Deep Learning in Forestry Using UAV-Acquired RGB Data: A Practical Review
Abstract
:1. Introduction
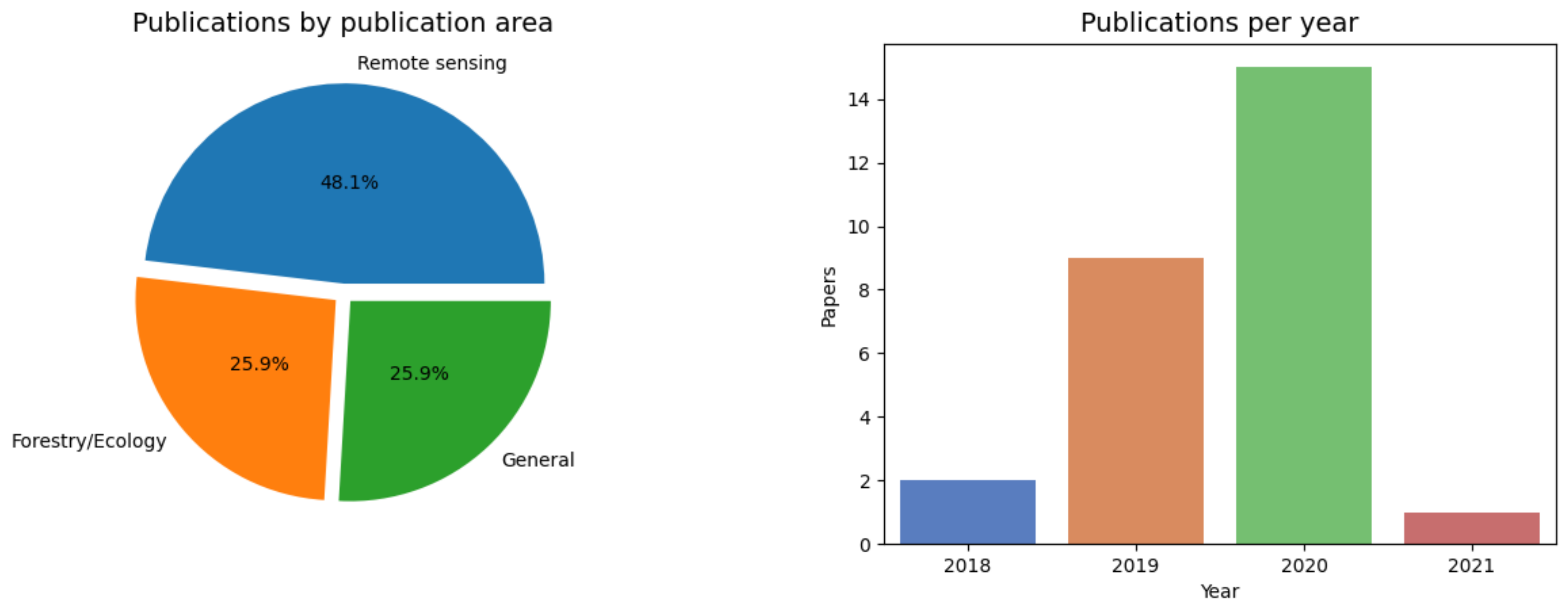
2. Review Methodology
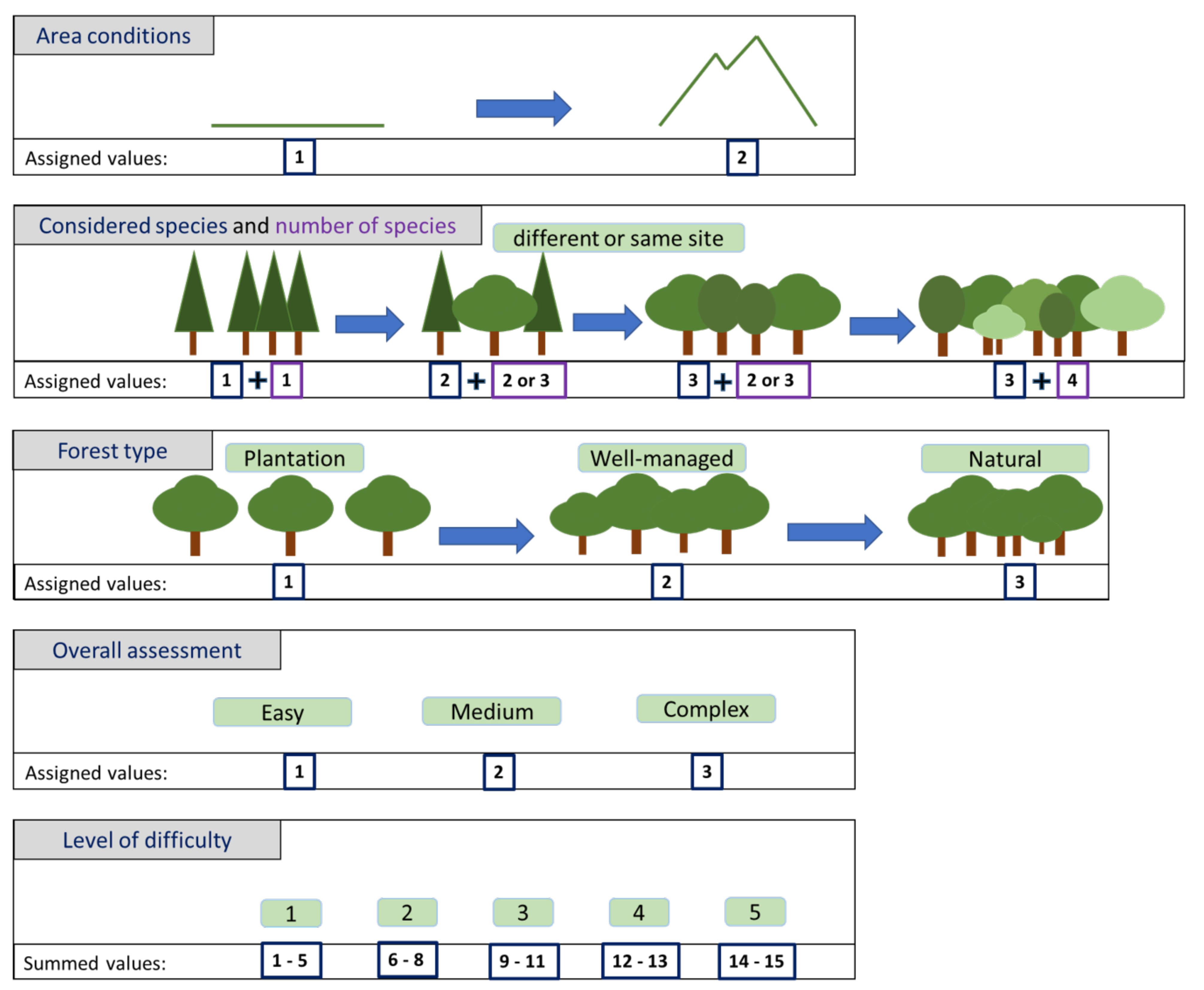
Evaluation of the Difficulty Level of Each Forestry Problem
3. Deep Learning for Image Analysis
3.1. Main Dataset Split Issues
- The training set is used to update the network weights during the training process. Therefore, these images are passed through the network to obtain an error measure using a “loss” function whose global minimum relates to the desired goal (for example overlap between the prediction and the real pixel-wise labels). Afterwards, the derivative of this loss function with respect to all the weights is computed, and the weights are updated according to this gradient. This process is commonly known as back-propagation.
