African Swine Fever Virus: A Review
Abstract
:1. Introduction
2. African Swine Fever Virus
3. Virion Structure
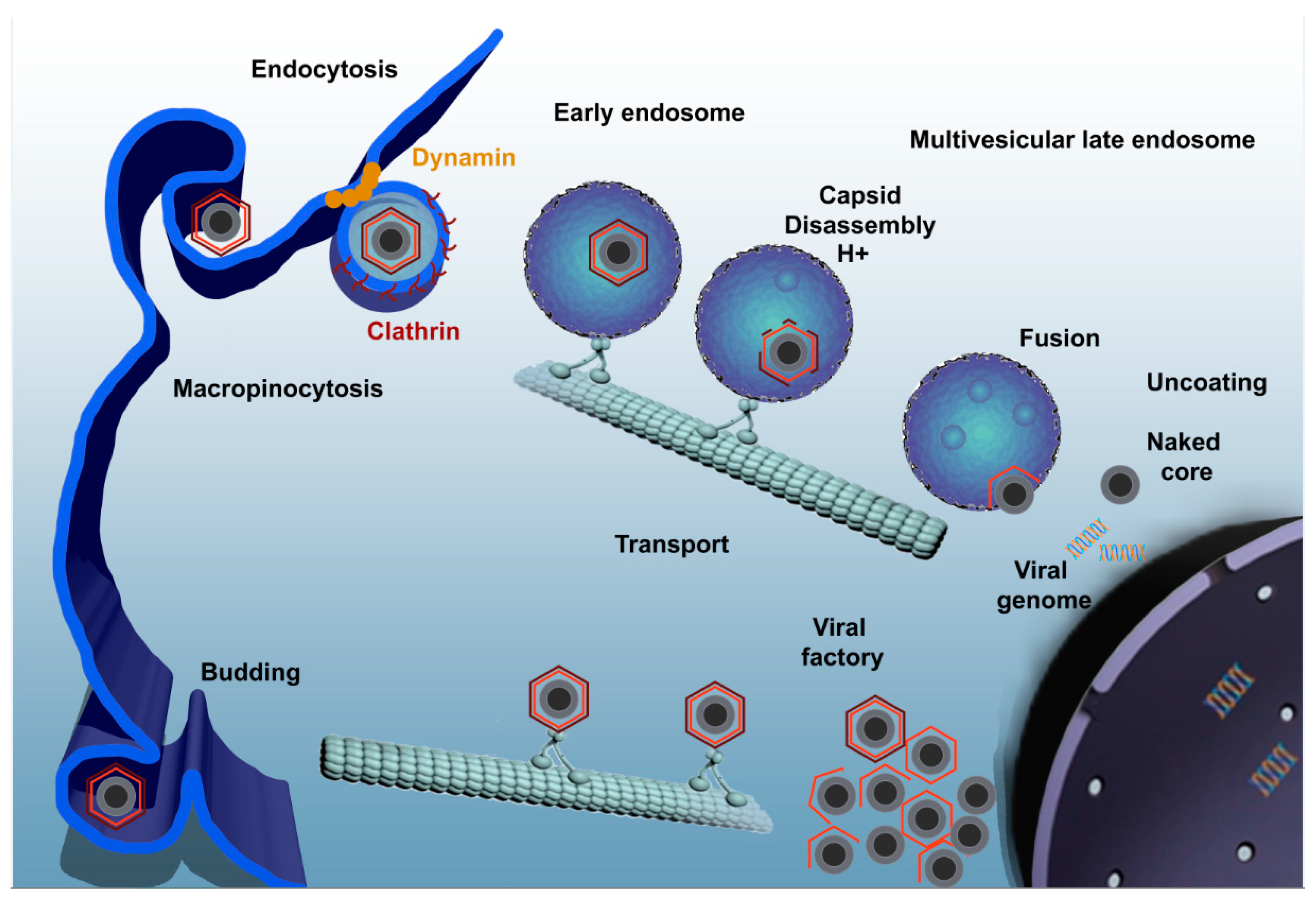
4. Viral Entry Mechanisms
5. ASFV Enters the Endosomal Pathway
6. ASFV Gene Expression and DNA Replication
7. Formation of the Viral Factory
8. ER Stress and Unfolded Protein Response
9. ASFV and Apoptosis
10. ASFV and Autophagy
11. ASFV Egress
12. Potential Vaccines and Antivirals
Acknowledgments
Conflicts of Interest
References
- Parker, J.; Plowright, W.; Pierce, M.A. The epizootiology of african swine fever in Africa. Vet. Rec. 1969, 85, 668–674. [Google Scholar] [PubMed]
- Thomson, G.R.; Gainaru, M.D.; van Dellen, A.F. Experimental infection of warthos (Phacochoerus aethiopicus) with african swine fever virus. Onderstepoort J. Vet. Res. 1980, 47, 19–22. [Google Scholar] [PubMed]
- Anderson, E.C.; Hutchings, G.H.; Mukarati, N.; Wilkinson, P.J. African swine fever virus infection of the bushpig (Potamochoerus porcus) and its significance in the epidemiology of the disease. Vet. Microbiol. 1998, 62, 1–15. [Google Scholar] [CrossRef]
- Penrith, M.L.; Vosloo, W. Review of african swine fever: Transmission, spread and control. J. S. Afr. Vet. Assoc. 2009, 80, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Dixon, L.K.; Escribano, J.M.; Martins, C.; Rock, D.L.; Salas, M.L.; Wilkinson, P.J. Asfarviridae. In Virus Taxonomy. VIIIth Report of the ICTV; Fauquet, C.M., Mayo, M.A., Maniloff, J., Desselberger, U., Ball, L.A., Eds.; Elsevier/Academic Press: London, UK, 2005; pp. 135–143. [Google Scholar]
- Kleiboeker, S.B.; Scoles, G.A.; Burrage, T.G.; Sur, J. African swine fever virus replication in the midgut epithelium is required for infection of ornithodoros ticks. J. Virol. 1999, 73, 8587–8598. [Google Scholar] [PubMed]
- Montgomery, R.E. On a form of swine fever occurring in british east Africa (Kenya Colony). J. Comp. Pathol. Ther. 1921, 34, 159–191. [Google Scholar] [CrossRef]
- Wozniakowski, G.; Kozak, E.; Kowalczyk, A.; Lyjak, M.; Pomorska-Mol, M.; Niemczuk, K.; Pejsak, Z. Current status of african swine fever virus in a population of wild boar in eastern Poland (2014–2015). Arch. Virol. 2016, 161, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Pejsak, Z.; Truszczynski, M.; Niemczuk, K.; Kozak, E.; Markowska-Daniel, I. Epidemiology of african swine fever in Poland since the detection of the first case. Pol. J. Vet. Sci. 2014, 17, 665–672. [Google Scholar] [CrossRef] [PubMed]
- OIE-WAHID. World Animal Health Information Database (Wahid) [Database on the Internet]. World Organisation for Animal Health (OIE), 2017. [Cited World Animal Health Information System (WAHIS)]. Available online: http://www.Oie.Int/wahis/public.Php?Page=home (accessed on 27 March 2017).
- Boshoff, C.I.; Bastos, A.D.; Gerber, L.J.; Vosloo, W. Genetic characterisation of african swine fever viruses from outbreaks in southern Africa (1973–1999). Vet. Microbiol. 2007, 121, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Bastos, A.D.; Penrith, M.L.; Cruciere, C.; Edrich, J.L.; Hutchings, G.; Roger, F.; Couacy-Hymann, E.; Thomson, G.R. Genotyping field strains of african swine fever virus by partial p72 gene characterisation. Arch. Virol. 