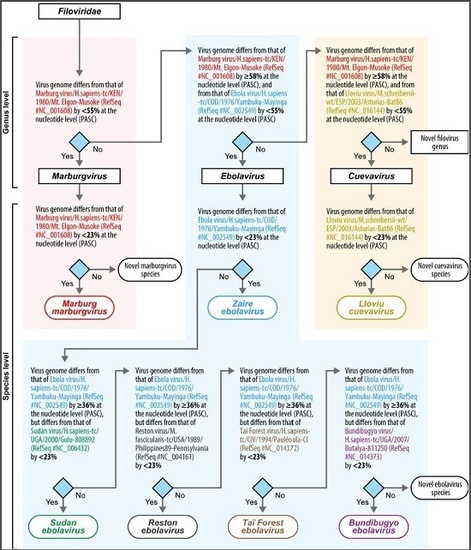
Implementation of Objective PASC-Derived Taxon Demarcation Criteria for Official Classification of Filoviruses
Abstract
:1. Introduction
2. Materials and Methods
3. Results
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Amarasinghe, G.K.; Bào, Y.; Basler, C.F.; Bavari, S.; Beer, M.; Bejerman, N.; Blasdell, K.R.; Bochnowski, A.; Briese, T.; Bukreyev, A.; et al. Taxonomy of the order Mononegavirales: Update 2017. Arch. Virol. 2017, 162. [Google Scholar] [CrossRef] [PubMed]
- Bukreyev, A.A.; Chandran, K.; Dolnik, O.; Dye, J.M.; Ebihara, H.; Leroy, E.M.; Mühlberger, E.; Netesov, S.V.; Patterson, J.L.; Paweska, J.T.; et al. Discussions and decisions of the 2012–2014 International Committee on Taxonomy of Viruses (ICTV) Filoviridae Study Group, January 2012–June 2013. Arch. Virol. 2014, 159, 821–830. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.H.; Becker, S.; Ebihara, H.; Geisbert, T.W.; Jahrling, P.B.; Kawaoka, Y.; Netesov, S.V.; Nichol, S.T.; Peters, C.J.; Volchkov, V.E.; et al. Family Filoviridae. In Virus Taxonomy—Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier/Academic Press: London, UK, 2011; pp. 665–671. [Google Scholar]
- Kuhn, J.H.; Becker, S.; Ebihara, H.; Geisbert, T.W.; Johnson, K.M.; Kawaoka, Y.; Lipkin, W.I.; Negredo, A.I.; Netesov, S.V.; Nichol, S.T.; et al. Proposal for a revised taxonomy of the family Filoviridae: Classification, names of taxa and viruses, and virus abbreviations. Arch. Virol. 2010, 155, 2083–2103. [Google Scholar] [CrossRef] [PubMed]
- Kiley, M.P.; Bowen, E.T.W.; Eddy, G.A.; Isaäcson, M.; Johnson, K.M.; McCormick, J.B.; Murphy, F.A.; Pattyn, S.R.; Peters, D.; Prozesky, O.W.; et al. Filoviridae: A taxonomic home for Marburg and Ebola viruses? Intervirology 1982, 18, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Francki, R.I.B.; Fauquet, C.M.; Knudson, D.L.; Brown, F. Classification and Nomenclature of Viruses—Fifth Report of the International Committee on Taxonomy of Viruses; Springer-Verlag: Vienna, Austria, 1991; Volume 2. [Google Scholar]
- Jahrling, P.B.; Kiley, M.P.; Klenk, H.-D.; Peters, C.J.; Sanchez, A.; Swanepoel, R. Family Filoviridae. In Virus Taxonomy—Sixth Report of the International Committee on Taxonomy of Viruses; Murphy, F.A., Fauquet, C.M., Bishop, D.H.L., Ghabrial, S.A., Jarvis, A.W., Martelli, G.P., Mayo, M.A., Summers, M.D., Eds.; Springer-Verlag: Vienna, Austria, 1995; Volume 10, pp. 289–292. [Google Scholar]
- Netesov, S.V.; Feldmann, H.; Jahrling, P.B.; Klenk, H.-D.; Sanchez, A. Family Filoviridae. In Virus Taxonomy—Seventh Report of the International Committee on Taxonomy of Viruses; van Regenmortel, M.H.V., Fauquet, C.M., Bishop, D.H.L., Carstens, E.B., Estes, M.K., Lemon, S.M., Maniloff, J., Mayo, M.A., McGeoch, D.J., Pringle, C.R., Eds.; Academic Press: San Diego, CA, USA, 2000; pp. 539–548. [Google Scholar]
