Agrobacterium tumefaciens-Mediated Genetic Transformation of Eclipta alba
Abstract
1. Introduction
2. Materials and Methods
2.1. Explant Source, Preparation and Culture Conditions
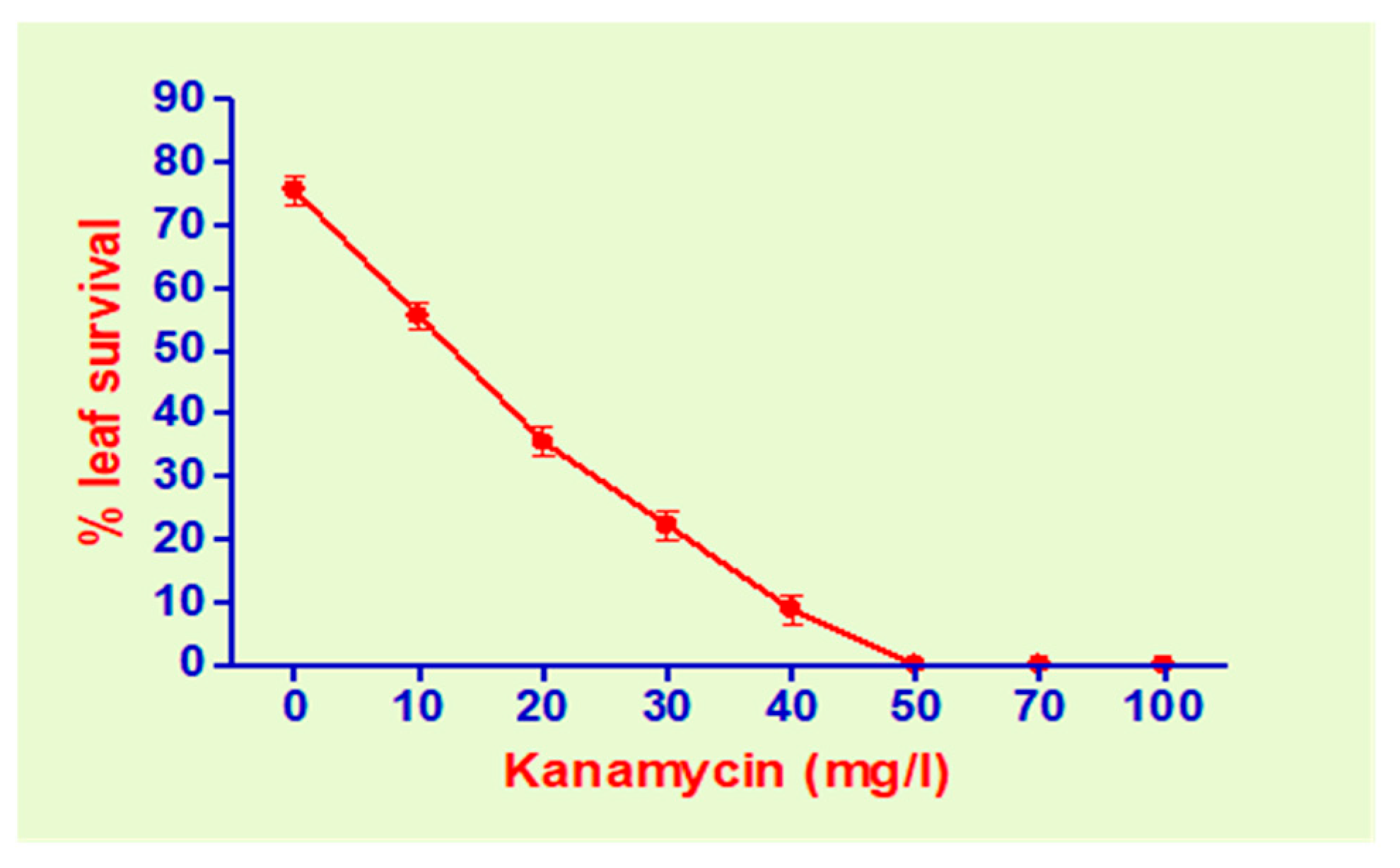
2.2. Agrobacterium Strain, Vector and Kanamycin Sensitivity Determination
2.3. Genetic Transformation
Co-Cultivation and Infection
2.4. Regeneration of Transformed Leaf Explants
2.5. Measurement of GUS Activity
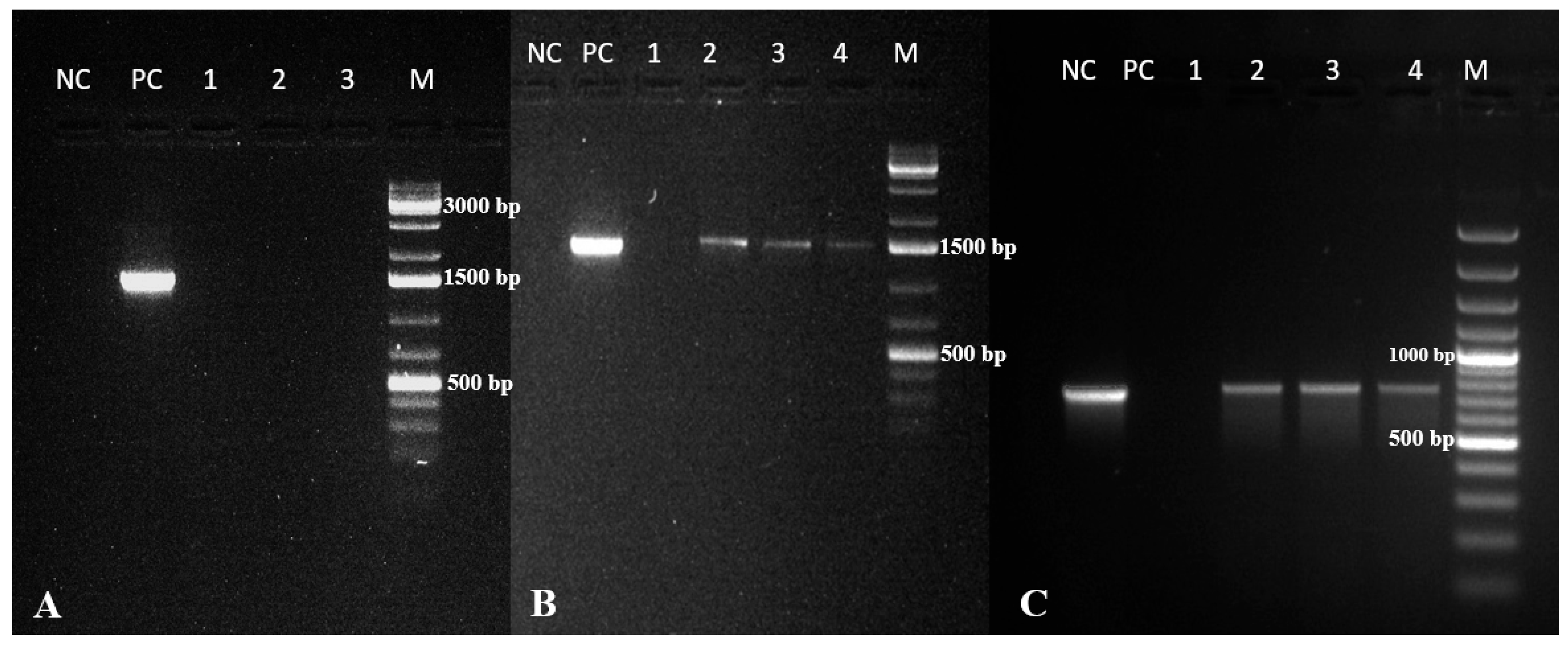
2.6. Molecular Analysis
2.7. Statistical Analysis
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Prasathkumar, M.; Anisha, S.; Dhrisya, C.; Becky, R.; Sadhasivam, S. Therapeutic and pharmacological efficacy of selective Indian medicinal plants—A review. Phytomedicine Plus 2021, 1, 100029. [Google Scholar] [CrossRef]
- Bhat, S.G. Medicinal Plants and Its Pharmacological Values. Natural Medicinal Plants; IntechOpen: London, UK, 2022. [Google Scholar] [CrossRef]
- Msomi, N.Z.; Simelane, B.C.M. Herbal Medicine. Herbal Medicine; IntechOpen: London, UK, 2019. [Google Scholar] [CrossRef]
- Atanasov, A.G.; Waltenberger, B.; Pferschy-Wenzig, E.M.; Linder, T.; Wawrosch, C.; Uhrin, P.; Temml, V.; Wang, L.; Schwaiger, S.; Heiss, E.H.; et al. Discovery and resupply of pharmacologically active plant-derived natural products: A review. Bi-Otechnol Adv. 2015, 33, 1582–1614. [Google Scholar] [CrossRef]
- Luqman, S.; Rizvi, S.I.; Beer, A.-M.; Khare, S.K.; Atukeren, P. Efficacy of herbal drugs in human diseases and disorders. Evidence-Based Complement. Altern. Med. 2014, 2014, 273676. [Google Scholar] [CrossRef] [PubMed]
- Uniyal, R.; Sandhu, S.; Chandok, J. Herbology. In the Ayurvedic Encyclopedia; Sri Satguru Publications: New Delhi India, 1998; p. 77. [Google Scholar]
- Aggarwal, D.; Datta, V.; Singh, R. Eclipta Alba (L.) Hassk.: An important medicinal plant for immunity and health. In Plants for immunity; Behl, R.K., Sharma, P.K., Arya, R.K., Chibbar, R.N., Eds.; Agrobios Publications: Jodhpur, India, 2022; pp. 181–191. [Google Scholar]
- Ma, R.; Yu, Z.; Cai, Q.; Li, H.; Dong, Y.; Oksman-Caldentey, K.-M.; Rischer, H. Agrobacterium-Mediated Genetic Transformation of the Medicinal Plant Veratrum dahuricum. Plants 2020, 9, 191. [Google Scholar] [CrossRef]
- Niazian, M. Application of genetics and biotechnology for improving medicinal plants. Planta 2019, 249, 953–973. [Google Scholar] [CrossRef] [PubMed]
