Antibacterial Effect of Sapindus mukorossi Aqueous Extract in Human Saliva—A Pilot Translational Study with an Ex Vivo Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of SMa
2.2. Ethics Statement and Patient Recruitment
2.3. Saliva Collection and Ex Vivo Incubation
2.4. Monitoring Oral Bacterial Growth
2.5. Salivary Microbiome Analysis
2.6. Statistical Analysis
3. Results
3.1. Ex Vivo Monitoring of Bacterial Growth
3.2. Salivary Metagenome Analysis
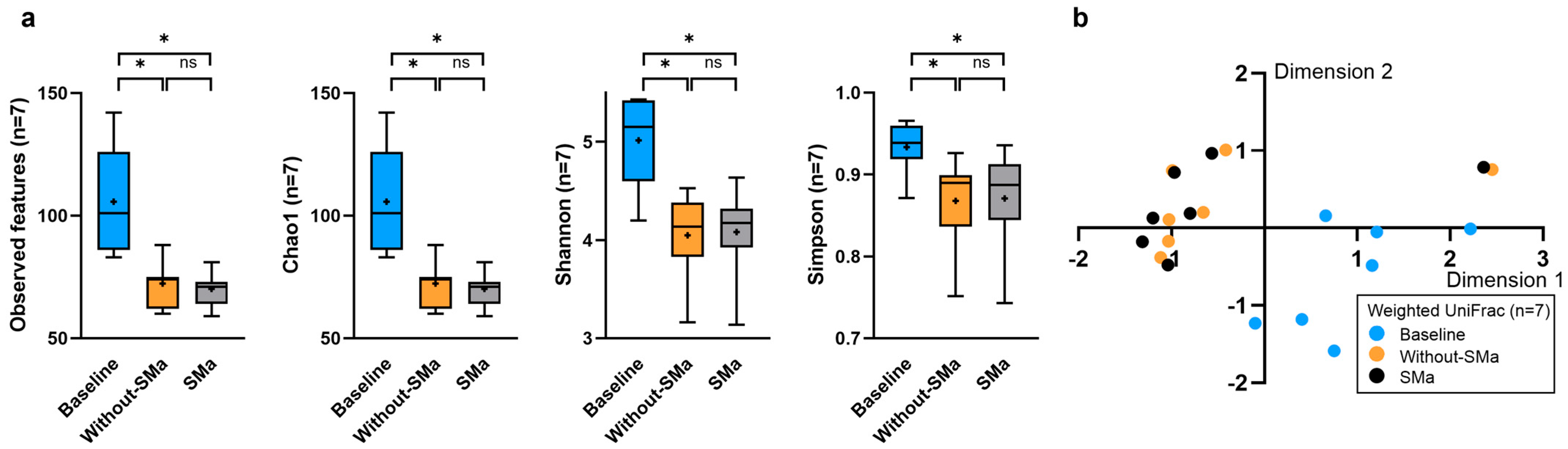
3.3. Alpha and Beta Diversity
3.4. Shifts in the Salivary Microbiome
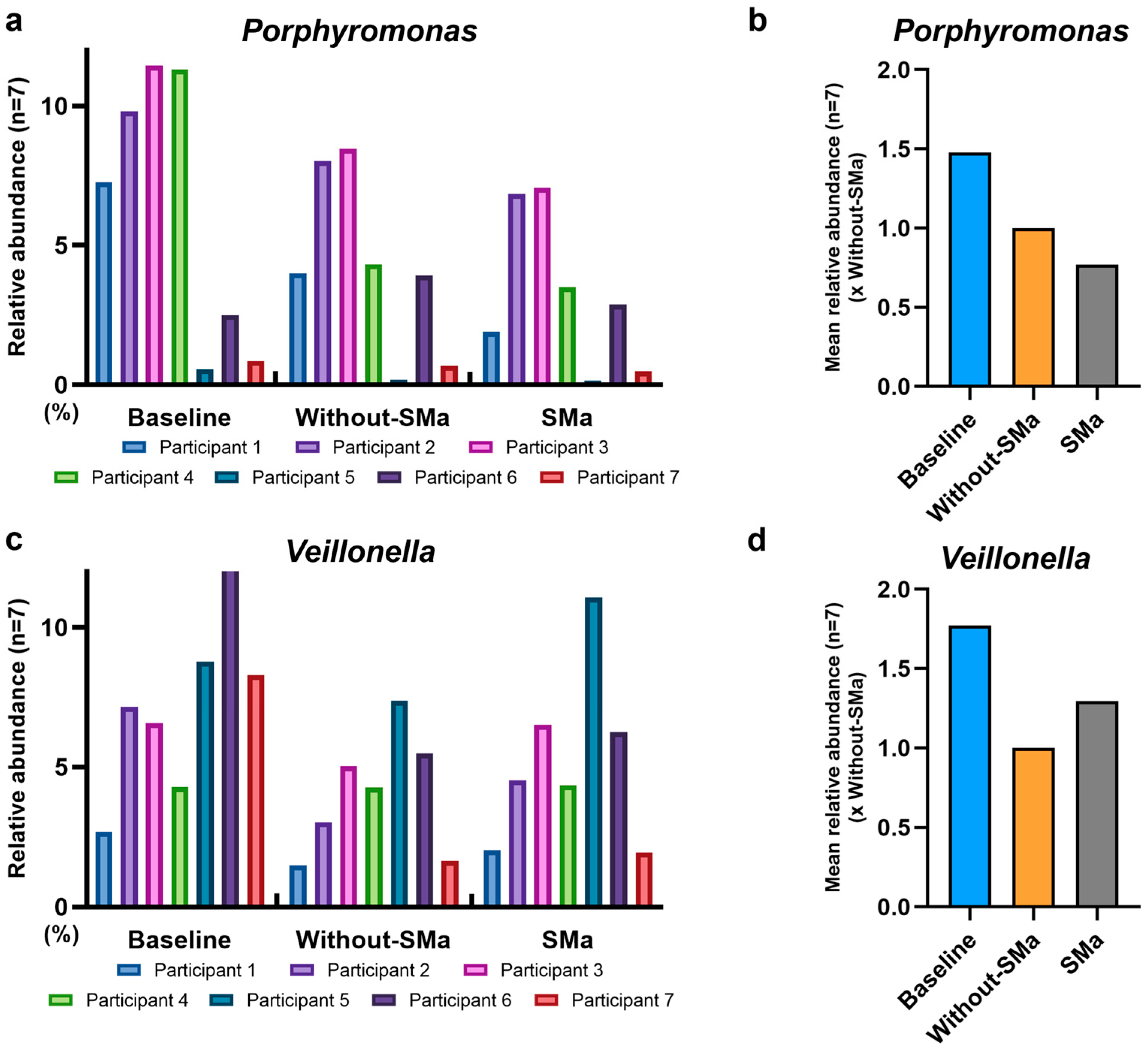
3.5. SMa Modulation of Porphyromonas and Veillonella
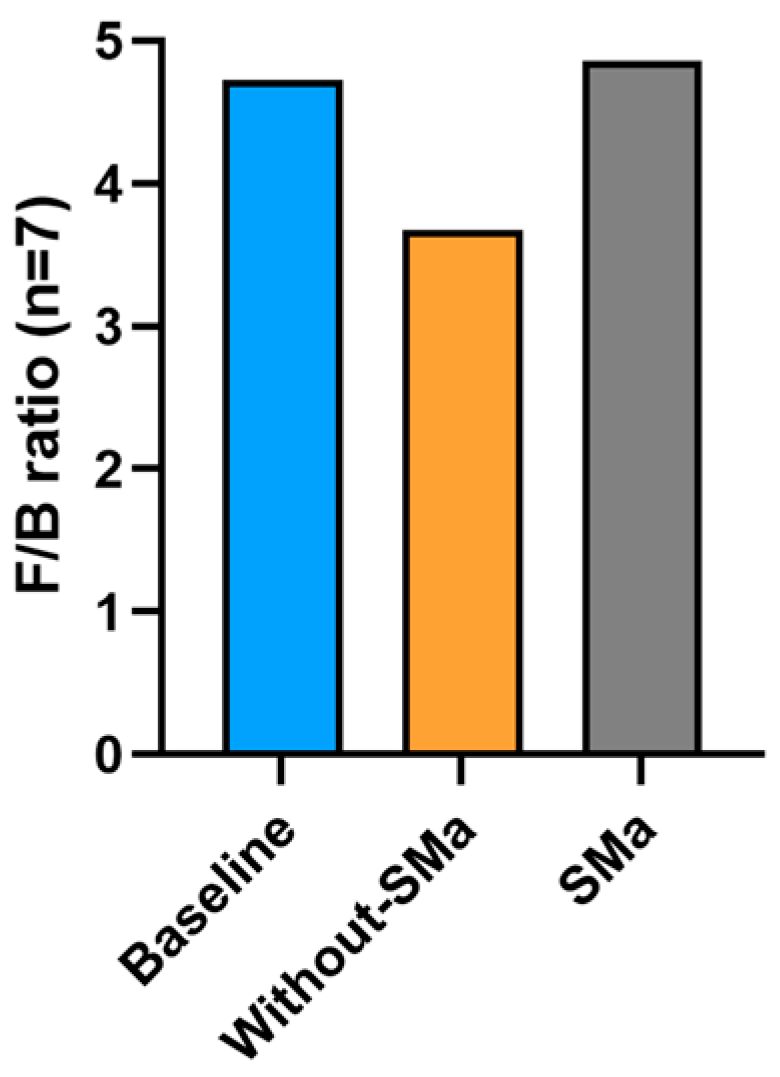
3.6. F/B Ratio
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Sm | Sapindus mukorossi |
| SMa | Sapindus mukorossi seed aqueous extract |
| ddH2O | Double-distilled water |
| OD | Optical density |
| NGS | Next generation sequencing |
| PERMANOVA | Permutational multivariate analysis of variance |
| F/B ratio | Firmicutes/Bacteroidetes ratio |
| OD600 | Optical density at 600 nm |
| ns | Not statistically significant |
| NMDS | Non-metric multidimensional scaling |
| Mn | Mean |
| H2O2 | Hydrogen peroxide |
| Hag1 | Hemaglutinin 1 |
| CHX | Chlorhexidine gluconate |
| CPC | Cetylpyridinium chloride |
References
- Sochacki, M.; Vogt, O. Triterpenoid Saponins from Washnut (Sapindus mukorossi Gaertn.)—A source of natural surfactants and other active components. Plants 2022, 11, 2355. [Google Scholar] [CrossRef]
- Upadhyay, A.; Singh, D.K. Pharmacological effects of Sapindus mukorossi. Revista do Instituto de Medicina Tropical de São Paulo 2012, 54, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Suhagia, B.N.; Rathod, I.S.; Sindhu, S. Sapindus mukorossi (Areetha): An overview. Int. J. Pharm. Sci. Res. 2011, 2, 1905. [Google Scholar]
- Ibrahim, M.; Khan, A.A.; Tiwari, S.K.; Habeeb, M.A.; Khaja, M.N.; Habibullah, C.M. Antimicrobial activity of Sapindus mukorossi and Rheum emodi extracts against H pylori: In vitro and in vivo studies. World J. Gastroenterol. 2006, 12, 7136. [Google Scholar] [CrossRef] [PubMed]
