Antiviral Screening of Multiple Compounds against Ebola Virus
Abstract
:1. Introduction
- 17-DMAG: An inhibitor of heat shock protein 90 (HSP90), which has been shown to reduce in vitro EBOV replication [7]
- BGB324: An inhibitor of Axl receptor tyrosine kinase, which appears to be involved with Ebola virus entry into host cells [8]
- JB1a: An antibody therapy, targeting beta-1 integrins, which have been proposed to facilitate the entry of filoviruses; treatment of target cells with the JB1a clone reduced infection using a vesicular stomatitis virus (VSIV) pseudotyped with EBOV glycoprotein [9]
- Omeprazole and esomeprazole magnesium: Members of the benzimidazoles that may stop viral entry via clathrin-mediated endocytosis by raising the endosomal pH. Both compounds were shown to inhibit lentivirus-based pseudotypes expressing EBOV glycoprotein [10]
- Gleevec and Tasigna (market names for imatinib mesylate and nilotinib, respectively): Specific tyrosine kinase inhibitors originally developed as anticancer compounds and proposed to inhibit phosphorylation of the VP40 matrix protein which is required for EBOV exit from cells [11]. During large-scale screens of antivirals against EBOV, other groups have identified Gleevec [12] and Tasigna [13] as potential EBOV inhibitors
- Aimspro (anti-inflammatory immuno-suppressive drug): Originally developed for the treatment of human immunodeficiency virus (HIV) by the production of hyperimmune serum in goats injected with inactivated HIV IIIB, the serum has revealed the presence of a range of components, including the cytokines interleukin (IL)-4 and IL-10, proopiomelanocortin, arginine vasopressin, β-endorphin and corticotropin-releasing factor [14]
- NCK-8 and D-LANA-14: Small molecules that mimic the properties of antimicrobial peptides, NCK-8 [15,16] and D-LANA-14 [17] have demonstrated potent activity against drug-resistant bacteria and their biofilms. The activity of this class of compounds is attributed to their membrane disrupting properties [18,19,20]. Peptide mimics [21] and several other small molecules have demonstrated activity against EBOV. Owing to the membrane-disrupting [22,23] modes of action of this class of compounds (e.g., NCK-8 and DLANA-14), they were expected to be active against EBOV
- Celgosivir and its prodrug castanospermine: Broad spectrum inhibitors of host glucosidases. Inhibitors of endoplasmic reticulum (ER) α-glucosidases have been shown to act as antivirals with several haemorrhagic fever viruses, including EBOV [24]
- Zidovudine, didanosine, stavudine, abacavir sulphate and entecavir: Compounds included in the study upon request of the Wellcome Trust
2. Materials and Methods
2.1. In Vitro Screening
2.1.1. Virus Assay
2.1.2. Molecular Assay
2.1.3. Toxicity Assay
2.1.4. Repeat Compound Screening
2.2. In Vivo Screening
3. Results
3.1. Selection of Compounds
3.2. Effects of Compounds against In Vitro EBOV Replication
3.2.1. Initial Screen at Recommended Concentration
3.2.2. Secondary Screening with a High Virus Inoculation
3.3. Screening of Compounds for Effects against Disease in EBOV-Infected Guinea Pigs
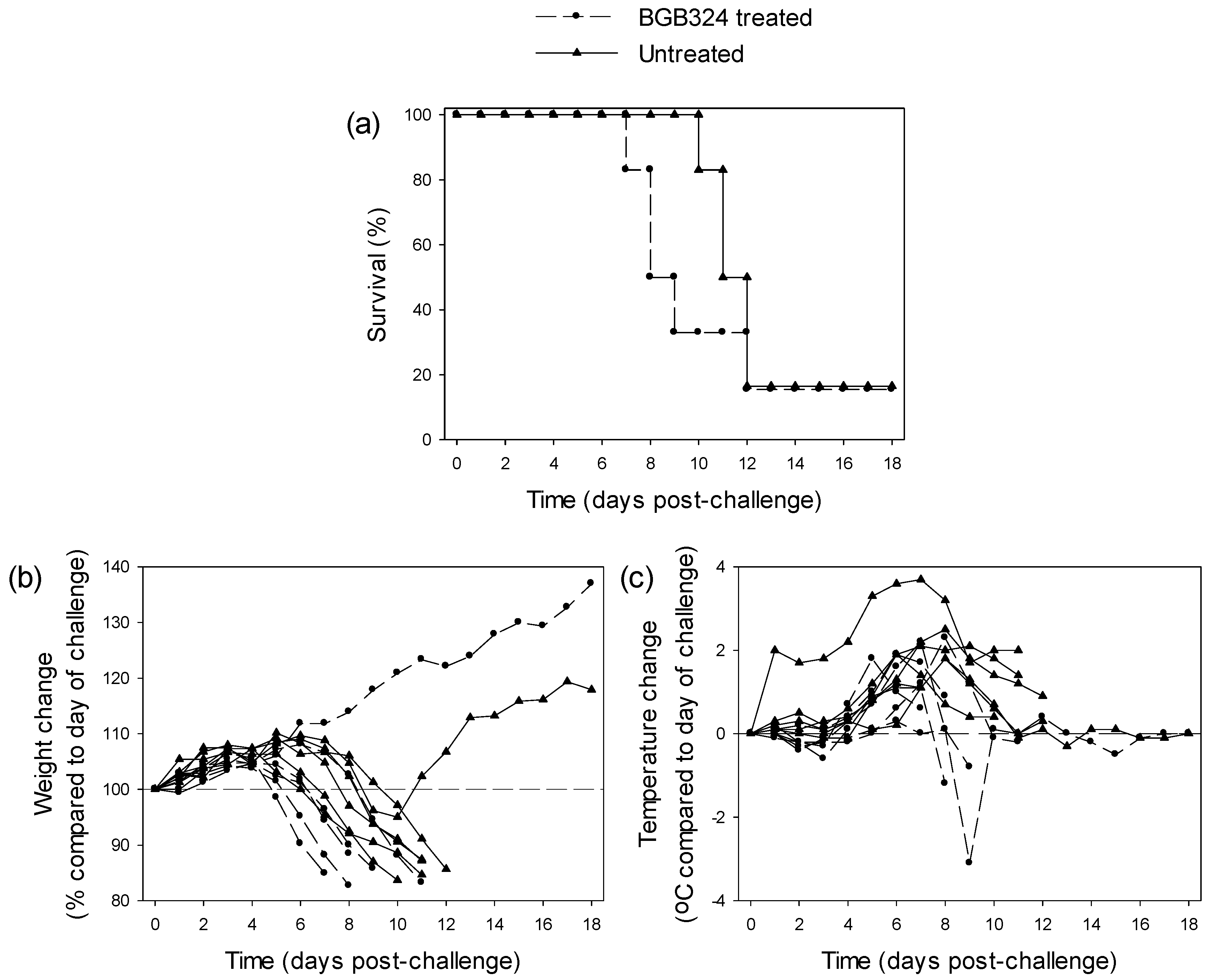
3.3.1. Testing of BGB324 in EBOV-Infected Guinea Pigs
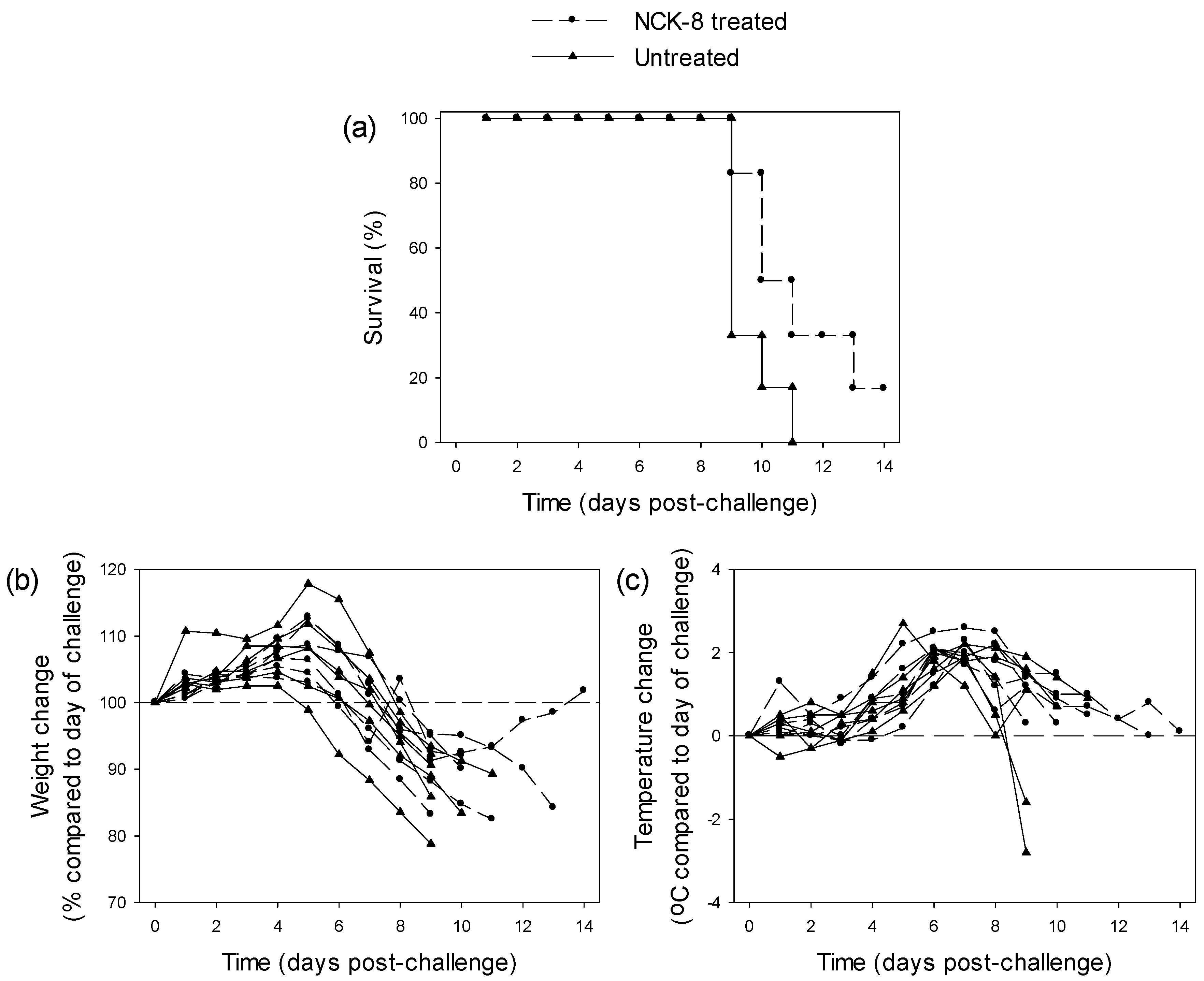
3.3.2. Testing of NCK-8 in EBOV-Infected Guinea Pigs
3.3.3. Testing of 17-DMAG in EBOV-Infected Guinea Pigs
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix
| U.S. Army Medical Research and Material Command (USAMRMC) equivalent TRL descriptions for drugs, biologics and vaccines (pharmaceuticals) synthesized from the Technology Readiness Assessment (TRA) Deskbook [46]. | |||
| TRL | Description | Decision Criterion | Supporting Information |
