A Fully Automated Three-Stage Procedure for Spatio-Temporal Leaf Segmentation with Regard to the B-Spline-Based Phenotyping of Cucumber Plants
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Acquisition
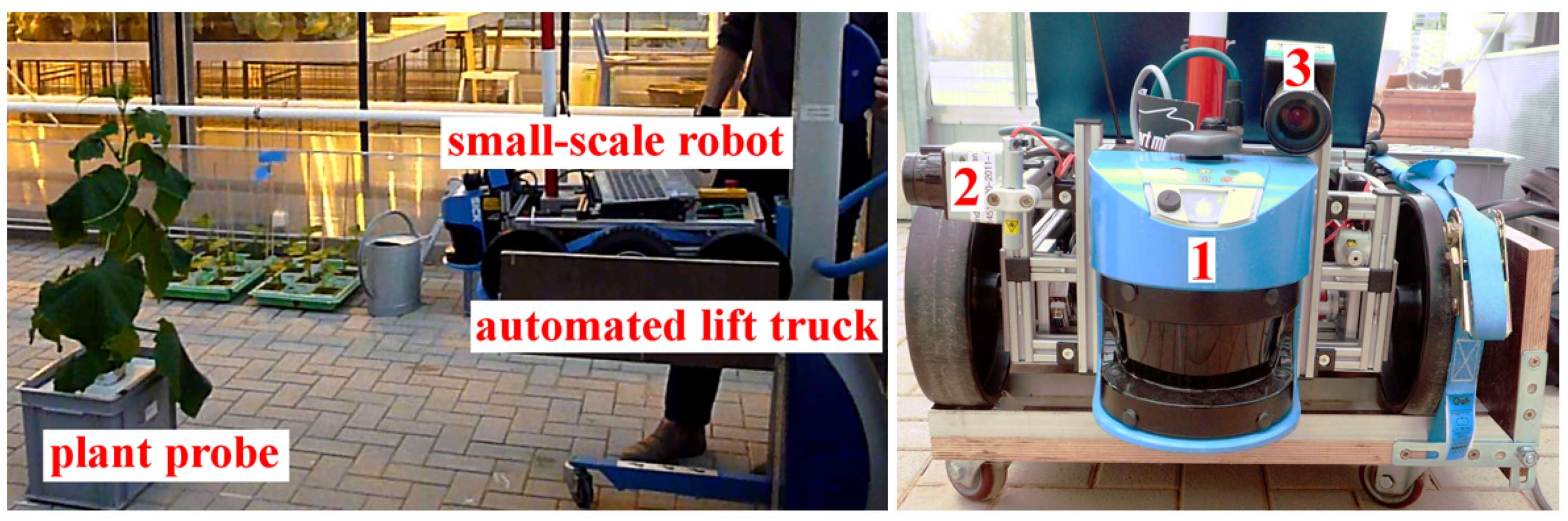
2.1.1. Multi-Sensor-System (MSS) for Plant Phenotyping
2.1.2. Reference Measurements by Means of a Geodetic Laser Scanner
2.1.3. Reference Measurements by Means of Standard Sensors of Crop Science
2.2. Spatio-Temporal Leaf Segmentation
2.2.1. Spatial Segmentation: Graph-Based Pre-Segmentation
- The internal difference measures the variation in the properties within a segment C and is defined as the maximum weight of the minimum spanning tree (MST) constructed by the nodes of C:
- The difference between two neighbouring segments and evaluates the variation between the properties of the two segments and is defined as the minimum of the weights belonging to all edges connecting and :
