi-scale spatial analysis with hierarchical clustering. The original LiDAR point cloud first underwent preprocessing, which involved extracting the 3D coordinates of each point and removing invalid data points using a dual-threshold filtering approach. Subsequently, the (CSF) algorithm was applied to separate ground points from non-ground points, resulting in a purified above-ground vegetation dataset for the subsequent tree–shrub classification. For the obtained non-ground point cloud, this study introduced a tree point detection method that combines multi-scale spatial analysis with a dynamic height threshold. Specifically, for each point in the cloud, the relative height and point cloud density features were calculated using three different neighborhood radii: 3.0 m, 5.0 m, and 7.0 m. To capture cross-scale spatial features ranging from individual tree crown details to the broader tree group environment, different spatial resolutions were applied: 3.0 m for resolving individual tree crowns or dense shrubs, 5.0 m for analyzing the local structure of tree clusters or the canopy of larger individual trees, and 7.0 m for characterizing tree groups, forest stand edges, or forest canopy gaps. The process is illustrated in
Figure 2. The formulas for calculating the relative height
and the point cloud density
are listed as follows:
where
is the height of point
,
denotes the set of points within the neighborhood of radius
r centered at
, and
is the number of points within this neighborhood.
The dynamic threshold was set with a base height threshold of 1.5 m, incorporating a density-adaptive adjustment mechanism. In sparse regions (low density), the threshold was appropriately increased to 1.8 m to suppress misclassification. The formula for calculating the dynamic threshold
is
This formula uses a density penalty term (coefficient 0.3) to reduce misjudgments in low-density areas. The density penalty coefficient was determined through an empirical optimization process. We evaluated a range of values on a representative subset of the study area, with each candidate coefficient assessed via visual inspection of the resulting separation accuracy. A value of 0.3 was optimally selected, as it effectively balanced sensitivity and specificity. It applied a sufficient penalty to raise the adaptive threshold in sparse regions, thereby reducing false inclusions from background vegetation, while remaining conservative enough to avoid incorrectly excluding genuine tree points in moderately dense areas. This calibrated value was subsequently applied consistently across the entire study area.
In such areas, where
is small,
is larger, increasing the density penalty term and thus raising the value of
. This means that stronger feature signals are required in low-density regions for a point to be classified as a tree. Simultaneously, the dynamic threshold adjustment allows for weak signals in sparse areas to be identified, thereby enabling the detection of isolated trees. The OPTICS algorithm is applied to the candidate tree points for hierarchical density-based clustering. The core distance and reachability distance for each point are calculated. The core distance
is defined as the minimum radius required to contain at least 10 neighboring points, calculated as
The reachability distance
is calculated as
Subsequently, a point ordering is generated based on the ascending reachability distance. Different density clusters are identified by analyzing the slope changes in the reachability plot. A new cluster is delineated when the increase in reachability distance between consecutive points exceeds three times the standard deviation. The maximum neighborhood radius is set to
. This forms an efficient multi-scale cascade analysis, where a 7.0 m radius is used at the front end to capture the macro-environment for optimized initial screening, while a 6.0 m radius is applied at the back end to focus on segmenting individual tree objects at typical crown scales, effectively avoiding under-segmentation. Finally, points belonging to the clusters identified in the OPTICS result are extracted as the tree point cloud. Shrub points are separated using a logical exclusion method, achieved by inversely selecting points that are neither classified as trees nor as ground points. Matrix operations are used to label candidate tree points, and their inverse yields the candidate shrub points:
Additionally, height constraints (Z < 3.0 m) and density constraints (point density in shrub areas is lower than in candidate tree areas) are applied to optimize the classification result. Morphological post-processing operations (such as opening) can be further employed to remove noise points and enhance the connectivity of shrub regions.
The advantage of this method lies in its multi-scale complementarity and dynamic trait optimization: a small radius (3 m) captures isolated trees, while a large radius (7 m) identifies extensive canopies. It adaptively adjusts search radii and threshold weights based on point density and validates the rationality of the classification results through height-position profile analysis. This method effectively improves the accuracy of tree–shrub separation in complex scenarios, providing a reliable data foundation for subsequent biomass estimation and ecological analysis.