| 1 | Lowest level of technology readiness. Maintenance of scientific awareness and generation of scientific and bioengineering knowledge base. Scientific findings are reviewed and assessed as a foundation for characterizing new technologies. | Scientific literature reviews and initial market surveys are initiated and assessed. Potential scientific application to defined problems is articulated. | Reviews of open, published scientific literature concerning basic principles. Findings from market surveys of the open literature. Privately funded research findings or market surveys are proprietary and rarely available to the public but could be made available upon request under confidentiality agreement. |
| 2 | Intense intellectual focus on the problem, with generation of scientific “paper studies” that review and generate research ideas, hypotheses, and experimental designs for addressing the related scientific issues. | Hypothesis(es) is (are) generated. Research plans and/or protocols are developed, peer reviewed, and approved. | Focused literature reviews are conducted and scientific discussions are held to generate research plans and studies that identify potential targets of opportunity for therapeutic intervention and to facilitate strategic planning. Supporting analyses provide scientific information and data for developing research proposals for filling in data gaps and identifying candidate concepts and/or therapeutic drugs. Documented by peer-reviewed approved protocol(s) or research plan(s). |
| 3 | Basic research, data collection, and analysis begin in order to test hypothesis, explore alternative concepts, and identify and evaluate critical technologies and components supporting research and eventual development of the pharmaceutical candidate, identification of sites and mechanisms of action (and potential correlates of protection for vaccines), as well as initial characterization of candidates. | Initial proof-of-concept for candidate constructs is demonstrated in a limited number of in vitro and in vivo research models. | Documentation of the results of laboratory studies demonstrates preliminary proof-of-concept (with candidate constructs) from in vitro and animal studies. |
| 4 | Non- good laboratory practice (GLP) research to refine hypothesis and identify relevant parametric data required for technological assessment in a rigorous (worst case) experimental design. Exploratory study of candidate drugs or of critical technologies for effective integration into candidate biologic/vaccine constructs. Candidate drugs (or biologics/vaccines) are evaluated in animal model(s) to identify and assess safety, toxicity and adverse/biological/side effects, and assays (and/or surrogate markers and endpoints) to be used during non-clinical and clinical studies to evaluate and characterize candidate pharmaceuticals are identified. | Proof-of-concept and safety of candidate drug formulations (or candidate biologic/vaccine constructs) are demonstrated in defined laboratory/animal model(s). | Documented proof-of-concept and safety of the candidate are demonstrated by results of formulation studies (or proposed production/purification methods of the biologic/vaccine), laboratory tests, pharmacokinetic studies, and selection of laboratory/animal models. |