- Shiu, S.T.; Lew, W.Z.; Lee, S.Y.; Feng, S.W.; Huang, H.M. Effects of Sapindus mukorossi seed oil on proliferation, osteogenetic/odontogenetic differentiation and matrix vesicle secretion of human dental pulp mesenchymal stem cells. Materials 2020, 13, 4063. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.K.; Wu, Y.F.; Chang, W.J.; Feng, S.W.; Huang, H.M. The treatment efficiency and microbiota analysis of Sapindus mukorossi seed oil on the ligature-induced periodontitis rat model. Int. J. Mol. Sci. 2022, 23, 8560. [Google Scholar] [CrossRef]
- Alshahrani, A.; Alhamdan, E.M.; Alsahhaf, A.; Shafqat, S.; Kumari, U.; Alyahya, A.M.; Abduljabbar, T. Influence of contemporary ceramo-metallic crown decontamination on the surface roughness, color stability, and disinfection. An in vitro analysis. Photodiagnosis Photodyn. Ther. 2023, 41, 103256. [Google Scholar] [CrossRef]
- Hariharan, A.; Krishnamurthi, S.; Thamarai, C.; Thanya, K.; Parameswari, D.; Krithika, A. Sapindus mukurossi–An Effective Biocleanser for Removable Dental Prostheses? An In vitro Study. J. Pharm. Bioallied Sci. 2024, 16, S1154–S1160. [Google Scholar] [CrossRef]
- Saglik, İ.; Tuncay, Ö.G.; Özhak, B. Investigation of the antimicrobial effects of Sapindus mukorossi on endodontic pathogens. J. Exp. Clin. Med. 2020, 37, 111–118. [Google Scholar]
- Kaliamoorthy, S.; Rayar, S.; SundarRaj, S.; Sayeeram, S.P.; Premkumar, V.; Muddappa, S.C.; Muthukumaran, V.; Raju, K.; Samidorai, A.; Sreeram, R. Comparative Evaluation of Dental Pulp Tissue Dissolution Ability of Sapindus mukorossi and Sodium Hypochlorite. Cureus 2024, 16, e51820. [Google Scholar] [CrossRef]
- Tahir, A.; Ur-Rehman Qazi, F.; Choudhry, Z.; Musheer, U.; Amin, M.; Malik, S.; AlMokhatieb, A.; Almadi, K.; Alkahtany, M.; Ahmed, M. Influence of Sapindus mukorossi extract in comparison to 17% EDTA as final root canal irrigant on the sealer penetration and microleakage of dentinal tubules. Eur. Rev. Med. Pharmacol. Sci. 2023, 27, 2724–2732. [Google Scholar]
- Aneja, K.R.; Joshi, R.; Sharma, C. In vitro antimicrobial activity of Sapindus mukorossi and Emblica officinalis against dental caries pathogens. Ethnobot. Leafl. 2010, 2010, 3. [Google Scholar]
- Alasqah, M.N. Efficacy of Adjunctive Fotoenticine Photodynamic Therapy and Sapindus mukorossi Therapy on Clinical, Radiographic, and Cytokine Profile of Diabetics with Peri-Implantitis. Photobiomodul. Photomed. Laser Surg. 2024, 42, 174–181. [Google Scholar] [CrossRef]
