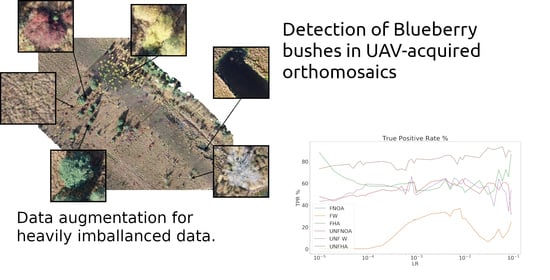
Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data
Abstract
1. Introduction
2. Materials and Methods
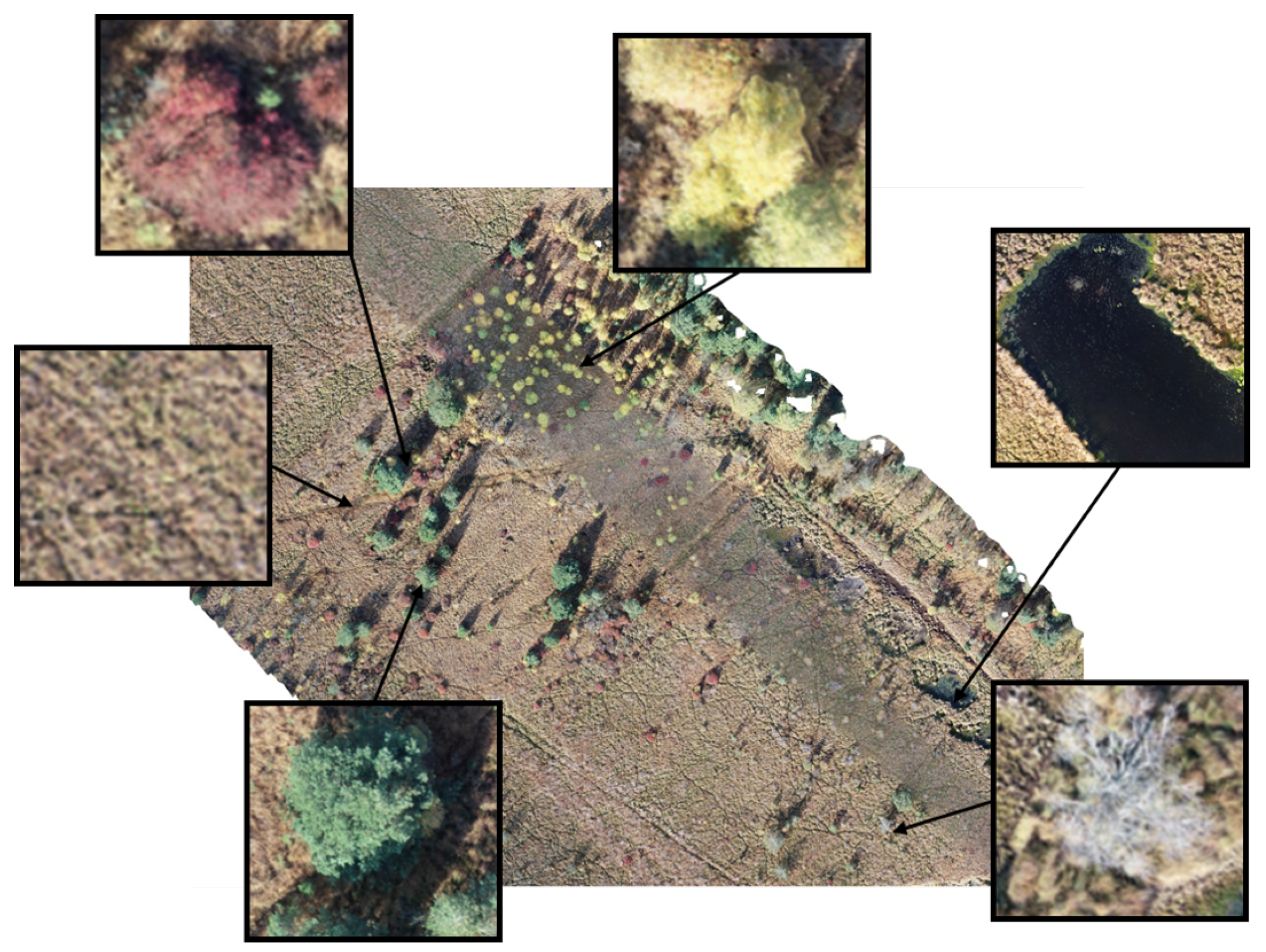
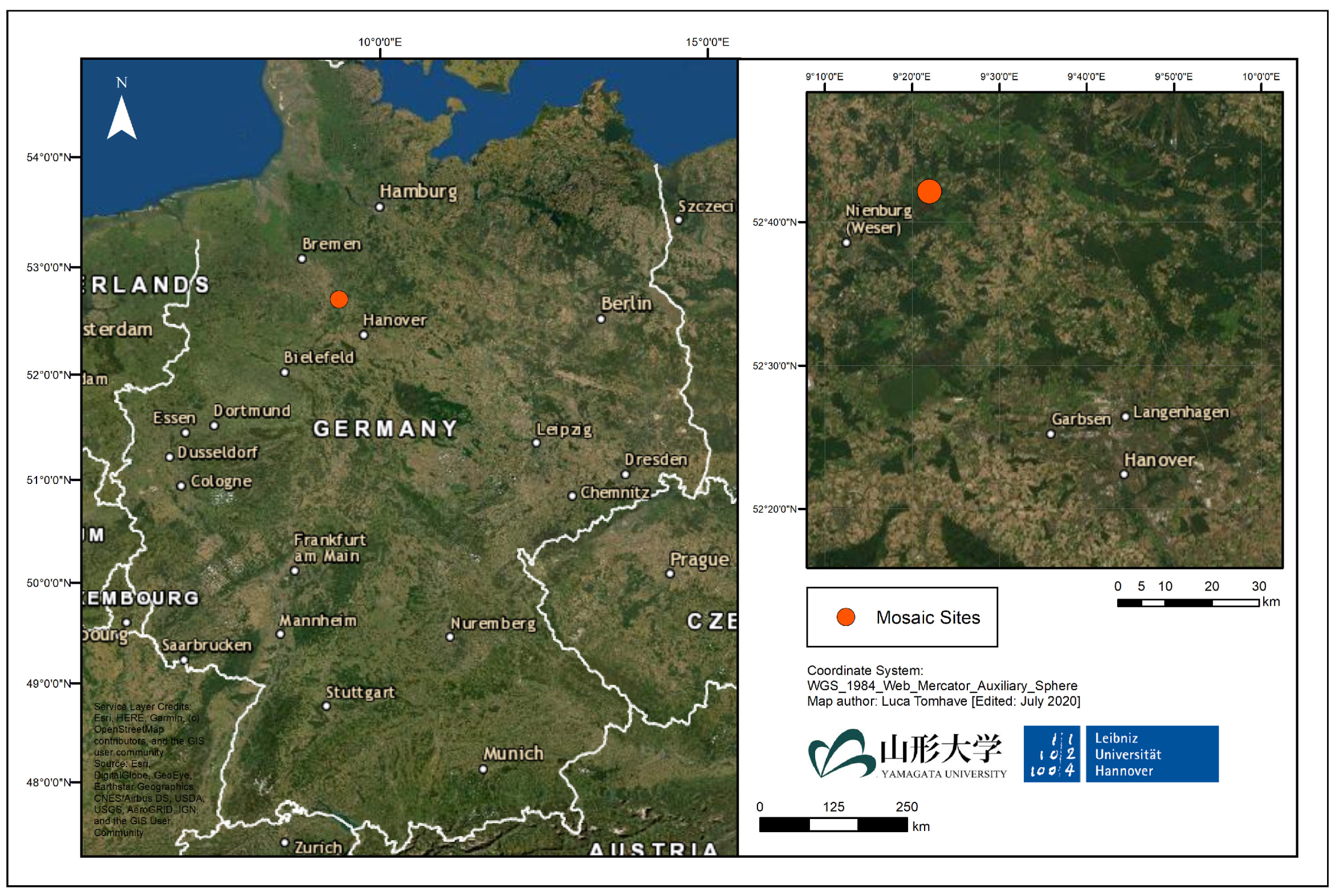
2.1. Data
Annotation and Dataset Construction
2.2. Definition of the DL Network
- Alexnet (alexnet) [18] is one of the first widely used convolutional neural networks, composed of eight layers (five convolutional layers sometimes followed by max-pooling layers and three fully connected layers). This network was the one that started the current DL trend after outperforming the current state-of-the-art method on the ImageNet data set by a large margin.
- VGG (vgg19_bn) [20] represents an evolution of the Alexnet network that allowed for an increased number of layers (19 with batch normalization in the version considered in our work) by using smaller convolutional filters.
- ResNet (resnet50, resnet152) [21] was one of the first DL architectures to allow higher number of layers by including blocks composed of convolution, batch normalization, and ReLU. Two versions with 50 and 152 layers, respectively, were used.
- Squeezenet (squeezenet1_0) [19] used so-called squeeze filters, including point-wise filter to reduce the number of parameters needed. A similar accuracy to Alexnet was claimed with fewer parameters.