- The validation set is generally used to tune specific hyperparameters of the network. These hyperparameters include the number of layers (depth), the number of weights per layer, the loss function, etc. However, in contrast with the training set, back-propagation is not used on these samples. Commonly, the results in the validation set are used to select a final model (the one that achieved the best performance), to stop the training process if a plateau is achieved (usually called early stopping) and, thus, to avoid over-fitting. Therefore, while the validation samples have no direct effect on the weights of the network, they have an impact on the final model and consequently should be treated as part of the training process.
- The testing set does not take part in the training process. Once the best model is trained, the testing samples are passed through the network to obtain a prediction that is then evaluated. The objective of these samples is to test the ability of the model to generalize to cases unseen during the training process.
- Wrong data split. Not splitting the dataset at the subject-level when defining the training, validation and testing sets can result in data from the same subject to appear in several sets. For example, having images of the same tree on different days split into the different sets might lead to a biased prediction (hereafter “Wrong Split” or WS).
- Late split. Procedures such as data augmentation or feature selection should always be performed after the split. If all the images are processed together, some information from the testing samples might be shared and potentially be involved in the training process. For example, if data augmentation is performed before isolating the test data from the training and validation data, augmented samples generated from the same unique image may be found in both sets, leading to a problem similar to having a wrong data split (hereafter “Late Split” or LS).
- Validation set for testing. The test set should only be used to evaluate the final performance of the models, not to choose the training hyperparameters (e.g., learning rate) of the model. A separate validation set must be used beforehand for hyperparameter optimization. We refer to studies using only one set for validation and testing into the category called “validation set for testing” or VST.
- Dependent test set. In our case, we only considered the test set to be properly independent if the surveyed region was physically separated. For example, we will consider acquiring data in several sites and designating one or more of them for testing acceptable. On the other hand, if the data from all the acquisition sites is randomly sampled to create the training/validation and testing sets (even if no physical overlap exists), we will consider the testing set not to be independent. By not separating the physical sites, images in the training and testing sets belonging to trees that are physically very close might have very similar characteristics and identical lighting conditions. Consequently, models trained under these conditions might not generalize well to other sites, even if the species are the same (hereafter “dependent test set” or DTS).
3.2. Computer Vision Paradigms and Deep Learning Architectures
3.2.1. Image Classification
3.2.2. Object Detection
3.2.3. Semantic Segmentation
3.2.4. Instance Segmentation and Panoptic Segmentation
3.3. Data Augmentation and Transfer Learning
- Small central rotations with a random angle. Depending on the orientation of the UAV, different orthomosaics acquired during different time frames might show different perspectives of the same trees. In order to introduce invariance to these differences, small rotations of the two main image axes can be applied to artificially increase the number of samples.
- Flips on the X and Y axes (up/down and left/right). Another way of addressing these differences is to mirror the image on their main axes (up/down, left/right).
- Gaussian blurring of the images. Due to the acquisition (movement, sensor characteristics, distance, etc.) and mosaicking process, some regions of the image might also present some blurring. A Gaussian kernel can be used to artificially expand the training dataset, simulate this blurring effect and improve generalization.
- Linear and small contrast changes. Similarly, different lightning conditions or shadows between regions of the image might also affect the results. By introducing contrast changes in the training set, these effects can be simulated to enlarge the number of training samples.
- Localized elastic deformation. Finally, elastic deformations can be applied to simulate possible different intra-species shapes.
3.4. Data Pre-Processing for UAV-Acquired Forest Images
4. Individual Tree Detection
- To study the characteristics of one tree or a small number of trees that are used for scaling up to whole forests where all trees are assumed to be similar [68].
- Large scale studies, where low resolution data encompassing whole forests is collected and the information from individual trees is inferred [69].
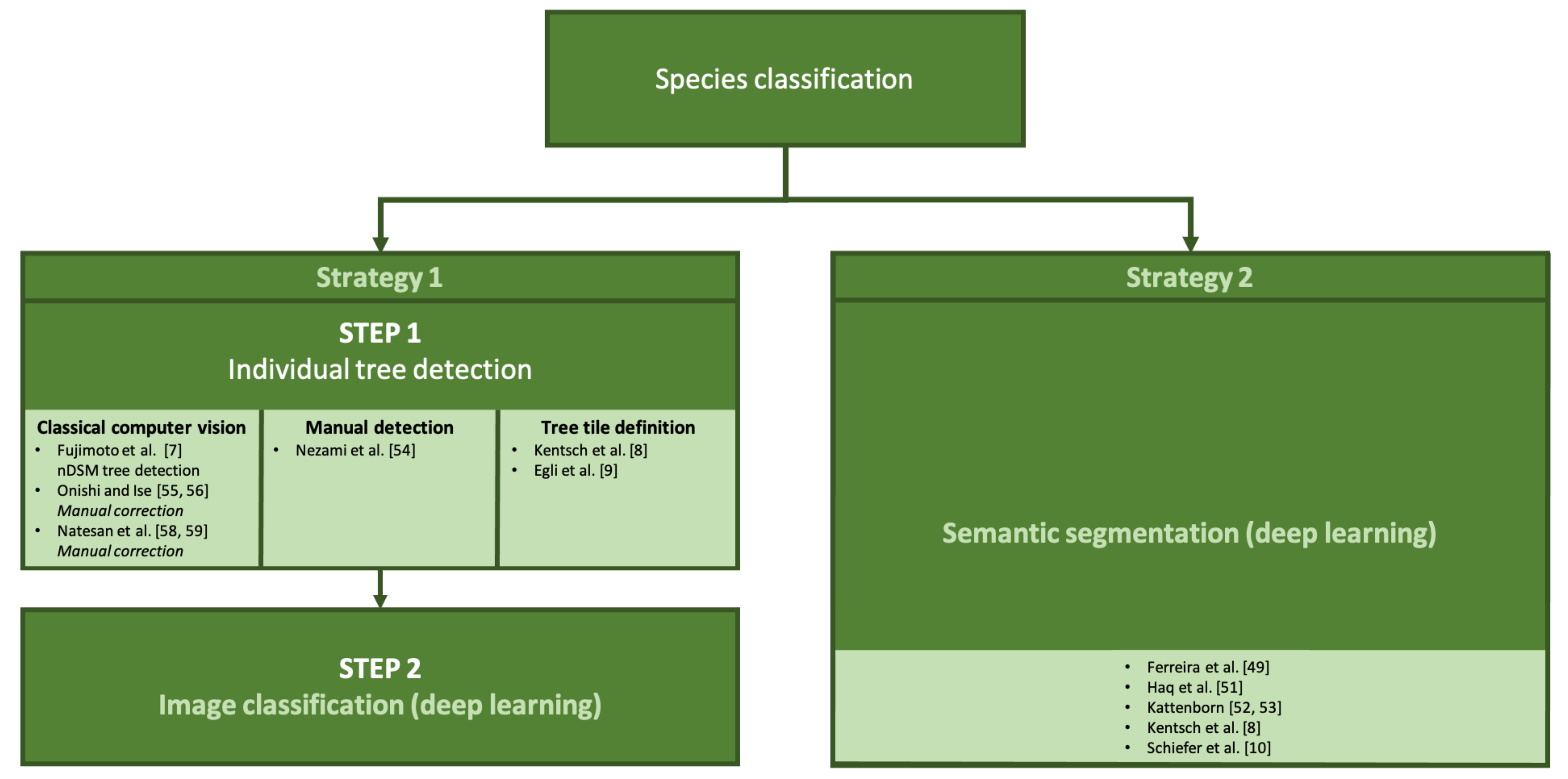
5. Tree Species Classification
5.1. Tree Detection + Image Classification for Species Classification
5.2. Semantic Segmentation for Tree Species Classification
6. Forest Anomaly Detection
6.1. Forest Fires
6.2. Tree Health Determination
7. Discussion
7.1. Individual Tree Detection
7.2. Tree Species Classification