| 5 | Intense period of non-clinical and preclinical research studies involving parametric data collection and analysis in well-defined systems, with pilot lots of candidate pharmaceuticals produced and further development of selected candidate(s). In the case of drug, results of research with pilot lots provide basis for a manufacturing process amenable to current good manufacturing practice (cGMP)-compliant pilot lot production. In the case of biologic/vaccine, research results support proposing a potency assay, proposing a manufacturing process amenable to cGMP-compliant pilot lot production, identifying and demonstrating proof-of-concept for a surrogate efficacy marker in an animal model(s) applicable to predicting protective immunity in humans, and demonstrating preliminary safety and efficacy against an appropriate route of challenge in a relevant animal model. Conduct GLP safety and toxicity studies in animal model systems. Identify endpoints of clinical efficacy or its surrogate. Conduct studies to evaluate pharmacokinetics (PK) and pharmacodynamics (PD) and/or immunogenicity as appropriate. Stability studies initiated. | A decision point is reached at which it is determined that sufficient data on the candidate pharmaceutical exist in the draft technical data package to justify proceeding with preparation of an investigational new drug (IND) application. | Reviewers confirm adequacy of information and data in draft technical data package to support preparation of an IND application. Documentation in the draft technical data package contains data from animal pharmacology and toxicology studies, proposed manufacturing information, and clinical protocols suitable for phase 1 clinical testing. |
| 6 | Pre-IND meeting (type B) held and IND application prepared and submitted to the Center for Drug Evaluation and Research (CDER) or the Center for Biologics Evaluation and Research (CBER). Phase 1 clinical trials are conducted to demonstrate safety of candidate in a small number of subjects under carefully controlled and intensely monitored clinical conditions. Evaluation of PK and PD (and/or immunogenicity) data to support the design of well-controlled, scientifically valid phase 2 studies. Production technologies for drug candidates are demonstrated through production-scale cGMP plant qualification, and surrogate efficacy models for biologics/vaccines are validated. | Data from phase 1 clinical trials meet clinical safety requirements and support proceeding to phase 2 clinical trials. | For phase 1 clinical trials to begin, the following are needed: the Food and Drug Administration (FDA)’s and sponsor’s summary minutes of pre-IND meeting document agreements and general adequacy of information and data to support submission of IND application. Review of the submitted IND application does not result in a FDA decision to put a clinical hold on phase 1 clinical trials with the candidate pharmaceutical. For entry into phase 2 clinical trials, the results from phase 1 clinical studies have to demonstrate safety of candidate pharmaceutical. An updated IND, amended with a new clinical protocol to support phase 2 clinical trials or surrogate test plan and submitted to the FDA, documents achieving this criterion. |
| 7 | Phase 2 clinical trials are conducted to determine activity/efficacy/immunogenicity/safety/toxicity of the pharmaceutical as appropriate. These and/or PK-PD data are used to establish product final dose, dose range, schedule, and route of administration. Data are collected, presented and discussed at pre-phase 3 (or surrogate efficacy] meeting (type B) with CDER (or CBER) in support of continued drug development of the drug (or biologic/vaccine), and clinical endpoints and/or surrogate efficacy markers and test plans agreed. | Phase 3 clinical study plan or surrogate test plan has been approved. | FDA’s summary minutes of pre-phase 3 meeting with sponsor discussing results of phase 1 and phase 2 trials, as well as protocols or test plans, provide a record of agreements and basis for sponsor to proceed with phase 3 clinical study or surrogate test plan. An updated IND application, amended with a new clinical protocol to support phase 3 clinical trials or surrogate test plan and submitted to the FDA, documents achieving this criterion. |