Two methods were employed for individual tree segmentation and to compare their performance: point cloud-based cluster segmentation and marker-controlled watershed segmentation based on a canopy height model. In the point cloud segmentation method, the raw point cloud data acquired by LiDAR were first systematically preprocessed. Noise filtering was achieved by statistical outlier removal, the improved fabric simulation filtering algorithm was used to separate ground and non-ground points, and the digital elevation model and digital surface model were generated based on the progressive triangular mesh encryption algorithm; then, the canopy height model was computed for the two differences, and the same specification of the digital elevation model was normalized with the point cloud data. In the selection of the segmentation algorithm, for the special morphology of seabuckthorn scrub, both the DBSCAN algorithm based on density clustering and the Euclidean clustering method were tested to achieve individual tree segmentation through three-dimensional spatial neighborhood analysis. Additionally, they were supplemented with vertical profile character analysis to detect the local extreme value points in the cross-section of 0.5 m interval of canopy height and to determine the position of tree tops by combining the characteristics of the curvature change [
57].
The marker-controlled watershed segmentation scheme was then unfolded based on the canopy height model [
58]. Firstly, anisotropic Gaussian filtering was applied to the 0.5 m resolution CHM to smooth out small undulations, and then local maximum detection in a dynamic window was used to locate potential tree tops. Considering the clumping nature of buckthorn scrub, morphological reconstruction techniques were introduced to eliminate pseudo-markers: firstly, small areas were removed by area-splitting operations, and then marker fusion algorithms based on canopy geometry features were applied.
The segmentation results generated by each algorithm were mainly examined by visual interpretation. We visually checked whether the segmented individual trees are reasonable, judged whether they were over-segmented or under-segmented, and checked the segmentation accuracy using precision, recall, over-segmentation rate and F1-score values.
where
TP,
FN, and
FP denote the number of correct segmentations, missed segmentations and over-segmentations, respectively.
2.4. Tree Height Trait Extraction
In this study, tree height was derived based on the results of high-precision individual tree segmentation. The point clouds of individual trees were normalized to eliminate the influence of terrain fluctuations by converting the absolute elevations into relative heights above the ground. Subsequently, the highest point within each normalized point cloud was identified and extracted. The elevation value of this peak point was directly taken as the tree height.
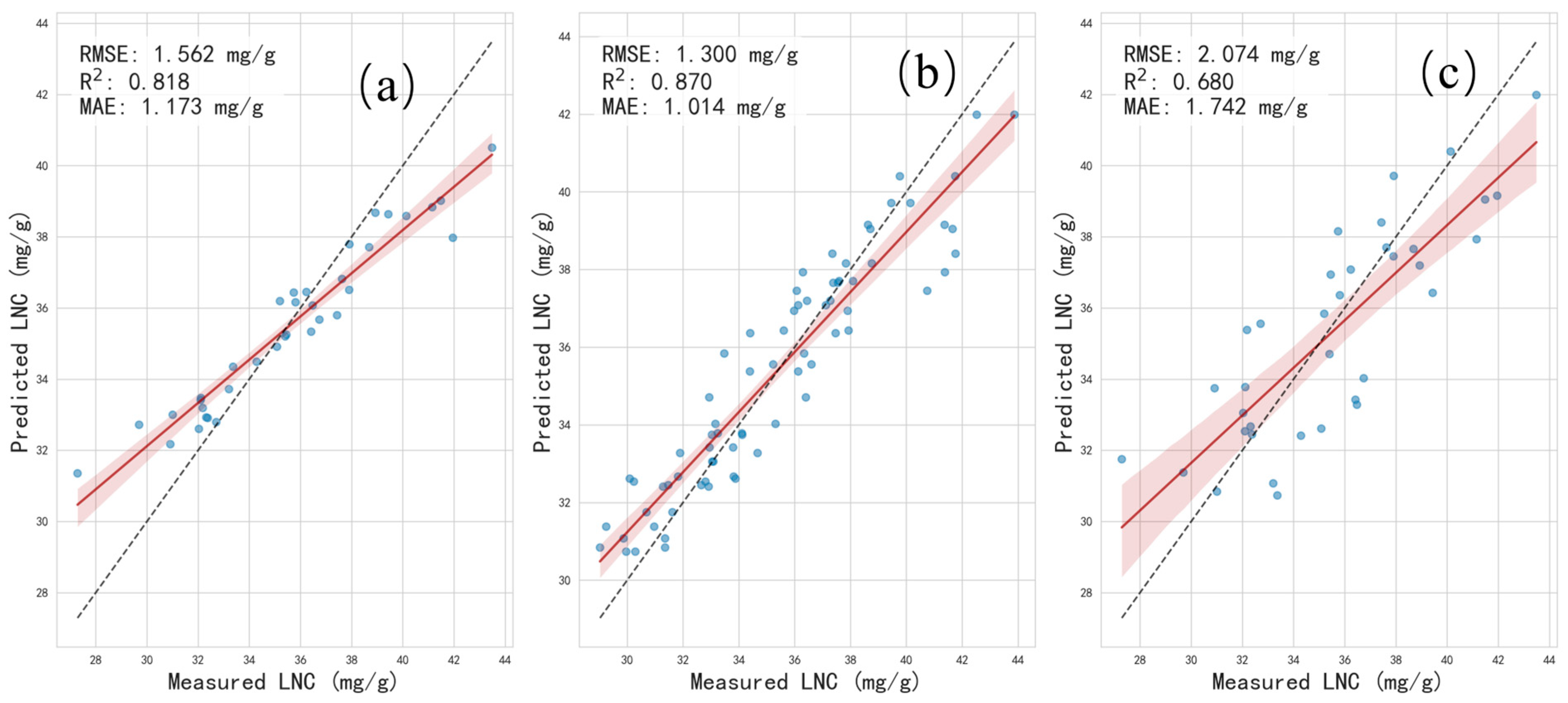
This study employs a stratified accuracy assessment framework to systematically evaluate the performance of a tree height prediction model. The samples were first divided into a low-error group (comprising the lower two-thirds of the absolute error distribution) and a high-error group (the upper one-third) based on the tertiles of absolute error. Additionally, a high-quality subset was defined using a relative error threshold of ≤20%, establishing a three-tier quality stratification system. The evaluation incorporated R2 and RMSE metrics, visualized through scatter plots showing the regression relationship between predicted and actual values (with 95% confidence intervals) against the y = x ideal line, along with boxplots comparing the distributions of actual values, predicted values, and absolute errors across the three groups. This multi-level assessment framework provides an overall accuracy measure and identifies the characteristics of high-quality predictions, offering targeted insights for model refinement and practical application under varying precision requirements.