7.3. Forest Anomaly Detection
7.4. Practical Aspects—Getting Started in the DL Analysis of UAV-Acquired Forest RGB Images
7.4.1. Available Datasets
Data Downloadable from the Web
- The dataset used in [8] for patch-wise segmentation and classification is available at https://zenodo.org/record/3693326#.YC8q2BGRXAI (accessed on 21 July 2021). The data includes orthomosaics and manual annotations (in the form of binary masks) for 7 winter mosaics and for 2 orthomosaics corresponding to the study of an invasive tree species growing in a pine forest taken in the summer season.
- The dataset used in [63] for the detection of fir trees affected by a bark beetle parasite is available for download at: https://zenodo.org/record/4054338#.YIuz1BGRXCI (accessed on 21 July 2021). The data includes 9 orthomosaics with manual annotations for the three classes considered as a binary mask that delineates the location of all the trees in the mosaics.
- The patches used to segment the Mauritia flexuosa palm tree from background used in [50] are available at http://didt.inictel-uni.edu.pe/dataset/MauFlex_Dataset.rar (accessed on 21 July 2021). Although the patches present an LS data problem mentioned in Section 5.2, this issue can potentially be fixed by automatically selecting every fourth image. The dataset contains RGB images and binary masks corresponding to the palm region of each image.
- The patches used in [47] to classify into cactus and non-cactus are also available for download at: https://www.kaggle.com/irvingvasquez/cactus-aerial-photos (accessed on 21 July 2021). The dataset is composed of a training and a validation folder each of them subdivided into cactus and non-cactus subfolders. Unfortunately, the video from which the patches were derived is not available, so addressing the data issues mentioned in Section 4 is not possible.
Data Available on Demand
- The authors of [56] shared a dataset with us containing images gathered in summer and autumn. Within each of these two folders, the images were divided into training, validation and testing. Each segmented tree canopy was stored as a separate image (with the non-canopy part left as background) with subfolders grouping each of the studied classes. An important detail is that these classes present an imbalance problem already detailed in Section 5.1: a majority of the images belong to the non-tree (“others”) class. Furthermore, the orthomosaics for each season were also provided with their corresponding prediction map as a shapefile. In the version that we had access to, no major description was provided for the dataset but it was relatively easy to understand from the description in the paper.
- The authors in [52,53] also readily provided data upon request: The shared dataset comprised the images of the Ulex europaeus and Pinus radiata species. Each folder contained the orthomosaics and DEMs for the different flights. Additionally, shapefiles with annotations of the target species and the AOI (area of interest) for each orthomosaic were also available.
7.4.2. Software Resources
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Paneque-Gálvez, J.; McCall, M.K.; Napoletano, B.M.; Wich, S.A.; Koh, L.P. Small Drones for Community-Based Forest Monitoring: An Assessment of Their Feasibility and Potential in Tropical Areas. Forests 2014, 5, 1481–1507. [Google Scholar] [CrossRef] [Green Version]
- Gambella, F.; Sistu, L.; Piccirilli, D.; Corposanto, S.; Caria, M.; Arcangeletti, E.; Proto, A.R.; Chessa, G.; Pazzona, A. Forest and UAV: A bibliometric review. Contemp. Eng. Sci. 2016, 9, 1359–1370. [Google Scholar] [CrossRef]
- Guimarães, N.; Pádua, L.; Marques, P.; Silva, N.; Peres, E.; Sousa, J.J. Forestry Remote Sensing from Unmanned Aerial Vehicles: A Review Focusing on the Data, Processing and Potentialities. Remote Sens. 2020, 12, 1046. [Google Scholar] [CrossRef] [Green Version]
- Banu, T.P.; Borlea, G.F.; Banu, C.M. The Use of Drones in Forestry. J. Environ. Sci. Eng. 2016, 5, 557–562. [Google Scholar]
- Chadwick, A.J.; Goodbody, T.R.H.; Coops, N.C.; Hervieux, A.; Bater, C.W.; Martens, L.A.; White, B.; Röeser, D. Automatic Delineation and Height Measurement of Regenerating Conifer Crowns under Leaf-Off Conditions Using UAV Imagery. Remote Sens. 2020, 12, 4104. [Google Scholar] [CrossRef]
- Ocer, N.E.; Kaplan, G.; Erdem, F.; Matci, D.K.; Avdan, U. Tree extraction from multi-scale UAV images using Mask R-CNN with FPN. Remote Sens. Lett. 2020, 11, 847–856. [Google Scholar] [CrossRef]
- Fujimoto, A.; Haga, C.; Matsui, T.; Machimura, T.; Hayashi, K.; Sugita, S.; Takagi, H. An End to End Process Development for UAV-SfM Based Forest Monitoring: Individual Tree Detection, Species Classification and Carbon Dynamics Simulation. Forests 2019, 10, 680. [Google Scholar] [CrossRef] [Green Version]
- Kentsch, S.; Lopez Caceres, M.L.; Serrano, D.; Roure, F.; Diez, Y. Computer Vision and Deep Learning Techniques for the Analysis of Drone-Acquired Forest Images, a Transfer Learning Study. Remote Sens. 2020, 12, 1287. [Google Scholar] [CrossRef] [Green Version]
- Egli, S.; Höpke, M. CNN-Based Tree Species Classification Using High Resolution RGB Image Data from Automated UAV Observations. Remote Sens. 2020, 12, 3892. [Google Scholar] [CrossRef]
- Schiefer, F.; Kattenborn, T.; Frick, A.; Frey, J.; Schall, P.; Koch, B.; Schmidtlein, S. Mapping forest tree species in high resolution UAV-based RGB-imagery by means of convolutional neural networks. ISPRS J. Photogramm. Remote Sens. 2020, 170, 205–215. [Google Scholar] [CrossRef]