- Densenet (densenet161) [22] uses a larger number of connections between layers to claim increased parameter efficiency and better feature propagation that allows them to work with even more layers (161 in this work).
- Wide ResNets (wide_resnet101_2) [24] tweak the basic architecture of regular ResNets to add more feature maps in each layer (increase width) while reducing the number of layers (network depth) in the hopes of ameliorating problems such as diminishing feature reuse.
- ResNeXt (resnext101_32x8d) [23] is a modification of the ResNet network that seeks to present a simple design that is easy to apply to practical problems. Specifically, the architecture has only a few hyper-parameters, with the most important being the cardinality (i.e., the number of independent paths, in the model).
2.3. Data Augmentation and Transfer Learning
- Small central rotations with a random angle. Depending on the orientation of the UAV, different orthomosaics acquired during different time frames might show different perspectives of the same trees. In order to introduce invariance to these differences, flips on the two main image axes can be applied to artificially increase the number of samples.
- Flips on the X and Y axes (up/down and left/right). Another way of addressing these differences is to mirror the image on their main axes (up/down, left/right).
- Gaussian blurring of the images. Due to the acquisition (movement, sensor characteristics, distance, etc.) and mosaicing process, some regions of the image might also present some blurring. Simulating these blurring with a Gaussian kernel to artificially expand the training dataset can also be used to simulate these issues and improve generalization.