| 8 | Implementation of expanded phase 3 clinical trials or surrogate tests to gather information relative to the safety and effectiveness of the candidate drug/biologic/vaccine. Trials are conducted to evaluate the overall risk–benefit of administering the candidate product and to provide an adequate basis for labelling. Process validation is completed and followed by lot consistency/reproducibility studies. In the case of a drug, New Drug Application (NDA) is submitted to CDER following pre-NDA meeting (type B). In the case of a biologic/vaccine, Biologics License Application (BLA) is prepared and submitted to CBER following pre-BLA meeting (type B). Facility Preapproval Inspection (PAI) is completed. | Approval of the NDA for drug by CDER, or approval of the BLA for biologics/vaccines by CBER. | FDA issuance of an approval letter after their review of the NDA or BLA application submitted by the sponsor for the pharmaceutical documents this criterion. |
| 9 | The pharmaceutical (i.e., drug/biologic/vaccine) can be distributed/marketed. Post-marketing studies (non-clinical or clinical) may be required and are designed after agreement with the FDA. Post-marketing surveillance. | None. Continue surveillance. | FDA transmits any requirement for post-marketing studies. Begin post-approval reporting requirements. Maintain cGMP compliance. |
| Points | Description | Decision Criteria | Supporting Info |
|---|---|---|---|
| 0 or 1 or 2 | Availability to make a difference to the current epidemic | Repurposed pharmaceutical or the ability to generate and supply sufficient material within two weeks for in vitro and in vivo studies, and if required subsequently, a reasonable number of therapeutic doses to make a difference to the current Public Health Emergency of International Concern declared by the World Health Organization (WHO). | Documentation to prove that the candidate is:
|
| 0 or 0.5 or 1 or 2 | Likely efficacy against the pathogenic microorganism of interest. | Prior efficacy data against the pathogenic microorganism (or a related agent) of interest, for example through reduction of load or host-immune response. | Scientific studies, reports, commercial-in-confidence information documenting evidence (e.g., theoretical or in silico, in vitro and/or in vivo data) that the candidate is likely to be efficacious against the pathogenic microorganism (or a related agent) of interest. Suggested score: in silico (0.5); in vitro (1); in vivo (2) |
| 0 or 1 or 2 | Practicality and cost-effectiveness (tie-breaker) | Is it a practical and cost-effective solution for frontline clinical response to the current, as well as future epidemics? | Documentation to prove that:
|
References
- Adams, M.J.; Carstens, E.B. Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2012). Arch. Virol. 2012, 157, 1411–1422. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.H.; Becker, S.; Ebihara, H.; Geisbert, T.W.; Johnson, K.M.; Kawaoka, Y.; Lipkin, W.I.; Negredo, A.I.; Netesov, S.V.; Nichol, S.T.; et al. Proposal for a revised taxonomy of the family Filoviridae: classification, names of taxa and viruses, and virus abbreviations. Arch. Virol. 2010, 155, 2083–2103. [Google Scholar] [CrossRef] [PubMed]
- Baize, S.; Pannetier, D.; Oestereich, L.; Rieger, T.; Koivogui, L.; Magassouba, N.; Soropogui, B.; Sow, M.S.; Keita, S.; De Clerck, H.; et al. Emergence of Zaire Ebola virus disease in Guinea. N. Engl. J. Med. 2014, 371, 1418–1425. [Google Scholar] [CrossRef] [PubMed]
- Mahanty, S.; Bray, M. Pathogenesis of filoviral haemorrhagic fevers. Lancet Infect. Dis. 2004, 4, 487–498. [Google Scholar] [CrossRef]