- Li, L.; Wei, M.-P.; Yu, H.; Xie, Y.-F.; Guo, Y.-H.; Cheng, Y.-L.; Yao, W.-R. Antifungal activity of Sapindus saponins against Candida albicans: Interruption of biofilm formation. J. Herb. Med. 2023, 42, 100776. [Google Scholar] [CrossRef]
- Yang, Y.-H.; Yu, J.-J.; Han, H.-Y.; Chang, W.-M.; Wang, C.-W. Ex-vivo investigation of human salivary microbial growth with lysogeny broth for translational research–A pilot study. J. Dent. Sci. 2025, 20, 437–443. [Google Scholar] [CrossRef]
- Benahmed, A.G.; Gasmi, A.; Doşa, A.; Chirumbolo, S.; Mujawdiya, P.K.; Aaseth, J.; Dadar, M.; Bjørklund, G. Association between the gut and oral microbiome with obesity. Anaerobe 2021, 70, 102248. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-C.; Nien, C.-J.; Chen, L.-G.; Huang, K.-Y.; Chang, W.-J.; Huang, H.-M. Effects of Sapindus mukorossi seed oil on skin wound healing: In vivo and in vitro testing. Int. J. Mol. Sci. 2019, 20, 2579. [Google Scholar] [CrossRef]
- Selvaraj, A.; Valliammai, A.; Premika, M.; Priya, A.; Bhaskar, J.P.; Krishnan, V.; Pandian, S.K. Sapindus mukorossi Gaertn. and its bioactive metabolite oleic acid impedes methicillin-resistant Staphylococcus aureus biofilm formation by down regulating adhesion genes expression. Microbiol. Res. 2021, 242, 126601. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.P.; Yu, H.; Guo, Y.H.; Cheng, Y.L.; Xie, Y.F.; Yao, W.R. Synergistic combination of Sapindoside A and B: A novel antibiofilm agent against Cutibacterium acnes. Microbiol. Res. 2022, 254, 126912. [Google Scholar] [CrossRef]
- Wei, M.-P.; Yu, H.; Guo, Y.-H.; Cheng, Y.-L.; Xie, Y.-F.; Yao, W.-R. Potent in vitro synergistic antibacterial activity of natural amphiphilic Sapindoside A and B against Cutibacterium acnes with destructive effect on bacterial membrane. Biochim. Biophys. Acta Biomembr. 2021, 1863, 183699. [Google Scholar] [CrossRef] [PubMed]
- Abusleme, L.; Dupuy, A.K.; Dutzan, N.; Silva, N.; Burleson, J.A.; Strausbaugh, L.D.; Gamonal, J.; Diaz, P.I. The subgingival microbiome in health and periodontitis and its relationship with community biomass and inflammation. ISME J. 2013, 7, 1016–1025. [Google Scholar] [CrossRef]