To systematically evaluate the accuracy of the tree height extraction, field-measured tree height data that were spatially aligned with the LiDAR data were used as reference values. The root–mean–square error (RMSE) and mean absolute error (MAE) were employed to quantify the magnitude of the estimation errors, while the coefficient of determination (R
2) was used to assess the linear agreement between the LiDAR-derived tree heights and the measured tree heights.
2.5. Diameter at Breast Height Prediction Model
In this study, an individual tree diameter at breast height (DBH) prediction model was developed using multiple regression, based on airborne LiDAR point cloud data and field-measured sample tree data. Tree height (H) and crown width (CW) were extracted as predictor variables from the point clouds of segmented individual trees. The DBH reference values were obtained from high-precision handheld LiDAR ground measurements. Tree height was calculated as the vertical difference between the highest point in the individual tree point cloud and the corresponding ground elevation, while crown width was derived from the major axis of the minimum bounding rectangle of the tree crown’s horizontal projection.
To systematically evaluate the effects of model structure and feature combinations on DBH prediction accuracy, three types of regression models were constructed: linear, quadratic polynomial, and power function models. Two variable sets were designed for the independent variables: the first used only tree height as the input variable (H model), and the second incorporated both tree height and crown width (H + CW model). This setup allowed for the additional contribution of crown width to be assessed to improve the DBH estimation.
where y is the dependent variable,
,
, ……
are different independent variables, and
,
are the coefficients corresponding to different independent variables.
The model validation is divided into two stages: internal validation is based on the handheld LiDAR data used for modeling, which is used to assess the model’s goodness-of-fit and stability; external validation uses an independent ground-truth chest diameter dataset to test the model’s generalization ability on unknown samples. Two indicators, namely the coefficient of determination (R2) and the root–mean–square error (RMSE), were selected for the accuracy evaluation to measure the prediction performance of the model.
By comparing the accuracy differences between different model structures and variable combinations, we focused on analyzing the degree of the contribution of tree height and crown width in the estimation of diameter at breast height (DBH), and explored the consistency of model performance between the training sample area and the extrapolation conditions. In addition, for samples with large prediction errors, the structural features of the point cloud were analyzed retrospectively to explore the potential causes of underestimation or overestimation (e.g., missing point cloud, branch and leaf shading, crown overlap), which provided the basis for improving the algorithms of individual tree segmentation and trait extraction.