- Linear and small contrast changes. Similarly, different lightning or shadows between regions of the image might also affect the results. By introducing these contrast changes, these effects can be stimulated and enlarge the number of training samples.
- Localized elastic deformation. Finally, elastic deformation were applied to simulate the possible different intra-species shapes of the blueberry patches.
2.4. Evaluation Criteria
3. Results
- First fold, testing: Orthomosaic 1, (6400 patches, with 2.53% blueberry), Training: Orthomosaics 2,3 (22,562 patches with 2.66 blueberry)
- Second fold, testing: Orthomosaic 2, (14,641 patches, with 2.58% blueberry), Training: Orthomosaics 1,3 (14,321 patches with 2.68 blueberry)
- Third fold, testing: Orthomosaic 3, (7921 patches, with 2.53% blueberry), Training: Orthomosaics 1,2 (21,041 patches with 2.57 blueberry)
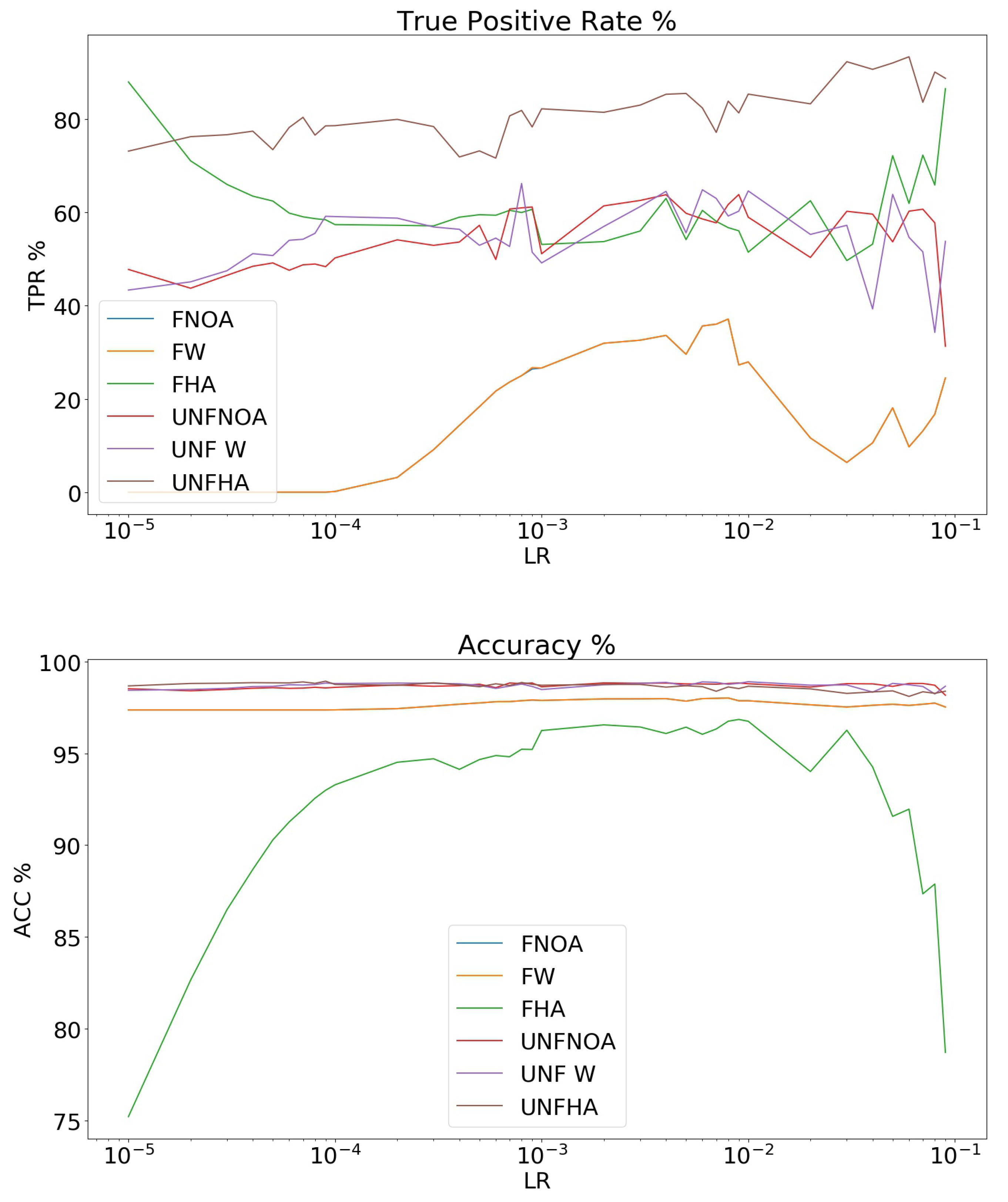
3.1. Data Balancing and TL
- Loss function Weighting. By giving different weights to the different classes in the loss function the relative importance of each class can be altered. However, this is not enough, as correctly detecting the presence of a class contributes the same as correctly detecting its absence: A network that does not predict the blueberry class in any patch will still be right over 97% of times. Consequently, even with loss function weighting, infrequent classes will remain underpredicted.
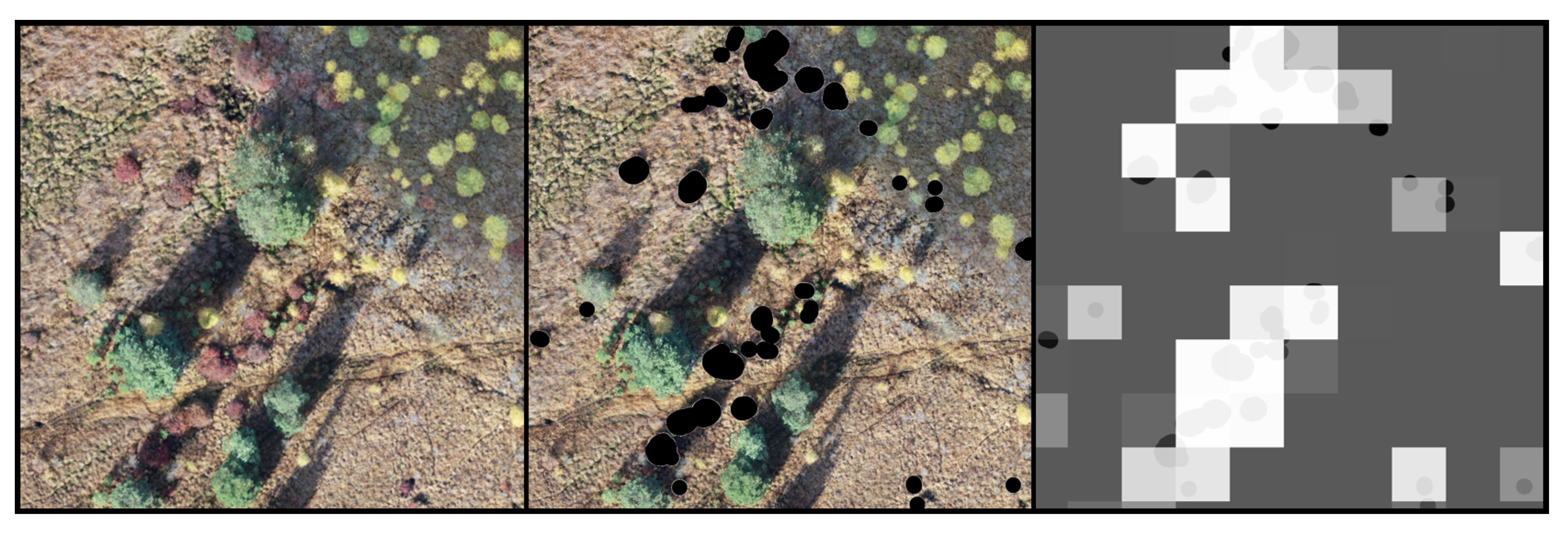
- Data augmentation: By making copies with simple transformations (see Section 2.3) of the blueberry patches the distribution of the training set can be altered and thus, increase the importance of classes in the loss function. This is expected to increase the classification accuracy of the patches containing the augmented classes while decreasing that of other classes.