- Bardar, S.; Kaul, V.K.; Kachhwaha, S.; Kothari, S. Nutrient optimization for improved in vitro plant regeneration in Eclipta alba (L.) Hassk. and assessment of genetic fidelity using RAPD analysis. Plant Tissue Cult. Biotechnol. 2014, 24, 223–234. [Google Scholar] [CrossRef]
- Dhaka, N.; Kothari, S.L. Micropropagation of Eclipta alba (L.) hassk—An important medicinal plant. Vitr. Cell. Dev. Biol. Plant 2005, 41, 658–661. [Google Scholar] [CrossRef]
- Prakash, P.; Sharumathy, D.; Sunkar, S.; Nandagopal, D.; Gopakumaran, N. Micropropagation of Eclipta alba using humic acid as media component. Plant Arch. 2015, 15, 181–185. [Google Scholar]
- Datta, V.; Sharma, L.; Aggarawal, D.; Sharma, A.K.; Dhama, K. Synergistic effect of plant growth regulators on micropropagation of Eclipta alba: A plant with diverse medicinal properties. J. Exp. Biol. Agric. Sci. 2022, 10, 1432–1440. [Google Scholar] [CrossRef]
- Murashige, T.; Skoog, F. A Revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: Beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef] [PubMed]
- Holsters, M.; De Waele, D.; Depicker, A.; Messens, E.; Van Montagu, M.; Schell, J. Transfection and transformation of Agro-bacterium tumefaciens. Mol. Gen Genet 1978, 163, 181–187. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.B.; Meng, L.S. Genetic transformation of Gentiana dahurica Fisch by Agrobacterium tumefaciens using zygotic embryo derived callus. Acta Physiol. Plant 2010, 32, 629–634. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, D.; Jaiswal, N.; Kumar, A.; Reddy, M.S. Factors affecting genetic transformation and shoot organogenesis of Bacopa monnieri (L.) Wettst. J. Plant Biochem Biotechnol 2013, 22, 382–391. [Google Scholar] [CrossRef]
- Gosal, S.S.; Wani, S.H. Plant genetic transformation and transgenic crops: Methods and applications. In Biotechnologies of Crop Improvement, Volume 2: Transgenic Approaches; Springer: Cham, Switzerland, 2018; pp. 1–23. [Google Scholar] [CrossRef]
- Gelvin, S.B. Agrobacterium-mediated plant transformation: The biology behind the ‘gene-jockeying’ tool. Microbiol Mol Biol Rev 2003, 67, 16–37. [Google Scholar] [CrossRef]
- Acanda, Y.; Canton, M.; Wu, H.; Zale, J. Kanamycin selection in temporary immersion bioreactors allows visual selection of transgenic citrus shoots. Plant Cell, Tissue Organ Cult. (PCTOC) 2017, 129, 351–357. [Google Scholar] [CrossRef]