- Tran, D.Q.; Park, M.; Jung, D.; Park, S. Damage-Map Estimation Using UAV Images and Deep Learning Algorithms for Disaster Management System. Remote Sens. 2020, 12, 4169. [Google Scholar] [CrossRef]
- Safonova, A.; Tabik, S.; Alcaraz-Segura, D.; Rubtsov, A.; Maglinets, Y.; Herrera, F. Detection of fir trees (Abies sibirica) damaged by the bark beetle in unmanned aerial vehicle images with deep learning. Remote Sens. 2019, 11, 643. [Google Scholar] [CrossRef] [Green Version]
- Balsi, M.; Esposito, S.; Fallavollita, P.; Nardinocchi, C. Single-tree detection in high-density LiDAR data from UAV-based survey. Eur. J. Remote Sens. 2018, 51, 679–692. [Google Scholar] [CrossRef] [Green Version]
- Qin, J.; Wang, B.; Wu, Y.; Lu, Q.; Zhu, H. Identifying Pine Wood Nematode Disease Using UAV Images and Deep Learning Algorithms. Remote Sens. 2021, 13, 162. [Google Scholar] [CrossRef]
- Kattenborn, T.; Leitloff, J.; Schiefer, F.; Hinz, S. Review on Convolutional Neural Networks (CNN) in vegetation remote sensing. ISPRS J. Photogramm. Remote Sens. 2021, 173, 24–49. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. In Proceedings of the 2015 IEEE International Conference on Computer Vision (ICCV), Santiago, Chile, 7–13 December 2015; pp. 1026–1034. [Google Scholar] [CrossRef] [Green Version]
- Wen, J.; Thibeau-Sutre, E.; Diaz-Melo, M.; Samper-González, J.; Routier, A.; Bottani, S.; Dormont, D.; Durrleman, S.; Burgos, N.; Colliot, O. Convolutional neural networks for classification of Alzheimer’s disease: Overview and reproducible evaluation. Med. Image Anal. 2020, 63, 101694. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Weng, Q. A survey of image classification methods and techniques for improving classification performance. Int. J. Remote Sens. 2007, 28, 823–870. [Google Scholar] [CrossRef]
- Everingham, M.; Van Gool, L.; Williams, C.K.; Winn, J.; Zisserman, A. The PASCALVisual Object Classes (VOC) Challenge. Int. J. Comput. Vis. 2010, 88, 303–338. [Google Scholar] [CrossRef] [Green Version]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In 25th International Conference on Neural Information Processing Systems—Volume 1; Curran Associates Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. arXiv 2015, arXiv:1512.03385. [Google Scholar]
- Iandola, F.N.; Moskewicz, M.W.; Ashraf, K.; Han, S.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50 × fewer parameters and <1 MB model size. arXiv 2016, arXiv:1602.07360. [Google Scholar]
- Huang, G.; Liu, Z.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2261–2269. [Google Scholar]
- Zagoruyko, S.; Komodakis, N. Wide Residual Networks. In Proceedings of the British Machine Vision Conference (BMVC); Richard, C., Wilson, E.R.H., Smith, W.A.P., Eds.; BMVA Press: York, UK, 2016; pp. 87.1–87.12. [Google Scholar]
- Xie, S.; Girshick, R.; Dollar, P.; Tu, Z.; He, K. Aggregated Residual Transformations for Deep Neural Networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception-v4, inception-resnet and the impact of residual connections on learning. In Proceedings of the AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; Volume 31. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]
- Howard, A.; Sandler, M.; Chu, G.; Chen, L.C.; Chen, B.; Tan, M.; Wang, W.; Zhu, Y.; Pang, R.; Vasudevan, V.; et al. Searching for mobilenetv3. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Seoul, Korea, 27–28 October 2019; pp. 1314–1324. [Google Scholar]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 2016, 39, 1137–1149. [Google Scholar] [CrossRef] [Green Version]
- Redmon, J.; Divvala, S.; Girshick, R.; Farhadi, A. You Only Look Once: Unified, Real-Time Object Detection. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 26 June–1 July 2016; pp. 779–788. [Google Scholar]
- Liu, W.; Anguelov, D.; Erhan, D.; Szegedy, C.; Reed, S.; Fu, C.Y.; Berg, A.C. SSD: Single Shot MultiBox Detector. In Computer Vision—ECCV 2016; Leibe, B., Matas, J., Sebe, N., Welling, M., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 21–37. [Google Scholar]
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-Decoder with Atrous Separable Convolution for Semantic Image Segmentation. In Computer Vision—ECCV 2018; Ferrari, V., Hebert, M., Sminchisescu, C., Weiss, Y., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 833–851. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. arXiv 2015, arXiv:1505.04597. [Google Scholar]
- Kirillov, A.; He, K.; Girshick, R.; Rother, C.; Dollár, P. Panoptic segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 9404–9413. [Google Scholar]
- He, K.; Gkioxari, G.; Dollar, P.; Girshick, R. Mask R-CNN. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017. [Google Scholar]
- Jung, A.B. Imgaug. 2020. Available online: https://github.com/aleju/imgaug (accessed on 1 July 2020).
- Agisoft LLC. Agisoft Metashape, Professional Edition. Version 1.5. 2018. Available online: http://agisoft.com/ (accessed on 12 June 2021).
- QGIS Geographic Information System. Open Source Geospatial Foundation Project. 2019. Available online: http://qgis.org/ (accessed on 12 June 2021).
- ESRI. ArcGIS Desktop v10.4 Software. Available online: https://www.esri.com/ (accessed on 12 June 2021).
- Toffain, P.; Benjamin, D.; Riba, E.; Mather, S.; Fitzsimmons, S.; Gelder, F.; Bargen, D.; Cesar de Menezes, J.; Joseph, D. OpendroneMap/ODM: 1.0.1. 2020. Available online: https://github.com/OpenDroneMap/ODM (accessed on 14 April 2021).