- No augmentation and no weighting of the loss function. This network was considered frozen FNOA and unfrozen UNFNOA.
- Only weighting of the loss function, with no data augmentation, FW and UNFW. In this case, the weights for the six classes were [6,2,2,1,2,2] in order to give more importance to the blueberry class and less to the soil class.
- Weighting of the loss function [8,2,2,1,2,2]. The blueberry class was, thus, assigned a weight of “8”, the soil class a weight of “1”, and the rest of classes a weight of “2”. A “high level” of data augmentation was used, naming the data sets FHA and UNFHA. Twelve new images for each image of the blueberry class was created.
Statistical Significance of the Results
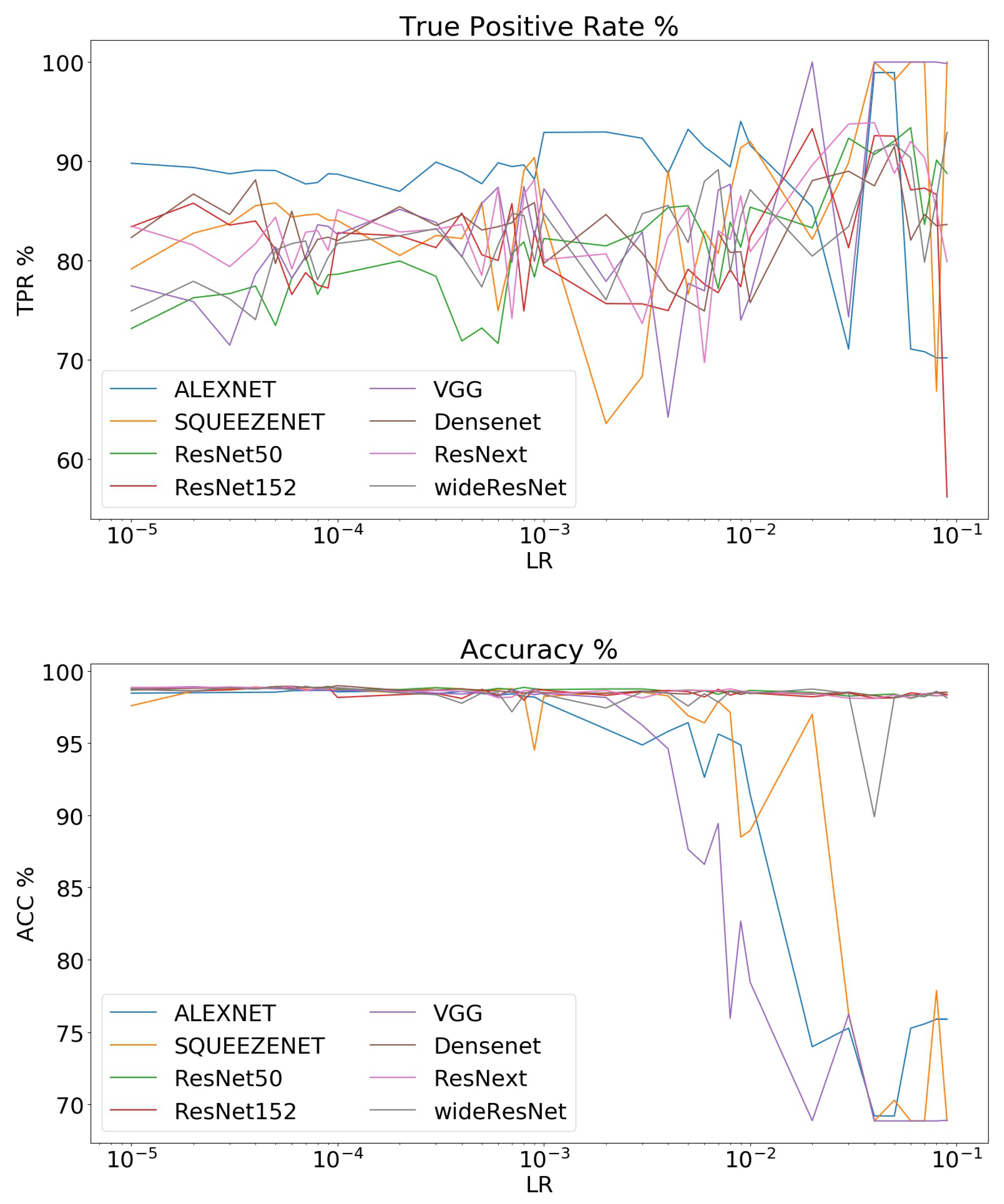
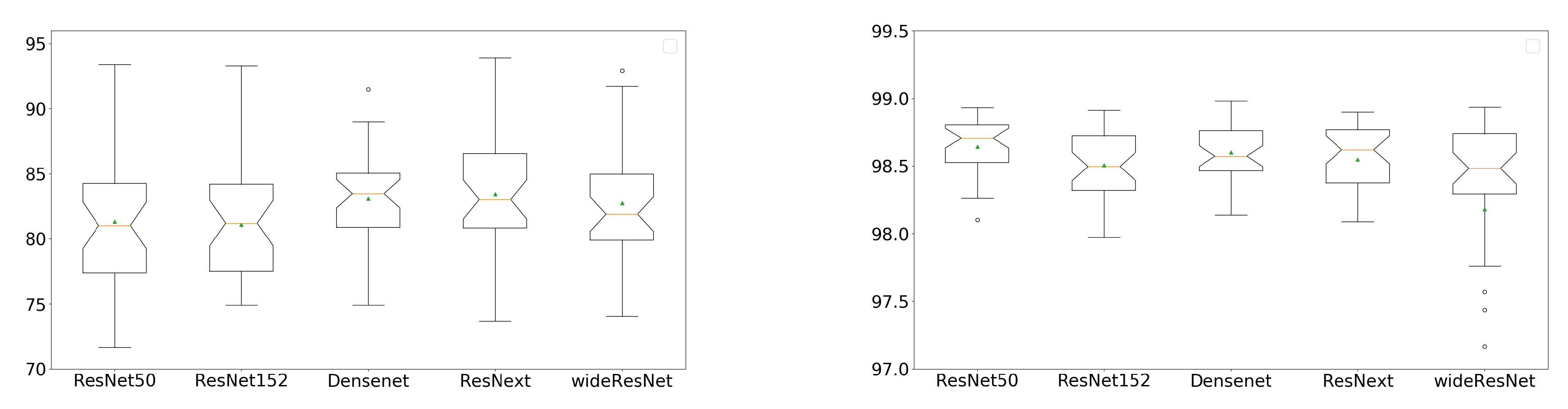
3.2. Comparison of Different Networks
4. Discussion
5. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
References
- Prentis, P.J.; Wilson, J.R.; Dormontt, E.E.; Richardson, D.M.; Lowe, A.J. Adaptive evolution in invasive species. Trends Plant Sci. 2008, 13, 288–294. [Google Scholar] [CrossRef] [PubMed]
- Pyšek, P.; Richardson, D.M. Invasive Species, Environmental Change and Management, and Health. Annu. Rev. Environ. Resour. 2010, 35, 25–55. [Google Scholar] [CrossRef]
- Pimentel, D.; Zuniga, R.; Morrison, D. Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol. Econ. 2005, 52, 273–288. [Google Scholar] [CrossRef]
- Grenzdorffer, G.; Teichert, B. The photogrammetric potential of low-cost UAVs in forestry and agriculture. Int. Arch. Photogramm. Remote Sens. Spatial Inf. Sci. 2008, 31, 1207–1214. [Google Scholar]
- Raparelli, E.; Bajocco, S. A bibliometric analysis on the use of unmanned aerial vehicles in agricultural and forestry studies. Int. J. Remote Sens. 2019, 40, 9070–9083. [Google Scholar] [CrossRef]
- Tang, L.; Shao, G. Drone remote sensing for forestry research and practices. J. For. Res. 2015, 26, 791–797. [Google Scholar] [CrossRef]
- Natesan, S.; Armenakis, C.; Vepakomma, U. Resnet-Based Tree Species Classification Using Uav Images. ISPRS-Int. Arch. Photogramm. Remote. Sens. Spat. Inf. Sci. 2019, XLII-2/W13, 475–481. [Google Scholar] [CrossRef]