2.6. Prediction of Leaf Nitrogen Content Within Individual Tree Canopies
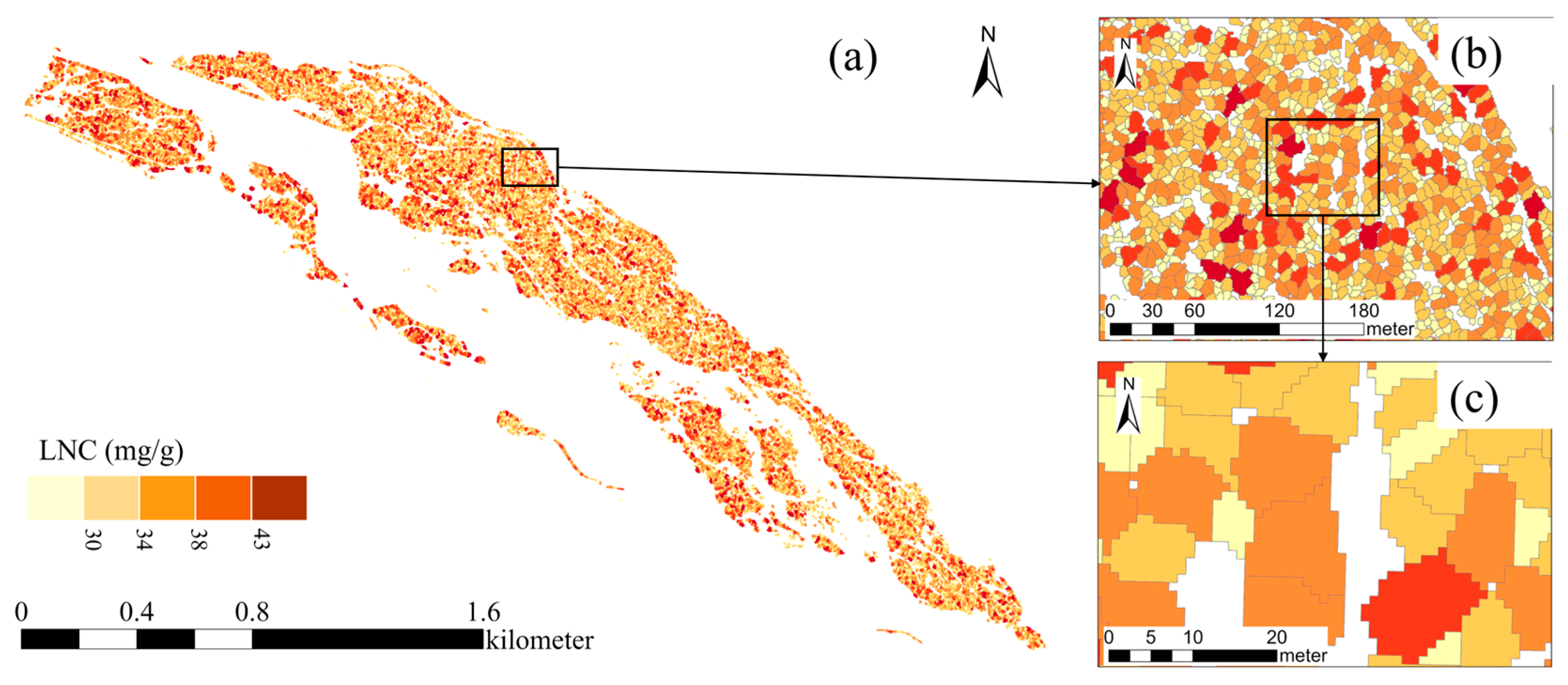
The accurate estimation of leaf biochemical traits, including nitrogen content, from canopy reflectance faces a fundamental constraint: the mixed-pixel problem. In this challenge, the spectral signal from a target tree crown is contaminated by contributions from surrounding vegetation, underlying soil, and adjacent canopy elements. To address this issue, we employed precise individual tree crown delineation, derived from our LiDAR-based segmentation workflow, to define spectrally pure regions of interest within the co-registered multispectral imagery. This method ensures that the reflectance data input into the N-PROSAIL inversion model primarily originate from the sunlit canopy of each target arborescent seabuckthorn individual. By isolating the spectral signal most directly associated with foliar properties, this approach minimizes contamination and enhances the physiological relevance of the retrieved trait estimates.
In this study, the N-PROSAIL model, a coupled model integrating the leaf optical model PROSPECT-PRO and the canopy radiative transfer model 4SAIL, was employed to perform forward simulations for constructing a synthetic dataset for leaf nitrogen content prediction in seabuckthorn forests. The N-PROSAIL model simulates directional canopy reflectance spectra based on input vegetation biochemical and biophysical traits, thereby establishing a physically meaningful mapping between traits and spectra. This process provides a large volume of physically interpretable training samples for machine learning models. The workflow for mapping leaf nitrogen content (LNC) at the individual plant level using the N-PROSAIL model combined with the Random Forest (RF) algorithm is illustrated in
Figure 3.
The dataset was generated through the following steps. First, key input traits affecting canopy reflectance and their plausible value ranges were identified. These include leaf biochemical traits, such as nitrogen content (N), chlorophyll content (Cab, μg/cm2), carotenoid content (Ccar, μg/cm2), equivalent water thickness (Cw, cm), and dry matter content (Cm, g/cm2), as well as canopy structural traits, such as leaf area index (LAI), average leaf inclination angle (ALIA), soil brightness coefficient (psoil), hotspot trait (hspot), and viewing geometry (solar zenith angle, view zenith angle, and relative azimuth angle).