2.2.2. Spatial Segmentation: Statistically-Based Region Merging
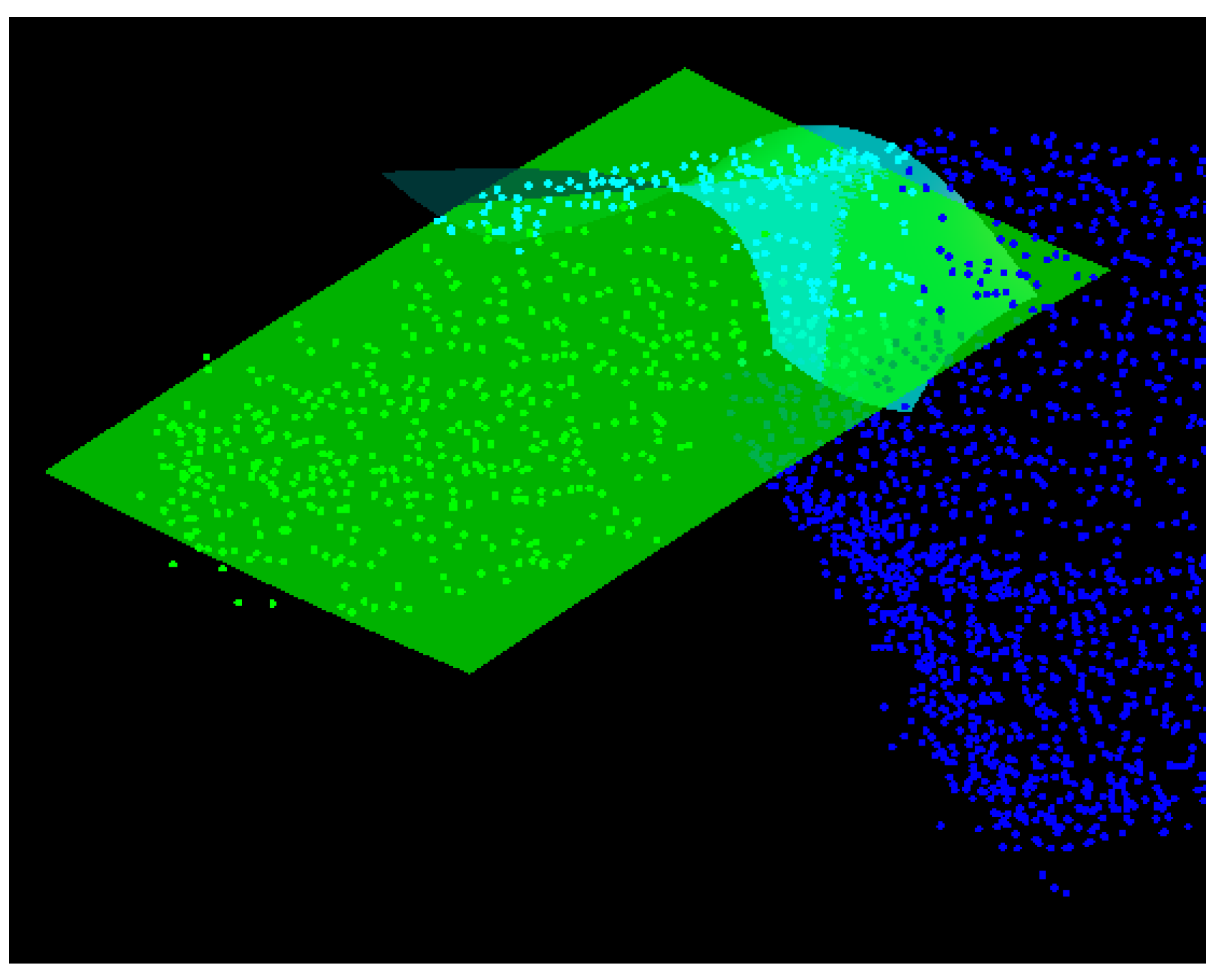
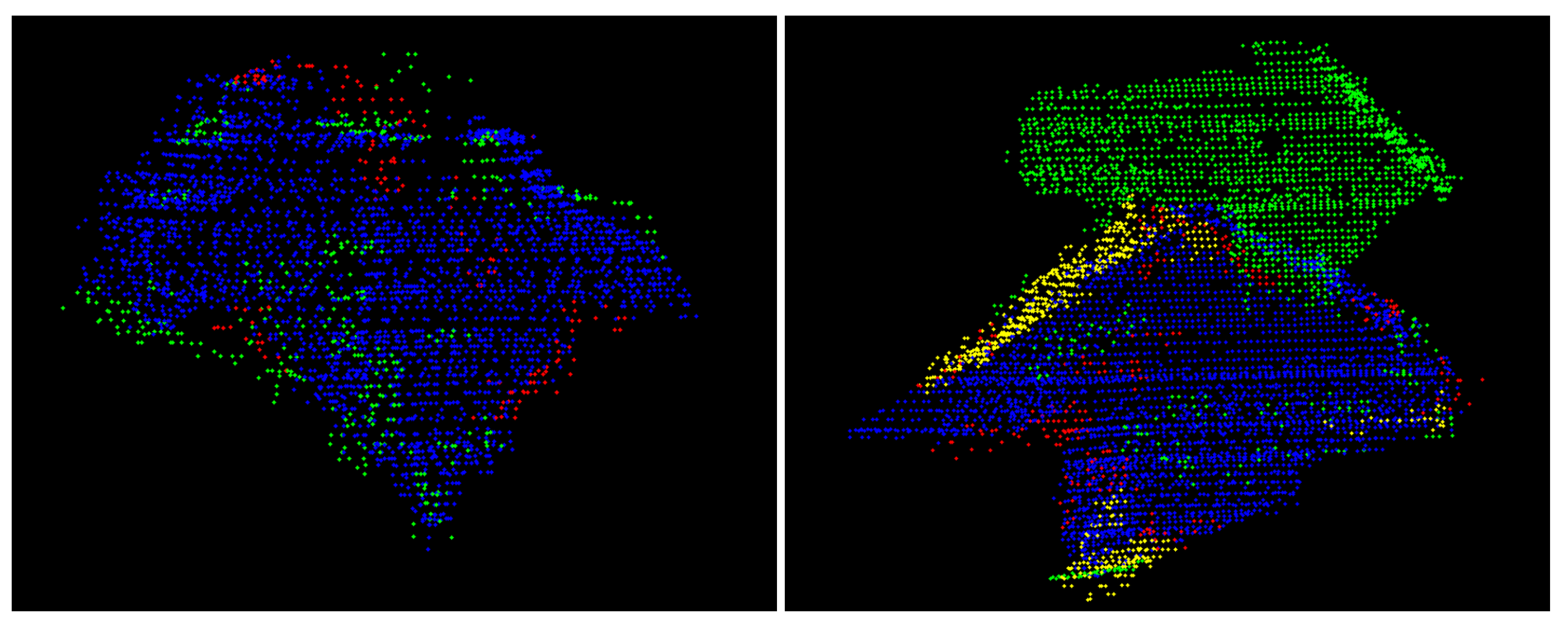
- The first test evaluates whether two neighbouring segments describe the same surface. For this purpose, a part of the super-segment (blue-coloured segment in Figure 6) as well as the neighbouring segment under investigation (green-coloured segment in Figure 6) are approximated by means of second-order surfaces (cyan- and green-coloured surfaces in Figure 6). Afterwards, the parameters of the best-fitting second order surfaces are statistically checked for equality according to the difference test described in Reference [28]. If the parameters do not differ significantly, the super-segment and the segment under investigation are merged.