- Gambella, F.; Sistu, L.; Piccirilli, D.; Corposanto, S.; Caria, M.; Arcangeletti, E.; Proto, A.R.; Chessa, G.; Pazzona, A. Forest and UAV: A bibliometric review. Contemp. Eng. Sci. 2016, 9, 1359–1370. [Google Scholar] [CrossRef]
- Kentsch, S.; Lopez Caceres, M.L.; Serrano, D.; Roure, F.; Diez, Y. Computer Vision and Deep Learning Techniques for the Analysis of Drone-Acquired Forest Images, a Transfer Learning Study. Remote Sens. 2020, 12, 1287. [Google Scholar] [CrossRef]
- Sa, I.; Chen, Z.; Popović, M.; Khanna, R.; Liebisch, F.; Nieto, J.; Siegwart, R. weedNet: Dense Semantic Weed Classification Using Multispectral Images and MAV for Smart Farming. IEEE Robot. Autom. Lett. 2018, 3, 588–595. [Google Scholar] [CrossRef]
- Deng, L.; Yu, R. Pest recognition system based on bio-inspired filtering and LCP features. In Proceedings of the 2015 12th International Computer Conference on Wavelet Active Media Technology and Information Processing (ICCWAMTIP), Chengdu, China, 18–20 December 2015; pp. 202–204. [Google Scholar] [CrossRef]
- Shiferaw, H.; Bewket, W.; Eckert, S. Performances of machine learning algorithms for mapping fractional cover of an invasive plant species in a dryland ecosystem. Ecol. Evol. 2019, 9, 2562–2574. [Google Scholar] [CrossRef] [PubMed]
- Stieper, L.C. Distribution of Wild Growing Cultivated Blueberries in Krähenmoor and Their Impact on Bog Vegetation and Bog Development. Bachelor’s Thesis, Institute of Physical Geography and Landscape Ecology, Leibniz University of Hannover, Hannover, Germany, 2018. [Google Scholar]
- Schepker, H.; Kowarik, I.; Grave, E. Verwilderung nordamerikanischer Kultur-Heidelbeeren (Vaccinium subgen. Cyanococcus) in Niedersachsen und deren Einschätzung aus Naturschutzsicht. Nat. Und Landsch. 1997, 72, 346–351. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25; Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Dupret, G.; Koda, M. Bootstrap re-sampling for unbalanced data in supervised learning. Eur. J. Oper. Res. 2001, 134, 141–156. [Google Scholar] [CrossRef]
- He, H.; Garcia, E.A. Learning from Imbalanced Data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2017, 60. [Google Scholar] [CrossRef]
- Iandola, F.N.; Moskewicz, M.W.; Ashraf, K.; Han, S.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and <1 MB model size. arXiv 2016, arXiv:1602.07360. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. arXiv 2015, arXiv:1512.03385. [Google Scholar]