The ecological representativeness of the look-up table (LUT) was ensured by defining prior parameter ranges from three sources, including field measurements from this study, the species-specific literature, and established model defaults (see
Table 1 for a complete summary). Parameter ranges were determined as follows.
Leaf nitrogen content was parameterized directly using the values obtained from the chemical analysis of 37 sampled trees (
Section 2.1). The leaf area index range was estimated by integrating hemispherical photography from sample plots with canopy gap fractions derived from the LiDAR point cloud [
61]. Ranges for key leaf biochemical constituents, specifically chlorophyll and carotenoid content, were synthesized from published studies on sea buckthorn leaf’s optical properties [
59,
60]. The soil brightness coefficient was bounded between 0 and 1 to reflect the observed spectral variation in bare soil within the study area. Finally, all observation geometry parameters were configured to match the specific solar zenith, view zenith, and relative azimuth angles recorded during the UAV multispectral data acquisition.
Subsequently, a global sensitivity analysis strategy based on Latin Hypercube Sampling (LHS) was adopted to efficiently and uniformly sample the multidimensional trait space. The LHS method ensures broad coverage of the trait space with a limited number of samples, avoiding the clustering effects common in simple random sampling. This approach ensures that the simulated dataset adequately captures spectral responses under diverse trait combinations. Each sampled trait set was input into the N-PROSAIL model to generate corresponding hyperspectral reflectance data covering the 400–2500 nm spectral range at approximately 1 nm resolution.
To align the simulated data with field-acquired multispectral imagery, the high-resolution reflectance spectra were convolved into four broad spectral bands matching the measured multispectral sensor. The spectral response function (SRF) of each band of the target sensor was obtained, which quantifies the relative sensitivity of the sensor to different wavelengths. The equivalent reflectance for band
i was calculated from the hyperspectral reflectance curve
R(
λ) using the following equation:
This convolution process yielded a simulated dataset with the same spectral characteristics as the field multispectral data. The final product is a large lookup table (LUT), where each row represents a sample consisting of a trait set and the corresponding simulated canopy reflectance. This dataset not only exceeds the sample size achievable through field surveys but also encompasses a wide range of trait combinations that are difficult to observe in practice.
The monitoring of vegetation traits via optical remote sensing inherently relies on modeling, whether through statistical or physical approaches. While empirical statistical models are commonly used for nitrogen content retrieval, and physical models often infer nitrogen indirectly via related traits, such as chlorophyll, the direct prediction of nitrogen content remains challenging. In response, this study employed random forest machine learning-based nonlinear regression methods to estimate leaf nitrogen content in seabuckthorn forests.
Random forest (RF) [
62] is a kind of tree structure integrated learning method that can be used for regression and classification. It was developed by the University of California, Berkeley Department of Statistics, by Professor Breiman, in 2001 [
62]; its essence is based on the CART (classification and regression tree) decision tree integrated learning algorithm [
63]. Using the bagging method from the training set of the self-help sampling method to generate a number of different sub-training sample sets, each sample is used to build a decision tree model, constituting a multi-classification model system, and finally, all the decision trees show the most voted results as the final prediction. The random forest algorithm demonstrates high predictive accuracy, strong resistance to noise, a low tendency to overfit, and excellent adaptability to diverse datasets. Due to its simple structure and excellent performance compared to other machine learning methods, it has been widely applied in various fields. In using the random forest regression method, n_estimators and max_features traits are two important traits to adjust. n_estimators defines the number of random trees. Increasing the number of trees generally leads to more reliable results, but also increases the computational cost.
The random forest regression model was configured with three vegetation indices as input features: the near-infrared band, the normalized blue-edge difference index (
nNDVIblue), and the MERIS terrestrial chlorophyll index (
MTCI).
These three indices were extracted from the multispectral imagery for each segmented individual tree crown. For the field-measured dataset, they were calculated directly from the UAV-acquired spectra. For the N-PROSAIL-simulated dataset, they were derived from the convolved broad-band reflectances to ensure spectral consistency. This common set of features allows for the direct comparison and integration of field and simulated data during model training.