- Oyetola, E.O.; Awosusi, O.O.; Agho, E.T.; Abdullahi, M.A.; Suleiman, I.K.; Egunjobi, S. Salivary Bacterial Count and its Implications on the Prevalence of Oral Conditions. J. Contemp. Dent. Pract. 2019, 20, 184–189. [Google Scholar] [CrossRef]
- Sakamoto, Y.; Tanabe, A.; Moriyama, M.; Otsuka, Y.; Funahara, M.; Soutome, S.; Umeda, M.; Kojima, Y. Number of bacteria in saliva in the perioperative period and factors associated with increased numbers. Int. J. Environ. Res. Public Health 2022, 19, 7552. [Google Scholar] [CrossRef]
- Tohara, T.; Kikutani, T.; Tamura, F.; Yoshida, M.; Kuboki, T. Multicentered epidemiological study of factors associated with total bacterial count in the saliva of older people requiring nursing care. Geriatr. Gerontol. Int. 2017, 17, 219–225. [Google Scholar] [CrossRef]
- Myers, J.A.; Curtis, B.S.; Curtis, W.R. Improving accuracy of cell and chromophore concentration measurements using optical density. BMC Biophys. 2013, 6, 4. [Google Scholar] [CrossRef]
- Beal, J.; Farny, N.G.; Haddock-Angelli, T.; Selvarajah, V.; Baldwin, G.S.; Buckley-Taylor, R.; Gershater, M.; Kiga, D.; Marken, J.; Sanchania, V. Robust estimation of bacterial cell count from optical density. Commun. Biol. 2020, 3, 512. [Google Scholar] [CrossRef] [PubMed]
- Montesinos-Cruz, V.; Somerville, G.A. Shining a Light on Spectrophotometry in Bacteriology. Antibiotics 2024, 13, 1164. [Google Scholar] [CrossRef]
- Jun, S.; Si, F.; Pugatch, R.; Scott, M. Fundamental principles in bacterial physiology—History, recent progress, and the future with focus on cell size control: A review. Rep. Prog. Phys. 2018, 81, 056601. [Google Scholar] [CrossRef] [PubMed]
- Nayfach, S.; Pollard, K.S. Average genome size estimation improves comparative metagenomics and sheds light on the functional ecology of the human microbiome. Genome Biol. 2015, 16, 51. [Google Scholar] [CrossRef] [PubMed]
- Charbon, G.; Mendoza-Chamizo, B.; Campion, C.; Li, X.; Jensen, P.R.; Frimodt-Møller, J.; Løbner-Olesen, A. Energy starvation induces a cell cycle arrest in Escherichia coli by triggering degradation of the DnaA initiator protein. Front. Mol. Biosci. 2021, 8, 629953. [Google Scholar] [CrossRef]