- Huang, G.; Liu, Z.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the 2017 IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2261–2269. [Google Scholar]
- Xie, S.; Girshick, R.; Dollar, P.; Tu, Z.; He, K. Aggregated Residual Transformations for Deep Neural Networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Zagoruyko, S.; Komodakis, N. Wide Residual Networks. In Proceedings of the British Machine Vision Conference (BMVC); Richard, C., Wilson, E.R.H., Smith, W.A.P., Eds.; BMVA Press: York, UK, 2016; pp. 87.1–87.12. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T. A survey on Image Data Augmentation for Deep Learning. J. Big Data 2019, 6, 1–48. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, J.; Tian, J.; Zhuo, L.; Zhang, J. Residual Dense Network Based on Channel-Spatial Attention for the Scene Classification of a High-Resolution Remote Sensing Image. Remote Sens. 2020, 12, 1887. [Google Scholar] [CrossRef]
- Masarczyk, W.; Głomb, P.; Grabowski, B.; Ostaszewski, M. Effective Training of Deep Convolutional Neural Networks for Hyperspectral Image Classification through Artificial Labeling. Remote Sens. 2020, 12, 2653. [Google Scholar] [CrossRef]
- Agisoft. Agisoft Metashape 1.5.5, Professional Edition. Available online: http://www.agisoft.com/downloads/installer/ (accessed on 19 August 2019).
- Team, T.G. GNU Image Manipulation Program. 2020. Available online: http://gimp.org (accessed on 29 June 2020).
- Shirokikh, B.; Zakazov, I.; Chernyavskiy, A.; Fedulova, I.; Belyaev, M. First U-Net Layers Contain More Domain Specific Information Than The Last Ones. In Proceedings of the DART workshop at the MICCAI Conference 2020, Lime, Peru, 4–8 October 2020. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Smith, L.N.; Topin, N. Super-Convergence: Very Fast Training of Neural Networks Using Large Learning Rates. arXiv 2017, arXiv:1708.07120. [Google Scholar]
- Jung, A.B.; Wada, K.; Crall, J.; Tanaka, S.; Graving, J.; Reinders, C.; Yadav, S.; Banerjee, J.; Vecsei, G.; Kraft, A.; et al. Imgaug. 2020. Available online: https://github.com/aleju/imgaug (accessed on 1 July 2020).