- Aggarwal, D.; Kumar, A.; Reddy, M.S. Agrobacterium tumefaciens mediated genetic transformation of selected elite clone(s) of Eucalyptus tereticornis. Acta Physiol. Plant. 2011, 33, 1603–1611. [Google Scholar] [CrossRef]
- Sun, Z.-L.; Zhou, W.; Yan, J.-D.; Gao, Y.-R.; Li, X.-W.; Sun, J.-C.; Fang, K.-F.; Zhang, Q.; Xing, Y.; Qin, L.; et al. Agrobacterium-mediated genetic transformation of Chinese chestnut (Castanea mollissima Blume). Plant Cell Tissue Organ Cult. (PCTOC) 2020, 140, 95–103. [Google Scholar] [CrossRef]
- Agarie, S.; Umemoto, M.; Sunagawa, H.; Anai, T.; Cushman, J.C. An Agrobacterium-mediated transformation via organogenesis regeneration of a facultative CAM plant, the common ice plant Mesembryanthemum crystallinum L. Plant Prod. Sci. 2020, 23, 343–349. [Google Scholar] [CrossRef]
- Bhatt, R.; Asopa, P.P.; Jain, R.; Kothari-Chajer, A.; Kothari, S.L.; Kachhwaha, S. Optimization of Agrobacterium Mediated Genetic Transformation in Paspalum scrobiculatum L. (Kodo Millet). Agronomy 2021, 11, 1104. [Google Scholar] [CrossRef]
- Beyaz, R.; Aycan, M.; Yildiz, M. The effect of explant position on Agrobacterium tumefaciens-mediated gene transfer in flax (Linum usitatissimum L.). J. Biotechnol. 2017, 256S, S17–S43. [Google Scholar] [CrossRef]
- Sadhu, S.K.; Jogam, P.; Gande, K.; Banoth, R.; Penna, S.; Peddaboina, V. Optimization of different factors for an Agrobacterium-mediated genetic transformation system using embryo axis explants of chickpea (Cicer arietinum L.). J. Plant Biotechnol. 2022, 49, 61–73. [Google Scholar] [CrossRef]
- Subramoni, S.; Nathoo, N.; Klimov, E.; Yuan, Z.-C. Agrobacterium tumefaciens responses to plant-derived signaling molecules. Front. Plant Sci. 2014, 5, 322. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.-Y.; Gelvin, S.B. T-DNA Binary Vectors and Systems. Plant Physiol. 2008, 146, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Wu, E.; Zhao, Z.Y. Agrobacterium-mediated sorghum transformation. Methods Mol. Biol. 2017, 1669, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Utami, E.S.W.; Hariyanto, S.; Manuhara, Y.S.W. Agrobacterium tumefaciens-mediated transformation of Dendrobium lasianthera J.J.Sm: An important medicinal orchid. J. Genet. Eng. Biotechnol. 2018, 16, 703–709. [Google Scholar] [CrossRef]