- Kuong, K.J.; Kuzminov, A. Disintegration of nascent replication bubbles during thymine starvation triggers RecA-and RecBCD-dependent replication origin destruction. J. Biol. Chem. 2012, 287, 23958–23970. [Google Scholar] [CrossRef] [PubMed]
- Münch, K.M.; Müller, J.; Wienecke, S.; Bergmann, S.; Heyber, S.; Biedendieck, R.; Münch, R.; Jahn, D. Polar fixation of plasmids during recombinant protein production in Bacillus megaterium results in population heterogeneity. Appl. Environ. Microbiol. 2015, 81, 5976–5986. [Google Scholar] [CrossRef]
- Janissen, R.; Arens, M.M.; Vtyurina, N.N.; Rivai, Z.; Sunday, N.D.; Eslami-Mossallam, B.; Gritsenko, A.A.; Laan, L.; de Ridder, D.; Artsimovitch, I. Global DNA compaction in stationary-phase bacteria does not affect transcription. Cell 2018, 174, 1188–1199.e1114. [Google Scholar] [CrossRef]
- Relvas, M.; Regueira-Iglesias, A.; Balsa-Castro, C.; Salazar, F.; Pacheco, J.J.; Cabral, C.; Henriques, C.; Tomás, I. Relationship between dental and periodontal health status and the salivary microbiome: Bacterial diversity, co-occurrence networks and predictive models. Sci. Rep. 2021, 11, 929. [Google Scholar] [CrossRef]
- Yasunaga, H.; Takeshita, T.; Shibata, Y.; Furuta, M.; Shimazaki, Y.; Akifusa, S.; Ninomiya, T.; Kiyohara, Y.; Takahashi, I.; Yamashita, Y. Exploration of bacterial species associated with the salivary microbiome of individuals with a low susceptibility to dental caries. Clin. Oral. Investig. 2017, 21, 2399–2406. [Google Scholar] [CrossRef]
- Yang, C.Y.; Yeh, Y.M.; Yu, H.Y.; Chin, C.Y.; Hsu, C.W.; Liu, H.; Huang, P.J.; Hu, S.N.; Liao, C.T.; Chang, K.P. Oral microbiota community dynamics associated with oral squamous cell carcinoma staging. Front. Microbiol. 2018, 9, 862. [Google Scholar] [CrossRef]
- Lombardo Bedran, T.B.; Marcantonio, R.A.C.; Spin Neto, R.; Alves Mayer, M.P.; Grenier, D.; Spolidorio, L.C.; Spolidorio, D.P. Porphyromonas endodontalis in chronic periodontitis: A clinical and microbiological cross-sectional study. J. Oral Microbiol. 2012, 4, 10123. [Google Scholar] [CrossRef]