- Feldmann, H.; Geisbert, T.W.; Jahrling, P.B.; Klenk, H.-D.; Netesov, S.V.; Peters, C.J.; Sanchez, A.; Swanepoel, R.; Volchkov, V.E. Family Filoviridae. In Virus Taxonomy—Eighth Report of the International Committee on Taxonomy of Viruses; Fauquet, C.M., Mayo, M.A., Maniloff, J., Desselberger, U., Ball, L.A., Eds.; Elsevier/Academic Press: San Diego, CA, USA, 2005; pp. 645–653. [Google Scholar]
- Kuhn, J.H.; Andersen, K.G.; Bào, Y.; Bavari, S.; Becker, S.; Bennett, R.S.; Bergman, N.H.; Blinkova, O.; Bradfute, S.; Brister, J.R.; et al. Filovirus RefSeq entries: Evaluation and selection of filovirus type variants, type sequences, and names. Viruses 2014, 6, 3663–3682. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Chetvernin, V.; Tatusova, T. PAirwise Sequence Comparison (PASC) and its application in the classification of filoviruses. Viruses 2012, 4, 1318–1327. [Google Scholar] [CrossRef] [PubMed]
- Lauber, C.; Gorbalenya, A.E. Genetics-based classification of filoviruses calls for expanded sampling of genomic sequences. Viruses 2012, 4, 1425–1437. [Google Scholar] [CrossRef] [PubMed]
- Ladner, J.T.; Beitzel, B.; Chain, P.S.G.; Davenport, M.G.; Donaldson, E.F.; Frieman, M.; Kugelman, J.R.; Kuhn, J.H.; O’Rear, J.; Sabeti, P.C.; et al. Standards for sequencing viral genomes in the era of high-throughput sequencing. MBio 2014, 5, e01360-14. [Google Scholar] [CrossRef] [PubMed]
- Ladner, J.T.; Kuhn, J.H.; Palacios, G. Standard finishing categories for high-throughput sequencing of viral genomes. Rev. Sci. Tech. 2016, 35, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Simmonds, P.; Adams, M.J.; Benkő, M.; Breitbart, M.; Brister, J.R.; Carstens, E.B.; Davison, A.J.; Delwart, E.; Gorbalenya, A.E.; Harrach, B.Z.; et al. Consensus statement: Virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 2017, 15, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Chetvernin, V.; Tatusova, T. Improvements to pairwise sequence comparison (PASC): A genome-based web tool for virus classification. Arch. Virol. 2014, 159, 3293–3304. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Kapustin, Y.; Tatusova, T. Virus classification by PAirwise Sequence Comparison (PASC). In Encyclopedia of Virology, 3rd ed.; Mahy, B.W.J., van Regenmortel, M.H.V., Eds.; Elsevier: Oxford, UK, 2008; Volume 5, pp. 342–348. [Google Scholar]
- Bào, Y.; Kuhn, J.H. Preliminary classification of novel hemorrhagic fever-causing viruses using sequence-based PAirwise Sequence Comparison (PASC) analysis. In Hemorrhagic Fever Viruses: Methods and Protocols; Salvato, M.S., Ed.; Humana Press: Totowa, NJ, USA, 2017; in press. [Google Scholar]
- Brister, J.R.; Ako-Adjei, D.; Bao, Y.; Blinkova, O. NCBI viral genomes resource. Nucleic Acids Res. 2015, 43, D571–D577. [Google Scholar] [CrossRef] [PubMed]
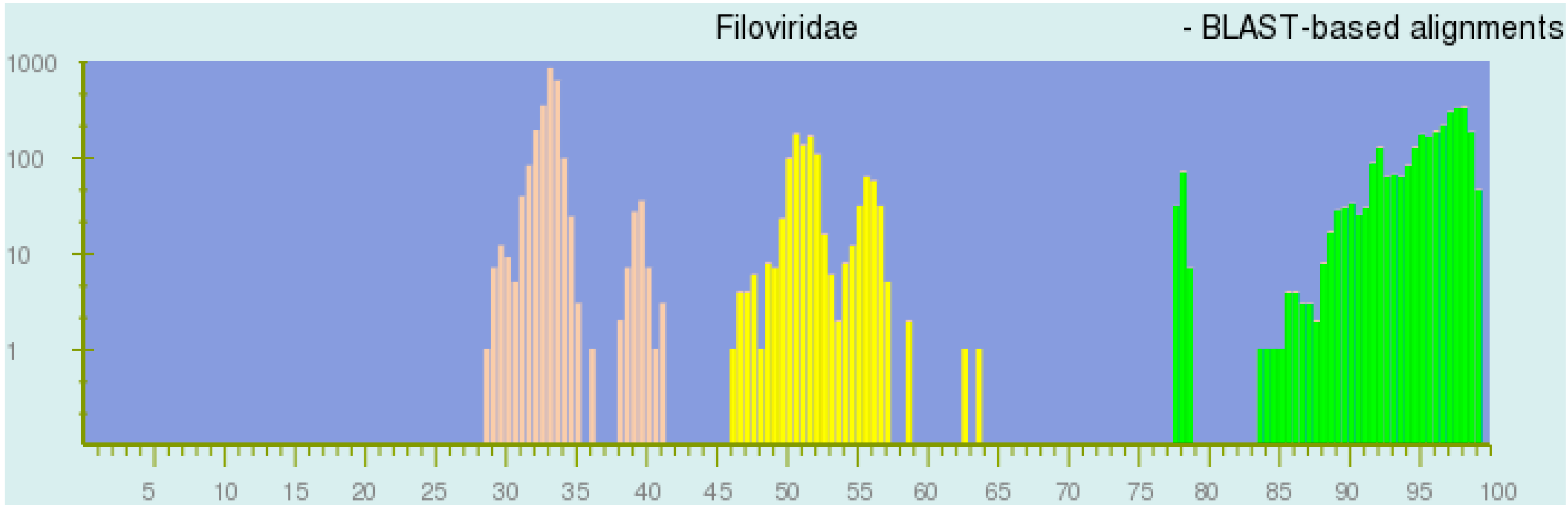
- National Center for Biotechnology Information. PASC—Filoviridae. List of Non-Redundant Sequences (Using BLAST-Based Alignments). 2017. Available online: https://www.ncbi.nlm.nih.gov/sutils/pasc/viridty.cgi?textpage=main&action=gilist&id=333 (accessed on 9 May 2017).