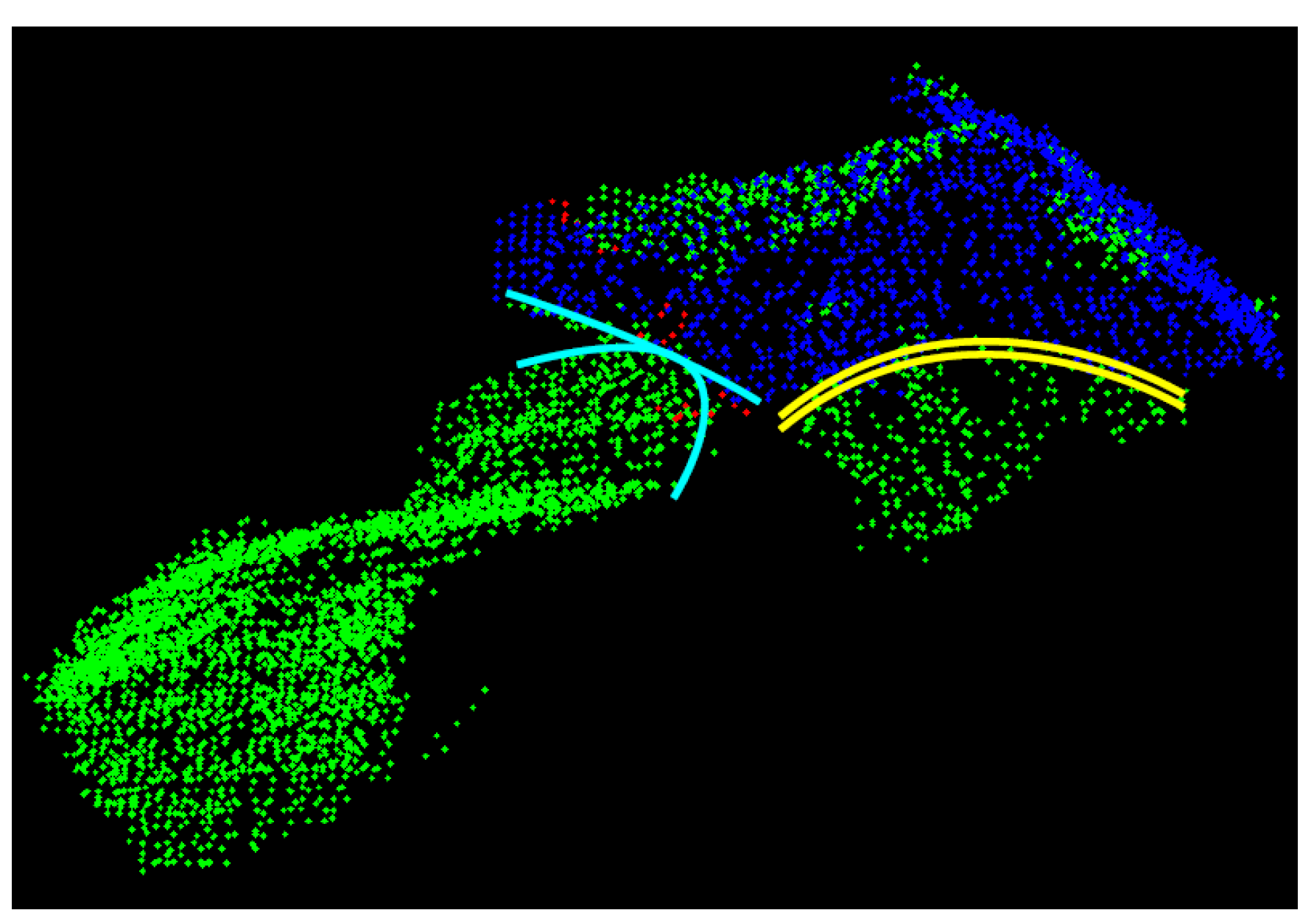
- In the second test the segments’ border edges are evaluated. As motivated by Figure 7, it can be assumed that two neighbouring segments describe the same leaf if the respective border edges describe the same space curve: This is the case in Figure 7 for the two segments with the yellow-coloured border edges, whereas the two segments with the cyan-coloured border edges describe two different, but touching leaves. In order to realize this second test, the edge points of the super-segment and of the respective neighbouring segments under investigation are automatically identified by means of a variant of the Douglas-Peucker-algorithm described in Reference [27]. Afterwards, the edge points describing a joint edge between two segments are approximated by means of space curves (cf. Figure 7). Analogously to the surface-based region merging, the parameters of the two resulting space curves are statistically checked for equality. If the estimated parameters do not differ significantly, the super-segment and the segment under investigation are merged.