2003, 148, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Achenbach, J.E.; Gallardo, C.; Nieto-Pelegrin, E.; Rivera-Arroyo, B.; Degefa-Negi, T.; Arias, M.; Jenberie, S.; Mulisa, D.D.; Gizaw, D.; Gelaye, E.; et al. Identification of a new genotype of african swine fever virus in domestic pigs from Ethiopia. Transbound. Emerg. Dis. 2016. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beato, R.; Salas, M.L.; Vinuela, E.; Salas, J. Role of the host cell nucleus in the replication of African swine fever virus DNA. Virology 1992, 188, 637–649. [Google Scholar] [CrossRef]
- Rojo, G.; Garcia-Beato, R.; Vinuela, E.; Salas, M.L.; Salas, J. Replication of african swine fever virus DNA in infected cells. Virology 1999, 257, 524–536. [Google Scholar] [CrossRef] [PubMed]
- Ballester, M.; Rodriguez-Carino, C.; Perez, M.; Gallardo, C.; Rodriguez, J.M.; Salas, M.L.; Rodriguez, F. Disruption of nuclear organization during the initial phase of african swine fever virus infection. J. Virol. 2011, 85, 8263–8269. [Google Scholar] [CrossRef] [PubMed]
- Almazan, F.; Rodriguez, J.M.; Andres, G.; Perez, R.; Vinuela, E.; Rodriguez, J.F. Transcriptional analysis of multigene family 110 of african swine fever virus. J. Virol. 1992, 66, 6655–6667. [Google Scholar] [PubMed]
- Rodriguez, J.M.; Salas, M.L.; Vinuela, E. Intermediate class of mRNAs in African swine fever virus. J. Virol. 1996, 70, 8584–8589. [Google Scholar] [PubMed]
- Almazan, F.; Rodriguez, J.M.; Angulo, A.; Vinuela, E.; Rodriguez, J.F. Transcriptional mapping of a late gene coding for the p12 attachment protein of African swine fever virus. J. Virol. 1993, 67, 553–556. [Google Scholar] [PubMed]
- Salas, M.L.; Kuznar, J.; Vinuela, E. Polyadenylation, methylation, and capping of the RNA synthesized in vitro by African swine fever virus. Virology 1981, 113, 484–491. [Google Scholar] [CrossRef]
- Salas, M.L.; Andres, G. African swine fever virus morphogenesis. Virus Res. 2013, 173, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Breese, S.S., Jr.; de Boer, C.J. Electron microscope observations of African swine fever virus in tissue culture cells. Virology 1966, 28, 420–428. [Google Scholar] [CrossRef]
- Schloer, G.M. Polypeptides and structure of African swine fever virus. Virus Res. 1985, 3, 295–310. [Google Scholar] [CrossRef]
- Valdeira, M.L.; Geraldes, A. Morphological study on the entry of African swine fever virus into cells. Biol. Cell 1985, 55, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Alcami, A.; Carrascosa, A.L.; Vinuela, E. Interaction of African swine fever virus with macrophages. Virus Res. 1990, 17, 93–104. [Google Scholar] [CrossRef]
- Alcami, A.; Carrascosa, A.L.; Vinuela, E. The entry of African swine fever virus into vero cells. Virology 1989, 171, 68–75. [Google Scholar] [CrossRef]
- Alcami, A.; Carrascosa, A.L.; Vinuela, E. Saturable binding sites mediate the entry of African swine fever virus into vero cells. Virology 1989, 168, 393–398. [Google Scholar] [CrossRef]
- Sanchez-Torres, C.; Gomez-Puertas, P.; Gomez-del-Moral, M.; Alonso, F.; Escribano, J.M.; Ezquerra, A.; Dominguez, J. Expression of porcine cd163 on monocytes/macrophages correlates with permissiveness to African swine fever infection. Arch. Virol. 2003, 148, 2307–2323. [Google Scholar] [CrossRef] [PubMed]