- Drone & UAV Mapping Platform DroneDeploy. Available online: http://www.dronedeploy.com/ (accessed on 14 April 2021).
- Trimble. eCognition Developer v9.0.0 Software. Available online: https://www.trimble.com/ (accessed on 12 June 2021).
- Team, T.G. GNU Image Manipulation Program. Available online: http://gimp.org (accessed on 19 August 2019).
- RectLabel. Available online: https://rectlabel.com/ (accessed on 14 April 2021).
- LabelImg. T.GitCode. 2015. Available online: http://github.com/tzutalin/labelImg (accessed on 14 April 2021).
- López-Jiménez, E.; Vasquez-Gomez, J.I.; Sanchez-Acevedo, M.A.; Herrera-Lozada, J.C.; Uriarte-Arcia, A.V. Columnar cactus recognition in aerial images using a deep learning approach. Ecol. Inform. 2019, 52, 131–138. [Google Scholar] [CrossRef]
- Fromm, M.; Schubert, M.; Castilla, G.; Linke, J.; McDermid, G. Automated Detection of Conifer Seedlings in Drone Imagery Using Convolutional Neural Networks. Remote Sens. 2019, 11, 2585. [Google Scholar] [CrossRef] [Green Version]
- Ferreira, M.P.; de Almeida, D.R.A.; de Almeida Papa, D.; Minervino, J.B.S.; Veras, H.F.P.; Formighieri, A.; Santos, C.A.N.; Ferreira, M.A.D.; Figueiredo, E.O.; Ferreira, E.J.L. Individual tree detection and species classification of Amazonian palms using UAV images and deep learning. For. Ecol. Manag. 2020, 475, 118397. [Google Scholar] [CrossRef]
- Morales, G.; Kemper, G.; Sevillano, G.; Arteaga, D.; Ortega, I.; Telles, J. Automatic Segmentation of Mauritia flexuosa in Unmanned Aerial Vehicle (UAV) Imagery Using Deep Learning. Forests 2018, 9, 736. [Google Scholar] [CrossRef] [Green Version]
- Haq, M.; Rahaman, G.; Baral, P.; Ghosh, A. Deep Learning Based Supervised Image Classification Using UAV Images for Forest Areas Classification. J. Indian Soc. Remote Sens. 2021, 49, 601–606. [Google Scholar] [CrossRef]
- Kattenborn, T.; Eichel, J.; Fassnacht, F.E. Convolutional Neural Networks enable efficient, accurate and fine-grained segmentation of plant species and communities from high-resolution UAV imagery. Sci. Rep. 2019, 9, 17656. [Google Scholar] [CrossRef]
- Kattenborn, T.; Eichel, J.; Wiser, S.; Burrows, L.; Fassnacht, F.E.; Schmidtlein, S. Convolutional Neural Networks accurately predict cover fractions of plant species and communities in Unmanned Aerial Vehicle imagery. Remote Sens. Ecol. Conserv. 2020, 6, 472–486. [Google Scholar] [CrossRef] [Green Version]
- Nezami, S.; Khoramshahi, E.; Nevalainen, O.; Pölönen, I.; Honkavaara, E. Tree Species Classification of Drone Hyperspectral and RGB Imagery with Deep Learning Convolutional Neural Networks. Remote Sens. 2020, 12, 1070. [Google Scholar] [CrossRef] [Green Version]
- Onishi, M.; Ise, T. Automatic classification of trees using a UAV onboard camera and deep learning. arXiv 2018, arXiv:1804.10390. [Google Scholar]
- Onishi, M.; Ise, T. Explainable identification and mapping of trees using UAV RGB image and deep learning. Sci. Rep. 2021, 11, 903. [Google Scholar] [CrossRef]
- Lin, C.; Ding, Q.; Tu, W.; Huang, J.; Liu, J. Fourier Dense Network to Conduct Plant Classification Using UAV-Based Optical Images. IEEE Access 2019, 7, 17736–17749. [Google Scholar] [CrossRef]
- Natesan, S.; Armenakis, C.; Vepakomma, U. Resnet-based tree species classification using UAV images. ISPRS Int. Arch. Photogramm. Remote. Sens. Spat. Inf. Sci. 2019, XLII-2/W13, 475–481. [Google Scholar] [CrossRef] [Green Version]
- Natesan, S.; Armenakis, C.; Vepakomma, U. Individual tree species identification using Dense Convolutional Network (DenseNet) on multitemporal RGB images from UAV. J. Unmanned Veh. Syst. 2020, 8, 310–333. [Google Scholar] [CrossRef]
- Barmpoutis, P.; Kamperidou, V.; Stathaki, T. Estimation of extent of trees and biomass infestation of the suburban forest of Thessaloniki (Seich Sou) using UAV imagery and combining R-CNNs and multichannel texture analysis. In Proceedings of the Twelfth International Conference on Machine Vision (ICMV 2019), Amsterdam, The Netherlands, 16–18 November 2019; Volume 11433, p. 114333C. [Google Scholar]
- Humer, C. Early Detection of Spruce Bark Beetles Using Semantic Segmentation and Image Classification. Ph.D. Thesis, Universitat Linz, Linz, Austria, 2020. [Google Scholar]
- Deng, X.; Tong, Z.; Lan, Y.; Huang, Z. Detection and Location of Dead Trees with Pine Wilt Disease Based on Deep Learning and UAV Remote Sensing. AgriEngineering 2020, 2, 294–307. [Google Scholar] [CrossRef]
- Nguyen, H.T.; Lopez Caceres, M.L.; Moritake, K.; Kentsch, S.; Shu, H.; Diez, Y. Individual Sick Fir Tree (Abies mariesii) Identification in Insect Infested Forests by Means of UAV Images and Deep Learning. Remote Sens. 2021, 13, 260. [Google Scholar] [CrossRef]
- Kim, S.; Lee, W.; Park, Y.s.; Lee, H.W.; Lee, Y.T. Forest fire monitoring system based on aerial image. In Proceedings of the 2016 3rd International Conference on Information and Communication Technologies for Disaster Management (ICT-DM), Vienna, Austria, 13–15 December 2016; pp. 1–6. [Google Scholar]
- Hossain, F.A.; Zhang, Y.M.; Tonima, M.A. Forest fire flame and smoke detection from UAV-captured images using fire-specific color features and multi-color space local binary pattern. J. Unmanned Veh. Syst. 2020, 8, 285–309. [Google Scholar] [CrossRef]