2.2.3. Temporal Segmentation: Dynamic Time Warping
2.3. B-Spline Based Determination of the Leaf Areas
2.3.1. B-Spline Based Leaf Modelling
2.3.2. Determination of Leaf Areas
3. Results
3.1. Segmentation Results
3.1.1. Results of the Spatial Segmentation
3.1.2. Results of the Temporal Segmentation
3.2. Phenotyping Results
3.2.1. Phenotyping Results Delivered by the Reference Methods
3.2.2. Results of the B-Spline-Based Phenotyping
4. Discussion
4.1. Discussion of the Segmentation Results
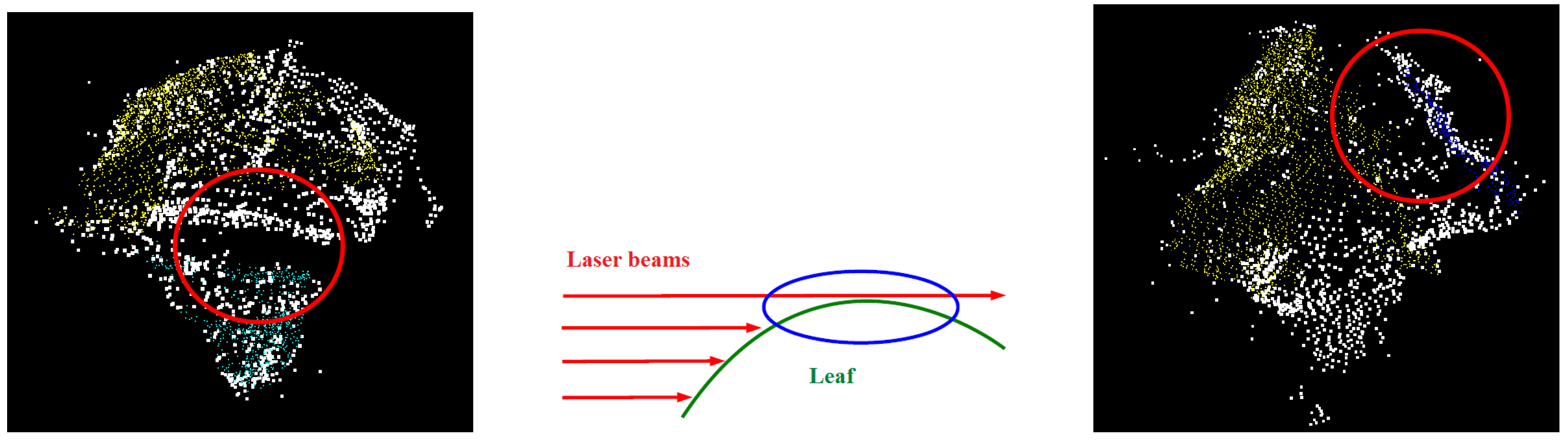
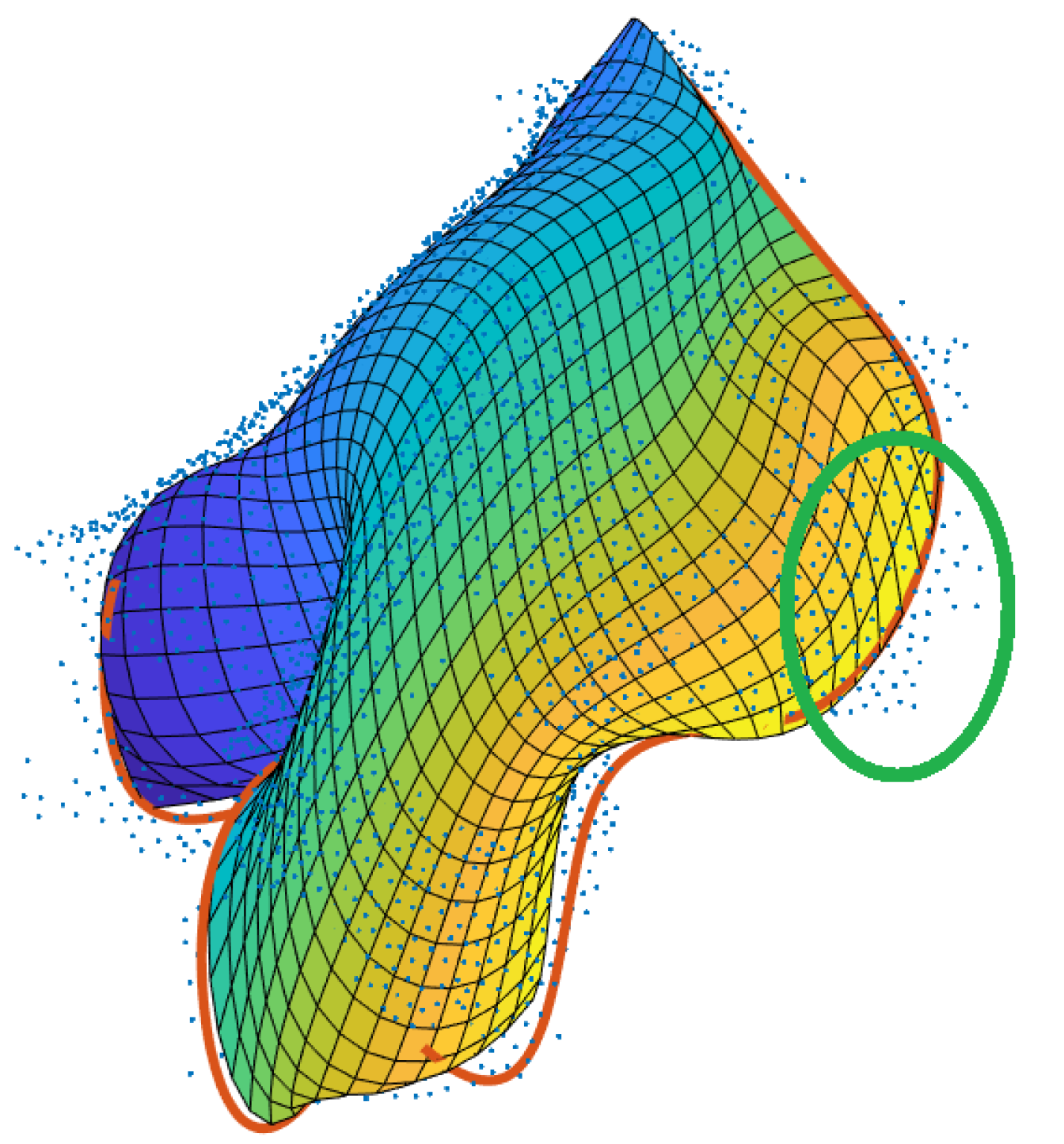
- Firstly, there is the obvious relation that the older the plant is, the larger the leaves become, leading to an increased amount of occlusion. In combination with the relatively low point density, these occlusions cause data gaps which impede the merging of neighbouring segments as can be seen in Figure 13 (left): the yellow- and the green-coloured segment obviously belong to the same leaf, but are not merged, as they are separated by a relatively large data gap (circled in red).