- He, B.; Feng, Y.; Zhang, H.; Xu, L.; Yang, W.; Zhang, Y.; Li, X.; Tu, C. Filovirus RNA in fruit bats, China. Emerg. Infect. Dis. 2015, 21, 1675–1677. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-L.; Zhang, Y.-Z.; Jiang, R.-D.; Guo, H.; Zhang, W.; Li, B.; Wang, N.; Wang, L.; Waruhiu, C.; Zhou, J.-H.; et al. Genetically diverse filoviruses in Rousettus and Eonycteris spp. bats, China, 2009 and 2015. Emerg. Infect. Dis. 2017, 23, 482–486. [Google Scholar] [CrossRef] [PubMed]
- International Committee on Taxonomy of Viruses. Taxonomy Proposal Templates. Available online: https://talk.ictvonline.org/files/taxonomy-proposal-templates/ (accessed on 9 May 2017).
| Current Taxonomy and Nomenclature |
|---|
| Order Mononegavirales Family Filoviridae Genus Marburgvirus Species Marburg Marburgvirus Virus 1: Marburg virus (MARV) Virus 2: Ravn virus (RAVV) Genus Ebolavirus Species Bundibugyo ebolavirus Virus: Bundibugyo virus (BDBV) Species Reston ebolavirus Virus: Reston virus (RESTV) Species Sudan ebolavirus Virus: Sudan virus (SUDV) Species Taï Forest ebolavirus Virus: Taï Forest virus (TAFV) Species Zaire ebolavirus Virus: Ebola virus (EBOV) Genus Cuevavirus Species Lloviu cuevavirus Virus: Lloviu virus (LLOV) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bào, Y.; Amarasinghe, G.K.; Basler, C.F.; Bavari, S.; Bukreyev, A.; Chandran, K.; Dolnik, O.; Dye, J.M.; Ebihara, H.; Formenty, P.; et al. Implementation of Objective PASC-Derived Taxon Demarcation Criteria for Official Classification of Filoviruses. Viruses 2017, 9, 106. https://doi.org/10.3390/v9050106
Bào Y, Amarasinghe GK, Basler CF, Bavari S, Bukreyev A, Chandran K, Dolnik O, Dye JM, Ebihara H, Formenty P, et al. Implementation of Objective PASC-Derived Taxon Demarcation Criteria for Official Classification of Filoviruses. Viruses. 2017; 9(5):106. https://doi.org/10.3390/v9050106
Chicago/Turabian StyleBào, Yīmíng, Gaya K. Amarasinghe, Christopher F. Basler, Sina Bavari, Alexander Bukreyev, Kartik Chandran, Olga Dolnik, John M. Dye, Hideki Ebihara, Pierre Formenty, and et al. 2017. "Implementation of Objective PASC-Derived Taxon Demarcation Criteria for Official Classification of Filoviruses" Viruses 9, no. 5: 106. https://doi.org/10.3390/v9050106
APA StyleBào, Y., Amarasinghe, G. K., Basler, C. F., Bavari, S., Bukreyev, A., Chandran, K., Dolnik, O., Dye, J. M., Ebihara, H., Formenty, P., Hewson, R., Kobinger, G. P., Leroy, E. M., Mühlberger, E., Netesov, S. V., Patterson, J. L., Paweska, J. T., Smither, S. J., Takada, A., ... Kuhn, J. H. (2017). Implementation of Objective PASC-Derived Taxon Demarcation Criteria for Official Classification of Filoviruses. Viruses, 9(5), 106. https://doi.org/10.3390/v9050106