4. Discussion
The primary contribution of this study resides in the design and validation of an integrated analytical workflow for resolving a persistent challenge in remote sensing of natural forests, namely the extraction of individual tree-level traits in monodominant stands where conspecifics exhibit a growth-form continuum from shrubs to trees. The key methodological advancement is the introduction of a dedicated tree–shrub separation step, which directly addresses the structural interference that has fundamentally limited previous approaches. Our empirical results demonstrate that this step is not merely beneficial but essential. When applied to raw point clouds, conventional segmentation algorithms performed poorly, with F1-scores as low as 33.09 percent. Following the application of our separation module, all methods showed a marked improvement, culminating in an optimal F1-score of 84.21 percent and the complete elimination of over-segmentation for hierarchical clustering. This sequence confirms that the synergistic integration of separation and segmentation, not the performance of any single algorithm, enables reliable individual tree mapping in such complex environments. Furthermore, by structurally coupling LiDAR-derived architecture with multispectral-derived biochemistry via radiative transfer modeling, the workflow delivers a comprehensive phenotypic portrait at the individual tree scale. This capacity directly supports downstream applications such as precision phenotyping for germplasm selection. Ultimately, the primary contribution of this work lies in integrating established components into a coherent workflow that addresses a specific ecological challenge, one that has proven difficult to tackle with conventional methods.
This study proposes a hierarchical and logically rigorous technical workflow for separating trees and shrubs from mixed seabuckthorn point clouds. Its core strength lies in a multi-step, progressive processing approach that effectively discriminates between the two vegetation types. The efficacy of the method stems mainly from the following key designs. First, the parallel multi-scale extraction of candidate feature points represents a major innovation. By detecting tree crown candidate points simultaneously at three scales (3 m, 5 m, and 7 m), the method adapts effectively to structural heterogeneity among stands. Given the considerable variation in crown size among species, a single scale would be insufficient to capture all treetops completely. The multi-scale strategy enables the synchronous detection of both pioneer species with smaller crowns and dominant species with larger crowns, significantly improving the completeness of crown detection and thereby supplying richer feature information for accurate tree–shrub separation. Second, the OPTICS clustering algorithm, incorporating both height and density constraints, is crucial for achieving fine classification. As a density-based clustering method, OPTICS performs well in identifying arbitrarily shaped clusters and handling non-uniformly distributed data, making it highly suitable for natural scenarios where sparse shrubs coexist with dense tree crowns. However, under complex conditions, relying solely on density may be inadequate for clearly distinguishing shrubs and low trees with significant vertical overlap. Our method introduces a height constraint, integrating the vertical vegetation structure into the clustering process. This allows low-lying, dense point clouds to be more reliably identified as shrubs, and taller, dense point clusters as trees. The dual “density + height” criterion considerably enhances the robustness and accuracy of classification.
The structural complexity inherent in natural seabuckthorn forests necessitated the two-stage methodology presented in this study. Unlike monocultural or plantation stands, these ecosystems are characterized by a continuous gradient of growth forms, where trees and shrubs are not merely adjacent but structurally interwoven. This heterogeneity presents a fundamental challenge for standard individual tree detection algorithms, which typically assume a distinct separation between target trees and the understory. Our findings indicate that applying such algorithms directly to the raw point cloud leads to systematic errors: under-segmentation due to the absorption of shrub points into tree crowns and over-segmentation arising from the complex internal architecture of large crowns. Consequently, the initial separation of tree points emerges not as an optional preprocessing step, but as a critical prerequisite for accurate delineation in structurally complex stands.
The efficacy of our approach lies in the complementary functions of its two stages. The first stage, tree-shrub separation via multi-scale spatial analysis and adaptive clustering, serves as a contextual filter. It explicitly addresses the primary source of commission error by removing non-tree vegetation, thereby transforming a mixed point cloud into a purified set of candidate tree points. The second stage, performing fine-scale individual crown segmentation, then operates on this optimized dataset. This sequential design allows each stage to specialize: the separation step mitigates noise, and the segmentation step maximizes fidelity in boundary detection. The quantitative improvements reported in
Table 2, notably the rise in F1-score for hierarchical clustering from 68.00% to 84.21% and the elimination of over-segmentation, directly validate this synergistic relationship. The separation stage resolves the coarse-scale confusion between growth forms, enabling the segmentation stage to achieve precision at the individual tree level.
This methodological framework carries significant implications for the remote sensing of structurally heterogeneous forests. It demonstrates that in ecosystems where biological form and spatial arrangement introduce significant noise, a single-algorithm solution is often insufficient. A hierarchical processing strategy, which decouples the problem of “what is a tree” from “where are its boundaries,” proves to be more robust. Future work could explore automating the adaptive thresholds used in the separation step or testing this two-stage paradigm on other forest types with similarly intermixed vegetation. Ultimately, this study underscores that advancing individual tree mapping in complex natural environments may depend less on refining a single algorithm and more on strategically orchestrating multiple analytical steps to mirror the ecological complexity on the ground.