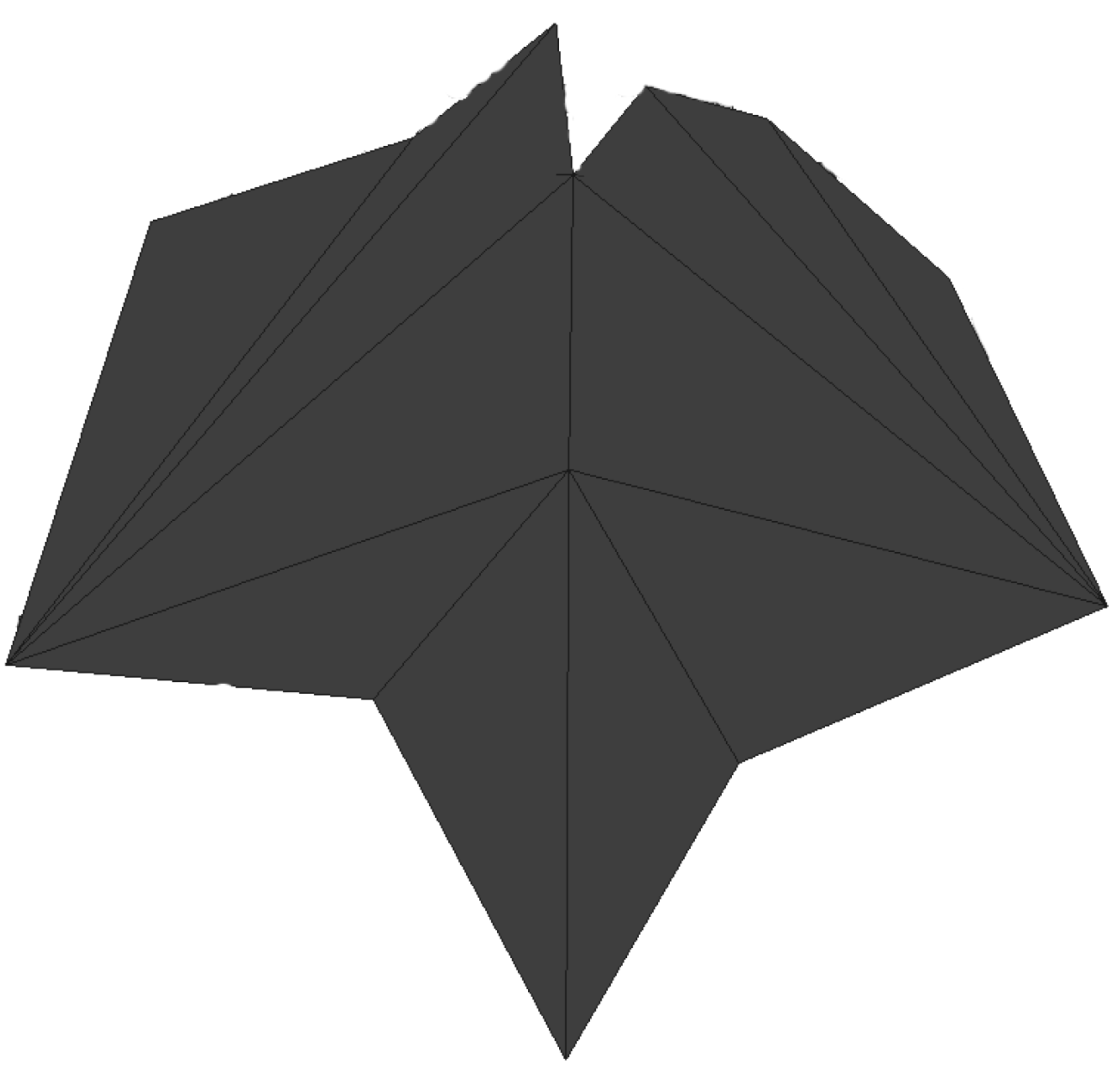
- Secondly, the plant raises its leaves from an almost vertical position to a nearly horizontal one when becoming older. As the laser beam is currently also horizontally oriented, those horizontal parts of the leaves cannot be acquired as is schematically shown in Figure 13 (middle). As a consequence, the leaves of later growth stages are incompletely acquired as can be seen in Figure 13 (right), presenting such a partly horizontally oriented leaf from above. The red circled part of the leaf is the critical part that impedes the merging of the blue- and the yellow-coloured segments.