- Popescu, L.; Gaudreault, N.N.; Whitworth, K.M.; Murgia, M.V.; Nietfeld, J.C.; Mileham, A.; Samuel, M.; Wells, K.D.; Prather, R.S.; Rowland, R.R. Genetically edited pigs lacking cd163 show no resistance following infection with the African swine fever virus isolate, georgia 2007/1. Virology 2017, 501, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Galindo, I.; Cuesta-Geijo, M.A.; Hlavova, K.; Munoz-Moreno, R.; Barrado-Gil, L.; Dominguez, J.; Alonso, C. African swine fever virus infects macrophages, the natural host cells, via clathrin and cholesterol-dependent endocytosis. Virus Res. 2015, 200, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, B.; Alonso, C. Dynamin and clathrin-dependent endocytosis in African swine fever virus entry. J. Virol. 2010, 84, 2100–2109. [Google Scholar] [CrossRef] [PubMed]
- Basta, S.; Gerber, H.; Schaub, A.; Summerfield, A.; McCullough, K.C. Cellular processes essential for African swine fever virus to infect and replicate in primary macrophages. Vet. Microbiol. 2010, 140, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, E.G.; Quintas, A.; Perez-Nunez, D.; Nogal, M.; Barroso, S.; Carrascosa, A.L.; Revilla, Y. African swine fever virus uses macropinocytosis to enter host cells. PLoS Pathog. 2012, 8, e1002754. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, B.; Guerra, M.; Salas, M.L.; Andres, G. African swine fever virus undergoes outer envelope disruption, capsid disassembly and inner envelope fusion before core release from multivesicular endosomes. PLoS Pathog. 2016, 12, e1005595. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Puertas, P.; Rodriguez, F.; Oviedo, J.M.; Brun, A.; Alonso, C.; Escribano, J.M. The african swine fever virus proteins p54 and p30 are involved in two distinct steps of virus attachment and both contribute to the antibody-mediated protective immune response. Virology 1998, 243, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Angulo, A.; Vinuela, E.; Alcami, A. Comparison of the sequence of the gene encoding African swine fever virus attachment protein p12 from field virus isolates and viruses passaged in tissue culture. J. Virol. 1992, 66, 3869–3872. [Google Scholar] [PubMed]
- Angulo, A.; Vinuela, E.; Alcami, A. Inhibition of African swine fever virus binding and infectivity by purified recombinant virus attachment protein p12. J. Virol. 1993, 67, 5463–5471. [Google Scholar] [PubMed]
- Cuesta-Geijo, M.A.; Galindo, I.; Hernaez, B.; Quetglas, J.I.; Dalmau-Mena, I.; Alonso, C. Endosomal maturation, Rab7 GTPase and phosphoinositides in African swine fever virus entry. PLoS ONE 2012, 7, e48853. [Google Scholar] [CrossRef] [PubMed]
- Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 2009, 10, 513–525. [Google Scholar] [CrossRef] [PubMed]
- Lozach, P.Y.; Mancini, R.; Bitto, D.; Meier, R.; Oestereich, L.; Overby, A.K.; Pettersson, R.F.; Helenius, A. Entry of bunyaviruses into mammalian cells. Cell Host Microbe 2010, 7, 488–499. [Google Scholar] [CrossRef] [PubMed]
- Pasqual, G.; Rojek, J.M.; Masin, M.; Chatton, J.Y.; Kunz, S. Old world arenaviruses enter the host cell via the multivesicular body and depend on the endosomal sorting complex required for transport. PLoS Pathog. 2011, 7, e1002232. [Google Scholar] [CrossRef]