- Fuerstenwerth, H. On the differences between ouabain and digitalis glycosides. Am. J. Ther. 2014, 21, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Dorival, I.; Wu, W.; Dowall, S.; Armstrong, S.; Touzelet, O.; Wastling, J.; Barr, J.N.; Matthews, D.; Carroll, M.; Hewson, R.; et al. Elucidation of the Ebola virus VP24 cellular interactome and disruption of virus biology through targeted inhibition of host-cell protein function. J. Proteome Res. 2014, 13, 5120–5135. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; McCarthy, S.; Chrovian, A.; Olinger, G.; Stossel, A.; Geisbert, T.W.; Hensley, L.E.; Connor, J.H. Inhibition of heat-shock protein 90 reduces Ebola virus replication. Antivir. Res. 2010, 87, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Shimojima, M.; Ikeda, Y.; Kawaoka, Y. The mechanism of Axl-mediated Ebola virus infection. J. Infect. Dis. 2007, 196 (Suppl. 2), S259–S263. [Google Scholar] [CrossRef] [PubMed]
- Takada, A.; Watanabe, S.; Ito, H.; Okazaki, K.; Kida, H.; Kawaoka, Y. Downregulation of beta1 integrins by Ebola virus glycoprotein: implication for virus entry. Virology 2000, 278, 20–26. [Google Scholar] [CrossRef] [PubMed]
- Long, J.; Wright, E.; Molesti, E.; Temperton, N.; Barclay, W. Antiviral therapies against Ebola and other emerging viral diseases using existing medicines that block virus entry. F1000Research 2015, 4, 30. [Google Scholar] [CrossRef] [PubMed]
- Garcia, M.; Cooper, A.; Shi, W.; Bornmann, W.; Carrion, R.; Kalman, D.; Nabel, G.J. Productive replication of Ebola virus is regulated by the c-Abl1 tyrosine kinase. Sci. Transl. Med. 2012, 4, 123ra24. [Google Scholar] [CrossRef] [PubMed]
- Uebelhoer, L.S.; Albarino, C.G.; McMullan, L.K.; Chakrabarti, A.K.; Vincent, J.P.; Nichol, S.T.; Towner, J.S. High-throughput, luciferase-based reverse genetics systems for identifying inhibitors of Marburg and Ebola viruses. Antivir. Res. 2014, 106, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Kouznetsova, J.; Sun, W.; Martinez-Romero, C.; Tawa, G.; Shinn, P.; Chen, C.Z.; Schimmer, A.; Sanderson, P.; McKew, J.C.; Zheng, W.; et al. Identification of 53 compounds that block Ebola virus-like particle entry via a repurposing screen of approved drugs. Emerg. Microbes Infect. 2014, 3, e84. [Google Scholar] [CrossRef] [PubMed]
- Quillinan, N.P.; McIntosh, D.; Vernes, J.; Haq, S.; Denton, C.P. Treatment of diffuse systemic sclerosis with hyperimmune caprine serum (AIMSPRO): a phase II double-blind placebo-controlled trial. Ann. Rheum. Dis. 2014, 73, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, C.; Manjunath, G.B.; Akkapeddi, P.; Yarlagadda, V.; Hoque, J.; Uppu, D.S.; Konai, M.M.; Haldar, J. Small molecular antibacterial peptoid mimics: the simpler the better! J. Med. Chem. 2014, 57, 1428–1436. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, C.; Manjunath, G.B.; Konai, M.M.; Uppu, D.S.; Hoque, J.; Paramanandham, K.; Shome, B.R.; Haldar, J. Aryl-Alkyl-Lysines: Agents That Kill Planktonic Cells, Persister Cells, Biofilms of MRSA and Protect Mice from Skin-Infection. PLoS ONE 2015, 10, e0144094. [Google Scholar] [CrossRef] [PubMed]
- Konai, M.M.; Haldar, J. Lysine-based small molecules that disrupt biofilms and kill both actively growing planktonic and nondividing stationary phase bacteria. ACS Infect. Dis. 2015, 1, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, C.; Manjunath, G.B.; Konai, M.M.; Uppu, D.S.; Hoque, J.; Paramanandham, K.; Shome, B.R.; Ravikumar, R.; Haldar, J. Aryl-alkyl-lysines: Membrane-active small molecules active against murine model of burn infection. ACS Infect. Dis. 2016, 2, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Konai, M.M.; Ghosh, C.; Yarlagadda, V.; Samaddar, S.; Haldar, J. Membrane active phenylalanine conjugated lipophilic norspermidine derivatives with selective antibacterial activity. J. Med. Chem. 2014, 57, 9409–9423. [Google Scholar] [CrossRef] [PubMed]