4.2. Discussion of the Phenotyping Results
5. Conclusions
5.1. Summary
- Each point cloud is pre-segmented by means of a graph-based segmentation algorithm.
- A statistically-based region merging procedure is applied to the undersegmented point clouds, yielding spatially segmented leaves.
- The leaves are tracked over time by means of a shape matching which simultaneously improves possible erroneous spatial segmentations by using information from temporally neighbouring point clouds.
5.2. Outlook
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Furbank, R.T.; Tester, M. Phenomics–technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 2011, 16, 635–644. [Google Scholar] [CrossRef] [PubMed]
- Paproki, A.; Sirault, X.; Berry, S.; Furbank, R.; Fripp, J. A novel mesh processing based technique for 3D plant analysis. BMC Plant Biol. 2012, 12, 63. [Google Scholar] [CrossRef] [PubMed]
- Becirevic, D.; Klingbeil, L.; Honecker, A.; Schumann, H.; Rascher, U.; Léon, J.; Kuhlmann, H. On the derivation of Crop heights from multitemporal UAV imagery. ISPRS Ann. Photogram. Remote Sens. Spat. Inf. Sci. 2019, IV-2/W5, 95–102. [Google Scholar] [CrossRef]
- Johansen, K.; Morton, M.J.L.; Malbeteau, Y.; Aragon, B.; Al-Mashharawi, S.; Ziliani, M.; Angel, Y.; Fiene, G.; Negrao, S.; Mousa, M.A.A.; et al. Predicting biomass and yield at harvest of salt-stressed tomato plants using UAV imagery. ISPRS Int. Arch. Photogram. Remote Sens. Spat. Inf. Sci. 2019, XLII-2/W13, 407–411. [Google Scholar] [CrossRef]
- Hétroy-Wheeler, F.; Casella, E.; Boltcheva, D. Segmentation of tree seedling point clouds into elementary units. Int. J. Remote Sens. 2016, 37, 2881–2907. [Google Scholar] [CrossRef]
- Lou, L.; Liu, Y.; Shen, M.; Han, J.; Corke, F.; Doonan, J.H. Estimation of Branch Angle from 3D Point Cloud of Plants. In Proceedings of the IEEE 2015 International Conference on 3D Vision, Lyon, France, 19–22 October 2015; pp. 554–561. [Google Scholar] [CrossRef]
- Kahlen, K.; Stützel, H. Estimation of Geometric Attributes and Masses of Individual Cucumber Organs Using Three-dimensional Digitizing and Allometric Relationships. J. Am. Soc. Hortic. Sci. 2007, 132, 439–446. [Google Scholar] [CrossRef]
- Eberius, M.; Lima-Guerra, J. High-Throughput Plant Phenotyping—Data Acquisition, Transformation, and Analysis. In Bioinformatics; Edwards, D., Stajich, J., Hansen, D., Eds.; Springer: New York, NY, USA, 2009; Volume 88, pp. 259–278. [Google Scholar] [CrossRef]
- Hartmann, A.; Czauderna, T.; Hoffmann, R.; Stein, N.; Schreiber, F. HTPheno: An image analysis pipeline for high-throughput plant phenotyping. BMC Bioinf. 2011, 12, 148. [Google Scholar] [CrossRef]
- Iyer-Pascuzzi, A.S.; Symonova, O.; Mileyko, Y.; Hao, Y.; Belcher, H.; Harer, J.; Weitz, J.S.; Benfey, P.N. Imaging and analysis platform for automatic phenotyping and trait ranking of plant root systems. Plant Physiol. 2010, 152, 1148–1157. [Google Scholar] [CrossRef]
- Quan, L.; Tan, P.; Zeng, G.; Yuan, L.; Wang, J.; Kang, S.B. Image-based plant modeling. ACM Trans. Graph. 2006, 25, 599. [Google Scholar] [CrossRef]
- Paulus, S.; Schumann, H.; Kuhlmann, H.; Léon, J. High-precision laser scanning system for capturing 3D plant architecture and analysing growth of cereal plants. Biosyst. Eng. 2014, 121, 1–11. [Google Scholar] [CrossRef]
- Elnashef, B.; Filin, S.; Lati, R.N. Tensor-based classification and segmentation of three-dimensional point clouds for organ-level plant phenotyping and growth analysis. Comput. Electr. Agric. 2019, 156, 51–61. [Google Scholar] [CrossRef]
- Gelard, W.; Devy, M.; Herbulot, A.; Burger, P. Model-based Segmentation of 3D Point Clouds for Phenotyping Sunflower Plants. In Proceedings of the 12th International Joint Conference on Computer Vision, Porto, Portugal, 1 January 2017; pp. 459–467. [Google Scholar] [CrossRef]
- Alenya, G.; Dellen, B.; Torras, C. 3D modelling of leaves from color and ToF data for robotized plant measuring. In Proceedings of the IEEE International Conference on Robotics and Automation 2011, Shanghai, China, 9–13 May 2011; pp. 3408–3414. [Google Scholar] [CrossRef]