- Cuesta-Geijo, M.A.; Chiappi, M.; Galindo, I.; Barrado-Gil, L.; Munoz-Moreno, R.; Carrascosa, J.L.; Alonso, C. Cholesterol flux is required for endosomal progression of african swine fever virions during the initial establishment of infection. J. Virol. 2015, 90, 1534–1543. [Google Scholar] [CrossRef] [PubMed]
- Alonso, C.; Miskin, J.; Hernaez, B.; Fernandez-Zapatero, P.; Soto, L.; Canto, C.; Rodriguez-Crespo, I.; Dixon, L.; Escribano, J.M. African swine fever virus protein p54 interacts with the microtubular motor complex through direct binding to light-chain dynein. J. Virol. 2001, 75, 9819–9827. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.M.; Salas, M.L. African swine fever virus transcription. Virus Res. 2013, 173, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, B.; Escribano, J.M.; Alonso, C. Visualization of the African swine fever virus infection in living cells by incorporation into the virus particle of green fluorescent protein-p54 membrane protein chimera. Virology 2006, 350, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Quetglas, J.I.; Hernaez, B.; Galindo, I.; Munoz-Moreno, R.; Cuesta-Geijo, M.A.; Alonso, C. Small Rho GTPases and cholesterol biosynthetic pathway intermediates in African swine fever virus infection. J. Virol. 2012, 86, 1758–1767. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, Z.G.; de Matos, A.P.; Rodrigues-Pousada, C. Association of African swine fever virus with the cytoskeleton. Virus Res. 1988, 11, 175–192. [Google Scholar] [CrossRef]
- De Matos, A.P.; Carvalho, Z.G. African swine fever virus interaction with microtubules. Biol. Cell 1993, 78, 229–234. [Google Scholar] [CrossRef]
- Heath, C.M.; Windsor, M.; Wileman, T. Aggresomes resemble sites specialized for virus assembly. J. Cell Biol. 2001, 153, 449–455. [Google Scholar] [CrossRef] [PubMed]
- Stefanovic, S.; Windsor, M.; Nagata, K.I.; Inagaki, M.; Wileman, T. Vimentin rearrangement during African swine fever virus infection involves retrograde transport along microtubules and phosphorylation of vimentin by calcium calmodulin kinase II. J. Virol. 2005, 79, 11766–11775. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Moreno, R.; Barrado-Gil, L.; Galindo, I.; Alonso, C. Analysis of HDAC6 and BAG3-aggresome pathways in African swine fever viral factory formation. Viruses 2015, 7, 1823–1831. [Google Scholar] [CrossRef] [PubMed]
- Galindo, I.; Hernaez, B.; Munoz-Moreno, R.; Cuesta-Geijo, M.A.; Dalmau-Mena, I.; Alonso, C. The ATF6 branch of unfolded protein response and apoptosis are activated to promote African swine fever virus infection. Cell Death Dis. 2012, 3, e341. [Google Scholar] [CrossRef] [PubMed]
- Netherton, C.L.; Parsley, J.C.; Wileman, T. African swine fever virus inhibits induction of the stress-induced proapoptotic transcription factor CHOP/GADD153. J. Virol. 2004, 78, 10825–10828. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Moon, A.; Childs, K.; Goodbourn, S.; Dixon, L.K. The African swine fever virus DP71L protein recruits the protein phosphatase 1 catalytic subunit to dephosphorylate eIF2alpha and inhibits chop induction but is dispensable for these activities during virus infection. J. Virol. 2010, 84, 10681–10689. [Google Scholar] [CrossRef] [PubMed]
- Rivera, J.; Abrams, C.; Hernaez, B.; Alcazar, A.; Escribano, J.M.; Dixon, L.; Alonso, C. The MyD116 African swine fever virus homologue interacts with the catalytic subunit of protein phosphatase 1 and activates its phosphatase activity. J. Virol. 2007, 81, 2923–2929. [Google Scholar] [CrossRef] [PubMed]