- Zhao, Y.; Ma, J.; Li, X.; Zhang, J. Saliency Detection and Deep Learning-Based Wildfire Identification in UAV Imagery. Sensors 2018, 18, 712. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Zhang, Y.; Jing, X.; Wang, G.; Mu, L.; Yi, Y.; Liu, H.; Liu, D. UAV Image-based Forest Fire Detection Approach Using Convolutional Neural Network. In Proceedings of the 2019 14th IEEE Conference on Industrial Electronics and Applications (ICIEA), Xi’an, China, 19–21 June 2019; pp. 2118–2123. [Google Scholar] [CrossRef]
- Lopez C, M.; Saito, H.; Kobayashi, Y.; Shirota, T.; Iwahana, G.; Maximov, T.; Fukuda, M. Interannual environmental-soil thawing rate variation and its control on transpiration from Larix cajanderi, Central Yakutia, Eastern Siberia. J. Hydrol. 2007, 338, 251–260. [Google Scholar] [CrossRef] [Green Version]
- Lopez C., M.; Gerasimov, E.; Machimura, T.; Takakai, F.; Iwahana, G.; Fedorov, A.; Fukuda, M. Comparison of carbon and water vapor exchange of forest and grassland in permafrost regions, Central Yakutia, Russia. Agric. For. Meteorol. 2008, 148, 1968–1977. [Google Scholar] [CrossRef]
- Diez, Y.; Kentsch, S.; Lopez-Caceres, M.L.; Nguyen, H.T.; Serrano, D.; Roure, F. Comparison of Algorithms for Tree-top Detection in Drone Image Mosaics of Japanese Mixed Forests. In ICPRAM 2020; INSTICC, SciTePress: Setubal, Portugal, 2020. [Google Scholar]
- Lecun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef] [Green Version]
- Dai, J.; Li, Y.; He, K.; Sun, J. R-fcn: Object detection via region-based fully convolutional networks. arXiv 2016, arXiv:1605.06409. [Google Scholar]
- Lin, T.Y.; Maire, M.; Belongie, S.; Hays, J.; Perona, P.; Ramanan, D.; Dollár, P.; Zitnick, C.L. Microsoft COCO: Common Objects in Context. In Computer Vision—ECCV 2014; Fleet, D., Pajdla, T., Schiele, B., Tuytelaars, T., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 740–755. [Google Scholar]
- Lin, T.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature Pyramid Networks for Object Detection. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 936–944. [Google Scholar]
- Weinstein, B.G.; Marconi, S.; Bohlman, S.; Zare, A.; White, E. Individual Tree-Crown Detection in RGB Imagery Using Semi-Supervised Deep Learning Neural Networks. Remote Sens. 2019, 11, 1309. [Google Scholar] [CrossRef] [Green Version]
- Weinstein, B.G.; Marconi, S.; Bohlman, S.A.; Zare, A.; White, E.P. Cross-site learning in deep learning RGB tree crown detection. Ecol. Inform. 2020, 56, 101061. [Google Scholar] [CrossRef]
- Bravo-Oviedo, A.; Pretzsch, H.; Ammer, C.; Andenmatten, E.; Barbati, A.; Barreiro, S.; Brang, P.; Bravo, F.; Coll, L.; Corona, P.; et al. European mixed forests: Definition and research perspectives. For. Syst. 2014, 23, 518–533. [Google Scholar] [CrossRef]
- Huuskonen, S.; Domisch, T.; Finér, L.; Hantula, J.; Hynynen, J.; Matala, J.; Miina, J.; Neuvonen, S.; Nevalainen, S.; Niemistö, P.; et al. What is the potential for replacing monocultures with mixed-species stands to enhance ecosystem services in boreal forests in Fennoscandia? For. Ecol. Manag. 2021, 479. [Google Scholar] [CrossRef]
- Fassnacht, F.E.; Latifi, H.; Stereńczak, K.; Modzelewska, A.; Lefsky, M.; Waser, L.T.; Straub, C.; Ghosh, A. Review of studies on tree species classification from remotely sensed data. Remote Sens. Environ. 2016, 186, 64–87. [Google Scholar] [CrossRef]
- Michałowska, M.; Rapiński, J. A Review of Tree Species Classification Based on Airborne LiDAR Data and Applied Classifiers. Remote Sens. 2021, 13, 353. [Google Scholar] [CrossRef]
- Kentsch, S.; Cabezas, M.; Tomhave, L.; Groß, J.; Burkhard, B.; Lopez Caceres, M.L.; Waki, K.; Diez, Y. Analysis of UAV-Acquired Wetland Orthomosaics Using GIS, Computer Vision, Computational Topology and Deep Learning. Sensors 2021, 21, 471. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; IEEE Computer Society: Los Alamitos, CA, USA, 2016; pp. 770–778. [Google Scholar]
- McGaughey, R.J. FUSION/LDV: Software for LIDAR Data Analysis and Visualization; US Department of Agriculture, Forest Service, Pacific Northwest Research Station: Seattle, WA, USA, 2009; Volume 123.
- Diez, Y.; Kentsch, S.; Caceres, M.L.L.; Moritake, K.; Nguyen, H.T.; Serrano, D.; Roure, F. A Preliminary Study on Tree-Top Detection and Deep Learning Classification Using Drone Image Mosaics of Japanese Mixed Forests. In Pattern Recognition Applications and Methods; De Marsico, M., Sanniti di Baja, G., Fred, A., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 64–86. [Google Scholar]
- Beucher, S.; Meyer, F. The Morphological Approach to Segmentation: The Watershed Transformation. Math. Morphol. Image Process. 1993, 34, 433–481. [Google Scholar]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Lin, Z.; Desmaison, A.; Antiga, L.; Lerer, A. Automatic Differentiation in PyTorch. In Proceedings of the NIPS Autodiff Workshop, Long Beach, CA, USA, 9 December 2017. [Google Scholar]
- Cooley, J.; Tukey, J. An Algorithm for the Machine Calculation of Complex Fourier Series. Math. Comput. 1965, 19, 297–301. [Google Scholar] [CrossRef]
- Code, P.W. CIFAR10 Classification Results. 2018. Available online: https://paperswithcode.com/sota/image-classification-on-cifar-10 (accessed on 8 April 2021).