- Konai, M.M.; Adhikary, U.; Samaddar, S.; Ghosh, C.; Haldar, J. Structure-Activity Relationship of Amino Acid Tunable Lipidated Norspermidine Conjugates: Disrupting Biofilms with Potent Activity against Bacterial Persisters. Bioconjugate Chem. 2015, 26, 2442–2453. [Google Scholar] [CrossRef] [PubMed]
- Clinton, T.R.; Weinstock, M.T.; Jacobsen, M.T.; Szabo-Fresnais, N.; Pandya, M.J.; Whitby, F.G.; Herbert, A.S.; Prugar, L.I.; McKinnon, R.; Hill, C.P.; et al. Design and characterization of ebolavirus GP prehairpin intermediate mimics as drug targets. Protein Sci. 2015, 24, 446–463. [Google Scholar] [CrossRef] [PubMed]
- Picazo, E.; Giordanetto, F. Small molecule inhibitors of ebola virus infection. Drug Discov. Today 2015, 20, 277–286. [Google Scholar] [CrossRef] [PubMed]
- Selakovic, Z.; Soloveva, V.; Gharibeh, D.N.; Wells, J.; Segan, S.; Panchal, R.G.; Solaja, B.A. Anti-Ebola activity of diazachrysene small molecules. ACS Infect. Dis. 2015, 1, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Warren, T.K.; Zhao, X.; Gill, T.; Guo, F.; Wang, L.; Comunale, M.A.; Du, Y.; Alonzi, D.S.; Yu, W.; et al. Small molecule inhibitors of ER alpha-glucosidases are active against multiple hemorrhagic fever viruses. Antivir. Res. 2013, 98, 432–440. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A. Analysis of filovirus entry into vero e6 cells, using inhibitors of endocytosis, endosomal acidification, structural integrity, and cathepsin (B and L) activity. J. Infect. Dis. 2007, 196 (Suppl. 2), S251–S258. [Google Scholar] [CrossRef] [PubMed]
- Muller, S.; Moller, P.; Bick, M.J.; Wurr, S.; Becker, S.; Gunther, S.; Kummerer, B.M. Inhibition of filovirus replication by the zinc finger antiviral protein. J. Virol. 2007, 81, 2391–2400. [Google Scholar] [CrossRef] [PubMed]
- Takada, A.; Ebihara, H.; Jones, S.; Feldmann, H.; Kawaoka, Y. Protective efficacy of neutralizing antibodies against Ebola virus infection. Vaccine 2007, 25, 993–999. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.H.; Lofts, L.L.; Kugelman, J.R.; Smither, S.J.; Lever, M.S.; van der Groen, G.; Johnson, K.M.; Radoshitzky, S.R.; Bavari, S.; Jahrling, P.B.; et al. Reidentification of Ebola Virus E718 and ME as Ebola Virus/H.sapiens-tc/COD/1976/Yambuku-Ecran. Genome Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Trombley, A.R.; Wachter, L.; Garrison, J.; Buckley-Beason, V.A.; Jahrling, J.; Hensley, L.E.; Schoepp, R.J.; Norwood, D.A.; Goba, A.; Fair, J.N.; et al. Comprehensive panel of real-time TaqMan polymerase chain reaction assays for detection and absolute quantification of filoviruses, arenaviruses, and New World hantaviruses. Am. J. Trop. Med. Hyg. 2010, 82, 954–960. [Google Scholar] [CrossRef] [PubMed]
- Dowall, S.; Taylor, I.; Yeates, P.; Smith, L.; Rule, A.; Easterbrook, L.; Bruce, C.; Cook, N.; Corbin-Lickfett, K.; Empig, C.; et al. Catheterized guinea pigs infected with Ebola Zaire virus allows safer sequential sampling to determine the pharmacokinetic profile of a phosphatidylserine-targeting monoclonal antibody. Antivir. Res. 2013, 97, 108–111. [Google Scholar] [CrossRef] [PubMed]
- Dowall, S.D.; Matthews, D.A.; Garcia-Dorival, I.; Taylor, I.; Kenny, J.; Hertz-Fowler, C.; Hall, N.; Corbin-Lickfett, K.; Empig, C.; Schlunegger, K.; et al. Elucidating variations in the nucleotide sequence of Ebola virus associated with increasing pathogenicity. Genome Biol. 2014, 15, 540. [Google Scholar] [CrossRef] [PubMed]
- Assistant Secretary of Defense for Research and Engineering. Technology Readiness Assessment (TRA) Guidance. Department of Defense, April 2011. [Google Scholar]