- Albano, C.; Bagarello, S.; Giordano, S.; Sanfilippo, M.F.; Comparato, C.; Scardino, G.; Garbo, V.; Boncori, G.; Condemi, A.; Cascio, A. Granulicatella spp., a causative agent of infective endocarditis in children. Pathogens 2022, 11, 1431. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Manoil, D.; Belibasakis, G.N.; Kotsakis, G.A. Veillonellae: Beyond bridging species in oral biofilm ecology. Front. Oral Health 2021, 2, 774115. [Google Scholar] [CrossRef] [PubMed]
- Magne, F.; Gotteland, M.; Gauthier, L.; Zazueta, A.; Pesoa, S.; Navarrete, P.; Balamurugan, R. The Firmicutes/Bacteroidetes ratio: A relevant marker of gut dysbiosis in obese patients? Nutrients 2020, 12, 1474. [Google Scholar] [CrossRef]
- Li, W.; Ma, Z. FBA ecological guild: Trio of Firmicutes-Bacteroidetes alliance against Actinobacteria in human oral microbiome. Sci. Rep. 2020, 10, 287. [Google Scholar] [CrossRef]
- Mao, X.; Hiergeist, A.; Auer, D.L.; Scholz, K.J.; Muehler, D.; Hiller, K.A.; Maisch, T.; Buchalla, W.; Hellwig, E.; Gessner, A. Ecological effects of daily antiseptic treatment on microbial composition of saliva-grown microcosm biofilms and selection of resistant phenotypes. Front. Microbiol. 2022, 13, 934525. [Google Scholar] [CrossRef]
- Yeon, L.S.; Young, L.S. Susceptibility of oral Streptococci to chlorhexidine and cetylpyridinium chloride. Biocontrol Sci. 2019, 24, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Bartsch, S.; Kohnert, E.; Kreutz, C.; Woelber, J.P.; Anderson, A.; Burkhardt, A.S.; Hellwig, E.; Buchalla, W.; Hiller, K.A.; Ratka-Krueger, P. Chlorhexidine digluconate mouthwash alters the oral microbial composition and affects the prevalence of antimicrobial resistance genes. Front. Microbiol. 2024, 15, 1429692. [Google Scholar] [CrossRef]
- Marchi, A.P.; Farrel Côrtes, M.; Vásconez Noguera, S.; Rossi, F.; Levin, A.S.; Figueiredo Costa, S.; Perdigão Neto, L.V. Chlorhexidine susceptibility and Eagle effect in planktonic cells and biofilm of nosocomial isolates. Eur. J. Clin. Microbiol. Infect. Dis. 2023, 42, 787–792. [Google Scholar] [CrossRef]
- Mao, X.; Auer, D.L.; Buchalla, W.; Hiller, K.A.; Maisch, T.; Hellwig, E.; Al-Ahmad, A.; Cieplik, F. Cetylpyridinium chloride: Mechanism of action, antimicrobial efficacy in biofilms, and potential risks of resistance. Antimicrob. Agents Chemother. 2020, 64. [Google Scholar] [CrossRef] [PubMed]