- Forzieri, G.; Girardello, M.; Ceccherini, G.; Spinoni, J.; Feyen, L.; Hartmann, H.; Beck, P.S.A.; Camps-Valls, G.; Chirici, G.; Mauri, A.; et al. Emergent vulnerability to climate-driven disturbances in European forests. Nat. Commun. 2021, 12, 1081. [Google Scholar] [CrossRef] [PubMed]
- Artes, T.; Oom, D.; de Rigo, D.; Durrant, T.; Maianti, P.; Libertà, G.; San-Miguel-Ayanz, J. A global wildfire dataset for the analysis of fire regimes and fire behaviour. Sci. Data 2019, 6. [Google Scholar] [CrossRef]
- Halofsky, J.; Peterson, D.; Harvey, B. Changing wildfire, changing forests: The effects of climate change on fire regimes and vegetation in the Pacific Northwest, USA. Fire Ecol. 2020, 16, 4. [Google Scholar] [CrossRef] [Green Version]
- Barmpoutis, P.; Papaioannou, P.; Dimitropoulos, K.; Grammalidis, N. A Review on Early Forest Fire Detection Systems Using Optical Remote Sensing. Sensors 2020, 20, 6442. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.; Zhang, Y.; Liu, Z. A survey on technologies for automatic forest fire monitoring, detection, and fighting using unmanned aerial vehicles and remote sensing techniques. Can. J. For. Res. 2015, 45, 783–792. [Google Scholar] [CrossRef]
- Axel, A.C. Burned Area Mapping of an Escaped Fire into Tropical Dry Forest in Western Madagascar Using Multi-Season Landsat OLI Data. Remote Sens. 2018, 10, 371. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; Rahman Siddiquee, M.M.; Tajbakhsh, N.; Liang, J. UNet++: A Nested U-Net Architecture for Medical Image Segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Stoyanov, D., Taylor, Z., Carneiro, G., Syeda-Mahmood, T., Martel, A., Maier-Hein, L., Tavares, J.M.R., Bradley, A., Papa, J.P., Belagiannis, V., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 3–11. [Google Scholar]
- Agne, M.C.; Beedlow, P.A.; Shaw, D.C.; Woodruff, D.R.; Lee, E.H.; Cline, S.P.; Comeleo, R.L. Interactions of predominant insects and diseases with climate change in Douglas-fir forests of western Oregon and Washington, U.S.A. For. Ecol. Manag. 2018, 409, 317–332. [Google Scholar] [CrossRef]
- Jactel, H.; Koricheva, J.; Castagneyrol, B. Responses of forest insect pests to climate change: Not so simple. Curr. Opin. Insect Sci. 2019, 35, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Przepióra, F.; Loch, J.; Ciach, M. Bark beetle infestation spots as biodiversity hotspots: Canopy gaps resulting from insect outbreaks enhance the species richness, diversity and abundance of birds breeding in coniferous forests. For. Ecol. Manag. 2020, 473, 118280. [Google Scholar] [CrossRef]
- Van Lierop, P.; Lindquist, E.; Sathyapala, S.; Franceschini, G. Global forest area disturbance from fire, insect pests, diseases and severe weather events. For. Ecol. Manag. 2015, 352, 78–88. [Google Scholar] [CrossRef] [Green Version]
- Thompson, I.; Mackey, B.; Mcnulty, S.; Mosseler, A. Forest Resilience, Biodiversity, and Climate Change. A Synthesis of the Biodiversity/Resilience/Stability Relationship in Forest Ecosystems; Secretariat of the Convention on Biological Diversity: Montreal, QC, Canada, 2009; p. 67. [Google Scholar]
- Cabezas, M.; Kentsch, S.; Tomhave, L.; Gross, J.; Caceres, M.L.L.; Diez, Y. Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data. Remote Sens. 2020, 12, 3431. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Van Rossum, G.; Drake, F.L., Jr. Python Tutorial; Centrum voor Wiskunde en Informatica: Amsterdam, The Netherlands, 1995. [Google Scholar]
- Bradski, G. The OpenCV Library. Dr. Dobb’s Journal of Software Tools. 2000. Available online: https://opencv.org/ (accessed on 15 August 2019).
- Chollet, F. Keras. 2015. Available online: https://keras.io (accessed on 12 June 2021).
- Howard, J.; Thomas, R.; Gugger, S. Fastai. 2018. Available online: https://github.com/fastai/fastai (accessed on 12 June 2021).




| Refs. | 1st Author | Problem Solved | Type of Data | Pre-Processing Software | Annotation |
|---|---|---|---|---|---|
| [47] | López-Jiménez | Cactus detection | Single images | N/A | N/A |
| [48] | Fromm | Seedling detection | image tiles | N/A | LabelImg |
| [5] | Chadwick | Tree detection segmentation | Point cloud Orthomosaic | Pix4D LAStools | N/A |
| [6] | Ocer | Tree detection counting | Orthomosaic | ArcMap | Pix4D |
| [49] | Ferreira | Palm tree detection | Orthomosaic | Pix4D | QGIS |
| [7] | Fujimoto | Species Classification | 3D point cloud | Metashape Fusion | N/A |
| [50] | Morales | Palm tree Detection | original images | N/A | N/A |
| [51] | Haq | Species Classification | Orthomosaic | Pix4D | N/A |
| [52] | Kattenborn | Species Classification | Orthomosaic DEM | Metashape | GIS-based |
| [53] | Kattenborn | Species Classification | Orthomosaic DEM | Metashape | GIS-based |
| [8] | Kentsch | Species Classification | Orthomosaic | Metashape | GIMP |
| [54] | Nezami | Species Classification | Dense point clouds orthomosaic | Metashape, | N/A |
| [55,56] | Onishi | Species Classification | Orthomosaic DSM | Metashape, | eCognition ArcGIS |
| [57] | Lin | Species Classification | Single images | N/A | N/A |
| [58,59] | Natesan | Species Classification | Orthomosaic DSM | Metashape | Local Max. watershed |
| [10] | Schiefer | Species Classification | Orthomosaic DSM, nDSM | Metashape | ArcGIS |
| [60] | Barmpoutis | Tree Health classification | Orthomosaic | Metashape | N/A |
| [61] | Humer | Tree Health classification | Single images | N/A | GIMP |
| [62] | Deng | Dead tree detection | Orthomosaic | Metashape | LabelImg |
| [63] | Nguyen | Tree Health classification | Orthomosaic nDSM | Metashape | GIMP |
| [12] | Safonova | Tree Health classification | Orthomosaic, DEM | Metashape | N/A |
| [64] | Kim | Forest fire detecton | Single images | N/A | N/A |
| [11] | Tran | Post fire mapping | Orthomosaic | DroneDeploy | Labelme |
| [65] | Hossain | Smoke/flame/ detection | Single images | N/A | N/A |