- Gunther, S.; Asper, M.; Roser, C.; Luna, L.K.; Drosten, C.; Becker-Ziaja, B.; Borowski, P.; Chen, H.M.; Hosmane, R.S. Application of real-time PCR for testing antiviral compounds against Lassa virus, SARS coronavirus and Ebola virus in vitro. Antivir. Res. 2004, 63, 209–215. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, S.D.; Majchrzak-Kita, B.; Racine, T.; Kozlowski, H.N.; Baker, D.P.; Hoenen, T.; Kobinger, G.P.; Fish, E.N.; Branch, D.R. A Rapid Screening Assay Identifies Monotherapy with Interferon-ss and Combination Therapies with Nucleoside Analogs as Effective Inhibitors of Ebola Virus. PLoS Negl. Trop. Dis. 2016, 10, e0004364. [Google Scholar] [CrossRef] [PubMed]
- Johansen, L.M.; Brannan, J.M.; Delos, S.E.; Shoemaker, C.J.; Stossel, A.; Lear, C.; Hoffstrom, B.G.; Dewald, L.E.; Schornberg, K.L.; Scully, C.; et al. FDA-approved selective estrogen receptor modulators inhibit Ebola virus infection. Sci. Transl. Med. 2013, 5, 190ra79. [Google Scholar] [CrossRef] [PubMed]
- Lind, T.; Rydberg, L.; Kyleback, A.; Jonsson, A.; Andersson, T.; Hasselgren, G.; Holmberg, J.; Rohss, K. Esomeprazole provides improved acid control vs. omeprazole In patients with symptoms of gastro-oesophageal reflux disease. Aliment. Pharmacol. Ther. 2000, 14, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Speed, B.; Bu, H.Z.; Pool, W.F.; Peng, G.W.; Wu, E.Y.; Patyna, S.; Bello, C.; Kang, P. Pharmacokinetics, distribution, and metabolism of [14C]sunitinib in rats, monkeys, and humans. Drug Metab. Dispos. 2012, 40, 539–555. [Google Scholar] [CrossRef] [PubMed]
- Gronchi, A.; Blay, J.Y.; Trent, J.C. The role of high-dose imatinib in the management of patients with gastrointestinal stromal tumor. Cancer 2010, 116, 1847–1858. [Google Scholar] [CrossRef] [PubMed]
- Cross, R.W.; Fenton, K.A.; Geisbert, J.B.; Mire, C.E.; Geisbert, T.W. Modeling the Disease Course of Zaire ebolavirus Infection in the Outbred Guinea Pig. J. Infect. Dis. 2015, 212, S305–S315. [Google Scholar] [CrossRef] [PubMed]
- Fleuren, E.D.; Hillebrandt-Roeffen, M.H.; Flucke, U.E.; Te Loo, D.M.; Boerman, O.C.; van der Graaf, W.T.; Versleijen-Jonkers, Y.M. The role of AXL and the in vitro activity of the receptor tyrosine kinase inhibitor BGB324 in Ewing sarcoma. Oncotarget 2014, 5, 12753–12768. [Google Scholar] [CrossRef] [PubMed]
- Uppu, D.S.; Akkapeddi, P.; Manjunath, G.B.; Yarlagadda, V.; Hoque, J.; Haldar, J. Polymers with tunable side-chain amphiphilicity as non-hemolytic antibacterial agents. Chem. Commun. (Camb.) 2013, 49, 9389–9391. [Google Scholar] [CrossRef] [PubMed]
- Egorin, M.J.; Lagattuta, T.F.; Hamburger, D.R.; Covey, J.M.; White, K.D.; Musser, S.M.; Eiseman, J.L. Pharmacokinetics, tissue distribution, and metabolism of 17-(dimethylaminoethylamino)-17-demethoxygeldanamycin (NSC 707545) in CD2F1 mice and Fischer 344 rats. Cancer Chemother. Pharmacol. 2002, 49, 7–19. [Google Scholar] [CrossRef] [PubMed]
- Bente, D.; Gren, J.; Strong, J.E.; Feldmann, H. Disease modeling for Ebola and Marburg viruses. Dis. Models Mech. 2009, 2, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Chen, T.; Hu, Y.; Zhou, Y.; Liu, Q.; Zhou, D.; Jin, X.; Huang, Z. An Ebola virus-like particle-based reporter system enables evaluation of antiviral drugs in vivo under non-BSL-4 conditions. J. Virol. 2016, 90, 8720–8728. [Google Scholar] [CrossRef] [PubMed]
- Edwards, M.R.; Pietzsch, C.; Vausselin, T.; Shaw, M.L.; Bukreyev, A.; Basler, C.F. High-Throughput Minigenome System for Identifying Small-Molecule Inhibitors of Ebola Virus Replication. ACS Infect. Dis. 2015, 1, 380–387. [Google Scholar] [CrossRef] [PubMed]
- Director, Research Directorate (DRD). Technology Readiness Assessment (TRA) Deskbook. Department of Defense, July 2009. [Google Scholar]
| Name | TRL Score 1 | Availability 2 | Efficacy 3 | Total |
|---|---|---|---|---|
| Ouabain | 4 | 2 | 1 | 7 |
| 17-DMAG | 4 | 2 | 1 | 7 |