- Benedict, C.A.; Norris, P.S.; Ware, C.F. To kill or be killed: Viral evasion of apoptosis. Nat. Immunol. 2002, 3, 1013–1018. [Google Scholar] [CrossRef] [PubMed]
- Brun, A.; Rivas, C.; Esteban, M.; Escribano, J.M.; Alonso, C. African swine fever virus gene a179l, a viral homologue of BCL-2, protects cells from programmed cell death. Virology 1996, 225, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Galindo, I.; Hernaez, B.; Diaz-Gil, G.; Escribano, J.M.; Alonso, C. A179l, a viral BCL-2 homologue, targets the core BCL-2 apoptotic machinery and its upstream BH3 activators with selective binding restrictions for bid and noxa. Virology 2008, 375, 561–572. [Google Scholar] [CrossRef] [PubMed]
- Chacon, M.R.; Almazan, F.; Nogal, M.L.; Vinuela, E.; Rodriguez, J.F. The African swine fever virus IAP homolog is a late structural polypeptide. Virology 1995, 214, 670–674. [Google Scholar] [CrossRef] [PubMed]
- Nogal, M.L.; Gonzalez de Buitrago, G.; Rodriguez, C.; Cubelos, B.; Carrascosa, A.L.; Salas, M.L.; Revilla, Y. African swine fever virus IAP homologue inhibits caspase activation and promotes cell survival in mammalian cells. J. Virol. 2001, 75, 2535–2543. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, C.I.; Nogal, M.L.; Carrascosa, A.L.; Salas, M.L.; Fresno, M.; Revilla, Y. African swine fever virus IAP-like protein induces the activation of nuclear factor kappa B. J. Virol. 2002, 76, 3936–3942. [Google Scholar] [CrossRef] [PubMed]
- Revilla, Y.; Callejo, M.; Rodriguez, J.M.; Culebras, E.; Nogal, M.L.; Salas, M.L.; Vinuela, E.; Fresno, M. Inhibition of nuclear factor kappa b activation by a virus-encoded I kappa b-like protein. J. Biol. Chem. 1998, 273, 5405–5411. [Google Scholar] [CrossRef] [PubMed]
- Ramiro-Ibanez, F.; Ortega, A.; Brun, A.; Escribano, J.M.; Alonso, C. Apoptosis: A mechanism of cell killing and lymphoid organ impairment during acute african swine fever virus infection. J. Gen. Virol. 1996, 77, 2209–2219. [Google Scholar] [CrossRef] [PubMed]
- Orvedahl, A.; Alexander, D.; Talloczy, Z.; Sun, Q.; Wei, Y.; Zhang, W.; Burns, D.; Leib, D.A.; Levine, B. HSV-1 ICP34.5 confers neurovirulence by targeting the Beclin 1 autophagy protein. Cell Host Microbe 2007, 1, 23–35. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, B.; Cabezas, M.; Munoz-Moreno, R.; Galindo, I.; Cuesta-Geijo, M.A.; Alonso, C. A179l, a new viral Bcl2 homolog targeting Beclin 1 autophagy related protein. Curr. Mol. Med. 2013, 13, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Klionsky, D.J. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 2016, 12, 1–222. [Google Scholar] [CrossRef] [PubMed]
- Jouvenet, N.; Monaghan, P.; Way, M.; Wileman, T. Transport of african swine fever virus from assembly sites to the plasma membrane is dependent on microtubules and conventional kinesin. J. Virol. 2004, 78, 7990–8001. [Google Scholar] [CrossRef] [PubMed]