| [66] | Zhao | Smoke/flame/ detection | Single images | N/A | N/A |
| [67] | Chen | Smoke/flame/ detection | Unprocessed Data Single images | N/A | N/A |
| Ref. 1st Author | Problem Solved | Assessed Difficulty | Type of Data | Resolution | Amount of Data | DL Network | Data Issues |
|---|---|---|---|---|---|---|---|
| [47] López-Jiménez | Cactus detection (Single Label Classification) | 1 | Unprocessed images video patches | N/A | 16,136 + 5364 labelled images | LeNet 5 | IINF |
| [48] Fromm | Seedling detection | 2 | Unprocessed images | 0.3 cm | 3940 seedlings in two sites 25 m long 9415 512 × 512 tiles (multiple captures) | Faster R-CNN R-FCN, SSD | DTS |
| [5] Chadwick | Tree crown delineation | 2 | Point cloud Orthomosaic | 3 cm | 18 plots, 2.2–24.6 ha, tree stem density 1500–6800 | Mask R-CNN | DTS |
| [6] Ocer | Tree detection/counting | 2 | Orthomosaic | 4–6.5 cm | 2 flights, 2897 trees | Mask R-CNN | |
| [49] Ferreira | Palm tree detection (Species Classification) | 3 | Orthomosaic | 4 cm | 28 plots (250 × 150 m), 1423 images | DeepLabv3 | DTS |
| Ref. aUI | Problem Solved | Assessed Difficulty | Type of Data | Resolution | Amount of Data | DL Network | Data Issues |
|---|---|---|---|---|---|---|---|
| [7] Fujimoto | Species Classification (Grayscale image classification) | 2 | 3D point cloud DEM | 2.3–3.1 cm | 0.81 ha, 129+152 images | CNN | DTS |
| [50] Morales | Palm tree Detection (semantic segmentation) | 2 | unprocessed images | 1.4–2.5 cm | 4 flights, 25,248 patches | Deeplab v3+ | LS DTS |
| [51] Haq | Species Classification (semantic segmentation) | 2 | Orthomosaic? | 11.86 cm | 2040 km² area, 60 images | Autoencoder | IINF |
| [52] Kattenborn | Species Classification (semantic segmentation) | 2/3 | Orthomosaic, DEM | 3–5 cm | Site 1: 21–37 ha Site 2: 20–50 ha | CNN- UNet | LS VST |
| [53] Kattenborn | Species Mapping (Object Detection) | 2/3/3 | Orthomosaic, DEM | 5 cm, 3 cm, 3 cm | S1: 20–50 ha, 7 flights S2: 21–37, 8 flights S3: 4.3 ha, 3 flights | CNN | VST |
| [8] Kentsch | Species Classification Winter Mosaics and Invasive Species (multi-label image Classification) | 2/4 | Orthomosaic | 2.74 cm | 8 flights, 6 sites, 233–1000 images, 3–8 ha | ResNet50 | DTS * |
| [54] Nezami | Species Classification (single-label image Classification) | 3 | Dense point cloud orthomosaic | 5–10 cm | 8 flights, 3039 labelled data, 803 test data | 3D-CNN | DTS |
| [49] Ferreira | Palm tree detection/Classification (semantic segmentation) | 3 | Orthomosaic | 4 cm | 28 plots (250 × 150 m), 1423 images | DeepLabv3 | DTS |
| [55,56] Onishi | Species Classification (single-label image Classification) | 3 | Orthomosaic, DSM | 5–10 cm | 2 flights, 11 ha | CNN | DTS |
| [57] Lin | Species Classification (single-label image Classification) | 3 | unprocessed images | 0.47–1.76 cm | 50–65 images | Fourier Dense | VST |
| [9] Egli | Species Classification (single-label image Classification) | 4 | unprocessed images | 0.27–54.78 cm | 1556 images, 477 trees | lightweight CNN | |
| [58,59] Natesan | Species Classification (single-label image Classification) | 4 | Orthomosaic, DSM | 20 ha, 3 flights | VGG16, ResNet-50 DenseNet | DTS | |
| [10] Schiefer | Species Classification (semantic segmentation) | 4/5 | Orthomosaic, DSM DTM and nDSM | 1.35 cm resampled to 2 cm | 135 plots (100 × 100 m), 51 orthomosaics | CNN (UNet) | DTS ** |
| Ref. 1st Author | Assessed Difficulty | Problem Solved | Type of Data | Resolution | Amount of Data | DL Network | Data Issues |
|---|---|---|---|---|---|---|---|
| [64] Kim | 1 | Fire/not fire image classification | Single images | varying | 126,849 fire images 75,889 non fire | CNN | |
| [11] Tran | 2 | Burnt region coarse segmentation | Orthomosaic | N/A | 2 plots 43/44 images | UNet | |
| [65] Hossain | 1 | Fire/smoke coarse segmentation | Single images | varying | 460 images | YOLOv3 | IINF |
| [66] Zhao | 2 | Fire/not fire image classification | Single images | varying | 1500 images | CNN | |
| [67] Chen | 1 | Fire/smoke/normal image classification | Single images | varying | 2100 images | CNN | IINF |
| Ref. 1st Author | Problem Solved | Assessed Difficulty | Type of Data | Resolution | Amount of Data | DL Network | Data Issues |
|---|---|---|---|---|---|---|---|
| [60] Barmpoutis | Tree Health classification | 2 | Orthomosaic | N/A | 4 plots in 60 ha area, >1500 infected trees | Faster R-CNN Mask R-CNN | IINF |
| [61] Humer | Tree Health classification | 3 | unprocessed images | N/A | 35 images | UNet | VST |
| [62] Deng | Dead tree detection | 3 | Orthomosaic | N/A | 1.7952 km, 340 data points, augmented 1700 | Faster R-CNN | LS DTS |
| [63] Nguyen | Tree Health classification | 4 | Orthomosaic | 1.45–2.6 cms/pixel | 18 Ha, 9 Orthomosaics | ResNet | |
| [12] Safonova | Tree Health classification | 5 | Orthomosaic | 5–10 cms/pixel | 1200 images, 4 Orthomosaics | Custom CNN |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Diez, Y.; Kentsch, S.; Fukuda, M.; Caceres, M.L.L.; Moritake, K.; Cabezas, M. Deep Learning in Forestry Using UAV-Acquired RGB Data: A Practical Review. Remote Sens. 2021, 13, 2837. https://doi.org/10.3390/rs13142837
Diez Y, Kentsch S, Fukuda M, Caceres MLL, Moritake K, Cabezas M. Deep Learning in Forestry Using UAV-Acquired RGB Data: A Practical Review. Remote Sensing. 2021; 13(14):2837. https://doi.org/10.3390/rs13142837
Chicago/Turabian StyleDiez, Yago, Sarah Kentsch, Motohisa Fukuda, Maximo Larry Lopez Caceres, Koma Moritake, and Mariano Cabezas. 2021. "Deep Learning in Forestry Using UAV-Acquired RGB Data: A Practical Review" Remote Sensing 13, no. 14: 2837. https://doi.org/10.3390/rs13142837
APA StyleDiez, Y., Kentsch, S., Fukuda, M., Caceres, M. L. L., Moritake, K., & Cabezas, M. (2021). Deep Learning in Forestry Using UAV-Acquired RGB Data: A Practical Review. Remote Sensing, 13(14), 2837. https://doi.org/10.3390/rs13142837