- Othmani, A.; Taleb, A.R.; Abdelkawy, H.; Hadid, A. Age estimation from faces using deep learning: A comparative analysis. Comput. Vis. Image Underst. 2020, 196, 102961. [Google Scholar] [CrossRef]
- Malik, H.; Farooq, M.S.; Khelifi, A.; Abid, A.; Nasir Qureshi, J.; Hussain, M. A Comparison of Transfer Learning Performance Versus Health Experts in Disease Diagnosis From Medical Imaging. IEEE Access 2020, 8, 139367–139386. [Google Scholar] [CrossRef]
- Van Rossum, G.; Drake, F.L., Jr. Python Tutorial; Centrum voor Wiskunde en Informatica: Amsterdam, The Netherlands, 1995. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems 32; Wallach, H., Larochelle, H., Beygelzimer, A., dAlchéBuc, F., Fox, E., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2019; pp. 8024–8035. [Google Scholar]
- Zaman-Allah, M.; Vergara, O.; Araus, J.L.; Tarekegne, A.; Magorokosho, C.; Zarco-Tejada, P.J.; Hornero, A.; Albà, A.H.; Das, B.; Craufurd, P.; et al. Unmanned aerial platform-based multi-spectral imaging for field phenotyping of maize. Plant Methods 2015, 11. [Google Scholar] [CrossRef]
- Frid-Adar, M.; Diamant, I.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing 2018, 321, 321–331. [Google Scholar] [CrossRef]
- Sandfort, V.; Yan, K.; Pickhardt, P.J.; Summers, R.M. Data augmentation using generative adversarial networks (CycleGAN) to improve generalizability in CT segmentation tasks. Nat. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef] [PubMed]


| TPR | FPR | ACC | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Best | Mean | Stdev | Best | Mean | Stdev | Best | Mean | Stdev | |
| FNOA | 37.13 | 15.86 | 13.05 | 0.00 | 0.13 | 0.14 | 98.01 | 97.66 | 0.24 |
| FW | 37.13 | 15.87 | 13.06 | 0.00 | 0.13 | 0.14 | 98.01 | 97.66 | 0.24 |
| FHA | 87.99 | 61.24 | 8.25 | 2.04 | 6.66 | 5.45 | 96.84 | 92.49 | 5.10 |
| UNFNOA | 63.83 | 54.55 | 7.05 | 0.02 | 0.13 | 0.04 | 98.83 | 98.68 | 0.15 |
| UNF W | 66.21 | 54.98 | 7.11 | 0.04 | 0.12 | 0.07 | 98.90 | 98.69 | 0.15 |
| UNFHA | 93.39 | 81.31 | 5.85 | 0.47 | 0.89 | 0.35 | 98.93 | 98.64 | 0.21 |
| TPR | FPR | ACC | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Best | Mean | Stdev | Best | Mean | Stdev | Best | Mean | Stdev | |
| ImageNet Weights | 93.28 | 81.31 | 5.85 | 0.47 | 0.88 | 0.34 | 98.92 | 98.64 | 0.21 |
| Random Weights | 92.58 | 83.88 | 7.66 | 0.79 | 1.33 | 0.46 | 98.63 | 98.27 | 0.53 |
| TPR | FPR | ACC | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Best | Mean | Stdev | Best | Mean | Stdev | Best | Mean | Stdev | |
| UNFNOA | 66.05 | 52.05 | 11.25 | 0.09 | 0.19 | 0.13 | 98.87 | 98.53 | 0.30 |
| UNFHA | 92.47 | 81.44 | 9.35 | 0.57 | 1.21 | 0.39 | 98.54 | 98.33 | 0.23 |
| Level 0 | Squeezenet | AlexNet | |
| Level 1 | Vgg | ||
| Level 2 | ResNet152 | ResNeXt | wideResNet |
| Level 3 | ResNet50 | Densenet | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cabezas, M.; Kentsch, S.; Tomhave, L.; Gross, J.; Caceres, M.L.L.; Diez, Y. Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data. Remote Sens. 2020, 12, 3431. https://doi.org/10.3390/rs12203431
Cabezas M, Kentsch S, Tomhave L, Gross J, Caceres MLL, Diez Y. Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data. Remote Sensing. 2020; 12(20):3431. https://doi.org/10.3390/rs12203431
Chicago/Turabian StyleCabezas, Mariano, Sarah Kentsch, Luca Tomhave, Jens Gross, Maximo Larry Lopez Caceres, and Yago Diez. 2020. "Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data" Remote Sensing 12, no. 20: 3431. https://doi.org/10.3390/rs12203431
APA StyleCabezas, M., Kentsch, S., Tomhave, L., Gross, J., Caceres, M. L. L., & Diez, Y. (2020). Detection of Invasive Species in Wetlands: Practical DL with Heavily Imbalanced Data. Remote Sensing, 12(20), 3431. https://doi.org/10.3390/rs12203431