A limitation of this study, however, is the relatively small amount of ground validation data. As a result, the evaluation relied primarily on visual interpretation and the manual labeling of sample area point clouds. Although this approach allows for a preliminary assessment of the reasonableness of the classification results, it has certain drawbacks. On the one hand, visual interpretation outcomes are susceptible to the interpreter’s experience and subjectivity, lacking a unified and objective quantitative evaluation standard. On the other hand, in the absence of high-precision ground truth data, such as precisely georeferenced individual tree survey data, it is difficult to perform a rigorous quantitative assessment of classification accuracy using metrics like overall accuracy, recall, and precision. In subsequent research, incorporating more sample plots and more detailed field survey data would improve the reliability and persuasiveness of the method validation, while also furnishing a more objective basis for parameter optimization.
The segmentation results demonstrate that the pre-processing step of separating trees and shrubs significantly enhances the accuracy of individual tree detection within seabuckthorn stands. This underscores the critical importance of distinguishing between growth forms prior to segmentation, as it effectively reduces commission errors and over-segmentation. However, it is important to note that the scope of this study is limited to the segmentation of trees; the shrub component, once separated, was not subjected to individual plant segmentation. Consequently, the performance of the proposed method for shrub identification remains uncertain. Future work should therefore include a dedicated evaluation of shrub segmentation to comprehensively assess its applicability across entire plant communities.
The comparative analysis confirms hierarchical clustering as the most effective segmentation method for our seabuckthorn forest data, achieving an optimal balance between detection accuracy and segmentation integrity (F1-score = 84.21%; over-segmentation rate = 0.00%). This performance advantage stems from a fundamental alignment between the algorithm’s design and the ecosystem’s structural complexity. Unlike DBSCAN, which is constrained by fixed density thresholds and thus prone to errors amid variable crown densities, or the marker-controlled watershed approach, whose over-segmentation (OR = 78.07%) reveals a critical sensitivity to local maxima in the CHM, hierarchical clustering operates through a flexible, proximity-based nested grouping. This method intrinsically accommodates key forest attributes: the growth-form continuum from shrub to tree, multi-axis crown architectures, variable within-crown point densities, and naturally clumped spatial distributions. Consequently, the selection of hierarchical clustering is justified not only by its quantitative superiority, but by its conceptual congruence with the ecological and structural reality of seabuckthorn stands, especially when deployed following the essential preprocessing step of tree–shrub separation, which removes understory interference and allows the algorithm to focus on genuine crown-scale topology.
To validate tree height accuracy, we stratified prediction errors using tertiles instead of the median or quartiles. This approach was chosen to create groups of markedly different sizes, which amplifies diagnostic contrast within an error distribution heavily skewed toward minor inaccuracies. The tertile-based stratification revealed a clear and interpretable pattern. The high-error group, which corresponds to the top tertile, consisted primarily of trees located in high-density canopy areas with small crowns. These conditions are known to challenge LiDAR-based height retrieval due to signal occlusion and limited crown definition. In contrast, the larger low-error group, comprising the bottom two tertiles, was dominated by open-grown trees with well-defined crowns, where the model performed reliably. Therefore, the tertile split served as a purposeful analytical tool rather than a mere statistical convention. It effectively separated the majority of accurately predicted trees from a distinct minority of problematic cases, with the latter directly linked to specific and ecologically meaningful forest structural conditions. Employing the median would have obscured this pattern by combining moderately accurate samples from dense areas with genuine algorithmic outliers. This method thereby enhances the diagnostic value of error analysis by cleanly isolating systematic failure modes for focused methodological investigation.
In constructing the DBH prediction model for seabuckthorn, tree height emerged as the most influential predictor, while the inclusion of crown width did not significantly enhance model accuracy. This could be attributed to the limited variability in crown size among seabuckthorn individuals or a weak inherent correlation between crown width and DBH. While the power function model yielded a lower root–mean–square error (RMSE) in certain cases, its coefficient of determination (R2) did not improve markedly, potentially due to sample distribution characteristics or the model’s sensitivity to extreme values. Future studies could incorporate additional structural attributes, such as the height to the first live branch or the crown volume, to further strengthen the robustness of DBH estimation. Moreover, the high consistency between validation results based on field-measured data and those derived from handheld device extraction confirms the feasibility of applying this method at a regional scale, though its stability across larger sample areas requires further verification.