- Manfroi, E.; Yamazaki-Lau, E.; Grando, M.F.; Roesler, E.A. Acetosyringone, pH and temperature effects on transient genetic transformation of immature embryos of Brazilian wheat genotypes by Agrobacterium tumefaciens. Genet. Mol. Biol. 2015, 38, 470–476. [Google Scholar] [CrossRef]
- Hwang, H.-H.; Yu, M.; Lai, E.-M. Agrobacterium-mediated plant transformation: Biology and applications. Arab. Book 2017, 15, e0186. [Google Scholar] [CrossRef]
- Song, S.; Yan, R.; Wang, C.; Wang, J.; Sun, H. Improvement of a Genetic transformation system and preliminary study on the function of LpABCB21 and LpPILS7 based on somatic embryogenesis in Lilium pumilum DC. Fisch. Int. J. Mol. Sci. 2020, 21, 6784. [Google Scholar] [CrossRef] [PubMed]
- Saini, S.; Sharma, I.; Kaur, N.; Pati, P.K. Auxin: A master regulator in plant root development. Plant Cell Rep. 2013, 32, 741–757. [Google Scholar] [CrossRef]
- Maheshwari, P.; Kovalchuk, I. Agrobacterium-mediated stable genetic transformation of populus angustifolia and populus balsamifera. Front. Plant Sci. 2016, 7, 296. [Google Scholar] [CrossRef]
- Cordeiro, D.; Alves, A.; Ferraz, R.; Casimiro, B.; Canhoto, J.; Correia, S. An efficient Agrobacterium-mediated genetic transformation method for Solanum betaceum Cav. embryogenic callus. Plants 2023, 12, 1202. [Google Scholar] [CrossRef] [PubMed]
- Zuker, A.; Ahroni, A.; Tzfira, T.; Ben-Meir, H.; Vainstein, A. Wounding by bombardment yields highly efficient Agrobacterium-mediated transformation of carnation (Dianthus caryophyllus L.). Mol. Breed. 1999, 5, 367–375. [Google Scholar] [CrossRef]
- Wang, J.; Sasse, A.; Sheridan, H. Traditional Chinese Medicine: From Aqueous Extracts to Therapeutic Formulae; Plant ex-tracts; IntachOpen: London, UK, 2019. [Google Scholar] [CrossRef]
- Tiwari, M.; Mishra, A.K.; Chakrabarty, D. Agrobacterium-mediated gene transfer: Recent advancements and layered immunity in plants. Planta 2022, 37, 256. [Google Scholar] [CrossRef]
- Li, Y.; Tang, D.; Liu, Z.; Chen, J.; Cheng, B.; Kumar, R.; Yer, H.; Li, Y. An Improved procedure for Agrobacterium-mediated transformation of ‘Carrizo’ citrange. Plants 2022, 11, 1457. [Google Scholar] [CrossRef] [PubMed]
- Mao, L.; Dai, Y.; Huang, Y.; Sun, H.; Li, Z.; Yang, B.; Zhang, Z.; Chen, W.; Ou, L.; Liu, Z.; et al. Effect of light intensity on gene expression in hypocotyl during the elongation in a leaf-yellowing mutant of pepper (Capsicum annuum L.). Agronomy 2022, 12, 2762. [Google Scholar] [CrossRef]