- Li, Y.; Fan, X.; Mitra, N.J.; Chamovitz, D.; Cohen-Or, D.; Chen, B. Analyzing growing plants from 4D point cloud data. ACM Trans. Graph. 2013, 32, 1–10. [Google Scholar] [CrossRef]
- Rist, F.; Herzog, K.; Mack, J.; Richter, R.; Steinhage, V.; Töpfer, R. High-Precision Phenotyping of Grape Bunch Architecture Using Fast 3D Sensor and Automation. Sensors 2018, 18, 763. [Google Scholar] [CrossRef] [PubMed]
- Prusinkiewicz, P.; Lindenmayer, A. The Algorithmic Beauty of Plants; Springer: New York, NY, USA, 1990. [Google Scholar] [CrossRef]
- Paulus, S.; Dupuis, J.; Mahlein, A.K.; Kuhlmann, H. Surface feature based classification of plant organs from 3D laserscanned point clouds for plant phenotyping. BMC Bioinf. 2013, 14, 238. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Short, T.H.; Fox, R.D.; Bauerle, W.L. Plant architectural parameters of a greenhouse cucumber row crop. Agric. Forest Meteorol. 1990, 51, 93–105. [Google Scholar] [CrossRef]
- Qian, T.; Zheng, X.; Guo, X.; Wen, W.; Yang, J.; Lu, S. Influence of temperature and light gradient on leaf arrangement and geometry in cucumber canopies: Structural phenotyping analysis and modelling. Inf. Process. Agric. 2019, 6, 224–232. [Google Scholar] [CrossRef]
- Paffenholz, J.A.; Harmening, C. Spatiotemporal monitoring of natural objects in occluded scenes. In 4th International Conference on Machine Control & Guidance; Schattenberg, J., Minßen, T.F., Eds.; Institut für mobile Maschinen und Nutzfahrzeuge: Braunschweig, Germany, 2014; pp. 63–74. [Google Scholar]
- Besl, P.J.; McKay, N.D. A method for registration of 3-D shapes. IEEE Trans. Pattern Anal. Mach. Intell. 1992, 14, 239–256. [Google Scholar] [CrossRef]
- SINOQUET, H. Characterization of the Light Environment in Canopies Using 3D Digitising and Image Processing. Ann. Botany 1998, 82, 203–212. [Google Scholar] [CrossRef]
- Wiechers, D.; Kahlen, K.; Stützel, H. A method to analyse the radiation transfer within a greenhouse cucumber canopy (Cucumis sativus L.). Acta Hortic. 2006, 75–80. [Google Scholar] [CrossRef]
- Felzenszwalb, P.F.; Huttenlocher, D.P. Efficient Graph-Based Image Segmentation. Int. J. Comput. Vis. 2004, 59, 167–181. [Google Scholar] [CrossRef]
- Harmening, C.; Brenner, C.; Paffenholz, J.A. Raumzeitliche Segmentierung von Pflanzen in stark verdeckten Szenen. In Photogrammetrie—Laserscanning—Optische 3D-Messtechnik; Luhmann, T., Müller, C., Eds.; Wichmann: Berlin, Germany, 2014; pp. 334–341. [Google Scholar]
- Heunecke, O.; Kuhlmann, H.; Welsch, W.; Eichhorn, A.; Neuner, H. Handbuch Ingenieurgeodäsie: Auswertung geodätischer Überwachungsmessungen, 2nd ed.; Wichmann: Heidelberg, Germany, 2008. [Google Scholar]
- Brendel, W.; Todorovic, S. Video object segmentation by tracking regions. In Proceedings of the IEEE 12th International Conference on Computer Vision (ICCV), Kyoto, Japan, 29 September–2 October 2009; pp. 833–840. [Google Scholar] [CrossRef]
- Müller, M. Information Retrieval for Music and Motion; Springer: Berlin/Heidelberg, Germny, 2007. [Google Scholar]
- Edelsbrunner, H.; Kirkpatrick, D.; Seidel, R. On the shape of a set of points in the plane. IEEE Trans. Inf. Theory 1983, 29, 551–559. [Google Scholar] [CrossRef]
- Edelsbrunner, H.; Mücke, E.P. Three-dimensional alpha shapes. ACM Trans. Graph. 1994, 13, 43–72. [Google Scholar] [CrossRef]
- Harmening, C.; Neuner, H. A constraint-based parameterization technique for B-spline surfaces. J. Appl. Geodesy 2015, 9, 143–161. [Google Scholar] [CrossRef]
- Beardsley, P.; Chaurasia, G. Editable Parametric Dense Foliage from 3D Capture. In Proceedings of the IEEE 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 5315–5324. [Google Scholar] [CrossRef]
- Piegl, L.A.; Tiller, W. The NURBS book. In Monographs In Visual Communications, 2nd ed.; Springer: Berlin, Germany; New York, NY, USA, 1997. [Google Scholar]