The finding that incorporating crown width did not significantly improve DBH estimation accuracy can be attributed to three interrelated factors rooted in the ecology and the remote sensing system. First, arborescent seabuckthorn in the high-altitude semi-arid study area exhibits a limited phenotypic range in terms of crown architecture. A conservative growth strategy prioritizing vertical and belowground investment over lateral crown expansion under harsh conditions naturally constrains crown width variation, thereby reducing its statistical utility as a predictive variable. Second, the allometric relationship between DBH and crown width is highly asymmetric and variable. For individuals of comparable DBH, crown width can differ substantially due to localized competition, microtopographic effects, and inherent genetic variation. This variability weakens the bivariate correlation and diminishes the marginal explanatory power of crown width beyond what is already captured by tree height. Third, the remote measurement of crown width introduces inherent uncertainty. While LiDAR-derived widths were validated against field data, delineations of the irregular and often asymmetric crowns of natural stands remain approximations. Representing such complex three-dimensional structures with a single planar metric inevitably oversimplifies the true functional allometry. This oversimplification, stemming from deriving a single diameter value from the major axis of a minimum bounding rectangle, further attenuates the predictive contribution of the crown width variable. Collectively, these ecological and methodological factors explain why crown width did not enhance model performance in this context.
Our methodology reveals a synergistic interdependence between structural segmentation and biochemical estimation. Reliable crown delineation serves not merely as a preprocessing step but as a critical prerequisite for accurate leaf nitrogen content retrieval. By ensuring that each spectral sample corresponds to a correctly isolated, single-tree crown, we substantially mitigate spectral contamination from mixed pixels, shadowed canopy components, and adjacent vegetation. This refinement of the input signal suppresses noise within the spectral–biochemical relationship, producing a prediction model that is both more accurate and more physically interpretable. Consequently, structural segmentation precision directly enhances biochemical credibility, integrating the two analytical streams into a coherent phenotyping framework whose combined value exceeds that of either component in isolation.
It is important to clarify the specific role of the N-PROSAIL model within our workflow to preempt concerns regarding undue complexity. The model was employed exclusively during the development phase of this study. Its primary function was not to establish a full, operational inversion pipeline, but rather to serve as a computational bridge, generating a synthetic training dataset to overcome the severe constraint of limited ground truth measurements. The final, practical output of this process is a calibrated Random Forest model, denoted RF_DA, which performs inference using only a small set of common vegetation indices. This design strategically encapsulates the underlying complexity offline, ultimately delivering a straightforward, transparent, and directly applicable tool for potential end-users. In our research, we employed traditional vegetation indices for regression prediction of LNC, but the results were unsatisfactory (specific details are provided in
Table A1). For future application in similar seabuckthorn stands, the requirement is simply to compute three standard spectral indices from the available UAV imagery, thereby eliminating any need for complex radiative transfer modeling or parameterization. Although the LNC prediction models, calibrated on field data and enriched with N-PROSAIL simulations, showed strong agreement with independent spectral measurements, the sampling approach presents a notable constraint. The collection of merely one sample per tree fails to represent potential intra-canopy nitrogen variation. Consequently, future work should prioritize stratified canopy samplings to account for vertical heterogeneity, thereby enhancing the granularity and reliability of nitrogen content estimates.
This study introduces a scalable technical workflow predicated on a critical preprocessing step: intraspecific growth-form separation, followed by individual tree segmentation and trait extraction. The framework is explicitly designed to overcome a persistent challenge in shrubland remote sensing, namely the difficulty in reliably delineating distinct woody individuals from a continuous and morphologically diverse vegetation matrix. Consequently, the approach shows considerable promise for application across other arid and semi-arid ecosystems dominated by monodominant woody species that exhibit a pronounced growth-form continuum, such as Tamarix spp., Caragana spp., or certain Artemisia species. The core methodology is inherently transferable, provided that the target species exhibits a polymorphic mixture of shrub-like and tree-like architectures within the same stand.
Successful operationalization in new contexts, however, will necessitate deliberate adaptation. First, the multi-scale spatial analysis and dynamic thresholding parameters central to the growth-form separation step must be recalibrated to reflect the characteristic crown dimensions and point cloud densities of the novel target species. Second, data acquisition parameters, particularly LiDAR point density and the selection of multispectral bands, should be optimized to capture the specific structural and physiological traits of interest. Third, while the N-PROSAIL model can be re-parameterized for different species, its efficacy as a data-augmentation tool hinges on the availability of reliable, species-specific leaf optical property libraries. In regions where such foundational biophysical data are scarce, its initial implementation might adopt a more pragmatic, empirical approach utilizing established vegetation indices as a precursor to full radiative transfer modeling.
In essence, the proposed workflow provides a generalizable conceptual architecture for individual-level mapping in complex woody communities. Its practical implementation, however, transforms it from a fixed protocol into an adaptable framework, whose underlying algorithms require careful tuning to the local species characteristics and data conditions. Future research should prioritize validating this framework across diverse biogeographic regions to develop robust, empirically grounded guidelines for parameter adaptation, thereby enhancing its utility for large-scale ecological monitoring.