| BGB324 | 4 | 2 | 1 | 7 |
| Zidovudine | 4 | 2 | 1 | 7 |
| Didanosine | 4 | 2 | 1 | 7 |
| Stavudine | 4 | 2 | 1 | 7 |
| Abacavir sulphate | 4 | 2 | 1 | 7 |
| Entecavir | 4 | 2 | 1 | 7 |
| JB1a | 3 | 2 | 1 | 6 |
| Omeprazole | 3 | 2 | 1 | 6 |
| Esomeprazole magnesium | 3 | 2 | 1 | 6 |
| Gleevec | 3 | 2 | 0.5 | 5.5 |
| Aimspro | 3 | 2 | 0 | 5 |
| NCK-8 | 3 | 2 | 0 | 5 |
| D-LANA-14 | 3 | 2 | 0 | 5 |
| Tasigna | 3 | 1 | 0.5 | 4.5 |
| Celgosivir | 2 | 2 | 0 | 4 |
| Castanospermine | 2 | 2 | 0 | 4 |
| Name | Concentration | MRC-5 | VeroE6 | ||
|---|---|---|---|---|---|
| Ct Difference 1 | Cell Appearance 2 | Ct Difference | Cell Appearance | ||
| Ouabain | 20 nM | 3.48 ± 0.21 | x | −3.73 ± 4.88 | x |
| 17-DMAG | 5 μM | 3.72 ± 0.18 | x | −0.63 ± 1.39 | x |
| BGB324 | 3 μM | 3.05 ± 0.75 | ✓ | −1.83 ± 1.13 | ✓ |
| Zidovudine | 5 μM | −3.12 ± 0.27 | ✓ | −7.91 ± 2.67 | ✓ |
| Didanosine | 5 μM | −0.43 ± 3.87 | ✓ | −2.52 ± 1.27 | ✓ |
| Stavudine | 5 μM | −2.87 ± 0.22 | ✓ | −3.93 ± 0.25 | ✓ |
| Abacavir sulphate | 5 μM | −1.54 ± 3.26 | ✓ | −3.95 ± 2.69 | ✓ |
| Entecavir | 5 μM | −3.08 ± 0.20 | ✓ | −4.44 ± 1.11 | ✓ |
| JB1a | 2 μg/mL | −4.02 ± 0.13 | ✓ | −5.48 ± 0.50 | ✓ |
| Omeprazole | 100 μM | 1.35 ± 1.35 | x | 2.21 ± 1.08 | x |
| Esomeprazole magnesium | 75 μM | 1.05 ± 0.79 | x | 1.62 ± 0.36 | x |
| Gleevec | 20 μM | 3.60 ± 0.63 | x | 3.49 ± 0.54 | x |
| Aimspro | Neat | −2.03 ± 0.95 | ✓ | −4.60 ± 1.15 | ✓ |
| NCK-8 | 1 mg/mL | >10 | * | >10 | * |
| D-LANA-14 | 1 mg/mL | >10 | * | >10 | * |
| Tasigna | 20 μM | 3.59 ± 0.57 | x | −0.13 ± 0.33 | ✓ |
| Celgosivir | 200 μM | −2.52 ± 0.21 | ✓ | −2.41 ± 0.12 | ✓ |
| Castanospermine | 200 μM | −1.58 ± 3.23 | ✓ | −0.26 ± 4.11 | ✓ |
| Name | Concentration | Ct Difference 1 |
|---|---|---|
| Ouabain | 20 nM | 0.06 ± 0.10 |
| 6.7 nM | 0.09 ± 0.23 | |
| 2.2 nM | 0.33 ± 0.35 | |
| 17-DMAG | 63.3 nM | 0.30 ± 0.20 |
| 21.1 nM | 0.26 ± 0.57 | |
| 7.0 nM | 0.44 ± 0.06 | |
| BGB324 | 1 μM | 0.90 ± 0.15 |
| 0.3 μM | 0.67 ± 0.09 | |
| 0.1 μM | 0.34 ± 0.05 | |
| Omeprazole | 100 μM | 0.70 ± 0.10 |
| 33.3 μM | 0.77 ± 0.31 | |
| 11.1 μM | 0.86 ± 0.22 | |
| Esomeprazole | 25 μM | 0.78 ± 0.25 |
| 8.3 μM | 0.50 ± 1.06 | |
| 0.93 μM | 0.17 ± 0.16 | |
| Gleevec | 6.7 μM | 1.55 ± 0.20 |
| 2.2 μM | 1.03 ± 0.42 | |
| 0.74 μM | 0.64 ± 0.03 | |
| NCK-8 | 150 μg/mL | 1.54 ± 0.44 |
| 50 μg/mL | 1.33 ± 0.09 | |
| 16.7 μg/mL | 1.09 ± 0.17 | |
| D-LANA-14 | 60 μg/mL | 0.96 ± 0.19 |
| 20 μg/mL | 0.37 ± 0.11 | |
| 6.7 μg/mL | 0.40 ± 0.32 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dowall, S.D.; Bewley, K.; Watson, R.J.; Vasan, S.S.; Ghosh, C.; Konai, M.M.; Gausdal, G.; Lorens, J.B.; Long, J.; Barclay, W.; et al. Antiviral Screening of Multiple Compounds against Ebola Virus. Viruses 2016, 8, 277. https://doi.org/10.3390/v8110277
Dowall SD, Bewley K, Watson RJ, Vasan SS, Ghosh C, Konai MM, Gausdal G, Lorens JB, Long J, Barclay W, et al. Antiviral Screening of Multiple Compounds against Ebola Virus. Viruses. 2016; 8(11):277. https://doi.org/10.3390/v8110277
Chicago/Turabian StyleDowall, Stuart D., Kevin Bewley, Robert J. Watson, Seshadri S. Vasan, Chandradhish Ghosh, Mohini M. Konai, Gro Gausdal, James B. Lorens, Jason Long, Wendy Barclay, and et al. 2016. "Antiviral Screening of Multiple Compounds against Ebola Virus" Viruses 8, no. 11: 277. https://doi.org/10.3390/v8110277
APA StyleDowall, S. D., Bewley, K., Watson, R. J., Vasan, S. S., Ghosh, C., Konai, M. M., Gausdal, G., Lorens, J. B., Long, J., Barclay, W., Garcia-Dorival, I., Hiscox, J., Bosworth, A., Taylor, I., Easterbrook, L., Pitman, J., Summers, S., Chan-Pensley, J., Funnell, S., ... Carroll, M. W. (2016). Antiviral Screening of Multiple Compounds against Ebola Virus. Viruses, 8(11), 277. https://doi.org/10.3390/v8110277