| Factor | Variable | % GUS Expression |
|---|---|---|
| A. tumefaciens strain | LBA 4404 | 52.3 ± 0.91 a |
| EHA 105 | 48.5 ± 0.4 b | |
| Pre-culture | 0 d | 36.8 ± 0.35 f |
| 1 d | 45.7 ± 0.20 d | |
| 2 d | 56.8 ± 0.26 a | |
| 3 d | 51.2 ± 0.2 b | |
| 4 d | 48.3 ± 0.20 c | |
| 5 d | 43.7 ± 0.28 e | |
| Method of injury | Intact | 36.7 ± 0.15 e |
| With hypodermic needle | 55.6 ± 0.36 a | |
| With surgical blade | 48.5 ± 0.1 b | |
| With carborundum | 44.5 ± 0.35 c | |
| With glass beads | 40.1 ± 0.35 d | |
| Acetosyringone | 0 | 43.5 ± 0.20 d |
| 100 | 58.6 ± 0.2 b | |
| 150 | 63.4 ± 0.25 a | |
| 200 | 57.4 ± 0.15 c | |
| pH of co-cultivation medium | 5.2 | 51.4 ± 0.25 d |
| 5.4 | 58.6 ± 0.15 a | |
| 5.6 | 53.3 ± 0.3 b | |
| 5.8 | 52.8 ± 0.2 c | |
| Optical density O.D Value | 0.2 | 34.6 ± 0.25 e |
| 0.4 | 42.6 ± 0.25 d | |
| 0.6 | 56.3 ± 0.20 a | |
| 0.8 | 53.8 ± 0.2 b | |
| 1.0 | 49.8 ± 0.15 c | |
| Co-cultivation period | 1 d | 43.5 ± 0.30 d |
| 2 d | 54.6 ± 0.25 a | |
| 3 d | 49.3 ± 0.25 b | |
| 4 d | 46.2 ± 0.37 c | |
| 5 d | 43.1 ± 0.26 d | |
| Infection Time | 5 min | 42.4 ± 0.3 c |
| 10 min | 46.6 ± 0.25 b | |
| 15 min | 53.3 ± 0.26 a | |
| 20 min | 42.3 ± 0.15 c | |
| 30 min | 38.8 ± 0.20 d | |
| Photoperiod | 24 h light | 46.3 ± 0.36 c |
| 24 h dark | 51.7 ± 0.36 b | |
| 16 h light/8 h dark | 56.5 ± 0.35 a |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aggarwal, D.; Datta, V.; Tuli, H.S.; Kumar, P.; Ramniwas, S. Agrobacterium tumefaciens-Mediated Genetic Transformation of Eclipta alba. Int. J. Plant Biol. 2024, 15, 641-651. https://doi.org/10.3390/ijpb15030047
Aggarwal D, Datta V, Tuli HS, Kumar P, Ramniwas S. Agrobacterium tumefaciens-Mediated Genetic Transformation of Eclipta alba. International Journal of Plant Biology. 2024; 15(3):641-651. https://doi.org/10.3390/ijpb15030047
Chicago/Turabian StyleAggarwal, Diwakar, Vasudha Datta, Hardeep Singh Tuli, Pawan Kumar, and Seema Ramniwas. 2024. "Agrobacterium tumefaciens-Mediated Genetic Transformation of Eclipta alba" International Journal of Plant Biology 15, no. 3: 641-651. https://doi.org/10.3390/ijpb15030047
APA StyleAggarwal, D., Datta, V., Tuli, H. S., Kumar, P., & Ramniwas, S. (2024). Agrobacterium tumefaciens-Mediated Genetic Transformation of Eclipta alba. International Journal of Plant Biology, 15(3), 641-651. https://doi.org/10.3390/ijpb15030047