- Cox, M.G. The Numerical Evaluation of B-Splines. IMA J. Appl. Math. 1972, 10, 134–149. [Google Scholar] [CrossRef]
- Boor, C.d. On calculating with B-splines. J. Approx. Theory 1972, 6, 50–62. [Google Scholar] [CrossRef]
- Schmitt, C.; Neuner, H. Knot estimation on B-Spline curves. Österreichische Z. Vermessung Geoinf. (VGI) 2015, 103, 188–197. [Google Scholar]
- Bureick, J.; Alkhatib, H.; Neumann, I. Robust Spatial Approximation of Laser Scanner Point Clouds by Means of Free-form Curve Approaches in Deformation Analysis. J. Appl. Geodesy 2016, 10, 27–35. [Google Scholar] [CrossRef]
- Harmening, C.; Neuner, H. Choosing the Optimal Number of B-spline Control Points (Part 1: Methodology and Approximation of Curves). J. Appl. Geodesy 2016, 10, 139–157. [Google Scholar] [CrossRef]
- Harmening, C.; Neuner, H. Choosing the optimal number of B-spline control points (Part 2: Approximation of surfaces and applications). J. Appl. Geodesy 2017, 11. [Google Scholar] [CrossRef]








| Min (res) | Max (res) | Std (res) | |
|---|---|---|---|
| Epoch 1 | mm | 10 mm | 4 mm |
| Epoch 2 | mm | 7 mm | 3 mm |
| Epoch 3 | mm | 12 mm | 4 mm |
| Name | Plant | Epoch | Stage | Sensor | Reference Measurement | Height [mm] | |
|---|---|---|---|---|---|---|---|
| E1 | early | MSS | Digitizer | ≈1200 | |||
| E2 | early | MSS | Digitizer | ≈1200 | |||
| E3 | early | MSS | Digitizer | ≈1200 | |||
| E4 | early | MSS | Digitizer | ≈1200 | |||
| E1 | late | MSS | Digitizer, Leaf area meter | ≈1400 | |||
| E2 | late | MSS | Digitizer, Leaf area meter | ≈1400 | |||
| E3 | late | MSS | Digitizer, Leaf area meter | ≈1400 | |||
| E4 | late | MSS | Digitizer, Leaf area meter | ≈1400 | |||
| E1 | late | TLS | Digitizer | ≈1400 |
| ID | [cm] | [cm] | [cm] | |
|---|---|---|---|---|
| 1 | 464.33 | 517.08 | 52.75 | 11.4% |
| 3 | 434.10 | 451.03 | 16.93 | 3.9% |
| 4 | 404.21 | 691.70 | 287.49 | 71.1% |
| 5 | 417.48 | 470.23 | 52.75 | 12.6% |
| 7 | 716.13 | 791.82 | 75.69 | 10.6% |
| 14 | 667.24 | 724.39 | 57.15 | 8.6% |
| ID | [cm] | [cm] | [cm] | |
|---|---|---|---|---|
| 4 | 410.67 | 23.9 | 16.54 | 4.2% |
| 5 | 497.87 | 15.6 | −5.98 | −1.2% |
| 6 | 489.06 | 6.6 | 0.002 | 0.0% |
| 7 | 401.17 | 12.4 | 29.74 | 8.0% |
| 8 | 395.67 | 9.2 | 6.51 | 1.7% |
| 9 | 345.81 | 7.9 | −12.11 | −3.4% |
| 10 | 419.56 | 34.9 | −60.89 | −12.7% |
| 11 | 449.23 | 20.2 | 0.13 | 0.0% |
| ID | [cm] | [cm] | [cm] | |
|---|---|---|---|---|
| 1 | 528.4275 | 26.9 | 11.34 | 2.1% |
| 3 | 326.8175 | 10.5 | −124.21 | −38.1% |
| 4 | 457.0825 | 31.2 | −234.62 | −51.3% |
| 5 | 466.7725 | 13.1 | −3.46 | −0.7% |
| 7 | 750.2975 | 50.1 | −41.52 | −5.5% |
| 14 | 705.7325 | 26.2 | −18.66 | −2.6% |
| ID | [cm] | [cm] | |
|---|---|---|---|
| 3 | 736.81 | 71.05 | 10.7% |
| 6 | 1094.20 | 27.66 | 2.6% |
| 8 | 1178.52 | 100.13 | 9.3% |
| 10 | 1529.59 | 151.03 | 11.0% |
| 13 | 1198.27 | 34.8 | 3.0% |
| 15 | 907.53 | −18.48 | −2% |
| 17 | 648.56 | 62.51 | 10.7% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harmening, C.; Paffenholz, J.-A. A Fully Automated Three-Stage Procedure for Spatio-Temporal Leaf Segmentation with Regard to the B-Spline-Based Phenotyping of Cucumber Plants. Remote Sens. 2021, 13, 74. https://doi.org/10.3390/rs13010074
Harmening C, Paffenholz J-A. A Fully Automated Three-Stage Procedure for Spatio-Temporal Leaf Segmentation with Regard to the B-Spline-Based Phenotyping of Cucumber Plants. Remote Sensing. 2021; 13(1):74. https://doi.org/10.3390/rs13010074
Chicago/Turabian StyleHarmening, Corinna, and Jens-André Paffenholz. 2021. "A Fully Automated Three-Stage Procedure for Spatio-Temporal Leaf Segmentation with Regard to the B-Spline-Based Phenotyping of Cucumber Plants" Remote Sensing 13, no. 1: 74. https://doi.org/10.3390/rs13010074
APA StyleHarmening, C., & Paffenholz, J.-A. (2021). A Fully Automated Three-Stage Procedure for Spatio-Temporal Leaf Segmentation with Regard to the B-Spline-Based Phenotyping of Cucumber Plants. Remote Sensing, 13(1), 74. https://doi.org/10.3390/rs13010074