- Bescos, R.; Ashworth, A.; Cutler, C.; Brookes, Z.L.; Belfield, L.; Rodiles, A.; Casas-Agustench, P.; Farnham, G.; Liddle, L.; Burleigh, M. Effects of chlorhexidine mouthwash on the oral microbiome. Sci. Rep. 2020, 10, 5254. [Google Scholar] [CrossRef] [PubMed]
- Choi, A.; Dong, K.; Williams, E.; Pia, L.; Batagower, J.; Bending, P.; Shin, I.; Peters, D.I.; Kaspar, J.R. Human saliva modifies growth, biofilm architecture, and competitive behaviors of oral streptococci. Msphere 2024, 9, e0077123. [Google Scholar] [CrossRef]
- Singh, R.; Kumari, N. Comparative determination of phytochemicals and antioxidant activity from leaf and fruit of Sapindus mukorrossi Gaertn.–A valuable medicinal tree. Ind. Crops Prod. 2015, 73, 1–8. [Google Scholar] [CrossRef]
- Uy, N.P.; Lee, H.D.; Ku, J.; Choi, K.; Lee, S. UPLC-QTOF-ESI-MS/MS-Based Comparative Study of Phytochemicals in Sapindus mukorossi. Horticulturae 2025, 11, 682. [Google Scholar] [CrossRef]
- Huang, H.C.; Liao, S.C.; Chang, F.R.; Kuo, Y.H.; Wu, Y.C. Molluscicidal saponins from Sapindus mukorossi, inhibitory agents of golden apple snails, Pomacea canaliculata. J. Agric. Food Chem. 2003, 51, 4916–4919. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.-H.; Yu, J.-J.; Chang, W.-M.; Huang, H.-M.; Wang, C.-W. Antibacterial Effect of Sapindus mukorossi Aqueous Extract in Human Saliva—A Pilot Translational Study with an Ex Vivo Model. Microbiol. Res. 2025, 16, 230. https://doi.org/10.3390/microbiolres16110230
Yang Y-H, Yu J-J, Chang W-M, Huang H-M, Wang C-W. Antibacterial Effect of Sapindus mukorossi Aqueous Extract in Human Saliva—A Pilot Translational Study with an Ex Vivo Model. Microbiology Research. 2025; 16(11):230. https://doi.org/10.3390/microbiolres16110230
Chicago/Turabian StyleYang, Yu-Hsin, Jing-Jie Yu, Wei-Min Chang, Haw-Ming Huang, and Chin-Wei Wang. 2025. "Antibacterial Effect of Sapindus mukorossi Aqueous Extract in Human Saliva—A Pilot Translational Study with an Ex Vivo Model" Microbiology Research 16, no. 11: 230. https://doi.org/10.3390/microbiolres16110230
APA StyleYang, Y.-H., Yu, J.-J., Chang, W.-M., Huang, H.-M., & Wang, C.-W. (2025). Antibacterial Effect of Sapindus mukorossi Aqueous Extract in Human Saliva—A Pilot Translational Study with an Ex Vivo Model. Microbiology Research, 16(11), 230. https://doi.org/10.3390/microbiolres16110230