- Andres, G.; Garcia-Escudero, R.; Vinuela, E.; Salas, M.L.; Rodriguez, J.M. African swine fever virus structural protein pE120R is essential for virus transport from assembly sites to plasma membrane but not for infectivity. J. Virol. 2001, 75, 6758–6768. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, V.; Risatti, G.R.; Holinka, L.G.; Krug, P.W.; Carlson, J.; Velazquez-Salinas, L.; Azzinaro, P.A.; Gladue, D.P.; Borca, M.V. Simultaneous deletion of the 9gl and uk genes from the African swine fever virus georgia 2007 isolate offers increased safety and protection against homologous challenge. J. Virol. 2017, 91, e01760-16. [Google Scholar]
- Lacasta, A.; Ballester, M.; Monteagudo, P.L.; Rodriguez, J.M.; Salas, M.L.; Accensi, F.; Pina-Pedrero, S.; Bensaid, A.; Argilaguet, J.; Lopez-Soria, S.; et al. Expression library immunization can confer protection against lethal challenge with african swine fever virus. J. Virol. 2014, 88, 13322–13332. [Google Scholar] [CrossRef] [PubMed]
- Reis, A.L.; Abrams, C.C.; Goatley, L.C.; Netherton, C.; Chapman, D.G.; Sanchez-Cordon, P.; Dixon, L.K. Deletion of African swine fever virus interferon inhibitors from the genome of a virulent isolate reduces virulence in domestic pigs and induces a protective response. Vaccine 2016, 34, 4698–4705. [Google Scholar] [CrossRef] [PubMed]
- Cubillos, C.; Gomez-Sebastian, S.; Moreno, N.; Nunez, M.C.; Mulumba-Mfumu, L.K.; Quembo, C.J.; Heath, L.; Etter, E.M.; Jori, F.; Escribano, J.M.; et al. African swine fever virus serodiagnosis: A general review with a focus on the analyses of african serum samples. Virus Res. 2013, 173, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Galindo, I.; Hernaez, B.; Berna, J.; Fenoll, J.; Cenis, J.L.; Escribano, J.M.; Alonso, C. Comparative inhibitory activity of the stilbenes resveratrol and oxyresveratrol on african swine fever virus replication. Antivir. Res. 2011, 91, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Fabregas, J.; Garcia, D.; Fernandez-Alonso, M.; Rocha, A.I.; Gomez-Puertas, P.; Escribano, J.M.; Otero, A.; Coll, J.M. In vitro inhibition of the replication of haemorrhagic septicaemia virus (VHSV) and African swine fever virus (ASFV) by extracts from marine microalgae. Antivir. Res. 1999, 44, 67–73. [Google Scholar] [CrossRef]
- Hurtado, C.; Bustos, M.J.; Sabina, P.; Nogal, M.L.; Granja, A.G.; Gonzalez, M.E.; Gonzalez-Porque, P.; Revilla, Y.; Carrascosa, A.L. Antiviral activity of lauryl gallate against animal viruses. Antivir. Ther. 2008, 13, 909–917. [Google Scholar] [PubMed]
- Mottola, C.; Freitas, F.B.; Simoes, M.; Martins, C.; Leitao, A.; Ferreira, F. In vitro antiviral activity of fluoroquinolones against African swine fever virus. Vet. Microbiol. 2013, 165, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, B.; Tarrago, T.; Giralt, E.; Escribano, J.M.; Alonso, C. Small peptide inhibitors disrupt a high-affinity interaction between cytoplasmic dynein and a viral cargo protein. J. Virol. 2010, 84, 10792–10801. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Galindo, I.; Alonso, C. African Swine Fever Virus: A Review. Viruses 2017, 9, 103. https://doi.org/10.3390/v9050103
Galindo I, Alonso C. African Swine Fever Virus: A Review. Viruses. 2017; 9(5):103. https://doi.org/10.3390/v9050103
Chicago/Turabian StyleGalindo, Inmaculada, and Covadonga Alonso. 2017. "African Swine Fever Virus: A Review" Viruses 9, no. 5: 103. https://doi.org/10.3390/v9050103
APA StyleGalindo, I., & Alonso, C. (2017). African Swine Fever Virus: A Review. Viruses, 9(5), 103. https://doi.org/10.3390/v9050103





